Introduction
Esophageal cancer (ESCA) is the 6th leading cause of
cancer-related mortality globally. Tens of thousands of individuals
are diagnosed with ESCA each year (1). Despite significant progress in
surgical, chemotherapy and radiation therapies for ESCA, the
fatality rate remains elevated, with a five-year survival rate not
exceeding 20% (2). An accumulating
body of studies shows that numerous factors, such as smoking
(3), alcohol consumption (4), gastroesophageal reflux disease
(5), the immune system (6) and obesity (7), are related to the initiation and
progression of ESCA. Accordingly, identifying promising biomarkers
for ESCA diagnosis and prognosis is urgent.
Angiopoietins (ANGPT) represent a family of secreted
factors that include ANGPT1/2/3/4 (in humans) (8–10).
ANGPT2 is an antagonistic ligand that maintains the constitutively
static endothelial ANGPT1/TIE2 signaling axis and has been
extensively studied as a candidate molecule for second-generation
anti-angiogenesis drugs (11).
ANGPT2 is not only strongly associated with cardiovascular disease
(CVD) but has also been established as a contributor to the
initiation and progression of lung (12), colon (13), melanoma (14), gastric (15) and renal cell carcinoma (16). Beyond its angiogenic functions,
ANGPT2 plays a documented role in immune modulation within the
tumor microenvironment (TME). Elevated serum ANGPT2 levels
correlate with therapeutic resistance, as they promote both
proangiogenic and immunosuppressive activity (17). Additionally, ANGPT2 sensitizes
TIE2+ M-myeloid-derived suppressor cells (MDSCs) to
impede TAA-specific T cells (18).
In addition, ANGPT2 and VEGFA have been reported to modulate ligand
programmed cell death ligand 1 (PD-L1) (19). However, studies on the prognostic
value and mechanism of ANGPT2 in ESCA are still lacking.
Additionally, the relationship between ANGPT2 and tumor immunity in
ESCA remains unclear.
In the present study, ANGPT2 expression and its
association with survival prediction were first analyzed in various
tumors. Subsequently, the expression of ANGPT2 was assessed in
various ESCA databases and ESCA clinical samples, examining its
clinical relevance and prognosis. An in vitro experiment was
then conducted to illustrate the role of ANGPT2 in ESCA, further
exploring the upstream regulatory factors of ANGPT2 to determine
possible regulatory microRNAs (miRNAs or miRs) and long non-coding
RNAs (lncRNAs). Finally, gene set enrichment analysis (GSEA)
suggested that ANGPT2 is related to immunity and subsequently
revealed the interconnection between ANGPT2 expression and TME
infiltrating characteristics, immune cell (IC) markers, and immune
checkpoints in ESCA. Furthermore, the interaction between ANGPT2
and chemotherapeutic drug sensitivity was analyzed. The findings of
the present study indicate that in patients with ESCA, high
expression of ANGPT2 is involved in tumor immunity and
chemotherapeutic drug sensitivity and suggest a poor prognosis for
patients.
Materials and methods
Patients and tumor tissues
In the present study, a retrospective analysis was
performed on 48 tumors and 21 surrounding tissues [collected
between January 2014 and December 2016 at the Affiliated Hospital
of Jiangsu University (Zhenjiang, China)] acquired from patients
with ESCA who did not receive chemotherapy or radiotherapy. The
present study was approved (approval no. KY201901) by the Ethics
Committee of The Affiliated Hospital of Jiangsu University
(Zhenjiang, China). Before undergoing surgical resection, all
patients signed an informed consent form. Furthermore, apart from
the verification by pathologists working independently, tissue
samples were preserved in liquid nitrogen at −80°C until they were
utilized for reverse transcription-quantitative PCR (RT-qPCR) and
western blotting. The inclusion and exclusion criteria for the
selection of patient samples with ESCA was listed as follows:
Inclusion criteria: (i) Aged ≥18 years old, with no
sex restriction; (ii) diagnosed with primary ESCA by
histopathological examination at the Affiliated Hospital of Jiangsu
University from 2014 to 2016, with a clear and confirmed
pathological diagnosis; (iii) scheduled for surgical resection of
ESCA, without receiving neoadjuvant chemotherapy, radiotherapy,
targeted therapy, immunotherapy or other antitumor treatments
before surgery; (iv) no history of malignant tumors, and no primary
malignant tumors in other organs or systems; (v) voluntarily signed
a written informed consent form before surgical treatment and
agreed to participate in the present study; (vi) tumor tissue
samples and paired adjacent normal esophageal tissue samples were
successfully collected, and the sample quality was verified to be
qualified by independent pathologists, which could be used for
subsequent experimental detection such as RT-qPCR, western blotting
and immunohistochemistry (IHC).
Exclusion criteria: (i) Under 18 years old; (ii)
unclear pathological; (iii) diagnosis, or diagnosed as metastatic
tumors involving the esophagus rather than primary ESCA; (iv)
complicated with primary malignant tumors in other organs or
systems, or with a history of malignant tumors; (v) the collected
tumor tissue or adjacent normal tissue samples were contaminated,
degraded, or insufficient in quantity, which could not meet the
requirements of subsequent experimental detection; (vi) refused to
sign the informed consent form, or voluntarily withdrew from the
study during the research process; (vii) complicated with severe
underlying diseases such as severe liver and kidney failure,
serious cardiovascular and cerebrovascular diseases, uncontrolled
infectious diseases, which may affect the collection of clinical
samples and the judgment of research results.
Cell lines
Cell lines TE-1 (cat. no. ZQ0235), KYSE30 (cat. no.
ZQ09963) and KYSE150 (cat. no. ZQ0449; all from Shanghai Zhongqiao
Xinnzhou Biotechnology Co., Ltd.) and the immortalized normal
esophageal squamous epithelium (Het-1A; cat. no. BNCC342346; BeNa
Culture Collection) were authenticated using Short Tandem Repeat
analysis. All cell lines were used for experiments at passage
numbers 8–15 to ensure consistent phenotypic and functional
stability. Mycoplasma contamination was routinely screened using
the Myco-Visible Mycoplasma LAMP Detection Kit (MP Biomedicals,
LLC) targeting the conserved 16S rRNA gene, and all cell lines
tested negative before experimentation. TE-1, KYSE30, KYSE150, 293T
and Het-1A cell lines were cultured in RPMI-1640 and DMEM media
with 10% fetal bovine serum (FBS; Biological Industries) and 100
IU/ml of penicillin and streptomycin (all from Gibco; Thermo Fisher
Scientific, Inc.) and incubated at 37°C in 5% CO2.
Analysis of RT-qPCR using quantitative
methods
Total RNA isolation from ESCA tissues and cells was
performed using the TRIzol reagent (Takara Biotechnology Co., Ltd.)
according to the manufacturer's protocols. To measure relative RNA
levels, the BioMate 3S Analyzer (Thermo Fisher Scientific, Inc.)
was used. The next step involved performing RT-qPCR reactions using
a reverse transcription kit, along with Cham Q SYBR master mix
(both from Vazyme Biotech Co., Ltd.). The listed primers were as
follows: ANGPT2 forward, 5′-AACTTTCGGAAGAGCATGGAC-3′ and reverse,
5′-CGAGTCATCGTATTCGAGCGG-3′; and β-actin forward,
5′-TCACCCACACTGTGCCCATCTACGA-3′ and reverse
5′-CAGCGGAACCGCTCATTGCCAATGG-3′. The thermal cycling conditions for
RT-qPCR were set as follows: initial denaturation at 95°C for 30
sec; followed by 40 cycles of denaturation at 95°C for 10 sec and
annealing/extension at 60°C for 30 sec; a subsequent melting curve
analysis was conducted (95°C for 15 sec, 60°C for 1 min, and 95°C
for 15 sec) to verify the specificity of the PCR amplicons, with
all procedures performed on a real-time PCR detection system. Then,
the 2−ΔΔCq methodology was applied to calculate the
relative expression of ANGPT2, using β-actin as an internal
reference gene (20). Each
experiment had three identical holes, and all experiments were
conducted thrice.
Western blotting
Cell lysis was performed in RIPA buffer containing
PMSF (Beijing Solarbio Science Technology Co., Ltd.) according to
the protocol. A BCA protein assay (CoWin Biosciences) was employed
to determine protein concentration, with the absorbance measured at
562 nm using a Multiskan SkyHigh Microplate Spectrophotometer
(Thermo Fisher Scientific, Inc.,) and protein concentration
calculated via SkanIt Software 6.0 (Thermo Fisher Scientific,
Inc.). Equal amounts of protein (20 µg) were loaded per lane and
separated using 8 and 12% SDS-PAGE and then transferred to a PVDF
membrane. A 1-h blockage of the membrane was conducted at room
temperature using a 5% BSA solution (Beijing Solarbio Science
Technology Co., Ltd.) in TBST containing 0.1% Tween-20 (Beijing
Solarbio Science Technology Co., Ltd.). The membranes were then
incubated overnight at 4°C with primary antibodies diluted in TBST
containing 5% BSA: ANGPT2 (1:1,000; cat. no. sc-74403; Santa Cruz
Biotechnology, Inc.) and GAPDH (1:1,000; cat. no. sc-47724; Santa
Cruz Biotechnology, Inc.). After washing with TBST three times (10
min each), the membranes were placed for 1 h at room temperature
with HRP-conjugated anti-rabbit or anti-mouse antibody (1:5,000;
cat. nos. 7074P2 and 7076P2, respectively; Cell Signaling
Technology, Inc.), subjected to enhanced ECL reagent (Vazyme
Biotech Co., Ltd.), and images were captured using a Chemis
Scope-4300 imager (Clinx Science Instruments Co., Ltd.).
Histological analysis
IHC staining was conducted on formalin-fixed,
paraffin-embedded tissue sections (4-µm-thick) according to
established protocols. Briefly, the incubation of sections was
conducted with a primary mouse monoclonal anti-ANGPT2 (1:50;
Beyotime Institute of Biotechnology; 0.5% BSA/PBS) and with an
HRP-conjugated anti-mouse secondary antibody (1:200; cat. no.
NB7539; R&D Systems, Inc.; 0.5% BSA/PBS). Antigen-antibody
complexes were observed with the DAB Substrate Kit (Abcam). Stained
slides were initially screened at low magnification (×100) to
determine the regions with the highest vascular density. A total of
four representative fields per slide were selected and
independently evaluated in a blinded manner by two investigators
using light microscopes (Olympus Corporation) coupled with
Image-Pro Plus 6.0 software (Media Cybernetics, Inc.). Staining
intensity was scored as: 0 (negative), 1 (weak), 2 (moderate), or 3
(strong). The positively stained cell proportion was scored as
follows: 0 (0%), 1 (<50%), 2 (50–75%), or 3 (>75%). The final
IHC score was computed by multiplying the intensity and proportion
scores. Tumors were classified as exhibiting low ANGPT2 expression
if the score was <6, and high expression if ≥6.
Small interfering RNA (siRNA)
transfection
The siRNAs for ANGPT2 and a negative control (NC) at
a final concentration of 50 nM which were then introduced into TE-1
cells and KYSE150 cells via Lipofectamine 2000 (Thermo Fisher
Scientific, Inc.) at 37°C for 6 h, were provided by Shanghai Gene
Pharma Co., Ltd. After transfection, the cells were cultured in
complete medium for an additional 48 h before subsequent
experimentation, and the transfected cells were further treated
with additional treatments for the required duration and then
subjected to functional determination. The sequences of si-ANGPT2
and si-NC are listed as follows: si-ANGPT2-1 sense,
5′-GCAUUCUGCUGUAUCUCUACCAUUU-3′ and antisense
5′-AAAUGGUAGAGAUACAGCAGAAUGC-3′; si-ANGPT2-2: sense
5′-GGAGAAUAUUGGCUGGGAATT−3′ and antisense,
5′-UUCCCAGCCAAUAUUCUCCTT−3′; si-ANGPT2-3 sense,
5′-GCAUCUACACGUUAACAUUTT−3′ and antisense
5′-AAUGUUAACGUGUAGAUGCTT−3′; and si-NC sense,
5′-UUCUCCGAACGUGUCACGUUU-3′ and antisense,
5′-ACGUGACACGUUCGGAGAAUU-3′. All siRNAs were chemically modified
with 2′-O-methyl (2′-OMe) to enhance stability and reduce
off-target effects, with HPLC purity ≥98%.
