Introduction
Osteosarcoma is the most common type of primary
tumor of the bone in childhood and adolescence (1). Adoption of neoadjuvant treatment
regimens that combine surgery and chemotherapy have improved
prognosis; however, survival rates for patients with metastatic
osteosarcoma remain at ~30% (2,3).
Therefore, identification of novel therapeutic targets and
development of alternative options are required.
Our previous study described development of a mouse
model of osteosarcoma by overexpressing c-MYC in bone marrow
stromal cells derived from Ink4a/Arf-null mice (4). When inoculated into C57BL/6 syngeneic
mice, these highly tumorigenic cells, designated accelerated bone
and tumor formation (AXT) cells, rapidly formed osteoid-rich
primary tumors and metastatic lesions that pathologically mimic
human osteosarcoma (5,6).
Anoikis is a type of cell death induced by
detachment from the extracellular matrix (ECM) (7,8).
Highly malignant tumor cells can escape anoikis, enabling them to
grow without the ECM and develop into metastatic lesions.
Consistent with their high metastatic capacity in vivo, AXT
cells also proliferate in non-adherent cultures, and under these
conditions, the expression levels of downstream target molecules of
hypoxia inducible factor-1 (HIF-1) are considerably
upregulated.
The HIF-1 complex, which comprises HIF-1α and HIF-1β
subunits, is a key regulator that enables malignant cells to
survive and continue to grow under hypoxic microenvironmental
conditions (9–11). HIF-1α transcriptionally activates
multiple target genes and interacts with several molecules to
promote progression, metastasis and therapeutic resistance of
various types of malignancy, including osteosarcoma. Analysis of
human osteosarcoma specimens suggests that the rate of positive
staining for HIF-1α in human osteosarcoma specimens is 34–88%
(12), and high expression of
HIF-1α is associated with clinical stage (12,13),
metastasis (12), overall survival
(12,14) and resistance to treatment (14,15),
making it a potential prognostic predictor of poor prognosis.
Several in vivo experimental studies demonstrate that HIF-1α
promotes osteosarcoma progression and metastasis directly (16–19).
Thus, HIF-1α and its associated pathways may be promising
therapeutic targets for osteosarcoma. However, the stage of
osteosarcoma development at which HIF-1α is expressed, and the role
it plays, is unclear. In particular, experimental verification of
the validity and problems of HIF-targeting therapy for osteosarcoma
remains unsolved. Using a syngeneic mouse model of osteosarcoma to
conduct depletion of HIF-1α and subsequent in vitro and
in vivo analyses may provide clues to resolving these
issues.
In the present study to clarify the role of HIF-1α
in the progression of osteosarcoma, HIF-1α was knocked out in AXT
cells using CRISPR-Cas9 with two different target sequences to
produce three HIF-1α-knockout (KO) clones. Using HIF-KO cells, cell
growth and cell cycle status was evaluated in vitro and the
effects of HIF-1α depletion in vivo on primary tumor
formation, metastasis, tumor angiogenesis and gene expression
changes was assessed.
Materials and methods
Reagents
Roxadustat (25 µM; 27 h), buparlisib (1 µM; 2 days)
(MedChemExpress), antimycin A (1 µM; 2 days), oligomycin (1 µM; 2
days) (Abcam), rotenone (1 µM; 2 days), mevalonate (200 µM; 17 h or
2 days) (MilliporeSigma), BAY-876 (3.3–30.0 µM), everolimus (1 or 5
µM, 19.5 h or 2 days), temsirolimus (1 or 5 µM; 19.5 h or 2 days),
shikonin (1 µM; 2 days), PT2385 (10 µM; 19.5 h), TC-S7009 (10 µM;
19.5 h) (Selleck Chemicals), simvastatin (0.5–3.0 µM; 4.5, 6, 14,
17, 21.5, 25 h or 2 days), atorvastatin (1–3 µM; 2 days) and
metformin (1–5 mM; 17 h or 2 days) (Tokyo Kasei Kogyo) were
reconstituted in dimethyl sulfoxide (DMSO; FUJIFILM Wako Pure
Chemical Corporation). 2-deoxy-D-glucose (3.3–30.0 mM; 2 days;
Tokyo Chemical Industry Co., Ltd.), Farnesyl Diphosphate (FPP; 50
µM; 2 days) and Geranylgeranyl Diphosphate (GGPP; 50 µM; 2 days;
Echelon) were dissolved in water. Reagents were applied to AXT,
control (Mock), or HIF-KO cells at the final concentrations for the
desired duration shown in parentheses at 5% CO2 at
37°C.
