Introduction
Oral squamous cell carcinoma (OSCC) is a common
malignant head and neck tumor with a poor prognosis, accounting for
90% of oral cancers (1,2). Despite the progress in comprehensive
treatments, the overall 5-year survival rate of patients with OSCC
remains unsatisfactory owing to high recurrence and metastasis
rates (3). Therefore, effective
treatments for OSCC are urgently needed.
Natural compounds play an important role in cancer
therapies. Oxymatrine is an analog of matrine, which is extracted
from the roots of the Chinese herb Sophora flavescens, also named
‘Ku-shen’ (4). Oxymatrine is
commonly used to treat hepatitis B and C viral infections (5). Its effects on hepatic fibrosis and
ischemia-reperfusion injury, as well as its analgesic and
cardioprotective effects, are well-defined (6,7).
Previously, an increasing number of studies have demonstrated the
antitumor activities of oxymatrine, including inhibition of cancer
cell proliferation, induction of apoptosis and reversal of
multidrug resistance (8–12). Therefore, oxymatrine is used as a
novel antitumor agent for the treatment of different types of
cancer. However, the antitumor effect of oxymatrine suppressor in
OSCC has not been well elucidated.
Dysregulation of the CXC chemokine receptor 4
(CXCR4) is implicated in multiple malignancies. It plays a major
role in tumor progression, such as angiogenesis, metastasis and
survival of cancer cells (13,14).
A previous study reported that CXCR4 downregulation inhibited tumor
growth by inducing cell apoptosis and cycle arrest (15). CXCR4 knockdown attenuates tumor
metastasis by inhibiting epithelial-mesenchymal transition
(16). Additionally, CXCR4 is
remarkably increased in OSCC and is associated with poor prognosis
(17,18). Therefore, CXCR4 overexpression is
critical for OSCC occurrence and development.
N6-methyladenosine (m6A), the most
abundant modification in eukaryotic mRNA, is involved in RNA
splicing, stability, translation and nucleation, and plays a role
in tumor progression (19,20). Methyltransferase-like protein 3
(METTL3) is an enzyme that catalyzes m6A and plays a key role in
promoting tumor progression in myeloid leukemia (21), breast (22) and colon cancer (23). In addition, METTL3 promotes tumor
progression by regulating m6A methylation of
B-cell-specific moloney murine leukemia virus insertion site 1
(BMI1) in OSCC, indicating that METTL3 may be an oncogenic factor
in OSCC (24).
The present study aimed to investigate the effect of
oxymatrine on OSCC and its underlying mechanisms. It was identified
that oxymatrine inhibits CXCR4 expression by reducing its
m6A modification level by downregulating METTL3, thereby
inhibiting the proliferation and migration of OSCC.
Materials and methods
Cell culture and transfection
Human normal squamous epithelial hNOK cells (cat.
no. CRL-2692) and OSCC cell lines SCC-15 (cat. no. CRL-1623) and
CAL-27 (cat. no. CRL-2095) were provided by the American Type
Culture Collection (ATCC). Both cell lines were cultured using
complete DMEM medium containing 10% fetal bovine serum (FBS; both
from Gibco; Thermo Fisher Scientific, Inc.) and 1%
penicillin/streptomycin solution (Procell Life Science &
Technology Co.) at 37°C with 5% CO2. METTL3- and
CXCR4-expressing plasmids (pcDNA3.1-METTL3 and pcDNA3.1-CXCR4) and
negative control were obtained from Shanghai GeneChem Co., Ltd.
Plasmids (~4 µg) were used for cell transfections and the
transfections were performed using the Lipofectamine®
3000 kit (cat. no. L3000015; Thermo Fisher Scientific, Inc.)
according to the manufacturer's protocol. After transfection, cells
were cultured at 37°C with 5% CO2 and collected for
further experiments 1-2 days later.
RNA isolation and reverse
transcription-quantitative PCR (RT-qPCR)
TRIzol® reagent (cat. no. 15596018;
Thermo Fisher Scientific, Inc.) was used for the extraction of
total RNA from OSCC cells. After RNA isolation, NanoDrop1000
(NanoDrop Technologies; Thermo Fisher Scientific, Inc.) was used to
detect the quality and concentration of total RNA at the wavelength
of 260/280 nm. Subsequently, Primescript RT Reagent (cat. no.
RR047A; TaKaRa Bio, Inc.) was used to reverse-transcribe the total
RNA into cDNAs according to the manufacturer's protocol. RT-qPCR
reactions were conducted on a LightCycler 480 (Roche Diagnostics)
using SYBR qPCR Master Mix (cat. no. Q711-02; Vazyme Biotech Co.,
Ltd.). For qPCR, the following thermocycling conditions were
applied: Initial denaturation at 95°C for 5 min, subsequently
denaturation at 95°C for 10 sec for 40 cycles, 60°C for 20 sec of
annealing and elongation and final extension at 72°C for 20 sec.
GAPDH was used as an internal reference gene. The 2−ΔΔCq
method was used to calculate the relative gene expression (25). The following primers were used in
the present study: METTL3 forward, 5′-AACAGAGCAAGAAGGTCGGG-3′ and
reverse, 5′-TCGGTCTGCACTGGAATCAC-3′; CXCR4 forward,
5′-ACTACACCGAGGAAATGGGCT-3′ and reverse,
5′-CCCACAATGCCAGTTAAGAAGA-3′; and GAPDH forward,
5′-GGAGCGAGATCCCTCCAAAAT-3′ and reverse,
5′-GGCTGTTGTCATACTTCTCATGG-3′.
Cell counting kit 8 (CCK-8)
Cell proliferation was evaluated using CCK-8 assays.
OSCC cells (~5,000) were seeded in a 96-well plate and cultured for
24-48 h. Afterwards, CCK-8 solution (10 µl; cat. no. CK04; Dojindo
Laboratories, Inc.) was added to each well. After incubation for ~1
h at 37°C, the absorbance of each well was measured at 450 nm by a
microplate reader (Infinite M200 PRO; Tecan Group, Ltd.).
Cell migration assay
Transwell assays were performed to detect cell
migration using chambers (Sigma-Aldrich; Merck KGaA). Cells were
first resuspended in DMEM (serum-free) and then ~2×104
cells were seeded into the upper chamber. Then, 500 µl DMEM
(containing 10% FBS) was added to the lower chamber. After 24 h,
migratory cells on the lower surface were fixed with 4%
paraformaldehyde (20 min) at room temperature. Cells were stained
with 0.1% crystal violet (Beyotime Institute of Biotechnology) for
30 min at room temperature. After washing with phosphate-buffered
saline (PBS) thrice, images of the cells that migrated through the
membrane were captured using a light microscope (magnification,
×20; Nikon Corporation).
Flow cytometry
The Annexin V-FITC Apoptosis Detection kit (cat. no.
556547; BD Biosciences) was used for early and late apoptosis
detection. Briefly, 5×105 OSCC cells were treated with
oxymatrine for ~24 h, harvested by trypsinization and centrifuged
(300 × g at room temperature) for 5 min. Cells were then washed
with ice-cold PBS, resuspended in 100 µl Annexin V-FITC binding
buffer, and mixed with Annexin V-FITC (5 µl) and propidium iodide
(5 µl). After incubation at room temperature in the dark (15 min),
300 µl of Annexin V-FITC binding buffer was added. Flow cytometry
(BD FACSAria III Flow Cytometer; BD Biosciences) was performed and
the results were analyzed using CellQuest™ software
(version 3.3; Becton Dickinson and Company).
