Introduction
Lung cancer is the most malignant cancer worldwide,
with ~1.76 million deaths annually (1). The two major subtypes of lung cancer
are non-small cell lung cancer (NSCLC), accounting for ~80% of all
lung cancer cases, and small cell lung cancer (2). At present, the common first-line
therapeutic regimens for lung cancer are chemotherapy,
immunotherapy and radiation therapies (3–5).
Despite continuous development of novel therapies, cisplatin (DDP)
remains an essential and widely used agent in the multidisciplinary
management of NSCLC. However, the 5-year-survival rate is only 17%
for patients with NSCLC (6). One of
the primary causes for this is DDP resistance, which attenuates the
effect of clinical treatment (7).
The underlying mechanisms of DDP resistance are still unclear.
Therefore, an improved understanding of the associated molecular
mechanisms involved in DDP resistance is essential for NSCLC
treatment. Aberrant repair of damaged DNA represents a key
mechanism contributing to DDP resistance (8). Genomic instability leads to continuous
cycles of DNA damage and repair in NSCLC cells. The DNA damage
response (DDR) is activated upon DNA injury to restore genomic
integrity (9). However, the
dysregulation of DDR pathways, such as through aberrant repair, can
promote malignant progression and exacerbate NSCLC development. DDR
has been associated with the resistance of cancer cells to UV
radiation and chemotherapy (10,11).
For instance, the upregulation of several key molecules of DDR,
such as BRCA1/2 and RAD51, can reduce the sensitivity of
chemotherapy and radiotherapy (12,13).
Therefore, a number of anticancer drugs targeting DDR-related
proteins have been developed to improve the sensitivity of cancer
cells to chemotherapeutic drugs (14). However, the genome of NSCLC is
highly unstable, and hence further investigation of the associated
mechanisms is necessary.
After DNA damage, the cell fate depends on the
recognition of DNA damage repair and tolerance factors, as well as
the activation of apoptosis, necrosis, autophagy and senescence
pathways (15). Specifically, the
phosphatidylinositol 3-kinase (PI3K)/Akt pathway plays a crucial
role in regulating cell behaviors such as proliferation and
apoptosis and hence is involved in determining cell fate after DNA
damage (16,17). Tripartite motif (TRIM)-containing
proteins, carrying an N-terminal RING finger, one or two B-boxes
and a coiled-coil domain, have been reported as regulators of the
PI3K/Akt signaling pathway (18).
TRIM13 and TRIM21 are reported to inhibit Akt signaling activation,
whereas TRIM11, TRIM14, TRIM26, TRIM27, TRIM59 and TRIM44 serve as
Akt pathway activators (18).
Moreover, TRIM family members have been found to be associated with
tumor progression. For instance, TRIM15 and TRIM24 contribute to
the progression of NSCLC and prostate cancer, respectively
(19,20). The knockdown of TRIM46 inhibits the
proliferation of breast cancer cells (21). TRIM46 plays an oncogenic role in
osteosarcoma by activating the nuclear factor κB (NF-κB) signaling
pathway (22). Recently, TRIM46 has
been found to promote lung adenocarcinoma growth and contribute to
chemoresistance (23). However, the
function and underlying mechanism of TRIM46 in NSCLC still need
exploration.
The present study was conducted to investigate the
role of TRIM46 in NSCLC. The difference in the expression of TRIM46
in NSCLC DDP-sensitive and DDP-resistant patients was evaluated.
DDP-induced apoptosis and the DNA damage degree were examined by
modulating the TRIM46 protein level in A549 wild-type and
DDP-resistant cells. Moreover, Akt signaling pathway activation and
the DNA damage-related markers were explored at different TRIM46
levels. The impact of TRIM46 on tumor growth was also assessed
in vivo.
Materials and methods
Reagents
The anti-TRIM46 (cat. no. Ab169044; 1:1,000 for
western blotting and 1:200 for immunohistochemical staining),
anti-caspase 3 (cat. no. Ab32351; 1:5,000 for western blotting),
anti-cleaved-caspase 3 (cat. no. Ab32042; 1:800 for western
blotting) and anti-RAD51 (cat. no. Ab133534; 1:1,000 for western
blotting) antibodies were purchased from Abcam. The anti-Akt (cat.
no. 9272; 1:1,000 for western blotting), anti-phosphorylated
(p-)Akt (cat. no. 4060; 1:1,000 for western blotting) and
anti-glyceraldehyde-3-phosphate dehydrogenase (GAPDH) antibodies
(cat. no. 5174; 1:3,000 for western blotting) were purchased from
Cell Signaling Technology, Inc. 3,3′-Diaminobenzidine (DAB)
detection kit (cat. no. 34002), Roswell Park Memorial Institute
1640 medium (RPMI-1640; cat. no. 11875101) and fetal bovine serum
(FBS; cat. no. 10099141C) were purchased from Gibco (Thermo Fisher
Scientific, Inc.). DDP (cat. no. S1166) was purchased from Selleck
Chemicals. LY294002 (cat. no. 440202) was purchased from Merck
KGaA. Horseradish peroxidase (HRP)-conjugated secondary antibodies
(cat. no. A0208 and A0216; 1:1,000 for western blotting), RIPA
lysis buffer (cat. no. P0013) and BCA Protein Assay Kit (cat. no.
P0011) were purchased from Beyotime Biotechnology. Immobilon
Western HRP Substrate (cat. no. WBKLS0100) was purchased from
MilliporeSigma.
Immunohistochemical staining
Tissue microarray (TMA) was purchased from Shanghai
Outdo Biotech Co., Ltd. (cat. no. HLugA180Su09). Tissues were fixed
with 4% paraformaldehyde and then embedded in paraffin. The tissue
sections (4-µm) were deparaffinized and rehydrated in xylene and a
descending series of ethanol concentrations. Antigen retrieval was
performed using sodium citrate acid buffer (pH 6.0) followed by
washing three times with PBS for 3 min. Then, 3% hydrogen peroxide
(100°C, 20 min) was used to inhibit endogenous peroxidase. The
sections were incubated with anti-TRIM46 antibody at 4°C overnight.
The sections were incubated with EliVision plus Polymer HRP
immunohistochemistry kit (Maxim Biotech, Inc.; cat. no. KIT-9903;
1:1,000) for 30 min at 25°C, then with DAB solution for coloration.
The sections were then counterstained with hematoxylin
(Sigma-Aldrich; Merk KGaA; cat. no. H9627) at 25°C for 3 min. A
NanoZoomer system (Hamamatsu Photonics K.K.) was used to collect
the images of the TMA. In total, two researchers evaluated the
immunoreactivity based on the H-score system, considering both the
percentage of positively stained cells (rated 0–4: 0, <5%; 1,
≥5% and <25%; 2, ≥25% and <50%; 3, ≥50% and <75%; 4, ≥75%)
and the intensity of staining (rated 0–3: 0, negative; 1, weak; 2,
moderate; 3, strong). This resulted in a total score ranging from 0
to12. Patients were then classified into high or low expression
groups based on an H-score cut-off point of 6.
Tumor formation assay in a nude mouse
model
A total of 24 female BALB/c nude mice (4 to 6 weeks
old, weighing 20–22 g) were purchased from Shanghai SLAC Laboratory
Animal Co., Ltd. The animals were maintained at a constant
temperature of 25°C with ~50% humidity under a regular 12:12-h
light/dark cycle with food and water available ad libitum.
After 1 week of adaptive feeding, the mice were randomly divided
into four groups (n=6 per group): A549/DDP cell-transplanted group
[short hairpin RNA negative control (shNC)], TRIM46 knockdown
A549/DDP cell-transplanted group (shTRIM46), A549/DDP
cell-transplanted and DDP-treated group (shNC + DDP) and TRIM46
knockdown A549/DDP cell-transplanted and DDP-treated group
(shTRIM46 + DDP). The tumor model was established by subcutaneously
injecting 100 µl 5×106 A549/DDP cells previously
transduced with shTRIM46 or shNC lentivirus into the right side of
the mice. After 12 days, the mice in the shNC + DDP and shTRIM46 +
DDP groups were treated with 5 mg/kg DDP once a week as previously
described (24). Tumor volume was
calculated every 3 days using the following formula: Volume=1/2×
(Length × width2). The maximum tumor volume and diameter
were 840 mm3 and 14.78 mm, respectively. After 3 weeks
of DDP injection, all mice were euthanized by cervical dislocation
under 5% isoflurane, and the tumors were harvested for the terminal
deoxynucleotidyl transferase-mediated dUTP nick end labeling
(TUNEL) and western blot assays. Confirmation of death was
established by the absence of a pedal/toe pinch reflex and the
cessation of breathing for ≥1 min. Animal experiments were
conducted in accordance with the Guide for the Care and Use of
Laboratory Animals and approved by the Animal Care and Use
Committee of Wenzhou Traditional Chinese Medicine Hospital of
Zhejiang Chinese Medical University (Wenzhou, China; approval
no.WTCM-KT-2020002).
