Neuroblastoma (NB) ranks among the most prevalent
malignant tumors in children, accounting for 10-15% of
cancer-related deaths in pediatric patients (1). Notably, NB exhibits marked
heterogeneity in terms of pathology, genetic alterations,
biological behavior and clinical presentation. While the
pathogenesis of NB remains complex, recent studies have highlighted
the critical role of the MYCN gene in both its initiation and
progression (2). The treatment of
NB typically involves surgery, radiotherapy and chemotherapy;
however, therapeutic responses vary substantially among patients
owing to the intrinsic heterogeneity of the tumor. Although
targeted therapies and immunotherapy have been incorporated into
clinical practice, the overall treatment outcomes for patients with
NB continue to be less than optimal (3). Consequently, a better understanding of
NB biology and the identification of novel molecular targets are
urgently needed to improve therapeutic efficacy.
Long non-coding RNAs (lncRNAs) are RNA molecules
>200 nucleotides long that function as independent
transcriptional units without protein-coding potential. In contrast
to protein-coding genes (PCGs), lncRNAs, along with their
protein-binding partners, perform intrinsic functions. These
molecules serve a critical role in regulating various cellular
processes, including chromatin structure, transcription, RNA
stability, RNA processing, protein synthesis and RNA/protein
modifications (4). Moreover,
lncRNAs can function as competitive endogenous RNAs (ceRNAs) and
act as molecular scaffolds for protein complexes. A growing body of
evidence has indicated that lncRNAs are frequently dysregulated
during tumorigenesis, where they influence both the progression of
tumors and their associated outcomes. In NB, aberrant lncRNA
expression has been closely associated with tumor initiation,
development, metastasis and prognosis, underscoring the potential
value for lncRNAs as diagnostic biomarkers and therapeutic targets
in clinical settings (5).
NB exhibits considerable variability in its
pathology, biology, genetics and clinical characteristics among
different patients (1–4). Unlike a number of other malignancies,
some cases of NB can undergo spontaneous regression or
differentiate into benign ganglioneuromas, whereas others display
highly aggressive behavior, characterized by rapid disease
progression or resistance to multimodal therapeutic regimens
(5–9). NB arises from the sympathetic nervous
system, specifically from sympathetic adrenal progenitor cells,
which possess the capacity to differentiate into adrenal chromaffin
cells as well as sympathetic ganglion cells (10–15).
The transition to NB cells is influenced by multiple
factors, including nerve growth factor, MYCN upregulation, SRY-box
transcription factor 10 (SOX10) and mammalian achaete-scute homolog
1 genes, anaplastic lymphoma kinase (ALK) mutations and MYCN
amplification. Transcription factors, such as SOX11, twist family
bHLH transcription factor 1 and achaete-scute complex-like 1, also
serve critical roles in the onset and progression of NB. The
amplification of MYCN leads to increased levels of N-Myc protein,
which has a key role in NB pathogenesis (6,7).
In addition, ALK expression is associated with a
poor prognosis and, in synergy with MYCN, accelerates the growth of
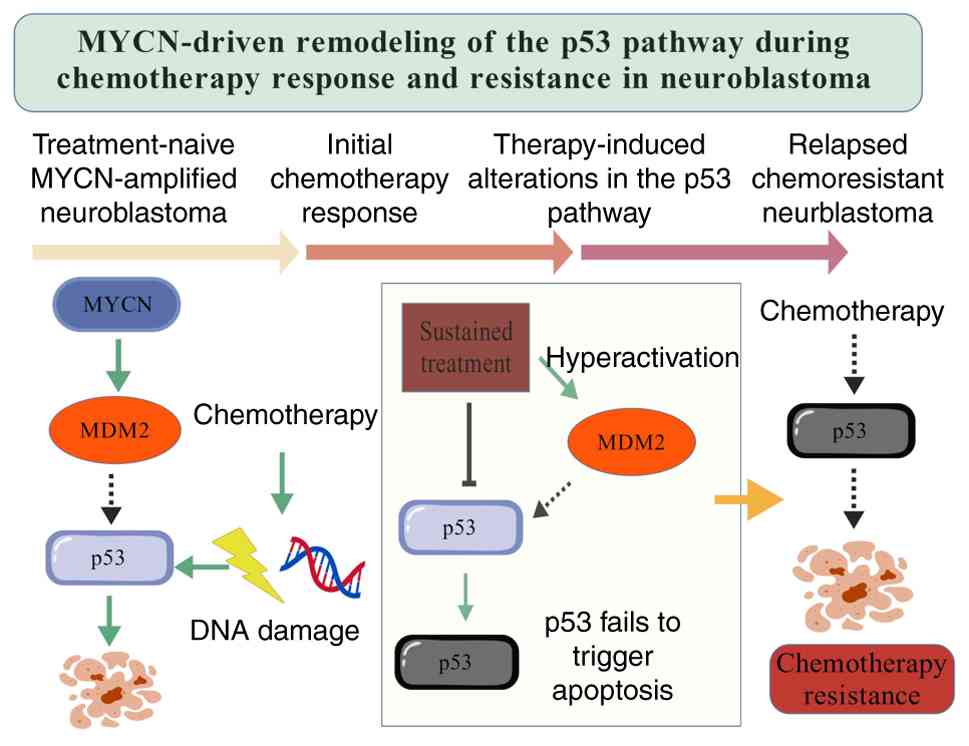
NB (16–22). As shown in Fig. 1, MYCN-driven oncogenic signaling
reshapes the p53 pathway during chemotherapy, contributing to an
initial treatment response followed by impaired p53-mediated
apoptosis and eventual chemoresistance, providing a mechanistic
context in which MYCN-ALK cooperation promotes aggressive disease
progression. Gaining a more profound understanding of these genes
and their underlying mechanisms is essential for the development of
targeted therapeutic strategies.
NB is treated using a risk-stratified approach,
similar to other types of cancer. Patients with low-risk NB
generally receive minimal treatment, with some achieving a cure
through surgery alone or undergoing spontaneous tumor regression.
Individuals classified as intermediate-risk typically receive
moderate-intensity chemotherapy in combination with surgical
resection. By contrast, high-risk NB is most commonly treated with
chemotherapy and radiotherapy (8).
The primary goal of chemotherapy is to reduce tumor
burden and control metastatic disease, commonly using agents such
as platinum compounds, etoposide and cyclophosphamide (23–28).
Additionally, immunotherapy is being investigated as a potential
treatment option for NB. While chemotherapy proves effective for
some patients, approximately one-third of high-risk children with
NB either do not respond adequately to it or face relapse (9). The 5-year survival rate for high-risk
NB remains at <50%, with outcomes ranging from complete
remission to drug resistance and severe toxicity (29–36).
Given the pronounced biological and clinical heterogeneity of NB,
the development of a universally effective treatment strategy
remains a major challenge (37–39).
The limitations of current therapies, primarily their lack of
targeting specificity, highlight the need for further research into
NB tumor biology and the development of novel therapeutic
approaches.
lncRNAs are RNA molecules >200 nucleotides long,
serving as independent transcription units that do not code for
proteins. With the advancement of high-throughput sequencing
technologies, research into lncRNAs genes has increased
substantially (40–44). The human genome is now annotated
with >20,000 lncRNA genes, a number comparable to that of PCGs.
While lncRNAs lack the ability to encode proteins, they exhibit
behaviors in biological processes that resemble those of mRNA. The
majority of lncRNAs are transcribed by RNA polymerase II and
undergo maturation through processes such as 5′capping, 3′end
cleavage, polyadenylation and splicing (45–49).
lncRNAs exhibit distinct and highly regulated
expression patterns. In contrast to mRNAs, only a limited number of
lncRNAs are expressed ubiquitously, whereas the majority show
tissue-specific or condition-dependent expression, highlighting
their functional significance. Rather than acting as templates for
protein synthesis, lncRNAs and their associated protein-binding
partners serve intrinsic biological roles (50–56).
These molecules are involved in the regulation of chromatin
organization, transcriptional control, RNA stability and
processing, protein synthesis and RNA/protein modifications, and
can also function as ceRNAs and molecular scaffolds. The mechanisms
through which lncRNAs exert their functions are diverse, including:
i) Chromatin regulation: lncRNAs interact with chromatin-modifying
enzymes (such as histone methyltransferases and
acetyltransferases), which influence gene expression by remodeling
chromatin. For example, certain lncRNAs recruit chromatin complexes
to either silence or activate target genes (57–59).
ii) Transcriptional regulation, in which lncRNAs directly interact
with transcription factors or RNA polymerase II to modulate gene
transcription (60–62). iii) RNA stability and processing:
lncRNAs regulate the stability of RNA by preventing its degradation
or facilitating the degradation of specific mRNAs. Additionally,
they serve a role in RNA splicing and post-transcriptional
modifications, such as capping and polyadenylation (63–65).
iv) Protein synthesis and modification, whereby certain lncRNAs
regulate translation through interactions with ribosomes or
specific proteins, and may also affect protein folding, trafficking
or post-translational modification (66–68).
v) ceRNA function: lncRNAs modulate gene expression by sequestering
microRNAs (miRNAs/miRs), which reduces their inhibitory effects on
target mRNAs. In this ceRNA mechanism, lncRNAs compete with other
RNA molecules for miRNAs, thereby influencing gene regulation
(69–71) (Table
I).
lncRNAs are frequently dysregulated during
tumorigenesis, where they act both as drivers and consequences of
tumor development (72–75). Oncogenic lncRNAs are often
upregulated in tumor tissues compared with in normal tissues, and
their silencing frequently suppresses tumor growth or induces
apoptosis. By contrast, lncRNAs with tumor-suppressive functions
are typically downregulated in cancer, and their depletion tends to
promote tumorigenesis. For example, activation of the lncRNA EPIC1
enhances tumorigenesis by modulating MYC target genes (40), whereas silencing of the lncRNA GAS5
accelerates cancer cell proliferation and facilitates tumor
development, highlighting its tumor-suppressive role. lncRNAs are
implicated not only in tumor initiation and progression, but also
in metastasis and prognosis. For example, lung cancer-associated
transcript 1 (LUCAT1) is markedly upregulated in both lung and
esophageal cancer, contributing to tumor progression, and higher
levels of LUCAT1 are associated with a poor prognosis (73). Collectively, these findings
underscore the pivotal roles of lncRNAs in tumor initiation,
progression and metastasis, supporting their potential utility as
clinically relevant biomarkers and therapeutic targets.
