Introduction
Endometrial cancer (EC) is the fifth most commonly
diagnosed malignancy worldwide and remains one of the most lethal
types of gynecological cancer (1).
Notably, ~75% of patients are diagnosed at an early stage (FIGO
stage I–II), with a favorable overall 5-year survival rate of ~80%.
By contrast, patients diagnosed at an advanced stage (FIGO stage
III) exhibit markedly poorer outcomes, with reported 5-year
survival rates ranging between 25 and 60% (2). Despite advances in therapeutic
approaches, including surgery, chemotherapy, radiotherapy and
hormonal therapy, the incidence and mortality of EC have continued
to increase (3). These trends
highlight an urgent need to elucidate the molecular mechanisms
underlying EC progression, and to identify novel biomarkers and
therapeutic targets to improve prognostic assessment and clinical
management.
DNA polymerase θ (POLQ, also known as Polθ), encoded
by the POLQ gene, is a key mediator of microhomology-mediated end
joining (MMEJ), a DNA double-strand break (DSB) repair pathway that
mediates the ligation of excised 3′ DNA ends using microhomologous
sequences (4,5). Structurally, POLQ consists of a
C-terminal DNA polymerase domain, an N-terminal helicase-like
domain with DNA-dependent ATPase activity, and a non-conserved
central domain (6). During the S
phase of the cell cycle, MMEJ directly competes with homologous
recombination (HR) for DNA end binding, thus serving a critical
role in DNA damage repair (7,8).
Tumors with HR deficiency are particularly dependent on
POLQ-mediated repair mechanisms (9). Aberrant POLQ expression has been
reported across multiple malignancies, including colorectal
(10), lung (11) and breast cancer (12,13),
where its increased expression is consistently associated with
unfavorable clinical outcomes (14–16).
Beyond its established functions in DNA replication, cell cycle
regulation and genome stability, POLQ has also been implicated in
immune regulation (17), cancer
stemness and ferroptosis (12).
However, to the best of our knowledge, the molecular mechanisms and
prognostic importance of POLQ in EC have not been previously
investigated.
Therefore, the current study aimed to systematically
analyze POLQ expression in EC, and to investigate its association
with the immune checkpoint protein programmed death-ligand 1
(PD-L1) using data from The Cancer Genome Atlas (TCGA), with
validation in EC patient specimens. Furthermore, the functional
effects of POLQ on EC cell proliferation and tumor progression, as
well as the underlying molecular mechanisms, were further
investigated through in vitro experiments. Collectively,
these findings may provide novel insights into the oncogenic role
of POLQ in EC, and support its potential value as both a prognostic
biomarker and a therapeutic target.
Materials and methods
Assessment of POLQ expression in EC
using publicly available databases
A total of two publicly available databases were
utilized: TCGA Uterine Corpus Endometrial Carcinoma (TCGA-UCEC)
project via the Genomic Data Commons portal (https://portal.gdc.cancer.gov/) and the TNMplot online
database (https://tnmplot.com/). Transcriptomic
profiles of 554 EC tissues and 35 adjacent non-tumor endometrial
tissues were retrieved from TCGA-UCEC. In parallel, expression data
from 469 EC tissues and 315 normal endometrial tissues from healthy
controls were retrieved from the TNMplot database, which integrates
RNA-sequencing (RNA-seq) data from TCGA, Genotype-Tissue Expression
and Gene Expression Omnibus cohorts (18).
Association between POLQ expression
and clinicopathological characteristics
To evaluate the association between POLQ expression
and clinicopathological features, association analysis using data
from TCGA-EC cohort was conducted. Patients were stratified into
high- and low-expression groups based on the median POLQ expression
levels (n=277/group). RNA-seq data and the corresponding clinical
information were retrieved from TCGA. Clinicopathological variables
included age, body mass index (BMI), histological subtype, tumor
grade, clinical stage and survival status. The prognostic
significance of POLQ expression was assessed using overall survival
(OS), disease-specific survival (DSS) and progression-free interval
(PFI). Survival curves were generated using the Kaplan-Meier method
and compared using the log-rank test. Furthermore, univariate Cox
proportional hazards regression and multivariate Cox regression
analyses were performed to determine whether POLQ expression was
independently associated with survival outcomes in TCGA-EC cohort.
All statistical analyses were conducted using R software (version
4.3.1; R Foundation for Statistical Computing; http://www.r-project.org/). Survival analyses were
performed using the ‘survival’ package (version 3.8–6; http://cran.r-project.org/package=survival) and
‘survminer’ package (version 0.5.1; http://cran.r-project.org/package=survminer) in R.
Immune infiltration and gene set
enrichment analysis (GSEA)
Immune infiltration analysis and GSEA were performed
using TCGA-UCEC cohort. The Estimation of Stromal and Immune Cells
in Malignant Tumor Tissues using Expression Data (ESTIMATE)
algorithm (19) was employed to
assess the relative abundance of stromal and immune cells within
tumor samples based on gene expression profiles. Immune
infiltration analyses, including the calculation of stromal, immune
and ESTIMATE scores, were carried out using the ‘estimate’ R
package (version 1.0.13; http://r-forge.r-project.org/projects/estimate/)
in R software (version 4.3.1), based on transcriptomic data from
TCGA-UCEC cohort. Briefly, the ESTIMATE algorithm calculates
stromal and immune scores based on single-sample GSEA of predefined
stromal- and immune-related gene signatures, and the ESTIMATE score
is derived from the combination of the stromal and immune scores.
Additionally, the relative abundance of specific immune cell
populations was assessed using the ‘GSVA’ R package (version
1.46.0) (20–22). In addition, GSEA, including Hallmark
pathway analysis, and Kyoto Encyclopedia of Genes and Genomes
(KEGG) pathway enrichment analyses were performed using R software
(version 4.3.1). GSEA was conducted using the ‘clusterProfiler’
package (version 4.6.2; http://bioconductor.org/packages/clusterProfiler/),
and gene sets were obtained from the Molecular Signatures Database
(MSigDB, version 7.5.1; http://www.gsea-msigdb.org/gsea/msigdb), including the
Hallmark gene set collection. KEGG pathway enrichment analysis was
also performed using ‘clusterProfiler’. Pathways with a false
discovery rate <0.25 were considered significantly enriched.
Spearman's correlation coefficients were calculated to evaluate the
associations between POLQ expression levels and pathway enrichment
scores, with a significance threshold set at rs>0.5 and
P<0.05.
Differential expression and Gene
Ontology (GO) enrichment analyses
Differential expression analysis between the
POLQ-high and POLQ-low groups was performed using the ‘limma’ R
package (version 3.54.0; http://bioconductor.org/packages/limma/) in R software
(version 4.3.1). RNA-seq data from TCGA-UCEC cohort were analyzed,
and patients were stratified into POLQ-high and POLQ-low groups
based on the median expression level of POLQ. Linear models were
fitted using the limma framework, and empirical Bayes moderation
was applied. Genes with P<0.05 and |log2FC |>1.5 were
considered significantly differentially expressed. Volcano plots
were generated using the ‘ggplot2’ R package (version 3.4.2;
http://cran.r-project.org/package=ggplot2), with
significance thresholds set at |log2FC|>1.5 and P<0.05. GO
enrichment analysis of the identified DEGs was conducted using the
‘clusterProfiler’ R package (version 4.6.2). Enriched GO terms in
the Biological Process categories were identified. GO terms with
P<0.05 were considered significantly enriched.
Clinical specimens and
immunohistochemistry (IHC)
A total of 113 formalin-fixed paraffin-embedded
(FFPE) tissue specimens, including 78 EC tissues and 35 paired
adjacent non-cancerous endometrial tissues, were collected from
Taihe Hospital (Shiyan, China) between July 2020 and October 2023.
The tissues were fixed in 10% neutral buffered formalin at room
temperature (20–25°C) for 24 h prior to routine dehydration and
paraffin embedding. All patients with EC were female, with a median
age of 59 years (range, 41–72 years). All diagnoses were confirmed
by senior pathologists, and none of the patients had received
preoperative radiotherapy or chemotherapy. FFPE tissue blocks were
cut into 3-µm sections, which were deparaffinized in xylene,
rehydrated through graded ethanol and subjected to heat-induced
antigen retrieval using EDTA buffer (pH 9.0) at 95–100°C for 15
min. Endogenous peroxidase activity was blocked following
incubation with 3% hydrogen peroxide for 10 min at room
temperature. After blocking with QuickBlock™ Blocking Buffer for
Immunol Staining (cat. no. P0260; Beyotime Biotechnology) at room
temperature for 15 min to reduce non-specific binding, the sections
were incubated at 4°C overnight with primary antibodies against
POLQ, Ki67 and PD-L1 (Table SI),
followed by incubation with horseradish peroxidase (HRP)-conjugated
secondary antibodies (ready-to-use; cat. no. PV-6000; Beijing
Zhongshan Jinqiao Biotechnology Co., Ltd.) at room temperature for
30 min. Immunoreactivity was visualized using DAB as the chromogen
and cell nuclei were counterstained with hematoxylin at room
temperature for 1 min. POLQ expression levels were
semi-quantitatively scored based on staining quantity (0–4) and
intensity (0–3), with final overall scores calculated as quantity ×
intensity (range, 0–12). Ki67 positive expression was quantified as
the percentage of Ki67+ cells at ×200 magnification.
PD-L1 IHC was carried out using a PD-L1 antibody (clone no. 22C3;
ready-to-use; cat. no. SK006; Dako; Agilent Technologies, Inc.),
which was used to incubate the sections at 4°C overnight. Detection
was carried out using the EnVision™ FLEX HRP detection kit (cat.
no. K8000; Dako; Agilent Technologies, Inc.), which includes an
HRP-conjugated secondary antibody polymer, according to the
manufacturer's instructions. PD-L1 expression was evaluated using a
semi-quantitative IHC scoring system. Staining intensity was scored
as 0 (negative), 1 (weak), 2 (moderate) or 3 (strong), and the
proportion of positive tumor cells was scored from 0 to 5 according
to the percentage of positive cells. The final IHC score was
calculated as the sum of the intensity and proportion scores,
yielding a total score ranging from 0 to 8 (23). All slides were independently
evaluated by at least two experienced pathologists using a
bright-field light microscope (Olympus Corporation).
