Medulloblastoma (MB) is the most common malignant
pediatric cancer, with ~1,800 new cases diagnosed in the United
States every year (1–3). MB is classified into four distinct
subgroups: Sonic hedgehog (SHH), wingless (WNT), Group 3 and Group
4 (4), with each subgroup
displaying distinct clinical presentation and genomic features
(5,6). Histological variants of MB are broadly
classified as classic (CMB), desmoplastic/nodular (DN), MB with
extensive nodularity (MBEN), and large cell/anaplastic (LC/A)
(7,8). Amplification of c-MYC is more
frequently observed in tumors with LC/A histology (9,10) and
in high-risk Group 3 and SHH tumors with TP53 abnormalities
(11). Notably, MYC-amplified Group
3 MBs exhibit a higher frequency of metastasis and have the worst
prognosis (12,13). Unfortunately, MYC inhibition has
proven unsuccessful in improving Group 3 MB patient outcomes
(14). As such, it is imperative to
expound on the mechanisms contributing to MB pathogenesis to
identify viable targets for developing novel therapeutics.
Initial studies on B7 homolog 3 (B7-H3) in cancer
primarily focused on its role in promoting immune evasion (15–17).
However, B7-H3 also exerts non-immunological functions to promote
tumorigenesis and metastasis (18–20).
For example, in hepatocellular carcinoma and melanoma, B7-H3
stimulates cell invasion and migration (21,22),
while in glioblastoma, B7-H3 promotes the epithelial-to-mesenchymal
transition (23). It was recently
demonstrated by the authors that B7-H3 is upregulated in Group 3
MBs and is associated with poor survival in MB patients (24). Further, it was showed that B7-H3
promotes angiogenesis and likely facilitates MB pathogenesis via
the exosome biogenesis (24,25).
As a result of its diverse functions, there has been increased
interest in therapeutically targeting B7-H3, with B7-H3 inhibitors
showing success in clinical trials (NCT01391143 and NCT02982941).
For instance, enoblituzumab (MGA271) is a humanized anti-B7H3
monoclonal antibody with an engineered Fc domain that is optimized
to augment antitumor effector-mediated functions (26). It accomplishes so by enhancing the
binding of CD16A (FcγRIIa) to trigger antibody-dependent
cell-mediated cytotoxicity (26)
and prevent the binding of CD32B (FcγRIIb), an inhibitory receptor
expressed on immune cells which dampens B-cell activation and
signaling (27).
In addition to B7-H3, MYC-amplified Group 3 MBs
overexpress the enhancer of zeste homolog 2 (EZH2) (28). EZH2 is the enzymatic catalytic
subunit of the polycomb repressive complex 2 (PRC2) (29). As a histone modifier, EZH2 is best
known for repressing gene transcription through the trimethylation
of H3K27 (H3K27me3) (30). Similar
to MYC, EZH2 can also indirectly regulate gene expression through
microRNAs (miRNAs or miRs) (31).
Of particular interest is the miR-29 family, which includes
miR-29a, miR-29b1/2 and miR-29c (32). For example, EZH2 directly inhibits
the transcription of miR-29b to upregulate the LOXL4 expression
(33). Notably, EZH2 has been shown
to bind to the promoter region of miR-29a/b1 and miR29b2/c to
facilitate MYC-mediated silencing of miR-29a-c expression (34). Notably, miR-29a directly binds to
the 3′-untranslated region (UTR) of B7-H3 transcripts to induce
translational repression (35). In
a previous study by the authors, it was found that overexpression
of miR-29a/b significantly reduced the protein levels of B7-H3.
Further, it was showed that B7-H3 expression was recused upon MYC
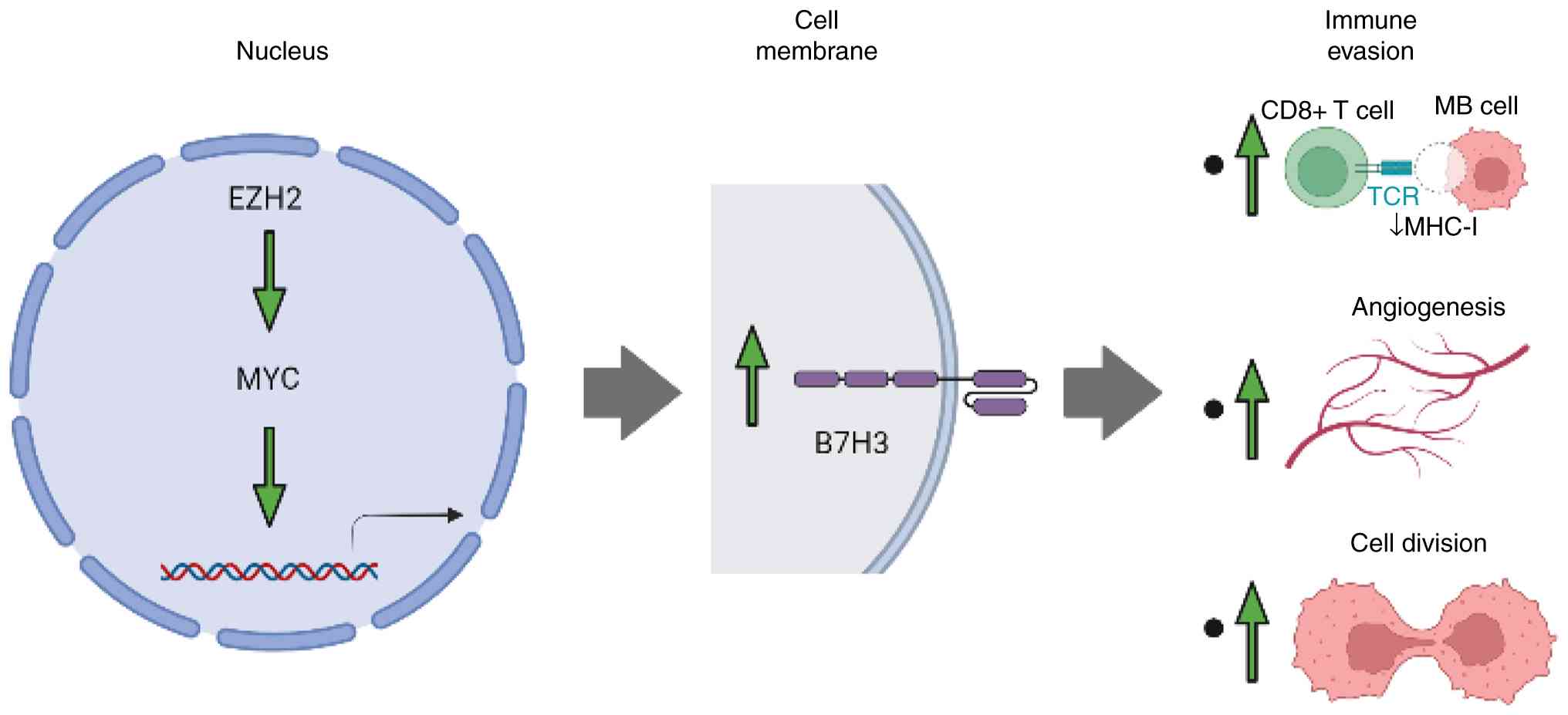
inhibition (24). Collectively,
these observations suggested that MYC and EZH2 may regulate B7-H3
expression directly or indirectly through miR-29 inhibition.
While research has supported the therapeutic
potential of B7-H3 inhibitors, such as enoblituzumab, suitable
therapies against B7-H3 remain in their infancy. As such,
investigating upstream regulators of B7-H3 expression may provide
more immediate effective therapies. One such regulator is miR-29.
However, the inability to successfully deliver miRNA-based
therapies to tumors and their inherent broad functionalities impede
the use of miRNA-based therapies in the clinical setting (35). Therefore, epigenetic therapies
targeting EZH2 may to lead to more effective methods of suppressing
B7-H3. For instance, studies have shown that Tazemetostat, a
small-molecule inhibitor of EZH2, demonstrates antitumor activity
in patients with advanced solid tumors (36), and has been recently approved by the
FDA for the treatment of relapsed/refractory follicular lymphoma
(37) (NCT01897571).
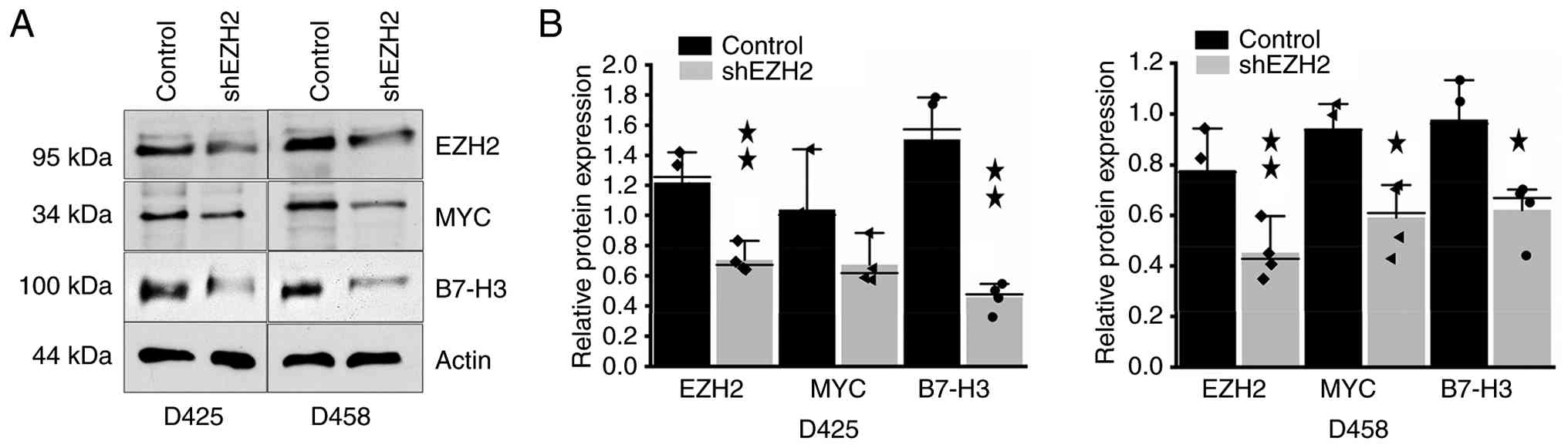
In the present study, it was reported that high
expression of EZH2 is associated with lower overall survival in
Group 3 MB patients. EZH2 silencing downregulated B7-H3 and MYC
transcript and protein expression, while pharmacological inhibition
of EZH2 significantly reduced clonogenic survival in MYC-amplified
Group 3 MB cells. It was further identified that both EZH2 and
B7-H3 inhibition decreased the viability of C17.2 cells stably
overexpressing EZH2. Importantly, it was demonstrated that EZH2
targeting markedly increased miR-29a expression in Group 3 MB
cells, suggesting that EZH2 inhibition facilitates miR-29-mediated
suppression of B7-H3.