Accumulating evidence has indicated that lncRNAs
serve critical roles in shaping the malignant phenotype of NB,
influencing processes such as proliferation, migration, invasion,
epithelial-mesenchymal transition (EMT) and programmed cell death
(PCD). These effects are mediated through interactions with
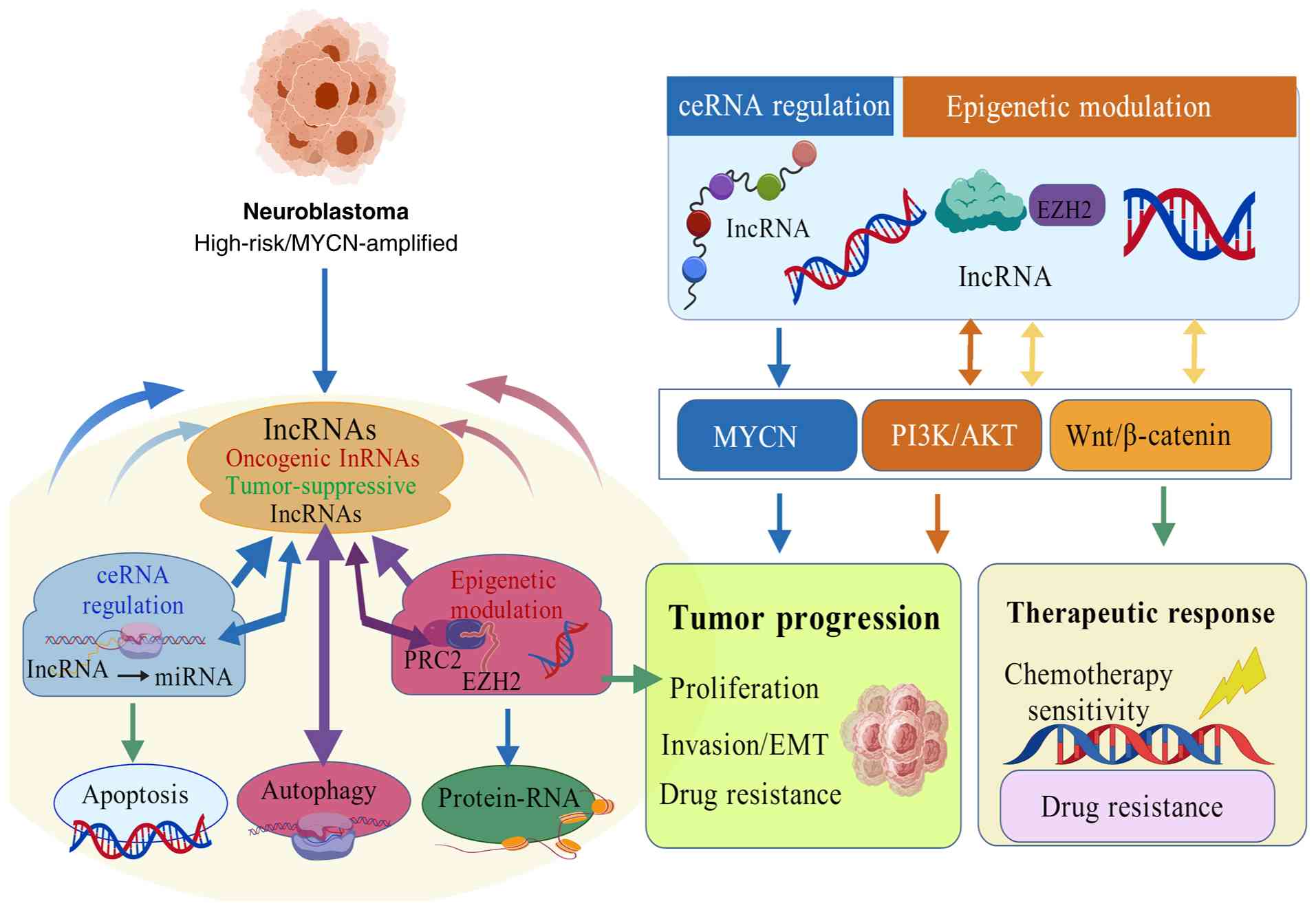
specific genes. As illustrated in Fig.
2, lncRNAs function through diverse molecular mechanisms,
including ceRNA regulation and epigenetic modulation, thereby
regulating key oncogenic pathways such as MYCN, PI3K/AKT and
Wnt/β-catenin signaling. The present review summarizes current
advances in understanding the involvement of lncRNAs in NB
pathogenesis, with an emphasis on their molecular mechanisms and
functional consequences.
Cell proliferation and invasion are essential
drivers of NB progression. Previous studies have identified several
lncRNAs that serve a role in regulating these processes,
contributing to the initiation and advancement of the tumor. For
example, Zhao et al (76)
demonstrated that the increased expression of the lncRNA zinc
finger protein 674 antisense RNA 1 (ZNF674-AS1) is associated with
poor prognosis and high-risk NB. In vivo experiments
revealed that targeting ZNF674-AS1 expression in NB cells
suppressed tumor growth. Mechanistically, ZNF674-AS1 was shown to
bind to the RNA-binding protein insulin-like growth factor 2
mRNA-binding protein 3, enhancing the stability of carbonic
anhydrase IX (CA9) mRNA, which leads to higher CA9 expression,
thereby promoting both the proliferation and invasion of NB cells
(76). In a similar study, Hsu
et al (54) identified the
lncRNA small nucleolar RNA host gene (SNHG)1, noting that its high
expression is associated with a poor prognosis in NB. Silencing
SNHG1 inhibited cell proliferation and invasion, and subsequent
experiments indicated that the interaction between SNHG1 and
HDAC1/2 modulates chromatin, thereby facilitating tumor progression
(54).
Further research has highlighted the critical role
of long intergenic non-protein coding RNA 1296 (LINC01296) in NB.
Compared with in normal tissues, LINC01296 and tripartite motif
containing 59 (TRIM59) have been found to be elevated in NB
tissues, whereas the levels of miR-584-5p and miR-34a-5p are
reduced. Depletion of LINC01296 was shown to suppress both cell
proliferation and invasion in NB cells by functioning as a sponge
for miR-584-5p and miR-34a-5p, which subsequently regulates TRIM59
expression (77). Unlike
traditional studies that focus on the interactions between lncRNAs
and other molecules to regulate tumor proliferation, Vaid et
al (81) focused on the
modification of lncRNAs. Their findings revealed that the lncRNA
telomeric repeat-containing RNA (TERRA) interacts with the telomere
region, modulating its structure and function, thus facilitating
tumor progression. By facilitating m6A methylation-dependent R-loop
formation, methyltransferase-like 3 can drive TERRA to target
telomeres, enhancing tumor cell proliferation and invasion
(78). These findings collectively
suggest that lncRNAs contribute to cancer progression through
interactions with proteins or RNAs, as detailed in Table II (54,76–78).
However, while these studies identify lncRNAs that promote tumor
proliferation and invasion, and describe their interactions with
specific proteins or RNAs, they lack comprehensive evidence
regarding the mechanisms through which these molecules regulate
cell proliferation and invasion.
lncRNAs are found in various forms, with some
promoting tumor growth and others inhibiting it. For example, Pan
et al (94) observed a
notable reduction in nuclear paraspeckle assembly transcript 1
(NEAT1) expression in NB tissues and cell lines, whereas
overexpression of NEAT1 inhibited cell proliferation and invasion.
Furthermore, NEAT1 was shown to interact directly with miR-183-5p,
negatively regulating its expression in NB. Notably, miR-183-5p
targets the 3′untranslated region (3′UTR) of forkhead box P1
(FOXP1) mRNA, reducing FOXP1 expression, and influencing cell
proliferation and invasion. Additionally, FOXP1 antagonizes the
effect of miR-183-5p on ERK/AKT phosphorylation, and FOXP1 small
interfering RNA (siRNA) further enhances the reduction of ERK/AKT
phosphorylation caused by miR-183-5p inhibitors in NB cells. This
previous study progressed from initially identifying NEAT1 as a
potential regulator of NB to uncovering its precise mechanistic
role.
Notably, while the majority of studies on lncRNAs in
NB focus on those that promote tumor growth, only a limited number
of studies have identified lncRNAs that inhibit tumor proliferation
and invasion, as summarized in Table
II (54,60,76–96).
EMT describes the biological process where
epithelial cells lose their polarity and the integrity of tight
junctions, adopting mesenchymal characteristics, such as altered
cell morphology, reduced adhesion and increased motility. EMT is a
key factor in the initiation and progression of tumors, driving
cellular migration and invasion, and enhancing both resistance to
treatments and metastatic potential (97–100).