Cell culture and transfection
HEC-1-B and Ishikawa cells were obtained from The
Cell Bank of Type Culture Collection of The Chinese Academy of
Sciences and cultured in RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) supplemented with 10% fetal bovine serum (FBS;
Gibco; Thermo Fisher Scientific, Inc.). Cells were maintained at
37°C in a humidified incubator with 5% CO2. Once they
had reached ~90% confluence, the cells were passaged and were then
seeded in 6-well plates for further experiments. Upon reaching ~30%
confluence, the cells were transfected with small interfering
(si)RNAs targeting POLQ (si-POLQ#1 and si-POLQ#2) or the
corresponding negative control siRNA (si-NC)(all synthesized by
Sangon Biotech Co., Ltd.), at a final concentration of 50 nM using
GenMute™ siRNA Transfection Reagent (cat. no. SL100568; SignaGen
Laboratories), according to the manufacturer's instructions. The
culture medium was replaced at 6 h post-transfection. Transfection
was performed at 37°C in a humidified incubator with 5%
CO2. To detect the mRNA expression levels of target
genes, cells were harvested at 24 h post-transfection, whereas
protein expression levels were assessed 48 h post-transfection. All
subsequent functional assays were performed 48 h after transfection
unless otherwise specified. All experiments were performed in
triplicate. The siRNA sequences used are listed in Table SII.
Reverse transcription-quantitative PCR
(RT-qPCR)
Total RNA was extracted from the transfected cells
using the RNAeasy™ Animal RNA Extraction Kit (cat. no. R0024;
Beyotime Biotechnology), which is designed for mammalian cells,
according to the manufacturer's instructions. cDNA was then
synthesized using the Evomak-Mlo Reverse Transcription Kit (cat.
no. AG11705; Hunan Accurate Bio-Medical Technology Co., Ltd.)
according to the manufacturer's protocol. qPCR was performed using
TB Green® Premix Ex Taq™ II (Tli RNaseH Plus) (cat. no.
RR820A; Takara Bio, Inc.) according to the manufacturer's
instructions. The thermocycling conditions were as follows: 95°C
for 30 sec, followed by 40 cycles at 95°C for 5 sec and 60°C for 30
sec. A melt curve analysis was performed to confirm amplification
specificity. Relative mRNA expression levels were calculated using
the 2−ΔΔCq method (24).
The primer sequences used are listed in Table SII.
Western blot analysis
Transfected cells were lysed on ice for 30 min using
Western and IP Cell Lysis Buffer (cat. no. P0037; Beyotime
Biotechnology) and cell lysates were centrifuged at 12,000 × g for
15 min at 4°C and the supernatants were collected for subsequent
analysis. Protein concentrations were calculated using a BCA assay
(cat. no. P0012S; Beyotime Biotechnology). Subsequently, equal
amounts of protein samples (30 µg per lane) were mixed with loading
buffer, separated by SDS-PAGE using different percentages of
polyacrylamide gels according to the molecular weights of the
target proteins (6–15%), and transferred onto PVDF membranes. After
blocking with 5% skimmed milk for 1 h at room temperature, the
membranes were incubated overnight at 4°C with primary antibodies
against the target proteins listed in Table SII. After washing with TBS-0.1%
Tween-20 three times, the membranes were incubated with
HRP-conjugated goat anti-rabbit IgG (H+L) (1:10,000; cat. no.
SA00001-2; Wuhan Sanying Biotechnology) or HRP-conjugated goat
anti-mouse IgG (H+L) (1:10,000; cat. no. SA00001-1; Wuhan Sanying
Biotechnology) for 1 h at room temperature. The protein bands were
then visualized using an enhanced chemiluminescence substrate (cat.
no. MA0187; Dalian Meilun Biology Technology Co., Ltd.) on a
Bio-Rad gel imaging system (Bio-Rad Laboratories, Inc.). The
detailed information of all primary antibodies used for western
blot analysis, including the antibody sources, catalog numbers and
dilutions, is provided in Table
SIII.
Cell Counting Kit 8 (CCK-8) assay
Cells were collected and, following transfection for
48 h, were counted and seeded in 96-well plates at a density of
2,000 cells/well. Cell viability was then assessed at 24, 48, 72
and 96 h after seeding. Following incubation for the indicated time
points, each well was supplemented with 10 µl CCK-8 reagent (cat.
no. BS350A; Biosharp Life Sciences), followed by incubation for an
additional 1 h at 37°C. The optical density in each well was then
measured at an absorbance of 450 nm using a microplate reader.
Colony formation assay
Transfected HEC-1-B and Ishikawa cells were
digested, counted using a hemocytometer (Neubauer chamber) and
seeded into 6-well plates at a density of 500 cells/well, with
three replicates per group. The cells were incubated for 7–14 days
depending on the growth rate of the cell lines, until visible
colonies (≥50 cells/colony) had formed. Subsequently, the plates
were washed with PBS and the colonies were fixed in 100% methanol
at room temperature for 15 min. After fixation, the colonies were
stained with 0.5% crystal violet solution (MilliporeSigma) at room
temperature for 20 min. Colonies were counted manually under a
bright-field light microscope (CX43; Olympus Corporation) and
representative images were captured using a smartphone camera
attached to the bright-field light microscope.
EdU proliferation assay
Transfected HEC-1-B and Ishikawa cells were
trypsinized, counted and seeded in 96-well plates at a density of
2×104 cells/well. The cells were then incubated with EdU
(final concentration: 10 µM; Beyotime Biotechnology) at 37°C for 2
h. Subsequently, cells were fixed with 4% paraformaldehyde at room
temperature for 15 min, followed by permeabilization with 0.3%
Triton X-100 at room temperature for 10 min, according to the
manufacturer's instructions. EdU incorporation was detected using
the Click reaction solution, and cell nuclei were counterstained
with Hoechst 33342, according to the BeyoClick™ EdU-594 Cell
Proliferation Assay Kit protocol (cat. no. C0078S; Beyotime
Biotechnology). Fluorescence images were captured under a
fluorescence microscope. EdU-positive proliferating cells displayed
red fluorescence, whereas total nuclei were visualized by blue
fluorescence. The proliferation rate was calculated as the ratio of
EdU-positive cells to the total number of cells.
Transwell assay
For the Transwell migration assays, transfected
HEC-1-B and Ishikawa cells were digested, washed twice with
serum-free RPMI-1640 medium, and resuspended at a density of
1.5×105 cells/well in 24-well Transwell plates (pore
size, 8 µm). Cell suspensions were seeded into the upper chamber of
Transwell inserts in serum-free medium, whereas 600 µl RPMI-1640
medium containing 10% FBS was added to the lower chamber as a
chemoattractant. Following incubation at 37°C for 24 h,
non-migrated cells on the upper surface of the membrane were gently
removed and the inserts were washed twice with PBS. Migrated cells
on the lower surface were fixed with 4% paraformaldehyde for 30 min
at 25°C and stained with 0.1% crystal violet for 20 min at 25°C.
Excess stain was then removed by washing three times with
double-distilled water, and, after drying, migrated cells were
visualized and images were using an inverted light microscope
(Olympus Corporation).
For the invasion assays, Transwell inserts were
precoated with Matrigel (cat. no. 356234; Corning, Inc.). Briefly,
pre-melted Matrigel was diluted and added to the upper surface of
the Transwell chamber, followed by incubation at 37°C overnight to
allow polymerization. Subsequently, the cells were harvested and
resuspended at a density of 7.5×105/well. The subsequent
procedures were performed as described for the Transwell migration
assays.
Wound healing assay
For wound healing assays, transfected HEC-1-B and
Ishikawa cells were seeded into 6-well plates and cultured until
they reached ~90% confluence. Subsequently, a straight scratch was
created in each well using a 200-µl pipette tip. The detached cells
were removed after washing twice with PBS and the cells were then
cultured in serum-free medium. Images of the wound area were
captured immediately after scratching (0 h) under an inverted
phase-contrast light microscope (magnification, ×10). Following
incubation at 37°C for 24 h in a humified atmosphere with 5%
CO2, images of the same wound areas were captured. The
wound area was semi-quantified using ImageJ software (version
1.53a; National Institutes of Health), and the cell migration rate
was calculated using the following formula: Migration rate
(%)=[(wound area at 0 h-wound area at 24 h)/wound area at 0 h]
×100.
Apoptosis assessment by flow
cytometry
Following transfection, the culture supernatants
were collected and adherent cells were detached using 0.25% trypsin
without EDTA at 37°C for 3 min; digestion was terminated by the
addition of complete medium. The cells were then pooled with the
collected supernatants and centrifuged at 300 × g for 5 min at 4°C
for further analysis. Both suspended and adherent cells were
pooled, washed twice with PBS and resuspended. A total of
1×104 cells/well were resuspended in 100 µl binding
buffer and incubated with 5 µl Annexin V-FITC and 2 µl propidium
iodide (PI) for 15 min at room temperature in the dark, according
to the manufacturer's instructions (Annexin V-FITC/PI Apoptosis
Detection Kit; cat. no. 556547; BD Biosciences). Staining was
terminated by adding 400 µl binding buffer, and the cell apoptosis
rate was subsequently analyzed by flow cytometry (FACSCalibur; BD
Biosciences) using FlowJo 7.6 software (BD Biosciences).
Cell cycle assay
Transfected cells were seeded into 6-well plates and
harvested when they reached ~80% confluence. The cells were then
digested with 0.25% trypsin without EDTA, washed with PBS and
resuspended in 300 µl cell cycle staining solution (Cell Cycle and
Apoptosis Analysis Kit; cat. no. CCS012; MultiSciences Biotech Co.,
Ltd.) containing PI and RNase A. Following incubation at 37°C for
30 min in the dark, cell cycle distribution was analyzed using a BD
FACSCanto II flow cytometer (BD Biosciences). The proportion of
cells in each phase of the cell cycle was quantified using FlowJo
7.6 software.