The Kaplan-Meier survival curve of all Group 3 MB
patients was plotted using the Cavalli dataset within the R2
program (http://r2.amc.nl) (24). Data mining of classic, large cell,
and nodular desmoplastic MB histology types was carried out using
the mixed MB xenograft dataset (Zhao-92-custom-ilmnhwg6v2)
available on the R2 software.
For western blots, cells were lysed in 1X RIPA
buffer (cat. no. J62725-AP; Thermo Fisher Scientific, Inc.)
containing protease inhibitor cocktail (cat. no. 11697498001; Roche
Diagnostics). Cell lysates were sonicated, 4×30 sec with 30 sec
intervals between each sonication, and centrifuged at 20,817 × g
for 15 min at 4°C. Supernatants were collected for protein
estimation as determined by BCA (cat. no. A53225; Thermo Fisher
Scientific, Inc.). Equal concentrations of cell lysates (40 µg per
load) were resolved by SDS-PAGE, and western blot analysis was
performed using nitrocellulose membrane and specific antibodies.
EZH2 and B7-H3 were run on 10% SDS gels, 12% SDS gels were used for
MYC and 15% gels were used to visualize H3K27me3. For H3K27me3,
separate gels were run for actin, using 10% gel; the lysates and
concentrations were unchanged. Blocking for all blots was performed
using 10% non-fat milk for 1 h at room temperature. After which,
the blots were treated with primary antibody at 4°C for overnight
followed by secondary antibody for 1 h at room temperature. Mouse
polyclonal antibodies against B7-H3 (1:1,000; cat. no. sc-376769;
Santa Cruz Biotechnology, Inc.) (44) and c-MYC (1:1,000; cat. no. sc-40;
Santa Cruz Biotechnology, Inc.) (45) and rabbit polyclonal antibodies
against EZH2 (1:1,000; cat. no. 4905; Cell Signaling Technology,
Inc.) (46) and H3K27me3 (1:1,000;
cat. no. sc-56616; Santa Cruz Biotechnology, Inc.) (47), were used as primary antibodies. With
the exception of H3K27me3, all blots presented in the manuscript
were stripped and incubated with anti-β-actin (1:10,000; cat. no.
3700; Cell Signaling Technology, Inc.) (48) as the loading control.
Anti-mouse/rabbit IgG HRP conjugated antibody (1:10,000; cat. no.
400111; Novus Biologicals, LLC) was used as a secondary antibody.
Immunodetection was accomplished using ECL reagents (cat. no.
1705060S; BioRad Laboratories, Inc). Densitometric analysis was
carried out using ImageJ software (version 1.53t 24; National
Institutes of Health).
Cell viability was determined by MTT assays (cat.
no. M5655-100MG; MilliporeSigma). D425 and D458 cell lines were
seeded in 96-well plates at a concentration of 5×103
cells/well. The C17.2 and C17.2-EZH2-OE cell lines were seeded in
96-well plates at a concentration of 3×103 cells/well.
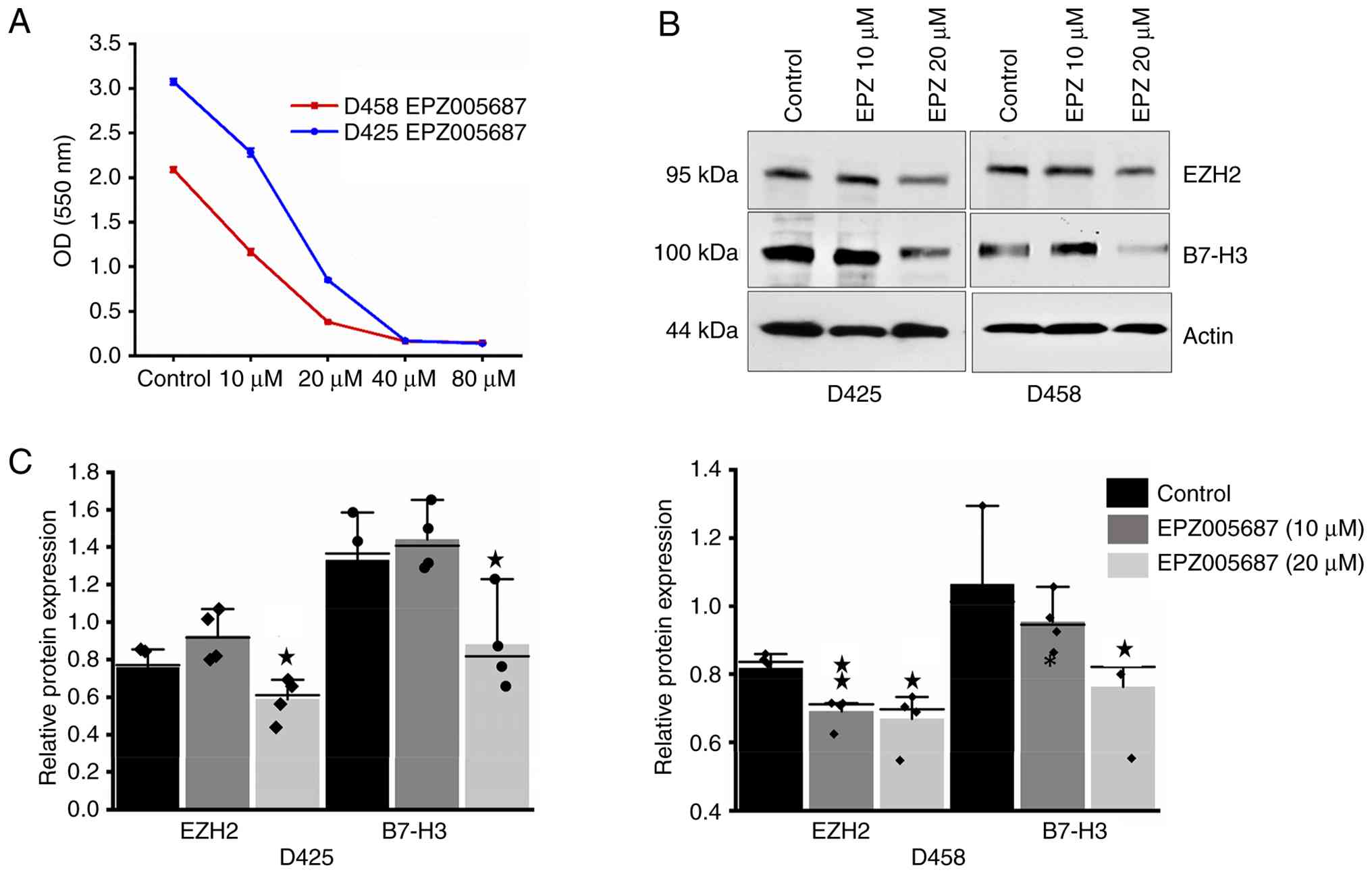
The D425 and D458 cell lines were treated with increasing
concentrations of EPZ005687 (0, 10, 20, 40 and 80 µM) for 5 days.
C17.2 and C17.2-EZH2-OE cell lines were treated with increasing
concentrations of EPZ005687 (0, 10, 15, 20, 25, 30 and 40 µM), IgG
(0, 2.5, 5 and 10 µg), or MJ18 (0, 2.5, 5, 10 µg) for 5 days. After
treatment, the medium was replaced with 100 µl sterile MTT solution
(cat. no. M6494; Thermo Fisher Scientific, Inc.) per well (0.5
µg/ml), and cells were incubated for 2 h in a cell culture
incubator at 37°C. The MTT crystals were solubilized in DMSO (cat.
no. D8418; MilliporeSigma) for 30 min. Absorbance was read using a
SpectraMax M5 spectrophotometer (Molecular Devices, LLC) at 550
nm.