Establishment of HIF-1α-KO (HIF-KO)
cells using CRISPR-Cas9
Mouse osteosarcoma AXT cells were immortalized cells
previously established by overexpressing c-MYC in bone marrow
stromal cells derived from Ink4a/Arf-null mice (4). The gRNA sequences targeting the HIF-1α
exon 3 (bHLH domain) and exon 5 (Oxygen-Dependent Degradation
Domain; ODDD) were searched using CHOPCHOP (version 3; http://chopchop.cbu.uib.no/; SCR_015723) (20) and two sequences were used: #1,
5′-AGATGTGAGCTCACATTGTGGGG−3′ for KO1-1 and 1–2; and
#2, 5′-GCTAACAGATGACGGCGACATGG−3′ for KO2 (the protospacer
adjacent motif (PAM) is underlined). The oligos were annealed and
ligated into the lentiCRISPRv2-puro vector (Addgene_98290; Addgene,
Inc.), and digested with BsmBI according to the protocol
provided by the Zhang lab (https://www.addgene.org/crispr/zhang/). The Cas9 is
integrated in the lentiCRISPRv2-puro vector. After confirmation of
the inserted sequence, the lentiCRISPRv2-puro vectors were
co-transfected along with the psPAX2 (cat. no. 12260; Addgene,
Inc.) and pCMV–VSV-G (Addgene_8454; Addgene, Inc.) vectors into
Lenti-X 293T cells (CVCL_4401; Takara Bio) using FuGENE HD (Promega
Corporation) to generate infectious lentivirus. The empty
lentiCRISPRv2-puro vector was used to establish control (Mock)
cells. The virus-containing medium was added to AXT cells, and
infected cells were selected with puromycin. To obtain knockout
cells, puromycin-resistant cells were subjected to single-cell
cloning.
Cell culture
Human osteosarcoma U2OS (cat. no. HTB-96) and SaOS-2
(cat. no. HTB-85) cell lines were purchased from American Type
Culture Collection. These cells were authenticated by examination
of in vitro growth characteristics and morphological
properties provided evidence of correct cell identity. AXT cells,
HIF-1α-knockout AXT cells or human SaOS-2 and U2OS cells were
cultured under 5% CO2 at 37°C in Iscove's Modified
Dulbecco's Medium (IMDM; Nacalai Tesque, Inc.) or McCoy's 5A medium
(Thermo Fischer Scientific, Inc.), respectively, supplemented with
10% FBS (MilliporeSigma). For non-adherent cell culture, 6 cm
ultra-low-attachment surface dishes (Corning, Inc.) were used.
Cell proliferation assay
AXT cells, including HIF-1α-knockout AXT cells were
treated with trypsin at 37°C for 2 min, collected and washed with
serum-free medium, and then transferred to 96-well cell culture
plates (1×103 cells per well in 50 µl of IMDM
supplemented with 10% FBS). Ultra-low-attachment surface plates
(Corning Inc.) were used for non-adherent culture. Cells were
incubated for 1 h at 37°C before addition of 50 µl of the
corresponding medium supplemented with agents at twice the desired
final concentrations: Buparlisib (1 µM), antimycin A (1 µM),
oligomycin (1 µM), rotenone (1 µM), mevalonate (200 µM), BAY-876
(3.3–30.0 µM), everolimus (1 µM), temsirolimus (1 µM), shikonin (1
µM), simvastatin (0.5–3.0 µM), atorvastatin (1–3 µM), metformin
(1–5 mM), 2-deoxy-D-glucose (3.3–30.0 mM), FPP (50 µM) and GGPP (50
µM). Hypoxic conditions were created by placing oxygen absorbers in
an airtight bag, and adjusting ventilation to maintain a constant
2% oxygen concentration using an oxygen monitor. (Bionix culture
kit; Sugiyamagen Co., Ltd.). After incubation for 2 days, cell
viability was measured using a Cell Titer Glo assay kit (Promega
Corporation). Assays were performed in at least triplicate, and
data are expressed as the mean±SD relative (fold-change) to the
corresponding control value for cells incubated in the absence of
the indicated agents.
Reverse transcription-quantitative PCR
(RT-qPCR)
Extraction of total RNA from cultured cells, RT and
qPCR were performed using the NucleoSpin RNA kit, PrimeScript
reverse transcriptase (Takara Bio, Inc.) and Thunderbird SYBR qPCR
mix (Toyobo Co., Ltd.). The sequences of the PCR primers are listed
in Table SI. To prepare samples
from the mouse tissue, the left lung was suspended in lysis buffer
(NucleoSpin RNA kit) and disrupted using a BioMasher (Nippi, Inc.).