During this transition, epithelial markers such as E-cadherin are
downregulated, whereas mesenchymal markers such as N-cadherin and
Vimentin are upregulated. This transition enables tumor cells to
detach from the primary tumor site, invade the bloodstream and
spread to distant organs, thus facilitating metastasis. EMT has
been strongly linked to the invasiveness and metastatic potential
of NB, influencing tumor progression and treatment outcomes
(101–106). Understanding the role of EMT in NB
is, therefore, essential for determining its biological behavior
and developing novel therapeutic approaches. However, the
mechanisms by which lncRNAs regulate EMT in NB cells remain
insufficiently explored.
In conclusion, although EMT is essential for tumor
development, the role of lncRNAs in regulating EMT in NB remains
under-researched. This represents an important direction for future
studies.
PCD is a process in which cells undergo controlled,
predetermined death via signaling pathways, which serves an
essential role in maintaining both physiological functions and
development. PCD comprises various forms, including apoptosis,
autophagy, ferroptosis, pyroptosis and programmed necrosis
(109–111). In the context of tumors, PCD is
crucial for their initiation and progression, as tumor cells often
evade immune detection and therapeutic interventions by suppressing
PCD, which in turn supports their survival, proliferation and
metastatic capabilities. Therefore, PCD not only contributes to
normal development and tissue repair, but also serves as an
important therapeutic target in cancer treatment (112–115). In NB, the regulation of PCD has a
notable impact on both tumor growth and metastasis. Research has
shown that NB cells can bypass normal cell cycle regulation by
modulating PCD pathways, thereby increasing the invasiveness and
resistance to treatment of the tumor (116–120). This highlights the importance of
investigating PCD in NB as it could be key to developing novel
treatment strategies. For example, Li et al (121) revealed that the lncRNA KCNQ1
opposite chain/antisense transcript 1 could regulate miR-296-5p.
The interaction between these two molecules was confirmed using RNA
immunoprecipitation and biotin precipitation assays, with
miR-296-5p inhibiting NB cell apoptosis in vitro and in
vivo. On the mechanistic level, miR-296-5p binds directly to
the 3′UTR of Bax mRNA, leading to suppression of Bax expression at
both mRNA and protein levels (121). Similarly, Yu et al
(122) revealed that elevated
expression of SNHG16 is associated with poor clinical outcomes. By
contrast, silencing SNHG16 can enhance apoptosis (122). Focusing on autophagy, Ye et
al (107) discovered that MEG3
markedly inhibits cell proliferation, migration and invasion.
Chromatin isolation by RNA purification analysis further suggested
that the anticancer effect of MEG3 is associated with autophagy and
the mTOR signaling pathway. Through LC3 fluorescence imaging and
western blotting, it was shown that MEG3 reduces autophagy by
inhibiting FOXO1 expression, without affecting the mTOR pathway.
Mechanistically, MEG3 and EZH2 interact through a negative feedback
loop to modulate autophagy in NB (107). Research into the role of lncRNAs
in PCD is still limited. Although necroptosis, ferroptosis and
necrosis have been linked to NB development (108–128), to the best of our knowledge, no
studies have identified specific lncRNAs that regulate these
processes to influence tumor initiation and progression; this
represents a promising area for future research.
Tumor resistance refers to the ability of tumor
cells to develop resistance to treatments such as chemotherapy,
immunotherapy and targeted therapies through various biological
mechanisms, which ultimately leads to reduced or completely
ineffective treatment outcomes (129–132). This phenomenon has a central role
in tumor progression and recurrence, as tumor cells escape
therapeutic interventions by either adapting to treatment pressure
or modifying their inherent characteristics, which eventually
results in clinical treatment failure. NB is primarily managed with
chemotherapy, targeted therapies and immunotherapy. Despite
chemotherapy being the cornerstone for high-risk NB treatment, its
effectiveness is often limited by its toxic effects on normal cells
and the emergence of tumor resistance. Targeted therapies aim to
inhibit key tumor-driving genes, such as MYCN and ALK, but
resistance remains a persistent challenge in clinical settings.
Although immunotherapy has demonstrated encouraging results in a
subset of patients, its overall success is still hindered by the
ability of the tumor to evade immune detection (133–140). Consequently, tumor resistance
substantially diminishes treatment efficacy, particularly in cases
of recurrent or metastatic NB, emphasizing the urgent need for
innovative therapeutic strategies. Zhao et al (76) reported that the lncRNA ZNF674-AS1
contributes to NB cell proliferation by upregulating CA9 and
preventing cisplatin-induced pyroptosis, which in turn leads to
cisplatin resistance. Similarly, Xiang et al (141) demonstrated that NUTM2A-AS1
expression in cisplatin-resistant NB cells increases in a manner
that is both time- and dose-dependent. By contrast, silencing
NUTM2A-AS1 was shown to boost cisplatin sensitivity and inhibit
metastatic behavior in NB cells. Additionally, the immune
checkpoint protein B7-H3 was identified as a target of NUTM2A-AS1
in these cells, with NUTM2A-AS1 reducing the degradation of B7-H3
(141). In another study, Wang
et al (79) discovered that
the lncRNA non-coding RNA activated by DNA damage targets
miR-144-3p to upregulate HDAC8, thereby accelerating NB progression
and contributing to resistance against doxorubicin. Tan et
al (95) explored the impact of
taurine upregulated gene 1 (TUG1) overexpression on tumor
immunotherapy, revealing that TUG1 knockdown markedly inhibited NB
cell proliferation, colony formation and migration when compared
with cytokine-induced killer (CIK) or dendritic cells co-cultured
with CIK (DC-CIK) therapies alone. Furthermore, TUG1 upregulation
was shown to strongly induce apoptosis and alter key molecules
involved in apoptosis and EMT. After transfection with TUG1, the
concentrations of IL-12, IL-2 and IFN-γ in the co-culture
supernatant were markedly elevated. These findings suggest that
TUG1 upregulation potentiates the antitumor effects of DC-CIK
immunotherapy (95). Although
previous reports have indicated that the knockdown of TUG1 can
inhibit tumor cell proliferation, colony formation and migration,
this study further demonstrated that TUG1 upregulation could
enhance the antitumor efficacy of DC-CIK immunotherapy, exerting a
synergistic tumor-suppressive effect through a gain-of-function
mechanism.
MYCN amplification is a hallmark of high-risk NB and
represents one of the most critical oncogenic drivers governing
tumor cell proliferation, metabolic reprogramming and resistance to
therapy. Beyond genomic amplification, MYCN expression and activity
are tightly regulated at transcriptional, post-transcriptional and
epigenetic levels, with lncRNAs emerging as key regulatory
components.
Several lncRNAs have been shown to directly or
indirectly modulate MYCN signaling. For example, MIAT acts as an
upstream regulator of N-Myc, and disruption of the MIAT/MYCN axis
induces cell death in MYCN-amplified NB cells, highlighting its
essential role in maintaining oncogenic MYCN activity (82). In addition, AC142119.1
epigenetically activates MYCN transcription by interacting with the
WDR5 protein and facilitating chromatin remodeling at the MYCN
promoter region, thus sustaining MYCN-driven transcriptional
programs (85).
Collectively, these findings suggest that lncRNAs
function as critical modulators of MYCN signaling by either
stabilizing MYCN expression or reshaping the epigenetic landscape
of MYCN target genes. Targeting MYCN-associated lncRNAs may thus
represent an alternative strategy to indirectly suppress MYCN
oncogenic activity in NB.
The PI3K/AKT signaling pathway serves a pivotal role
in NB cell survival, proliferation, metabolism and chemoresistance.
Aberrant activation of this pathway is frequently observed in
aggressive NB and is associated with poor prognosis (10). Emerging evidence has indicated that
lncRNAs can modulate PI3K/AKT signaling through ceRNA mechanisms
and transcriptional regulation.
For example, NEAT1 negatively regulates NB cell
proliferation and migration by suppressing ERK/AKT phosphorylation
via the miR-183-5p/FOXP1 axis. In this regulatory cascade, NEAT1
functions as a ceRNA that sequesters miR-183-5p, thereby restoring
FOXP1 expression and attenuating downstream AKT signaling (94). Similarly, other lncRNAs indirectly
influence PI3K/AKT signaling by regulating upstream growth factor
receptors or metabolic regulators.
These observations underscore that lncRNAs do not
act as isolated regulators but rather fine-tune PI3K/AKT signaling
outputs by integrating miRNA networks and transcriptional control,
ultimately shaping NB cell fate decisions.
The p53 signaling pathway is a central mediator of
DNA damage response and PCD, and its dysregulation is closely
linked to chemotherapy resistance and relapse in high-risk NB.
Although TP53 mutations are relatively rare at diagnosis,
functional inactivation of p53 signaling frequently occurs during
disease progression, where lncRNAs appear to serve contributory
roles (11).
These findings suggest that lncRNAs may contribute
to NB drug resistance by attenuating p53-dependent cell death
pathways, thereby facilitating tumor persistence and relapse.
Targeting p53-related lncRNAs networks could enhance therapeutic
responsiveness in refractory NB.
The Wnt/β-catenin signaling pathway is a key driver
of tumor invasion, metastasis and EMT in NB. Aberrant activation of
this pathway promotes tumor cell motility and invasive behavior,
contributing to disease progression and poor clinical outcomes
(13).
Several lncRNAs have been implicated in regulating
Wnt/β-catenin signaling in NB. Notably, double homeobox A
pseudogene 8 facilitates NB progression by activating the
Wnt/β-catenin pathway through the miR-29/nucleolar protein 4 like
axis (84). This activation
enhances EMT-associated phenotypes and increases the invasive
capacity of tumor cells. Through miRNA sponging and downstream
target activation, lncRNAs effectively reprogram Wnt/β-catenin
signaling networks.