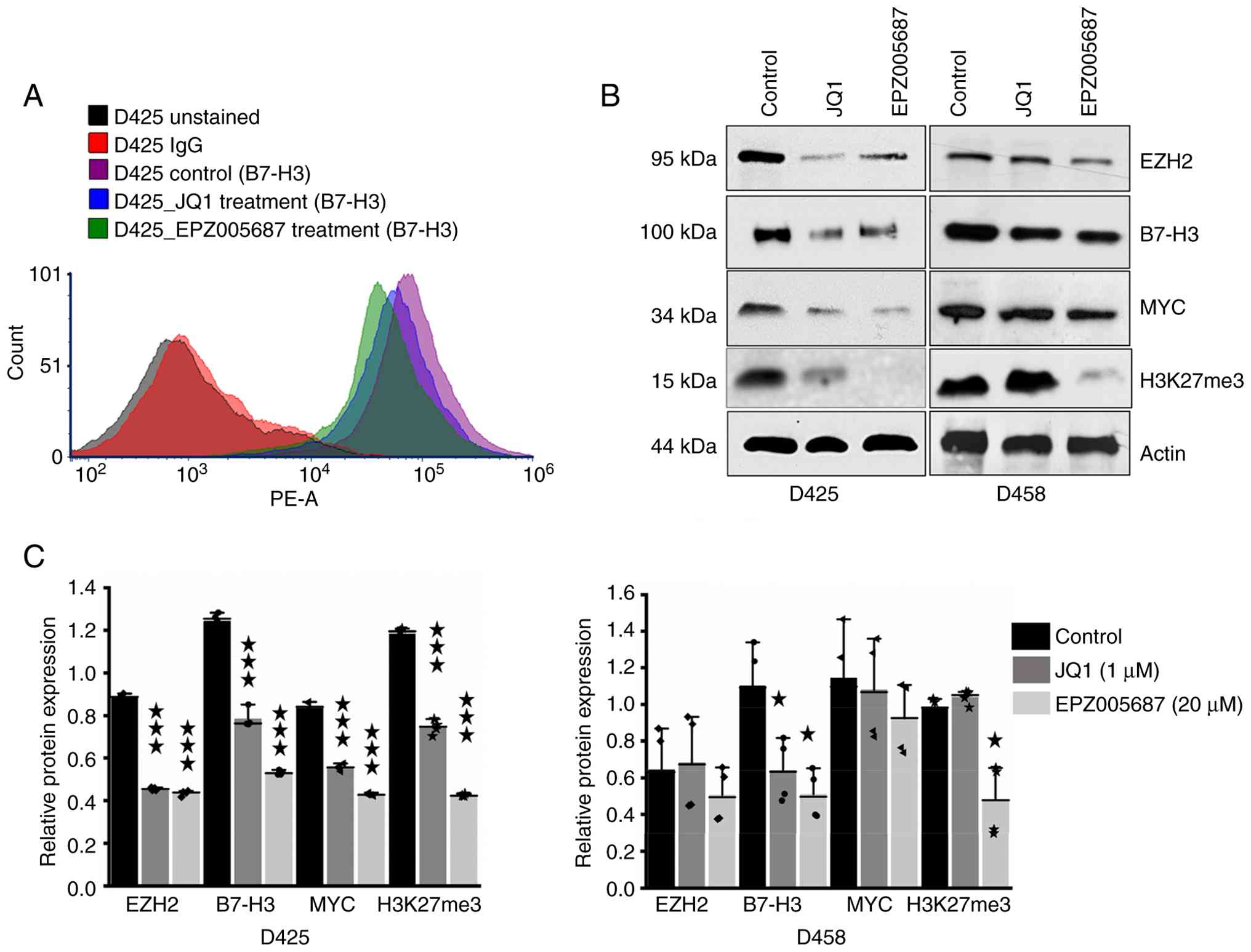
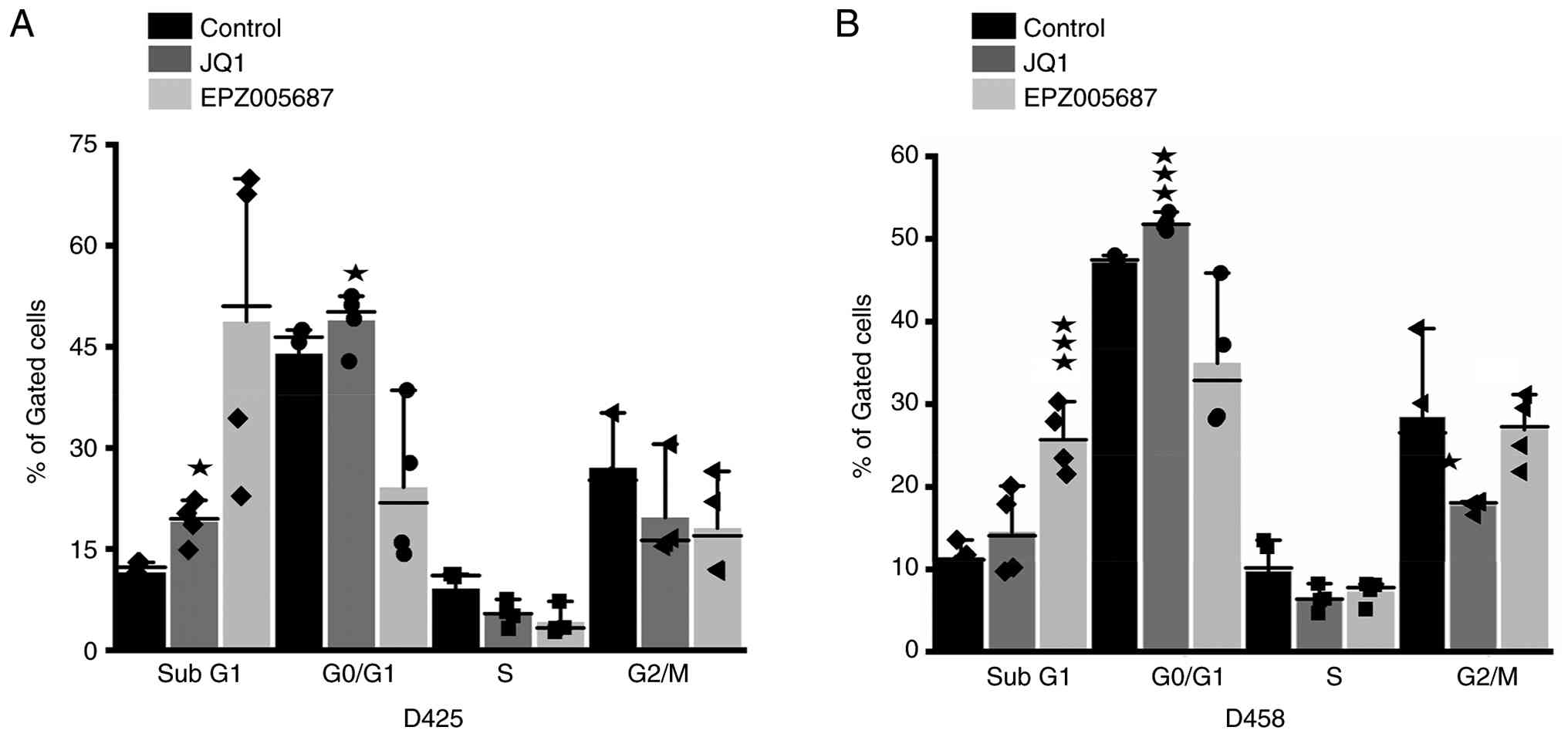
For PE staining, D425 cells were cultured in 100-mm
dishes to 80% confluency. A total of 24 h after inoculation, the
cells were randomized and treated with JQ1 (1 µM) or EPZ005687 (20
µM) for 24 h. Harvested cells were fixed in ice-cold (−20°C) 70%
ethanol overnight. Cells were then washed twice with 1X PBS and
labeled with 5 µl (0.25 µg/million cells) PE anti-human CD276
(B7-H3) antibody (cat. no. 331605; BioLegend, Inc.). For the cell
cycle analysis, D425, D458, C17.2 and C17.2 EZH2-OE cells
(1×106, n=4) were seeded in 60-mm plates. After 24 h,
the media was removed and replaced with media alone (control), JQ1
(1 µM), or EPZ005687 (20 µM) and incubated further for 24 h. Cells
were harvested, washed with 1X PBS, and then fixed in 1 ml ice cold
70% ethanol at −20°C overnight. Fixed cells were then washed with
PBS and resuspended in 500 µl of propidium iodide staining solution
(50 µg/ml) (cat. no. 1032; BioSure). The cell cycle was analyzed by
FACS Caliber System (Becton Dickinson and Company) following
standard protocol (49). The
proportion of cells in each phase of the cell cycle was assessed
using Cell Quest software (Becton, Dickinson and Company). For the
clonogenic assay (colony formation assay), D425 and D458 cells were
seeded in 60-mm plates at 50 and 100 cells for 24 h. After which,
the cells were treated with JQ1 (1 µM) or EPZ005687 (20 µM). The
cells were allowed to form colonies over a period of 14 days.
Colonies were measured as a cluster of 50 or more cells. Media and
inhibitor treatments were changed every three days. Colonies were
fixed with 4% paraformaldehyde for 20 min and stained using crystal
violet (0.5%) overnight. The samples were then quantified by manual
inspection.
Statistical analysis and graphing were performed
using Origin version 9.0 software (Microcal Software Inc.).
Statistical significance for independent samples was calculated
using one-way ANOVA with Tukey's post hoc test. Data are expressed
as the median ± SD. P<0.05 was considered to indicate a
statistically significant difference.
B7-H3 levels are significantly elevated in patients
with hepatocellular carcinoma, non-small cell lung cancer, renal
cell carcinoma and osteosarcoma, with greater levels of B7-H3
associated with higher tumor stage, tumor size, and metastasis
status (50–52). Similarly, EZH2 expression increases
with tumor grade and is associated with poor patient prognosis
(53), prompting interest in
targeting epigenetic modifiers such as EZH2 in MB (53,54).