Western blot analysis
To isolate total cell proteins, OSCC cells were
lysed using RIPA buffer (Beyotime Institute of Biotechnology). A
BCA protein assay kit (cat. no. P0012S; Beyotime Institute of
Biotechnology) was used to assess the concentrations of the
isolated proteins. Equal amounts of protein (60 µg) were separated
using 10% SDS-PAGE. Subsequently, the separated proteins were
transferred onto PVDF membranes. Membranes were blocked with 5%
fat-free milk for ~1 h at room temperature and then incubated with
primary antibodies overnight at 4°C. TBST (with 0.1% Tween) was
used to wash the membranes thrice. The membranes were then
incubated with the corresponding secondary antibodies, HRP-labeled
goat anti-mouse (1:2,000; cat. no. A0216) and HRP-labeled goat
anti-rabbit (1:2,000; cat. no. A0208) diluted in TBST (cat. no.
ST671; Beyotime Institute of Biotechnology), at room temperature
for 1 h. Finally, the membranes were exposed to enhanced
chemiluminescence (ECL; Thermo Fisher Scientific, Inc.). Primary
antibodies against anti-CXCR4 (1:1,000; cat. no. ab181020), Bax
(1:1,000; cat. no. ab182734), anti-Bcl-2 (1:1,000; cat. no.
ab32124), anti-METTL3 (1:1,000; cat. no. ab195352), anti-alkB
homolog 5 (ALKBH5; 1:1,000; cat. no. ab195377), anti-WT1-associated
protein (WTAP; 1:1,000; cat. no. ab195380), anti-METTl14, (1:1,000;
cat. no. ab220030), anti-fat mass and obesity associated (FTO;
1:2,000; cat. no. ab126605) and β-actin (1:2,500; cat. no. ab8226)
were obtained from Abcam. Image-Pro Plus software (version 6.0;
Media Cybernetics, Inc.) was used to semi-quantify relative protein
expression levels, and β-actin was used as the loading control.
RNA-binding protein
immunoprecipitation (RIP) analysis
A Magna RIP kit (cat. no. 17-704; MilliporeSigma)
was used for the RIP assay in accordance with the manufacturer's
protocol. Briefly, total cell lysate was mixed with RIP buffer.
Then 50 µl Protein A + G magnetic beads were cultured with the
mixture at 4°C for ~12 h. The beads were conjugated with
anti-METTL3, anti-Ago2 (cat. no. ab186733) or anti-IgG (cat. no.
ab172730; both from Abcam), respectively. Finally, RT-qPCR was
performed to quantify the immunoprecipitated RNAs using the RIP
assay.
Methylated RNA immunoprecipitation
(MeRIP)
MeRIP was used to detect m6A levels in
CXCR4 mRNA. Total RNA was extracted from OSCC cells by using
DynabeadsTM mRNA Purification kit (cat. no. 61006;
Thermo Fisher Scientific, Inc.). The isolated RNAs were treated
with DNase R (Qiagen GmbH). Anti-m6A antibody (5 µg,
cat. no. ab208577; Abcam) was used to immunoprecipitate chemically
fragmented RNA. After being washed thrice with IP buffer, elution
buffer was used to elute RNA from the beads at 4°C for 1 h. RT-qPCR
was used to detect the RNA expression in input and
immunoprecipitation (IP) groups.
m6A enrichment assay
The enrichment of m6A in cells and
tissues was measured by using EpiQuik™m6A RNA
Methylation Quantification kit (cat. no. P-9005; EpiGentek)
according to the manufacturer's protocol. Briefly, total RNA was
isolated from cells and tissues firstly. RNA (~200 ng) was added to
interact with strip wells by using RNA high binding solution,
followed up with incubation at 37°C for 90 min. Afterwards, the
wells were washed with wash buffer for three times. Each well was
then added with 50 µl capture antibody and cultured at room
temperature for 1 h. Subsequently, 50 µl detection antibody was
added into each well for 30 min incubation at room temperature.
After being washed with wash buffer, each well was added with 50 µl
enhanced solution and incubated for 30 min at room temperature.
Then, 100 µl developer solution was added into each well for
another incubation for 10 min in the dark at room temperature and
the stop solution was subsequently added. Finally, the detected
signal was enhanced and then quantified calorimetrically by
measuring the absorbance at optical density (OD)=450 nm in a
microplate reader (Infinite M200 PRO, Tecan Group, Ltd.). The
amount of m6A is proportional to the OD intensity
measured.
RNA stability assay
SCC-15 and CAL-27 cells (~5×105) were
treated with actinomycin D (Act-D, 5 µg/ml; MedChemExpress) for 0,
2, 4, and 6 h. At the indicated times, total RNA was isolated and
quantified by qPCR. Linear regression analysis was used to estimate
the mRNA degradation rate.
Animal model
A total of 12 BALB/c female nude mice (age, 4-weeks
old; weight, ~20 g) were obtained from GemPharmatech. All mice were
raised in a standard barrier environment, under
specific-pathogen-free conditions at 22°C with a 12 h light/dark
cycle and free access to food and water. Mice were randomly divided
into two groups (control and oxymatrine-treated). CAL-27 cells
(~1×106; PBS solution was used for suspension) were
subcutaneously injected into the bilateral hind legs of the nude
mice. After a week, one group was injected with PBS as a control,
and the other was treated with oxymatrine (30–50 mg/kg) every 3
days. Tumor size was recorded every 3 days according to the
following formula: V=(width2 × length)/2. At total of 4
weeks later, the mice were euthanized by tail vein injection of
sodium pentobarbital (200 mg/kg). Tumor tissues were harvested from
mice and preserved in −80°C for further study. All animal
experiments were conducted in accordance with the Guidelines for
the Care and Use of Laboratory Animals. The present study was
approved (approval no. 20190612) by the Ethics Committee of the
Guangzhou Hospital of Integrated Traditional and Western Medicine
(Guangzhou, China).
Immunohistochemistry analysis
(IHC)
For IHC, the slides were heated at 60°C for 2 h,
followed by the removal of paraffin using xylene. After being
rinsed with ethanol, antigen recovery was conducted for 10 min and
the samples were immersed in 3% hydrogen peroxide and subsequently
blocked with 10% serum (cat. no. SL034; Beijing Solarbio Science
& Technology Co., Ltd.) at room temperature for 15 min. The
slides were incubated with primary antibodies anti-CXCR4 (1:1,000;
cat. no. ab181020) and anti-METTL3 (1:1,000; cat. no. ab195352;
both from Abcam) at 4°C overnight and then incubated with the
corresponding secondary antibody, HRP-labeled goat anti-rabbit
(1:2,000) for 1 h at room temperature. The slides were then stained
with DAB work solution (cat. no. P0202; Beyotime Institute of
Biotechnology) at room temperature, counterstained with hematoxylin
work solution (cat. no. C0107; Beyotime Institute of Biotechnology)
at room temperature and identified with 1% hydrochloric acid
alcohol. Finally, slides were fixed with neutral balsam (cat. no.
G8590; Solarbio Science & Technology Co., Ltd.) at room
temperature and visualized under a light microscope (magnification,
×20; Nikon Corporation).