Cell lines and cell culture
Human NSCLC cell lines A549 and A549/DDP and the
human bronchial epithelial cell line 16HBE were purchased from
BioLeaf Biotech (Shanghai Baili Biotechnology Co., Ltd.) and
cultured in RPMI-1640 medium supplemented with 10% FBS. A549 cells
were treated with various concentrations of DDP (0, 5 or 10 µM) or
20 µM LY294002, an inhibitor of Akt or vehicle (80 mg/ml DMSO).
A549/DDP cells were treated with 0, 50 or 100 µM DDP. All cells
were maintained at 37°C in a humidified incubator with 5%
CO2.
Western blot analysis
Total protein was extracted from the tumor tissue
homogenate or cell lines using RIPA lysis buffer containing an
inhibitor cocktail (Bio-Rad Laboratories, Inc.) on ice for 10 min.
Total protein concentration was measured using a BCA Protein Assay
kit. Subsequently, the proteins (20 µg per lane) were separated by
10 or 15% sodium dodecyl sulfate-polyacrylamide gel electrophoresis
and transferred to polyvinylidene difluoride membranes
(MilliporeSigma). After blocking with 5% non-fat milk at 25°C for 1
h, the membranes were incubated with primary antibodies at 4°C
overnight, then with the corresponding HRP-conjugated secondary
antibodies for 1 h at 25°C. Finally, the membranes were visualized
using Immobilon Western HRP Substrate and the protein levels were
semi-quantified by band densitometry (Quantity One software,
version 4.62; Bio-Rad Laboratories, Inc.).
RNA isolation and reverse
transcription-quantitative polymerase chain reaction (PCR)
analyses
Following treatment, the cells were harvested using
TRIzol (Invitrogen; Thermo Fisher Scientific, Inc.) reagent to
extract total RNA. After transcribing RNA to cDNA using the
RevertAid First Strand cDNA Synthesis Kit (cat. no. K1622; Thermo
Fisher Scientific Inc.) according to the manufacturer's
instructions, a SYBR Green Kit (cat. no. 4309155; Thermo Fisher
Scientific Inc.) was used to quantify the mRNA level on an ABI7500
system. The PCR cycling conditions were 95°C for 10 min, followed
by 40 cycles at 95°C for 15 sec and 60°C for 45 sec, followed by a
final extension step of 95°C for 15 sec, 60°C for 1 min, 95°C for
15 sec and 60°C for 15 sec. The relative expression of TRIM46
(forward: 5′-GTCCGCATCAGTGCTTTGGG-3′ and reverse:
5′-TCAGGTTCCGGAAAAGCCC-3′) was analyzed using the 2−ΔΔCq
method (25), using GAPDH (forward:
5′-TCAGACACCATGGGGAAGGT-3′ and reverse: 5′-TCCCGTTCTCAGCCATGTAG-3′)
as an internal control.
Construction and packaging of
lentivirus carrying TRIM46 and shTRIM46
The coding sequence of TRIM46 (NM_001282378.2) was
amplified using the following primers: Forward 5′-CGGAATTCATGGCAGAGGGTGAGGATATG-3′
and reverse: 5′-CGGGATCCTCTGCATATCCTCACCCTCTG-3′
and engineered into the pLVX-Puro lentiviral vector (Shanghai
Lianmai Biotechnology Co., Ltd.). The shRNA sequences targeting
TRIM46 (shTRIM46-1, 5′-GGTGAGGATATGCAGACCT-3′; shTRIM46-2,
5′-GATATGCAGACCTTCACTT-3′; and shTRIM46-3,
5′-AGACCTTCACTTCCATCAT-3′), as well as the shNC
(5′-GATACATACTAGATCGACT-3′) vector, were purchased from Shanghai
GeneChem Co., Ltd and engineered into the pLKO.1 puro lentiviral
vector (Addgene, Inc.). The recombinant plasmid pLVX-Puro-TRIM46
(1,000 ng) or pLKO.1 puro-shTRIM46 (1,000 ng), together with psPAX2
(100 ng; Addgene, Inc.) and pMD2G (900 ng; Addgene, Inc.), were
co-transfected into 293T cells (ATCC) seeded in a 9-well plate
(1×105 cells/well) for 6 h at 37°C. The transfection
procedures used Lipofectamine 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Subsequently, 48 h after transfection, the recombinant lentivirus
in the cell supernatant was collected by centrifugation at 5,000 ×
g for 5 min and the purification and titration of recombinant
lentivirus was performed as previously described (24). A549 or A549/DDP cells were plated in
a 6-well plate (5×105 cells/well) and infected with the
recombinant lentivirus-transducing units at an MOI of 20 in the
presence of 8 µg/ml polybrene (Sigma-Aldrich; Merck KGaA) for 24 h
at 37°C. Stable cells were selected by puromycin (3 µg/ml; Thermo
Fisher Scientific) for 4 more days. pLKO.1-scrambled shRNA (shNC)
and blank pLVX-Puro (vector) were used as the negative
controls.
Cell apoptosis
Following treatment, the cells were harvested and
co-stained with Annexin V-fluorescein isothiocyanate and propidium
iodide following the instructions on the Annexin V apoptosis
detection kit (BD Biosciences). The apoptotic cells were then
quantitatively analyzed by flow cytometry (Accuri™ C6; BD
Biosciences) using CellQuest Pro software, version 3.3 (Becton,
Dickinson and Company).
Comet assay
The degree of DDP-induced DNA damage was examined by
the comet assay, as previously described (26). Briefly, A549 cells were transfected
with TRIM46-overexpressing/knockdown lentivirus or the
corresponding control lentivirus and treated with DDP. Then, the
cells were collected, washed with cold phosphate-buffered saline,
mixed with 140 µl of 0.5% agarose at 43°C and then seeded onto
slides previously coated with 1.0% agarose. After solidification at
4°C, the slides were immersed in a cold lysis buffer (2.5 M NaCl,
100 mM EDTA and 10 mM Tris, with 1 ml of Triton X-100 per 100 ml)
in the dark for 1 h at 4°C. Next, the slides were transferred to an
alkaline electrophoresis solution (0.3 M sodium hydroxide and 1 mM
EDTA) for a 40-min incubation at 4°C. Electrophoresis was conducted
at 4°C under alkaline conditions for 20 min at 25 V/cm. The slides
were washed thrice with neutralizing buffer (0.4 M Tris-HCl, pH
7.5) and stained with DAPI. A fluorescence microscope (Nikon
Corporation) was used to obtain the images and the percentage of
tail DNA was determined using CometScore 1.5 software (TriTek
Corp).
Cell proliferation
A total of 5×103 cells were seeded into
96-well culture plates and cultured overnight. Cell proliferation
was measured using a Cell Counting Kit 8 (CCK-8) kit (Bio-Rad
Laboratories, Inc.). Briefly, at the end of the pre-set treatment,
the culture medium was replaced with a mixture of 10 µl CCK-8
solution and 90 µl RPMI-1640. After incubation for 1 h, the optical
density at 450 nm was determined using a microplate reader (Bio-Rad
Laboratories, Inc.).
TUNEL
The tumor samples obtained from each group of mice
were fixed with 4% paraformaldehyde at 25°C for 48 h. Subsequently,
the samples were embedded in paraffin and cut into slices with a
thickness of 4 µm. The TUNEL assay was performed following the
protocols of a TUNEL staining kit (cat. no. 11684817910; Roche
Diagnostics). In brief, the TUNEL reaction was carried out at 37°C
for 1 h. After the reaction, the sections were washed three times
with PBS, incubated with anti-fluorescein antibody-peroxidase for
30 min, stained with DAB for 10 min, washed three times with PBS
and then re-stained with hematoxylin. The tissue sections were then
examined by light microscopy (Olympus Corporation) and the number
of TUNEL-positive cells was counted using ImageJ software version
1.61 (National Institutes of Health). According to the distribution
of apoptotic cells, five positive visual fields from each treatment
group were imaged.
Statistical analysis
The data are presented as mean ± standard deviation.