Statistical analysis
All statistical analyses were performed using
GraphPad Prism 8.0 (Dotmatics). Data are presented as the mean ±
standard deviation, and all in vitro experiments were
conducted in triplicate. For normally distributed continuous
variables, paired or unpaired Student's t-tests were used to
compare the differences between two groups. Non-normally
distributed data were analyzed using the Wilcoxon rank-sum test.
Comparisons among multiple groups were carried out using one-way
ANOVA followed by Bonferroni post hoc test. Categorical variables
were analyzed using the χ2 test, as all expected
frequencies were ≥5. Correlations were assessed using Spearman's
rank correlation coefficient. Survival analyses were performed
using the Kaplan-Meier method and compared using the log-rank test.
Two-tailed P<0.05 was considered to indicate a statistically
significant difference.
Results
POLQ is upregulated in EC, and is
associated with adverse clinicopathological features and a poor
prognosis
Comprehensive bioinformatics analyses using multiple
publicly available tumor databases, including TNMplot and TCGA-EC
cohort, demonstrated that POLQ mRNA expression was significantly
increased in EC tissues compared with that in adjacent normal
tissues (Fig. 1A-C).
Clinicopathological analyses further revealed that high POLQ
expression was significantly associated with patient age,
histological subtype and grade, and clinical stage, whereas no
significant associations were observed with BMI or residual tumor
status (Table SIV). Notably, POLQ
expression was markedly increased in patients with advanced
clinical stage EC (Fig. 1D), a
higher histological grade (Fig.
1E), a more aggressive histological subtype (Fig. 1F) and tumor invasion (Fig. 1G), compared with their respective
control groups. Correlation analysis based on TCGA-EC data further
showed a positive association between POLQ expression and the
proliferation marker Ki67 (Fig.
1H), suggesting a potential role for POLQ in promoting tumor
cell proliferation. Additionally, survival analyses further
revealed that high POLQ expression was significantly associated
with unfavorable OS (Fig. 1I), DSS
(Fig. 1J) and PFI (Fig. 1K). Univariate Cox regression
analysis demonstrated that POLQ expression was significantly
associated with overall survival in patients with EC. However, this
association was not maintained in the multivariate Cox regression
analysis (P=0.393), indicating that POLQ expression was not an
independent prognostic factor (Table
SV). To further validate the aforementioned findings at the
protein level, IHC analysis was performed on tissue microarrays
comprising 78 EC specimens and 35 paired adjacent non-cancerous
endometrial tissues. POLQ expression was semi-quantitatively
evaluated based on both staining intensity and quantity. Consistent
with the transcriptomic data, POLQ protein expression levels were
markedly elevated in EC tissues compared with those in adjacent
non-cancerous controls (Fig. 1L and
M), thus supporting the oncogenic role of POLQ in EC. Taken
together, these findings suggest that POLQ is upregulated in EC,
and is associated with aggressive clinicopathological features and
poor prognosis, potentially contributing to tumor progression,
although it does not serve as an independent prognostic factor.
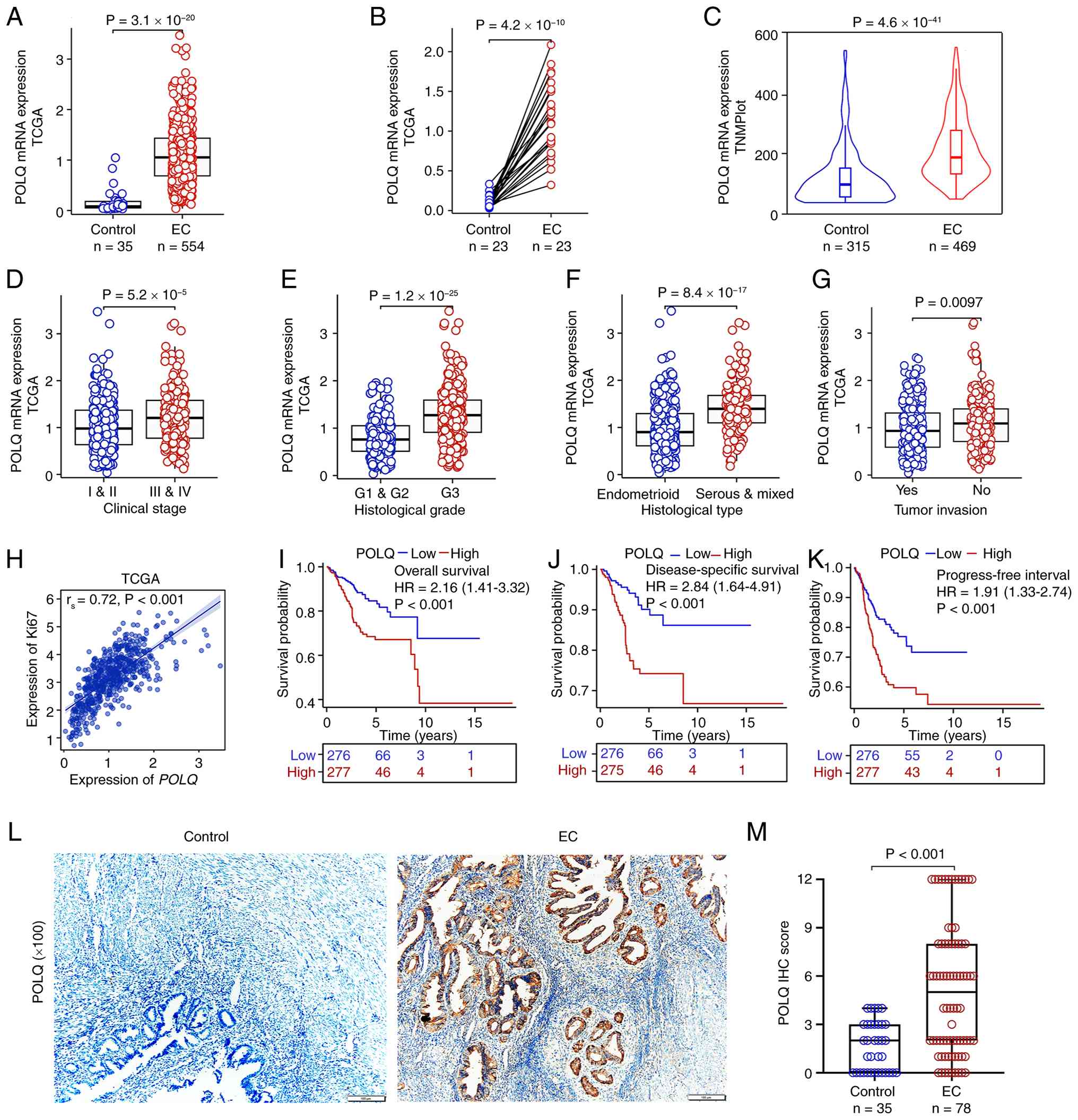 | Figure 1.Aberrant expression of POLQ in EC.
Expression levels of POLQ in EC tissues compared with control
tissues based on (A) unpaired TCGA, (B) paired TCGA and (C) TNMplot
datasets. Association between POLQ expression and
clinicopathological features in EC, including (D) clinical stage,
(E) histological grading, (F) histological type and (G) tumor
invasion. (H) Correlation analysis of POLQ expression and Ki67
expression based on TCGA-EC data. Prognostic analysis of POLQ
expression in EC based on TCGA data, including (I) overall
survival, (J) disease-specific survival and (K) progression-free
interval. (L) IHC images of POLQ in EC (n=78) and adjacent
non-cancerous control tissues (n=35); ×100 magnification. (M) IHC
score of POLQ in EC and adjacent non-cancerous control tissues. EC,
endometrial cancer; IHC, immunohistochemistry; POLQ, DNA polymerase
θ; TCGA, The Cancer Genome Atlas. |
POLQ expression is associated with an
immunosuppressive tumor microenvironment in EC
To further elucidate the biological role of POLQ in
EC, its association with the tumor immune microenvironment was
evaluated using TCGA data. GSEA suggested that aberrant POLQ
expression could affect the infiltration of multiple immune cell
populations, with significantly reduced infiltration of T cells and
natural killer (NK) cells in the high POLQ expression group
compared with the low POLQ expression group (Fig. 2A). Consistently, analysis using the
ESTIMATE algorithm revealed that stromal, immune and composite
ESTIMATE scores were significantly decreased in EC samples with
high POLQ expression (Fig. 2B).
Furthermore, correlation analyses demonstrated that POLQ expression
was weakly negatively correlated with CD8+ T-cell
infiltration, and was also negatively correlated with the
infiltration of NK cells and dendritic cells, while showing a
positive correlation with T helper 2 cells (Fig. 2C). Notably, POLQ expression
exhibited a weak but statistically significant positive correlation
with immune checkpoint-related molecules, particularly with CD274
(PD-L1; Fig. 2D). Consistent with
these findings, the expression levels of CD274, lymphocyte
activation gene 3, programmed cell death 1 ligand 2, and T cell
immunoreceptor with Ig and ITIM domains were significantly
increased in POLQ-high EC samples compared with in POLQ-low samples
(Fig. 2E). Further correlation
analyses confirmed a weak but statistically significant association
between POLQ and CD274 (PD-L1) expression (Fig. 2F), as well as a notable association
between POLQ expression and microsatellite instability score
(Fig. 2G) . Collectively, these
findings suggest that elevated POLQ expression is associated with
an immunosuppressive tumor microenvironment in EC, characterized by
reduced cytotoxic immune cell infiltration and increased expression
of immune checkpoint-related molecules.