Overall, lncRNAs participate in NB progression not
merely as individual regulators but as integral components of key
oncogenic signaling networks. By modulating MYCN, PI3K/AKT, p53 and
Wnt/β-catenin pathways, lncRNAs orchestrate tumor growth, survival,
metastasis and therapeutic resistance, highlighting their potential
as novel biomarkers and therapeutic targets in NB.
lncRNAs serve an essential role in the initiation,
progression and prognosis of NB. Research has shown that lncRNAs
exhibit distinct expression patterns in NB, where they regulate
various genes and signaling pathways involved in processes such as
cell proliferation, migration, PCD and resistance to chemotherapy.
These regulatory activities markedly contribute to tumor growth,
invasion and metastasis. The expression levels of specific lncRNAs
are closely associated with the malignancy, prognosis and
therapeutic response of NB, positioning them as promising
biomarkers for predicting clinical outcomes and treatment
effectiveness.
To provide a clearer overview of the prognostic and
diagnostic relevance of lncRNAs in NB, representative lncRNAs with
reported clinical significance are summarized in Table III (97–106),
including their expression patterns, molecular targets or signaling
pathways, associated clinical outcomes and validation status.
Notably, although a growing number of lncRNAs have
been associated with NB prognosis, only a subset has been validated
using patient-derived samples or independent clinical cohorts,
whereas a number of findings remain based primarily on
retrospective datasets or experimental models.
Furthermore, RNA interference (RNAi) targeting of
lncRNAs provides a potential therapeutic approach for NB. RNAi
utilizes exogenous double-stranded RNA to degrade target RNA, and
several studies (142–144) have utilized lncRNA short hairpin
RNAs (shRNAs) delivered via viral infection for treatment purposes
(144). Although no drugs
targeting lncRNAs have been developed for NB, some drugs (such as
ribociclib) have been found to modulate its progression by
interacting with lncRNAs (145,146). Taken together, these findings
highlight the translational potential of lncRNAs as biomarkers for
early diagnosis, prognostic stratification and therapeutic response
prediction in NB, while also underscoring the need for further
large-scale clinical validation.
Although lncRNAs represent promising diagnostic
biomarkers and therapeutic targets in NB, their translation into
routine clinical practice remains challenging (76). Several critical barriers must be
addressed before lncRNA-based therapies can be safely and
effectively applied in patients.
One of the primary challenges in lncRNA-targeted
therapy lies in efficient and tumor-specific delivery. lncRNA-based
therapeutics, including antisense oligonucleotides (ASOs), siRNAs
and shRNAs, are inherently susceptible to enzymatic degradation in
circulation and often exhibit limited stability in vivo.
Moreover, achieving efficient intracellular delivery, particularly
nuclear localization for functionally relevant lncRNAs, remains a
notable hurdle (147).
In addition, the lack of tumor-specific delivery
systems may lead to insufficient accumulation at tumor sites and
unintended uptake by normal tissues. This issue is particularly
relevant in NB, where systemic administration of nucleic acid-based
therapeutics may result in off-target distribution to healthy
organs, increasing the risk of adverse effects (147). Although viral vectors and
nanoparticle-based delivery systems have been explored to improve
delivery efficiency (14), concerns
regarding immunogenicity, biodistribution and long-term safety
persist.
Another major limitation arises from the complex and
context-dependent functions of lncRNAs. Numerous lncRNAs exhibit
pleiotropic regulatory roles and may participate in multiple
signaling pathways across different cell types. Consequently,
therapeutic modulation of a single lncRNA may unintentionally
disrupt normal cellular processes, leading to off-target effects
(4).
Furthermore, the tissue-specific and developmental
stage-specific expression patterns of lncRNAs complicate
therapeutic targeting. In pediatric tumors such as NB, unintended
interference with normal developmental programs represents a
particularly important safety concern (54). These factors collectively underscore
the need for precise targeting strategies and comprehensive
functional validation before clinical application.
Despite encouraging preclinical data, no
lncRNA-targeted therapies have yet been approved for the treatment
of NB. Safety concerns remain a major obstacle to clinical
translation. Potential risks include immune activation, insertional
mutagenesis (in the case of viral delivery) and long-term toxicity
resulting from sustained lncRNAs modulation (9).
To overcome these challenges, future efforts should
focus on the development of safer and more efficient delivery
systems, such as chemically modified oligonucleotides,
ligand-targeted nanoparticles and tumor-specific promoters. In
parallel, comprehensive toxicological evaluations and longitudinal
studies are required to assess long-term safety. An improved
understanding of lncRNA biology and context-dependent functions
will be essential for translating lncRNA-based strategies into
clinically viable therapies for NB.
A growing body of evidence underscores the pivotal
roles of lncRNAs in tumor initiation and progression. Aberrant
lncRNA expression has been closely associated with key oncogenic
processes, including DNA damage responses, immune evasion and
metabolic dysregulation in cancer cells. Depending on their
biological context, specific lncRNAs may function as oncogenes or
tumor suppressors, and their expression levels are associated with
clinical outcomes, including disease progression and overall
survival in patients with NB.
Furthermore, an increasing number of therapeutic
agents have been reported to modulate NB cell proliferation,
progression and drug resistance, at least in part through
lncRNA-associated regulatory pathways. Collectively, these findings
highlight lncRNAs as promising molecular targets for NB prognosis,
diagnosis and treatment. Targeted modulation of lncRNA expression
may be achieved through multiple strategies, including ASOs, RNAi,
CRISPR/Cas9-based genome editing, viral delivery of lncRNA and
small-molecule inhibitors. Taken together, lncRNAs represent a
highly promising class of therapeutic targets for the development
of molecularly guided and precision-based treatments for NB.
Not applicable.
Funding: No funding was received.
Not applicable.
ZX was responsible for the original draft
preparation, research design and methodology development, and
participated in the review and editing of the manuscript. ZX made
the following specific contributions to the study:
Conceptualization of the topic, core framework and chapter
structure design of the review; developing the systematic strategy
for literature searching, screening and data extraction; and
independently drafting the full initial manuscript along with the
figures and tables. HZ and LZ jointly reviewed and revised the
paper. CW provided supervision and guidance throughout the research
process. CW made decisive contributions to the conceptual framework
design of the study, the interpretation of key conclusions and the
revision of the final manuscript. Data authentication is not
applicable. All authors commented on previous versions of the
manuscript, and read and approved the final manuscript.
Not applicable.
Not applicable.
The authors declare that they have no competing
interests.
|
1
|
Alexander F: Neuroblastoma. Urol Clin
North Am. 27383–392. (vii)2000. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Quamine AE, Dray EL, Mohrdieck NR, King
CA, Griggs AA, Kline JM, Cho MM, Rinella SP, Tippins KE, Bates PD,
et al: TIM-3 blockade enhances ex vivo stimulated allogeneic NK
cell therapy for relapsed murine neuroblastoma after hematopoietic
cell transplant. J Immunother Cancer. 13:e0102392025. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Xiang S, Chen P, Shi X, Cai H, Shen Z, Liu
L, Xu A, Zhang J, Zhang X, Bing S, et al: Disruption of the
KLHL37-N-Myc complex restores N-Myc degradation and arrests
neuroblastoma growth in mouse models. J Clin Invest.
135:e1766552025. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Körber V, Stainczyk SA, Kurilov R, Henrich
KO, Hero B, Brors B, Westermann F and Höfer T: Neuroblastoma arises
in early fetal development and its evolutionary duration predicts
outcome. Nat Genet. 55:619–630. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lunghi G, Pedroli C, Tagliabue I, Dobi D,
Ciampa MG, Mauri L, Rouvière L, Henriques A, Callizot N, Sonnino S,
et al: GM1 oligosaccharide-mediated rescue in GBA-linked
Parkinson's disease via modulation of lysosomal and mitochondrial
dysfunctions. Glycoconj J. 42:159–171. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Li CH, Sharma S, Heczey AA, Woods ML,
Steffin DHM, Louis CU, Grilley BJ, Thakkar SG, Wu M, Wang T, et al:
Long-term outcomes of GD2-directed CAR-T cell therapy in patients
with neuroblastoma. Nat Med. 31:1125–1129. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Li Q, Tian T, Geng W, Huang X, Xu X, Adeli
M, Wang X, Cheng L, Ma T, Luo H, et al: Neuroblastoma-Targeting
π-Conjugated COP nanostructure with multiple enzyme-mimetic actions
for sonochemodynamic immunotherapies. Adv Mater. 37:e25032612025.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Jansky S, Sharma AK, Körber V, Quintero A,
Toprak UH, Wecht EM, Gartlgruber M, Greco A, Chomsky E, Grünewald
TGP, et al: Single-cell transcriptomic analyses provide insights
into the developmental origins of neuroblastoma. Nat Genet.
53:683–693. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Giudice AM, Roth SL, Matlaga S,
Cresswell-Clay E, Mishra P, Schürch PM, Boateng-Antwi KAM, Samanta
M, Pascual-Pasto G, Zecchino V, et al: Reprogramming the
neuroblastoma tumor immune microenvironment to enhance GPC2 CAR T
cells. Mol Ther. 33:4552–4569. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Sainero-Alcolado L, Mushtaq M, Liaño-Pons
J, Rodriguez-Garcia A, Yuan Y, Liu T, Ruiz-Pérez MV, Schlisio S,
Bedoya-Reina O and Arsenian-Henriksson M: Expression and activation
of nuclear hormone receptors result in neuronal differentiation and
favorable prognosis in neuroblastoma. J Exp Clin Cancer Res.