Cell migration and invasion
assays
Following siRNA transfection, TE-1 cells and KYSE150
cells were harvested, resuspended, and plated at a density of
7×104 cells/well into the top chambers (8-µm pore
polycarbonate membrane; Corning, Inc.) in serum-free medium. The
bottom chamber medium included 10% FBS as a chemoattractant. For
invasion assays, membranes were pre-coated with 60 µg of Matrigel
(Becton, Dickinson and Company) at 4°C and subsequently incubated
at 37°C for 2 h to allow for gel solidification. Post-incubation at
37°C in a 5% CO2 incubator for 24 or 48 h, the cells
were fixed with 4% paraformaldehyde at room temperature for 15 min
and subsequently stained with 0.5% crystal violet at room
temperature for 20 min. Migratory and invasive cells were
quantified by counting five randomly selected microscopic
fields/well under an inverted light microscope.
Wound healing assay
The siRNA-transfected cells were plated in 6-well
dishes until they reached 80–85% confluence. Afterward, a straight
scratch was made with a 200-µl pipette tip, and then the cells were
cultured in serum-free medium for 24 h. Images of cells were
subsequently captured using an inverted light microscope (Olympus
Corporation).
Dual-luciferase reporter assay
For the dual-luciferase reporter assay, wild-type
(WT) and mutant (MUT) sequences of ANGPT2 (NM_001118887.2),
MAPKAPK5-AS1 (NR_015404.2), and SNHG1 (ENST00000535076.6) were
first synthesized via PCR-based Accurate Synthesis (PAS) with
full-length overlap primers. The MUT constructs harbored a targeted
mutation of ‘AACTGGA’ to ‘TTGACCT’. These sequences were cloned
into the psicheck2.0 vector (cat. no. ZVE1012; Zaiji Biotechnology;
http://www.zolgene.com) at the Xho I and
NotI restriction sites, inserted downstream of the
Renilla luciferase (RLU) gene with Firefly luciferase (FLU)
serving as an internal control. Recombinant vectors were
transformed into DH5α competent cells, and positive clones were
validated by Xho-ApaI double digestion and Sanger
sequencing. 293T cells (cat. no. ZCL1005; Zaiji Biotechnology) were
cultured in DMEM-H medium (cat. no. C11995500bt; Gibco; Thermo
Fisher Scientific, Inc.) supplemented with 10% FBS (cat. no.
P30-3302; PAN Biotech UK, Ltd.), 1X Penicillin-Streptomycin (cat.
no. 15140122; Gibco; Thermo Fisher Scientific, Inc.), 1X
L-Glutamine (cat. no. 25030-081; Gibco; Thermo Fisher Scientific,
Inc.), 1X Sodium Pyruvate (cat. no. 11360-070; Gibco; Thermo Fisher
Scientific, Inc.) and 1X MEM NEAA (cat. no. 11140-050; Gibco;
Thermo Fisher Scientific, Inc.) at 37°C in a humidified atmosphere
with 5% CO2. Cells were passaged at 80% confluence at a
1:3 ratio following trypsinization (0.25% Trypsin-EDTA; cat. no.
25200056; Gibco; Thermo Fisher Scientific, Inc.) and seeded into
6-well plates 24 h prior to transfection to achieve 60–70%
confluency at the time of transfection. Transfection was performed
using Lipofectamine3000 (cat. no. L3000-015; Invitrogen; Thermo
Fisher Scientific, Inc.) with two concentration gradients of
hsa-miR-145-5p mimic or miR-negative control (miR-NC; 10 µl/well,
20 µM). Experimental groups included WT/MUT recombinant vector +
hsa-miR-145-5p, while control groups consisted of recombinant
vector + miR-NC and empty psicheck2.0 vector +
hsa-miR-145-5p/miR-NC. Culture medium was replaced with fresh
complete medium 6 h post-transfection. A total of 48 h after
transfection, cells were lysed with 1X Lysis Buffer (from
dual-luciferase reporter assay kit; cat. no. DD1205, Vazyme Biotech
Co., Ltd.), and luciferase activities were measured using a
SpectraMax iD5 multi-mode microplate reader (Molecular Devices,
LLC) with freshly prepared FLU working solution (150 µg/ml) and RLU
working solution (100 µM). RLU activity was calculated as the ratio
of RLU activity to FLU activity. All experiments were performed in
triplicate, and statistical significance was determined by
Student's t-test with P<0.05 considered to indicate a
statistically significant difference.
The Cancer Genome Atlas (TCGA)
database download
Data on gene expression and associated prognostic
and clinicopathological information were obtained by accessing TCGA
(https://genome-cancer.ucsc.edu/) for the
following cancer types: bladder urothelial carcinoma (BLCA), breast
invasive carcinoma (BRCA), cholangiocarcinoma (CHOL), esophageal
carcinoma (ESCA), glioblastoma multiforme (GBM), head and neck
squamous cell carcinoma (HNSC), kidney renal clear cell carcinoma
(KIRC), kidney renal papillary cell carcinoma (KIRP), liver
hepatocellular carcinoma (LIHC), lung adenocarcinoma (LUAD), lung
squamous cell carcinoma (LUSC), pancreatic adenocarcinoma (PRAD),
rectum adenocarcinoma (READ), stomach adenocarcinoma (STAD), kidney
chromophobe (KICH), thyroid carcinoma (THCA), colon adenocarcinoma
(COAD) and uterine corpus endometrial carcinoma (UCEC). The
fragments per kilobase million (FPKM) measurement was transformed
to one million transcripts per kilobase (TPM), which represented
the same transcript quantity as the microarray.
Gene expression profiling interactive
analysis (GEPIA) database analysis
Using TCGA and GTEx data, GEPIA (http://gepia.cancer-pku.cn/) was employed to analyze
tumor and normal gene expression profiling and to analyze the
survival of ANGPT2 in 7 human tumors, including overall survival
(OS) and disease-free survival (DFS), with P<0.05 considered to
indicate a statistically significant difference.
Gene Expression Omnibus (GEO;
http://www.ncbi.nlm.nih.gov/) database
analysis
GEO databases, including GSE20347, GSE38129,
GSE45670 and GSE70409, were deployed to analyze ANGPT2 mRNA levels
between ESCA cancer and normal tissues. Single-cell sequencing data
(GSE160269) were used to analyze ANGPT2 expression in the TME, with
P<0.05 considered to indicate a statistically significant
difference.
Nomogram scoring system
A prognostic nomogram was constructed using clinical
characteristics and risk scores derived from multivariate survival
analysis, implemented using the ‘rms’ package. The nomogram's
calibration was assessed by plotting the predicted probabilities of
1-, 3-, and 5-year OS against the corresponding observed survival
rates, allowing for a visual evaluation of predictive accuracy
across different time points.
StarBase database download
StarBase [ENCORI: The Encyclopedia of RNA
Interactomes (sysu.edu.cn)] is used to predict miRNAs upstream of
ANGPT and IncRNAs upstream of miR-145-5p. The regulatory network of
the predicted miRNAs and ANGPT2 was visualized using Cytoscape
software (v3.9.1; Cytoscape Consortium; http://cytoscape.org/).
GSEA
The study initially sorted all genes based on their
correlation with ANGPT2 expression and employed GSEA to elucidate
the notable disparity in survival rates between the high and low
ANGPT2 groups. Each analysis involved executing 1,000 permutations
on the genome. ANGPT2 expression was utilized as a phenotypic
marker. Each phenotype was classified based on the enrichment
pathways using the nominal p-value and the normalized enrichment
score (NES).
Evaluation of the immune infiltration
and TME in ANGPT2 expression cohorts
Using single-sample GSEA (ssGSEA), 23 subgroups of
infiltrating ICs were quantified in both expression groups to
assess the tumor-infiltrating IC proportion in the TME. The
‘ESTIMATE’ was used to assess the TME scores (immune, stromal and
estimated) of both groups.
Assessment of the immune infiltration
and TME between ANGPT2 expression groups
To evaluate the tumor-infiltrating immune cell
(TIIC) proportion in the TME, ssGSEA was utilized to determine the
abundance of 23 infiltrating IC subgroups from both expression
groups. The ‘ESTIMATE’ was performed to assess the TME scores of
both groups.
Drug sensitivity analysis
To evaluate the differential sensitivity to
chemotherapeutic agents between high-and low-ANGPT2 expression
groups in ESCA, the pRRophetic algorithm was utilized to predict
the half-maximal inhibitory concentration (IC50) values
of clinically commonly used chemotherapeutic drugs for ESCA.
Briefly, transcriptomic data (FPKM format) of patients with ESCA
from TCGA database were first converted to TPM values for
standardization, followed by calibration of the pRRophetic model
using the Cancer Cell Line Encyclopedia (CCLE) dataset to ensure
the reliability of IC50 predictions. The preprocessed
TCGA-ESCA transcriptomic data were then input into the calibrated
model to predict IC50 values of first-line
chemotherapeutic drugs for ESCA, including cisplatin, docetaxel,
doxorubicin, erlotinib, paclitaxel and vinorelbine. Statistical
comparison of the predicted IC50 values between the two
ANGPT2 expression groups was performed using the Mann-Whitney U
test with Holm-Bonferroni correction for multiple comparisons,
where a lower IC50 value indicates higher sensitivity to
the corresponding chemotherapeutic drug.
TIMER2.0 database analysis
The correlation between ANGPT2 expression in ESCA
and IC infiltration level or immune checkpoint expression was
analyzed using TIMER 2.0 [TIMER2.0 (comp-genomics.org)], with
P<0.05 deemed to indicate a statistically significant
difference.
Statistical analysis
Statistical analyses were performed using R software
(v.4.0.5; Foundation for Statistical Computing; http://www.r-project.org/) and GraphPad Prism
(v.9.0.0; Dotmatics). The Wilcoxon rank-sum test (unpaired) or
Wilcoxon signed-rank test (paired) was used to compare mRNA, miRNA,
lncRNA, or protein expression levels between tumor and normal
tissues. The relationship between ANGPT2 expression and
clinicopathological characteristics was analyzed via logistic
regression. Kaplan-Meier curves were plotted to visualize OS and
DFS, with differences compared using the log-rank test. The Cox
proportional hazards model was used for univariate and multivariate
analyses to identify independent prognostic factors, with the
proportional hazards' assumption verified using Schoenfeld
residuals. Spearman's correlation coefficient was applied for
correlation analyses between ANGPT2 expression and immune cell
infiltration, immune checkpoints, or target miRNAs/lncRNAs.
For multiple comparison analyses, appropriate
correction methods were applied to control the family-wise error
rate (FWER) or false discovery rate (FDR): (i) When comparing
ANGPT2 expression across multiple tumor types (TCGA pan-cancer
analysis) or among different clinicopathological subgroups (for
example, tumor grade and lymph node status), Bonferroni correction
was used to adjust the raw P-values for FWER. For correlation
analyses involving multiple miRNAs/lncRNAs or multiple immune cell
markers/immune checkpoints, Benjamini-Hochberg (BH) correction was
implemented to control the FDR. (ii) In drug sensitivity analysis,
Holm-Bonferroni correction was adopted for comparing
IC50 values of multiple chemotherapeutic agents.
For in vitro functional assays and western
blotting, data were presented as the mean ± SD from at least three
independent biological replicates. Outliers were excluded using the
Grubbs test (α=0.05), and differences between groups were assessed
by unpaired Student's t-test or Mann-Whitney U test, with
appropriate multiple comparison correction as specified. ssGSEA for
immune infiltration was performed with 1,000 permutations, and
nomogram calibration was evaluated using the Hosmer-Lemeshow test
and C-index (bootstrap resampling, n=1,000). P<0.05 was
considered to indicate a statistically significant difference, with
all P-values adjusted for multiple comparisons unless otherwise
stated.
Results
Pan-cancer analysis of ANGPT2
expression
Using the TCGA pan-cancer transcriptomic dataset,
the analysis illustrated that ANGPT2 was significantly upregulated
in 13 tumor types: BRCA, CHOL, COAD, ESCA, GBM, HNSC, LUAD, KIRC,
KIRP, LIHC, READ, STAD and THCA, compared with normal tissues
(Fig. 1A). No significant
differential expression was detected in BLCA, KICH, LUSC, PRAD and
UCEC tumors. Validation via the GEPIA platform further confirmed
elevated ANGPT2 expression in CHOL, ESCA, HNSC, GBM, KICH, KIRC and
STAD relative to normal tissues (Fig.