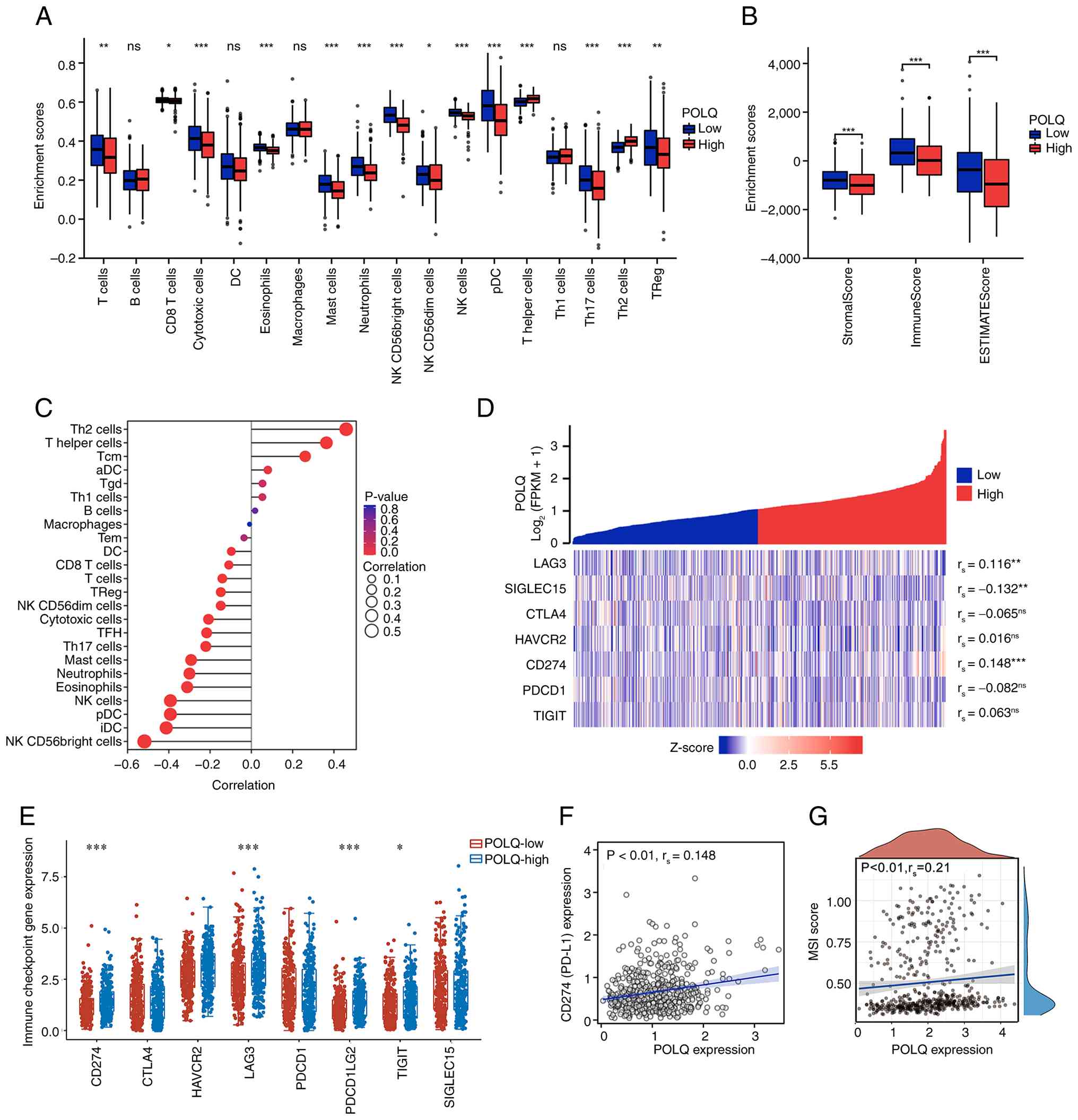 | Figure 2.Immune infiltration analysis of POLQ
in EC. (A) Immune cell abundance of high- or low-POLQ expression
groups in EC. (B) Immune infiltration analyses of the ESTIMATE
algorithm, including stromal score, immune score and ESTIMATE score
in EC with high- and low-POLQ expression. (C) Correlation analyses
of immune cells and POLQ expression. (D) Correlation analysis
between immune checkpoint-related molecules and POLQ expression.
(E) Expression levels of immune checkpoint-related genes in high-
and low-POLQ expression groups. Correlation analysis (F) between
CD274 (PD-L1) and POLQ expression, and (G) between MSI score and
POLQ expression in EC. *P<0.05, **P<0.01, ***P<0.001. aDC,
activated DC; DC, dendritic cell; EC, endometrial cancer; ESTIMATE,
Estimation of Stromal and Immune Cells in Malignant Tumor Tissues
using Expression Data; iDC, immature DC; MSI, microsatellite
instability; NK, natural killer; ns, not significant; pDC,
plasmacytoid DC; PD-L1, programmed death-ligand 1; POLQ, DNA
polymerase θ; THF, T follicular helper; Th, T helper; TReg,
regulatory T. |
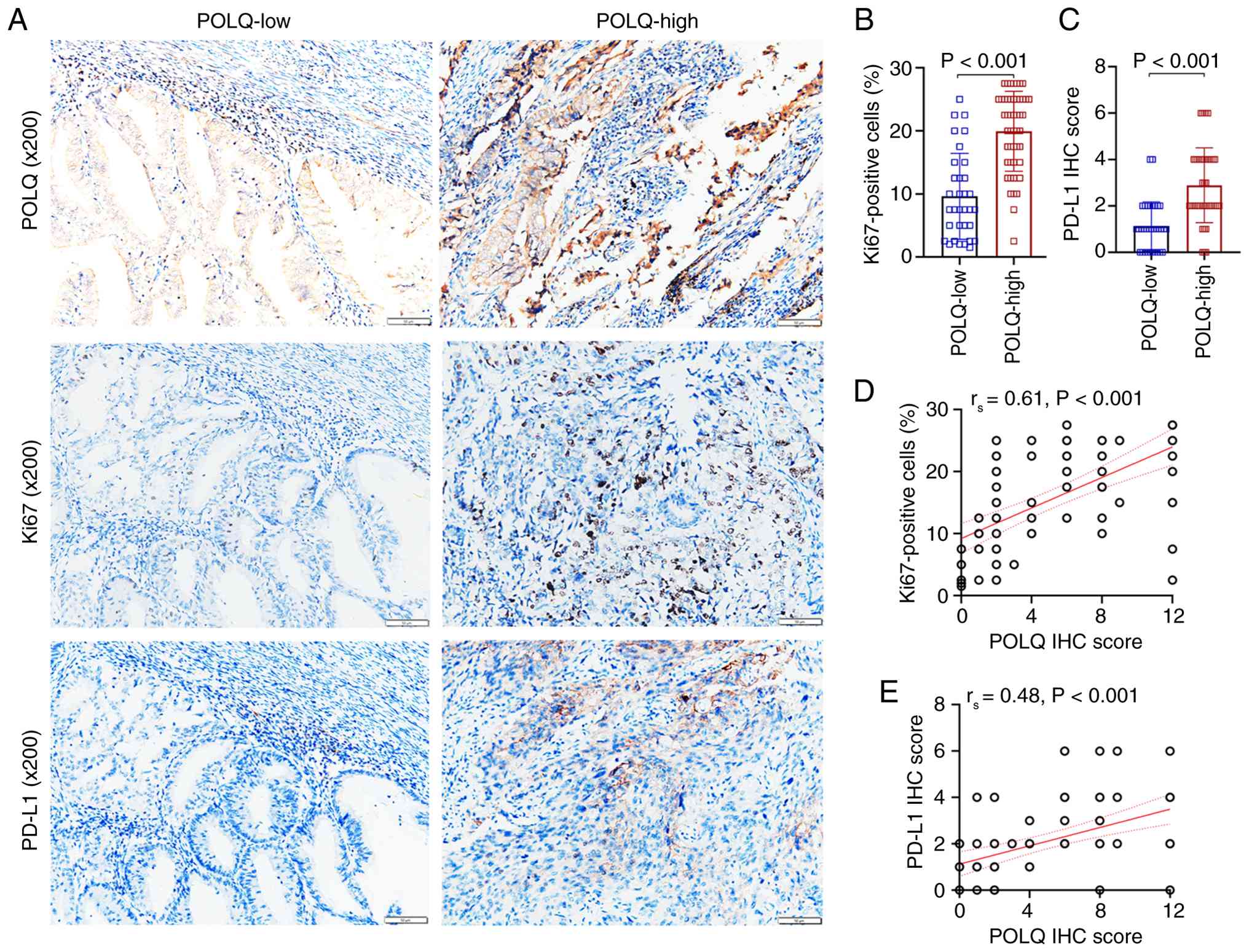
POLQ expression is positively
associated with Ki67+ cells and PD-L1 expression in
EC
Ki67, a well-established marker of cellular
proliferation, was employed to investigate the association between
POLQ expression and tumor growth in EC. In parallel, the expression
levels of CD274 (PD-L1), an immune checkpoint molecule with
immunosuppressive functions, were also detected. Using IHC assays,
the association between POLQ expression, proliferative activity and
immunosuppressive signaling was assessed in EC tissues. The results
demonstrated that the proportion of Ki67+ cells was
significantly higher in the POLQ-high subgroup compared with that
in the POLQ-low subgroup (Fig. 3A and
B). Consistently, POLQ expression levels, quantified by IHC
scoring, showed a significant positive correlation with the
percentage of Ki67+ cells (Fig. 3D). In addition, PD-L1 expression was
markedly increased in POLQ-high EC tissue samples compared with
that in POLQ-low tissues (Fig. 3A and
C), and POLQ expression was positively associated with PD-L1
levels (Fig. 3E), thus indicating
that POLQ upregulation could be involved in the establishment of an
immunosuppressive tumor microenvironment in EC.
