Introduction
Covalent histone methylation of lysine residues
carries out a key role in regulating chromatin dynamics and
functions (1–3). Methylation of lysines on histone H3
and H4 activates or represses gene transcription, depending on the
position of the modified residues (1,4). In
general, methylated histone H3 lysine 4 (H3K4) is associated with
active or balanced gene states, whereas methylated H3K9 and H3K27
are gene repressive (5). The
removal of methyl groups from lysine residues on histones is
regulated by histone lysine demethylases (KDM) (6,7).
To date, two distinct families of demethylases have
been described the flavin-dependent KDM1 and the JmjC
domain-containing KDM2-8 subfamilies. The first family, KDM1,
catalyzes the demethylation of mono- and di-methylated lysine
residues (Kme1 and Kme2). The second family of KDMs (KDM2-8) is
capable of demethylating Kme1, Kme2 and Kme3 (8). In addition, KDMs demethylate
non-histone substrates and also have several
demethylase-independent functions (9). The KDM expression profile varies in
different cells and tissues. Altered expression of KDMs, especially
those targeting H3K4 and H3K27, is common in several types of human
cancer (9). The KDM5 subfamily,
that has 4 members (KDM5A-D), is capable of removing tri- and di-
methyl marks from H3K4 (6). This
subfamily is deregulated in several types of cancer and can
modulate chemoresistance by numerous mechanisms including
autophagy, epithelial-mesenchymal transition (EMT), stemness,
metabolism and DNA repair (6,9). The
role of KDM5s in the development of chemoresistance has been
described in various types of cancer (10) KDM5A (11–14);
KDM5B (15–21) and KDM5C (22–24).
The KDM5D gene (also known as JARID1D
or SMCY) is located on the Y chromosome and it is the only
male-specific KDM5. It is expressed in all male tissues, and
carries out a role in spermatogenesis (25,26).
KDM5D has been described as an important tumor-suppressor in
castration-resistant prostate cancer and its low expression is
associated with a worse prognosis (27). In prostate cancer, KDM5D regulates
matrix metalloproteinase family genes associated with invasion, and
loss of KDM5D with increased H3K4me3 levels in promoter regions of
relevant genes increases invasiveness and metastatic ability
(28,29). Moreover, in clear cell renal cell
carcinoma (ccRCC), KDM5D is downregulated by loss of the Y
chromosome, which contributes to the pathogenesis of ccRCC
(30). In papillary renal cell
carcinoma, KDM5D also facilitates demethylation of CDK4 and
promotes proliferation of cancer cells (31). Chen et al (32) described the
function of KDM5D in the development of CDDP tolerance in head and
neck squamous cell carcinoma (33).
Decreased KDM5D expression was also observed in gastric, colorectal
and hepatocellular carcinomas (34–36).
Moreover, in an analysis of the Cancer Genome Atlas database, Duan
et al (37) showed that
KDM5D was notably downregulated in 24 different cancer types (such
as breast, pancreatic and prostate cancer) compared with adjacent
tissues. To the best of our knowledge, little is known about the
importance of KDM5D in neuroblastoma. However, it has been reported
that low KDM5D expression is associated with a worse prognosis in
some tumors (38,39). We hypothesize that this is due to
the decrease of cell junctions and therefore increase the ability
to metastasize and due to the decrease of the presentation of
antigens and therefore the immune response to the tumor (38,39).
CUL4A is a protein of the cullin family that acts as
a scaffold for cullin RING ligase 4 complexes that promote
ubiquitination of various substrates. It carries out an important
role in DNA repair and replication, chromatin restructuring, cell
cycle regulation, embryogenesis, hematopoiesis and spermatogenesis
(40,41). There are a growing number of studies
associating overexpression or amplification of CUL4A to
increased growth, progression and metastasis in cancer (42–53).
The CUL4A gene is located on 13q34, an area prone to
amplification in some types of cancer (45–53).
Shen et al (34)
demonstrated that KDM5D carries out an important role in the
induction of EMT of gastric cancer cells through demethylation in
the promoter of CUL4A in male patients. In addition, another
study suggests a relationship between the CUL4A gene and the
sensitivity of colorectal cancer cells to CDDP (53).
Neuroblastoma is a malignancy of the sympathetic
nervous system and is the most common malignant extracranial tumor
of childhood. It is characterized by high degree of heterogeneity,
which may account for the wide range of clinical presentations and
variable response to treatment (54). The combination of clinical and
genetic factors allows stratification of patients into very low,
low, intermediate and high-risk groups (55). High-risk neuroblastoma is
characterized by the development of acquired chemoresistance
(56). To the best of our
knowledge, there is currently little information regarding the role
of KDM5D in neuroblastoma and the contribution of KDM5D to
CDDP-chemoresistance.
The present study aims to investigate the importance
of KDM5D expression for the proliferation of neuroblastoma cells
and their chemoresistance to CDDP. On the basis of the
aforementioned studies that demonstrated an association between
KDM5D and CUL4A and their contribution to chemoresistance to CDDP,
the present study investigated whether changes of CUL4A expression
could be mediated by KDM5D in neuroblastoma cell lines.
Materials and methods
Cell culture and chemicals
Human high-risk neuroblastoma cell lines UKF-NB-3
and the derived CDDP-resistant line UKF-NB-3CDDP were
provided by Prof. Jindrich Cinatl, Goethe University, Frankfurt am
Main, Germany. IMR-32 was purchased from MillporeSigma and SK-N-F1
by American Type Culture Collection. The CDDP-chemoresistant cell
lines IMR-32CDDP and SK-N-F1CDDP were derived
in our laboratory from their chemosensitive parental cell lines
(IMR-32 and SK-N-F1) after long-term cultivation with increasing
CDDP (Sandoz Group AG) concentration (57,58).
All tested cell lines were of male origin. Cells were cultured at
37°C and 5% CO2 in Iscove's Modified Dulbecco's Medium
(IMDM) supplemented with 10% (v/v) fetal bovine serum (both Thermo
Fisher Scientific, Inc.). KDOAM-25 citrate (400 nM; HY-102047B;
MedChemExpress) was used to inhibit KDM5s, where it was added to
cells and incubated for 48 h at 37°C.
KDM5 inhibition using KDOAM-25
For the inhibition of KDM5 demethylases, cells were
treated with KDOAM-25 citrate for 24–48 h at 37°C and 5%
CO2 (cat. no. HY-102047B; MedChemExpress) at a final
concentration of 400 nM. This concentration was selected based on
preliminary optimization experiments demonstrating efficient
reduction of H3K4 trimethylation without affecting cell viability
(data not shown). For supplementary validation experiments,
KDOAM-25 was added 24 h before CDDP treatment, whereas for all main
experiments presented in the manuscript, KDOAM-25 was added 48 h
before CDDP administration and maintained throughout the treatment
period.
Cell proliferation assay
To determine cell proliferation, cells were placed
in 24-well plates (1×105 cells per well) or 96-well
plates (1×104 cells per well) and seeded for 24 h at
37°C.and then cells were treated with CDDP at a final concentration
of 0.6-300 µM for 48 h at 37°C. Cells were then incubated with
PrestoBlue® Cell Viability Reagent (Thermo Fisher
Scientific, Inc.) according to the manufacturer's protocol.
Fluorescence was measured at an excitation wavelength of 560 nm and
an emission wavelength of 590 nm using the SpectraMax®
i3× Multi-Mode Microplate Reader (Molecular Devices, LLC). Each
sample was analyzed in triplicate. The optical density of the
medium was read as background, and the value of the optical density
of the control cells was taken as 100%. IC50 values were
calculated using SOFTmax® Pro 7.2 GxP software (Agilent
Technologies, Inc.).
Transfection
On-Targetplus small interfering (si)RNA (Revvity
Discovery Limited), a smart pool of three siRNAs [12.5 or 25 nM;
cat. no. L-007948-00-0005; National Center for Biotechnology
Information (NCBI) accession nos. NM_001146705.2, NM_001146706.2
and NM_004653.5] was used for silencing of KDM5D and On-Targetplus
Non-Targeting control siRNA (12.5 or 25 nM; cat. no.
D-001320-01-20), a smart pool of four siRNAs, was used as a
negative control (Revvity Discovery Limited). For initial
validation experiments, siRNA was tested at 12.5 and 25 nM. Based
on these results, a final concentration of 25 nM was used for all
subsequent functional experiments. siRNAs were transfected with
Dharmafect transfection reagent (cat. no. T-2001-03; Revvity
Discovery Limited) for 48 h according to the manufacturer's
protocol. Following 48 h of incubation at 37°C, cells were
harvested and used for further analysis.
To ectopically express KDM5D,
UKF-NB-3CDDP cells were transfected for 48 h with the
GenEZTM ORF clone plasmid KDM5D pCMV-3Tag1a (OHu18895C;
Genscript) designed for transcript variant 1 of KDM5D (8 and 16
ng/µl; NCBI accession number NM_001146705.1; GenScript Biotech
Corporation) and pCMV6-AC-GFP, mammalian vector with C-terminal
tGFP tag (20 nM; OriGene Technologies, Inc.) was used as a negative
control. For initial validation experiments, siRNA was tested at 8
and 16 ng/µl. Based on these results, a final concentration of 16
ng/µl was used for all subsequent functional experiments. The KDM5D
ORF clone plasmid or control pCMV6-AC-GFP was transfected using
Dharmafect transfection reagent (T-2001-03; Revvity Discovery
Limited). The transfected cells were selected by Gibco geneticin™
Selective Antibiotic (cat. no. G418 Sulfate; 50 mg/ml) (Thermo
Fisher Scientific, Inc.) in a final concentration 400 µg/ml for 72
h at 37°C. Overexpression efficiency was determined by reverse
transcription-quantitative polymerase chain reaction (RT-qPCR) and
flow cytometry respectively.
The exact siRNA/shRNA sequences used in the present
study are proprietary to Revvity Discovery Limited and are not
publicly available. They can be obtained for manufacturer upon
reasonable request (www.dharmacon.com).
RT-qPCR
mRNA from all NBL cell lines was isolated using the
PureLink™ RNA Mini Kit (Thermo Fisher Scientific, Inc.) according
to the manufacturer's protocol. The quality of the extracted mRNA
was measured using the NanoDrop One spectrophotometer (260/280 nm
ratio) (Thermo Fisher Scientific, Inc.). Complementary DNA was
synthesized from 1.0 µg of mRNA using the Generi Biotech Reverse
Transcription Kit according to the manufacturer's instructions
(GENERI BIOTECH). RT-qPCR was performed using the gb Easy PCR
Master Mix (cat. no. 3006; GENERI BIOTECH) according to the
manufacturer's instructions, with custom primers (Generi custom
oligo synthesis) and hydrolysis probes. The RT-qPCR assays are
identified by Generi Biotech IDs. These assays can be ordered at ID
(ID: ‘hKDM5B_Q1’ and ‘hKDM5D_Q1’) from GENERI BIOTECH. Expression
levels of target genes and the internal control POLR2A (ID, hPOLR2A
‘hPOLR2A_Q1’), which is homogeneously and uniformly expressed in
neuroblastoma cell lines (59),
were analyzed by RT-qPCR on a QuantStudio 3 Real-Time PCR System
(Thermo Fisher Scientific, Inc.). Each sample was analyzed in
triplicate. The thermocycling conditions were: 95°C 3 min, 50
cycles of 95°C for 10 sec, 60°C 20 sec. The relative differences in
gene expression were expressed as fold change and were obtained
with the 2−ΔΔCq method (Relative Expression Software
Tool REST 2009 software, v2.0.11; QIAGEN) (60). The exact primer sequences used in
the present study are proprietary to GENERI BIOTECH and are not
publicly available. They can be obtained for manufacturer upon
reasonable request (www.generi-biotech.com).