41:2262022. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gawne PJ, Bryant HE, DuBois SG, George SL,
Gray J, Knox L, Matchett KB, Peet C, Vallis KA, Wallace HJ, et al:
Theranostics for neuroblastoma: Making molecular radiotherapy work
better. J Nucl Med. 66:490–496. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Langenberg KPS, van Hooff SR, Koopmans B,
Strijker JGM, Kholosy WM, Ober K, Zwijnenburg DA, van der Hoek JJF,
Keller KM, Vernooij L, et al: Exploring high-throughput drug
sensitivity testing in neuroblastoma cell lines and patient-derived
tumor organoids in the era of precision medicine. Eur J Cancer.
218:1152752025. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Hennchen M, Stubbusch J, Abarchan-El
Makhfi I, Kramer M, Deller T, Pierre-Eugene C, Janoueix-Lerosey I,
Delattre O, Ernsberger U, Schulte JB and Rohrer H: Lin28B and Let-7
in the control of sympathetic neurogenesis and neuroblastoma
development. J Neurosci. 35:16531–16544. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Hanna R, Abdallah J and Abou-Antoun T: A
novel mechanism of 17-AAG therapeutic efficacy on HSP90 inhibition
in MYCN-amplified neuroblastoma cells. Front Oncol. 10:6245602021.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Casey MJ and Stewart RA: Zebrafish as a
model to study neuroblastoma development. Cell Tissue Res.
372:223–232. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Rebernick RJ, Choi JE, Hesse C, Hosseini
N, Chu A, Zhou J, Ning Y, Wang R, Cao X, Irwin M, et al:
Chromosomal instability degrades developmental phenotypes essential
for anti-GD2 immunotherapy outcomes in high-risk neuroblastoma.
Cell Rep Med. 6:1023752025. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hagemann S, Misiak D, Bell JL, Fuchs T,
Lederer MI, Bley N, Hämmerle M, Ghazy E, Sippl W, Schulte JH and
Hüttelmaier S: IGF2BP1 induces neuroblastoma via a druggable
feedforward loop with MYCN promoting 17q oncogene expression. Mol
Cancer. 22:882023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Floros KV, Cai J, Jacob S, Kurupi R,
Fairchild CK, Shende M, Coon CM, Powell KM, Belvin BR, Hu B, et al:
MYCN-amplified neuroblastoma is addicted to iron and vulnerable to
inhibition of the system Xc-/glutathione axis. Cancer Res.
81:1896–1908. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yang Y, Zhao J, Zhang Y, Feng T, Yv B,
Wang J, Gao Y, Yin M, Tang J and Li Y: MYCN protein stability is a
better prognostic indicator in neuroblastoma. BMC Pediatr.
22:4042022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Suk Y and Singh SK: Safety and efficacy of
lorlatinib against ALK-driven refractory or relapsed neuroblastoma.
Cell Rep Med. 4:1010712023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Schmitt-Hoffner F, van Rijn S, Toprak UH,
Mauermann M, Rosemann F, Heit-Mondrzyk A, Hübner JM, Camgöz A,
Hartlieb S, Pfister SM, et al: FOXR2 stabilizes MYCN protein and
identifies Non-MYCN-amplified neuroblastoma patients with
unfavorable outcome. J Clin Oncol. 39:3217–3228. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Chen Y, Zhuo R, Sun L, Tao Y, Li G, Zhu F,
Xu Y, Wang J, Li Z, Yu J, et al: Super-enhancer-driven IRF2BP2
enhances ALK activity and promotes neuroblastoma cell
proliferation. Neuro Oncol. 26:1878–1894. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bhoopathi P, Mannangatti P, Emdad L, Das
SK and Fisher PB: The quest to develop an effective therapy for
neuroblastoma. J Cell Physiol. 236:7775–7791. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
ElHarouni D, Hernansaiz-Ballesteros R,
Peterziel H, Balasubramanian GP, Previti C, Schramm K,
Blattner-Johnson M, Kabbe R, Jones BC, Oppermann S, et al:
Integrative multiomics and drug sensitivity profiling reveal
potential biomarkers and therapeutic strategies in pediatric solid
tumors. Cancer Res. 86:773–784. 2026. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Krols S, Rishfi M, Martens F, Van
Hauwermeiren A, Sanders E, De Sutter PJ, Vermeulen A, De Wever K,
Bekaert SL, Dolman MEM, et al: Second-generation AURKA-targeting
PROTACs: Structural optimization toward in vivo degradation in
neuroblastoma. J Med Chem. 68:23962–23976. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Anderson J, Majzner RG and Sondel PM:
Immunotherapy of neuroblastoma: Facts and hopes. Clin Cancer Res.
28:3196–3206. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Shen WT, Zhang JA, Yu Y, Zhang SD, Sun L,
Kai M, Gao W and Zhang L: Dual-action cellular nanoparticles for
effective lead (Pb2+) detoxification. ACS Nano.
19:37142–37153. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Miao Y, Chen H, Li Y, Li L, Ye J, Zhang J,
Wang J, Wu H, Li G, Chen Y, et al: A small molecule selectively
targets N-Myc to suppress neuroblastoma cancer progression. Int J
Biol Sci. 21:4895–4907. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Herter S, Emperador M, Smyrilli K, Kocher
D, Celikyürekli S, Zeiser C, Gerloff X, Kreth S, Henrich KO, Maaß
KK, et al: High content-imaging drug synergy screening identifies
specific senescence-related vulnerabilities of mesenchymal
neuroblastomas. Cell Death Dis. 16:6442025. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hanssen KM, Murray J, Pandher R, Alfred S,
Gamble LD, Brand J, Mosmann E, Kusuma FK, Mak C, Kearns A, et al:
Arginine depletion potentiates standard-of-care chemo-immunotherapy
in preclinical models of high-risk neuroblastoma. J Exp Clin Cancer
Res. 44:2392025. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zadran SK, Facchinello N, De Rosa P,
Saporetti R, Costantini PE, Ulfo L, Nigro M, Petrosino A,
Pappagallo L, Aloisi S, et al: Systematic targeting of GD2-positive
neuroblastoma tumors with a photooncolytic phage nanovector
platform. Adv Sci (Weinh). 12:e153562025. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Montuori G, Tu F, Qin D, Schmargon R,
Rodriguez-Fos E, Helmsauer K, Hui H, Mandal S, Purshouse K,
Fankhänel L, et al: Extrachromosomal DNA-driven oncogene dosage
heterogeneity promotes rapid adaptation to therapy in
MYCN-amplified cancers. Cancer Discov. 15:2054–2077. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Desai AV, Applebaum MA, Karrison TG,
Oppong A, Yuan C, Berg KR, MacQuarrie K, Sokol E, Hall AG, Pinto N,
et al: Efficacy of post-induction therapy for high-risk
neuroblastoma patients with end-induction residual disease. Cancer.
128:2967–2977. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liaño-Pons J, Garde-Lapido E, Fahrig FL,
Jäckering M, Yuan Y, Andersson S, Schort L, Esteve M, Mohlin S,
Bedoya-Reina OC and Arsenian-Henriksson M: Combined targeting of
PRDX6 and GSTP1 as a potential differentiation strategy for
neuroblastoma treatment. Proc Natl Acad Sci USA.
122:e24272111222025. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Gelineau NU, Bozsaky E, van Zogchel LMJ,
Rifatbegovic F, Lazic D, Ziegler A, Javadi A, Zappeij-Kannegieter
L, Pötschger U, Fiocco M, et al: Sensitive detection of minimal
residual disease and immunotherapy targets by multi-modal bone
marrow analysis in high-risk neuroblastoma-a multi-center study. J
Exp Clin Cancer Res. 44:2242025. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pearson AD, Chi S, Laetscht TW, Marshall
L, Raetz E, George RE, Chesler L, Karres D, Scobie N, Knoderer H,
et al: Paediatric strategy forum for medicinal product development
of Cyclin-dependent kinase inhibitors in children and adolescents
ACCELERATE in collaboration with the European Medicines Agency With
participation of the Food and Drug Administration. Eur J Cancer.
226:1156292025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Lode HN, Siebert N, Valteau-Couanet D,
Garaventa A, Canete A, Anderson J, Yaniv I, Ash S, Gray J,
Klingebiel T, et al: Fcγ receptor polymorphism in patients with
Relapsed/Refractory High-risk neuroblastoma correlates with
outcomes in the SIOPEN dinutuximab beta Long-term infusion trial.
Clin Cancer Res. 31:3692–3701. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Salemi F, Alam W, Hassani MS, Hashemi SZ,
Jafari AA, Mirmoeeni SMS, Arbab M, Mortazavizadeh SMR and Khan H:
Neuroblastoma: Essential genetic pathways and current therapeutic
options. Eur J Pharmacol. 926:1750302022. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Qadir MI, Ahmed B and Noreen S: Advances
in the management of neuroblastoma. Crit Rev Eukaryot Gene Expr.
34:1–13. 2024. View Article : Google Scholar
|
|
40
|
Mondal S, Liu PY, Seneviratne J, De Weck
A, Venkat P, Mayoh C, Wu J, Maag J, Chen J, Wong M, et al: The
super Enhancer-driven long noncoding RNA PRKCQ-AS1 promotes
neuroblastoma tumorigenesis by interacting with MSI2 protein and is
targetable by small molecule compounds. Adv Sci (Weinh).
12:e24125202025. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Herman AB, Tsitsipatis D and Gorospe M:
Integrated lncRNAs function upon genomic and epigenomic regulation.