To evaluate circulating tumor cells (CTCs), total RNA was extracted
from 200 µl blood obtained from mice by cardiac blood sampling
after euthanasia using a NucleoSpin RNA blood kit (Takara Bio,
Inc.). Since GFP is expressed by AXT cells, tumor cells were
quantitated based on the level of Gfp mRNA expression
relative to Actb mRNA expression. Thermal cycler StepOne
(Thermo Fisher Scientific, Inc.) was used for qPCR analysis with
the 2step protocol; 60 sec at 95°C, then 40 cycles of 15 sec at
95°C and 60 sec at 60°C with the melting and dissociation curve
process. The fold change of gene expression was determined using
the relative quantification 2ΔΔCq method (21).
Western blot analysis
Cell lysates from Mock and HIF-KO cells were
prepared with 2× Laemmli sample buffer (Bio-Rad Laboratories, Inc.)
supplemented with 5% β-mercaptoethanol or radioimmunoprecipitation
assay (RIPA) buffer (Nacalai Tesque, Inc.) supplemented with
protease and phosphatase inhibitor cocktails (Nacalai Tesque,
Inc.). Protein concentrations were determined using a BCA assay kit
(Thermo Fisher Scientific, Inc.). Equal amounts of protein (30 µg
per lane) were used, and western blot analyses were conducted
according to standard semidry transfer procedures using 5–20%
gradient precast polyacrylamide gels (ATTO Corporation) and PVDF
membranes (ATTO Corporation). Membranes were blocked with 5% skim
milk (Nacalai Tesque, Inc.) for 1 h at room temperature and
subsequently incubated overnight at 4°C with the primary antibodies
listed in Table SII. After washing
4 times, the membranes were incubated with HRP-conjugated secondary
antibodies (1:3,000; listed in Table
SII) for 1 h at room temperature. Detection procedures were
performed using a Clarity Western ECL substrate (Bio-Rad
Laboratories, Inc.) and Amersham Hyperfilm ECL (Cytiva). Signal
intensities of bands were quantitated with ImageJ software (version
1.49; National Institutes of Health) (22).
GTP-RhoA assay
Mock and HIF-KO cells were treated with or without 1
µM simvastatin for 4.5 h at 37°C. Cells were lysed with a
magnesium-containing buffer (Rho activation assay kit; cat. no.
17-294 MilliporeSigma) and GTP-RhoA was isolated from each lysate
containing 1,183 µg of protein with beads conjugated with a GST
fusion protein containing the Rho binding domain of Rhotekin. A
part of the same lysate was used for evaluation of total RhoA.
Cell cycle analysis
Cells were trypsinized, washed with PBS and fixed
with 70% ethanol for ≥48 h at −20°C. Subsequently, the cells were
washed twice with ice-cold PBS, and stained with PBS containing 10
µg/ml propidium iodide and 20 µg/ml RNase A (MilliporeSigma). The
DNA content of at least 10,000 singlet cells was analyzed by flow
cytometry (FACSVerse; BD Biosciences).
Cholesterol measurement
Total cholesterol was extracted from
1×106 Mock and HIF-KO cells treated with or without 1 µM
simvastatin for 10 h at 37°C (n=3) by the traditional Bligh-Dyer
method using acetate, methanol and chloroform (23). In the final step, the dried
chloroform layer was dissolved in 50 µl of isopropanol and
quantified using a cholesterol measurement kit (LabAssay
Cholesterol; FUJIFILM Wako Pure Chemical Corporation).
Animal care
All animal care and procedures were conducted in
accordance with the Guiding Principles for the Care and Use of
Laboratory Animals at Hoshi University, as adopted by the
Institutional Animal Care and Use Committee on Animal Research of
Hoshi University (approved no. P24-091). A total of 81 mice were
used in this study; 7-week-old female syngeneic C57BL/6J mice
(Sankyo Labo Service Corporation) (total number, 39; weight ~18 g)
or 42 12-week-old C57BL/6 SCID mice (strain no. 001913; The Jackson
Laboratory; 29 female mice; weight ~21 g and 13 male mice; weight
~25 g). Mice were housed under specific pathogen-free conditions in
ventilated cages (floor area 501 cm2; five mice per
cage) with ALPHA-dri bedding (Shepherd Specialty Papers) and fed a
standard chow diet with access to water ad libitum. Animals
were inspected daily to ensure that they were not distressed during
the experiments. Rooms were temperature controlled at 22°C and kept
on a 12-h light/dark cycle. All procedures were performed under
inhalational anesthesia using Narcobit-E (Natsume Seisakusho Co.,
Ltd.) with isoflurane (FUJIFILM Wako Pure Chemical Corporation) at
a concentration of 4% for induction and 2% for maintenance. Before
analysis, the mice were euthanized by intraperitoneal injection of
a lethal dose (100 mg/kg) of pentobarbital sodium (Tokyo Kasei
Kogyo Co., Ltd.). After administering pentobarbital, mortality was
confirmed by both respiratory arrest (arrest of the chest wall
movement) and no response to a toe pinch. The chest cavity was then
opened to confirm cardiac arrest and death.