Cell cycle analysis
Cell cycle analysis was carried out using
FxCycleTM Violet Ready FlowTM reagent (Thermo
Fisher Scientific, Inc.). After treatment with CDDP and/or
transfection, harvested neuroblastoma cells with 0.25% trypsin
(Thermo Fisher Scientific, Inc.) were pelleted by centrifugation
(300 × g) for 3 min at room temperature, resuspended in 100 µl of
3.6% paraformaldehyde (Biogen) and incubated at 20°C for 15 min.
After this incubation, the suspension was then centrifuged (300 ×
g) for 3 min at room temperature, and the pellet was washed twice
with phosphate-buffered saline (PBS; Thermo Fisher Scientific,
Inc.). Permeabilization was performed with 90% methanol (PENTA) for
1 h at −20°C. Pellets were washed with PBS, resuspended in 500 µl
of PBS and a drop of FxCycleTM Violet Ready
FlowTM reagent was added. After 30 min of incubation at
room temperature, cell cycle analysis was performed using a BD
FACSCelesta (BD Biosciences) and data were analyzed using Flowlogic
software, version 8 (Inivai Technologies).
Determination of protein levels and
histone H3K4 methylation status by flow cytometry
After treatment with CDDP and/or transfection,
harvested neuroblastoma cells (UKF-NB-3; UKF-NB-3CDDP)
were washed with cold PBS (Thermo Fisher Scientific, Inc.),
trypsinized with 0.25% trypsin (Thermo Fisher Scientific, Inc.) and
collected by centrifugation (300 × g) for 3 min at room
temperature. Cell pellets were washed with PBS and fixed in 3.6%
paraformaldehyde for 15 min at room temperature. The cell pellets
were then washed with PBS and permeabilized with 90% methanol for 1
h at −20°C. The pellets were then washed three times with 0.5%
bovine serum albumin (BSA; Roth) in PBS. After washing, the cell
pellets were blocked with 5% BSA in PBS for 1 hour at room
temperature After blocking, the pellets were incubated with the
primary antibody anti-JARID1D rabbit mAB at a dilution of 1:100
(cat. no. PA5-100844; Invitrogen, Thermo Fisher Scientific, Inc.),
anti-JARID1B rabbit mAB at dilution of 1:400 (cat. no. 15327S; Cell
Signaling Technology, Inc.), anti-trimethyl histone H3 (Lys4)
rabbit (cat. no. 07-473; MilliporeSigma) at a dilution of 1:400,
Cleaved-Caspase 3 (Asp175) Rabbit mAB at a dilution of 1:50 (cat.
no. 9602S; Cell Signaling Technology, Inc.) or CUL4A Rabbit mAB at
a dilution of 1:50 (cat. no 2699; Cell Signaling Technology, Inc.)
for 1 h at laboratory temperature. Cell pellets were then washed
with 0.5% BSA (Roth) and incubated in fluorochrome-conjugated
secondary antibody Anti-Rabbit IgG (H+L) Alexa Fluor®
647 Conjugate (cat. no. A21245; Thermo Fisher Scientific, Inc.)
diluted 1:500 and incubated for 30 min at room temperature in the
dark. Cell pellets incubated with secondary antibody (1:500) only
were used as a control. Washed and resuspended cells were measured
using a BD FACSCelesta (BD Biosciences), and data were analyzed
using Flowlogic software, version 8 (Inivai Technologies).
Cell migration monitoring
Real-time monitoring of cell migration of sensitive
cells (UKF-NB-3) and their derived chemo-resistant cells
(UKF-NB-3CDDP) was carried out using the xCELLigence
RTCA DP instrument (Agilent Technologies, Inc.) in a humidified
incubator at 37°C and 5% CO2. For sensitive cells, KDM5D
silencing (siKDM5D) and for CDDP-resistant UKF-NB-3CDDP
ORD cDNA transfection were performed. Cells were serum-starved for
2 h in IMDM without FBS at 37°C and 5% CO2. After the
starvation period cells were trypsinized and seeded at a density of
1×104 cells/well into upper chamber of the 16-well
electronically integrated Boyden chamber for invasion/migration
assays-CIM-plate 16 (RTCA, Agilent) containing starvation media.
The wells of the lower chamber were loaded with IMDM with 10% FBS.
As the cells migrated towards the chemoattractant across
microelectronics sensors integrated at the bottom side of upper
chamber, the impedance was measured every 30 min for 168 h at 37°C.
The measurements were recorded and analyzed using Real-Time Cell
Analysis Software 1.2 (Agilent Technologies, Inc.).
Cell proliferation monitoring
Real-time monitoring of cell proliferation was
carried out using the xCELLigence RTCA DP instrument (Agilent
Technologies, Inc.) in a humidified incubator at 37°C with 5%
CO2. Sensitive cells and their derived chemoresistant
cells (UKF-NB-3, UKF-NB-3CDDP, IMR-32,
IMR-32CDDP, SK-N-F1 or SK-N-F1CDDP) were
seeded at a concentration 8,000 cells per well in wells of 16-well
E plates for impedance-based detection. For sensitive cells KDM5D
silencing (siKDM5D) was carried out. After 48 h of transfection
(siKDM5D), cells were seeded at a concentration 8,000 cells per
well in wells of 16-well E plates for impedance-based detection.
The cell index was monitored every 30 min for 96 h at 37°C and data
were recorded using the xCELLigence RTCA software Pro version 2.8
(Agilent Technologies Inc.) provided.
R2 genomics analysis and visualization
platform
The R2 Genomics Platform (https://r2.amc.nl) is an open-access online genomic
analysis and visualization platform that is publicly available to
analyze and interpret clinical and genomic data. Several
neuroblastoma datasets are available for survival analysis. The
present study used three different datasets: i) Tumor neuroblastoma
from Kocak (649; custom; ID: Agilent 44K microarray, Kocak dataset,
Wolf normalization-ag44kcwolf;), which contains gene expression
profiles from 649 patient-derived neuroblastoma tumors (https://hgserver1.amc.nl/cgi-bin/r2/main.cgi R2
internal identifier: ps_avgpres_gse45547geo649_ag44kcwolf)
(61); ii) tumor neuroblastoma from
SEQC (498; RPM; ID: SEQC/MAQC-III neuroblastoma dataset - seqcnb1),
which contains expression data from gene expression microarrays for
498 patient-derived neuroblastoma tumors
(https://hgserver1.amc.nl/cgi-bin/r2/main.cgi R2 internal
identifier: ps_avgpres_gse62564geo498_seqcnb1) (62) and iii) tumor neuroblastoma from
Oberthuer (251; user-defined; ID: Oberthuer neuroblastoma
gene-expression dataset - amexp255), which consists of gene
expression profiles from 251 patient-derived neuroblastoma tumors
(https://hgserver1.amc.nl/cgi-bin/r2/main.cgi R2 internal
identifier: ps_avgpres_nb251_amexp255) (63), (Table
SI). To analyze the prognostic significance of the KDM5D
and CUL4A genes Kaplan-Meier curves were generated comparing
overall survival of patients with low and high expression of KDM5D
or CUL4A using the integrated plotting function with default
settings on R2: Genomics Analysis and Visualization Platform
(Department of Oncogenomics, Amsterdam University Medical Centers
(AMC), University of Amsterdam; accessed in 2024). Median
expression cutoff mode and Bon-Ferroni correction for multiple
testing was performed for all analyses. Statistical significance of
survival differences was determined using the log-rank (Mantel-Cox)
test.
Western blot analysis
Proteins were extracted from cultured neuroblastoma
cells. Extraction of proteins was conducted using the ReadyPrep™
Protein Extraction Kit (Bio-Rad Laboratories, Inc.), with the
addition of a complete protease inhibitor cocktail (Roche Applied
Science). The concentration of proteins was subsequently measured
using a BCA Protein Assay kit (Thermo Fisher Scientific Inc.).
Samples (40 µg) were resolved on 4–20% precast gradient
SDS-polyacrylamide gels (Mini-PROTEAN® TGX™, Bio-Rad
Laboratories, Inc.), transferred onto PVDF membranes (Bio-Rad
Laboratories, Inc.) and blotted on PVDF membranes (Bio-Rad
Laboratories, Inc.). Membranes were blocked in 5% BSA in TBS
containing 0.1% Tween-20 for 1 h at room temperature, followed by
incubation with primary antibodies overnight at 4°C. The following
primary antibodies were used: KDM5D Rabbit pAb (cat. no.
PA5-100844; Thermo Fisher Scientific, Inc.), was diluted to a
concentration of 1:500, while CUL4A Rabbit pAb (cat. no. 2699S;
Cell Signaling Technology Inc.) was diluted to 1:1,000. The PARP
Rabbit pAb (cat. no. 9542S; Cell Signaling Technology Inc.) and the
Caspase-3 Rabbit pAb (cat. no. 9622S; Cell Signaling Technology
Inc.) were both diluted to 1:1,000. β-tubulin Mouse mAb (cat. no.
86298S, Cell Signaling Technology Inc.) was diluted to 1:1,000 and
used as a loading control. Secondary antibodies, AlexaFluor 488
Mouse, Rabbit (cat. nos. A-11008 and A28175; Thermo Fisher
Scientific Inc.) were diluted 1:2,000 and the incubation conditions
were 1 h at room temperature. The membranes were then subjected to
visualization via the ChemiDoc MP imaging system (Bio-Rad
Laboratories, Inc.). The analysis was conducted utilizing ImageJ
1.52a software (National Institutes of Health).
For the verification of sustained KDM5D silencing,
UKF-NB-3 cells were transfected with ON-TARGETplus KDM5D siRNA as
described above and lysed at multiple time points up to 7 days
after transfection. KDM5D protein levels were analyzed by western
blotting using the KDM5D antibody and β-tubulin as a loading
control.
Statistical analysis
All experiments were repeated independently at least
three times, and data are expressed as mean ± standard error of the
mean. ANOVA with post hoc Tukey honestly significant difference was
used to compare different groups. One-way ANOVA with Tukey's post
hoc test was used when a single independent variable (such as
treatment, transfection or time) was analyzed. Two-way ANOVA with
Tukey's post hoc test was applied when two independent variables
(such as cell line and treatment) were analyzed simultaneously.
Differences were considered statistically significant when. The
exact statistical tests used for each analysis are indicated in the
respective figure legends.