Statistical analysis
GraphPad Prism version 7 (GraphPad Software, Inc.)
was used for statistical analysis. Data are from three independent
experiments and were presented as the mean ± SD. The differences
between two groups were analyzed using unpaired Student's t-test.
One-way ANOVA followed by Turkey's multiple comparison test was
applied to analyze differences among multiple groups. P<0.05 was
considered to indicate a statistically significant difference.
Results
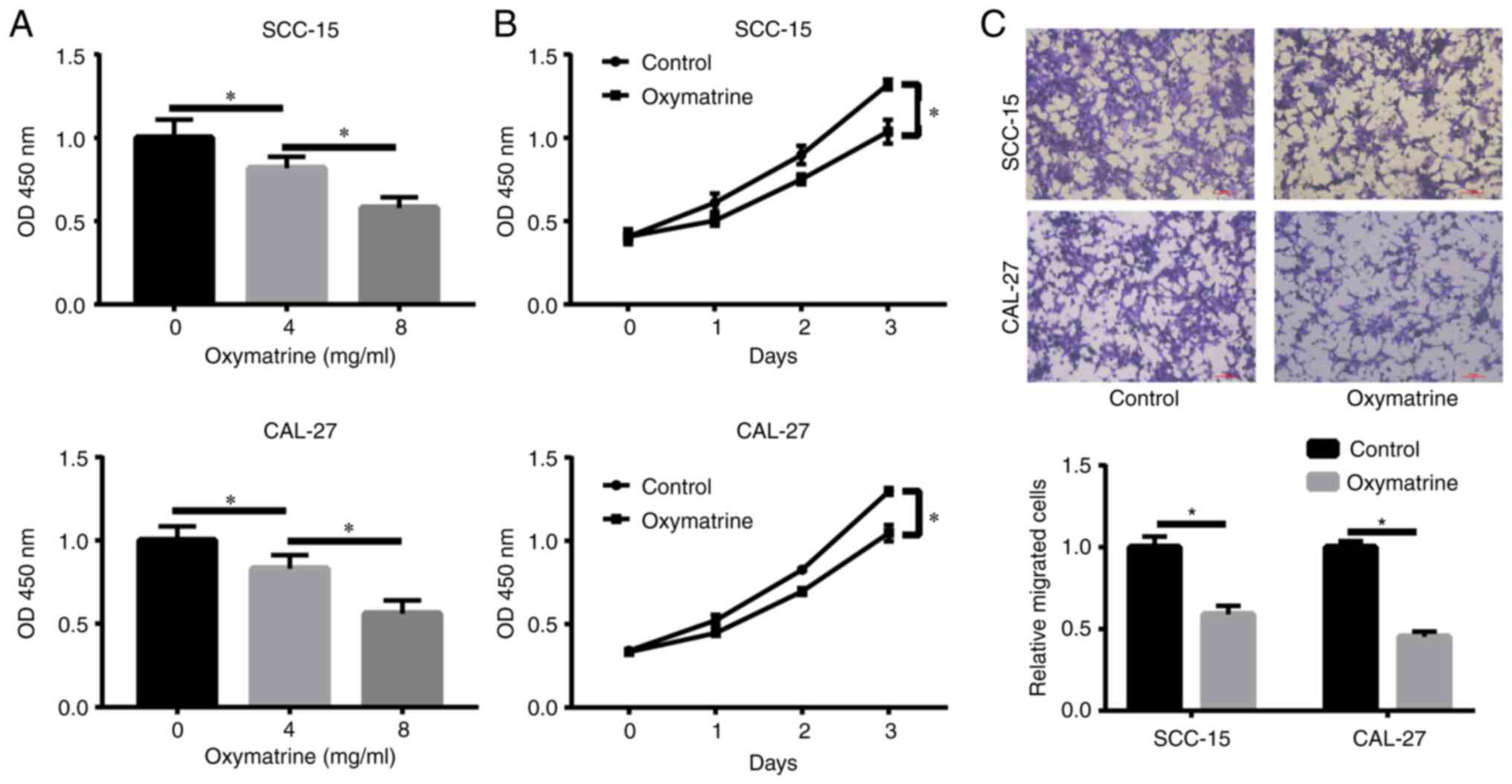
Oxymatrine inhibits proliferation and
migration of OSCC cells
A total of 2 OSCC cell lines, SCC-15 and CAL-27,
were cultured with different concentrations of oxymatrine (0, 4 and
8 mg/ml) for 24 h and cell proliferation was assessed using CCK-8
assays. The results revealed that oxymatrine significantly reduced
cell proliferation in a dose-dependent manner (Fig. 1A). A concentration of 8 mg/ml was
selected for further experiments owing to its strong effects. As
revealed in Fig. 1B, oxymatrine
inhibited proliferation of OSCC cells at the indicated time points.
To explore the effect of oxymatrine on tumor metastasis, a
Transwell assay was conducted, and the results showed that
oxymatrine attenuated SCC-15 and CAL-27 cell migration (Fig. 1C). Notably, the same concentration
(8 mg/ml) of oxymatrine did not promote apoptosis or inhibit the
proliferation (Fig. S1A and B) of
the human normal squamous epithelial (hNOK) cells, indicating that
oxymatrine exerts antitumor effects but has no side effects on
normal cells.