Tumor xenograft model
To establish tumor xenografts, AXT cells (including
Mock and HIF-KO cells) suspended in 50 µl of IMDM or 50 µl of
Matrigel (Corning, Inc.) were injected subcutaneously into the
flanks of 39 7-week-old female syngeneic C57BL/6J mice (Sankyo Labo
Service Corporation) or 42 12-week-old C57BL/6 SCID mice (strain
no. 001913; The Jackson Laboratory), respectively. Unless otherwise
specified, 1×106 cells were transplanted per site. Mice
were assigned randomly into experimental groups. For blinding
purposes, researchers performing the cell inoculation were aware of
group assignments, while outcome assessment was conducted by other,
blinded researchers. No exclusion criteria were set and data from
all mice were presented. The criteria for endpoints were as
follows: i) The mean tumor diameter was not >20 mm; ii) the
combined tumor burden was <15% of body weight (10-week-old mice
had a body weight of ~20 g); iii) there was no ulceration,
infection or necrosis of the tumor; iv) body weight loss was
>20% of the baseline weight. No mice reached the endpoint
criteria in the present study. Daily inspection confirmed that
growing tumors did not meet the endpoint criteria. The major and
minor axes of the tumors were measured, and the estimated tumor
weight was calculated from the following formula, with reference to
the guidelines of Washington State University (https://iacuc.wsu.edu/documents/2017/12/tumor-burden-guidelines.pdf/):
estimated tumor weight (mg)=tumor volume
(mm3)=d2 × D/2, where d and D are the
shortest and longest diameters in mm, respectively. The volume of
tumors with the maximum diameter in each mouse was listed in
Table SIII. After euthanasia,
tumors were collected, imaged and weighed for data collection. None
of the mice unexpectedly died during the study.
Gene set enrichment analysis (GSEA) of
adherent vs. non-adherent cells
Total RNA was collected from AXT cells cultured
under adherent or non-adherent conditions for 21 h at 37°C.
Comparison of these cells was performed using a microarray (Takara
Bio, Inc.). GSEA (24,25) was performed using the fgsea package
(26) in R software (version 4.3.1)
(27), based on gene expression
data obtained from microarray analysis. Genes were ranked according
to the log2-fold-change in expression between
non-adherent cells and adherent cells (n=1 each) and this ranked
list was used for GSEA. Gene sets were obtained from the Molecular
Signatures Database (28). An
enrichment plot for the ‘QI_HYPOXIA’ gene set (29) was generated using the plotEnrichment
function in fgsea.
GSEA and visualization of
significantly associated genes
Using total RNA collected from mice inoculated with
Matrigel embedded Mock and HIF-KO cells, RNA-seq was performed by
Rhelixa, Inc. Reads obtained from next generation sequencing were
mapped to the mouse reference genome (GRCm38) using HISAT2 (version
2.1.0) (30). Transcript-level read
counts were quantified using featureCounts (SCR_012919; version
1.6.3) (31). Transcripts with low
expression were also included in the analyses. GSEA was performed
using the fgsea package (26), with
genes ranked by log2-fold-change in expression between
HIF-1α knockout samples (n=3; KO1-1, KO1-2 and KO2) and Mock
controls (n=1). Enrichment results with a false discovery rate
(FDR) <0.2 were visualized using the plotGseaTable function in
fgsea. To examine the expression patterns of genes identified as
enriched by GSEA, a heatmap was generated using the pheatmap
package (SCR_016418) (32),
annotated with their corresponding gene sets. RNA-seq datasets in
this paper are available on the Gene expression omnibus site
(https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE307480).
The accession number is GSE307480.