Mol Cell. 82:2252–2266. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Bridges MC, Daulagala AC and Kourtidis A:
LNCcation: LncRNAs localization and function. J Cell Biol.
220:e2020090452021. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Guerra M, Meola L, Lattante S, Conte A,
Sabatelli M, Sette C and Bernardini C: Characterization of SOD1-DT,
a divergent long non-coding RNA in the locus of the SOD1 human
gene. Cells. 12:20582023. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Grammatikakis I and Lal A: Significance of
lncRNAs abundance to function. Mamm Genome. 33:271–280. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang Y, Wang X, Hu C and Yi H: Shiny
transcriptional junk: LncRNAs-derived peptides in cancers and
immune responses. Life Sci. 316:1214342023. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Núñez-Martínez HN and Recillas-Targa F:
Emerging functions of lncRNAs loci beyond the transcript itself.
Int J Mol Sci. 23:62582022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Wang X, Guo Y, Chen G, Fang E, Wang J, Li
Q, Li D, Hu A, Bao B, Zhou Y, et al: Therapeutic targeting of FUBP3
phase separation by GATA2-AS1 inhibits malate-aspartate shuttle and
neuroblastoma progression via modulating SUZ12 activity. Oncogene.
42:2673–2687. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Senmatsu S and Hirota K: Roles of lncRNAs
transcription as a novel regulator of chromosomal function. Genes
Genet Syst. 95:213–223. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Ye M, Gao R, Chen S, Bai J, Chen J, Lu F,
Gu D, Shi X, Yu P, Tian Y, et al: FAM201A encodes small protein
NBASP to inhibit neuroblastoma progression via inactivating MAPK
pathway mediated by FABP5. Commun Biol. 6:7142023. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Srinivas T, Siqueira E and Guil S:
Techniques for investigating lncRNAs transcript functions in
neurodevelopment. Mol Psychiatry. 29:874–890. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Zhao J, Le M, Li J, Huang Q, Chen H, Zhang
W, Mao H, Sun Q, Li A, Zhao Y, et al: LINC00938 alleviates hypoxia
ischemia encephalopathy induced neonatal brain injury by regulating
oxidative stress and inhibiting JNK/p38 MAPK signaling pathway. Exp
Neurol. 367:1144492023. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhao X, Li D, Huang D, Song H, Mei H, Fang
E, Wang X, Yang F, Zheng L, Huang K and Tong Q: Retraction notice
to: Risk-associated long noncoding RNA FOXD3-AS1 inhibits
neuroblastoma progression by repressing PARP1-mediated activation
of CTCF. Mol Ther. 31:18572023. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Zhou Y, Wang X, Yao L and Zhu M:
LDAformer: predicting lncRNAs-disease associations based on
topological feature extraction and Transformer encoder. Brief
Bioinform. 23:bbac3702022. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Hsu CL, Yin CF, Chang YW, Fan YC, Lin SH,
Wu YC, Huang HC and Juan HF: LncRNA SNHG1 regulates neuroblastoma
cell fate via interactions with HDAC1/2. Cell Death Dis.
13:8092022. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Peng L, Wang C, Tian X, Zhou L and Li K:
Finding lncRNAs-Protein interactions based on deep learning with
Dual-net neural architecture. IEEE/ACM Trans Comput Biol Bioinform.
19:3456–346. 2028. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Much C, Lasda EL, Pereira IT, Vallery TK,
Ramirez D, Lewandowski JP, Dowell RD, Smallegan MJ and Rinn JL: The
temporal dynamics of lncRNAs Firre-mediated epigenetic and
transcriptional regulation. Nat Commun. 15:68212024. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Su PP, Liu DW, Zhou SJ, Chen H, Wu XM and
Liu ZS: Down-regulation of Risa improves podocyte injury by
enhancing autophagy in diabetic nephropathy. Mil Med Res.
9:232022.PubMed/NCBI
|
|
58
|
Yuan J, Zhu Q, Zhang X, Wen Z, Zhang G, Li
N, Pei Y, Wang Y, Pei S, Xu J, et al: Ezh2 competes with p53 to
license lncRNAs Neat1 transcription for inflammasome activation.
Cell Death Differ. 29:2009–2023. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Yao L, Peng P, Ding T, Yi J and Liang J:
mA-Induced lncRNAs MEG3 promotes cerebral Ischemia-reperfusion
injury via modulating oxidative stress and mitochondrial
dysfunction by hnRNPA1/Sirt2 axis. Mol Neurobiol. 61:6893–6908.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Liu X, Chen J, Zhang S, Liu X, Long X, Lan
J, Zhou M, Zheng L and Zhou J: LINC00839 promotes colorectal cancer
progression by recruiting RUVBL1/Tip60 complexes to activate NRF1.
EMBO Rep. 23:e541282022. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Zou L, Wang X and Han X: lncRNAs MALAT
1/miR-625-3p/HIF-1α axis regulates the EMT of hypoxia-induced RPE
cells by activating NF-κB/snail signaling. Exp Cell Res.
429:1136502023. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Sethi SC, Singh R, Sahay O, Barik GK and
Kalita B: Unveiling the hidden gem: A review of long non-coding RNA
NBAT-1 as an emerging tumor suppressor and prognostic biomarker in
cancer. Cell Signal. 126:1115252025. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Jing W, Tuxiu X, Xiaobing L, Guijun J,
Lulu K, Jie J, Lu Y, Liying Z, Xiaoxing X and Jingjun L: lncRNAs
GAS5/miR-137 is a Hypoxia-responsive axis involved in cardiac
arrest and cardiopulmonary cerebral resuscitation. Front Immunol.
12:7907502022. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Desideri F, Grazzi A, Lisi M, Setti A,
Santini T, Colantoni A, Proietti G, Carvelli A, Tartaglia GG,
Ballarino M and Bozzoni I: CyCoNP lncRNA establishes cis and trans
RNA-RNA interactions to supervise neuron physiology. Nucleic Acids
Res. 52:9936–9952. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Zhu SF, Yuan W, Du YL and Wang BL:
Research progress of lncRNAs and miRNA in hepatic
ischemia-reperfusion injury. Hepatobiliary Pancreat Dis Int.
22:45–53. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Zhang C, Zhou B, Gu F, Liu H, Wu H, Yao F,
Zheng H, Fu H, Chong W, Cai S, et al: Micropeptide PACMP inhibition
elicits synthetic lethal effects by decreasing CtIP and
poly(ADP-ribosyl)ation. Mol Cell. 82:1297–1312.e8. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
He X, Yang T, Lu YW, Wu G, Dai G, Ma Q,
Zhang M, Zhou H, Long T, Yan Y, et al: The long noncoding RNA
CARDINAL attenuates cardiac hypertrophy by modulating protein
translation. J Clin Invest. 134:e1691122024. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Zheng SM, Feng YC, Zhu Q, Li RQ, Yan QQ,
Teng L, Yue YM, Han MM, Ye K, Zhang SN, et al: MILIP binding to
tRNAs promotes protein synthesis to drive Triple-negative breast
cancer. Cancer Res. 84:1460–1474. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Su K, Wang N, Shao Q, Liu H, Zhao B and Ma
S: The role of a ceRNA regulatory network based on lncRNAs MALAT1
site in cancer progression. Biomed Pharmacother. 137:1113892021.
View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Chen L, Wei K, Li J, Li Y, Cao H and Zheng
Z: Integrated analysis of lncRNAs-mediated ceRNA network in
calcific aortic valve disease. Cells. 11:22042022. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Bazrgar M, Mirmotalebisohi SA, Ahmadi M,
Azimi P, Dargahi L, Zali H and Ahmadiani A: Comprehensive analysis
of lncRNAs-associated ceRNA network reveals novel potential
prognostic regulatory axes in glioblastoma multiforme. J Cell Mol
Med. 28:e183922024. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Fang D, Ou X, Sun K, Zhou X, Li Y, Shi P,
Zhao Z, He Y, Peng J and Xu J: m6A modification-mediated lncRNAs
TP53TG1 inhibits gastric cancer progression by regulating CIP2A
stability. Cancer Sci. 113:4135–4150. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Liu Y, Zhang J, Cao F, Dong X, Li J, Cao
Y, Li Z, Guo Y, Yan J, Liu Y and Zhao Q:
N6-methyladenosine-mediated overexpression of long noncoding RNA
ADAMTS9-AS2 triggers neuroblastoma differentiation via regulating
LIN28B/let-7/MYCN signaling. JCI Insight. 8:e1657032023. View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Wang J, Su Z, Lu S, Fu W, Liu Z, Jiang X
and Tai S: lncRNAs HOXA-AS2 and its molecular mechanisms in human
cancer. Clin Chim Acta. 485:229–233. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Yao ZT, Yang YM, Sun MM, He Y, Liao L,
Chen KS and Li B: New insights into the interplay between long
non-coding RNAs and RNA-binding proteins in cancer. Cancer Commun
(Lond). 42:117–140. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Zhao K, Wang X, Jin Y, Zhu X, Zhou T, Yu
Y, Ji X, Chang Y, Luo J, Ni X, et al: LncRNA ZNF674-AS1 drives cell
growth and inhibits cisplatin-induced pyroptosis via up-regulating
CA9 in neuroblastoma. Cell Death Dis. 15:52024. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Xiao H, Li Y, Zhang Y and Wang P: Long
noncoding RNA LINC01296 regulates the cell proliferation, migration
and invasion in neuroblastoma. Metab Brain Dis. 37:1247–1258. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Yang H, Guo JF, Zhang ML and Li AM:
LncRNAs SNHG4 promotes neuroblastoma proliferation, migration, and
invasion by sponging miR-377-3p. Neoplasma. 67:1054–1062. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Wang B, Xu L, Zhang J, Cheng X, Xu Q, Wang
J and Mao F: lncRNAs NORAD accelerates the progression and
doxorubicin resistance of neuroblastoma through up-regulating HDAC8
via sponging miR-144-3p. Biomed Pharmacother. 129:1102682020.