Statistical analysis was performed using GraphPad Prism 5.0
software (Dotmatics). Significant differences between groups were
evaluated using one-way analysis of variance followed by Tukey's
multiple comparisons test or using unpaired Student's t-test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
TRIM46 expression is associated with
DDP resistance in NSCLC tissues
The function of TRIM46 was investigated by first
determining whether TRIM46 expression was associated with DDP
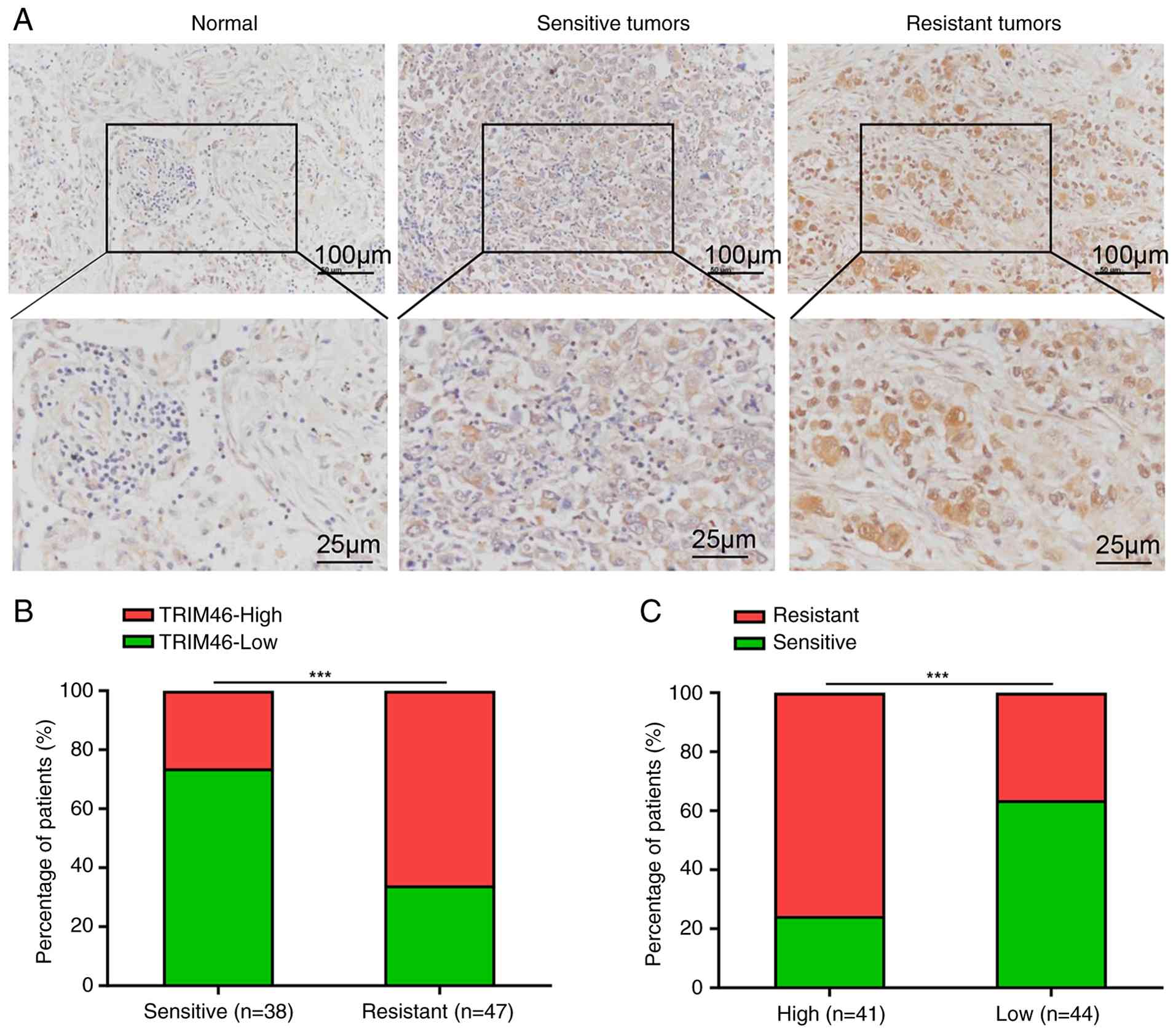
resistance in human samples. The immunohistochemical staining
results of the TMA showed that the TRIM46 level was elevated in
NSCLC tissues compared with normal tissues. Moreover, TRIM46
expression further increased in DDP-resistant tissues compared with
DDP-sensitive tissues (Fig. 1A and
B). The association between TRIM46 and DDP resistance was
explored by dividing the patients into two groups based on low or
high TRIM46 expression. It was found that the number of resistant
tissues significantly increased in the high-TRIM46-expression group
compared with the low-TRIM46-expression group (Fig. 1C). These results indicated that
TRIM46 expression may be associated with DDP resistance in NSCLC
tissues.
TRIM46 upregulation contributes to DDP
resistance in NSCLC cells
TRIM46 expression in bronchial epithelial cells
(16HBE) and DDP-sensitive (A549) and DDP-resistant (A549/DDP) NSCLC
cells was measured to explore the involvement of TRIM46 in DDP
resistance. As shown in Fig. 2A,
the TRIM46 level was significantly elevated in NSCLC cells compared
with 16HBE cells. Furthermore, TRIM46 expression was significantly
upregulated in A549/DDP cells compared with A549 cells, indicating
the possible involvement of TRIM46 in DDP resistance. A549 cells
were infected with blank lentivirus or TRIM46-overexpressing
lentivirus to directly determine the effect of TRIM46 on DDP
resistance. The results revealed that TRIM46 expression was
significantly upregulated at both the mRNA and protein levels upon
infection with TRIM46-overexpressing lentivirus (Fig. 2B). As shown in Fig. 2C and D, DDP induced the apoptosis of
A549 cells in a concentration-dependent manner. However, TRIM46
overexpression significantly suppressed the apoptosis induced by
DDP in A549 cells. Furthermore, the comet assay was employed to
investigate whether TRIM46 was related to DNA damage. The formation
of a ‘comet’ in the DNA of cells can be visualized using
single-cell gel electrophoresis and indicates DNA strand breaks as
the damaged DNA migrates at a different rate than the non-damaged
DNA. The results demonstrated an increase in the percentage of tail
DNA in DDP-treated A549 cells compared with the control group.
However, TRIM46 overexpression showed a decrease in the percentage
of tail DNA in A549 cells treated with DDP (Fig. 2E and F). These results indicated
that TRIM46 alleviated DNA damage and the cell apoptosis of A549
cells treated with DDP.