Immunohistochemistry
Tumors were collected, fixed with 4%
paraformaldehyde at room temperature for 2 days and embedded in
paraffin. Deparaffinized and hydrophilized tissue sections (5-µm
thickness) were put into citrate buffer (0.01 mol/l, pH 6.0) and
heated in an oven at 90°C for 1 h for antigen retrieval. After a
brief wash with PBS, tissue sections were put into 3%
H2O2-methanol at room temperature for 8 min
for inactivation of intrinsic peroxidase activity. After a brief
wash with PBS, tissue sections were blocked with 3% bovine serum
albumin (BSA; FUJIFILM Wako Pure Chemical Corporation)-PBS at room
temperature for 1 h and subsequently incubated with the primary
antibodies listed in Table SII
diluted with 1.5% BSA-PBS at 4°C overnight. After washing with PBS
3 times, samples were treated with a horseradish peroxidase
(HRP)-conjugated Histofine simple stain kit (1:1 dilution, listed
in Table SII; Nichirei
Biosciences, Inc.) as the secondary antibody at room temperature
for 1 h. After washing with PBS, staining was developed using the
3,3′diaminobenzidine substrate (Impact DAB; Vector Laboratories,
Inc). Then Mayer's hematoxylin solution (FUJIFILM Wako Pure
Chemical Corporation) was used for nuclear staining at room
temperature for 1 min. For H&E staining, deparaffinized and
hydrophilized sections were stained with hematoxylin solution for 4
min at room temperature, washed with water and stained with 1%
eosin Y solution (FUJIFILM Wako Pure Chemical Corperation) for 2
min at room temperature.
Hypoxic regions were evaluated using a rabbit IgG
polyclonal anti-pimonidazole antibody (cat. no. PAb2627, NPI Inc.);
mice were injected intraperitoneally with pimonidazole
hydrochloride (60 mg/kg; NPI Inc.) 1 h before tumor collection. For
quantitative analysis of blood vessels, CD31 immunostaining was
observed under a BZ-X800 multifunctional microscope (Keyence
Corporation). Microvessel density was quantified as the
CD31-positive area normalized to the total solid tumor area using
BZ-X800 Analyzer exe 1.1.2.4 software (Keyence Corporation).
Statistical analysis
Unless indicated otherwise, quantitative data are
expressed as the mean ± SD relative to the control value and data
were analyzed by one-way analysis of variance (ANOVA) with the
Dunnet's post hoc test. For the angiogenesis analysis (Fig. 5D), the data did not meet the
assumption of normality using the Shapiro-Wilk test, therefore,
differences among groups were evaluated using the Kruskal-Wallis
test followed by Dunn's multiple comparison test. Shapiro-Wilk test
to assess normality was applied only for angiogenesis quantitation.
All analyses were performed using GraphPad Prism 9 (Dotmatics).
P<0.05 was considered to indicate a statistically significant
difference (*P<0.05, **P<0.01, ***P<0.005 and NS, not
significant). All assays were performed at least in triplicate. To
evaluate the magnitude of differences in tumor weight between
groups, effect sizes were calculated for each comparison between
the control (Mock) and knockout groups (KO1-1, KO1-2 and KO2) using