View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Yang L, Pei L and Yi J: LINC00839
regulates proliferation, migration, invasion, apoptosis and
glycolysis in neuroblastoma cells through miR-338-3p/GLUT1 Axis.
Neuropsychiatr Dis Treat. 17:2027–2040. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Vaid R, Thombare K, Mendez A,
Burgos-Panadero R, Djos A, Jachimowicz D, Lundberg KI, Bartenhagen
C, Kumar N, Tümmler C, et al: METTL3 drives telomere targeting of
TERRA lncRNAs through m6A-dependent R-loop formation: A therapeutic
target for ALT-positive neuroblastoma. Nucleic Acids Res.
52:2648–267. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Feriancikova B, Feglarova T, Krskova L,
Eckschlager T, Vicha A and Hrabeta J: MIAT is an upstream regulator
of NMYC and the disruption of the MIAT/NMYC axis induces cell death
in NMYC amplified neuroblastoma cell lines. Int J Mol Sci.
22:33932021. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Yang H, Zhang X, Zhao Y, Sun G, Zhang J,
Gao Y, Liu Q, Zhang W and Zhu H: Downregulation of lncRNAs XIST
represses tumor growth and boosts radiosensitivity of neuroblastoma
via modulation of the miR-375/L1CAM axis. Neurochem Res.
45:2679–2690. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Nie L, Li C, Zhao T, Wang Y and Liu J:
LncRNAs double homeobox A pseudogene 8 (DUXAP8) facilitates the
progression of neuroblastoma and activates Wnt/β-catenin pathway
via microRNA-29/nucleolar protein 4 like (NOL4L) axis. Brain Res.
1746:1469472020. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Yang R, Liu N, Li T, Liu F, Zhang J, Zhao
H, Zou L and He X: LncRNAs AC142119.1 facilitates the progression
of neuroblastoma by epigenetically initiating the transcription of
MYCN. J Transl Med. 21:6592023. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Xu Y, Qiu Z, Chen J, Huang L, Zhang J and
Lin J: LINC00460 promotes neuroblastoma tumorigenesis and cisplatin
resistance by targeting miR-149-5p/DLL1 axis and activating Notch
pathway in vitro and in vivo. Drug Deliv Transl Res. 14:2003–2018.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Wang J, Wang Z, Lin W, Han Q, Yan H, Yao
W, Dong R, Jia D, Dong K and Li K: LINC01296 promotes neuroblastoma
tumorigenesis via the NCL-SOX11 regulatory complex. Mol Ther
Oncolytics. 24:834–848. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Zhang Q, Wei J, Li N and Liu B: LINC00839
promotes neuroblastoma progression by Sponging miR-454-3p to
Up-Regulate NEUROD1. Neurochem Res. 47:2278–2293. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Hu Y, Sun H, Hu J and Zhang X: lncRNAs
DLX6-AS1 promotes the progression of neuroblastoma by activating
STAT2 via targeting miR-506-3p. Cancer Manag Res. 12:7451–7463.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Xu Z, Sun Y, Wang D, Sun H and Liu X:
SNHG16 promotes tumorigenesis and cisplatin resistance by
regulating miR-338-3p/PLK4 pathway in neuroblastoma cells. Cancer
Cell Int. 20:2362020. View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Mi J, Han Y, Zhang J, Hao X, Xing M and
Shang C: Long noncoding RNA LINC01410 promotes the tumorigenesis of
neuroblastoma cells by sponging microRNA-506-3p and modulating
WEE1. Cancer Med. 9:8133–8143. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Jia P, Wei E, Liu H, Wu T and Wang H:
Silencing of long non-coding RNA DLX6-AS1 weakens neuroblastoma
progression by the miR-513c-5p/PLK4 axis. IUBMB Life. 72:2627–2636.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Chen M, Zhao M, Hou Y and Zhu B:
Expression of lncRNAs CCAT2 in children with neuroblastoma and its
effect on cancer cell growth. Mol Cell Biochem. 476:1871–1879.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
94
|
Pan W, Wu A, Yu H, Yu Q, Zheng B, Yang W,
Tian D, Gao Y and Li P: NEAT1 negatively regulates cell
proliferation and migration of neuroblastoma cells by
miR-183-5p/FOXP1 via the ERK/AKT pathway. Cell Transplant.
29:9636897209436082020. View Article : Google Scholar : PubMed/NCBI
|
|
95
|
Tan WQ, Yuan L, Cao X, Wu XY, Xing YQ and
Ye M: Overexpression of lncRNAs TUG1 enhances the efficacy of
DC-CIK immunotherapy in neuroblastoma in vitro and in vivo. Cancer
Biomark. 36:53–61. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Zhou X, Lu H, Li F, Han L, Zhang H, Jiang
Z, Dong Q and Chen X: lncRNAs cancer susceptibility candidate
(CASC7) upregulates phosphatase and tensin homolog by
downregulating miR-10a to inhibit neuroblastoma cell proliferation.
Neuroreport. 31:381–386. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Lamouille S, Xu J and Derynck R: Molecular
mechanisms of epithelial-mesenchymal transition. Nat Rev Mol Cell
Biol. 15:178–196. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Dongre A and Weinberg RA: New insights
into the mechanisms of Epithelial-mesenchymal transition and
implications for cancer. Nat Rev Mol Cell Biol. 20:69–84. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
99
|
Mittal V: Epithelial mesenchymal
transition in tumor metastasis. Annu Rev Pathol. 13:395–412. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
100
|
Taki M, Abiko K, Ukita M, Murakami R,
Yamanoi K, Yamaguchi K, Hamanishi J, Baba T, Matsumura N and Mandai
M: Tumor immune microenvironment during Epithelial-mesenchymal
transition. Clin Cancer Res. 27:4669–4679. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Manfioletti G and Fedele M:
Epithelial-mesenchymal transition (EMT). Int J Mol Sci.
24:113862023. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Zhang Y and Weinberg RA:
Epithelial-to-mesenchymal transition in cancer: Complexity and
opportunities. Front Med. 12:361–373. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Piera-Velazquez S and Jimenez SA:
Endothelial to mesenchymal transition: Role in physiology and in
the pathogenesis of human diseases. Physiol Rev. 99:1281–1324.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Lee J, You JH, Kim MS and Roh JL:
Epigenetic reprogramming of epithelial-mesenchymal transition
promotes ferroptosis of head and neck cancer. Redox Biol.
37:1016972020. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Sarrand J and Soyfoo MS: Involvement of
Epithelial-mesenchymal transition (EMT) in autoimmune diseases. Int
J Mol Sci. 24:144812023. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
van Staalduinen J, Baker D, Ten Dijke P
and van Dam H: Epithelial-mesenchymal-transition-inducing
transcription factors: New targets for tackling chemoresistance in
cancer? Oncogene. 37:6195–6211. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Ye M, Lu H, Tang W, Jing T, Chen S, Wei M,
Zhang J, Wang J, Ma J, Ma D and Dong K: Downregulation of MEG3
promotes neuroblastoma development through FOXO1-mediated autophagy
and mTOR-mediated epithelial-mesenchymal transition. Int J Biol
Sci. 16:3050–3061. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Ge Y, Tan S, Bi J, Rao M, Yu Y and Tian L:
SNHG16 knockdown inhibits tumorigenicity of neuroblastoma in
children via miR-15b-5p/PRPS1 axis. Neuroreport. 31:1225–1235.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
109
|
Tower J: Programmed cell death in aging.
Ageing Res Rev. 23:90–100. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Nguyen TT, Wei S, Nguyen TH, Jo Y, Zhang
Y, Park W, Gariani K, Oh CM, Kim HH, Ha KT, et al:
Mitochondria-associated programmed cell death as a therapeutic
target for age-related disease. Exp Mol Med. 55:1595–1619. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Kari S, Subramanian K, Altomonte IA,
Murugesan A, Yli-Harja O and Kandhavelu M: Programmed cell death
detection methods: A systematic review and a categorical
comparison. Apoptosis. 27:482–508. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Peng F, Liao M, Qin R, Zhu S, Peng C, Fu
L, Chen Y and Han B: Regulated cell death (RCD) in cancer: Key
pathways and targeted therapies. Signal Transduct Target Ther.
7:2862022. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Liu L, Li H, Hu D, Wang Y, Shao W, Zhong
J, Yang S, Liu J and Zhang J: Insights into N6-methyladenosine and
programmed cell death in cancer. Mol Cancer. 21:322022. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Hsu SK, Li CY, Lin IL, Syue WJ, Chen YF,
Cheng KC, Teng YN, Lin YH, Yen CH and Chiu CC: Inflammation-related
pyroptosis, a novel programmed cell death pathway, and its
crosstalk with immune therapy in cancer treatment. Theranostics.
11:8813–8835. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Dai X, Wang D and Zhang J: Programmed cell
death, redox imbalance, and cancer therapeutics. Apoptosis.
26:385–414. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Hassannia B, Wiernicki B, Ingold I, Qu F,
Van Herck S, Tyurina YY, Bayır H, Abhari BA, Angeli JPF, Choi SM,
et al: Nano-targeted induction of dual ferroptotic mechanisms
eradicates high-risk neuroblastoma. J Clin Invest. 128:3341–3355.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
117
|
Valter K, Zhivotovsky B and Gogvadze V:
Cell death-based treatment of neuroblastoma. Cell Death Dis.