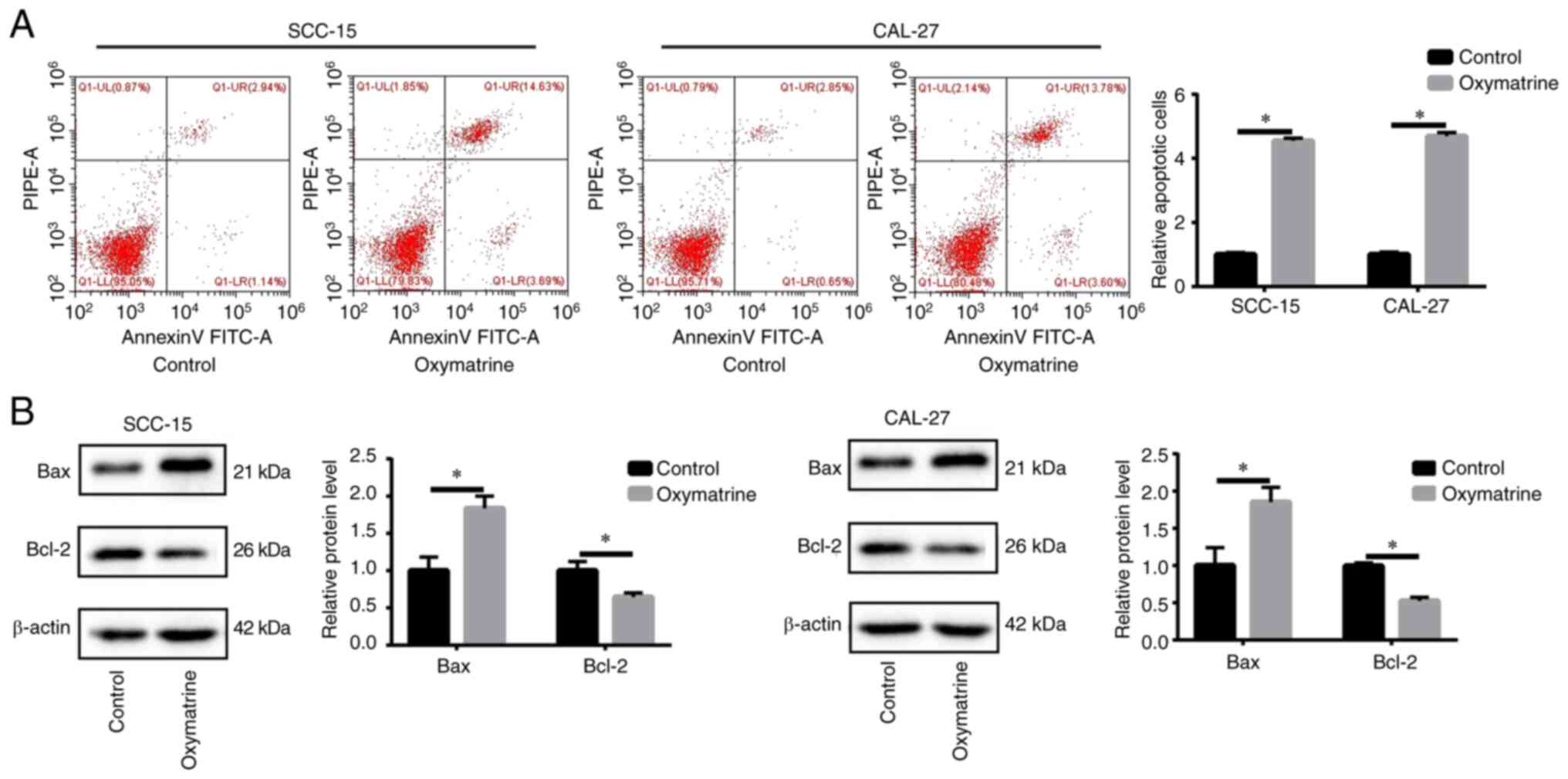
Oxymatrine induces OSCC cell
apoptosis
Apoptosis was assessed using flow cytometric
analysis. It was identified that, compared with the control group,
oxymatrine induced higher rates of apoptosis (Fig. 2A). Moreover, oxymatrine increased
the expression of the proapoptotic protein Bax but significantly
reduced the antiapoptotic protein Bcl-2 (Fig. 2B). These results indicated that
oxymatrine acts as a tumor suppressor in OSCC by inhibiting cell
proliferation and migration and promoting cell apoptosis.
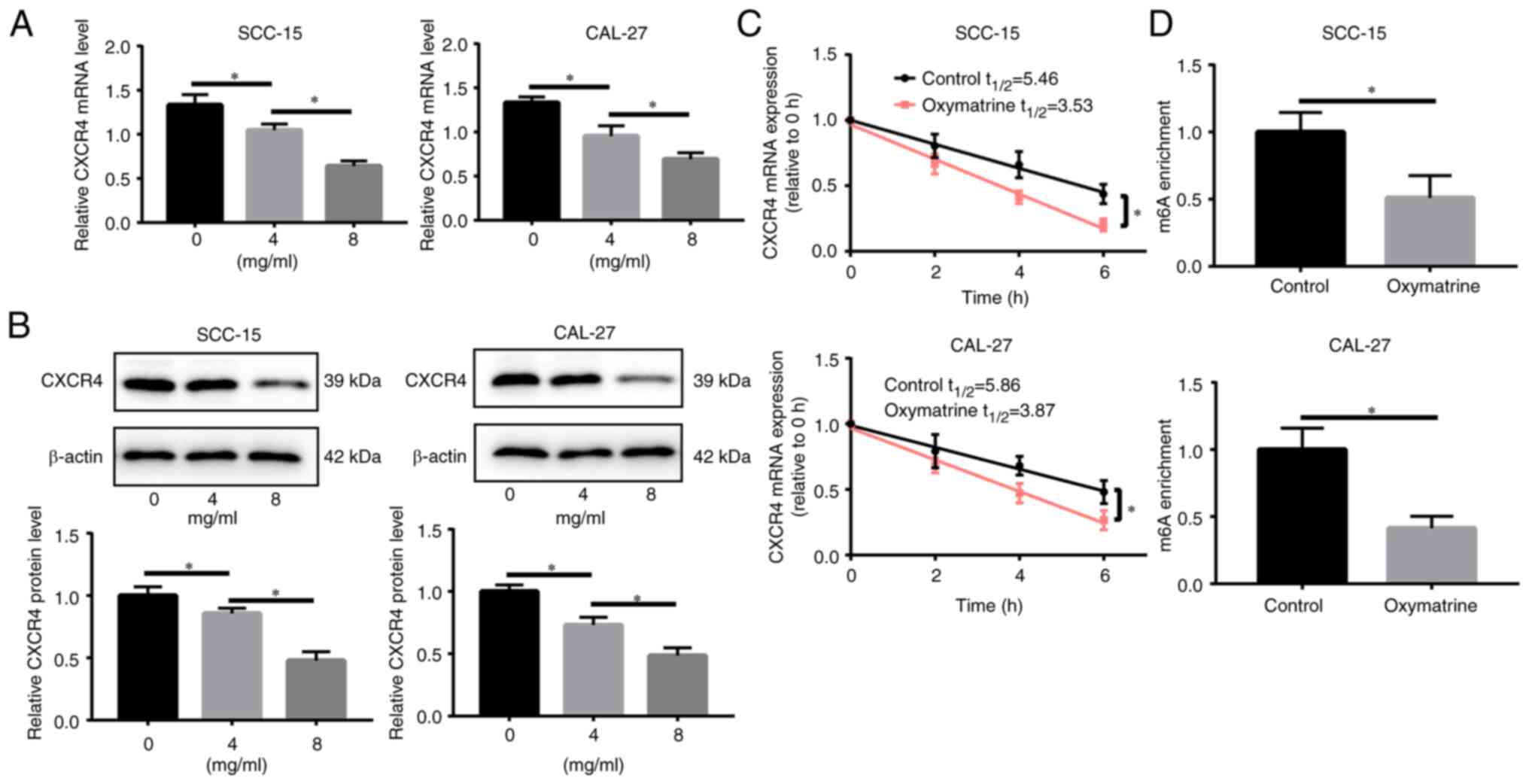
Oxymatrine inhibits CXCR4
expression
CXCR4 is markedly increased in OSCC and promotes the
occurrence and development of OSCC (17,18);
therefore, it was investigated whether oxymatrine treatment in OSCC
affects CXCR4 expression. Oxymatrine treatment reduced CXCR4
expression level in a concentration-dependent manner in SCC-15 and
CAL-27 cells (Fig. 3A and B).
Considering m6A is the most abundant modification in
eukaryotic mRNA, which is closely related to the RNA regulation, as
well as the stability of RNA, it was hypothesized that oxymatrine
may regulate CXCR4 expression and the mRNA stability by modulating
the m6A modification level of CXCR4 mRNA. Notably,
oxymatrine treatment (8 mg/ml) shortened the half-life of CXCR4
mRNA in both SCC-15 and CAL-27 cells (Fig. 3C). Furthermore, the MeRIP assay
revealed that oxymatrine reduced m6A modification of
CXCR4 mRNA in SCC-15 and CAL-27 cells (Fig. 3D). These data suggested that
oxymatrine reduces CXCR4 expression by downregulating the
m6A level of CXCR4 mRNA.