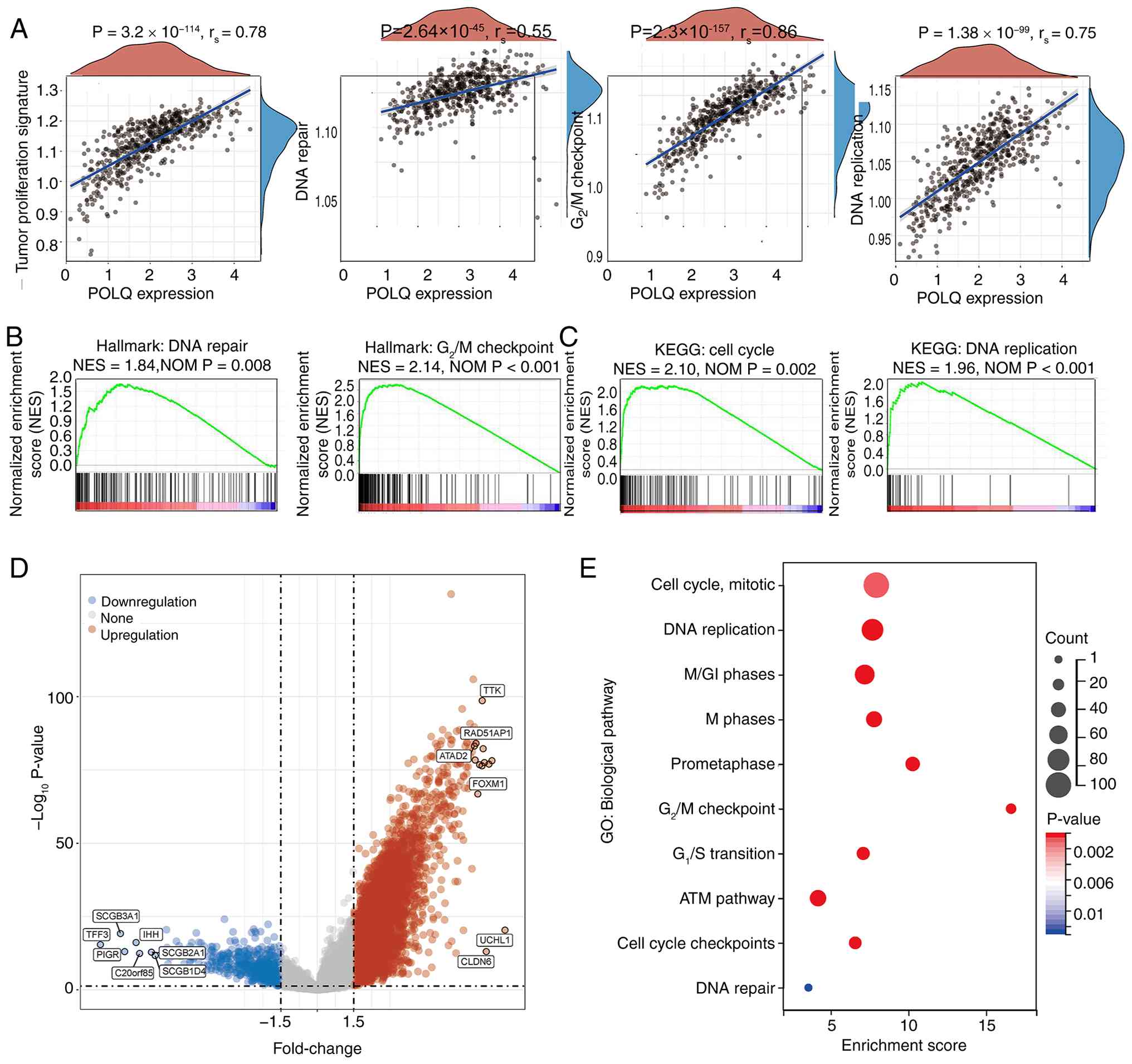
POLQ upregulation is associated with
cell cycle progression and DNA repair pathways in EC
To elucidate the potential molecular mechanisms
underlying the effects of POLQ on EC, GSEA and KEGG enrichment
analyses were performed to identify molecular pathways associated
with differential POLQ expression. GSEA revealed that EC samples
with high POLQ expression were significantly enriched in pathways
associated with tumor cell proliferation, DNA repair, the
G2/M cell cycle checkpoint and DNA replication (Fig. 4A). Hallmark pathway analysis further
confirmed strong associations between POLQ upregulation and DNA
repair [normalized enrichment score (NES)=1.84; P=0.008], as well
as the G2/M checkpoint (NES=2.14; P<0.001; Fig. 4B). Consistently, KEGG pathway
analysis revealed that high POLQ expression was significantly
associated with ‘cell cycle’ (NES=2.10; P=0.002) and ‘DNA
replication’ (NES=1.96; P<0.001), both of which are critical
drivers of tumor cell proliferation and progression (Fig. 4C). Differentially expressed genes
(DEGs) between POLQ-high and POLQ-low groups are presented in
Fig. 4D. Subsequent GO enrichment
analysis of these DEGs further indicated that aberrant POLQ
expression predominantly affected EC tumorigenesis and progression
through regulation of the ‘Cell Cycle, Mitotic’, ‘DNA Replication’
and ‘ATM pathway’ terms (Fig. 4E).
These findings suggested that POLQ could promote EC progression by
enhancing DNA repair capacity and facilitating cell cycle
progression.
POLQ promotes EC cell proliferation,
migration, invasion and epithelial-mesenchymal transition (EMT) in
vitro
To investigate the biological role of POLQ in EC
in vitro, the HEC-1-B and Ishikawa cell lines were selected.
Firstly, POLQ expression at both the mRNA and protein levels was
assessed following cell transfection with si-NC, si-POLQ#1 or
si-POLQ#2 for 48 h. The results showed that both si-POLQ#1 and
si-POLQ#2 efficiently suppressed POLQ expression compared with that
in the si-NC group (Fig. 5A and B).
Functional assays demonstrated that POLQ knockdown markedly
inhibited HEC-1-B and Ishikawa cell proliferation, as evidenced by
the CCK-8 assays (Fig. 5C) and
reduced EdU incorporation (Fig.
5D). Consistently, colony formation ability was significantly
reduced in POLQ-silenced cells (Fig.
5E). Transwell assays further revealed that POLQ knockdown
notably impaired both the migratory and invasive capacities of
HEC-1-B and Ishikawa cells (Fig.
5F), while wound healing assays confirmed a pronounced
reduction in cell migration following POLQ silencing (Fig. 5G). At the molecular level, western
blot analysis demonstrated that POLQ knockdown modulated the
expression levels of EMT-related proteins, characterized by
E-cadherin upregulation, and downregulation of N-cadherin and
vimentin (Fig. 5H). These results
suggested that POLQ could promote EC progression by facilitating
EMT.
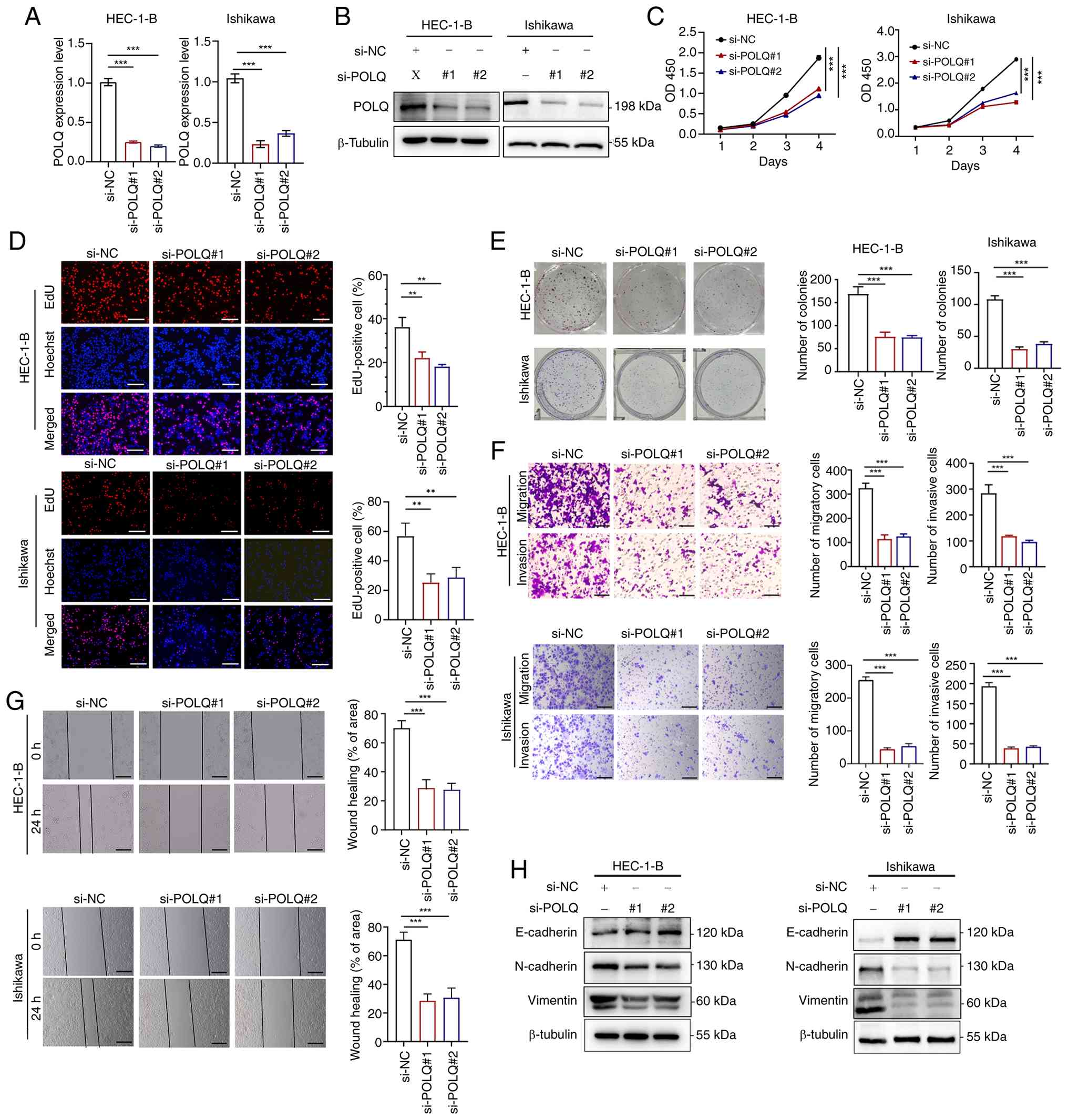 | Figure 5.Aberrant POLQ expression affects cell
proliferation and invasion in endometrial cancer cell lines. (A)
mRNA and (B) protein expression levels of POLQ in HEC-1-B and
Ishikawa cells transfected with si-NC, si-POLQ#1 and si-POLQ#2.