9:1132018. View Article : Google Scholar : PubMed/NCBI
|
|
118
|
Matthews CC and Feldman EL: Insulin-like
growth factor I rescues SH-SY5Y human neuroblastoma cells from
hyperosmotic induced programmed cell death. J Cell Physiol.
166:323–331. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
119
|
Locquet MA, Ichim G, Bisaccia J, Dutour A,
Lebecque S, Castets M and Weber K: Caspase-8 deficiency induces a
switch from TLR3 induced apoptosis to lysosomal cell death in
neuroblastoma. Sci Rep. 11:106092021. View Article : Google Scholar : PubMed/NCBI
|
|
120
|
Rakshit J, Mallick A, Roy S, Sarbajna A,
Dutta M and Bandyopadhyay J: Iron-induced apoptotic cell death and
autophagy dysfunction in human neuroblastoma cell line SH-SY5Y.
Biol Trace Elem Res. 193:138–151. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
121
|
Li MM, Liu XH, Zhao YC, Ma XY, Zhou YC,
Zhao YX and Liu XY: Long noncoding RNA KCNQ1OT1 promotes apoptosis
in neuroblastoma cells by regulating miR-296-5p/Bax axis. FEBS J.
287:561–577. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
122
|
Yu Y, Chen F, Yang Y, Jin Y, Shi J, Han S,
Chu P, Lu J, Tai J, Wang S, et al: lncRNAs SNHG16 is associated
with proliferation and poor prognosis of pediatric neuroblastoma.
Int J Oncol. 55:93–102. 2019.PubMed/NCBI
|
|
123
|
Nicolai S, Pieraccioli M, Peschiaroli A,
Melino G and Raschellà G: Neuroblastoma: Oncogenic mechanisms and
therapeutic exploitation of necroptosis. Cell Death Dis.
6:e20102015. View Article : Google Scholar : PubMed/NCBI
|
|
124
|
Dong Y, Gong W, Hua Z, Chen B, Zhao G, Liu
Z, Thiele CJ and Li Z: Corrigendum: Combination of rapamycin and
MK-2206 induced cell death via autophagy and necroptosis in
MYCN-amplified neuroblastoma cell lines. Front Pharmacol.
15:14250392024. View Article : Google Scholar : PubMed/NCBI
|
|
125
|
Jing Chu: Development of a prognostic
model for children with neuroblastoma based on necroptosis-related
genes. Front Genet. 13:9470002022. View Article : Google Scholar : PubMed/NCBI
|
|
126
|
Alborzinia H, Flórez AF, Kreth S, Brückner
LM, Yildiz U, Gartlgruber M, Odoni DI, Poschet G, Garbowicz K, Shao
C, et al: MYCN mediates cysteine addiction and sensitizes
neuroblastoma to ferroptosis. Nat Cancer. 3:471–485. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
127
|
Yang X and Thiele CJ: Targeting the tumor
necrosis factor-related apoptosis-inducing ligand path in
neuroblastoma. Cancer Lett. 197:137–143. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
128
|
Murphy C, Devis-Jauregui L, Struck R,
Boloix A, Gallagher C, Gavin C, Cottone F, Fernandez AS, Madden S,
Roma J, et al: In vivo cisplatin-resistant neuroblastoma metastatic
model reveals tumour necrosis factor receptor superfamily member 4
(TNFRSF4) as an independent prognostic factor of survival in
neuroblastoma. PLoS One. 19:e03036432024. View Article : Google Scholar : PubMed/NCBI
|
|
129
|
Cao Y: Adipocyte and lipid metabolism in
cancer drug resistance. J Clin Invest. 129:3006–3017. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
130
|
Burke MR, Smith AR and Zheng G: Overcoming
cancer drug resistance utilizing PROTAC technology. Front Cell Dev
Biol. 10:8727292022. View Article : Google Scholar : PubMed/NCBI
|
|
131
|
Hu XY, Song Z, Yang ZW, Li JJ, Liu J and
Wang HS: Cancer drug resistance related microRNAs: Recent advances
in detection methods. Analyst. 147:2615–2632. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
132
|
Hussain S, Singh A, Nazir SU, Tulsyan S,
Khan A, Kumar R, Bashir N, Tanwar P and Mehrotra R: Cancer drug
resistance: A fleet to conquer. J Cell Biochem. 120:14213–14225.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
133
|
Gao L, Wu ZX, Assaraf YG, Chen ZS and Wang
L: Overcoming anti-cancer drug resistance via restoration of tumor
suppressor gene function. Drug Resist Updat. 57:1007702021.
View Article : Google Scholar : PubMed/NCBI
|
|
134
|
Mabe NW, Huang M, Dalton GN, Alexe G,
Schaefer DA, Geraghty AC, Robichaud AL, Conway AS, Khalid D, Mader
MM, et al: Transition to a mesenchymal state in neuroblastoma
confers resistance to anti-GD2 antibody via reduced expression of
ST8SIA1. Nat Cancer. 3:976–993. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
135
|
Naiditch JA, Jie C, Lautz TB, Yu S, Clark
S, Voronov D, Chu F and Madonna MB: Mesenchymal change and drug
resistance in neuroblastoma. J Surg Res. 193:279–288. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
136
|
Grossmann LD, Chen CH, Uzun Y, Thadi A,
Wolpaw AJ, Louault K, Goldstein Y, Surrey LF, Martinez D, Calafatti
M, et al: Identification and characterization of
Chemotherapy-resistant High-Risk neuroblastoma persister cells.
Cancer Discov. 14:2387–2406. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
137
|
Çoku J, Booth DM, Skoda J, Pedrotty MC,
Vogel J, Liu K, Vu A, Carpenter EL, Ye JC, Chen MA, et al: Reduced
ER-mitochondria connectivity promotes neuroblastoma multidrug
resistance. EMBO J. 41:e1082722022. View Article : Google Scholar : PubMed/NCBI
|
|
138
|
Harvey H, Piskareva O, Creevey L, Alcock
LC, Buckley PG, O'Sullivan MJ, Segura MF, Gallego S, Stallings RL
and Bray IM: Modulation of chemotherapeutic drug resistance in
neuroblastoma SK-N-AS cells by the neural apoptosis inhibitory
protein and miR-520f. Int J Cancer. 136:1579–1588. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
139
|
Clark-Corrigall J, Myssina S, Michaelis M,
Cinatl J Jr, Ahmed S and Carr-Wilkinson J: Elevated expression of
LGR5 and WNT signaling factors in neuroblastoma cells with acquired
drug resistance. Cancer Invest. 41:173–182. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
140
|
Berlak M, Tucker E, Dorel M, Winkler A,
McGearey A, Rodriguez-Fos E, da Costa BM, Barker K, Fyle E, Calton
E, et al: Mutations in ALK signaling pathways conferring resistance
to ALK inhibitor treatment lead to collateral vulnerabilities in
neuroblastoma cells. Mol Cancer. 21:1262022. View Article : Google Scholar : PubMed/NCBI
|
|
141
|
Xiang T, Li Y, Liu G and Li X:
NR1D1-transactivated lncRNAs NUTM2A-AS1 promotes chemoresistance
and immune evasion in neuroblastoma via inhibiting B7-H3
degradation. J Cell Mol Med. 28:e183602024. View Article : Google Scholar : PubMed/NCBI
|
|
142
|
Su Z, Lu H, Wu Y, Li Z and Duan L:
Predicting potential lncRNAs biomarkers for lung cancer and
neuroblastoma based on an ensemble of a deep neural network and
LightGBM. Front Genet. 14:12380952023. View Article : Google Scholar : PubMed/NCBI
|
|
143
|
Yerukala Sathipati S, Sahu D, Huang HC,
Lin Y and Ho SY: Identification and characterization of the lncRNAs
signature associated with overall survival in patients with
neuroblastoma. Sci Rep. 9:51252019. View Article : Google Scholar : PubMed/NCBI
|
|
144
|
Yang H, Pu Y, Hu X, Zhang H, Li S, Zhang
W, Chen M, Zhang H, Jin R and Nie Y: ATRA derived lipid
nanoparticles for co-delivery of siCYP26A1 to overcome metabolic
limitations of differentiation therapy in solid tumors. J Control
Release. 384:1139332025. View Article : Google Scholar : PubMed/NCBI
|
|
145
|
Croteau N, Nuchtern J and LaQuaglia MP:
Management of neuroblastoma in pediatric patients. Surg Oncol Clin
N Am. 30:291–304. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
146
|
Zafar A, Wang W, Liu G, Xian W, McKeon F,
Zhou J and Zhang R: Targeting the p53-MDM2 pathway for
neuroblastoma therapy: Rays of hope. Cancer Lett. 496:16–29. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
147
|
Chronopoulos A, Vemula CK, Zamloot V,
Chavez I, Kennedy R, Kim W, Bell D, Pan Y, Moghimi B, Kim J, et al:
Extracellular matrix topography drives adrenergic to mesenchymal
transition in neuroblastoma. Adv Mater. 37:e25015262025. View Article : Google Scholar : PubMed/NCBI
|
|
148
|
Mitra S, Muralidharan SV, Di Marco M,
Juvvuna PK, Kosalai ST, Reischl S, Jachimowicz D, Subhash S,
Raimondi I, Kurian L, et al: Subcellular distribution of p53 by the
p53-Responsive lncRNA NBAT1 determines chemotherapeutic response in
neuroblastoma. Cancer Res. 81:1457–1471. 2021. View Article : Google Scholar : PubMed/NCBI
|