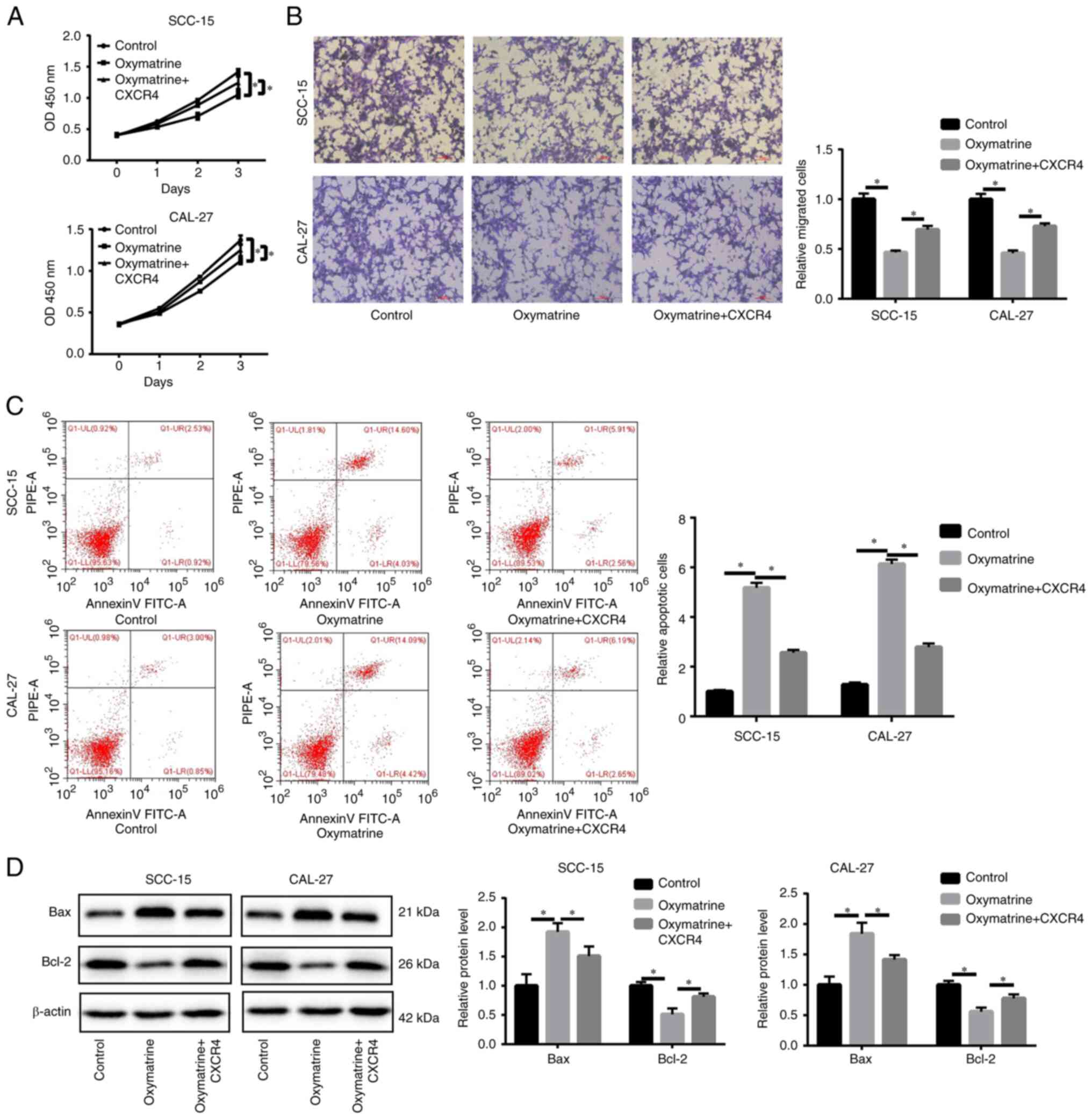
Oxymatrine exerts antitumor effects in
OSCC by inhibiting CXCR4 expression
To validate whether oxymatrine functions as a tumor
suppressor in OSCC by inhibiting CXCR4, rescue experiments were
conducted. Firstly, the transfection efficiency of CXCR4 plasmid
was determined at mRNA level by RT-qPCR, and it was found that the
transfection of CXCR4 plasmid significantly upregulated the CXCR4
expression in OSCC cells (Fig.
S1D). CCK-8 and Transwell assays revealed that CXCR4
overexpression partly reversed the inhibitory effects of oxymatrine
on cell proliferation and migration (Fig. 4A and B). CXCR4 upregulation
alleviated apoptosis (Fig. 4C). In
addition, in oxymatrine-treated SCC-15 and CAL-27 cells, CXCR4
upregulation reduced Bax protein expression level and increased
Bcl-2 protein expression level (Fig.
4D).
Oxymatrine downregulates CXCR4 by
decreasing the level of m6A modification via inhibiting
METTL3 expression
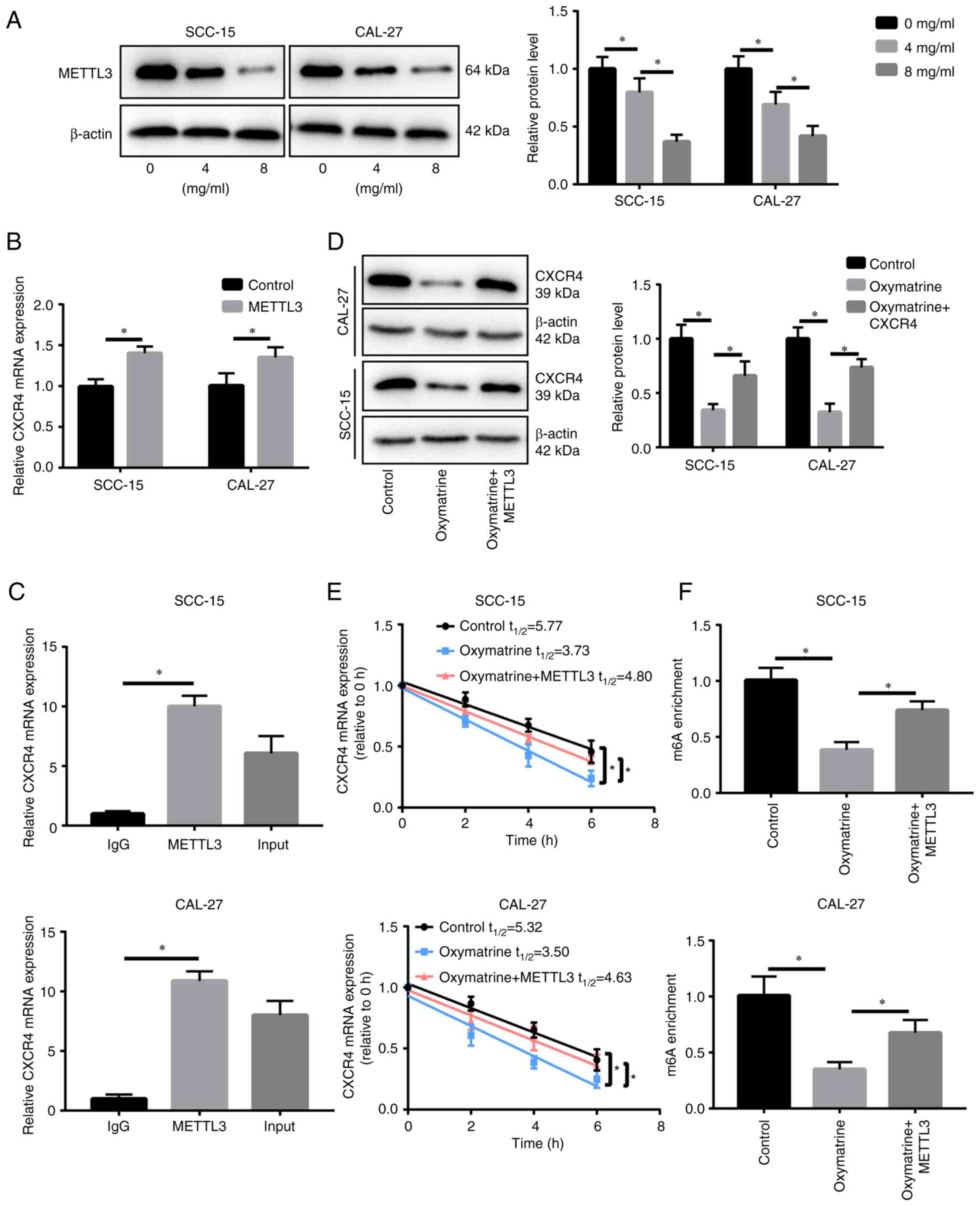
In order to clarify how oxymatrine regulates the
m6A modification level of CXCR4, it was first detected
whether the expression of m6A methylase (METTL3, METTL14
and WTAP) and transferase (ALKBH5 and FTO) are influenced by
oxymatrine. It was identified that the expression of METTL14, WTAP,
ALKBH5 and FTO was not significantly changed when treated with
oxymatrine (Fig. S1C), whereas
METTL3 was downregulated significantly in a dose-dependent manner
(Fig. 5A) indicating that
m6A methylase METTL3 may participate in oxymatrine's
regulation of CXCR4. Firstly, the transfection efficiency of METTL3
plasmid was determined at mRNA level by RT-qPCR, and it was
observed that the transfection of METTL3 plasmid significantly
upregulated the METTL3 expression in OSCC cells (Fig. S1D). Furthermore, METTL3
overexpression promoted CXCR4 expression (Fig. 5B). The RIP assay demonstrated that
METTL3 binds to CXCR4 mRNA (Fig.