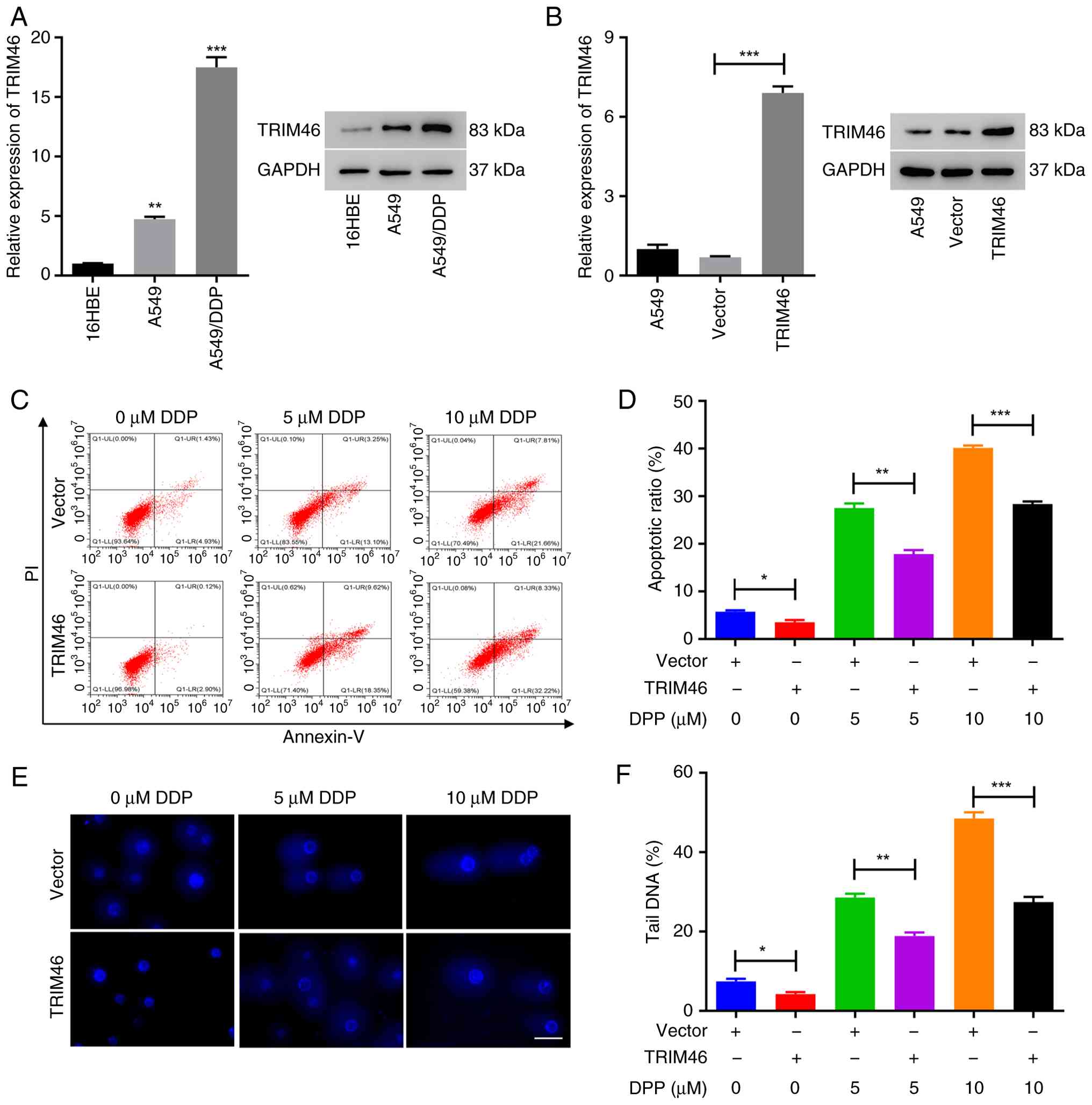 | Figure 2.TRIM46 upregulation contributes to
DDP resistance in non-small cell lung cancer cells. (A) RT-qPCR
(left) and western blot (right) analysis of TRIM46 expression in
16HBE, A549 and A549/DDP cells. (B) RT-qPCR (left) and western blot
(right) analysis of TRIM46 expression in A549 cells infected with
TRIM46 overexpression lentivirus. (C) Cell apoptosis was measured
after transduction with TRIM46-overexpression lentivirus and/or DDP
treatment (0, 5 and 10 µM) in A549 cells. (D) Quantitative analysis
of cell apoptosis. (E) Representative images of the comet assay
showing DNA fragmentation after transduction with
TRIM46-overexpression lentivirus and/or DDP treatment (0, 5 and 10
µM) in A549 cells (scale bar, 50 µm; magnification, ×400). (F)
Quantification of tail DNA (%) revealing DNA damage after
transduction with TRIM46-overexpression lentivirus and/or DDP
treatment (0, 5 and 10 µM) in A549 cells. *P<0.05, **P<0.01,
***P<0.001. TRIM46, tripartite motif 46; DDP, cisplatin;
RT-qPCR, reverse transcription-quantitative polymerase chain
reaction; PI, propidium iodide. |
TRIM46 knockdown alleviates DDP
resistance in NSCLC cells
The function of TRIM46 was further explored using
lentiviruses to knock down its expression in A549/DDP cells. As
expected, all three lentiviruses significantly decreased TRIM46
expression, and the knockdown efficiency of shTRIM46-2 was the most
significant (Fig. 3A). Knockdown of
TRIM46 using shTRIM46-2 lentivirus promoted the apoptosis of
A549/DDP cells treated with or without DDP (Fig. 3B and C). Meanwhile, the comet assay
results also showed that TRIM46 knockdown significantly enhanced
DNA damage in A549/DDP cells treated with DDP (Fig. 3D and E).
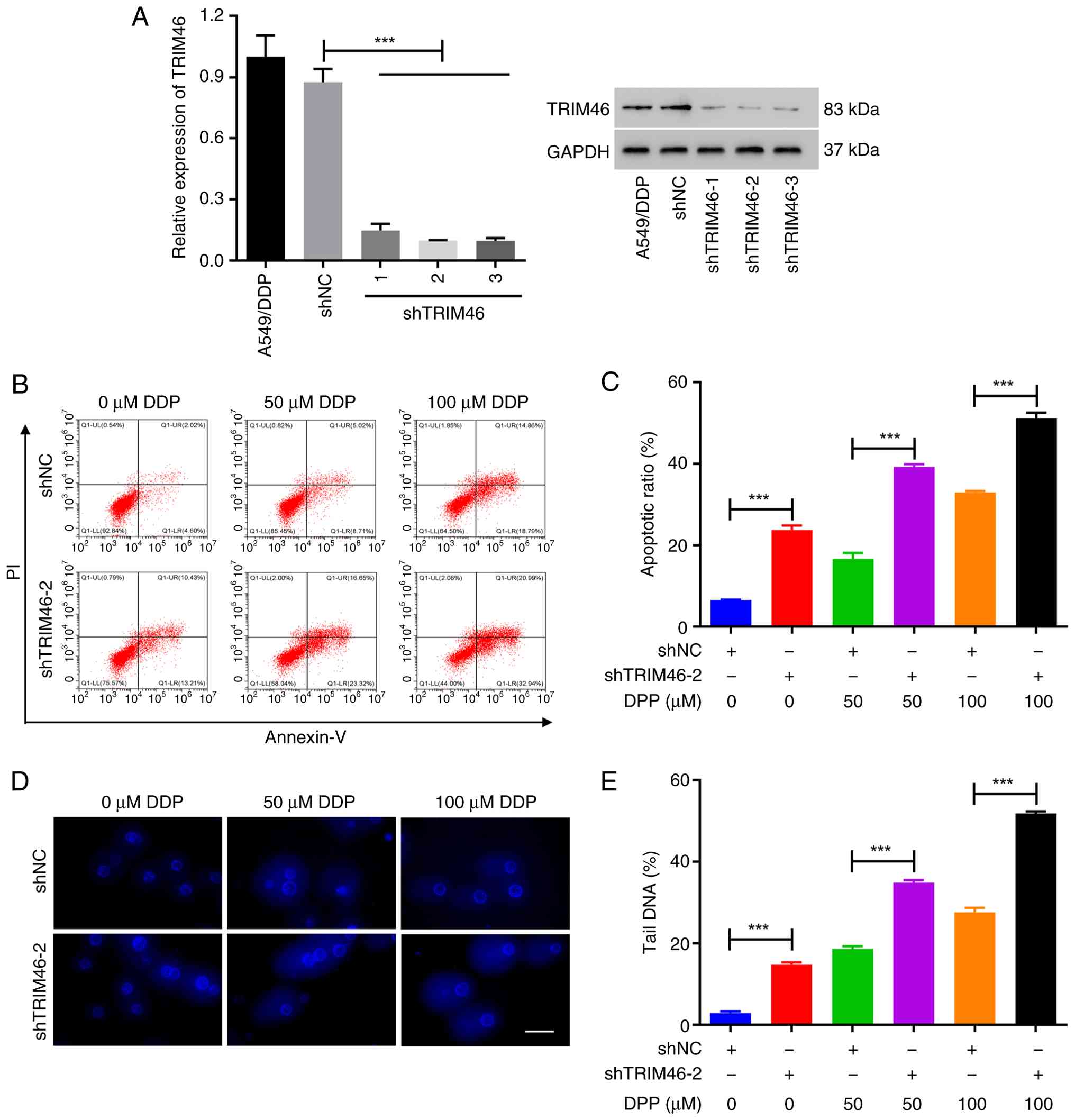 | Figure 3.TRIM46 knockdown alleviates DDP
resistance in non-small cell lung cancer cells. (A) Reverse
transcription-quantitative polymerase chain reaction (left) and
western blot (right) analysis of TRIM46 expression in A549/DPP
cells infected with TRIM46 knockdown lentiviruses. (B) Cell
apoptosis was measured after transduction with TRIM46 knockdown
lentivirus and/or DDP treatment (0, 50 and 100 µM) in A549/DDP
cells. (C) Quantitative analysis of cell apoptosis. (D)
Representative images of the comet assay showing DNA fragmentation
after transduction with TRIM46 knockdown lentivirus and/or DDP
treatment (0, 50 and 100 µM) in A549/DDP cells (scale bar, 50 µm;
magnification, ×400). (E) Quantification of tail DNA (%) revealing
DNA damage after transduction with TRIM46 knockdown lentivirus
and/or DDP treatment (0, 50 and 100 µM) in A549/DDP cells.