1B-H). ANGPT2 was upregulated in CHOL, ESCA, HNSC, GBM, KIRC
and STAD, implicating it as a potential driver of oncogenesis and
disease progression in these contexts.
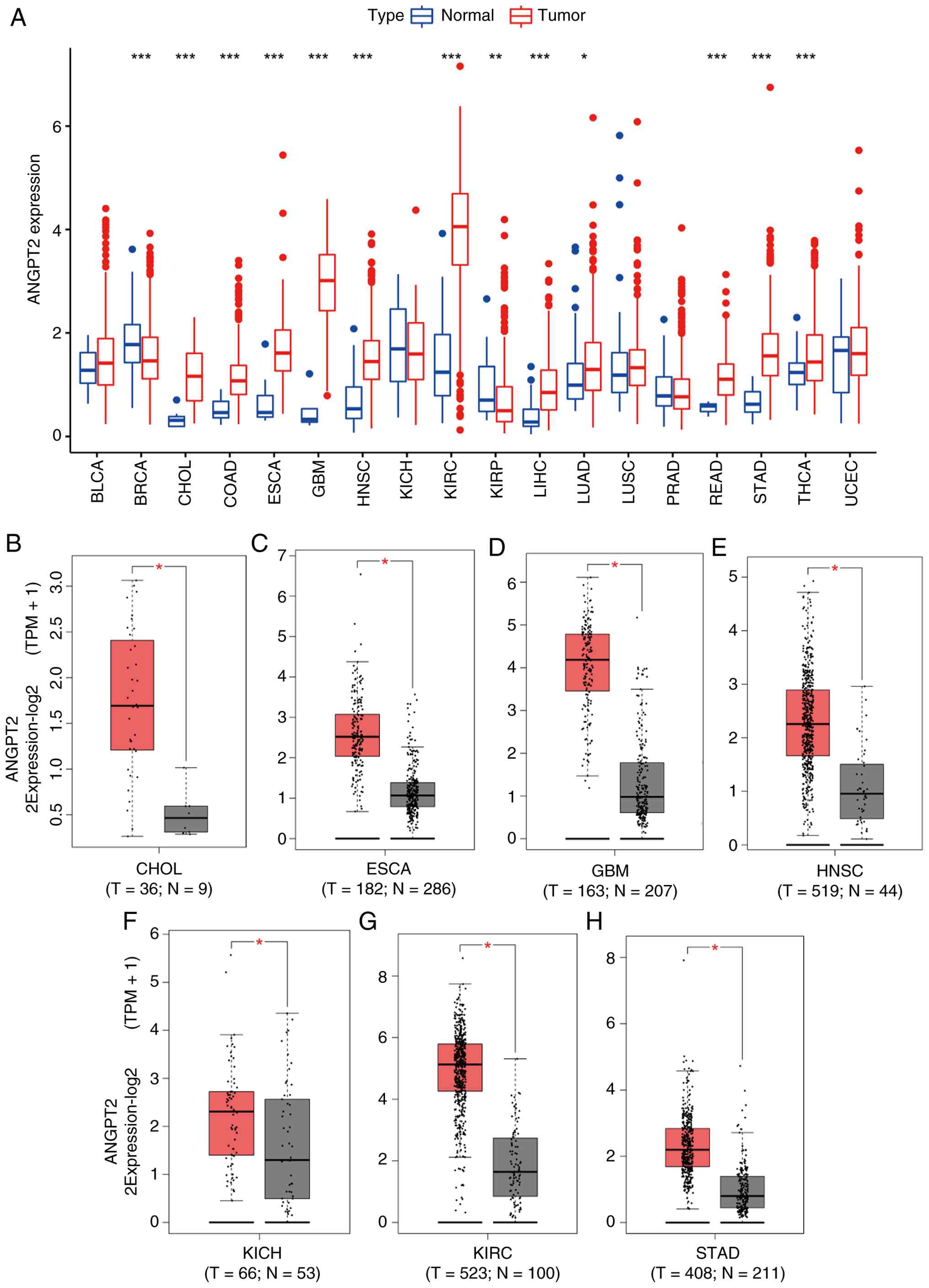 | Figure 1.ANGPT2 expression in multiple tumors.
(A) ANGPT2 levels in 18 human tumors using TCGA tumor and normal
data. (B-H) ANGPT2 levels in TCGA; (B) CHOL, (C) ESCA, (D) GBM, (E)
HNSC, (F) KICH, (G) KIRC and (H) STAD tissues vs. the matching TCGA
and GTEx normal tissues. *P<0.05, **P<0.01 and ***P<0.001;
all P-values are adjusted for multiple comparisons where
applicable. ANGPT2, angiopoietin-2; TCGA, The Cancer Genome Atlas;
CHOL, cholangiocarcinoma; ESCA, esophageal cancer; GBM,
glioblastoma multiforme; HNSC, head and neck squamous cell
carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell
carcinoma; STAD, stomach adenocarcinoma. |
Prognostic analysis of ANGPT2 in
tumors
Through the GEPIA database, the ANGPT2 prognostic
value was ascertained in CHOL, ESCA, HNSC, GBM, KIRC and STAD,
assessing both OS and DFS, revealing that overexpressed ANGPT2 was
significantly linked to worse OS in ESCA and STAD (Fig. 2A-G). By contrast, for DFS, high
ANGPT2 expression predicted adverse outcomes exclusively in ESCA
(Fig. 2H-N). Further analysis of
TCGA clinical data (Table I)
confirmed a significant link between ANGPT2 levels and survival
outcomes, specifically in ESCA. These findings suggest that ANGPT2,
particularly when combined with OS and DFS metrics, serves as a
robust biomarker of poor prognosis in patients with ESCA.
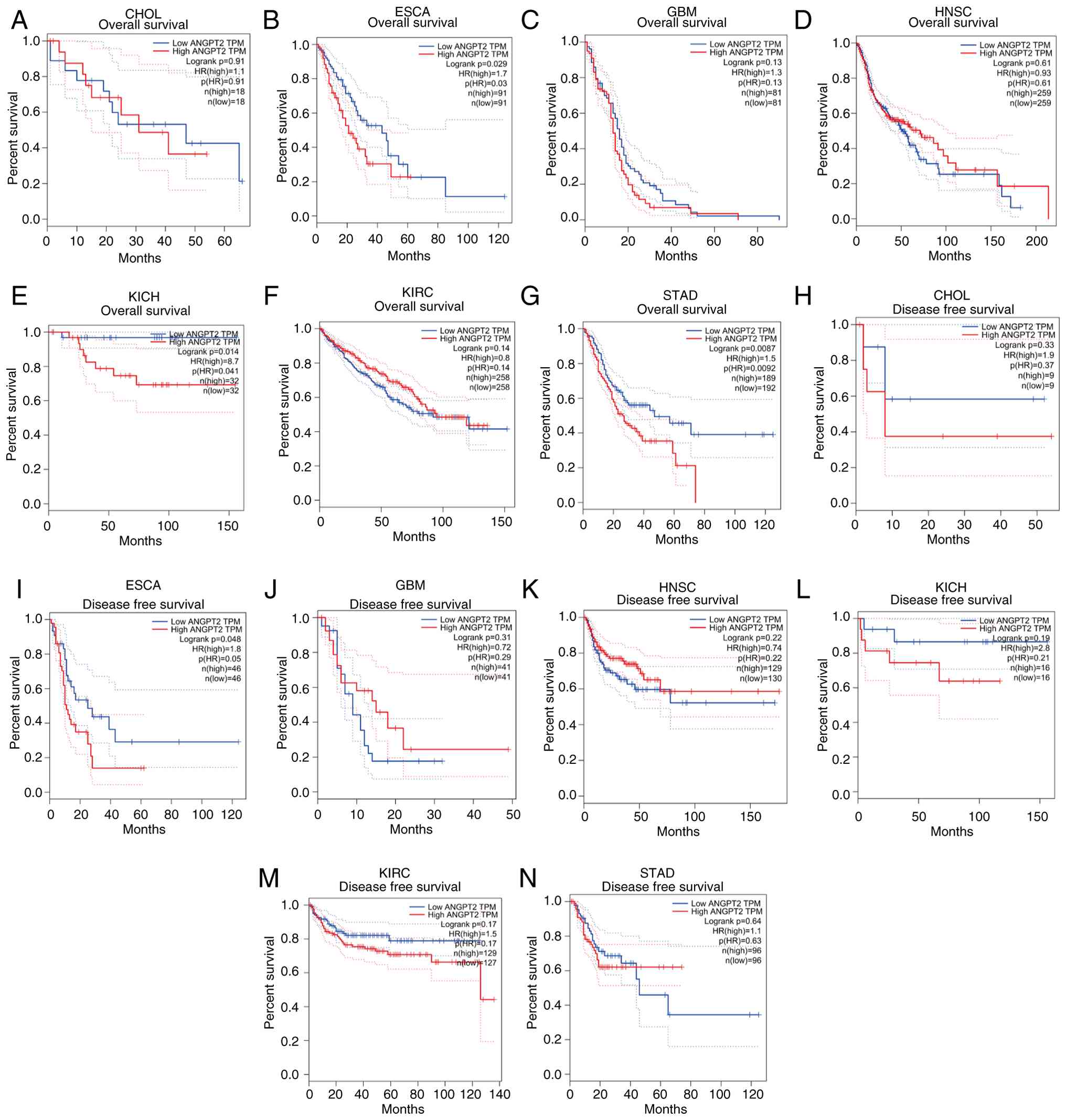 | Figure 2.Gene expression profiling interactive
analysis database: OS and DFS analysis of ANGPT2 in diverse human
tumors. (A-G) OS plot: ANGPT2 in (A) CHOL, (B) ESCA, (C) GBM, (D)
HNSC, (E) KICH, (F) KIRC and (G) STAD. (H-N) DFS plot: ANGPT2 in
(H) CHOL, (I) ESCA, (J) GBM, (K) HNSC, (L) KICH, (M) KIRC and (N)
STAD. OS, Overall survival; DFS, disease-free survival; ANGPT2,
angiopoietin-2; CHOL, cholangiocarcinoma; ESCA, esophageal cancer;
GBM, glioblastoma multiforme; HNSC, head and neck squamous cell
carcinoma; KICH, kidney chromophobe; KIRC, kidney renal clear cell
carcinoma; STAD, stomach adenocarcinoma. |
 | Table I.Relationship between differentially
expressed genes and patient survival in esophageal cancer. |
Table I.
Relationship between differentially
expressed genes and patient survival in esophageal cancer.
| Gene | KM | HR | HR.95L | HR.95H | Cox P-value |
|---|
| GPER1 |
5.8×10−4 | 0.56 | 0.34 | 0.94 |
3.0×10−2 |
| EFNA1 |
9.3×10−4 | 1.51 | 1.08 | 2.13 |
1.8×10−2 |
| ABRACL |
2.1×10−3 | 1.88 | 1.30 | 2.71 |
7.5×10−4 |
| PIGU |
5.5×10−3 | 1.88 | 1.06 | 3.31 |
3.0×10−2 |
| MT1E |
6.5×10−3 | 0.84 | 0.73 | 0.97 |
1.7×10−2 |
| PGK1 |
9.2×10−3 | 1.81 | 1.12 | 2.91 |
1.4×10−2 |
| LINC00365 |
1.0×10−2 | 1.62 | 1.06 | 2.50 |
1.5×10−2 |
| SNRPB |
1.5×10−2 | 1.72 | 1.11 | 2.67 |
1.4×10−2 |
| HSPD1 |
1.6×10−2 | 1.74 | 1.22 | 2.48 |
2.3×10−3 |
| ANGPT2 |
1.8×10−2 | 1.35 | 1.00 | 1.82 |
4.8×10−2 |
| GLA |
2.2×10−2 | 1.65 | 1.17 | 2.34 |
4.5×10−3 |
| HSP90AB1 |
2.3×10−2 | 1.45 | 1.17 | 1.98 |
1.8×10−2 |
| SPDL1 |
2.5×10−2 | 1.60 | 1.02 | 2.50 |
3.8×10−2 |
| CACYBP |
2.5×10−2 | 1.74 | 1.07 | 2.82 |
2.3×10−2 |
| H1-4 |
2.8×10−2 | 1.40 | 1.06 | 1.87 |
1.9×10−2 |
| PTGSE3 |
3.0×10−2 | 2.59 | 1.47 | 4.57 |
1.0×10−3 |
| MRPL47 |
3.7×10−2 | 1.57 | 1.03 | 2.39 |
3.5×10−2 |
Patient characteristics
To assess ANGPT2 expression in ESCA, clinical and
transcriptomic data were retrieved from 183 patients with ESCA in
the TCGA database, which included their demographics and
clinicopathological characteristics, as shown in Table II.
 | Table II.Clinical characteristics of patients
with esophageal carcinoma from The Cancer Genome Atlas. |
Table II.