As previously revealed, B7-H3 amplifications are associated with
reduced survival in MB patients (24). Previous studies suggested that EZH2
may regulate B7-H3 expression (34,55),
thus the expression of EZH2 and B7-H3 in Group 3 MB patient
survival was compared by data mining using the publicly available
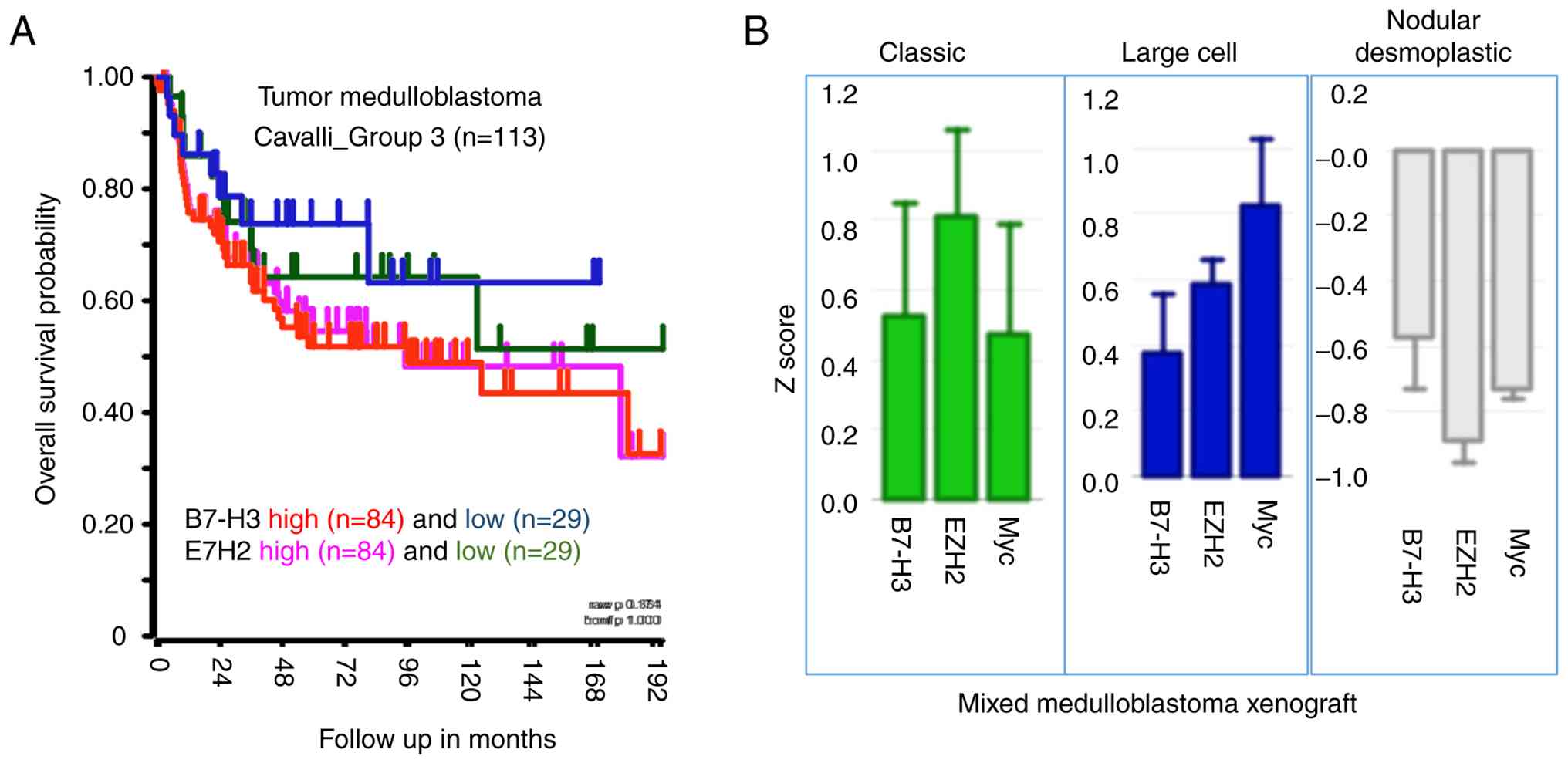
MB patient datasets. Kaplan-Meier survival curves plotted using the
Cavalli dataset revealed a significant association between Group 3
MB patients with high EZH2 expression (n=84) and poor overall
survival when compared with patients with low EZH2 expression
(n=29, P<0.001). Similarly, MB patients with high B7-H3
expression (n=84) had lower overall survival when compared with
patients with low B7-H3 expression (n=29, P<0.05) (Fig. 1A), highlighting the clinical
significance of targeting EZH2 in MYC amplified Group 3 MB. To
further identify the relationship among EZH2, B7-H3, and MYC
transcript expression in MB, a data mining analysis (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi) of
xenografts representative of different MB histological variants was
performed; classic (n=14), large cell (n=8) and
desmoplastic/nodular (n=18) (Fig.
1B). For this analysis, the relative transcript levels among
EZH2, B7-H3 and MYC in the different MB histological variants were
compared by their respective Z score. Similar to Group 3 MB, the
large cell/anaplastic MB variant is frequently associated with MYC
amplification (9,10,56,57).
In agreement with previous studies, the present analysis showed
that both EZH2 and MYC transcript levels were highly expressed in
both classic and large cell MB variants (58,59).
By contrast, nodular desmoplastic MB xenografts exhibited lower
EZH2, MYC and B7-H3 transcript levels.
It has been reported that EZH2 directly interacts
with MYC and MYCN, with EZH2 depletion resulting in reduced MYC
expression and transcriptional activity (38). A similar study revealed that EZH2
and MYC induce each other's expression through miRNA targeting
(34). To investigate whether such
mechanisms may participate in MB, RT-qPCR analysis was performed on
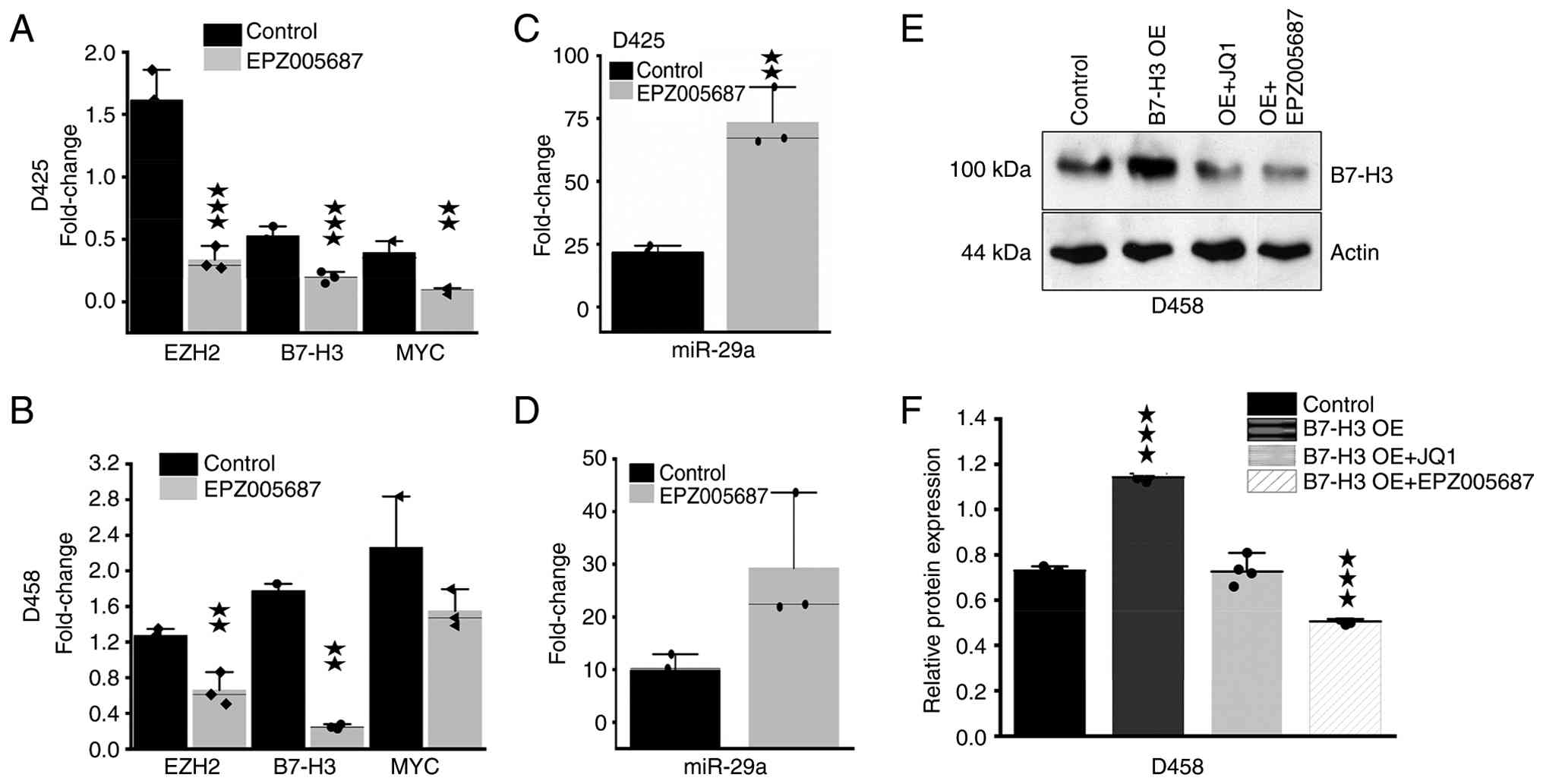
D425 and D458 cells treated with or without EPZ005687 for 24 h. The
results demonstrated that in D425 cells, EPZ005687 treatment
significantly reduced the mRNA levels of EZH2 (P<0.001), B7-H3
(P<0.001) and MYC (P<0.01) when compared with controls
(Fig. 5A). EPZ005687 treatment also
led to significant downregulation of EZH2 (P<0.01) and B7-H3
(P<0.01) mRNA levels in D458 cells (Fig. 5B). These results indicated that
EPZ005687 reduces B7-H3 expression in Group 3 MB cells at the
post-transcriptional level.
In addition to EZH2 and MYC, miR-29a is known to
regulate B7-H3 expression through translational suppression
(35,73). Importantly, both EZH2 and MYC can
bind to the promoter regions of miR-29a/b1 and miR29b2/c, leading
to miR-29 suppression (34). Thus,
it stands to reason that EZH2 and MYC may indirectly augment B7-H3
expression through targeting miR-29a/b. This notion is supported by
a previous study by the authors which demonstrated that the
endogenous levels of miR-29a are low in D425 and D458 cells;
However, JQ1 significantly increased miR-29a/b transcript levels,
whereas forced expression of miR-29a/b significantly reduced B7-H3
levels (24). As EZH2 inhibition
downregulated B7-H3 protein levels (Fig. 3C), it was wondered whether EZH2 may
regulate B7-H3 expression indirectly through miR-29a/b suppression.