Results of the (C) CCK-8 assay, (D) EdU experiment (scale bar, 50
µm), (E) colony formation assay, (F) Transwell assays (×10
magnification; scale bar, 200 µm) and (G) wound healing assay of
transfected HEC-1-B and Ishikawa cells (×10 magnification; scale
bar, 200 µm. (H) Western blotting of proteins related to
epithelial-mesenchymal transition in HEC-1-B and Ishikawa cells
transfected with si-NC, si-POLQ#1 and si-POLQ#2. For each cell
line, western blot analyses were performed using samples derived
from the same experimental panel and normalized to the same loading
control antibody (β-tubulin). All experiments were performed in
triplicate. **P<0.01, ***P<0.001. NC, negative control; POLQ,
DNA polymerase θ; si, small interfering. |
POLQ regulates apoptosis and cell
cycle progression via the ataxia-telangiectasia mutated (ATM)/P53
pathway in EC cells
Flow cytometric analysis revealed that POLQ
knockdown significantly promoted HEC-1-B and Ishikawa cell
apoptosis compared with that in the si-NC group (Fig. 6A). In addition, POLQ silencing
significantly altered cell cycle distribution, with a decreased
proportion of cells in the G2/M phase, indicating
impaired cell cycle progression (Fig.
6B). Consistently, RT-qPCR analysis demonstrated that knockdown
of POLQ markedly reduced the expression levels of key cell
cycle-related regulators, including P21, cyclin-dependent kinase 1
(CDK1) and cyclin B1 (CCNB1) (Fig.
6C). At the protein level, western blot analysis confirmed that
POLQ knockdown could markedly decrease the protein expression
levels of phosphorylated (P)-P53, P21, CDK1, CCNB1, P-ATM and
P-checkpoint kinase 2 (CHK2) in both HEC-1-B and Ishikawa cells
(Fig. 6D). Collectively, these
results indicated that POLQ could regulate cell cycle progression
and apoptosis in EC cells by modulating the ATM-mediated P53
signaling pathway.
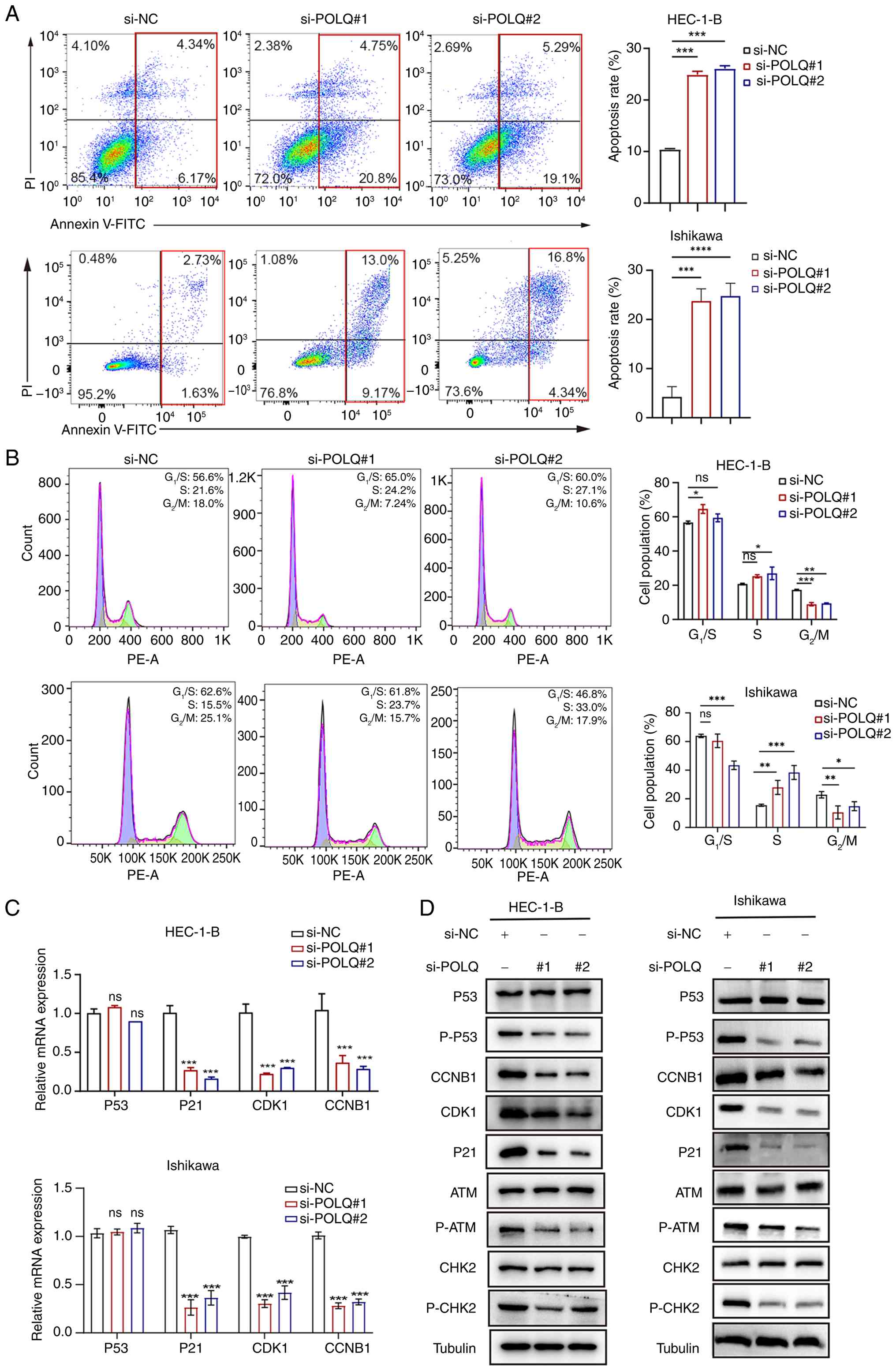 | Figure 6.Silencing POLQ promotes apoptosis,
induces S-phase accumulation with a decreased proportion of cells
in the G2/M-phase, and attenuates ATM-CHK2-mediated DNA
damage response signaling in endometrial cancer cell lines. (A)
Apoptosis analysis and (B) cell cycle analysis of HEC-1-B and
Ishikawa cells transfected with si-NC, si-POLQ#1 and si-POLQ#2. (C)
mRNA and (D) protein expression levels of G2/M-related
genes and the ATM signaling pathway in HEC-1-B and Ishikawa cells.
For each cell line, western blot analyses were performed using
samples derived from the same experimental panel and normalized to
the same loading control antibody (β-tubulin). All experiments were
performed in triplicate. *P<0.05, **P<0.01, ***P<0.001,
****P<0.0001vs. si-NC or as indicated. ATM,
ataxia-telangiectasia mutated; CCNB1, cyclin B1; CDK1,
cyclin-dependent kinase 1; CHK2, checkpoint kinase 2; NC, negative
control; ns, not significant; P-, phosphorylated; PI, propidium
iodide; POLQ, DNA polymerase θ; si, small interfering. |
Discussion
EC is one of the most common gynecological
malignancies, with a steadily increasing global incidence (25). Its pronounced histological and
molecular heterogeneity is reflected by distinct genomic
alterations, including TP53 and POLE mutations, as well as
recurrent somatic copy number alterations such as HER-2
amplification (26,27). It has been reported that POLQ, a
highly conserved DNA polymerase that mediates an alternative DSB
repair pathway, known as POLQ-mediated end joining, is upregulated
in several types of cancer, including gastric cancer, cervical
cancer, pancreatic carcinoma, breast cancer and lung
adenocarcinoma, thus conferring survival advantages to cancer cells
and promoting tumor progression (10,12,14,16,28,29).
The results of the present study revealed significant POLQ
upregulation in EC tumors compared with in normal endometrial
tissues, as evidenced by integrated bioinformatics analysis and
validation in a human tissue cohort. These findings suggested that
POLQ could represent a promising therapeutic strategy for
inhibiting EC initiation and progression.
Tumor cells are frequently subjected to DNA damage,
including DSBs and mutations, which should be efficiently repaired
to maintain cellular function (30). POLQ serves a critical role in
repairing such DNA injuries, thereby supporting tumor cell
survival, continuous proliferation and resistance to cell death.
Acting as a tumor promoter, POLQ can affect several cellular
processes through distinct regulatory networks. For example, Yao
et al (10) demonstrated
that POLQ knockdown could inhibit cell cycle progression by
suppressing the expression of proteins associated with the
G1/M and S/M phases of the cell cycle in colorectal
cancer. In addition, Pan et al (31) reported that POLQ depletion could
disrupt the progression of hepatocellular carcinoma by impairing
cell proliferation, whereas Li et al (32) showed that POLQ silencing suppressed
tumor growth via activation of the cyclic GMP/AMP synthase
(cGAS)/stimulator of interferon genes (STING)/interferon-stimulated
genes pathway in esophageal squamous cell carcinoma. Furthermore,
POLQ has been reported to be involved in maintaining cancer
stemness and conferring resistance to ferroptosis in gastric cancer
cells (12). Collectively, the
aforementioned studies from other malignancies suggest that POLQ
upregulation may be associated with poor survival in EC, primarily
via enhancing DNA repair capacity, sustaining replication and
promoting G2/M cell cycle progression.
Consistent with the aforementioned observations, the
analysis of TCGA-EC cohort in the present study revealed a
significant association between POLQ expression and
clinicopathological features, including clinical stage and
histological grade. Elevated POLQ expression was also significantly
associated with worse OS and DSS, thus highlighting its potential
prognostic value in EC. Consistently, POLQ expression was
positively associated with Ki67+ proliferation index,
further supporting its role in promoting tumor cell proliferation.