***P<0.001. TRIM46, tripartite motif 46; DDP, cisplatin; sh,
short hairpin; NC, negative control; PI, propidium iodide. |
TRIM46 depletion reduces cell
proliferation by regulating the Akt signaling pathway
Cell proliferation was detected in the A549/DDP
cells transduced with shTRIM46-2 and shTRIM46-3. The results showed
that TRIM46 depletion effectively reduced the proliferation of
A549/DDP cells (Fig. 4A). The
protein levels of p-Akt (Ser 473) were measured to elucidate the
role of TRIM46. It was found that TRIM46 knockdown resulted in a
significant decrease in the level of p-Akt (Fig. 4B). Based on the aforementioned
result depicted in Fig. 3, where
TRIM46 depletion induced DNA damage, the expression of RAD51, a key
protein involved in DNA repair, was further detected. As shown in
Fig. 4B, TRIM46 knockdown not only
decreased the protein level of RAD51 but also increased the protein
levels of caspase 3 and cleaved-caspase 3 in A549/DDP cells. These
findings collectively indicated that TRIM46 knockdown induced DNA
damage, thereby suppressing cell proliferation. The Akt pathway is
involved in regulating cell behaviors such as proliferation and
apoptosis; it also plays an important role in DNA damage repair
(16,17). To examine the role of the Akt
pathway in regulating TRIM46-mediated DDP resistance, LY294002, an
inhibitor of Akt, was applied to A549 cells infected with
TRIM46-overexpression lentivirus. The CCK-8 assay results showed
that TRIM46 overexpression significantly enhanced the proliferation
of A549 cells compared with the vector-transfected group. Moreover,
the promotive effect of cell proliferation induced by TRIM46
overexpression was alleviated by LY294002 (Fig. 4C). As shown in Fig. 4D, LY294002 treatment decreased the
p-Akt level and also partially blocked the increase in p-Akt level
caused by TRIM46 overexpression. Moreover, LY294002 treatment
effectively alleviated the increase in RAD51 level and the decrease
in the levels of caspase 3 and cleaved-caspase3 induced by TRIM46
overexpression in A549 cells. Taken together, these findings
emphasized that TRIM46 knockdown may induce DNA damage by
regulating the Akt signaling pathway, thereby inhibiting the
proliferation of NSCLC cells.
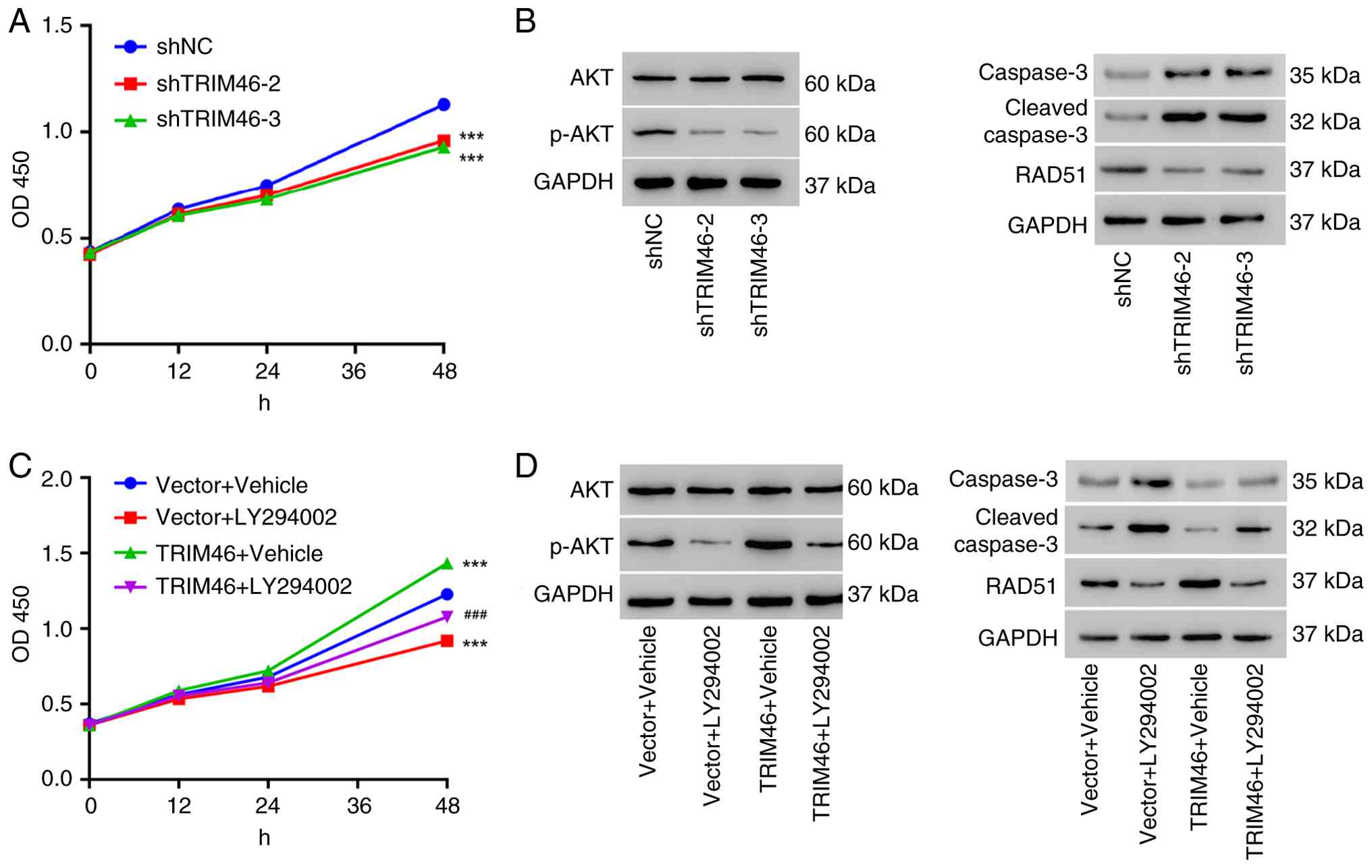 | Figure 4.TRIM46 depletion inhibits cell
proliferation by regulating the Akt signaling pathway. (A) Cell
proliferation of A549/DDP cells transduced with TRIM46-knockdown
lentiviruses. (B) Western blot analysis of p-Akt, Akt, caspase 3,
cleaved-caspase 3 and RAD51 expression in A549/DDP cells transduced
with TRIM46-knockdown lentiviruses. (C) Cell proliferation of A549
cells transduced with TRIM46-overexpression lentivirus and/or
treated with LY294002 (20 µM) or vehicle. (D) Western blot analysis