Results
Loss of KDM5D expression in CDDP
resistant neuroblastoma cell line
Chemotherapy is one of the main treatment modalities
for high-risk neuroblastoma, and CDDP is part of the majority of
treatment protocols. The occurrence of chemoresistance is a
considerable negative sign and contributes fundamentally to
treatment failure. In this context, cell lines with intrinsic
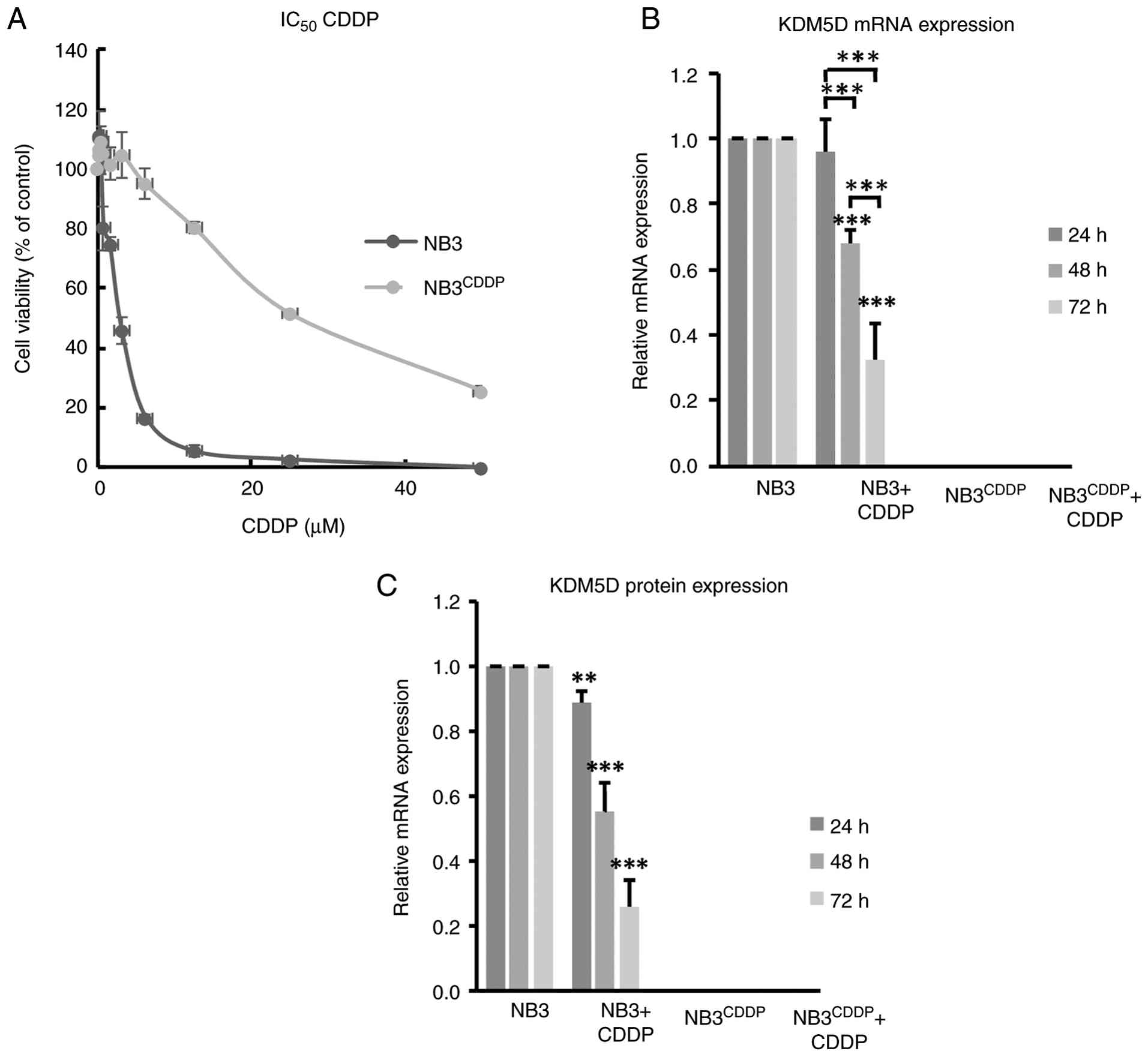
induced resistance to CDDP were analyzed. The IC50 of
the CDDP-resistant cell lines (UKF-NB-3CDDP,
IMR-32CDDP and SK-N-F1CDDP) was significantly
higher compared with the drug-sensitive parental lines UKF-NB-3,
IMR-32 and SK-N-F1 (Figs. 1A and
S1). In this group of cell lines,
the mRNA and protein levels of KDM5D were examined in relation to
CDDP resistance and CDDP treatment. The RT-qPCR results showed a
loss of KDM5D expression in all CDDP-resistant cell lines. In
addition, CDDP treatment decreased the expression of KDM5D in
sensitive cells and this decrease was dependent on the duration of
incubation with CDDP. The present study observed that no expression
was present in drug-resistant cell lines even after treatment with
CDDP (Figs. 1B and S1). The same results were observed at the
protein level, as determined by means of flow cytometry (Figs. 1C and S1, S2,
S3, S4, S5,
S6, S7). For further experiments, UKF-NB-3 and
UKF-NB-3CDDP cell lines were selected, as they were used
previously to conduct a study with this cell line on the importance
of KDM5B for CDDP-resistance (18).
KDM5D expression decreases histone
H3K4 trimethylation in neuroblastoma cells
H3K4me3 is frequently associated with
transcriptional activation of neighboring genes and therefore could
activate oncogenes (64,65). We observed the change in
trimethylation of H3K4 after CDDP treatment, silencing or
overexpression of KDM5D in neuroblastoma cell lines (Fig. S8). To demonstrate the importance of
KDM5D for methylation, the present study monitored H3K4
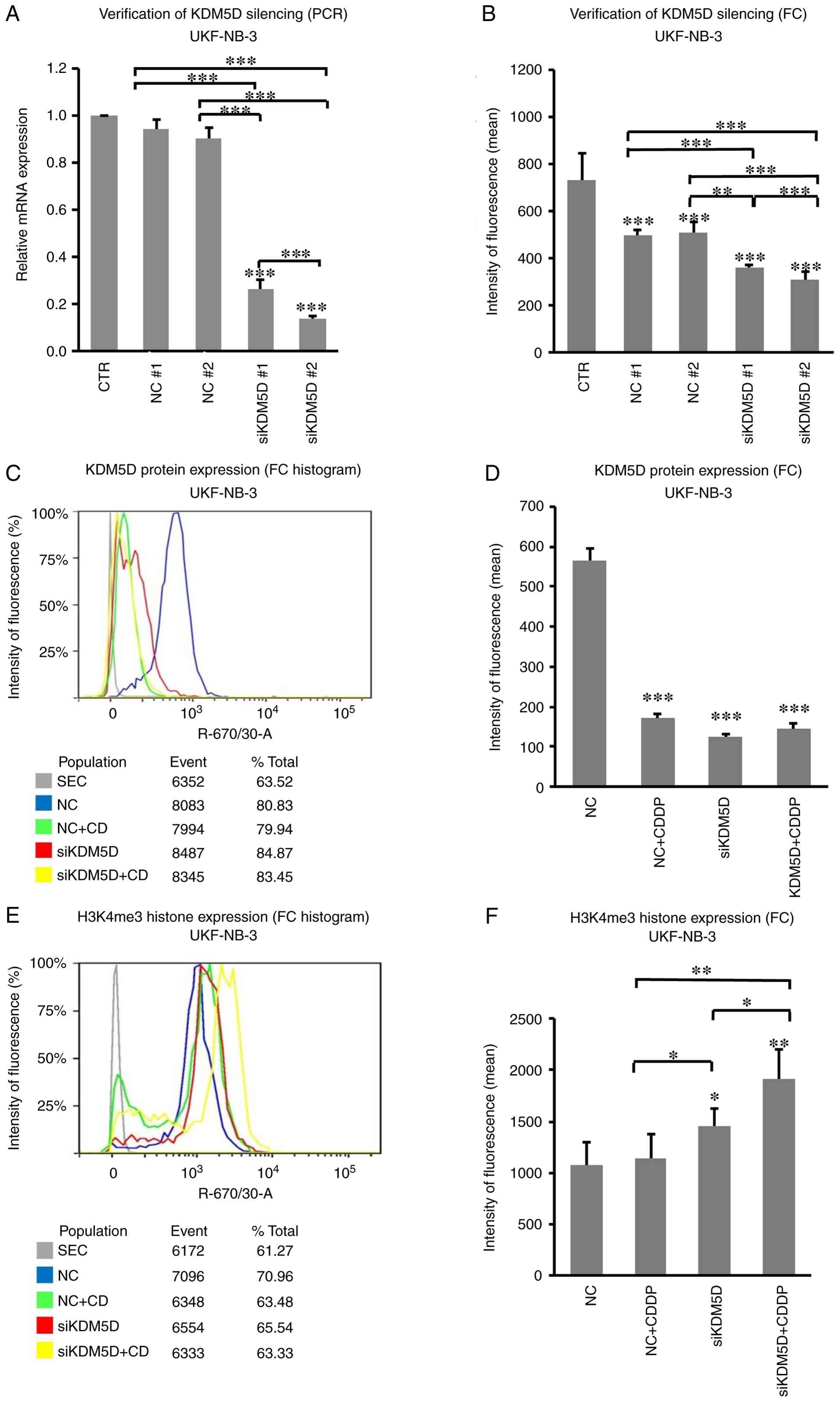
trimethylation after KDM5D silencing and overexpression. siRNA
against KDM5D in the cell line UKF-NB-3 decreased expression
levels of KDM5D mRNA (measured by RT-PqCR; Fig. 2A) and protein (measured by flow
cytometry; Figs. 2B and S9) compared with non-coding
siRNA-transfected control cells. Treatment of UKF-NB-3 with CDDP
reduced KDM5D expression in controls, but did not further reduce
expression in siRNA-transfected cells (Figs. 2C, D and S10). In addition, trimethylation of
histone H3K4 was significantly increased in the KDM5D knockdown
cells compared with the UKF-NB-3 control cells as shown by flow
cytometry results, whereas the treatment with CDDP increased
trimethylation only in the transfected cells, but not in control
(Figs. 2E, F and S11).
The CDDP-resistant cell line
UKF-NB-3CDDP, which does not express KDM5D, was
transfected with the KDM5D ORF clone plasmid pCMV-3Tag1 carrying
the KDM5D gene for 48 h, resulting in overexpression of
KDM5D at the protein level compared with cells transfected with the
control plasmid (Figs. 3A and
S12). In addition, treatment with
CDDP decreased the expression of KDM5D in transfected cells
(Figs. 3B, C and S13). Trimethylation of histone H3K4 was
significantly decreased in cells transfected with KDM5D
compared with control, while treatment with CDDP increased H3K4
trimethylation in both control and transfected cells (Figs. 3D, E and S14).