5C). Although oxymatrine treatment inhibited CXCR4 expression,
METTL3 upregulation reversed this effect (Fig. 5D). In addition, METTL3 prolonged
the half-life of CXCR4 mRNA, which was reduced by oxymatrine
treatment in SCC-15 and CAL-27 cells (Fig. 5E). The MeRIP assay showed that
METTL3 overexpression reversed the reduction in the level of CXCR4
mRNA m6A methylation, which was induced by oxymatrine in
SCC-15 and CAL-27 cells (Fig.
5F).
To elucidate whether other m6A enzymes
participate in regulating the level of CXCR4 mRNA m6A
methylation, the expression of other m6A enzymes was
detected, including METTL14, WTAP, ALKBH5 and FTO. It was found
that the protein expression levels of these enzymes were not
significantly changed when treated with oxymatrine (Fig. S1C), indicating that they may not
participate in regulating the level of CXCR4 mRNA m6A
methylation. Therefore, oxymatrine inhibits CXCR4 expression by
reducing the m6A methylation level via downregulating
METTL3.
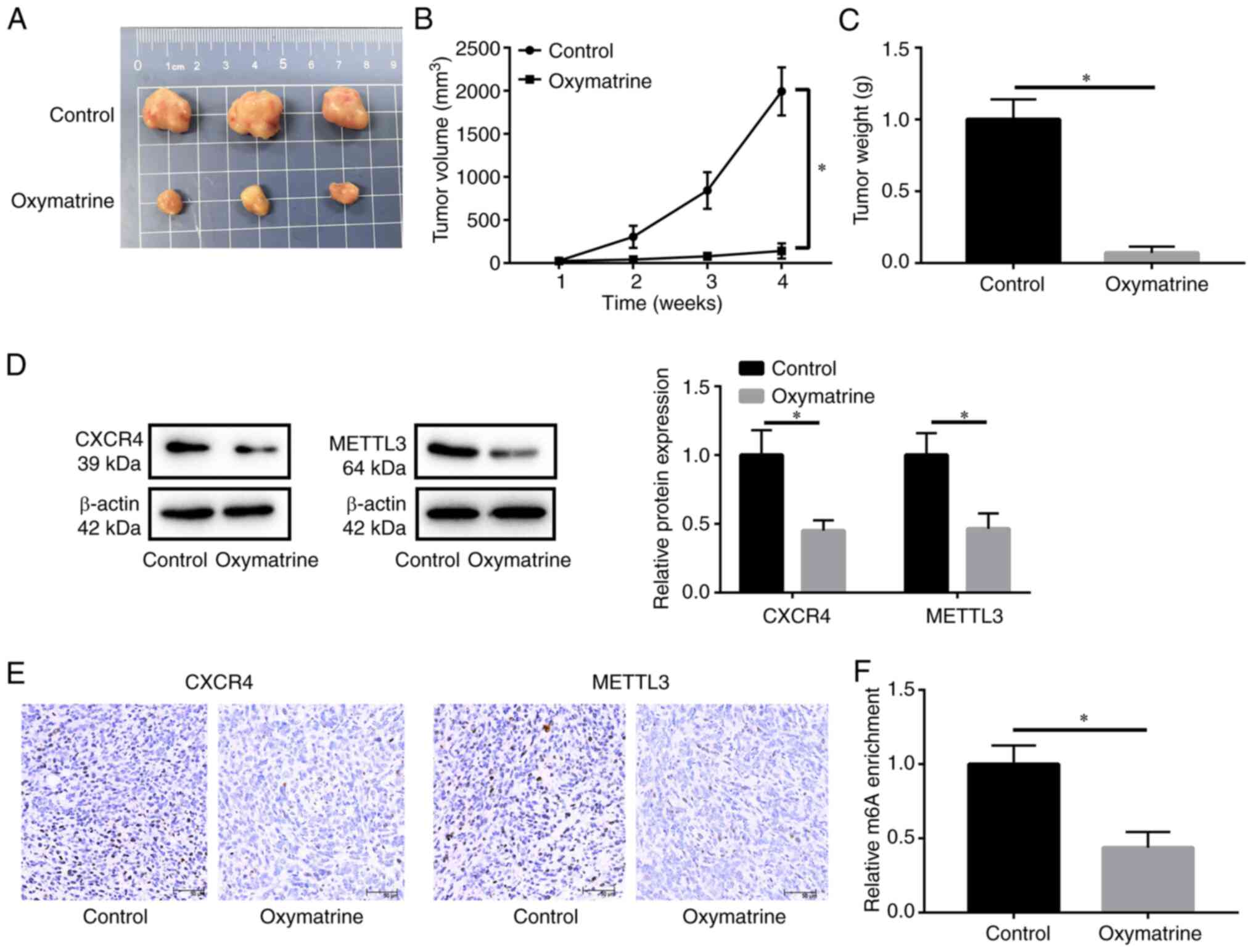
Oxymatrine inhibits tumor growth in
vivo
To further evaluate the efficacy of oxymatrine in
the treatment of OSCC, xenograft tumors were established in nude
mice. Compared with the control group, oxymatrine treatment
inhibited tumor growth and reduced tumor volume and weight
(Fig. 6A-C). Western blotting and
IHC assays revealed that CXCR4 and METTL3 were downregulated in the
oxymatrine-treated group compared with the control group (Fig. 6D and E). Additionally,
m6A levels in the oxymatrine-treated group were much
lower than those in the control group (Fig. 6F).