Functional experiments further verified that POLQ can enhance EC
cell proliferation and progression. Nevertheless, the relatively
limited sample size underscores the need for validation in larger
cohorts, as well as in vivo studies and potential clinical
trials to fully elucidate the therapeutic relevance of POLQ in
EC.
Tumor-immune homeostasis serves a critical role in
tumor initiation and progression. Activation of the PD-L1 signaling
pathway can enable tumor immune evasion, whereas its inhibition can
enhance endogenous antitumor immunity (33). Notably, POLQ inhibition has been
reported to activate the cGAS/STING pathway, thereby promoting
immune cell infiltration (17,32).
In bladder cancer, the combined assessment of POLQ and PD-L1
expression could serve as a predictor of immunotherapy efficacy
(34), thus suggesting a role for
POLQ in modulating the tumor microenvironment. In line with these
findings, the present study revealed that high POLQ expression was
associated with reduced immune and stromal scores, and diminished
infiltration of antitumor immune cells, including CD8+ T
cells and NK cells. Furthermore, POLQ expression was positively
correlated with PD-L1 levels in both TCGA dataset and validation
cohort, although the correlation in the TCGA dataset was weak,
suggesting a potential association between POLQ and the
immunosuppressive tumor microenvironment in EC. The present study
primarily focused on characterizing the immunosuppressive landscape
associated with POLQ expression, with particular attention to PD-L1
expression and the infiltration of T cells and NK cells. Although
these analyses provide initial insights into the potential
immunoregulatory role of POLQ, a comprehensive evaluation of
immunotherapy response and additional immune cell subtypes was
beyond the scope of the current work. Therefore, further studies
incorporating functional assays, broader immune profiling and
immunotherapy-related models will be required to fully elucidate
the role of POLQ in tumor immune regulation.
GSEA indicated that POLQ was primarily involved in
cell cycle regulation, DNA repair and DNA replication, consistent
with its key role in the DNA damage response (DDR). Previous
studies have suggested that POLQ inhibitors enhance
radiosensitivity in human colorectal cancer (HCT116), non-small
cell lung cancer (H460) and bladder cancer (T24) cell lines, as
well as in mouse xenograft models in preclinical settings (35), highlighting its pivotal function in
DNA repair. DNA damage commonly arising from metabolic processes
and environmental factors can activate DDR mechanisms designated to
maintain genomic stability (36).
In the current study, POLQ expression was associated with the ATM
signaling pathway, a key regulator of multiple DDR cascades. Upon
the occurrence of DSBs, ATM is activated through
autophosphorylation at Ser1981 (37). In vitro experiments
demonstrated that POLQ knockdown inhibited EC cell proliferation,
invasion and metastasis. Mechanistically, POLQ depletion reduced
the phosphorylation of ATM and CHK2, suggesting impaired activation
of the DNA damage response pathway, which may subsequently
influence downstream signaling pathways, including the P53
signaling pathway and its downstream effectors, such as P21, CDK1
and CCNB1. Consequently, POLQ inhibition altered cell cycle
distribution, characterized by a reduced proportion of cells in the
G2/M phase, and promoted apoptosis.
Synthetic lethality represents a key strategy in
anticancer drug development. Preclinical studies have shown that
POLQ inhibitors can act synergistically with poly(ADP-ribose)
polymerase (PARP) inhibitors to overcome resistance to
PARP-targeted therapies (9,38). Recently, POLQ-targeting compounds,
including novobiocin, have been identified as potential anticancer
agents (39). Therefore,
investigating the role of POLQ in EC could provide novel insights
into the development of efficient synthetic lethal therapies, thus
offering novel therapeutic strategies for patients with EC.
Collectively, the current study provided a
comprehensive analysis of POLQ upregulation in EC, demonstrating
its association with poor prognosis based on multiple public
databases and clinical specimens. POLQ expression was positively
correlated with Ki67 and PD-L1, thus supporting its involvement in
tumor cell proliferation and immunosuppression. In vitro
experiments further showed that POLQ could facilitate
G2/M-phase cell cycle progression, activate the
ATM-mediated P53 signaling pathway and promote EMT in EC cells, as
summarized in Fig. 7. Taken
together, these findings highlighted POLQ and its associated
pathways as potential prognostic biomarkers and therapeutic targets
in EC. Although elevated POLQ expression was associated with a poor
prognosis, multivariate Cox regression analysis indicated that POLQ
was not an independent prognostic factor. This represents an
important limitation of the present study, suggesting that POLQ
alone may not be sufficient as a standalone prognostic biomarker
and may instead function as part of a broader regulatory network.
Future studies incorporating larger, independent cohorts,
integrated biomarker analyses and mechanistic investigations will
be required to further define the prognostic relevance of POLQ. In
addition, prospective studies are needed to assess whether POLQ may
have clinical value in combination with other established
prognostic factors.
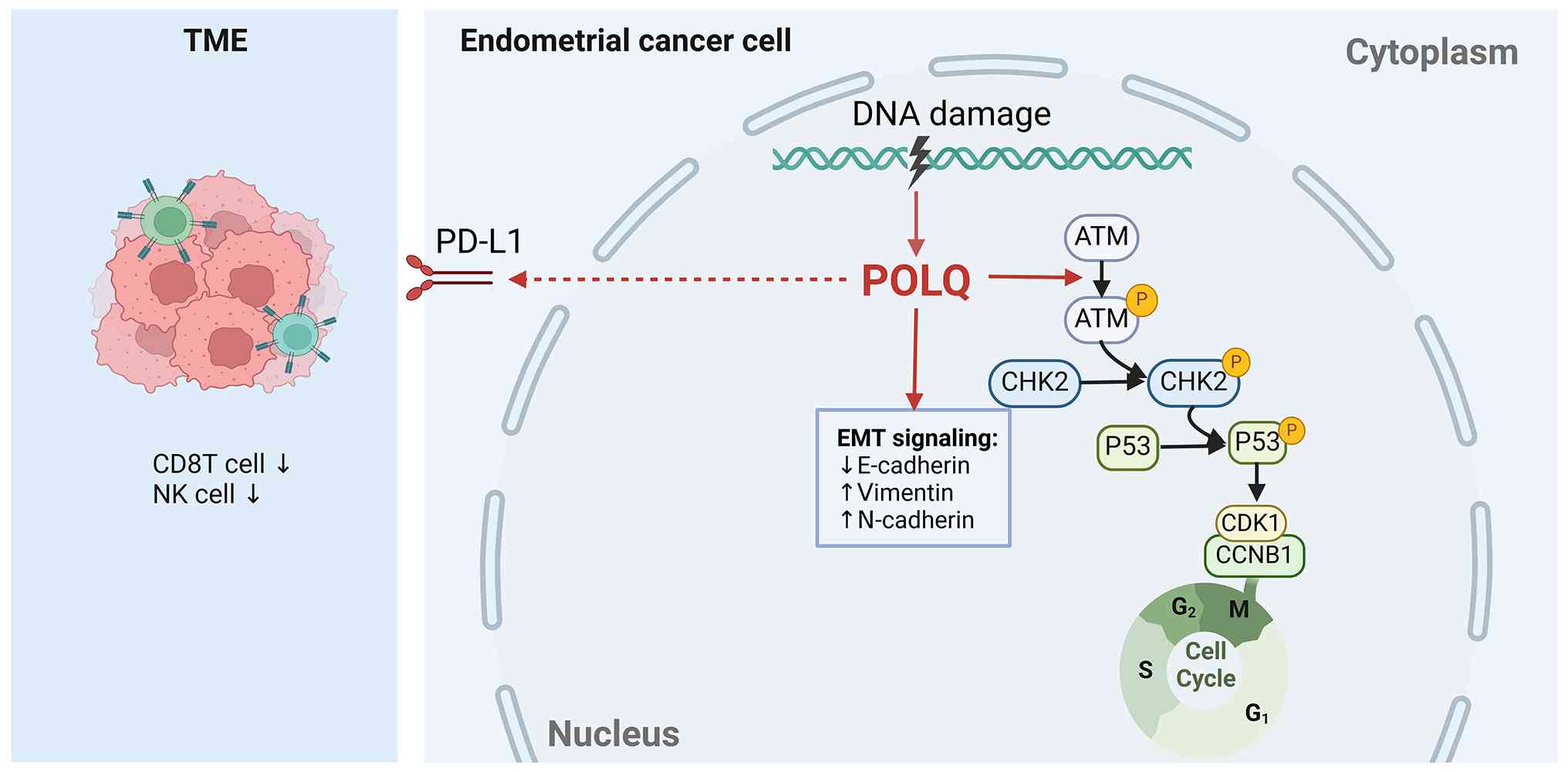 | Figure 7.Schematic diagram. POLQ is
upregulated in EC and drives tumor progression by activating
ATM/P53 signaling, inducing EMT through cadherin switching, and
promoting an immunosuppressive microenvironment via PD-L1
upregulation. POLQ upregulation is associated with aggressive
clinicopathological features in EC, indicating its potential as a
prognostic biomarker and therapeutic target. ATM,
ataxia-telangiectasia mutated; CCNB1, cyclin B1; CDK1,
cyclin-dependent kinase 1; CHK2, checkpoint kinase 2; EMT,
epithelial-mesenchymal transition; NK, natural killer; PD-L1,
programmed death-ligand 1; POLQ, DNA polymerase θ; TME, tumor
microenvironment. |
Supplementary Material
Supporting Data
Acknowledgments
The authors would like to thank Dr Hanmao Tong and
Dr Xianbin Tang (Department of Pediatrics, Affiliated Taihe
Hospital of Hubei University of Medicine, Shiyan, Hubei, China) for
their administrative support in facilitating the ethics approval
process.
Funding
The present study was supported by the Bethune Charity
Foundation (grant no. 4-1-281).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
NZ developed the bioinformatics analysis strategy,
performed the experiments and conducted the statistical analyses.