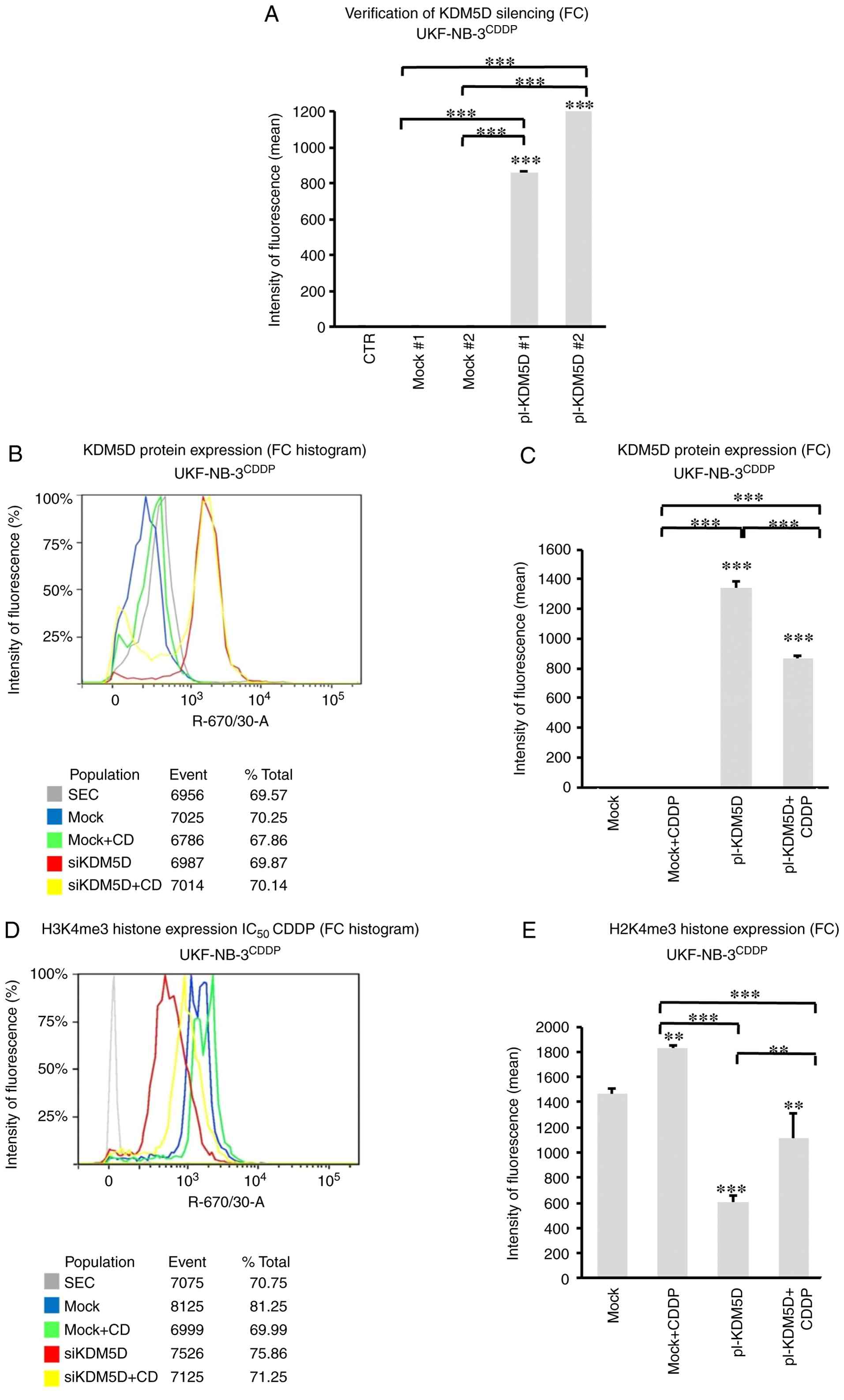 | Figure 3.Aberrant expression of KDM5D affects
histone H3K4 trimethylation in the CDDP-resistant neuroblastoma
cell line UKF-NB-3CDDP. (A) Flow cytometry showed
increased levels of KDM5D protein in the UKF-NB-3CDDP
cell line transfected with the KDM5D ORF-clone plasmid pCMV-3Tag1,
expressing the KDM5D gene (pl-KDM5D #, 8 ng/µl; or pl-KDM5D #2, 16
ng/µl) compared with control cells transfected with the control
plasmid pCMV6-AC-GFP (mock #1, 8 ng/µl; or mock #2, 16 ng/µl),
while control cells did not express KDM5D (P<0.001). (B) Flow
cytometry histogram showed increased expression of KDM5D protein
after KDM5D overexpression (48 h). (C) Flow cytometry proved
increased protein level of KDM5D in KDM5D transfected
UKF-NB-3CDDP cells and showed that CDDP treatment in
transfected cells decreased protein expression of KDM5D
(P<0.001). (D) Flow cytometry histogram showed that histone H3K4
trimethylation decreased after KDM5D overexpression (48 h). (E)
KDM5D overexpression decreased histone H3K4 trimethylation in
UKF-NB-3CDDP (P<0.001) but treatment with CDDP
increased H3K4me3 trimethylation in transfected cells (P<0.05).
Data are shown as mean ± standard deviation from three independent
experiments. Statistical significance was determined using one-way
(A) and two-way ANOVA (C, E) with Tukey's post hoc test
**P<0.01; ***P<0.001. CTR, control; pl-ORF-clone plasmid;
SEC, control with only the secondary antibody; Mock/NC, control
cells with non-coding RNA; pl-KDM5D, control cells transfected with
pl-KDM5D; FC, flow cytometry. |
KDM5D expression increases the cell
sensitivity to CDDP
To determine the effect of KDM5D expression on
sensitivity to CDDP in neuroblastoma, the percentage of viable
cells following treatment with CDDP at varying concentrations and
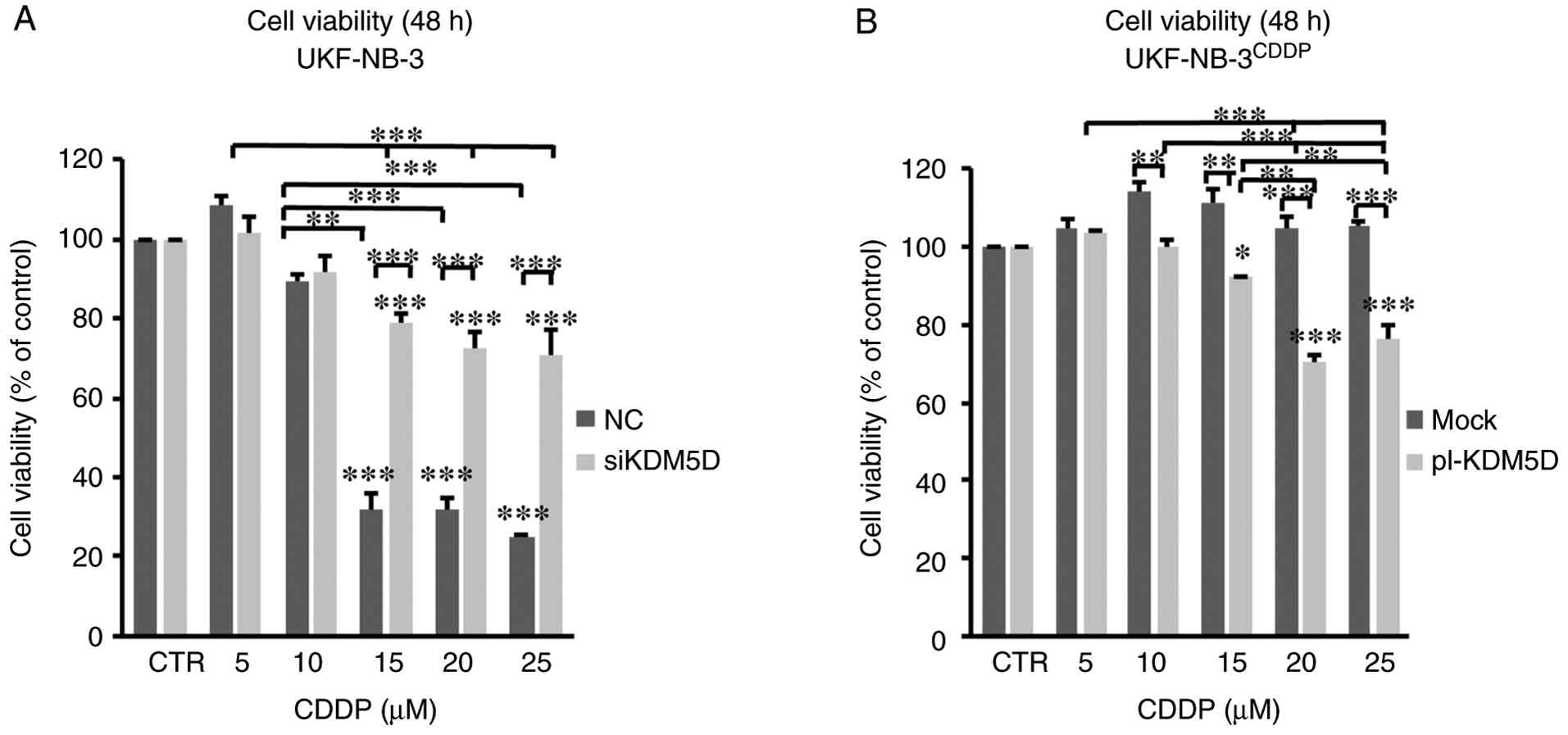
time intervals was monitored. KDM5D knockdown did not affect
the viability of cells in UKF-NB-3 after 24 h of transfection
(Fig. S15) or after 48 h of
transfection (Fig. 4A). The
viability of KDM5D knockdown UKF-NB-3 cells decreased after
incubation with CDDP depending on incubation time and
concentration, but the decrease in viability of NC transfected
cells after treatment with CDDP was more substantial (Figs. 4A and S16).
To compare the effect of inhibition of KDM5s
demethylases on the viability and sensitivity of UKF-NB-3 and
UKF-NB3CDDP cells to CDDP, KDOAM-25 citrate was used as
an inhibitor of the KDM5 demethylases subfamily (66), and therefore a concentration that
effectively reduced H3K4 trimethylation without influencing cell
survival (400 nM) was selected (data not shown). Inhibition of KDM5
by KDOAM-25 protected both sensitive (UKF-NB-3) and resistant
(UKF-NB-3CDDP) cells from the effects of CDDP (Fig. S16).
Overexpression of KDM5D did not alter cell
viability in UKF-NB-3CDDP after 24 h, even after
incubation with CDDP (Fig. S17).
However, after 48 h of incubation with CDDP, cell viability
decreased with increasing CDDP concentration compared with
mock-transfected controls, whose viability did not change after
treatment with CDDP (Fig. 4B).
The decreased sensitivity to CDDP after incubation
with the inhibitor was consistent with the downregulation of KDM5D
by siRNA, and conversely, the increase in sensitivity to CDDP in
the resistant line with induced KDM5D expression suggests the
importance of KDM5D in the anticancer efficacy of CDDP.