Discussion
Oxymatrine has been revealed to inhibit the growth
of several types of cancer and has gained increasing attention in
recent decades. Oxymatrine induces apoptosis in pancreatic cancer
cells by influencing the ratio of Bax/Bcl2 and the release of
mitochondrial cytochrome c (26).
In bladder cancer, oxymatrine suppresses tumor progression by
inducing apoptosis and cell cycle arrest (27). However, little is known about the
role of oxymatrine in OSCC. In the present study, it was found that
oxymatrine attenuated the progression of OSCC by inhibiting tumor
cell proliferation and migration and promoting cell apoptosis. The
antitumor effect of oxymatrine in OSCC in vivo was further
confirmed.
CXCR4, a highly conserved seven-span transmembrane
G-protein-coupled receptor that binds to the ligand CXCL12
(28), plays an important role in
the survival, proliferation, angiogenesis and metastasis of various
tumors (14). Dysregulation of
CXCR4 has been observed in OSCC. Additionally, OSCC progression is
regulated by the miR-139-5p/CXCR4 axis (29). Zerumbone, a bioactive monocyclic
sesquiterpene isolated from the rhizomes of tropical ginger,
reduces OSCC cell motility and proliferation by targeting CXCR4
(30). Therefore, it was
hypothesized that oxymatrine may function as a tumor suppressor by
downregulating CXCR4. The present findings showed that oxymatrine
treatment reduced the mRNA and protein expression levels of CXCR4
in a dose-dependent manner in OSCC cells. Furthermore, the rescue
experiments indicated that ectopic CXCR4 expression partly restored
the suppressive effect of oxymatrine on OSCC cell viability and
migration, but alleviated oxymatrine-induced cell apoptosis.
Therefore, oxymatrine inhibits OSCC tumor progression by inhibiting
CXCR4 expression.
m6A is the most common mRNA modification
(31,32). Accumulating evidence has revealed
that m6A regulators, such as METTL3, are dysregulated in
numerous tumors, including OSCC and METTL3 plays a role in tumor
cell proliferation, migration and apoptosis by stabilizing its
downstream genes (33–37). In the present study, it was
demonstrated that oxymatrine reduced the stability of CXCR4 mRNA,
which was confirmed by the Act-D assay, suggesting that oxymatrine
may influence CXCR4 m6A methylation level in OSCC cells.
MeRIP results revealed that oxymatrine treatment reduced the
m6A methylation level of CXCR4 mRNA. Moreover, it was
found that oxymatrine also decreased METTL3 expression, indicating
that oxymatrine may decrease the m6A methylation level
of CXCR4 mRNA by regulating METTL3 expression. The RIP assay showed
that METTL3 binds to CXCR4 mRNA and the MeRIP assay identified that
METTL3 knockdown reduces CXCR4 m6A mRNA methylation
level. Furthermore, oxymatrine treatment in OSCC cells
overexpressing METTL3 increased the stability and m6A
methylation level of CXCR4 mRNA. Collectively, the results of the
present study confirmed that oxymatrine induced a reduction in
m6A methylation levels in CXCR4 mRNA by inhibiting
METTL3 expression.
In summary, the present study demonstrated for the
first time, to the best of our knowledge, that oxymatrine
attenuates OSCC progression by inhibiting cell proliferation and
migration and inducing cell apoptosis. Mechanistically, it was
revealed that oxymatrine directly destabilizes CXCR4 mRNA via
inhibiting METTL3-mediated m6A methylation. Therefore,
the present findings revealed the therapeutic value of oxymatrine
in treating OSCC. However, further studies are needed to determine
the validity, safety and pharmacokinetics of oxymatrine.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The datasets in the present study are available from
the corresponding author on reasonable request.
Authors' contributions
JS and ZL designed the present study. RL, YL and LX
performed the experiments. RL and YL analyzed the data. JS, RL and
YL prepared the draft of the manuscript. All authors read and
approved the final version of the manuscript. ZL and RL confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
The protocols of animal studies were approved
(approval no. 20190612) by The Animal Care and Use Committee of the
Guangzhou hospital of integrated traditional and West medicine
(Guangzhou, China).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
OSCC
|
oral squamous cell carcinoma
|
|
CXCR4
|
CXC chemokine receptor 4
|
|
METTL3
|
methyltransferase-like protein 3
|
|
METTL14
|
methyltransferase-like protein 14
|
|
m6A
|
N6-methyladenosine
|
|
CCK-8
|
Cell Counting Kit-8
|
|
RT-qPCR
|
reverse transcription-quantitative
PCR
|
|
MeRIP
|
methylated RNA immunoprecipitation
|
|
BMI1
|
B-cell-specific moloney murine
leukemia virus insertion site 1
|
|
Bax
|
Bcl-2 associated X
|
|
Bcl-2
|
B-cell lymphoma-2
|
|
ALKBH5
|
alkB homolog 5
|
|
FTO
|
fat mass and obesity associated
|
|
WTAP
|
WT1-associated protein
|
References
|
1
|
Sung H, Ferlay J, Siegel RL, Laversanne M,
Soerjomataram I, Jemal A and Bray F: Global cancer statistics 2020:
GLOBOCAN estimates of incidence and mortality worldwide for 36
cancers in 185 countries. CA Cancer J Clin. 71:209–249. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Sun S, Wang J, Liu J, Yin F, Xin C, Zeng
X, Li J and Chen Q: MiR-302b suppresses tumor metastasis by
targeting frizzled 6 in OSCC. J Dent Res. 100:739–745. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hartner L: Chemotherapy for oral cancer.
Dent Clin North Am. 62:87–97. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rabea EI, Nasr HM and Badawy ME: Toxic
effect and biochemical study of chlorfluazuron, oxymatrine, and
spinosad on honey bees (Apis mellifera). Arch Environ Contam
Toxicol. 58:722–732. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Wu XN and Wang GJ: Experimental studies of
oxymatrine and its mechanisms of action in hepatitis B and C viral
infections. Chin J Dig Dis. 5:12–16. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Shen XC, Yang YP, Xiao TT, Peng J and Liu
XD: Protective effect of oxymatrine on myocardial fibrosis induced
by acute myocardial infarction in rats involved in TGF-β1-Smads
signal pathway. J Asian Nat Prod Res. 13:215–224. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Wang Y, Yuan J, Yuan X, Wang W, Pei X,
Zhao Q, Cao H, Xu M and Liu Z: Observation of antinociceptive
effects of oxymatrine and its effect on delayed rectifier K+
currents (Ik) in PC12 cells. Neurochem Res. 37:2143–2149. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
He X, Fang J, Huang L, Wang J and Huang X:
Sophora flavescens Ait.: Traditional usage, phytochemistry and
pharmacology of an important traditional Chinese medicine. J
Ethnopharmacol. 172:10–29. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Guo B, Zhang T, Su J, Wang K and Li X:
Oxymatrine targets EGFR(p-Tyr845) and inhibits EGFR-related
signaling pathways to suppress the proliferation and invasion of
gastric cancer cells. Cancer Chemother Pharmacol. 75:353–363. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang X, Liu C, Wang J, Fan Y, Wang Z and
Wang Y: Oxymatrine inhibits the migration of human colorectal
carcinoma RKO cells via inhibition of PAI-1 and the TGF-β1/Smad
signaling pathway. Oncol Rep. 37:747–753. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wu C, Huang W, Guo Y, Xia P, Sun X, Pan X
and Hu W: Oxymatrine inhibits the proliferation of prostate cancer
cells in vitro and in vivo. Mol Med Rep. 11:4129–4134. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Li M, Su BS, Chang LH, Gao Q, Chen KL, An
P, Huang C, Yang J and Li ZF: Oxymatrine induces apoptosis in human
cervical cancer cells through guanine nucleotide depletion.