To evaluate this possibility, miR-29a/b expression upon EPZ005687
treatment was assessed. The results from RT-qPCR analysis revealed
that in D425 cells, miR-29 transcript levels were significantly
(P<0.01) increased following EZH2 inhibition (Fig. 5C). miR-29a/b mRNA levels were also
increased in EPZ005687-treated D458 cells (Fig. 5D). As EPZ005687 treatment
downregulated B7-H3 protein levels with corresponding increases in
miR-29a expression, this suggests that EZH2 inhibition may
facilitate miR-29a-mediated B7-H3 suppression.
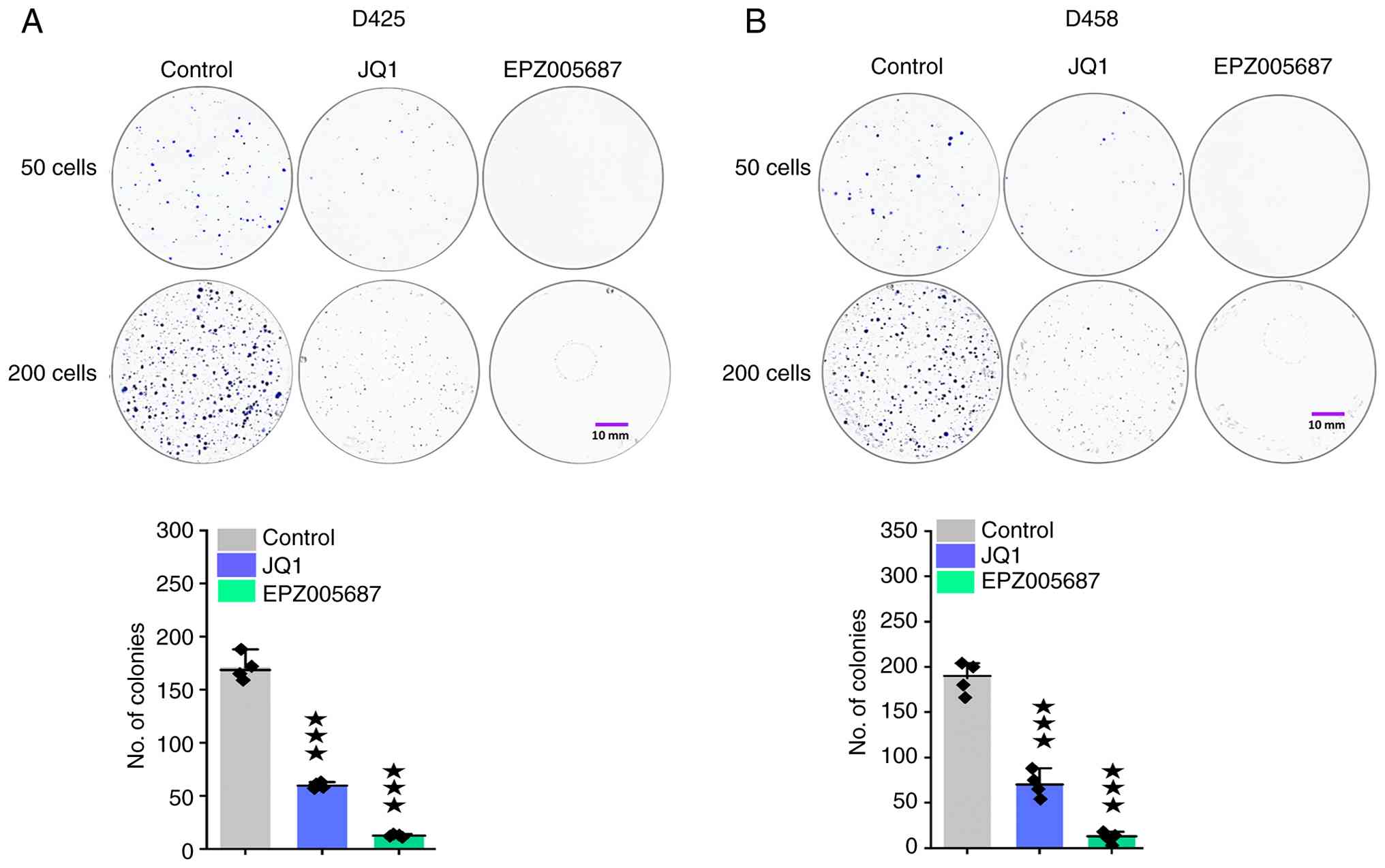
To examine the long-term impacts of EZH2 inhibition,
colony-forming ability in D425 and D458 cells was studied using
clonogenic assay. MB cells were plated at densities of 50 and 200
cells and allowed to proliferate over 14 days. Colonies were
measured as a cluster of 50 or more cells (Fig. 7A and B). Although both JQ1 and
EPZ005687 treatment significantly reduced the number of colonies
compared with controls, it was observed that cells treated with
EPZ005687 formed fewer visible colonies than those treated with
JQ1. Taken together, the present results highlighted the
therapeutic potential of EZH2 inhibition in treating aggressive
MYC-amplified Group 3 MBs.
MB is considered to arise from undifferentiated
neural stem/progenitor cells found in the external granule layer of
the cerebellum (80). In addition
to MYC, human MBs also express significantly elevated levels of
REST/NRSF, a transcriptional suppressor of neuronal differentiation
genes (81,82). NSCs transduced with the MYC oncogene
do not form tumors in mice because they express the endogenous
mouse REST transcripts only under proliferation conditions, which
cease upon neuronal differentiation. By contrast, murine NSCs
stably expressing c-MYC and transduced with the human REST
transgene, continue to express REST/NRSF as they enter neuronal
differentiation. This prevents terminal differentiation into mature
neurons thereby conferring a proliferative advantage and
tumorigenic ability (74). As such,
both c-MYC and REST/NRSF overexpression are requirements for NSCs
to produce cerebellar tumors. As with other NSC models, C17.2 NSCs
do not display anchorage-dependent growth; however, transformation
induced by oncogenes such as REST, confers the ability to grow on
soft agar (75). In MB, inhibition
of EZH2 in REST-transformed C17.2 NSCs impairs their ability to
undergo anchorage-dependent growth on soft agar (28), indicating that EZH2 activity is
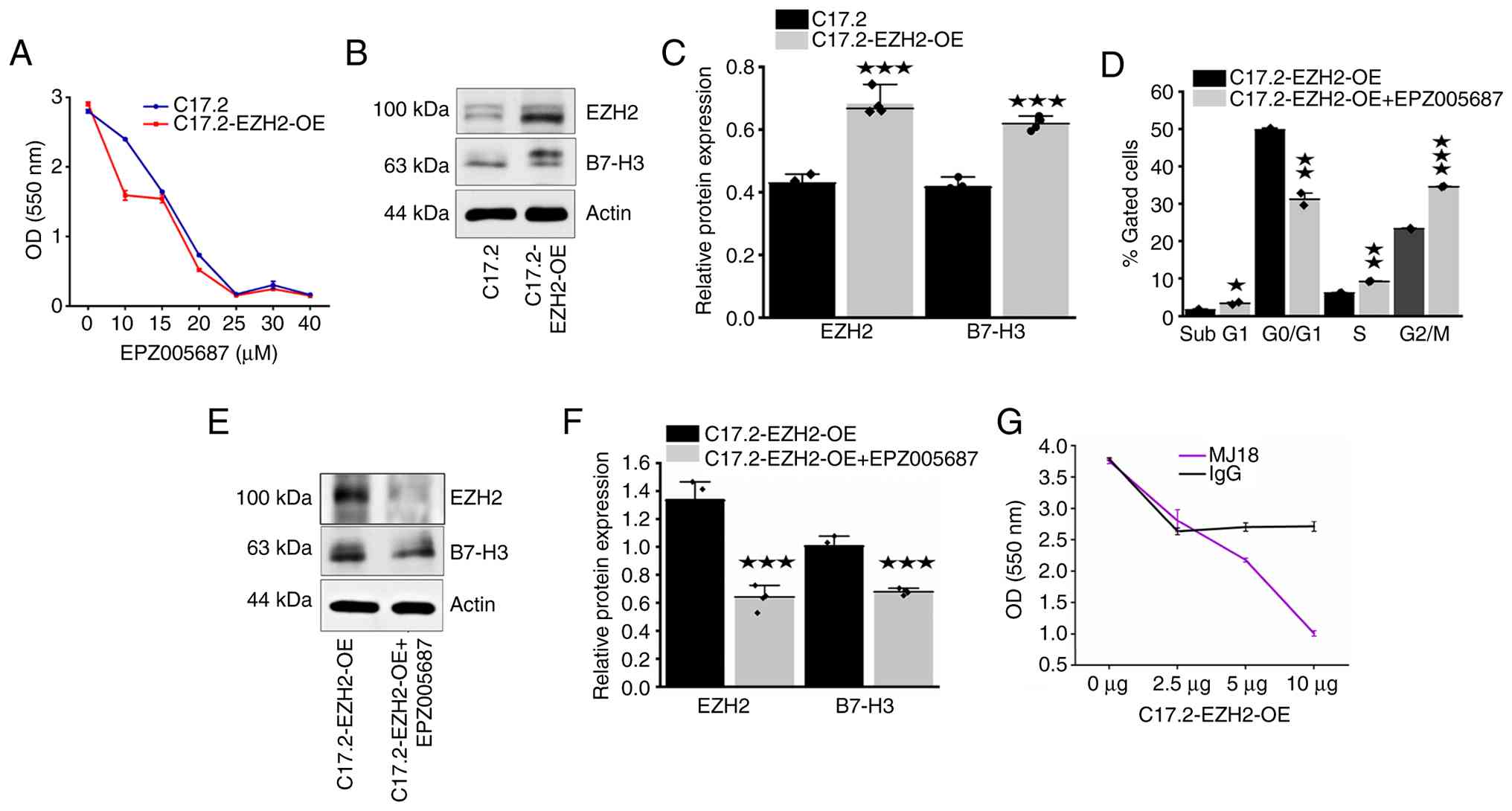
required for this behavior. As such, the C17.2-EZH2-OE NSC line
used in the present study serves as a model to test the efficacy of
EZH2 inhibition in combating MB tumorigenesis. In the current
study, it was found that EPZ005687 treatment significantly reduced
the viability of EZH2-amplified NSCs, in addition to downregulating
B7-H3 expression, supporting the authors' hypothesis that EZH2
promotes MB tumorigenesis by upregulating B7-H3 expression.