Expression of KDM5D increases CDDP
induced apoptosis in neuroblastoma cells
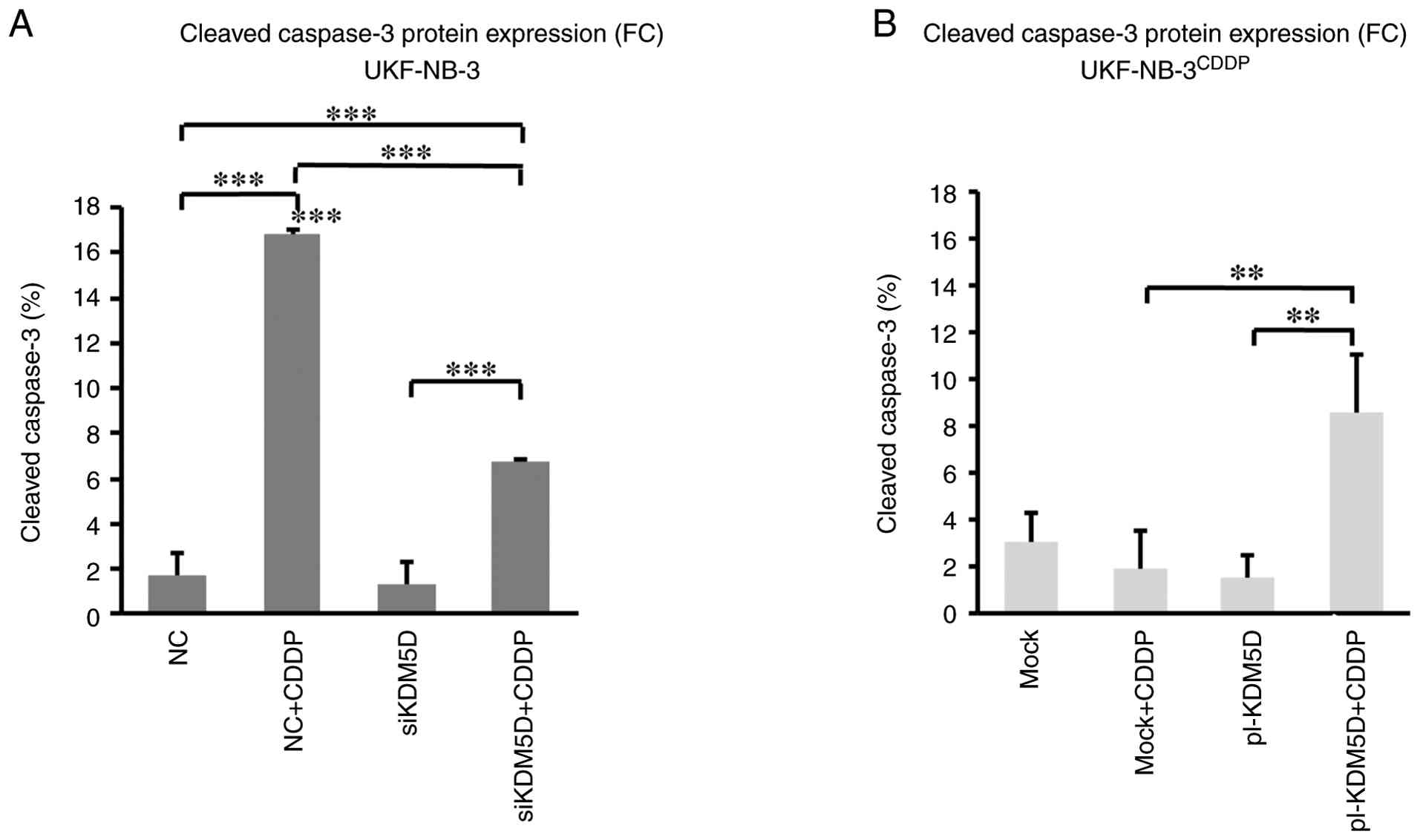
To determine whether overexpression of KDM5D
in resistant cells and CDDP treatment induces apoptosis, activated
caspase-3 and cleavage of PARP was examined to determine the
relationship between the expression of KDM5D and CUL4A. Control
cells had increased levels of activated caspase-3 after treatment
with CDDP. Silencing of KDM5D in the sensitive cell line
UKF-NB-3 did not change the amount of activated caspase-3 compared
with control cells. Treatment with CDDP increased activated
caspase-3, but the level of cleaved caspase-3 was significantly
lower in cells with KDM5D knockdown compared with controls
after CDDP treatment (Figs. 5A,
S18 and S19). The KDM5 pan-inhibitor KDOAM-25
citrate had a similar effect as KDM5D silencing (Fig. S20, Fig. S21, Fig. S22). Overexpression of KDM5D
resulted in an increase in activated caspase-3 level after
treatment with CDDP in contrast with control cells (Figs. 5B and S23). Taken together, KDM5D is involved in
the sensitivity to CDDP induced apoptosis in neuroblastoma
cells.
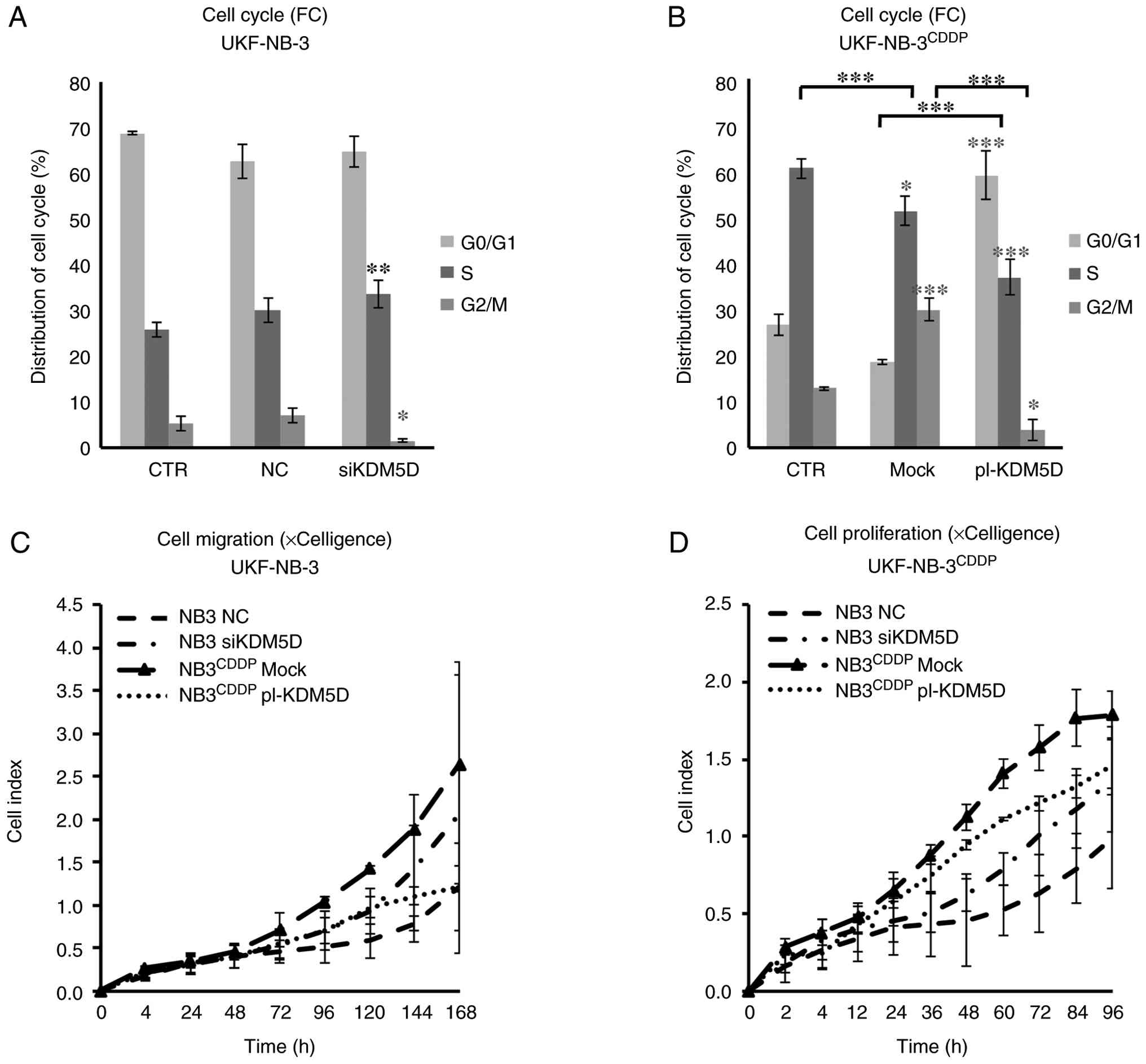
KDM5D expression decreased S-phase,
cell proliferation and migration
Because inhibition of KDM5D protects cells from the
effect of CDDP (Figs. 4A, 5A, S15
and S16) and artificial
expression, on the contrary, increases the effect of this
cytostatic agent (Figs. 4B,
5B and S17), the present study focused on the
significance of KDM5D for cell cycle and proliferation. In the
sensitive cells, silencing of KDM5D resulted in an increase
of cells in S phase and a decrease in G2/M phase compared with the
control (Figs. 6A, S24 and C). Transfection of the plasmid
with KDM5D in the cell line UKF-NB-3CDDP
decreased the percentage of cells in S phase and increased G0/G1
phase compared with control (Figs.
6B, S25A and E). Treatment
with CDDP resulted in cell cycle arrest at checkpoint G0/G1 and
G2/M in the sensitive cell line, and therefore the cell cycle was
not evaluable (Fig. S24B, D,
F).
The xCELLigence system was used to monitor cell
migration (Figs. 6C and S26A) and proliferation (Figs. 6D and S26B-D) in real time, which showed that
the CDDP-resistant UKF-NB-3CDDP had a higher cell index
of migration and proliferation compared with the CDDP-sensitive
UKF-NB-3 with KDM5D expression. In addition, KDM5D knock down
increased cell proliferation even in IMR-32 and SK-N-F1 lines
(Fig. S26C-D). The silencing of
KDM5D in UKF-NB-3 increased cell migration and proliferation, while
KDM5D overexpression in UKF-NB-3CDDP cells decreased
migration and proliferation compared with controls. KDM5D silencing
was verified over a period of 7 days using western blotting
(Fig. S26E).
KDM5D plays a role in the development
of chemoresistance to CDDP in neuroblastoma through regulation of
the CUL4A gene
To assess the relationship between CUL4A and KDM5D
(Fig. S18), its expression in
CDDP-sensitive and resistant cells was examined. The CDDP-sensitive
cell line UKF-NB-3 showed reduced expression of CUL4A protein
compared with the CDDP-resistant UKF-NB-3CDDP, and the
expression of CUL4A increased after treatment with CDDP in the
sensitive but not in the resistant cells (Figs. 7A, B S27 and S28). In addition, knockdown of
KDM5D increased CUL4A expression Figs. 7B and S27). In UKF-NB-3CDDP cells,
overexpression of KDM5D decreased CUL4A expression compared
with control, while CDDP treatment of cells with overexpressed
KDM5D increased CUL4A expression levels. However, in the
control, CDDP treatment did not alter CUL4A expression in resistant
cells (Figs. 7C and S28). Inhibition of all KDM5s members by
KDOAM-25 in sensitive cells increased level of CUL4A, compared with
control but did not change CUL4A expression levels in
chemoresistant cells that do not express KDM5D (Figs. S11 and S29, S30,
S31).