of p-Akt, Akt, caspase 3, cleaved-caspase 3 and RAD51 in A549 cells
transduced with TRIM46-overexpression lentivirus and/or treated
with LY294002 (20 µM) or vehicle. ***P<0.001 vs. shNC or Vector
+ Vehicle; ###P<0.001 vs. TRIM46 + Vehicle. TRIM46,
tripartite motif 46; DDP, cisplatin; sh, short hairpin; NC,
negative control; OD, optical density; p-, phosphorylated. |
TRIM46 knockdown increases the
sensitivity of xenograft tumors to DDP treatment
A549/DDP cells transduced with shTRIM46 or shNC
lentiviruses were subcutaneously injected into nude mice to
validate the function of TRIM46 in vivo. The mice were
treated with DDP after 12 days of injection. As illustrated in
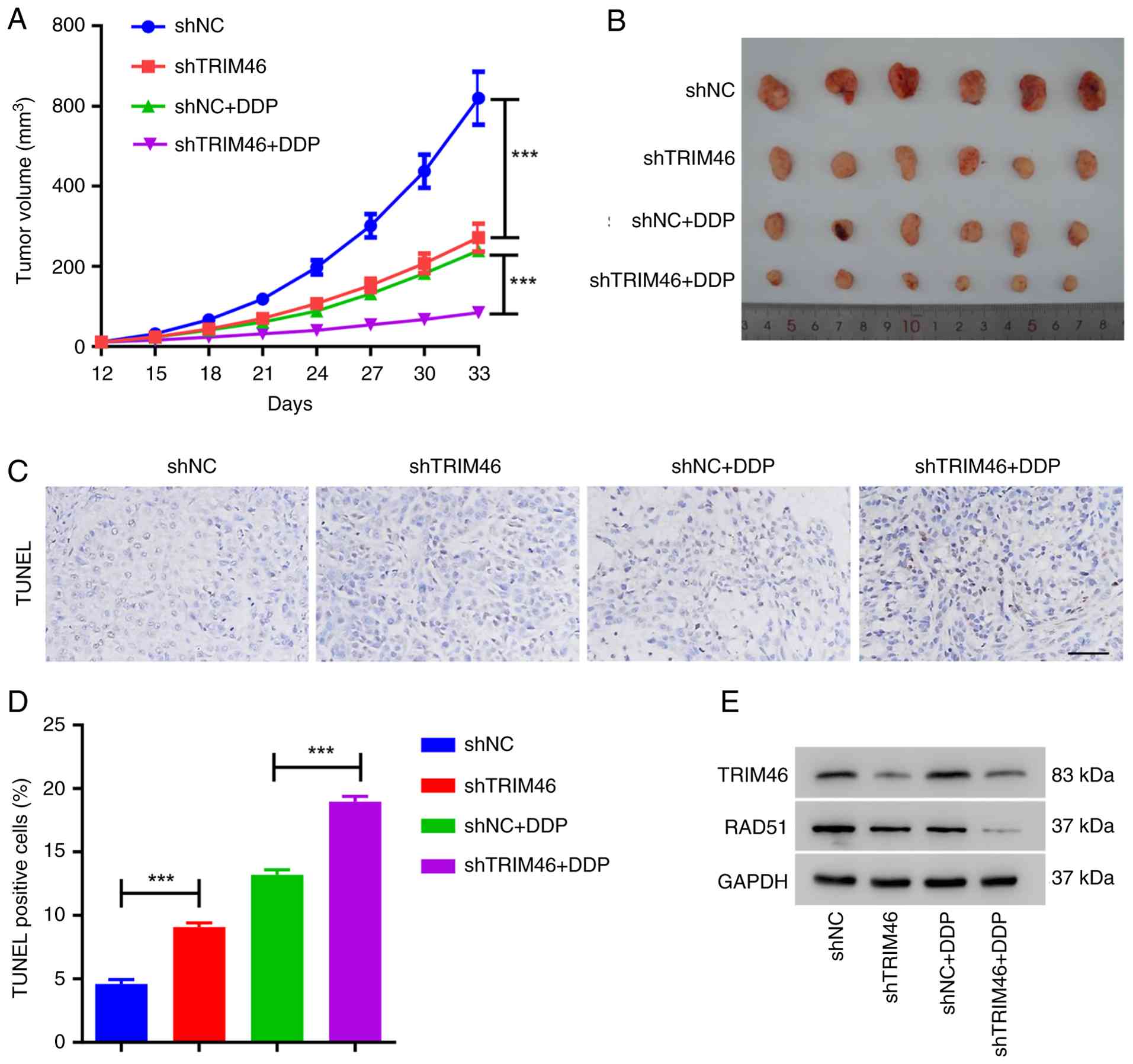
Fig. 5A and B, mice receiving
shTRIM46 cell injections, as well as those injected with shNC cells
and treated with DDP, exhibited smaller tumor volumes compared with
the shNC group. The lowest tumor volumes were observed in the mice
co-treated with shTRIM46 and DDP. The TUNEL assay revealed a higher
number of apoptotic cells in the shTRIM46 group compared with the
shNC group. Furthermore, the group co-treated with shTRIM46 and DDP
exhibited more apoptotic cells than the group co-treated with shNC
and DDP (Fig. 5C and D), suggesting
that TRIM46 depletion enhanced DDP-induced apoptosis. Additionally,
the group co-treated with shTRIM46 and DDP displayed the lowest
levels of RAD51 (Fig. 5E). These
findings collectively indicated that TRIM46 knockdown enhanced the
sensitivity of xenograft tumors to DDP treatment in
vivo.
Discussion
DDP was the first-in-class drug for NSCLC therapy;
it activates various cytoplasmic substrates and nuclear DNA and
leads to DNA damage, thus inducing tumor cell death by activating
the apoptotic signaling pathway (27–29).
Despite the efficacy of initial treatment, patients inevitably
become resistant to DDP. DDR is one of the primary mechanisms
underlying DDP resistance against NSCLC, although it is not fully
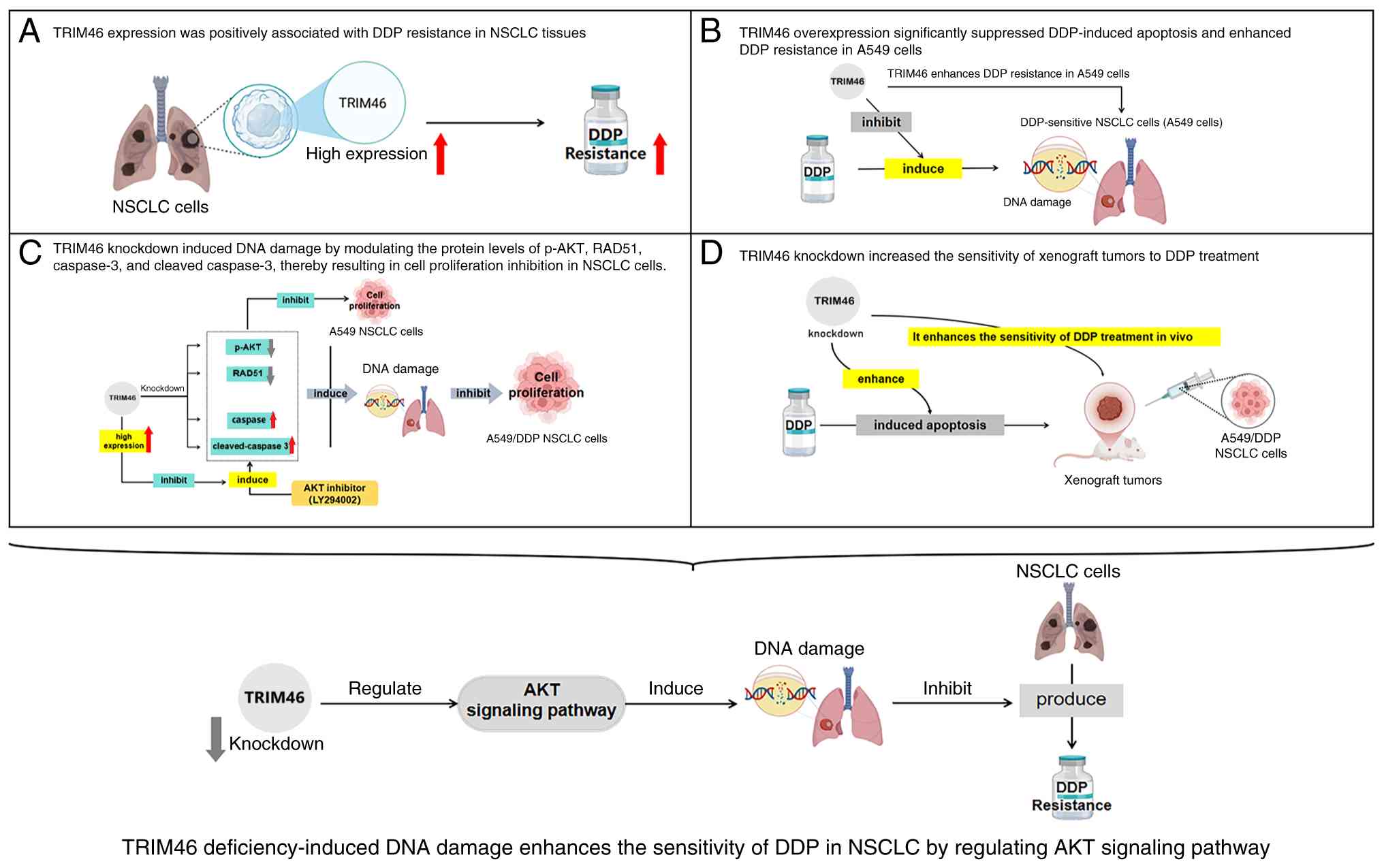
understood (27). The present study
showed that TRIM46 contributed to DDP resistance in NSCLC via
activating Akt signaling and reducing DNA damage. Additionally,
TRIM46 deficiency restored the function of DDP against NSCLC tumors
(Fig. 6).
The results of the present study revealed that
TRIM46 was highly expressed in the tissues of DDP-resistant NSCLC
tumors. Additionally, the level of TRIM46 was associated with DDP
resistance. A number of TRIM family members are involved in the
development of lung cancer. Accumulating evidence suggests that
TRIM proteins, including TRIM8, TRIM13, TRIM67, TRIM28, TRIM19 and
TRIM45, regulate the abundance and activity of p53, which is a
critical cancer suppressor (30).