Overexpression of EZH2 was first reported in
prostate cancer, with high EZH2 expression being linked with poor
patient prognosis (83). Later
studies found that EZH2 is also overexpressed in breast, gastric
and esophageal cancer, and endometrial carcinoma (84–87),
with EZH2 overexpression being associated with features of
aggressive tumor subgroups, clinical progression and reduced
survival (84,86). Notably, EZH2 mRNA is
significantly higher in MB tumor tissues when compared with normal
tissues (88) and has been proposed
as a potential therapeutic target for the treatment of high-risk MB
tumors (28). Thus, understanding
the oncogenic and regulatory mechanisms of EZH2 has the potential
to improve treatment options for a wide range of cancers.
The authors acknowledge Wade Smith,and Collin
Bomstad, (University of Illinois College of Medicine at Peoria
(UICOMP), IL, USA) for initiating this work in the lab as a part of
the Summer 2018 Craig fellowship program at UICOMP. The authors
also acknowledge Dr. Janardhan Avilala (UICOMP) for his technical
assistance. The authors would like to thank UICOMP administrative
assistant, Christina Constantinidou for her help with the
manuscript formatting.
The present study was supported by William E. McElroy Charitable
Foundation (Springfield, Illinois).
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
SA, IP and KV conceptualized the present study as
part of the McElroy grant. SA, KS, IP and SV performed experiments.
SA, KV and KS performed the data mining studies. SA and KS confirm
the authenticity of all the raw data. SV and RV provided the C17.2
and C17.2-EZH2_OE cells. The methods and materials used in the
publication were previously established by SA. All authors
contributed to the critical discussions and SA, KS and IP
contributed greatly to the analysis of the results. RV made
substantial contributions to the conception of the present study
and data interpretation. KS and SA wrote the manuscript and IP
helped in editing the manuscript. All authors reviewed the
manuscript. All authors read and approved the final manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
|
1
|
Ostrom QT, Gittleman H, Truitt G, Boscia
A, Kruchko C and Barnholtz-Sloan JS: CBTRUS Statistical report:
Primary brain and other central nervous system tumors diagnosed in
the united states in 2011-2015. Neuro Oncol. 20 (Suppl_4):iv1–iv86.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Vladoiu MC, El-Hamamy I, Donovan LK,
Farooq H, Holgado BL, Sundaravadanam Y, Ramaswamy V, Hendrikse LD,
Kumar S, Mack SC, et al: Childhood cerebellar tumours mirror
conserved fetal transcriptional programs. Nature. 572:67–73. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Hovestadt V, Smith KS, Bihannic L, Filbin
MG, Shaw ML, Baumgartner A, DeWitt JC, Groves A, Mayr L, Weisman
HR, et al: Resolving medulloblastoma cellular architecture by
single-cell genomics. Nature. 572:74–79. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Taylor MD, Northcott PA, Korshunov A,
Remke M, Cho YJ, Clifford SC, Eberhart CG, Parsons DW, Rutkowski S,
Gajjar A, et al: Molecular subgroups of medulloblastoma: The
current consensus. Acta Neuropathol. 123:465–472. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ramaswamy V, Remke M, Bouffet E, Faria CC,
Perreault S, Cho YJ, Shih DJ, Luu B, Dubuc AM, Northcott PA, et al:
Recurrence patterns across medulloblastoma subgroups: An integrated
clinical and molecular analysis. Lancet Oncol. 14:1200–1207. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Northcott PA, Korshunov A, Witt H,
Hielscher T, Eberhart CG, Mack S, Bouffet E, Clifford SC, Hawkins
CE, French P, et al: Medulloblastoma comprises four distinct
molecular variants. J Clin Oncol. 29:1408–1414. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Louis DN, Ohgaki H, Wiestler OD, Cavenee
WK, Burger PC, Jouvet A, Scheithauer BW and Kleihues P: The 2007
WHO classification of tumours of the central nervous system. Acta
Neuropathol. 114:97–109. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Louis DN, Perry A, Reifenberger G, von
Deimling A, Figarella-Branger D, Cavenee WK, Ohgaki H, Wiestler OD,
Kleihues P and Ellison DW: The 2016 World Health Organization
classification of tumors of the central nervous system: A summary.
Acta Neuropathol. 131:803–820. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Eberhart CG, Kratz J, Wang Y, Summers K,
Stearns D, Cohen K, Dang CV and Burger PC: Histopathological and
molecular prognostic markers in medulloblastoma: c-myc, N-myc,
TrkC, and anaplasia. J Neuropathol Exp Neurol. 63:441–449. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ellison DW, Kocak M, Dalton J, Megahed H,
Lusher ME, Ryan SL, Zhao W, Nicholson SL, Taylor RE, Bailey S and
Clifford SC: Definition of disease-risk stratification groups in
childhood medulloblastoma using combined clinical, pathologic, and
molecular variables. J Clin Oncol. 29:1400–1407. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Kool M, Jones DT, Jager N, Northcott PA,
Pugh TJ, Hovestadt V, Piro RM, Esparza LA, Markant SL, Remke M, et
al: Genome sequencing of SHH medulloblastoma predicts
genotype-related response to smoothened inhibition. Cancer Cell.
25:393–405. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Millard NE and De Braganca KC:
Medulloblastoma: WHO 2021 and beyond. J Child Neurol. 31:1341–1353.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Remke M, Hielscher T, Northcott PA, Witt
H, Ryzhova M, Wittmann A, Benner A, von Deimling A, Scheurlen W,
Perry A, et al: Adult medulloblastoma comprises three major
molecular variants. J Clin Oncol. 29:2717–2723. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Pugh TJ, Weeraratne SD, Archer TC,
Pomeranz Krummel DA, Auclair D, Bochicchio J, Carneiro MO, Carter
SL, Cibulskis K, Erlich RL, et al: Medulloblastoma exome sequencing
uncovers subtype-specific somatic mutations. Nature. 488:106–110.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Leitner J, Klauser C, Pickl WF, Stöckl J,
Majdic O, Bardet AF, Kreil DP, Dong C, Yamazaki T, Zlabinger G, et
al: B7-H3 is a potent inhibitor of human T-cell activation: No
evidence for B7-H3 and TREML2 interaction. Eur J Immunol.
39:1754–1764. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chapoval AI, Ni J, Lau JS, Wilcox RA,
Flies DB, Liu D, Dong H, Sica GL, Zhu G, Tamada K and Chen L:
B7-H3: A costimulatory molecule for T cell activation and IFN-gamma
production. Nat Immunol. 2:269–274. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hashiguchi M, Kobori H, Ritprajak P,
Kamimura Y, Kozono H and Azuma M: Triggering receptor expressed on
myeloid cell-like transcript 2 (TLT-2) is a counter-receptor for
B7-H3 and enhances T cell responses. Proc Natl Acad Sci USA.
105:10495–10500. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen YW, Tekle C and Fodstad O: The
immunoregulatory protein human B7H3 is a tumor-associated antigen
that regulates tumor cell migration and invasion. Curr Cancer Drug
Targets. 8:404–413. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Han S, Shi X, Liu L, Zong L, Zhang J, Chen
Q, Qian Q, Chen L, Wang Y, Jin J, et al: Roles of B7-H3 in Cervical
Cancer and Its Prognostic Value. J Cancer. 9:2612–2624. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lin L, Cao L, Liu Y, Wang K, Zhang X, Qin
X, Zhao D, Hao J, Chang Y, Huang X, et al: B7-H3 promotes multiple
myeloma cell survival and proliferation by ROS-dependent activation
of Src/STAT3 and c-Cbl-mediated degradation of SOCS3. Leukemia.
33:1475–1486. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kang FB, Wang L, Jia HC, Li D, Li HJ,
Zhang YG and Sun DX: B7-H3 promotes aggression and invasion of
hepatocellular carcinoma by targeting epithelial-to-mesenchymal
transition via JAK2/STAT3/Slug signaling pathway. Cancer Cell Int.
15:452015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang J, Chong KK, Nakamura Y, Nguyen L,
Huang SK, Kuo C, Zhang W, Yu H, Morton DL and Hoon DS: B7-H3
associated with tumor progression and epigenetic regulatory
activity in cutaneous melanoma. J Invest Dermatol. 133:2050–2058.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhong C, Tao B, Chen Y, Guo Z, Yang X,
Peng L, Xia X and Chen L: B7-H3 regulates glioma growth and cell
invasion through a JAK2/STAT3/Slug-Dependent signaling pathway.
Onco Targets Ther. 13:2215–2224. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Purvis IJ, Avilala J, Guda MR,
Venkataraman S, Vibhakar R, Tsung AJ, Velpula KK and Asuthkar S:
Role of MYC-miR-29-B7-H3 in medulloblastoma growth and
angiogenesis. J Clin Med. 8:11582019. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Purvis IJ, Velpula KK, Guda MR, Nguyen D,
Tsung AJ and Asuthkar S: B7-H3 in medulloblastoma-Derived exosomes;
A novel tumorigenic role. Int J Mol Sci. 21:70502020. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Loo D, Alderson RF, Chen FZ, Huang L,
Zhang W, Gorlatov S, Burke S, Ciccarone V, Li H, Yang Y, et al:
Development of an Fc-enhanced anti-B7-H3 monoclonal antibody with
potent antitumor activity. Clin Cancer Res. 18:3834–3845. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Karnell JL, Dimasi N, Karnell FG III,
Fleming R, Kuta E, Wilson M, Wu H, Gao C, Herbst R and Ettinger R:
CD19 and CD32b differentially regulate human B cell responsiveness.