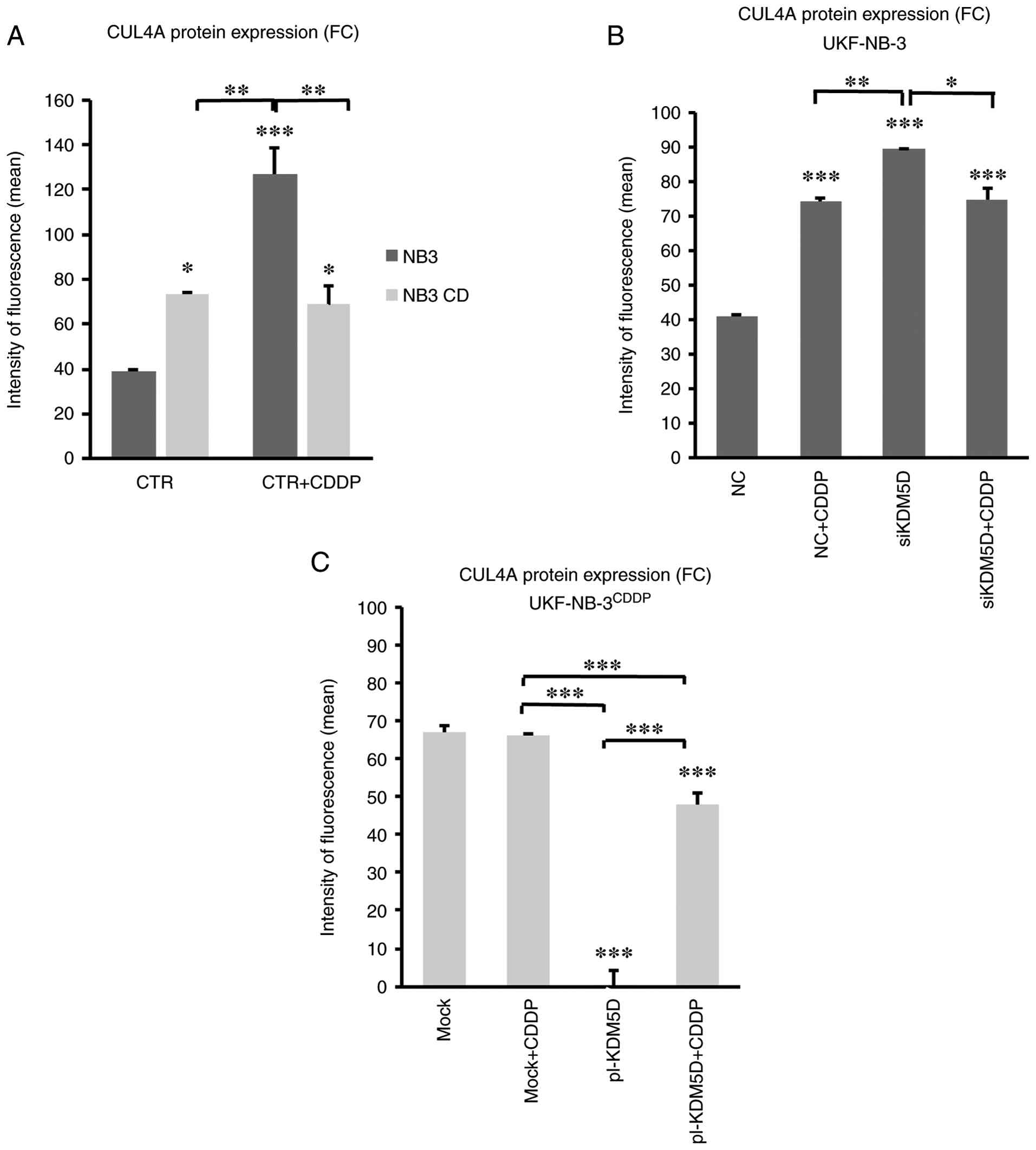 | Figure 7.KDM5D and CDDP treatment affect
expression of CUL4A in UKF-NB-3 and UKF-NB-3CDDP. Flow
cytometry analysis in (A) UKF-NB-3 and UKF-NB-3CDDP,
where sensitive cell line UKF-NB-3 showed lower expression of CUL4A
gene compared with the resistant cell line (P<0.05) and this
expression increased in sensitive cells after CDDP treatment
(P<0.001) compared with control. (B) Silencing of KDM5D
increased of CUL4A level (P<0.001). (C) In the CDDP-resistant
cell line UKF-NB-3CDDP, KDM5D overexpression decreased
CUL4A expression compared with mock and control plasmid
(P<0.001). CDDP treatment of overexpressed cells with KDM5D
increased the level of CUL4A (P<0.001), while in control cells
CDDP treatment did not change the level of CUL4A. Data are shown as
mean ± standard deviation from three independent experiments.
Statistical significance was determined using two-way ANOVA with
Tukey's post hoc test. *P<0.05; **P<0.01; ***P<0.001
(ANOVA with Tukey's post hoc test). CTR, control; NC, non-coding
RNA; si, short interfering RNA; pl, ORF-clone plasmid; NB3,
UKF-NB-3; Mock/NC, control cells; siKDM5D/pl-KDM5D, control cells
transfected with siKDM5D/pl-KDM5D; FC, flow cytometry. |
Reduced expression of KDM5D is
associated with poor overall survival
To investigate the role of KDM5D and
CUL4A in neuroblastoma, patient data was screened from the
R2 database: Genomics Analysis and Visualization Platform
(http://r2.amc.nl). The association between
KDM5D expression and overall survival was compared in male
patients with two different neuroblastoma risk groups according to
International Neuroblastoma Staging System (INSS) stages:
non-high-risk (st1, st2, st3 and st4S) and high-risk (st4)
(67). This analysis showed that
low KDM5D expression was associated with poor survival rates
in the group with st4 (Figs. 8B and
S32). Analysis of three different
datasets of patients with neuroblastoma from the R2 platform
indicated a statistically significant correlation between
KDM5D expression and higher survival rates in high-risk
neuroblastoma patients. In the non-high-risk group, there was not a
significant difference in all data sets (Figs. 8A and S32). These observations are consistent
with our hypothesis that low KDM5D concentration in neuroblastoma
has tumor-promoting effects and is associated with
chemoresistance.
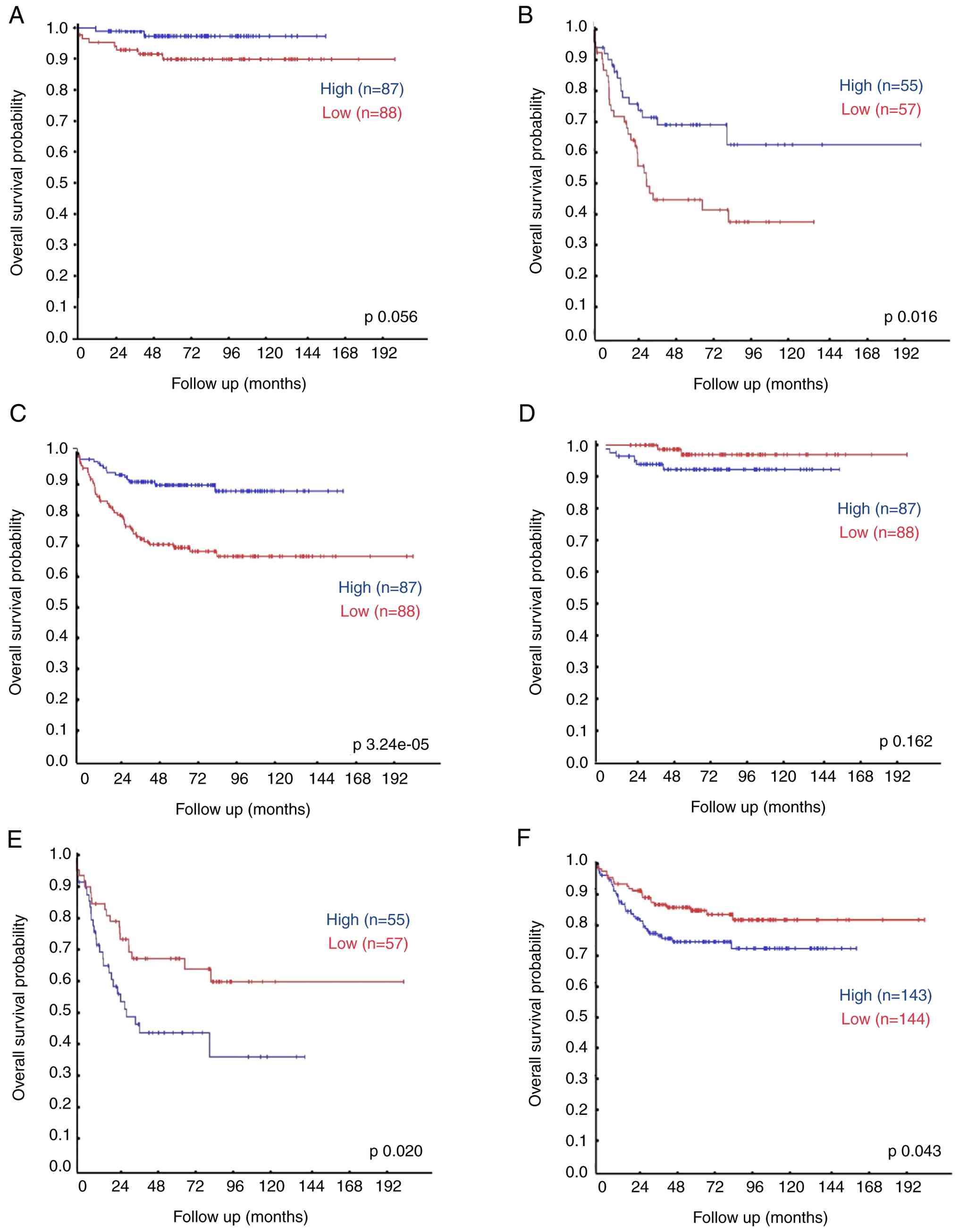 | Figure 8.Kaplan-Meier curves analyzing KDM5D
(A, B, C) and CUL4A (D, E, F) expression data and overall survival
of patients in the SEQC neuroblastoma dataset (498; RPM; seqcnb1):
(A) Male patients with stages 1, 2, 3 and 4S, KDM5AD expression.
(B) Male patients with stage 4, KDM5D expression. (C) All male
patients according to the International Neuroblastoma Staging
System, KDM5D expression. (D) Male patients with stages 1, 2, 3 and
4S, CUL4A expression. (E) Male patients with stage 4, CUL4A
expression. (F) All male patients according to the International
Neuroblastoma Staging System, CUL4A expression. Statistical
significance of survival differences was determined using the
log-rank (Mantel-Cox) test. R2: Genomics Analysis and Visualization
Platform revealed that low relative expression of KDM5D and high
expression of CUL4A is associated with worse survival. |
To uncover the relationship between the expression
of KDM5D and CUL4A, the present study further
analyzed the data from the aforementioned databases in R2: for an
association between the expression of CUL4A and overall
survival of neuroblastoma patients. Kaplan-Meier analysis in these
datasets revealed that high relative expression of CUL4A is
often associated with worse survival in two datasets of patients
with stages 1, 2, 3 or 4S (Figs. 8D
and S33); moreover, this
association is significant in stage 4 (Figs. 8E and S32), however, this was not true in the
Oberthuer dataset.
Discussion
It is well known that in cancer, aberrant epigenetic
modifications, which include histone methylation, contribute to
various stages of neoplastic development, including initiation,
promotion, invasion, metastasis, EMT and chemoresistance (1–5). There
is increasing evidence for the importance of histone lysine
demethylase function, the dysregulation of which has been described
in several types of cancer (8,9).
Aberrant expression of KDM5D has been observed in a number of types
of cancer (27–33). According to the results of analysis
of publicly available genomics data, the present study found that
low expression of KDM5D is associated with worse survival in male
patients with stage 4 neuroblastoma. As drug resistance often
develops in high-risk neuroblastoma (53,54),
the present study investigated the role of KDM5D in the development
of chemoresistance to CDDP, which is commonly used in therapy of
the high-risk neuroblastoma (68).
The present study demonstrated that expression of
KDM5D was lost in all tested CDDP-resistant neuroblastoma cell
lines compared with sensitive parental cells. Silencing of
KDM5D in the sensitive neuroblastoma cell lines resulted in
increased trimethylation of H3K4, decreased sensitivity of cells to
CDDP, inhibition of apoptosis induced by CDDP, increased
proliferation and proportion of cells in S phase and acceleration
of cell migration. To date, limited studies have been published
describing the relationship between KDM5D and chemoresistance.
Komura et al (27) found
that KDM5D plays an important role as a tumor-suppressor in
castration-resistant prostate cancer, where it activates the
androgen receptor, and low KDM5D expression associates with
docetaxel insensitivity, aggressiveness and worse prognosis. KDM5D,
along with five other Y-linked genes, is reported to be
downregulated in 12 major non-reproductive types of cancer,
suggesting selection against their activity and their function as
tumor-suppressors (69,70). Moreover, loss of the Y chromosome
was described as being associated with shorter overall survival and
resistance to radiotherapy and cisplatin-based chemotherapeutics in
male neck squamous cell carcinoma (66,71).
The present study also observed that the CDDP-resistant
neuroblastoma cell line had increased expression of CUL4A compared
with the sensitive parental cells and that KDM5D knockdown in the
sensitive cells increased CUL4A.