Hedges'g (33), a bias-corrected
standardized mean difference. Effect sizes and their 95% confidence
intervals (not shown) were estimated with the cohen.d function in
the effsize R package (34), with
missing values removed listwise. Group means and standard
deviations were calculated from the available numeric
observations.
Results
Osteosarcoma cells upregulate
downstream targets of HIF-1α under non-adherent conditions
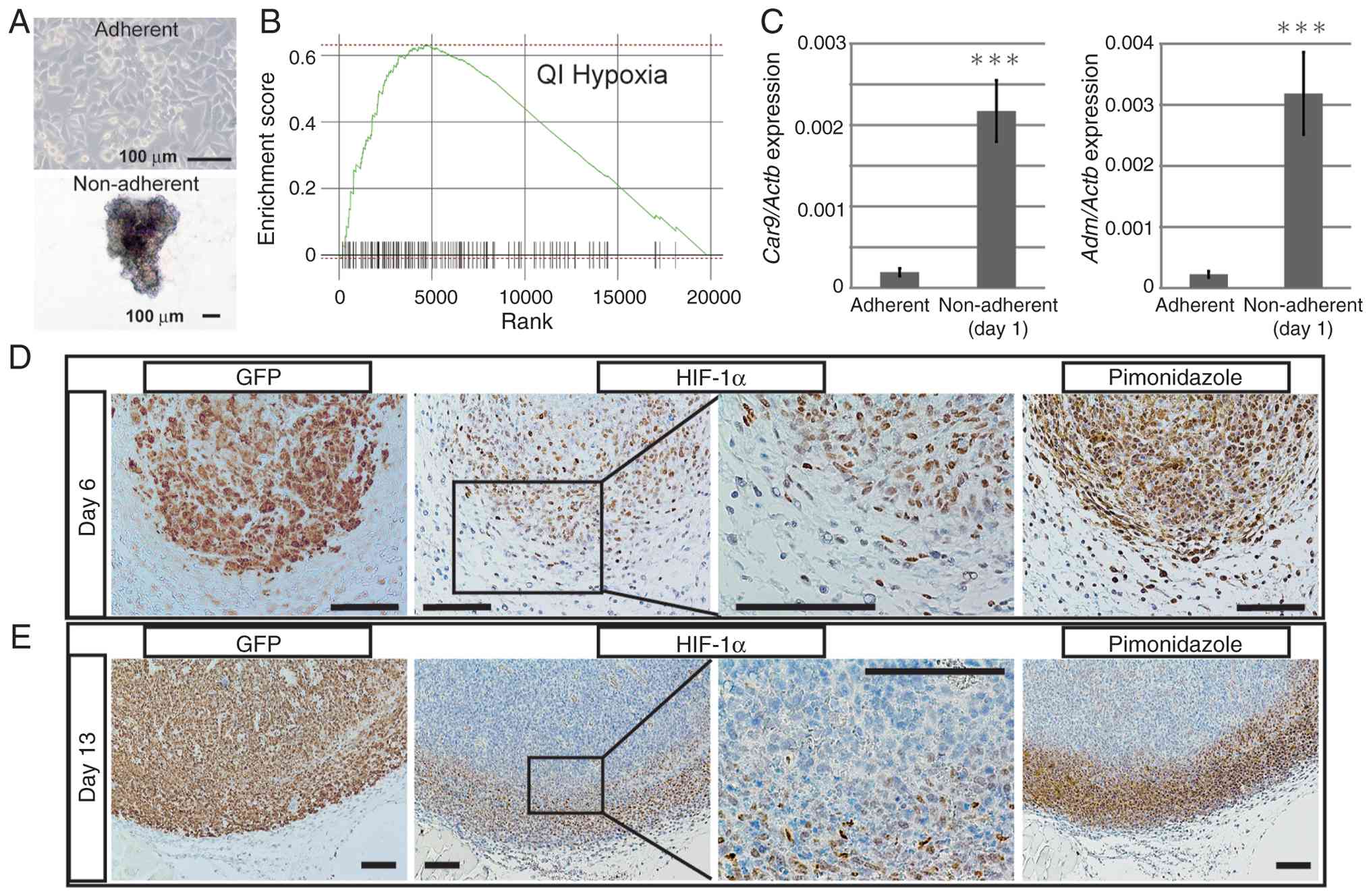
The ability of adherent cells to grow under
non-adherent anchorage-independent conditions is one of the
hallmarks of malignancy (7,8). Our previous study developed mouse
osteosarcoma AXT cells that proliferate both under adherent and
non-adherent conditions (Fig. 1A)
(35). Analysis of metabolite
levels suggested that non-adherent conditions mimic the in
vivo environment more closely when compared with adherent
normal culture conditions (35).
Therefore, therapeutic approaches that inhibit the mechanisms or
molecules enabling non-adherent cell growth might exert an
anti-tumor effects on osteosarcoma in vivo.
GSEA was used to analyze changes
(log2-fold) in expression of genes by adherent and
non-adherent AXT cells. Only ‘QI_HYPOXIA’, which comprises genes
upregulated under hypoxic conditions in prostate cancer cells
(29), was enriched significantly
in non-adherent cells (normalized enrichment score=2.05, FDR
<0.001; Fig. 1B). These results
suggest that hypoxia-related transcriptional programs may be more
activated in non-adherent cells than in adherent cells. Expression
of carbonic anhydrase 9 (Car9) and adrenomedullin
(Adm), both transcriptional targets of HIF-1α, in AXT cells
was higher under non-adherent conditions when compared with
adherent conditions (Fig. 1C).
Similarly, expression of HIF-1α transcriptional target genes
NDRG1 and ADM by human osteosarcoma cell lines Saos2
and U2OS was higher under non-adherent conditions compared with
adherent conditions (Fig. S1A and
B).
In vivo expression of HIF-1α in osteosarcoma
was analyzed by examining serial sections of tumors derived from
AXT cells. AXT cells express GFP, which allows direct detection of
tumor areas (Fig. 1D and E). In
small tumors on Day 6 post-inoculation, the nucleus of the majority
of tumor cells was positive for HIF-1α. Hypoxic areas in tissues
were visualized by administering pimonidazole, which is reductively
activated in hypoxic cells and forms stable covalent adducts with
thiol (sulphydryl) groups within amino acids (36). On Day 6 post-implantation, the
majority of tumor cells were positive for reductively activated
pimonidazole, which was detectable by a specific antibody,
indicating that the entire tumor was hypoxic at Day 6 (Fig. 1D). At 13 days post-implantation,
HIF-1α was highly expressed in peripheral tumor areas in which
osteosarcoma cells proliferate and infiltrate the surrounding
tissue. Importantly, areas with high expression of HIF-1α were
hypoxic (Fig. 1E). These results
suggest that hypoxic regions emerge during growth and progression
of osteosarcoma in vivo, and that HIF-1α may play an
important role.