Moreover, a study has suggested that TRIM proteins activate the
NF-κB pathway in all types of lung cancer (18). Over the years, TRIM proteins have
been found to control the PI3K/Akt signaling pathway. TRIM10,
TRIM11, TRIM26 and TRIM37 inhibit Akt signaling activation
(31). By contrast, TRIM22, TRIM24,
TRIM25, TRIM27, TRIM59 and TRIM44 promote Akt pathway activation
(31). These results collectively
suggested the complicated roles of TRIM proteins in lung cancer.
Exploring the role of other TRIM members can help understand the
development of lung cancer. The present study found that patients
with a lower level of TRIM46 were sensitive to DDP. Vice versa,
patients with a higher level of TRIM46 were resistant to DDP
treatment. In line with these findings, previous studies have
reported that TRIM46 expression is increased in NSCLC tissues
compared with normal lung tissues and its upregulation is
associated with NSCLC cancer progression and chemoresistance by
regulating the Akt or NF-κB signaling pathways (23,32,33).
Moreover, TRIM46 expression was also shown to be increased in the
tissues and cell lines of pancreatic cancer, ovarian cancer and
osteosarcoma as well as found to promote cancer progression by
regulating the Wnt or NF-κB signaling pathways (15,34,35).
However, a previous study showed downregulation of TRIM46 in NSCLC
tumor cells compared with normal tissues (36). This apparent paradox suggests a
context-dependent role of TRIM46 in NSCLC pathogenesis; its
expression and function may be differentially regulated during
tumorigenesis vs. acquired therapy resistance. Cancer cells could
selectively upregulate TRIM46 as an adaptive survival mechanism
under chemotherapeutic stress such as DDP treatment, leveraging its
ability to activate pro-survival Akt signaling and enhance DNA
repair, even if its basal expression differs. These results imply
distinct roles of TRIM46 in different conditions such as therapy
response status.
The present study found that DDP resistance in NSCLC
may be associated with TRIM46-mediated Akt signaling regulation. At
present, the PI3K/Akt pathway has been found to phosphorylate
multiple proteins, thereby stimulating tumor cell growth,
suppressing apoptosis and facilitating DNA damage repair after
chemotherapy (37). The
PI3K/Akt/NF-κB (17) and
PI3K/Akt/mTOR (16) pathways have
been found to be related to DDP treatment failure. Moreover,
PI3K/Akt participates in the induction of the ATP-binding cassette
transporter, as well as the regulation of aerobic glycolysis to
increase the energy supply (38,39).
Among the varied reported Akt-related TRIM proteins, TRIM18 has
been reported to negatively regulate protein phosphatase 2
phosphatase activator, resulting in Akt activation (40). Accordingly, another C-1 subfamily
member, TRIM46, also resulted in Akt activation in lung cancer
(23) and ovarian cancer cells
(41). In agreement with these
findings, the present study observed that the knockdown of TRIM46
reduced p-Akt levels and that TRIM46 overexpression activated Akt
signaling. These results at least partially indicated that TRIM46
may contribute to DDP resistance by activating Akt signaling. These
results also rendered a rationale to combine TRIM46 knockdown and
an Akt pathway inhibitor to treat DDP-resistant NSCLC.
A previous study has shown that circular RNA VMP1
depletion counteracts DDP resistance, thus attenuating NSCLC tumor
growth in vivo (42).
Additionally, TRIM59, a family member of TRIM46, has been reported
to enhance DDP resistance in NSCLC tumor cell lines. Blocking the
expression of TRIM59 also significantly suppressed tumor growth
(43). Furthermore, targeting DDP
resistance by TRIM65 deficiency in A549/DDP cells was shown to
effectively decrease tumor volume (44). These studies demonstrated that
targeting DDP resistance is an important therapeutic strategy
against NSCLC. In the present study, it was found that TRIM46
expression was positively associated with DDP-resistance in NSCLC
tumor cells. Additionally, TRIM46 knockdown in A549/DDP cells
significantly attenuated tumor growth in vivo, indicating
TRIM46 as a candidate therapeutic target of DDP-resistant NSCLC.
However, more cell lines are needed to validate the function of
TRIM46. Moreover, further investigations are necessary to explore
the functions and mechanisms of TRIM46 in DDP resistance in
vitro and in vivo.
The present study revealed that TRIM46 could act as
a positive regulator of DDP resistance in NSCLC via activating Akt
and reducing DNA damage. Targeting TRIM46, potentially through
small-molecule inhibitors or RNA interference-based strategies
(such as small interfering RNA or shRNA delivery via nanocarriers),
could sensitize resistant tumors to conventional DDP chemotherapy.
Combining TRIM46 inhibition with Akt pathway inhibitors might
represent a particularly effective synergistic strategy, as
suggested by the results using LY294002. However, this warrants
further investigation.
The present study had several limitations. First,
the investigation was primarily conducted using the A549 cell line
model. Validating the role of TRIM46 in additional NSCLC cell lines
with inherent or acquired DDP resistance can strengthen the
generalizability of the conclusions. Second, it was established
that TRIM46 activated Akt signaling. However, the precise molecular
mechanism by which TRIM46 (potentially through its E3 ubiquitin
ligase activity) regulates Akt activation warrants further
elucidation. Future studies should focus on identifying the
specific substrates of TRIM46 involved in this pathway.
Acknowledgements
Not applicable.
Funding
This study was supported by the Joint TCM Science &
Technology Projects of National Demonstration Zones for
Comprehensive TCM Reform (grant no. GZY-KJS-ZJ-2025-093) and the
Zhejiang Chinese Medical University special research projects of
affiliated hospitals (grant no. 2022FSYYZZ29).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
SJ and DZ performed the experiments, collected data
and drafted the manuscript. ZL, LY and JZ performed the statistical
analysis and participated in the study design. YW, PJ, MP and YL
participated in the analysis or interpretation of data and drafted
the manuscript. All authors read and approved the final version of
the manuscript. SJ, DZ and JZ confirm the authenticity of all the
raw data.
Ethics approval and consent to
participate
Animal experiments were conducted in accordance with
the Guide for the Care and Use of Laboratory Animals and approved
by the Animal Care and Use Committee of Wenzhou Traditional Chinese
Medicine Hospital of Zhejiang Chinese Medical University (Wenzhou,
China; approval no. WTCM-KT-2020002). This study used commercially
available tissue microarrays with samples obtained from fully
anonymized and consented donors and involved only retrospective
analysis. The institutional ethics committee reviewed the protocol
and granted an exemption from ethics approval.
Patient consent for publication
Not applicable.
Competing interests
All authors declare that they have no competing
interests.
References
|
1
|
Thai AA, Solomon BJ, Sequist LV, Gainor JF
and Heist RS: Lung cancer. Lancet. 398:535–554. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Torre LA, Siegel RL and Jemal A: Lung
cancer statistics. Adv Exp Med Biol. 893:1–19. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Suay G, Garcia-Cañaveras JC, Aparisi F,
Garcia J, Juan-Vidal O and Lahoz A: Immune checkpoint inhibitors as
first-line treatment for brain metastases in stage IV NSCLC
patients without driver mutations. Cancer Lett. 606:2173172024.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhou Y, Zhai W, Sun W, Han Y, Lin Z, Liu
D, Zheng Y, Luo X, Zhao Z, Feng S, et al: Safety and necessity of
omitting mediastinal lymph node dissection in cN0/N1 non-small cell
lung cancer after neoadjuvant immunotherapy. Front Immunol.
16:15876582025. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zhao L, Li M, Shen C and Luo Y, Hou X, Qi
Y, Huang Z, Li W, Gao L, Wu M and Luo Y: Nano-assisted radiotherapy
strategies: New opportunities for treatment of non-small cell lung
cancer. Research (Wash D C). 7:04292024.PubMed/NCBI
|
|
6
|
Miller KD, Siegel RL, Lin CC, Mariotto AB,
Kramer JL, Rowland JH, Stein KD, Alteri R and Jemal A: Cancer
treatment and survivorship statistics, 2016. CA Cancer J Clin.
66:271–289. 2016.PubMed/NCBI
|
|
7
|
Ashrafizadeh M, Zarrabi A, Hushmandi K,
Hashemi F, Moghadam ER, Owrang M, Hashemi F, Makvandi P, Goharrizi
MASB, Najafi M and Khan H: Lung cancer cells and their
sensitivity/resistance to cisplatin chemotherapy: Role of microRNAs
and upstream mediators. Cell Signal. 78:1098712021. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Du J, Wang X, Li Y, Ren X, Zhou Y, Hu W,
Zhou C, Jing Q, Yang C, Wang L, et al: DHA exhibits synergistic
therapeutic efficacy with cisplatin to induce ferroptosis in
pancreatic ductal adenocarcinoma via modulation of iron metabolism.