The present study concludes that low expression of
KDM5D is associated with worse survival in male patients with
neuroblastoma, as shown by analysis of publicly available genomics
data. Accordingly, the present study examined neuroblastoma cell
lines with genotype XY and showed that all CDDP-resistant cell
lines had KDM5D expression below the detection limit. Silencing of
KDM5D results in increased trimethylation of histone H3K4 or
vice versa overexpression decreased H3K4 trimethylation in
CDDP-resistant cells. Our previous study demonstrated an increase
in H3K4me3 after silencing of KDM5B (18), therefore all members of the KDM5
subfamily may potentially have an important function in regulating
of H3K4 methylation.
Results of the present study demonstrate that KDM5D
expression affects proliferation, migration and sensitivity of
neuroblastoma cells to CDDP. Both the reduction of KDM5D expression
in cells due to cytostatics and the more selective toxicity of
cytostatics towards cells with increased expression of KDM5D and
thus the selection of cells with lower expression can be applied.
The concentration of CDDP (20 µM) used markedly reduced cell
viability compared with controls after a 24 h incubation, and KDM5D
expression values measured by RT-qPCR at this time interval were
reduced insignificantly. From this, it is difficult to draw
conclusions about which of the mechanisms applies.
Finally, the present study examined the expression
of the gene CUL4A. Sensitive cell lines showed reduced
expression of CUL4A compared with resistant ones. Moreover,
this expression increased after silencing of KDM5D, CDDP or
KDOAM-25 treatment in CDDP-sensitive but not in resistant cells,
which do not express KDM5D. These findings suggest that CDDP and
KDOAM-25 affect CUL4A expression via KDM5D. The relationship
between the expression of KDM5D and CUL4A was described by Shen
et al (34) in a study
demonstrating that KDM5D regulates the methylation of H3K4 in the
promoter of CUL4A and represses the expression of CUL4A. This study
revealed that CUL4A expression plays an important role in
metastasis and EMT of gastric cancer cells but to the best of our
knowledge, there is no information regarding the importance of
CUL4A in neuroblastoma.
Englinger et al (53) showed that loss of CUL4A expression
leads to hypersensitivity to CDDP in colon cancer cells.
Downregulation of CUL4A mediated a lack of nucleotide
excision repair that caused trabedectin resistance and collateral
CDDP hypersensitivity in colorectal carcinomas.
Overall, in CDDP-resistant neuroblastoma cell
lines, there is a loss of KDM5D expression that leads to a more
aggressive phenotype of neuroblastoma by promoting cell
proliferation and migration, evading cell death, promoting S phase
of the cell cycle and desensitizing susceptible cells to CDDP. The
data of the present study may suggest that the change in expression
is caused by demethylation of H3K4 in the CUL4A gene. The
precise mechanisms by which KDM5D and CUL4A are involved in the
development of chemoresistance remain to be elucidated. To the best
of our knowledge, the relationship between KDM5D and CUL4A in
neuroblastoma has not yet been studied. We hypothesize that KDM5D
and/or CUL4A may be a useful biomarker for detection of
chemoresistance in clinical practice and that inhibition of CUL4A
may be a potential therapeutic approach in neuroblastoma after
further preclinical and clinical studies.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
This research was funded by the Grant Agency of Charles
University, Czech Republic (grant no. 822219) and by Ministry of
Health of the Czech Republic (grant no. NW24-03-00101).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
TE conceived and led the present study. NP
performed sample preparation, RT-qPCR, flow cytometry, cell
proliferation assay, siRNA and ORF cDNA transfection, cell
proliferation monitoring, data analysis, and the statistical
analysis, and wrote the manuscript. JH contributed to the
acquisition of data by performing part of the flow cytometry
experiments and assisting in data interpretation. MB contributed to
the acquisition of data by preparing biological samples and
performing part of the RT-qPCR experiments, and assisted in data
interpretation. NP, TE, JH and MB confirm the authenticity of all
the raw data. All authors have reviewed the manuscript and read and
approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Barski A, Cuddapah S, Cui K, Roh TY,
Schones DE, Wang Z, Wei G, Chepelev I and Zhao K: High-resolution
profiling of histone methylations in the human genome. Cell.
129:823–837. 2007. View Article : Google Scholar
|
|
2
|
Mikkelsen TS, Ku M, Jaffe DB, Issac B,
Lieberman E, Giannoukos G, Alvarez P, Brockman W, Kim TK, Koche RP,
et al: Genome-wide maps of chromatin state in pluripotent and
lineage-committed cells. Nature. 448:553–560. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Tsukada YI, Fang J, Erdjument-Bromage H,
Warren ME, Borchers CH, Tempst P and Zhang Y: Histone demethylation
by a family of JmjC domain-containing proteins. Nature.
439:811–816. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Shukla A, Chaurasia P and Bhaumik SR:
Histone methylation and ubiquitination with their cross-talk and
roles in gene expression and stability. Cell Mol Life Sci.
66:1419–1433. 2009. View Article : Google Scholar
|
|
5
|
Jambhekar A, Dhall A and Shi Y: Roles and
regulation of histone methylation in animal development. Nat Rev
Mol Cell Biol. 20:625–641. 2019. View Article : Google Scholar
|
|
6
|
Plch J, Hrabeta J and Eckschlager T: KDM5
demethylases and their role in cancer cell chemoresistance. Int J
Cancer. 144:221–231. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ruthenburg AJ, Allis CD and Wysocka J:
Methylation of lysine 4 on histone H3: Intricacy of writing and
reading a single epigenetic mark. Mol Cell. 25:15–30. 2007.
View Article : Google Scholar
|
|
8
|
Arifuzzaman S, Khatun MR and Khatun R:
Emerging of lysine demethylases (KDMs): From pathophysiological
insights to novel therapeutic opportunities. Biomed Pharmacother.
129:1103922020. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sterling J, Menezes SV, Abbassi RH and
Munoz L: Histone lysine demethylases and their functions in cancer.
Int J Cancer. 148:2375–2388. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang N, Ma T and Yu B: Targeting
epigenetic regulators to overcome drug resistance in cancers.
Signal Transduct Target Ther. 8:692023. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wu H, Xu L and Hu X: KDM5A regulates the
growth and gefitinib drug resistance against human lung
adenocarcinoma cells 3. Biotech. 12:972022.
|
|
12
|
Banelli B, Carra E, Barbieri F, Würth R,
Parodi F, Pattarozzi A, Carosio R, Forlani A, Allemanni G, Marubbi
D, et al: The histone demethylase KDM5A is a key factor for the
resistance to temozolomide in glioblastoma. Cell Cycle.
14:3418–3429. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Feng T, Wang Y, Lang Y and Zhang Y: KDM5A
promotes proliferation and EMT in ovarian cancer and closely
correlates with PTX resistance. Mol Med Rep. 16:3573–3580. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hou J, Wu J, Dombkowski A, Zhang K,
Holowatyj A, Boerner JL and Yang ZQ: Genomic amplification and a
role in drug-resistance for the KDM5A histone demethylase in breast
cancer. Am J Transl Res. 4:247–256. 2012.PubMed/NCBI
|
|
15
|
Roesch A, Vultur A, Bogeski I, Wang H,
Zimmermann KM, Speicher D, Körbel C, Laschke MW, Gimotty PA,
Philipp SE, et al: Overcoming intrinsic multidrug resistance in
melanoma by blocking the mitochondrial respiratory chain of
slow-cycling JARID1B high cells. Cancer Cell. 23:811–825. 2013.
View Article : Google Scholar
|
|
16
|
Xu W, Zhou B, Zhao X, Zhu L, Xu J, Jiang
Z, Chen D, Wei Q, Han M, Feng L, et al: KDM5B demethylates H3K4 to
recruit XRCC1 and promote chemoresistance. Int J Biol Sci.
14:1122–1132. 2018. View Article : Google Scholar
|
|
17
|
Liu J and Nie C: KDM5B regulates the
PTEN/PI3K/Akt pathway to increase sorafenib-resistance in
hepatocellular carcinoma. Anticancer Drugs. 33:840–849. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Belhajova M, Podhorska N, Vicha A and
Eckschlager T: KDM5B expression in cisplatin resistant
neuroblastoma cell lines. Oncol Lett. 24:3652022. View Article : Google Scholar
|
|
19
|
Kuo YT, Liu YL, Adebayo BO, Shih PH, Lee
WH, Wang LS, Liao YF, Hsu WM, Yeh CT and Lin CM: JARID1B expression
plays a critical role in chemoresistance and stem cell-like
phenotype of neuroblastoma cells. PLoS One. 10:e01253432015.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li L, Shou H, Wang Q and Liu S:
Investigation of the potential theranostic role of KDM5B/miR-29c
signaling axis in paclitaxel resistant endometrial carcinoma. Gene.
694:76–82. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang L, Mao Y, Du G, He C and Han S:
Overexpression of JARID1B is associated with poor prognosis and
chemotherapy resistance in epithelial ovarian cancer. Tumor Biol.
36:2465–2472. 2015. View Article : Google Scholar
|
|
22
|
Lin H, Yang G, Yu J, Wang J, Li Q, Guo S
and Cao B: KDM5C inhibits multidrug resistance of colon cancer cell
line by down-regulating ABCC1. Biomed Pharmacother. 107:1205–1209.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Hong Z, Wu G, Xiang ZD, Xu CD, Huang SS,
Li C, Shi L and Wu DL: KDM5C is transcriptionally regulated by BRD4
and promotes castration-resistance prostate cancer cell
proliferation by repressing PTEN. Biomed Pharmacother.
114:1087932019. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Outchkourov NS, Muiño JM, Kaufmann K, van
IJcken WF, Groot Koerkamp MJ, van Leenen D, de Graaf P, Holstege
FC, Grosveld FG and Timmers HT: Balancing of histone H3K4
methylation states by the Kdm5c/SMCX histone demethylase modulates
promoter and enhancer function. Cell Rep. 3:1071–1079. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Agulnik AI, Longepied G, Ty MT and Bishop
CE: Mitchell M0020: Mouse H-Y encoding Smcy gene and its X
chromosomal homolog Smcx. Mamm Genome. 10:926–929. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Akimoto C, Kitagawa H, Matsumoto T and
Kato S: Spermatogenesis-specific association of SMCY and MSH5.
Genes Cells. 13:623–633. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Komura K, Jeong SH, Hinohara K, Qu F, Wang
X, Hiraki M, Azuma H, Lee GS, Kantoff PW and Sweeney CJ: Resistance
to docetaxel in prostate cancer is associated with androgen
receptor activation and loss of KDM5D expression. Proc Natl Acad
Sci USA. 113:6259–6264. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Jangravi Z, Tabar MS, Mirzaei M,
Parsamatin P, Vakilian H, Alikhani M, Shabani M, Haynes PA,
Goodchild AK, Gourabi H, et al: Two splice variants of Y
chromosome-located lysine-specific Demethylase 5D have distinct
function in prostate cancer cell line (DU-145). J Proteome Res.
14:3492–3502. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li N, Dhar SS, Chen TY, Kan PY, Wei Y, Kim
JH, Chan CH, Lin HK, Hung MC and Lee MG: JARID1D is a suppressor
and prognostic marker of prostate cancer invasion and metastasis.