Clinical characteristics of patients
with esophageal carcinoma from The Cancer Genome Atlas.
| Clinical
characteristics | Total (n=183) | Percentage, % |
|---|
| Age, years |
|
|
|
≤60 | 91 | 49.7 |
|
>60 | 92 | 50.3 |
| Sex |
|
|
|
Female | 27 | 14.8 |
|
Male | 156 | 85.2 |
| Grade |
|
|
| G1 | 19 | 13.3 |
| G2 | 76 | 53.1 |
| G3 | 48 | 33.6 |
| Stage |
|
|
| I | 18 | 11.3 |
| II | 78 | 48.7 |
|
III | 55 | 34.4 |
| IV | 9 | 5.6 |
| T |
|
|
| T1 | 31 | 18.8 |
| T2 | 43 | 26.1 |
| T3 | 86 | 52.1 |
| T4 | 5 | 3 |
| N |
|
|
| N0 | 76 | 46.6 |
| N1 | 68 | 41.7 |
| N2 | 12 | 7.4 |
| N3 | 7 | 4.3 |
| M |
|
|
| M0 | 134 | 93.7 |
| M1 | 9 | 6.3 |
Correlation between clinical analysis
and ANGPT2 expression
To evaluate ANGPT2 expression in ESCA,
transcriptomic data from four independent GEO datasets (GSE20347,
GSE38129, GSE45670 and GSE70409) were first analyzed. In all
cohorts, ANGPT2 mRNA levels displayed significant overexpression in
ESCA tissues relative to neighboring healthy tissues (GSE20347:
P=0.0003; GSE38129: P=0.0067; GSE45670: P=0.0032; GSE70409:
P=0.0003; Fig. 3A-D). Afterwards,
the TCGA database was utilized to analyze 171 ESCA samples,
resulting in the discovery of ANGPT2 expression data for every
individual. According to the paired t-test, the ANGPT2 expression
level in tumors was elevated compared with healthy tissues
(P=0.002; Fig. 3E). The outcomes
illustrated a significant relationship between ANGPT2 levels and
the histological tumor grade (P=0.033; Fig. 3F). The survival analysis indicated
that ESCA with elevated levels of ANGPT2 had a more unfavorable
prognosis compared with ESCA with lower levels of ANGPT2 (P=0.044;
Fig. 3G). In the analysis that
considered multiple variables, ANGPT2 continued to be linked to OS
independently, with a hazard ratio (HR) of 1.06 (Confidence
interval: 1.00–1.1; P=0.038), in addition to lymph node status and
sex (Fig. 3H). Next, a nomogram
that included ANGPT2 expression and clinicopathological factors was
developed for predicting the survival rates at 1, 3, and 5 years
(Fig. 3I). IHC analyses of ANGPT2
protein levels in ESCA tumors and normal tissues revealed that
ANGPT2 expression was significantly higher in cancerous tissues
(Fig. 3J and K). Furthermore,
IHC-based analysis further linked elevated ANGPT2 protein levels to
lymph node metastasis and advanced histological grade (Table III). Concordantly, ANGPT2 mRNA and
protein expression levels were consistently and significantly
upregulated in ESCA tumors relative to normal tissues (Fig. 3L and M). To investigate whether
ANGPT2 has prognostic value in patients with ESCA, data from The
Affiliated Hospital of Jiangsu University (Zhenjiang, China) were
utilized. The findings indicated that individuals diagnosed with
ESCA who had elevated levels of ANGPT2 exhibited a poorer prognosis
(P=0.0002; Fig. 3N).
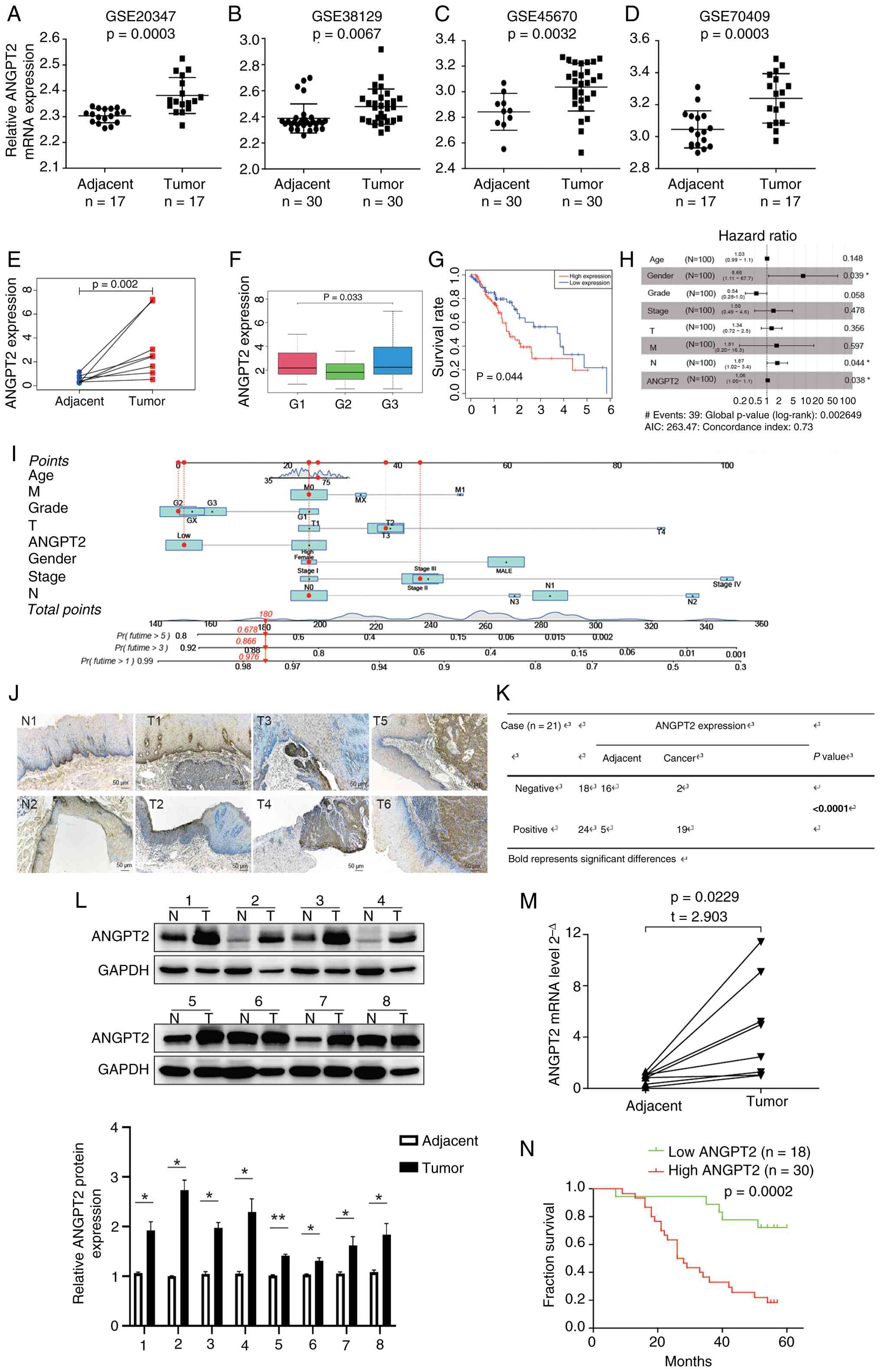 | Figure 3.Association of ANGPT2 expression with
clinical analysis in ESCA. (A-D) ANGPT2 expression in the GSE20347,
GSE38129, GSE45670 and GSE70409 datasets. (E) Paired ANGPT2
expression in adjacent and tumor tissues. (F) Correlation between
ANGPT2 expression and grade. (G) Impact of ANGPT2 expression on OS
in patients with ESCA. (H) Multivariate Cox regression: Connection
with OS and clinicopathologic characteristics in patients with
ESCA. (I) Nomogram for the 1-, 3-, and 5-yr OS prediction of
patients with ESCA. (J) Representative images and (K) statistical
analysis: ANGPT2 expression in ESCA tumors and normal tissues.
Scale bars, 50 µm. (K) Number of ANGPT2 expression negative or
positive cases in normal tissues and ESCC tissues based on the IHC
staining score results. The results demonstrated that 16 cases were
negative and only 5 cases were positive for ANGPT2 expression in
normal tissues, while merely 2 cases were negative and 19 cases
were positive for ANGPT2 expression in ESCC tissues. (L) ANGPT2
protein expression in paired tissues obtained from patients with
ESCA. (M) ANGPT2 mRNA expression levels in ESCA and paired adjacent
normal tissues. (N) Kaplan-Meier: Prognosis of OS according to the
IHC scores of ANGPT2. Statistical analyses: Paired Student's t-test
was used for comparisons between paired tumor and normal tissues
(E, L and M); unpaired Student's t-test was used for comparison of
IHC scores between tumor and normal tissues (K); one-way ANOVA was
used for analyzing ANGPT2 expression across different tumor grades
(F); log-rank test was used for survival analyses (G and N); Cox
proportional hazards model was used for multivariate regression
analysis (H). *P<0.05 and **P<0.01; all P-values are adjusted
for multiple comparisons where applicable. ANGPT2, angiopoietin-2;
ESCA, esophageal cancer; OS, overall survival; ESCC, esophageal
squamous cell carcinoma; IHC, immunohistochemical. |
 | Table III.Specimens assayed for ANGPT2
expression. |
Table III.
Specimens assayed for ANGPT2
expression.
|
|
| Expression level of
ANGPT2 |
|
|---|
|
|
|
|
|
|---|
| Clinicopathological
characteristics | Total (n=48) | Low | High | P-value |
|---|
| Sex |
|
|
| 0.1018 |
|
Male | 31 | 9 | 22 |
|
|
Female | 17 | 9 | 8 |
|
| Age |
|
|
| 0.8776 |
|
<60 | 18 | 7 | 11 |
|
|
≥60 | 30 | 11 | 19 |
|
| Tumor location |
|
|
| 0.8811 |
|
Upper/Middle | 26 | 10 | 16 |
|
|
Lower | 22 | 8 | 14 |
|
|
Differentiation |
|
|
| 0.0214 |
|
Poor | 13 | 3 | 10 |
|
|
Moderately | 11 | 8 | 3 |
|
|
High | 24 | 7 | 17 |
|
| TNM stage |
|
|
| 0.0736 |
|
I–II | 24 | 12 | 12 |
|
|
III–IV | 24 | 6 | 18 |
|
| Lymphatic
metastasis |
|
|
| 0.0110 |
| No | 26 | 14 | 12 |
|
|
Yes | 22 | 4 | 18 |
|
| Distant
metastasis |
|
|
| 0.0913 |
| No | 33 | 15 | 18 |
|
|
Yes | 15 | 3 | 12 |
|
| Recurrence |
|
|
| 0.0538 |
| No | 35 | 16 | 19 |
|
|
Yes | 13 | 2 | 11 |
|
In vitro, ANGPT2 modulates migration
and invasion of TE-1 and KYSE150 cells
The results revealed a significant increase in
ANGPT2 levels in KYSE30, KYSE150 and TE-1 cells, unlike Het-1A
cells (Fig. S1A). To examine the
biological role of ANGPT2 in a laboratory setting, TE-1 cells and
KYSE150 cells were subjected to si-ANGPT2 to suppress ANGPT2. Among
the three ANGPT2-targeting siRNAs transfected into TE-1 and KYSE150
cells, two exhibited significantly higher knockdown efficacy
compared with the non-targeting si-NC group. Notably, si-ANGPT2-3
achieved the most robust silencing effect, with an interference
efficiency exceeding 70% at protein levels (Fig. S1B). The si-ANGPT2 group (Fig. S1C) exhibited a significant
reduction in cells' migratory and invasive abilities, as
demonstrated by Transwell assays. By contrast, the si-ANG2 group
exhibited a reduction in cell migration during wound healing
analysis compared with the control (Fig. S1D). The outcomes demonstrated that
ANGPT2 had a notable impact on diminishing the movement and
infiltration of ESCC cells.