J Immunol. 192:1480–1490. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Alimova I, Venkataraman S, Harris P,
Marquez VE, Northcott PA, Dubuc A, Taylor MD, Foreman NK and
Vibhakar R: Targeting the enhancer of zeste homologue 2 in
medulloblastoma. Int J Cancer. 131:1800–1809. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Pasini D and Di Croce L: Emerging roles
for Polycomb proteins in cancer. Curr Opin Genet Dev. 36:50–58.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Simon JA and Lange CA: Roles of the EZH2
histone methyltransferase in cancer epigenetics. Mutat Res.
647:21–29. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chang TC, Yu D, Lee YS, Wentzel EA, Arking
DE, West KM, Dang CV, Thomas-Tikhonenko A and Mendell JT:
Widespread microRNA repression by Myc contributes to tumorigenesis.
Nat Genet. 40:43–50. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Kriegel AJ, Liu Y, Fang Y, Ding X and
Liang M: The miR-29 family: Genomics, cell biology, and relevance
to renal and cardiovascular injury. Physiol Genomics. 44:237–244.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Yin H, Wang Y, Wu Y, Zhang X, Zhang X, Liu
J, Wang T, Fan J, Sun J, Yang A and Zhang R: EZH2-mediated
Epigenetic Silencing of miR-29/miR-30 targets LOXL4 and contributes
to tumorigenesis, metastasis, and immune microenvironment
remodeling in breast cancer. Theranostics. 10:8494–8512. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhang X, Zhao X, Fiskus W, Lin J, Lwin T,
Rao R, Zhang Y, Chan JC, Fu K, Marquez VE, et al: Coordinated
silencing of MYC-mediated miR-29 by HDAC3 and EZH2 as a therapeutic
target of histone modification in aggressive B-Cell lymphomas.
Cancer Cell. 22:506–523. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Ganju A, Khan S, Hafeez BB, Behrman SW,
Yallapu MM, Chauhan SC and Jaggi M: miRNA nanotherapeutics for
cancer. Drug Discov Today. 22:424–432. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Italiano A, Soria J, Toulmonde M, Michot
JM, Lucchesi C, Varga A, Coindre JM, Blakemore SJ, Clawson A,
Suttle B, et al: Tazemetostat, an EZH2 inhibitor, in relapsed or
refractory B-cell non-Hodgkin lymphoma and advanced solid tumours:
A first-in-human, open-label, phase 1 study. Lancet Oncol.
19:649–659. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Morschhauser F, Tilly H, Chaidos A, McKay
P, Phillips T, Assouline S, Batlevi CL, Campbell P, Ribrag V, Damaj
GL, et al: Tazemetostat for patients with relapsed or refractory
follicular lymphoma: An open-label, single-arm, multicentre, phase
2 trial. Lancet Oncol. 21:1433–1442. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Zhao Y, Li T, Tian S, Meng W, Sui Y, Yang
J, Wang B, Liang Z, Zhao H, Han Y, et al: Effective inhibition of
MYC-Amplified Group 3 medulloblastoma through targeting EIF4A1.
Cancer Manag Res. 12:12473–12485. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Venkataraman S, Birks DK, Balakrishnan I,
Alimova I, Harris PS, Patel PR, Handler MH, Dubuc A, Taylor MD,
Foreman NK and Vibhakar R: MicroRNA 218 acts as a tumor suppressor
by targeting multiple cancer phenotype-associated genes in
medulloblastoma. J Biol Chem. 288:1918–1928. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Parker MA, Anderson JK, Corliss DA,
Abraria VE, Sidman RL, Park KI, Teng YD, Cotanche DA and Snyder EY:
Expression profile of an operationally-defined neural stem cell
clone. Exp Neurol. 194:320–332. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Asuthkar S, Velpula KK, Nalla AK, Gogineni
VR, Gondi CS and Rao JS: Irradiation-induced angiogenesis is
associated with an MMP-9-miR-494-syndecan-1 regulatory loop in
medulloblastoma cells. Oncogene. 33:1922–1933. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Asuthkar S, Gondi CS, Nalla AK, Velpula
KK, Gorantla B and Rao JS: Urokinase-type plasminogen activator
receptor (uPAR)-mediated regulation of WNT/β-catenin signaling is
enhanced in irradiated medulloblastoma cells. J Biol Chem.
287:20576–20589. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Asuthkar S, Venkataraman S, Avilala J,
Shishido K, Vibhakar R, Veo B, Purvis IJ, Guda MR and Velpula KK:
SMYD3 promotes cell cycle progression by inducing cyclin D3
transcription and stabilizing the cyclin D1 protein in
medulloblastoma. Cancers (Basel). 14:16732022. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Huang Y, Zhang HL, Li ZL, Du T, Chen YH,
Wang Y, Ni HH, Zhang KM, Mai J, Hu BX, et al: FUT8-mediated
aberrant N-glycosylation of B7H3 suppresses the immune response in
triple-negative breast cancer. Nat Commun. 12:26722021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Kim K, Yoo HC, Kim BG, Kim S, Sung Y, Yoon
I, Yu YC, Park SJ, Kim JH, Myung K, et al: O-GlcNAc modification of
leucyl-tRNA synthetase 1 integrates leucine and glucose
availability to regulate mTORC1 and the metabolic fate of leucine.
Nat Commun. 13:29042022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Li Y, Zhang M, Feng H and Mahati S: The
tumorigenic properties of EZH2 are mediated by MiR-26a in uveal
melanoma. Front Mol Biosci. 8:7135422021. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Girault V, Gilard V, Marguet F, Lesueur C,
Hauchecorne M, Ramdani Y, Laquerrière A, Marret S, Jégou S,
Gonzalez BJ, et al: Prenatal alcohol exposure impairs autophagy in
neonatal brain cortical microvessels. Cell Death Dis. 8:e26102017.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Yang L, Li Z and Ouyang Y: Taurine
attenuates ER stressassociated apoptosis and catabolism in nucleus
pulposus cells. Mol Med Rep. 25:e26102017.
|
|
49
|
Asuthkar S, Nalla AK, Gondi CS, Dinh DH,
Gujrati M, Mohanam S and Rao JS: Gadd45a sensitizes medulloblastoma
cells to irradiation and suppresses MMP-9-mediated EMT. Neuro
Oncol. 13:1059–1073. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Zhang G, Xu Y, Lu X, Huang H, Zhou Y, Lu B
and Zhang X: Diagnosis value of serum B7-H3 expression in non-small
cell lung cancer. Lung Cancer. 66:245–249. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Xu F, Yi J, Wang F, Wang W, Wang Z, Xue J
and Luan X: Involvement of soluble B7-H3 in combination with the
serum inflammatory cytokines interleukin-17, −8 and −6 in the
diagnosis of hepatocellular carcinoma. Oncol Lett. 14:8138–8143.
2017.PubMed/NCBI
|
|
52
|
Wang L, Kang FB, Zhang GC, Wang J, Xie MF
and Zhang YZ: Clinical significance of serum soluble B7-H3 in
patients with osteosarcoma. Cancer Cell Int. 18:1152018. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Miele E, Valente S, Alfano V, Silvano M,
Mellini P, Borovika D, Marrocco B, Po A, Besharat ZM, Catanzaro G,
et al: The histone methyltransferase EZH2 as a druggable target in
SHH medulloblastoma cancer stem cells. Oncotarget. 8:68557–68570.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Zwergel C, Romanelli A, Stazi G, Besharat
ZM, Catanzaro G, Tafani M, Valente S and Mai A: Application of
small epigenetic modulators in pediatric medulloblastoma. Front
Pediatr. 6:3702018. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Xu H, Cheung IY, Guo HF and Cheung NK:
MicroRNA miR-29 modulates expression of immunoinhibitory molecule
B7-H3: Potential implications for immune based therapy of human
solid tumors. Cancer Res. 69:6275–6281. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Brown HG, Kepner JL, Perlman EJ, Friedman
HS, Strother DR, Duffner PK, Kun LE, Goldthwaite PT and Burger PC:
‘Large cell/anaplastic’ medulloblastomas: A pediatric oncology
group study. J Neuropathol Exp Neurol. 59:857–865. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Leonard JR, Cai DX, Rivet DJ, Kaufman BA,
Park TS, Levy BK and Perry A: Large cell/anaplastic
medulloblastomas and medullomyoblastomas: Clinicopathological and
genetic features. J Neurosurg. 95:82–88. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Liu H, Sun Q, Sun Y, Zhang J, Yuan H, Pang
S, Qi X, Wang H, Zhang M, Zhang H, et al: MELK and EZH2 cooperate
to regulate medulloblastoma cancer stem-like cell proliferation and
differentiation. Mol Cancer Res. 15:1275–1286. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Alassiri AH, Alsufiani FM, Almutairi AA,
Almohini IA, Aldosari MA and Essa MF: Spectrum of medulloblastoma
subtypes and frequency of MYC amplification; Experience from a
tertiary care center in Saudi Arabia. Neurosciences (Riyadh).