Knocking out HIF-1α attenuates cell
growth under hypoxic and non-adherent conditions
AXT cells, in which HIF-1α was knocked out using
CRISPR-Cas9 with two different target sequences, were used to
evaluate the role of HIF-1α in growth of osteosarcoma. Two KO
clones (KO1-1 and KO1-2) and one clone (KO2) were obtained using
target sequences #1 and #2, respectively (Fig. 2A). Each cell type had a different
morphology. KO1-1 cells were similar to control (Mock) cells,
whereas KO1-2 cells were round and KO2 cells were larger when
compared with the Mock, KO1-1 and KO1-2 cells. In Mock cells,
HIF-1α protein was detectable under normoxic conditions, and
addition of the HIF-PH inhibitor roxadustat (37) led to accumulation of HIF-1α. By
contrast, no HIF-1α protein was detected in KO cells (Fig. 2B). Expression of Car9, a
transcriptional target of HIF-1α, was upregulated significantly in
Mock cells under hypoxic (2% O2) conditions compared
with normoxic conditions (Fig. 2C).
By contrast, no increase in Car9 was observed in HIF-KO
cells under hypoxic conditions. The effects of HIF-1α depletion on
cell proliferation were evaluated under adherent and non-adherent
culture conditions, as well as under normoxic and hypoxic
conditions. Under adherent and normoxic conditions, KO cells tended
to proliferate slightly more slowly when compared with Mock cells
(Fig. 2D). Under hypoxic
conditions, the difference was more pronounced, although the growth
rate of KO cells still remained high (Fig. 2E). Notably, the proliferation rate
of KO1-2 was almost equivalent with that of Mock cells. Under
non-adherent conditions, proliferation of KO1-1 and KO2 cells was
suppressed significantly, while proliferation of KO1-2 cells was
similar to that of Mock cells (Fig.
2F). There were differences between the tumor cell clones,
suggesting that the intracellular mechanism in KO1-2 cells allowed
proliferation even in the absence of HIF-1α. Under non-adherent
hypoxic conditions, proliferation of all knockout cells was lower
when compared with that of Mock cells (Fig. 2G). Cell cycle analysis showed that
under normoxic and adherent conditions, there was no reduction in
the number of S-phase KO cells compared with Mock cells, and no
significant induction of apoptosis (Fig. S2A-D). As non-adherent cells were
fragile and the majority of cells were physically damaged during
sample preparation, only live cells were gated and analyzed
(Figs. S2E, 2H and I). Consistent
with cell growth under non-adherent normoxic conditions, the
S-phase population of KO1-1 and KO2 cells, but not KO1-2 cells, was
lower when compared with that of Mock cells (Fig. 2H and J). The S-phase population of
Mock cells was lower under non-adherent hypoxic conditions when
compared with under normoxic conditions (Fig. 2J and K). The S-phase population of
KO1-1 and KO2 cells was smaller when compared with that of Mock
cells, whereas the S-phase population of KO1-2 was the same as that
of Mock cells, while the number of cells in G2/M was lower when
compared with that of Mock cells (Fig.
2I and K). Consistent with these results, expression of mRNA
encoding Cyclin D1 in KO1-1 and KO2 under non-adherent hypoxic
culture was significantly lower when compared with that in Mock
cells, while that in KO1-2 tended to be lower when compared with
that in Mock cells, although the difference was not significant
(Fig. 2L).