Forecasting and examination of
ANGPT2′s upstream regulatory elements
The non-coding RNAs are well-established modulators
of gene expression. To identify potential upstream regulators of
ANGPT2, StarBase was employed to predict miRNAs with binding
potential to the ANGPT2 3′ untranslated region (3′ UTR), requiring
consensus across at least two prediction algorithms. This yielded
10 candidate miRNAs, which were visualized in a regulatory network
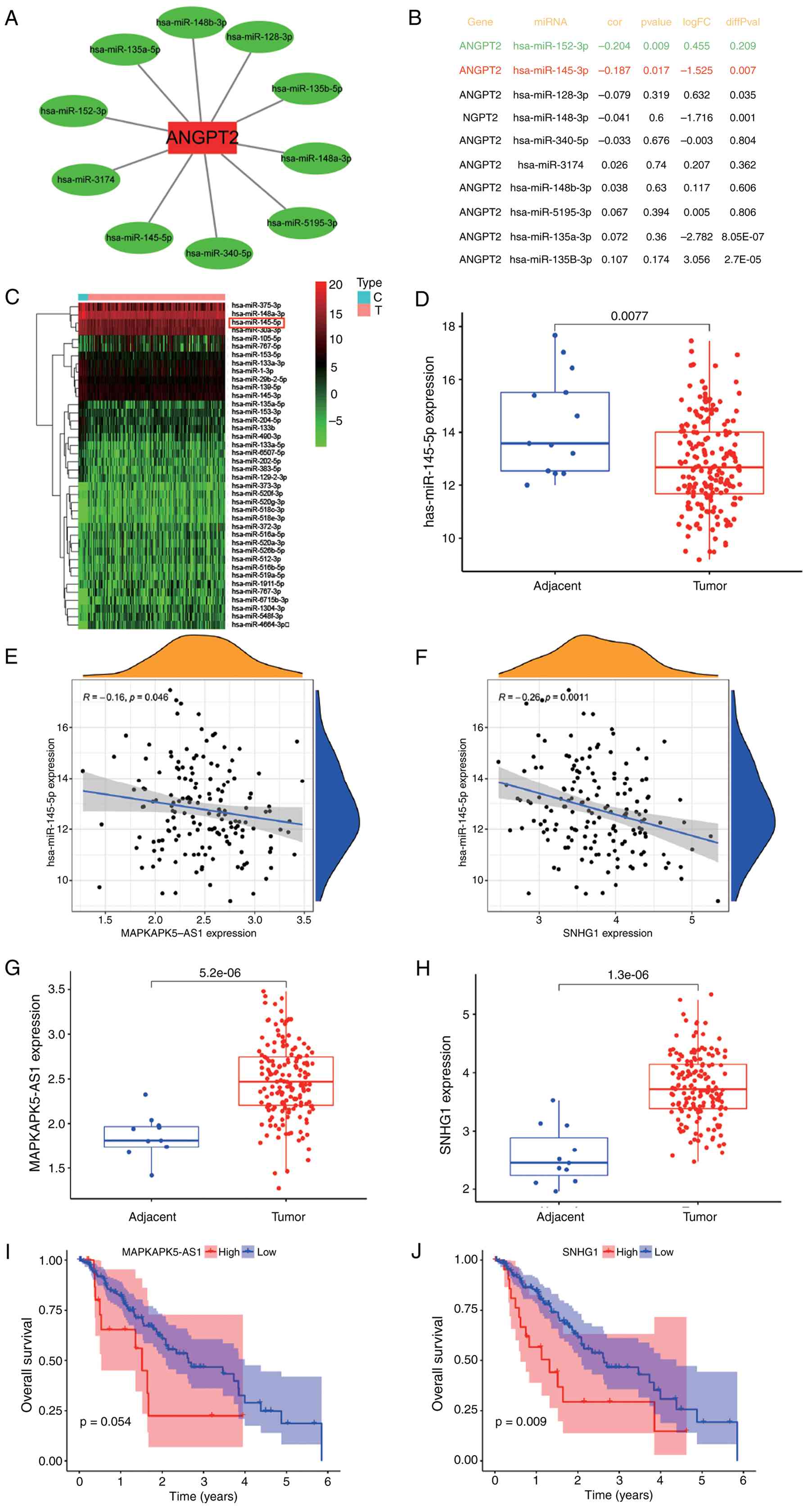
using Cytoscape (v3.9.1; http://cytoscape.org/). (Fig. 4A). Given that miRNAs exert
regulatory control by suppressing their target genes (20), an inverse correlation between
candidate miRNAs and ANGPT2 was hypothesized. Correlation analysis
using TCGA-ESCA data revealed that only miR-152-3p and miR-145-5p
exhibited significant negative correlations with ANGPT2 expression
(Fig. 4B). Of these, only
miR-145-5p was significantly impaired in ESCA tissues (P=0.0077;
Fig. 4C and D), positioning it as
the most plausible direct regulator of ANGPT2 in ESCA. StarBase was
then utilized to forecast the upstream lncRNA of miR-145-5p, to
identify the regulatory factors influencing its expression.
According to the present results, 38 kinds of lncRNAs could bind
miRNAs. miRNAs can induce gene silencing through their binding to
mRNAs, while IncRNAs can regulate gene expression through their
competitive binding to miRNAs. IncRNAs can bind to miRNAs through
miRNA response elements, thereby impacting gene silencing caused by
miRNAs. Hence, an inverse relationship is expected to exist between
lncRNAs and miRNAs. Through the TCGA analysis (Fig. 4E and F), only MAPKAPK5-AS1 (P=0.046)
and SNHG1 (P=0.0011) displayed a negative link with miR-145-5p.
Eventually, the significance and predictive capabilities of
MAPKAPK5-AS1 and SNHG1 were explored in ESCA. As demonstrated in
Fig. 5G-J, MAPKAPK5-AS1 was
significantly upregulated in ESCA (P=5.2×10−6), but its
upregulation did not have a significant influence on patient
prognosis (P=0.054). SNHG1 was also significantly upregulated in
ESCA (P=1.3×10−6), and SNHG1 overexpression in ESCA
suggested that patients had a poor prognosis (P=0.009). Combined
with the aforementioned analysis, MAPKAPK5-AS1 and SNHG1 may
negatively regulate miR-145-5p and promote ANGPT2 expression. To
validate this regulatory axis (MAPKAPK5-AS1/SNHG1 → miR-145-5p →
ANGPT2), dual-luciferase reporter assays were performed. WT and MUT
recombinant vectors of ANGPT2 3′UTR, MAPKAPK5-AS1 and SNHG1 were
constructed, with the binding region sequence mutated from
‘AACTGGA’ to ‘TTGACCT’ (Fig.
S3A-C). The dual-luciferase reporter assay results demonstrated
that miR-145-5p mimic significantly reduced the luciferase activity
of the ANGPT2 3′UTR-WT vector, but had no obvious effect on the MUT
vector; meanwhile, both MAPKAPK5-AS1-WT and SNHG1-WT vectors could
significantly inhibit the inhibitory effect of miR-145-5p mimic on
luciferase activity, while the MUT vectors lost this regulatory
effect (Fig. S3D-F). These
findings confirmed the direct binding of miR-145-5p to the 3′UTR of
ANGPT2, and the specific binding of MAPKAPK5-AS1 and SNHG1 to
miR-145-5p, verifying the existence of the
MAPKAPK5-AS1/SNHG1/miR-145-5p/ANGPT2 ceRNA regulatory axis in
ESCA.
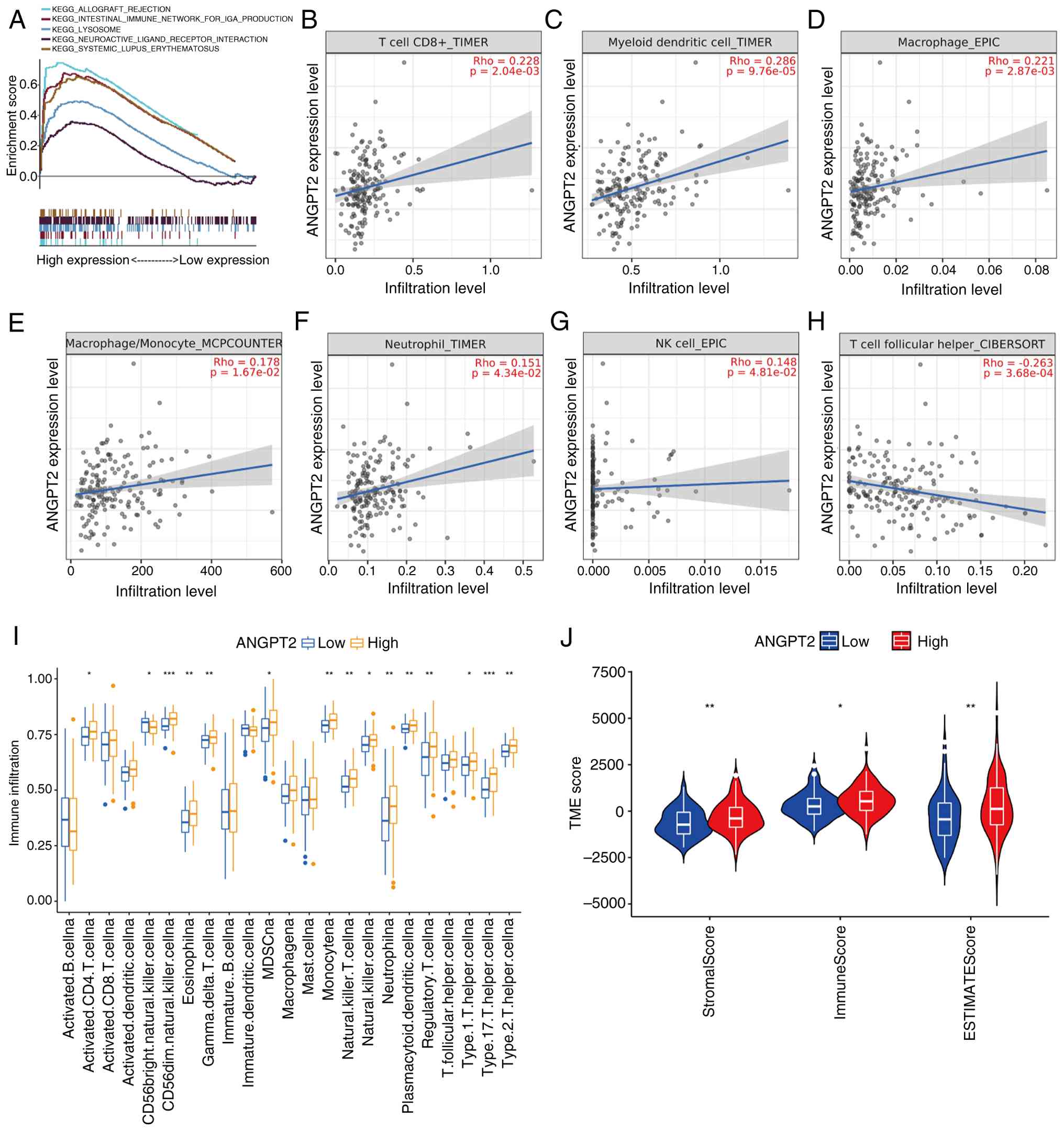 | Figure 5.Connection of ANGPT2 expression with
immunity in ESCA. (A) GSEA enrichment plots showing signaling
pathways significantly enriched in the ANGPT2 high-expression group
(1,000 permutations were performed; pathways include
‘ALLOGRAFT_REJECTION’ and ‘INTERFERON_GAMMA_RESPONSE’, consistent
with immune-related functions). (B-H) Spearman's correlation
analysis between ANGPT2 expression and infiltration levels of
specific immune cell subsets in ESCA (adjusted for tumor purity via
TIMER2.0 or indicated algorithms): (B) CD8+ T cells
(TIMER, Rho=0.220, P=2.04×10−3); (C) myeloid dendritic
cells (TIMER, Rho=0.286, P=9.76×10−5); (D) macrophages
(EPIC, Rho=0.211, P=2.87×10−3); (E) macrophage/monocytes
(MCPCOUNTER, Rho=0.178, P=1.67×10−2), (F) neutrophils
(TIMER, Rho=0.132, P=4.34×10−2), (G) NK cells (EPIC,
Rho=0.148, P=4.81×10−2), (H) T follicular helper cells
(CIBERSORT, Rho=0.260, P=3.68×10−4). (I) Single-sample
GSEA showing the abundance of 23 infiltrating immune cell subgroups
in ANGPT2 high- vs. low-expression groups. (J) ESTIMATE algorithm
analysis of Stromal Score, Immune Score and ESTIMATE Score in
ANGPT2 high- vs. low-expression groups (P<0.01 for Stromal
Score; P<0.05 for Immune Score; P<0.01 for ESTIMATE Score, as
indicated by symbols). *P<0.05, **P<0.01 and ***P<0.001;
all P-values are adjusted for multiple comparisons where
applicable. ANGPT2, angiopoietin-2; ESCA, esophageal cancer; GSEA,
Gene Set Enrichment Analysis; NK, natural killer. |
Connection between ANGPT2 expression
and immunity
To ascertain the differentially activated signaling
pathways in ESCA, a GSEA was conducted on the datasets of ANGPT2
with high and low expression. The signaling pathway with the most
significant enrichment was selected based on its NES. The ANGPT2
high-expression phenotype was differentially enriched in allogeneic
rejection, IGA-producing intestinal immune network, neuroactive
ligand receptor interaction, lysosome and systemic lupus
erythematosus (SLE) (Fig. 5A).