Anticancer Drugs. 25:161–173. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Barbieri F, Bajetto A, Stumm R, Pattarozzi
A, Porcile C, Zona G, Dorcaratto A, Ravetti JL, Minuto F, Spaziante
R, et al: Overexpression of stromal cell-derived factor 1 and its
receptor CXCR4 induces autocrine/paracrine cell proliferation in
human pituitary adenomas. Clin Cancer Res. 14:5022–5032. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhou Y, Cao HB, Li WJ and Zhao L: The
CXCL12 (SDF-1)/CXCR4 chemokine axis: Oncogenic properties,
molecular targeting, and synthetic and natural product CXCR4
inhibitors for cancer therapy. Chin J Nat Med. 16:801–810.
2018.PubMed/NCBI
|
|
15
|
Yu T, Wu Y, Huang Y, Yan C, Liu Y, Wang Z,
Wang X, Wen Y, Wang C and Li L: RNAi targeting CXCR4 inhibits tumor
growth through inducing cell cycle arrest and apoptosis. Mol Ther.
20:398–407. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Duan Y, Zhang S, Wang L, Zhou X, He Q, Liu
S, Yue K and Wang X: Targeted silencing of CXCR4 inhibits
epithelial-mesenchymal transition in oral squamous cell carcinoma.
Oncol Lett. 12:2055–2061. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang X, Li F and Zhou X: miR-204-5p
regulates cell proliferation and metastasis through inhibiting
CXCR4 expression in OSCC. Biomed Pharmacother. 82:202–207. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xia J, Chen N, Hong Y, Chen X, Tao X,
Cheng B and Huang Y: Expressions of CXCL12/CXCR4 in oral
premalignant and malignant lesions. Mediators Inflamm.
2012:5163952012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Barbieri I and Kouzarides T: Role of RNA
modifications in cancer. Nat Rev Cancer. 20:303–322. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Huang H, Weng H and Chen J: m6A
modification in coding and non-coding RNAs: Roles and therapeutic
implications in cancer. Cancer Cell. 37:270–288. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Barbieri I, Tzelepis K, Pandolfini L, Shi
J, Millan-Zambrano G, Robson SC, Aspris D, Migliori V, Bannister
AJ, Han N, et al: Promoter-bound METTL3 maintains myeloid leukaemia
by m6A-dependent translation control. Nature.
552:126–131. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Cai X, Wang X, Cao C, Gao Y, Zhang S, Yang
Z, Liu Y, Zhang X, Zhang W and Ye L: HBXIP-elevated
methyltransferase METTL3 promotes the progression of breast cancer
via inhibiting tumor suppressor let-7g. Cancer Lett. 415:11–19.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Xu J, Chen Q, Tian K, Liang R, Chen T,
Gong A, Mathy NW, Yu T and Chen X: m6A methyltransferase METTL3
maintains colon cancer tumorigenicity by suppressing SOCS2 to
promote cell proliferation. Oncol Rep. 44:973–986. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Liu L, Wu Y, Li Q, Liang J, He Q, Zhao L,
Chen J, Cheng M, Huang Z, Ren H, et al: METTL3 promotes
tumorigenesis and metastasis through BMI1 m6A
methylation in oral squamous cell carcinoma. Mol Ther.
28:2177–2190. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chen D, Wang H, Chen J, Li Z, Li S, Hu Z,
Huang S, Zhao Y and He X: MicroRNA-129-5p regulates glycolysis and
cell proliferation by targeting the glucose transporter SLC2A3 in
gastric cancer cells. Front Pharmacol. 9:5022018. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ling Q, Xu X, Wei X, Wang W, Zhou B, Wang
B and Zheng S: Oxymatrine induces human pancreatic cancer PANC-1
cells apoptosis via regulating expression of Bcl-2 and IAP
families, and releasing of cytochrome c. J Exp Clin Cancer Res.
30:662011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li S, Zhang Y, Liu Q, Zhao Q, Xu L, Huang
S, Huang S and Wei X: Oxymatrine inhibits proliferation of human
bladder cancer T24 cells by inducing apoptosis and cell cycle
arrest. Oncol Lett. 13:4453–4458. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Caruz A, Samsom M, Alonso JM, Alcami J,
Baleux F, Virelizier JL, Parmentier M and Arenzana-Seisdedos F:
Genomic organization and promoter characterization of human CXCR4
gene. FEBS Lett. 426:271–278. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Jiang Q, Cao Y, Qiu Y, Li C, Liu L and Xu
G: Progression of squamous cell carcinoma is regulated by
miR-139-5p/CXCR4. Front Biosci (Landmark Ed). 25:1732–1745. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Zainal NS, Gan CP, Lau BF, Yee PS, Tiong
KH, Abdul Rahman ZA, Patel V and Cheong SC: Zerumbone targets the
CXCR4-RhoA and PI3K-mTOR signaling axis to reduce motility and
proliferation of oral cancer cells. Phytomedicine. 39:33–41. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Nombela P, Miguel-López B and Blanco S:
The role of m6A, m5C and Ψ RNA modifications
in cancer: Novel therapeutic opportunities. Mol Cancer. 20:182021.
View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Pan Y, Ma P, Liu Y, Li W and Shu Y:
Multiple functions of m6A RNA methylation in cancer. J
Hematol Oncol. 11:482018. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Li F, Yi Y, Miao Y, Long W, Long T, Chen
S, Cheng W, Zou C, Zheng Y, Wu X, et al:
N6-methyladenosine modulates nonsense-mediated mRNA
decay in human glioblastoma. Cancer Res. 79:5785–5798. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Yao QJ, Sang L, Lin M, Yin X, Dong W, Gong
Y and Zhou BO: Mettl3-Mettl14 methyltransferase complex regulates
the quiescence of adult hematopoietic stem cells. Cell Res.
28:952–954. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zeng C, Huang W, Li Y and Weng H: Roles of
METTL3 in cancer: Mechanisms and therapeutic targeting. J Hematol
Oncol. 13:1172020. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhao W, Cui Y, Liu L, Ma X, Qi X, Wang Y,
Liu Z, Ma S, Liu J and Wu J: METTL3 facilitates oral squamous cell
carcinoma tumorigenesis by enhancing c-Myc stability via
YTHDF1-mediated m6A modification. Mol Ther Nucleic
Acids. 20:1–12. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Ban Y, Tan P, Cai J, Li J, Hu M, Zhou Y,
Mei Y, Tan Y, Li X, Zeng Z, et al: LNCAROD is stabilized by m6A
methylation and promotes cancer progression via forming a ternary
complex with HSPA1A and YBX1 in head and neck squamous cell
carcinoma. Mol Oncol. 14:1282–1296. 2020. View Article : Google Scholar : PubMed/NCBI
|