NZ, JW and XZ performed formal analysis and investigation. NZ and
JW prepared the original draft. NZ, JW and XZ reviewed and edited
the manuscript. YH acquired funding, contributed to the study
conception and critically revised the manuscript. LG was
responsible for providing key experimental materials and technical
resources, and contributed to data acquisition and interpretation.
KT supervised the study, contributed to the experimental design and
interpretation of the data, and critically revised the manuscript.
NZ and JW confirm the authenticity of all the raw data. All authors
contributed to the experiments, and read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Affiliated Taihe Hospital of Hubei University of
Medicine (approval no. 2023KS58; Shiyan, China). All procedures
involving human participants were conducted in accordance with The
Declaration of Helsinki. Written informed consent for the use of
residual pathological specimens for this research and future
related studies was obtained from all patients. All samples were
rigorously de-identified.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Siegel RL, Giaquinto AN and Jemal A:
Cancer statistics, 2024. CA Cancer J Clin. 74:12–49.
2024.PubMed/NCBI
|
|
2
|
Makker V, MacKay H, Ray-Coquard I, Levine
DA, Westin SN, Aoki D and Oaknin A: Endometrial cancer. Nat Rev Dis
Primers. 7:882021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
van den Heerik A, Horeweg N, de Boer SM,
Bosse T and Creutzberg CL: Adjuvant therapy for endometrial cancer
in the era of molecular classification: radiotherapy,
chemoradiation and novel targets for therapy. Int J Gynecol Cancer.
31:594–604. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Wood RD and Doublié S: DNA polymerase θ
(POLQ), double-strand break repair, and cancer. DNA Repair (Amst).
44:22–32. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yousefzadeh MJ, Wyatt DW, Takata K, Mu Y,
Hensley SC, Tomida J, Bylund GO, Doublié S, Johansson E, Ramsden
DA, et al: Mechanism of suppression of chromosomal instability by
DNA polymerase POLQ. PLoS Genet. 10:e10046542014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Seki M, Masutani C, Yang LW, Schuffert A,
Iwai S, Bahar I and Wood RD: High-efficiency bypass of DNA damage
by human DNA polymerase Q. EMBO J. 23:4484–4494. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ceccaldi R, Liu JC, Amunugama R, Hajdu I,
Primack B, Petalcorin MI, O'Connor KW, Konstantinopoulos PA,
Elledge SJ, Boulton SJ, et al: Homologous-recombination-deficient
tumours are dependent on Polθ-mediated repair. Nature. 518:258–262.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mateos-Gomez PA, Gong F, Nair N, Miller
KM, Lazzerini-Denchi E and Sfeir A: Mammalian polymerase θ promotes
alternative NHEJ and suppresses recombination. Nature. 518:254–257.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Zatreanu D, Robinson HMR, Alkhatib O,
Boursier M, Finch H, Geo L, Grande D, Grinkevich V, Heald RA,
Langdon S, et al: Polθ inhibitors elicit BRCA-gene synthetic
lethality and target PARP inhibitor resistance. Nature Commun.
12:36362021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yao Q, Gao S, Sun Q, Liuhua W, Ren J and
Wang D: POLQ knockdown inhibits proliferation, migration, and
invasion by inducing cell cycle arrest in colorectal cancer. Discov
Oncol. 15:6332024. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Shinmura K, Kato H, Kawanishi Y, Yoshimura
K, Tsuchiya K, Takahara Y, Hosokawa S, Kawase A, Funai K and
Sugimura H: POLQ Overexpression Is associated with an increased
somatic mutation load and PLK4 overexpression in lung
adenocarcinoma. Cancers (Basel). 11:7222019. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Peng Y, Zheng W, Chen Y, Lei X, Yang Z,
Yang Y, Liang W, Sun K, Li G and Yu J: POLQ inhibition attenuates
the stemness and ferroptosis resistance in gastric cancer cells via
downregulation of dihydroorotate dehydrogenase. Cell Death Dis.
15:2482024. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kunihisa T, Inubushi S, Tanino H and
Hoffman RM: Induction of the DNA-repair gene POLQ only in
BRCA1-mutant breast-cancer cells by methionine restriction. Cancer
Genomics Proteomics. 21:399–404. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zang Y, Zhao R, Wang T, Gao Y, Chen L, Liu
S, Wang Y and Xue F: Identification of POLQ as a key gene in
cervical cancer progression using integrated bioinformatics
analysis and experimental validation. Mol Med Rep. 27:1152023.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Patterson-Fortin J and D'Andrea AD:
Targeting polymerase theta (POLθ) for cancer therapy. Cancer Treat
Res. 186:285–298. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Del Puerto-Nevado L, Fernández-Aceñero MJ,
Cebrián A, Fatych Y, Díez-Valladares LI, Pérez-Aguirre E, de la
Serna S, García-Botella A, Martínez-Useros J, García-Foncillas J
and Mateos-Gómez PA: POLQ immunostaining behaves as a prognostic
factor for pancreatic carcinoma. Front Oncol. 14:14331792024.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Oh G, Wang A, Wang L, Li J, Werba G,
Weissinger D, Zhao E, Dhara S, Hernandez RE, Ackermann A, et al:
POLQ inhibition elicits an immune response in homologous
recombination-deficient pancreatic adenocarcinoma via cGAS/STING
signaling. J Clin Invest. 133:e1659342023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Bartha Á and Győrffy B: TNMplot.com: A web
tool for the comparison of gene expression in normal, tumor and
metastatic tissues. Int J Mol Sci. 22:26222021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Yoshihara K, Shahmoradgoli M, Martínez E,
Vegesna R, Kim H, Torres-Garcia W, Treviño V, Shen H, Laird PW,
Levine DA, et al: Inferring tumour purity and stromal and immune
cell admixture from expression data. Nat Commun. 4:26122013.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Hänzelmann S, Castelo R and Guinney J:
GSVA: Gene set variation analysis for microarray and RNA-seq data.
BMC Bioinformatics. 14:72013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Huber W, Carey VJ, Gentleman R, Anders S,
Carlson M, Carvalho BS, Bravo HC, Davis S, Gatto L, Girke T, et al:
Orchestrating high-throughput genomic analysis with bioconductor.
Nat Methods. 12:115–121. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci U S A. 102:15545–15550. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Pasanen A, Loukovaara M and Bützow R:
Clinicopathological significance of deficient DNA mismatch repair
and MLH1 promoter methylation in endometrioid endometrial
carcinoma. Mod Pathol. 33:1443–1452. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-ΔΔCT method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Crosbie EJ, Kitson SJ, McAlpine JN,
Mukhopadhyay A, Powell ME and Singh N: Endometrial cancer. Lancet.
399:1412–1428. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Huvila J, Pors J, Thompson EF and Gilks
CB: Endometrial carcinoma: Molecular subtypes, precursors and the
role of pathology in early diagnosis. J Pathol. 253:355–365. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Urick ME and Bell DW: Clinical
actionability of molecular targets in endometrial cancer. Nat Rev
Cancer. 19:510–521. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Higgins GS, Harris AL, Prevo R, Helleday
T, McKenna WG and Buffa FM: Overexpression of POLQ confers a poor
prognosis in early breast cancer patients. Oncotarget. 1:175–184.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Rao X, Xing B, Wu Z, Bin Y, Chen Y, Xu Y,
Zhou D, Zhou X, Wu C, Ye W, et al: Targeting polymerase θ impairs
tumorigenesis and enhances radiosensitivity in lung adenocarcinoma.
Cancer Sci. 114:1943–1957. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Hanahan D: Hallmarks of Cancer: New
dimensions. Cancer Discov. 12:31–46. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Pan Q, Wang L, Liu Y, Li M, Zhang Y, Peng
W, Deng T, Peng ML, Jiang JQ, Tang J, et al: Knockdown of POLQ
interferes the development and progression of hepatocellular
carcinoma through regulating cell proliferation, apoptosis and
migration. Cancer Cell Int. 21:4822021. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Li J, Ko JM, Dai W, Yu VZ, Ng HY, Hoffmann
JS and Lung ML: Depletion of DNA polymerase theta inhibits tumor
growth and promotes genome instability through the cGAS-STING-ISG
pathway in esophageal squamous cell carcinoma. Cancers (Basel).
13:32042021. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Kornepati AVR, Vadlamudi RK and Curiel TJ:
Programmed death ligand 1 signals in cancer cells. Nat Rev Cancer.
22:174–189. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Liu G, Jin K, Liu Z, Su X, Xu Z, Li B, Xu
J, Chang Y, Wang Y, Zhu Y, et al: POLQ identifies a better response
subset to immunotherapy in muscle-invasive bladder cancer with high
PD-L1. Cancer Med. 13:e69622024. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Checcucci E, Bignante G, Volpi G, Liguori
S, Alessio P, Sica M, Ortenzi M, Amparore D, Saliva A, Piana A, et
al: Small-molecule Polθ inhibitors provide safe and effective tumor
radiosensitization in preclinical models. Clin Cancer Res.
29:1631–1642. 2023. View Article : Google Scholar
|
|
36
|
Chatterjee N and Walker GC: Mechanisms of
DNA damage, repair, and mutagenesis. Environ Mol Mutagen.
58:235–263. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Jin MH and Oh DY: ATM in DNA repair in
cancer. Pharmacol Ther. 203:1073912019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Seed G, Beije N, Yuan W, Bertan C, Goodall
J, Lundberg A, Tyler M, Figueiredo I, Pereira R, Baker C, et al:
Elucidating acquired PARP inhibitor resistance in advanced prostate
cancer. Cancer Cell. 42:2113–2123.e2114. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Syed A, Filandr F, Patterson-Fortin J,
Bacolla A, Ravindranathan R, Zhou J, McDonald DT, Albuhluli ME,
Verway-Cohen A, Newman JA, et al: Novobiocin blocks nucleic acid
binding to Polθ and inhibits stimulation of its ATPase activity.
Nucleic Acids Res. 51:9920–9937. 2023. View Article : Google Scholar : PubMed/NCBI
|