These enrichment results suggested that ANGPT2 was related to
immunity. The ICs infiltrating tumors can profoundly affect tumor
progression. The correlation between ANGPT2 expression in ESCA and
the analyzed ICs, such as CD8+ T cells, dendritic cells
(DCs), monocytes, macrophages (MPh), neutrophils, natural killer
cells (NKs) and T cell follicular helper cells, was significant
(Fig. 5B-H). To quantify
differences in immune infiltration across the broader immune
landscape, ssGSEA was used to compare 23 distinct IC subsets
between high- and low-ANGPT2 groups. Results demonstrated
significant disparities in immune infiltration profiles, with
high-ANGPT2 tumors exhibiting elevated infiltration across multiple
IC subtypes (Fig. 5I). Further
characterization of the TME using the ESTIMATE algorithm manifested
that ANGPT2 overexpression correlated with significantly higher
stromal, immune and ESTIMATE scores (Fig. 5J), indicating a more immune- and
stroma-rich TME, consistent with enhanced immune recruitment or
remodeling. To probe the mechanistic link between ANGPT2 and immune
activation, correlations between ANGPT2 levels and established IC
marker genes in ESCA were analyzed using TCGA data (Table IV). ANGPT2 showed strong positive
correlations with markers of T cell (CD3G and CD4), T follicular
helper cell (ICOS), M1-MPh (NOS2, IRF5 and PTGS2), M2-MPh (VSIG4,
CD163 and MS4A4A), neutrophil cell (FCGR3B), DC (HLA-DPB1, HLA-DRA,
NRP1, HLA-DPA1 and ITGAX), NKs (NKG7, FCGR3A) and monocyte cell
(S100A8, S100A9, CD68 and LYZ). ANGPT2 is highly correlated with
immune invasion, but its expression in ICs is very low. The results
of single-cell sequencing of ESCA are depicted in Fig. S2. The aforementioned results
partially support that ANGPT2 is positively correlated with
immunity.
 | Table IV.Correlation analysis between
angiopoietin-2 and immune cell biomarkers in esophageal cancer
determined by The Cancer Genome Atlas. |
Table IV.
Correlation analysis between
angiopoietin-2 and immune cell biomarkers in esophageal cancer
determined by The Cancer Genome Atlas.
| Immune cell | Biomarker | Cor | P-value |
|---|
| B cell | CD19 | 0.01 |
8.8×10−1 |
|
| CD79A | −0.7 |
3.2×10−1 |
| T cell | CD3D | 0.13 |
8.3×10−2 |
|
| CD3G | 0.18 |
1.9×10−2 |
|
| CD4 | 0.28 |
3.0×10−4 |
| Tfh cell | ICOS | 0.21 |
5.0×10−3 |
| M1 macrophage | NOS2 | 0.18 |
1.7×10−2 |
|
| IRF5 | −0.24 |
1.9×10−3 |
|
| PTGS2 | 0.20 |
7.4×10−3 |
| M2 macrophage | CD163 | 0.30 |
1.0×10−4 |
|
| VSIG4 | 0.21 |
6.5×10−3 |
|
| MS4A4A | 0.29 |
1.1×10−4 |
| Neutrophil | FCGR3B | 0.23 |
2.4×10−3 |
|
| CXCR2 | −0.09 |
2.5×10−1 |
| Dendritic cell | HLA-DPB1 | 0.22 |
4.8×10−3 |
|
| HLA-DQB1 | 0.15 |
4.5×10−2 |
|
| HLA-DRA | 0.25 |
1.2×10−3 |
|
| HLA-DPA1 | 0.26 |
6.4×10−4 |
|
| CD1C | −0.05 |
5.2×10−1 |
|
| NRP1 | 0.28 |
2.1×10−4 |
|
| ITGAX | 0.26 |
6.2×10−4 |
| Natural killer
cell | NKG7 | 0.04 |
5.3×10−1 |
|
| FCGR3A | 0.23 |
2.3×10−3 |
| Monocyte cell | S100A8 | −0.28 |
2.1×10−4 |
|
| S100A9 | −0.27 |
3.4×10−4 |
|
| CD68 | 0.16 |
3.6×10−2 |
|
| LYZ | 0.35 |
5.1×10−6 |
Relationship between ANGPT2 and immune
checkpoints and chemotherapeutic sensitivity in ESCA
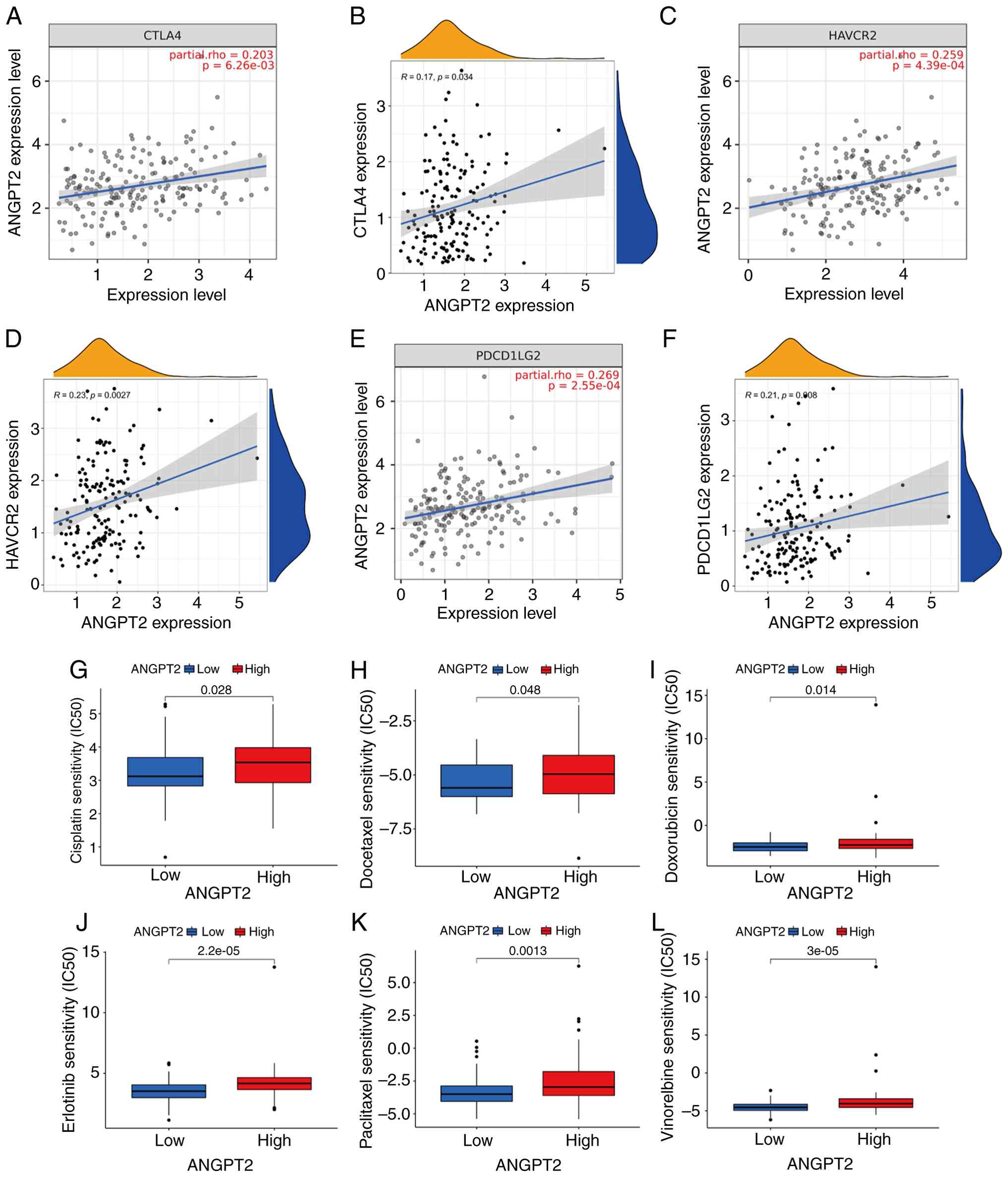
Tumor cells can modify the TME and facilitate immune
system evasion by the tumor (21).
The relation between 70 immune checkpoints and ANGPT2 was analyzed
via the TCGA database (Table SI;
Fig. 6B, D and F). Immune
checkpoints, including CTLA-4 (22), HAVCR2 (23) and PDCD1LG2 (24), play a key role in cancer
progression. Since ANGPT2 may play a carcinogenic role in ESCA, the
relationship between ANGPT2 and CTLA-4, as well as HAVCR2 and
PDCD1LG2, was also evaluated using TIMER2.0. ANGPT2 expression in
ESCA was positively related to CTLA-4, HAVCR2 and PDCD1LG2
(Fig. 6A, C and D). The findings
suggest that the evasion of the immune system by tumors may be
involved in ESCA development through the action of ANGPT2. To
explore its potential clinical relevance in therapeutic response,
the sensitivity of ESCA tumors stratified by ANGPT2 expression
levels to commonly used chemotherapeutic agents, was evaluated.
Notably, the low ANGPT2 expression group exhibited lower
IC50 values for chemotherapeutics, including cisplatin,
docetaxel, doxorubicin, erlotinib, paclitaxel and vinorelbine. To
summarize, the findings suggest a correlation between drug
sensitivity and ANGPT2 (Fig.
6G-L).
Discussion
To date, despite the comprehensive treatment
available for ESCA, the prognosis is still unsatisfactory.
Exploring the molecular mechanism of ESCA carcinogenesis and
identifying promising prognostic biomarkers are extremely critical.
ANGPT2 has been reported to be crucial not only in CVD but also in
initiation and progression of human tumors. However, its role in
ESCA remains underexplored, necessitating further mechanistic and
clinical investigation.