Cell Death Dis. 12:7052021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Srinivas US, Tan BWQ, Vellayappan BA and
Jeyasekharan AD: ROS and the DNA damage response in cancer. Redox
Biol. 25:1010842019. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Carusillo A and Mussolino C: DNA damage:
From threat to treatment. Cells. 9:16652020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Li L, Kumar AK, Hu Z and Guo Z: Small
molecule inhibitors targeting key proteins in the DNA damage
response for cancer therapy. Curr Med Chem. 28:963–985. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Hu XY, Hou PF, Li TT, Quan HY, Li ML, Lin
T, Liu JJ, Bai J and Zheng JN: The roles of Wnt/β-catenin signaling
pathway related lncRNAs in cancer. Int J Biol Sci. 14:2003–2011.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Xin R, Hu B, Qu D and Chen D: Oncogenic
lncRNA MALAT-1 recruits E2F1 to upregulate RAD51 expression and
thus promotes cell autophagy and tumor growth in non-small cell
lung cancer. Pulm Pharmacol Ther. 20:1021992023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ma J, Setton J, Lee NY, Riaz N and Powell
SN: The therapeutic significance of mutational signatures from DNA
repair deficiency in cancer. Nat Commun. 9:32922018. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Roos WP, Thomas AD and Kaina B: DNA damage
and the balance between survival and death in cancer biology. Nat
Rev Cancer. 16:20–33. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gong T, Cui L, Wang H, Wang H and Han N:
Knockdown of KLF5 suppresses hypoxia-induced resistance to
cisplatin in NSCLC cells by regulating HIF-1alpha-dependent
glycolysis through inactivation of the PI3K/Akt/mTOR pathway. J
Transl Med. 16:1642018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Xu S, Yang Z, Jin P, Yang X, Li X, Wei X,
Wang Y, Long S, Zhang T, Chen G, et al: Metformin suppresses tumor
progression by inactivating stromal fibroblasts in ovarian cancer.
Mol Cancer Ther. 17:1291–1302. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Huang N, Sun X, Li P, Liu X, Zhang X, Chen
Q and Xin H: TRIM family contribute to tumorigenesis, cancer
development, and drug resistance. Exp Hematol Oncol. 11:752022.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Liang M, Wang L, Sun Z, Chen X, Wang H,
Qin L, Zhao W and Geng B: E3 ligase TRIM15 facilitates non-small
cell lung cancer progression through mediating Keap1-Nrf2 signaling
pathway. Cell Commun Signal. 20:622022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Groner AC, Cato L, de Tribolet-Hardy J,
Bernasocchi T, Janouskova H, Melchers D, Houtman R, Cato ACB,
Tschopp P, Gu L, et al: TRIM24 is an oncogenic transcriptional
activator in prostate cancer. Cancer Cell. 29:846–858. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang L, Li X, Dong W, Sun C, Guo D and
Zhang L: Mmu-miR-1894-3p inhibits cell proliferation and migration
of breast cancer cells by targeting Trim46. Int J Mol Sci.
17:6092016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Jiang W, Cai X, Xu T, Liu K, Yang D, Fan
L, Li G and Yu X: Tripartite Motif-containing 46 promotes viability
and inhibits apoptosis of osteosarcoma cells by activating NF-B
signaling through ubiquitination of PPAR. Oncol Res. 28:409–421.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Tantai J, Pan X, Chen Y, Shen Y and Ji C:
TRIM46 activates AKT/HK2 signaling by modifying PHLPP2
ubiquitylation to promote glycolysis and chemoresistance of lung
cancer cells. Cell Death Dis. 13:2852022. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
He S, Ma X, Zheng N, Wang G, Wang M, Xia W
and Yu D: PRDM14 mediates chemosensitivity and glycolysis in
drug-resistant A549/cisplatin cells and their progenitor A549 human
lung adenocarcinoma cells. Mol Med Rep. 23:1492021. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Chen CC, Chen CY, Ueng SH, Hsueh C, Yeh
CT, Ho JY, Chou LF and Wang TH: Corylin increases the sensitivity
of hepatocellular carcinoma cells to chemotherapy through long
noncoding RNA RAD51-AS1-mediated inhibition of DNA repair. Cell
Death Dis. 9:5432018. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Chen SH and Chang JY: New insights into
mechanisms of cisplatin resistance: From tumor cell to
microenvironment. Int J Mol Sci. 20:41362019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Romani AMP: Cisplatin in cancer treatment.
Biochem Pharmacol. 206:1153232022. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sancho-Martinez SM, Prieto-Garcia L,
Prieto M, López-Novoa JM and López-Hernández FJ: Subcellular
targets of cisplatin cytotoxicity: An integrated view. Pharmacol
Ther. 136:35–55. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Liu J, Zhang C, Wang X, Hu W and Feng Z:
Tumor suppressor p53 cross-talks with TRIM family proteins. Genes
Dis. 8:463–474. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhan W and Zhang S: TRIM proteins in lung
cancer: Mechanisms, biomarkers and therapeutic targets. Life Sci.
268:1189852021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Pan L, Zhang D, Shao Q, Cheng M, Liao Z,
Yu L, Wang Y, Jia P and Zhang J: Panax notoginseng improves the
sensitivity of non-small cell lung cancer to cisplatin by
inhibiting Akt signaling. Cancer Biomark. 42:187585922413033772025.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhong Y, Luo B, Hong M, Hu S, Zou D, Yang
Y, Wei S, Faruque MO, Dong S, Zhu X, et al: Oxymatrine induces
apoptosis in non-small cell lung cancer cells by downregulating
TRIM46. Toxicon. 244:1077732024. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu S, Xiao C, Rong Y, Liu M, Yang K, Tang
J and Wang Z: Comprehensive analysis of ferroptosis-related genes
indicates that TRIM46 is a novel biomarker and promotes the
progression of ovarian cancer via modulating ferroptosis and Wnt
signaling pathway. Am J Cancer Res. 14:4686–4707. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Xu L, Wan X, Shan X, Zha W, Shi Y and Fan
R: ONECUT3 activates the TRIM46-NF-κB pathway to promote the
development of pancreatic cancer. Biochem Biophys Res Commun.
759:1517052025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhan W, Han T, Zhang C, Xie C, Gan M, Deng
K, Fu M and Wang JB: TRIM59 promotes the proliferation and
migration of non-small cell lung cancer cells by upregulating cell
cycle related proteins. PLoS One. 10:e01425962015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Liu R, Chen Y, Liu G, Li C, Song Y, Cao Z,
Li W, Hu J, Lu C and Liu Y: PI3K/AKT pathway as a key link
modulates the multidrug resistance of cancers. Cell Death Dis.
11:7972020. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Lee JH, Liu R, Li J, Wang Y, Tan L, Li XJ,
Qian X, Zhang C, Xia Y, Xu D, et al: EGFR-phosphorylated platelet
isoform of phosphofructokinase 1 promotes PI3K activation. Mol
Cell. 70:197–210.e7. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Ke B, Wei T, Huang Y, Gong Y, Wu G, Liu J,
Chen X and Shi L: Interleukin-7 Resensitizes Non-small-cell lung
cancer to cisplatin via inhibition of ABCG2. Mediators Inflamm.
2019:72414182019. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Zhang L, Li J, Lv X, Guo T, Li W and Zhang
J: MID1-PP2A complex functions as new insights in human lung
adenocarcinoma. J Cancer Res Clin Oncol. 144:855–864. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Peng L, Liu L, Xu L, Chen J, Wang X, Zhang
M and Ge Y: TRIM46 promotes chemoresistance of ovarian cancer via
activating PHLPP2/PI3K/AKT pathway. Biochem Cell Biol. 103:1–9.
2025. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Xie H, Yao J, Wang Y and Ni B:
Exosome-transmitted circVMP1 facilitates the progression and
cisplatin resistance of non-small cell lung cancer by targeting
miR-524-5p-METTL3/SOX2 axis. Drug Deliv. 29:1257–1271. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
He R and Liu H: TRIM59 knockdown blocks
cisplatin resistance in A549/DDP cells through regulating
PTEN/AKT/HK2. Gene. 747:1445532020. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Pan X, Chen Y, Shen Y and Tantai J:
Knockdown of TRIM65 inhibits autophagy and cisplatin resistance in
A549/DDP cells by regulating miR-138-5p/ATG7. Cell Death Dis.
10:4292019. View Article : Google Scholar : PubMed/NCBI
|