Cancer Res. 76:831–843. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Arseneault M, Monlong J, Vasudev NS,
Laskar RS, Safisamghabadi M, Harnden P, Egevad L, Nourbehesht N,
Panichnantakul P, Holcatova I, et al: Loss of chromosome Y leads to
down regulation of KDM5D and KDM6C epigenetic modifiers in clear
cell renal cell carcinoma. Sci Rep. 7:448762017. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhu M, Zhang RN, Zhang H, Qu CB, Zhang XC,
Ren LX, Yang Z and Gu JF: PCGF6/MAX/KDM5D facilitates MAZ/CDK4 axis
expression and pRCC progression by hypomethylation of the DNA
promoter. Epigenetics Chromatin. 16:92023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Chen TM, Huang CM, Setiawan SA, Hsieh MS,
Sheen CC and Yeh CT: KDM5D histone demethylase identifies
platinum-tolerant head and neck cancer cells vulnerable to mitotic
catastrophe. Int J Mol Sci. 24:53102023. View Article : Google Scholar
|
|
33
|
Dunford A, Weinstock DM, Savova V,
Schumacher SE, Cleary JP, Yoda A, Sullivan TJ, Hess JM, Gimelbrant
AA, Beroukhim R, et al: Tumor-suppressor genes that escape from
X-inactivation contribute to cancer sex bias. Nature Genet.
49:10–16. 2017. View Article : Google Scholar
|
|
34
|
Shen X, Hu K, Cheng G, Xu L, Chen Z, Du P
and Zhuang Z: KDM5D inhibit epithelial-mesenchymal transition of
gastric cancer through demethylation in the promoter of Cul4A in
male. J Cell Biochem. 120:12247–12258. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Liu M and Gao N: KDM5D inhibits the
transcriptional activation of FKBP4 by suppressing the expression
of E2F1 in colorectal cancer in males. Biochem Pharmacol.
194:1148142021. View Article : Google Scholar
|
|
36
|
Li S, Wu Z, Li Q, Liang Q, Zhou H, Shi Y,
Zhang R and Pan H: The prognostic value of AT-rich interaction
domain (ARID) family members in patients with hepatocellular
carcinoma. Evid Based Complement Alternat Med.
2022:11503902022.PubMed/NCBI
|
|
37
|
Duan Y, Du Y, Gu Z, Zheng X and Wang C:
Expression, prognostic value, and functional mechanism of the KDM5
family in pancreatic cancer. Front Cell Dev Biol. 10:8873852022.
View Article : Google Scholar
|
|
38
|
Abdel-Hafiz HA, Schafer JM, Chen X, Xiao
T, Gauntner TD, Li Z and Theodorescu D: Y chromosome loss in cancer
drives growth by evasion of adaptive immunity. Nature. 619:624–631.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Li J, Lan Z, Liao W, Horner JW, Xu X, Liu
J, Yoshihama Y, Jiang S, Shim HS, Slotnik M, et al: Histone
demethylase KDM5D upregulation drives sex differences in colon
cancer. Nature 619: 632–639, 2023. Hannah J and Zhou P: Distinct
and overlapping functions of the cullin E3 ligase scaffolding
proteins CUL4A and CUL4B. Gene. 573:33–45. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Jin J, Arias EE, Chen J, Harper JW and
Walter JC: A family of diverse Cul4-Ddb1-interacting proteins
includes Cdt2, which is required for S phase destruction of the
replication factor Cdt1. Mol Cell. 23:709–721. 2006. View Article : Google Scholar
|
|
41
|
Yasui K, Arii S, Zhao C, Imoto I, Ueda M,
Nagai H, Emi M and Inazawa J: TFDP1, CUL4A, and CDC16 identified as
targets for amplification at 13q34 in hepatocellular carcinomas.
Hepatology. 35:1476–1484. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Jia L, Yan F, Cao W, Chen Z, Zheng H, Li
H, Pan Y, Narula N, Ren X, Li H and Zhou P: Dysregulation of CUL4A
and CUL4B Ubiquitin ligases in lung cancer. J Biol Chem.
292:2966–2978. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Hannah J and Zhou PB: The CUL4A ubiquitin
ligase is a potential therapeutic target in skin cancer and other
malignancies. Chin J Cancer. 32:478–482. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Chen LC, Manjeshwar S, Lu Y, Moore D,
Ljung BM, Kuo WL, Dairkee SH, Wernick M, Collins C and Smith HS:
The human homologue for the Caenorhabditis elegans cul-4 gene is
amplified and overexpressed in primary breast cancers. Cancer Res.
58:3677–3683. 1998.PubMed/NCBI
|
|
45
|
Hung MS, Mao JH, Xu Z, Yang CT, Yu JS,
Harvard C, Lin YC, Bravo DT, Jablons DM and You L: Cul4A is an
oncogene in malignant pleural mesothelioma. J Cell Mol Med.
15:350–358. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Ren S, Xu C, Cui Z, Yu Y, Xu W, Wang F, Lu
J, Wei M, Lu X, Gao X, et al: Oncogenic CUL4A determines the
response to thalidomide treatment in prostate cancer. J Mol Med
(Berl). 90:1121–1132. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Shiyanov P, Nag A and Raychaudhuri P:
Cullin 4A associates with the UV-damaged DNA-binding protein DDB. J
Biol Chem. 274:35309–35312. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Nag A, Bondar T, Shiv S and Raychaudhuri
P: The xeroderma pigmentosum group E gene product DDB2 is a
specific target of cullin 4A in mammalian cells. Mol Cell Biol.
21:6738–6747. 2001. View Article : Google Scholar
|
|
49
|
Kapetanaki MG, Guerrero-Santoro J, Bisi
DC, Hsieh CL, Rapić-Otrin V and Levine AS: The DDB1-CUL4ADDB2
ubiquitin ligase is deficient in xeroderma pigmentosum group E and
targets histone H2A at UV-damaged DNA sites. Proc Natl Acad Sci
USA. 103:2588–2593. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Shinomiya T, Mori T, Ariyama Y, Sakabe T,
Fukuda Y, Murakami Y, Nakamura Y and Inazawa J: Comparative genomic
hybridization of squamous cell carcinoma of the esophagus: The
possible involvement of the DP1 gene in the 13q34 amplicon. Genes
Chromosomes Cancer. 24:337–344. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Dohna M, Reincke M, Mincheva A, Allolio B,
Solinas-Toldo S and Lichter P: Adrenocortical carcinoma is
characterized by a high frequency of chromosomal gains and
high-level amplifications. Genes Chromosomes Cancer. 28:145–152.
2000. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Michiels EM, Weiss MM, Hoovers JM, Baak
JP, Voûte PA, Baas F and Hermsen MA: Genetic alterations in
childhood medulloblastoma analyzed by comparative genomic
hybridization. J Pediatr Hematol Oncol. 24:205–210. 2002.
View Article : Google Scholar
|
|
53
|
Englinger B, Mair M, Miklos W, Pirker C,
Mohr T, van Schoonhoven S, Lötsch D, Körner W, Ferk F, Knasmüller
S, et al: Loss of CUL4A expression is underlying cisplatin
hypersensitivity in colorectal carcinoma cells with acquired
trabectedin resistance. Br J Cancer. 116:489–500. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Ngan ES: Heterogeneity of neuroblastoma.
Oncoscience. 2:837–838. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Louis CU and Shohet JM: Neuroblastoma:
Molecular pathogenesis and therapy. Annu Rev Med. 66:49–63. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Nader JH, Bourgeois F, Bagatell R, Moreno
L, Pearson ADJ and DuBois SG: Systematic review of clinical drug
development activities for neuroblastoma from 2011 to 2020. Pediatr
Blood Cancer. 70:e301062023. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Bedrnicek J, Vicha A, Jarosova M,
Holzerova M, Cinatl J, Michaelis M, Cinatl J and Eckschlager T:
Characterization of drug-resistant neuroblastoma cell lines by
comparative genomic hybridization. Neoplasma. 52:415–419. 2005.
|
|
58
|
Procházka P, Libra A, Zemanová Z,
Hřebačková J, Poljaková J, Hraběta J, Bunček M, Stiborová M and
Eckschlager T: Mechanisms of ellipticine-mediated resistance in
UKF-NB-4 neuroblastoma cells. Cancer Sci. 103:334–341. 2012.
View Article : Google Scholar
|
|
59
|
Bachetti T, Di Paolo D, Di Lascio S,
Mirisola V, Brignole C, Bellotti M, Caffa I, Ferraris C, Fiore M,
Fornasari D, et al: PHOX2B-mediated regulation of ALK expression:
In vitro identification of a functional relationship between two
genes involved in neuroblastoma. PLoS One. 5:e131082010. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Pfaffl MW, Horgan GW and Dempfle L:
Relative expression software tool (REST) for group-wise comparison
and statistical analysis of relative expression results in
real-time PCR. Nucleic Acids Res. 30:e362002. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Kocak H, Ackermann S, Hero B, Kahlert Y,
Oberthuer A, Juraeva D, Roels F, Theissen J, Westermann F, Deubzer
H, et al: Hox-C9 activates the intrinsic pathway of apoptosis and
is associated with spontaneous regression in neuroblastoma. Cell
Death Dis. 4:e5862013. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Su Z, Fang H, Hong H, Shi L, Zhang W,
Zhang W, Zhang Y, Dong Z, Lancashire LJ, Bessarabova M, et al: An
investigation of biomarkers derived from legacy microarray data for
their utility in the RNA-seq era. Genome Biol. 15:5232014.
View Article : Google Scholar
|
|
63
|
Oberthuer A, Berthold F, Warnat P, Hero B,
Kahlert Y, Spitz R, Ernestus K, König R, Haas S, Eils R, et al:
Customized oligonucleotide microarray gene expression-based
classification of neuroblastoma patients outperforms current
clinical risk stratification. J Clin Oncol. 24:5070–5078. 2006.
View Article : Google Scholar
|
|
64
|
Wysocka J, Swigut T, Xiao H, Milne TA,
Kwon SY, Landry J, Kauer M, Tackett AJ, Chait BT, Badenhorst P, et
al: A PHD finger of NURF couples histone H3 lysine 4 trimethylation
with chromatin remodelling. Nature. 442:86–90. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Li S, Shen L and Chen KN: Association
between H3K4 methylation and cancer prognosis: A meta-analysis.
Thoracic Cancer. 9:794–799. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Tumber A, Nuzzi A, Hookway ES, Hatch SB,
Velupillai S, Johansson C, Kawamura A, Savitsky P, Yapp C,
Szykowska A, et al: Potent and selective KDM5 inhibitor stops
cellular demethylation of H3K4me3 at transcription start sites and
proliferation of MM1S myeloma cells. Cell Chem Biol. 24:371–380.
2017. View Article : Google Scholar
|
|
67
|
Brodeur GM, Pritchard J, Berthold F,
Carlsen NL, Castel V, Castelberry RP, De Bernardi B, Evans AE,
Favrot M, Hedborg F, et al: Revisions of the international criteria
for neuroblastoma diagnosis, staging and response to treatment. J
Clin Oncol. 11:1466–1477. 1993. View Article : Google Scholar
|
|
68
|
Cheung NK, Kushner BH, LaQuaglia M, Kramer
K, Gollamudi S, Heller G, Gerald W, Yeh S, Finn R, Larson SM, et
al: N7: A novel multi-modality therapy of high-risk neuroblastoma
in children diagnosed over 1 year of age. Med Pediatr Oncol.
36:227–230. 2001. View Article : Google Scholar
|
|
69
|
Haupt S, Caramia F, Klein SL, Rubin JB and
Haupt Y: Sex disparities matter in cancer development and therapy.
Nat Rev Cancer. 21:393–407. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Hollows R, Wei W, Cazier JB, Mehanna H,
Parry G, Halford G and Murray P: Association between loss of Y
chromosome and poor prognosis in male head and neck squamous cell
carcinoma. Head Neck. 41:993–1006. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Huang RS, Kistner EO, Bleibel WK, Shukla
SJ and Dolan ME: Effect of population and gender on
chemotherapeutic agent-induced cytotoxicity. Mol Cancer Ther.
6:31–36. 2007. View Article : Google Scholar : PubMed/NCBI
|