The current study conducted a pan-cancer analysis of
ANGPT2 expression through the TCGA database, with validation
performed via the GEPIA. Survival analysis showcased that patients
with ESCA with overexpressed ANGPT2 exhibited significantly worse
OS. Multivariate clinical data analysis further indicated that
ANGPT2 overexpression may serve as an independent prognostic factor
in ESCA. In vitro functional assays demonstrated that
knocking down ANGPT2 significantly suppressed ESCA migration and
invasion. Collectively, ANGPT2 plays an oncogenic role in ESCA
progression. ANGPT2 exhibits significant stage-dependent
overexpression in ESCA. In TCGA and GEO cohorts, its mRNA levels
escalate with TNM stages (I–IV), showing marked differences between
adjacent stages (all P<0.05). Local IHC data confirm higher
protein scores in advanced (III–IV, median=7.8) vs. early stages
(I–II, median=3.2, P<0.001), correlating with T/N/M
classifications. ANGPT2 distinguishes early from advanced ESCA with
area under the curves (AUCs) of 0.78 (mRNA) and 0.86 (protein),
improving to 0.92 when combined with CEA/SCC. It also aids early
diagnosis, with AUCs of 0.73 (mRNA) and 0.80 (protein) vs. normal
tissues. Mechanistically, stage-dependent upregulation via the
MAPKAPK5-AS1/SNHG1/hsa-miR-145-5p axis drives angiogenesis, TME
remodeling and immune checkpoint (CTLA-4, HAVCR2 and PDCD1LG2)
activation. Clinically, it refines TNM staging (C-index=0.78) and
enables non-invasive monitoring via serum levels (AUC=0.83). These
data support ANGPT2 as a valuable diagnostic biomarker for ESCA
stage stratification and early detection.
miRNAs and lncRNAs are key regulators of gene
expression. To identify potential upstream miRNAs targeting ANGPT2,
StarBase was used to predict miRNA-ANGPT2 interactions. Finally, 10
miRNAs were obtained, and some of the predicted miRNAs had been
verified to have a tumor suppressor role in human tumors. For
instance, hsa-miR-148a-3p is associated with drug resistance and
aggressiveness in ESCA (20), and
miR-152 functions as a tumor suppressor in human BRCA by targeting
PIK3CA (21). After correlation and
expression analyses, hsa-miR-145-5p was chosen as the most
promising upstream tumor suppressor miRNA of ANGPT2. High
hsa-miR-145-5p expression in ESCA is related to prognosis (22). In addition, lncRNAs potentially
interacting with hsa-miR-145-5p were predicted, identifying 38
candidate lncRNAs. Through correlation and expression analyses, as
well as survival prediction analysis, two lncRNAs, MAPKAPK5-AS1 and
SNHG1, were prioritized. Both have been implicated in the
pathogenesis of multiple malignancies. For instance, MAPKAPK5-AS1
drives LIHC progression via a MAPKAPK5-AS1/PLAGL2/HIF-1α signaling
axis (23), while SNHG1 knockdown
in ESCA suppresses migration and invasion and promotes apoptosis
through miR-204 upregulation and HOXC8 downregulation (24). In summary, the MAPKAPK5-AS1 and
SNHG1/hsa-miR-145-5p/ANGPT2 axes may represent novel regulatory
signaling contributing to ESCA pathogenesis. The dual-luciferase
reporter assay results have confirmed the direct binding between
the predicted lncRNAs (MAPKAPK5-AS1 and SNHG1) and miR-145-5p, as
well as between miR-145-5p and the 3′UTR of ANGPT2, providing
experimental evidence for the post-transcriptional regulatory
mechanism of this axis. However, RNA immunoprecipitation assays to
verify the endogenous physical interaction between these lncRNAs
and miR-145-5p in ESCA cells have not been performed in the current
study; this experiment is planned for follow-up research to fully
characterize the competing endogenous RNA regulatory network in
ESCA and confirm the in vivo functional relevance of this
axis.
Emerging evidence indicates that overexpressed
ANGPT2 is associated with increased all-cause and cardiovascular
mortality in the general population, and increased death in
patients with cardiogenic shock (25,26).
ANGPT2 levels are highly expressed in human malignancies and CVD,
including heart failure, ischemic myocardial injury, and other
complications secondary to chronic kidney impairment, diabetes and
hypertension. Despite distinct clinical phenotypes, cancer and CVD
share key ANGPT2-driven pathogenic mechanisms, particularly poor
vascular network remodeling, inflammation and epithelial (or
endothelial) to mesenchymal transition. ANGPT2-dependent vascular
instability is a well-known underlying mechanism behind malignancy
and CVD progression (27). In
addition, ANGPT2 has been shown to exert immunomodulatory effects
by inducing PD-L1 overexpression in tumor-associated MPh, thereby
impairing T-cell-mediated immune surveillance and antitumor
cytotoxicity. These findings position ANGPT2 not only as a
functional contributor to immune evasion but also as a probable
biomarker and therapeutic target within the immune checkpoint
landscape (28). Studies have shown
that endothelial cell- and MPh-derived Angpt2 aggravates cardiac
hypoxia and inflammation by promoting abnormal vascular remodeling,
enhancing neutrophil infiltration, and pro-inflammatory MPh
polarization after myocardial ischemia and myocardial infarction
(29).
In the present study, the phenotype of ANGPT2
overexpression is correlated with allogeneic rejection, neuroactive
ligand receptor interaction, the IGA-producing intestinal immune
network, lysosome function and SLE, as determined by GSEA, all of
which underscore its immunological relevance. Given the
accumulating evidence that immune dysregulation contributes to
tumorigenesis, the findings of the present study further
demonstrate that ANGPT2 has a high correlation with ICs, including
CD8+ T cells, MPh, DCs, monocytes, neutrophils, NK cells
and T cell follicular helper cells. Moreover, the TME attributes
and the relative abundance of 23 TIICs differed significantly
between ANGPT2 expression groups. Additionally, ANGPT2
overexpression in ESCA is closely related to the immune checkpoints
CLAT-4, HAVCR2 and PDCD1LG2, indicating the implication of tumor
immunity in the carcinogenesis of ESCA mediated by ANGPT2.
Moreover, ANGPT2 was related to drug sensitivity.
Notably, the current findings on the association
between ANGPT2 and tumor immunity provide a strong rationale for
exploring its potential value in ESCA immunotherapy, especially in
combination with PD-1/PD-L1 inhibitors. Immune checkpoint
inhibitors (ICIs) targeting the PD-1/PD-L1 axis have transformed
the treatment of various cancers, but their efficacy in ESCA
remains limited, with only ~20–30% of patients achieving durable
responses (30). This limited
efficacy is largely attributed to the immunosuppressive TME, which
restricts T cell infiltration and functional activation. The
present data reveal that ANGPT2 overexpression is positively
correlated with PDCD1LG2 (PD-L2) expression, a key ligand of PD-1,
and the enrichment of immunosuppressive cells such as M2
macrophages and myeloid-derived suppressor cells MDSCs in ESCA.
This aligns with previous findings demonstrating that ANGPT2 can
induce PD-L1 upregulation on tumor-associated macrophages and tumor
cells, thereby fostering immune evasion (19). Collectively, these observations
suggest that ANGPT2 may be a critical driver of ICI resistance in
ESCA, and targeting ANGPT2 could synergize with PD-1/PD-L1 blockade
to enhance therapeutic efficacy.
Preclinical studies in other cancer types have
already validated the synergistic effects of ANGPT2 inhibition and
PD-1/PD-L1 inhibitors. For example, in non-small cell lung cancer,
neutralizing ANGPT2 normalized tumor vasculature, increased
intratumoral CD8+ T cell infiltration, and enhanced the
antitumor activity of anti-PD-1 therapy (18). In hepatocellular carcinoma, ANGPT2
knockdown reversed the immunosuppressive TME by reducing regulatory
T cell accumulation and restoring effector T cell function, thereby
sensitizing tumors to PD-L1 inhibitors (31). Extrapolating these findings to ESCA,
it was hypothesized that ANGPT2 inhibition could remodel the
immunosuppressive TME characterized by high PDCD1LG2 expression and
abnormal immune cell infiltration. Specifically, targeting ANGPT2
may downregulate PD-L1/PD-L2 expression on tumor cells and
macrophages, reduce the recruitment of immunosuppressive cells, and
enhance the cytotoxic activity of CD8+ T cells,
ultimately overcoming ICI resistance and improving treatment
outcomes.
The clinical implications of this combination
strategy are substantial. Patients with ESCA with high ANGPT2
expression, who exhibit poor prognosis and compromised tumor
immunity, may be ideal candidates for dual ANGPT2/PD-1/PD-L1
targeting. ANGPT2 could serve as a predictive biomarker to identify
patients most likely to benefit from ICI-based combination therapy,
addressing the unmet need for personalized immunotherapy in ESCA.
Furthermore, the observation of the present study that ANGPT2
expression correlates with chemotherapeutic sensitivity suggests
that triple therapy (ANGPT2 inhibition + PD-1/PD-L1 blockade +
chemotherapy) may yield even greater clinical benefits.
Chemotherapy can induce immunogenic cell death, release
tumor-associated antigens, and synergize with immunotherapy to
amplify antitumor immune responses (32), while ANGPT2 inhibition may further
enhance this synergy by remodeling the TME.
However, several considerations must be addressed
before clinical translation. First, the specific molecular
mechanisms by which ANGPT2 regulates PD-L1/PD-L2 expression in ESCA
require further validation, for example, whether ANGPT2 directly
modulates the transcriptional activity of PDCD1LG2 or acts through
downstream signaling pathways such as PI3K/Akt or NF-κB. Second,
preclinical studies using ESCA cell line-derived xenografts and
patient-derived xenografts (PDXs) are needed to confirm the
synergistic efficacy of ANGPT2 inhibitors (for example,
neutralizing antibodies and siRNA) and PD-1/PD-L1 blockers. Third,
the optimal dosage, administration schedule and safety profile of
ANGPT2-targeting agents in combination with ICIs must be evaluated
in preclinical models to minimize toxicity while maximizing
efficacy. Finally, clinical trials are warranted to assess the
safety and efficacy of this combination strategy, with correlative
studies to monitor TME remodeling (for example, CD8+ T
cell infiltration and PD-L1 expression) and validate ANGPT2 as a
predictive biomarker.
In addition to immunotherapy, the crosstalk between
ANGPT2 and other oncogenic pathways in ESCA merits further
exploration. For instance, ANGPT2 has been reported to interact
with VEGFA to promote angiogenesis and immune suppression (19), and dual targeting of ANGPT2 and
VEGFA has shown promising results in preclinical models (33). Whether this dual anti-angiogenic
strategy can synergize with immunotherapy in ESCA deserves
investigation. Furthermore, the current identification of the
MAPKAPK5-AS1/SNHG1/hsa-miR-145-5p/ANGPT2 regulatory axis provides
additional therapeutic targets. Targeting MAPKAPK5-AS1 or SNHG1
could downregulate ANGPT2 expression, offering an alternative
approach to modulate ANGPT2 activity in ESCA. However, the
potential off-target effects of lncRNA-targeting therapies need to
be carefully evaluated, and more specific targeting strategies (for
example, antisense oligonucleotides and small molecule inhibitors)
should be developed.
It is also important to acknowledge the limitations
of the present study. First, the in vitro functional assays
were performed in only two ESCA cell lines (TE-1 and KYSE150), and
validation in additional cell lines and in vivo models (for
example, PDXs) is necessary to confirm the oncogenic role of ANGPT2
in ESCA. Second, the correlation between ANGPT2 expression and
immune infiltration was based on bioinformatics analysis of public
databases, and experimental validation using clinical samples (for
example, flow cytometry and multiplex immunofluorescence) is
required to confirm these findings. Third, the mechanism by which
ANGPT2 regulates the TME and immune checkpoints in ESCA remains
partially elusive, and further studies are needed to dissect the
detailed signaling pathways. Finally, the drug sensitivity analysis
was based on the pRRophetic algorithm, and experimental validation
using ESCA cell lines and patient-derived organoids is needed to
confirm the correlation between ANGPT2 expression and
chemotherapeutic sensitivity.
In summary, ANGPT2 is overexpressed in multiple
tumors, including ESCA, and functions as an independent prognostic
biomarker. It was found that MAPKAPK5-AS1 and SNHG1/hsa-miR-145-5p
can regulate ANGPT2. In addition, tumor immunity may be involved in
ESCA carcinogenesis, which is modulated by ANGPT2. Notably, ANGPT2
holds significant potential as a therapeutic target for enhancing
the efficacy of PD-1/PD-L1 inhibitors in ESCA, providing a
rationale for combination immunotherapy strategies. While these
results provide a compelling mechanistic and clinical framework,
they require further validation through functional in vitro
and in vivo studies, as well as prospective, large-scale,
multicenter clinical trials, to confirm their translational
relevance and therapeutic potential.
Finally, the present study has certain limitations
that should be acknowledged. First, consistent with the focused
research design centered on ANGPT2, the expression levels of the
upstream regulatory molecules (MAPKAPK5-AS1, SNHG1 and miR-145-5p)
were not validated in the clinical samples of patients with ESCA.