25:218–221. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Habib S, Ariatti M and Singh M: Anti-c-myc
RNAi-Based Onconanotherapeutics. Biomedicines. 8:6122020.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Webb MS, Tortora N, Cremese M, Kozlowska
H, Blaquiere M, Devine DV and Kornbrust DJ: Toxicity and
toxicokinetics of a phosphorothioate oligonucleotide against the
c-myc oncogene in cynomolgus monkeys. Antisense Nucleic Acid Drug
Dev. 11:155–163. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Gjorgjieva M, Sobolewski C, Dolicka D,
Correia de Sousa M and Foti M: miRNAs and NAFLD: From
pathophysiology to therapy. Gut. 68:2065–2079. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Zhang H, Zhu D, Zhang Z, Kaluz S, Yu B,
Devi NS, Olson JJ and Van Meir EG: EZH2 targeting reduces
medulloblastoma growth through epigenetic reactivation of the
BAI1/p53 tumor suppressor pathway. Oncogene. 39:1041–1048. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Natsumeda M, Liu Y, Nakata S, Miyahara H,
Hanaford A, Ahsan S, Stearns D, Skuli N, Kahlert UD, Raabe EH, et
al: Inhibition of enhancer of zest homologue 2 is a potential
therapeutic target for high-MYC medulloblastoma. Neuropathology.
39:71–77. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Li Z, Takenobu H, Setyawati AN, Akita N,
Haruta M, Satoh S, Shinno Y, Chikaraishi K, Mukae K, Akter J, et
al: EZH2 regulates neuroblastoma cell differentiation via NTRK1
promoter epigenetic modifications. Oncogene. 37:2714–2727. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Duan R, Du W and Guo W: EZH2: A novel
target for cancer treatment. J Hematol Oncol. 13:1042020.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Miranda TB, Cortez CC, Yoo CB, Liang G,
Abe M, Kelly TK, Marquez VE and Jones PA: DZNep is a global histone
methylation inhibitor that reactivates developmental genes not
silenced by DNA methylation. Mol Cancer Ther. 8:1579–1588. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Tan J, Yang X, Zhuang L, Jiang X, Chen W,
Lee PL, Karuturi RK, Tan PB, Liu ET and Yu Q: Pharmacologic
disruption of Polycomb-repressive complex 2-mediated gene
repression selectively induces apoptosis in cancer cells. Genes
Dev. 21:1050–1063. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Hubaux R, Thu KL, Coe BP, MacAulay C, Lam
S and Lam WL: EZH2 promotes E2F-driven SCLC tumorigenesis through
modulation of apoptosis and cell-cycle regulation. J Thorac Oncol.
8:1102–1106. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Cao Z, Wu W, Wei H, Zhang W, Huang Y and
Dong Z: Downregulation of histone-lysine N-methyltransferase EZH2
inhibits cell viability and enhances chemosensitivity in lung
cancer cells. Oncol Lett. 21:262021.PubMed/NCBI
|
|
71
|
Zhang J, Liu L, Han S, Li Y, Qian Q, Zhang
Q, Zhang H, Yang Z and Zhang Y: B7-H3 is related to tumor
progression in ovarian cancer. Oncol Rep. 38:2426–2434. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Zhang W, Zhang L, Qian J, Lin J, Chen Q,
Yuan Q, Zhou J, Zhang T, Shi J and Zhou H: Expression
characteristic of 4Ig B7-H3 and 2Ig B7-H3 in acute myeloid
leukemia. Bioengineered. 12:11987–12002. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Wang J, Chen X, Xie C, Sun M, Hu C, Zhang
Z, Luan L, Zhou J, Zhou J, Zhu X, et al: MicroRNA miR-29a inhibits
colon cancer progression by downregulating B7-H3 Expression:
Potential molecular targets for colon cancer therapy. Mol
Biotechnol. 63:849–861. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Su X, Gopalakrishnan V, Stearns D, Aldape
K, Lang FF, Fuller G, Snyder E, Eberhart CG and Majumder S:
Abnormal expression of REST/NRSF and Myc in neural stem/progenitor
cells causes cerebellar tumors by blocking neuronal
differentiation. Mol Cell Biol. 26:1666–1678. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Venkataraman S, Alimova I, Fan R, Harris
P, Foreman N and Vibhakar R: MicroRNA 128a increases intracellular
ROS level by targeting Bmi-1 and inhibits medulloblastoma cancer
cell growth by promoting senescence. PLoS One. 5:e107482010.
View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Yamato I, Sho M, Nomi T, Akahori T,
Shimada K, Hotta K, Kanehiro H, Konishi N, Yagita H and Nakajima Y:
Clinical importance of B7-H3 expression in human pancreatic cancer.
Br J Cancer. 101:1709–1716. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Sander S, Bullinger L, Klapproth K,
Fiedler K, Kestler HA, Barth TF, Möller P, Stilgenbauer S, Pollack
JR and Wirth T: MYC stimulates EZH2 expression by repression of its
negative regulator miR-26a. Blood. 112:4202–4212. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Koh CM, Iwata T, Zheng Q, Bethel C,
Yegnasubramanian S and De Marzo AM: Myc enforces overexpression of
EZH2 in early prostatic neoplasia via transcriptional and
post-transcriptional mechanisms. Oncotarget. 2:669–683. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Salvatori B, Iosue I, Djodji Damas N,
Mangiavacchi A, Chiaretti S, Messina M, Padula F, Guarini A,
Bozzoni I, Fazi F and Fatica A: Critical role of c-Myc in acute
myeloid leukemia involving direct regulation of miR-26a and histone
methyltransferase EZH2. Genes Cancer. 2:585–592. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Eberhart CG and Burger PC: Anaplasia and
grading in medulloblastomas. Brain Pathol. 13:376–385. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Ballas N, Grunseich C, Lu DD, Speh JC and
Mandel G: REST and its corepressors mediate plasticity of neuronal
gene chromatin throughout neurogenesis. Cell. 121:645–657. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Chen ZF, Paquette AJ and Anderson DJ:
NRSF/REST is required in vivo for repression of multiple neuronal
target genes during embryogenesis. Nat Genet. 20:136–142. 1998.
View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Varambally S, Dhanasekaran SM, Zhou M,
Barrette TR, Kumar-Sinha C, Sanda MG, Ghosh D, Pienta KJ, Sewalt
RG, Otte AP, et al: The polycomb group protein EZH2 is involved in
progression of prostate cancer. Nature. 419:624–629. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Bachmann IM, Halvorsen OJ, Collett K,
Stefansson IM, Straume O, Haukaas SA, Salvesen HB, Otte AP and
Akslen LA: EZH2 expression is associated with high proliferation
rate and aggressive tumor subgroups in cutaneous melanoma and
cancers of the endometrium, prostate, and breast. J Clin Oncol.
24:268–273. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Qiu BQ, Lin XH, Ye XD, Huang W, Pei X,
Xiong D, Long X, Zhu SQ, Lu F, Lin K, et al: Long non-coding RNA
PSMA3-AS1 promotes malignant phenotypes of esophageal cancer by
modulating the miR-101/EZH2 axis as a ceRNA. Aging (Albany NY).
12:1843–1856. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Gan L, Xu M, Hua R, Tan C, Zhang J, Gong
Y, Wu Z, Weng W, Sheng W and Guo W: The polycomb group protein EZH2
induces epithelial-mesenchymal transition and pluripotent phenotype
of gastric cancer cells by binding to PTEN promoter. J Hematol
Oncol. 11:92018. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Krill L, Deng W, Eskander R, Mutch D,
Zweizig S, Hoang B, Ioffe O, Randall L, Lankes H, Miller DS and
Birrer M: Overexpression of enhance of Zeste homolog 2 (EZH2) in
endometrial carcinoma: An NRG oncology/gynecologic oncology group
study. Gynecol Oncol. 156:423–429. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Xue P, Huang S, Han X, Zhang C, Yang L,
Xiao W, Fu J, Li H and Zhou Y: Exosomal miR-101-3p and miR-423-5p
inhibit medulloblastoma tumorigenesis through targeting FOXP4 and
EZH2. Cell Death Differ. 29:82–95. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Roussel MF and Robinson GW: Role of MYC in
Medulloblastoma. Cold Spring Harb Perspect Med. 3:a0143082013.
View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Dhall G, Grodman H, Ji L, Sands S, Gardner
S, Dunkel IJ, McCowage GB, Diez B, Allen JC, Gopalan A, et al:
Outcome of children less than three years old at diagnosis with
non-metastatic medulloblastoma treated with chemotherapy on the
‘Head Start’ I and II protocols. Pediatr Blood Cancer.
50:1169–1175. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Huether R, Dong L, Chen X, Wu G, Parker M,
Wei L, Ma J, Edmonson MN, Hedlund EK, Rusch MC, et al: The
landscape of somatic mutations in epigenetic regulators across
1,000 paediatric cancer genomes. Nat Commun. 5:36302014. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Feinberg AP, Koldobskiy MA and Gondor A:
Epigenetic modulators, modifiers and mediators in cancer aetiology
and progression. Nat Rev Genet. 17:284–299. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Veo B, Danis E, Pierce A, Sola I, Wang D,
Foreman NK, Jin J, Ma A, Serkova N, Venkataraman S and Vibhakar R:
Combined functional genomic and chemical screens identify SETD8 as
a therapeutic target in MYC-driven medulloblastoma. JCI Insight.
4:e1229332019. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Cheng Y, He C, Wang M, Ma X, Mo F, Yang S,
Han J and Wei X: Targeting epigenetic regulators for cancer
therapy: Mechanisms and advances in clinical trials. Signal
Transduct Target Ther. 4:622019. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Pei Y, Liu KW, Wang J, Garancher A, Tao R,
Esparza LA, Maier DL, Udaka YT, Murad N, Morrissy S, et al: HDAC
and PI3K antagonists cooperate to inhibit growth of MYC-Driven
medulloblastoma. Cancer Cell. 29:311–323. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Robinson G, Parker M, Kranenburg TA, Lu C,
Chen X, Ding L, Phoenix TN, Hedlund E, Wei L, Zhu X, et al: Novel
mutations target distinct subgroups of medulloblastoma. Nature.
488:43–48. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Jones DT, Jäger N, Kool M, Zichner T,
Hutter B, Sultan M, Cho YJ, Pugh TJ, Hovestadt V, Stütz AM, et al:
Dissecting the genomic complexity underlying medulloblastoma.
Nature. 488:100–105. 2012. View Article : Google Scholar : PubMed/NCBI
|