Although the regulatory relationships among these molecules were
confirmed by in silico analysis and in vitro
functional assays, the clinical relevance and expression patterns
of this lncRNA/miRNA axis in patient tissues remain to be
elucidated. Future studies with expanded sample sizes are warranted
to validate the expression of this regulatory axis and explore its
clinical prognostic value in ESCA.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the National Natural Science
Foundation of China (grant no. 81572956), the Jiangsu Provincial
Science and Technology Supporting Program (grant no. BE2017096),
the Medical Innovation Team of Jiangsu (grant no. CXTDC2016009),
the Guiding Science and Technology Plan Project for Social
Development in Zhenjiang (grant nos. FZ2023049 and FZ2024059) and
the Student Innovation Training Program Projects of Jiangsu
University (grant no. 201810299262W).
Availability of data and materials
The data generated in the present study may be found
in the Gene Expression Omnibus under accession numbers GSE20347,
GSE38129, GSE45670, GSE70409 and GSE160269 or at the following URL:
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE20347;
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE38129;
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE45670;
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE70409;
https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE160269.
The data generated in the present study may be requested from the
corresponding author.
Authors' contributions
XT and RL conceived and designed the study. ZZ, ZX,
YuF, YY and YiF performed the experimental procedures and acquired
the research data. ZZ, YuF, YiF, SL, CC, KZ, DC and YY analyzed and
interpreted the experimental data. ZZ, XT and RL drafted the
original manuscript. DC, KZ, CC and SL critically revised the
manuscript for important intellectual content. All authors read and
approved the final version of the manuscript and agreed to be
accountable for all aspects of the work in ensuring that questions
related to the accuracy or integrity of any part of the work are
appropriately investigated and resolved. ZZ and XT confirm the
authenticity of all the raw data.
Ethics approval and consent to
participate
The present study was approved (approval no.
KY201901) by the Ethics Committee of The Affiliated Hospital of
Jiangsu University (Zhenjiang, China). Written informed consent was
provided by all participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
BLCA
|
bladder urothelial carcinoma
|
|
BRCA
|
breast invasive carcinoma
|
|
CHOL
|
cholangiocarcinoma
|
|
COAD
|
colon adenocarcinoma
|
|
GBM
|
glioblastoma multiforme
|
|
HNSC
|
head and neck squamous cell
carcinoma
|
|
KICH
|
kidney chromophobe
|
|
KIRC
|
kidney renal clear cell carcinoma
|
|
KIRP
|
kidney renal papillary cell
carcinoma
|
|
LIHC
|
liver hepatocellular carcinoma
|
|
LUAD
|
lung adenocarcinoma
|
|
LUSC
|
lung squamous cell carcinoma
|
|
PRAD
|
pancreatic adenocarcinoma
|
|
READ
|
rectum adenocarcinoma
|
|
STAD
|
stomach adenocarcinoma
|
|
THCA
|
thyroid carcinoma
|
|
UCEC
|
uterine corpus endometrial
carcinoma
|
References
|
1
|
Wang M, Sun X, Xin H, Wen Z and Cheng Y:
SPP1 promotes radiation resistance through JAK2/STAT3 pathway in
esophageal carcinoma. Cancer Med. 11:4526–4543. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
He Y, Li D, Shan B, Liang D, Shi J, Chen W
and He J: Incidence and mortality of esophagus cancer in China,
2008–2012. Chin J Cancer Res. 31:426–434. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Wu S, Zhang L, Deng J, Guo B, Li F, Wang
Y, Wu R, Zhang S, Lu J and Zhou Y: A novel micropeptide encoded by
Y-Linked LINC00278 links cigarette smoking and AR signaling in male
esophageal squamous cell carcinoma. Cancer Res. 80:2790–2803. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhou Z, Tang J, Lu Y, Jia J, Luo T, Su K,
Dai X, Zhang H and Liu O: Prognosis-related molecular subtyping in
head and neck squamous cell carcinoma patients based on
glycolytic/cholesterogenic gene data. Cancer Cell Int. 23:372023.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Hao Y, Wang M, Jiang X, Zheng Y, Ran Q, Xu
X, Zou B, Wang J, Liu N and Qin B: Non-acid reflux and esophageal
dysmotility is associated with early esophageal squamous cell
carcinoma. J Cancer Res Clin Oncol. 149:8327–8334. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Shi X, Zhao H, Yu J, Cai P, Zhou S, Yang N
and Li D: Changes in PD-1 expression on T lymphocyte subsets and
related immune indicators before and after definitive
chemoradiotherapy for esophageal squamous cell carcinoma. Ann Med.
57:24451902025. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Nan Y, Liu S, Luo Q, Wu X, Zhao P, Chang
W, Zhang R, Li Y and Liu Z: m(6)A demethylase FTO stabilizes LINK-A
to exert oncogenic roles via MCM3-mediated cell-cycle progression
and HIF-1α activation. Cell Rep. 42:1132732023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Valenzuela DM, Griffiths JA, Rojas J,
Aldrich TH, Jones PF, Zhou H, McClain J, Copeland NG, Gilbert DJ,
Jenkins NA, et al: Angiopoietins 3 and 4: diverging gene
counterparts in mice and humans. Proc Natl Acad Sci USA.
96:1904–1909. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Maisonpierre PC, Suri C, Jones PF,
Bartunkova S, Wiegand SJ, Radziejewski C, Compton D, McClain J,
Aldrich TH, Papadopoulos N, et al: Angiopoietin-2, a natural
antagonist for Tie2 that disrupts in vivo angiogenesis. Science.
277:55–60. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Davis S, Aldrich TH, Jones PF, Acheson A,
Compton DL, Jain V, Ryan TE, Bruno J, Radziejewski C, Maisonpierre
PC and Yancopoulos GD: Isolation of angiopoietin-1, a ligand for
the TIE2 receptor, by secretion-trap expression cloning. Cell.
87:1161–1169. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hultström M, Fromell K, Larsson A, Persson
B, Nilsson B, Quaggin SE, Betsholtz C, Frithiof R, Lipcsey M and
Jeansson M: Angiopoietin-2 inhibition of thrombomodulin-mediated
anticoagulation-A novel mechanism that may contribute to
hypercoagulation in critically Ill COVID-19 patients. Biomedicines.
10:13332022. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Marola OJ, Onos KD, Diemler C, Keezer KJ,
Sasner M and Howell GR: Identifying genetic risk factors driving
cerebral amyloid angiopathy using novel mouse models. Basic Science
and Pathogenesis. Alzheimer's Dement. 20 (Suppl
1):e0914462024.https://doi.org/10.1002/alz.091446 View Article : Google Scholar
|
|
13
|
Urosevic J, Blasco MT, Llorente A,
Bellmunt A, Berenguer-Llergo A, Guiu M, Cañellas A, Fernandez E,
Burkov I, Clapés M, et al: ERK1/2 signaling induces upregulation of
ANGPT2 and CXCR4 to mediate liver metastasis in colon cancer.
Cancer Res. 80:4668–4680. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Kim B, Park YY and Lee JH: CXCL10 promotes
melanoma angiogenesis and tumor growth. Anim Cells Syst (Seoul).
28:453–465. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yang YC, Ho KH, Pan KF, Hua KT, Tung MC,
Ku CC, Chen JQ, Hsiao M, Chen CL, Lee WJ and Chien MH: ESM1
facilitates the EGFR/HER3-triggered epithelial-to-mesenchymal
transition and progression of gastric cancer via modulating
interplay between Akt and angiopoietin-2 signaling. Int J Biol Sci.
20:4819–4837. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Gu J, Zhang Y, Han Z, Gao L, Cui J, Sun Y,
Niu Y, You B, Huang CP, Chang C, et al: Targeting the
ERβ/Angiopoietin-2/Tie-2 signaling-mediated angiogenesis with the
FDA-approved anti-estrogen Faslodex to increase the Sunitinib
sensitivity in RCC. Cell Death Dis. 11:3672020. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhu J, Wu Y, Yu Y, Li Y, Shen J and Zhang
R: MYBL1 induces transcriptional activation of ANGPT2 to promote
tumor angiogenesis and confer sorafenib resistance in human
hepatocellular carcinoma. Cell Death Dis. 13:7272022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lauret Marie Joseph E, Laheurte C, Jary M,
Boullerot L, Asgarov K, Gravelin E, Bouard A, Rangan L, Dosset M,
Borg C and Adotévi O: Immunoregulation and clinical implications of
ANGPT2/TIE2(+) M-MDSC signature in non-small cell lung cancer.
Cancer Immunol Res. 8:268–279. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
He J, Jiang M, Liu J, Zhu R, Lv W, Lian R,
Yang Y and Wang R: Homeobox D9 drives the malignant phenotypes and
enhances the Programmed death ligand-1 expression in non-small cell
lung cancer cells via binding to Angiopoietin-2 promoter. World J
Surg Oncol. 21:932023. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cao D, Ge S and Li M: MiR-451a promotes
cell growth, migration and EMT in osteosarcoma by regulating
YTHDC1-mediated m6A methylation to activate the AKT/mTOR signaling
pathway. J Bone Oncol. 33:1004122022. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang Y, Zhang J, Zhao W, Wang D, Ma W,
Shang S, Feng C and Yu H: MicroRNA expression in esophageal
squamous cell carcinoma: Novel diagnostic and prognostic
biomarkers. Mol Med Rep. 15:3833–3839. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shi P, Liu Y, Yang D, Wu Y, Zhang L, Jing
S, Shi H and Geng C: CircRNA ZNF609 promotes the growth and
metastasis of thyroid cancer in vivo and in vitro by downregulating
miR-514a-5p. Bioengineered. 13:4372–4384. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Li HM, Yu YK, Liu Q, Wei XF, Zhang J,
Zhang RX, Sun HB, Wang ZF, Xing WQ and Li Y: LncRNA SNHG1 regulates
the progression of esophageal squamous cell cancer by the
miR-204/HOXC8 Axis. Onco Targets Ther. 13:757–767. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Slywitch E, Savalli C, Duarte AC and
Escrivão MAMS: Obese vegetarians and omnivores show different
metabolic changes: Analysis of 1340 individuals. Nutrients.
14:22042022. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Varricchi G, Loffredo S, Bencivenga L,
Ferrara AL, Gambino G, Ferrara N, de Paulis A, Marone G and Rengo
G: Angiopoietins, vascular endothelial growth factors and secretory
phospholipase A(2) in ischemic and non-ischemic heart failure. J
Clin Med. 9:19282020. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Huang X, Zheng S, Li S, Huang Y, Zhang W,
Liu F and Cao Q: Machine learning-based pathomics model predicts
ANGPT2 expression and prognosis in hepatocellular carcinoma. Am J
Pathol. 195:561–574. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Wu X, Giobbie-Hurder A, Liao X, Connelly
C, Connolly EM, Li J, Manos MP, Lawrence D, McDermott D, Severgnini
M, et al: Angiopoietin-2 as a biomarker and target for immune
checkpoint therapy. Cancer Immunol Res. 5:17–28. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lee SJ, Lee CK, Kang S, Park I, Kim YH,
Kim SK, Hong SP, Bae H, He Y, Kubota Y and Koh GY: Angiopoietin-2
exacerbates cardiac hypoxia and inflammation after myocardial
infarction. J Clin Invest. 128:5018–5033. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Luo T, Yang H, Ran X, Xu B, Zhang Y, Zhang
L, Zhang C and Fu M: Analysis of the safety and efficacy of
PD-1/PD-L1 inhibitors combined with chemotherapy in the treatment
of locally advanced resectable esophageal squamous cell carcinoma:
A systematic review and meta-analysis based on four randomized
controlled trials. Front Oncol. 15:15901112025. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Kawagishi N, Suda G, Yamamoto Y, Baba M,
Furuya K, Maehara O, Ohnishi S, Yoshida S, Fu Q, Yang Z, et al:
Serum angiopoietin-2 predicts the occurrence and recurrence of
hepatocellular carcinoma after direct-acting antiviral therapy for
hepatitis C. Viruses. 15:1812023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sagie S, Babu T, Weller C, Tabachnik C,
Livneh I, Quast NP, Gumpert N, Shomuradova A, Raybould MI, Levy R,
et al: Lymphodepleting chemotherapy potentiates neoantigen-directed
T cell therapy by enhancing antigen presentation. Cell Rep Med.
6:1025062025. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Popov V, Raycheva G and Grudeva-Popova Z:
Dynamics of VEGF-А, Аngiopoietin-2 and HIF-1α levels in patients
with brain metastases treated with cyberknife radiosurgery. Technol
Cancer Res Treat. 24:153303382513139452025. View Article : Google Scholar : PubMed/NCBI
|