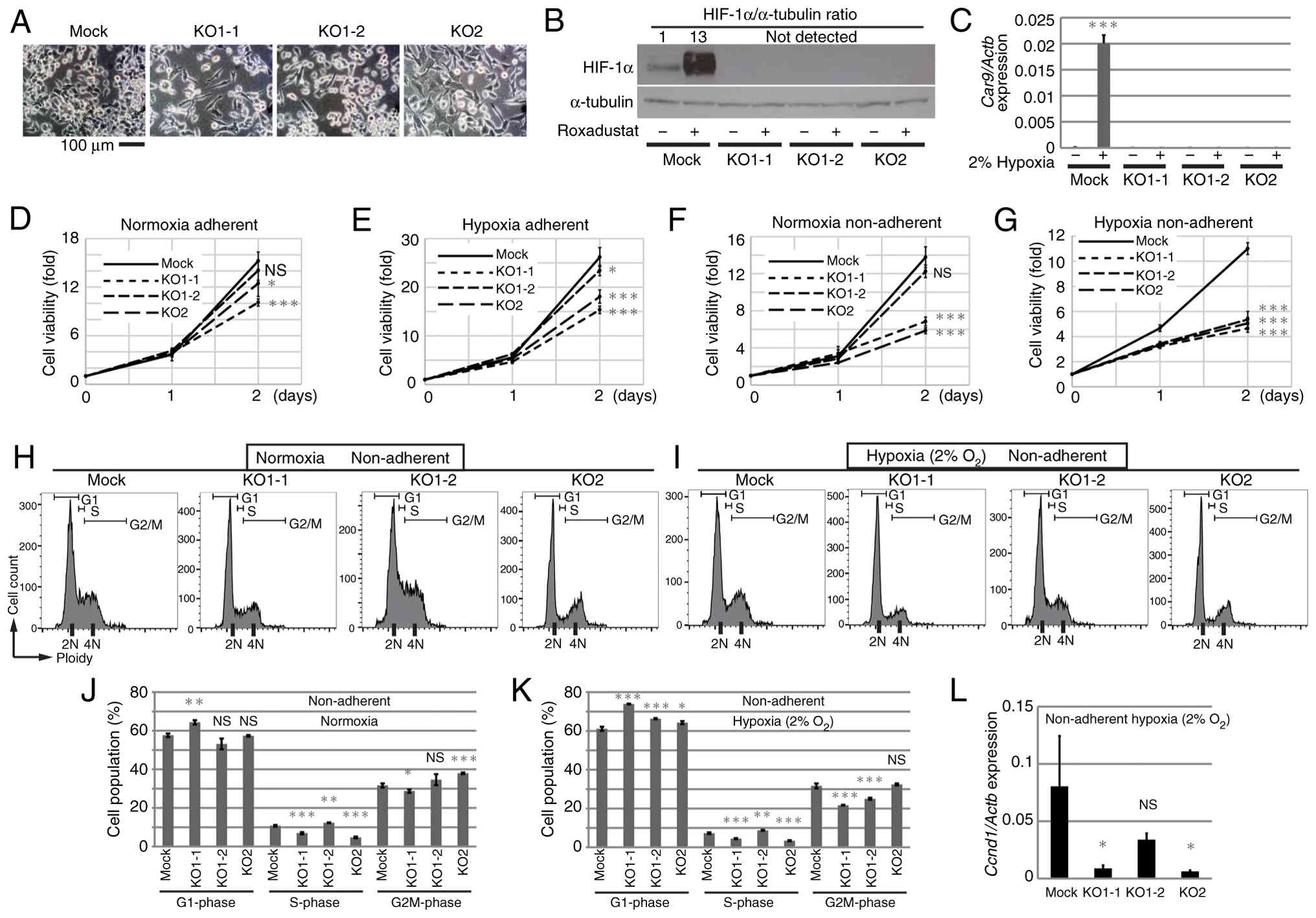 | Figure 2.Attenuation of cell growth in
HIF-1α-knockout cells in vitro. (A) Morphology of control
(Mock) and HIF-1α-knockout AXT (HIF-KO, KO1-1, 1–2 and KO2) cells.
(B) Western blot analysis of HIF-1α expression in Mock and HIF-KO
cells in the presence/absence of 25 µM roxadustat for 27 h. The
relative (fold) values of HIF-1α against the corresponding control
value normalized to the intensities α-tubulin bands are shown.
HIF-1α bands of HIF-KO cells were not detected. (C) Reverse
transcription-quantitative PCR analysis of Car9 in Mock and
HIF-KO cells under normoxic or hypoxic (2% O2)
conditions for 20 h. Data are normalized to the corresponding
levels of Actb mRNA, and are shown as the mean±SD of
triplicate values. Growth of Mock and HIF-KO cells under (D)
normoxia and adherent conditions, (E) hypoxia and adherent
conditions, (F) normoxia and non-adherent conditions and (G)
hypoxia and non-adherent conditions. The ratio relative to the
value for Day 0 was calculated for each data point (n=4; D-G). Flow
cytometry analysis of DNA content in Mock and HIF-KO cells under
(H) normoxia and non-adherent conditions and (I) hypoxia and
non-adherent conditions for 31 h. (J) The population of each cell
fraction in (H) are shown (n=3). (K) The population of each cell
fraction in (I) are shown (n=3). (L) Reverse
transcription-quantitative PCR analysis of Ccnd1 in Mock and
HIF-KO cells under non-adherent or hypoxic (2% O2)
conditions for 21 h (n=3). HIF, hypoxia inducible factor; KO,
knockout. *P<0.05, **P<0.01, ***P<0.005 and NS, not
significant. |
These results suggest that HIF-1α supports
proliferation under non-adherent hypoxic conditions in
vitro; however, the mechanism used by KO1-2 cells might
maintain high proliferation even in the absence of HIF-1α.
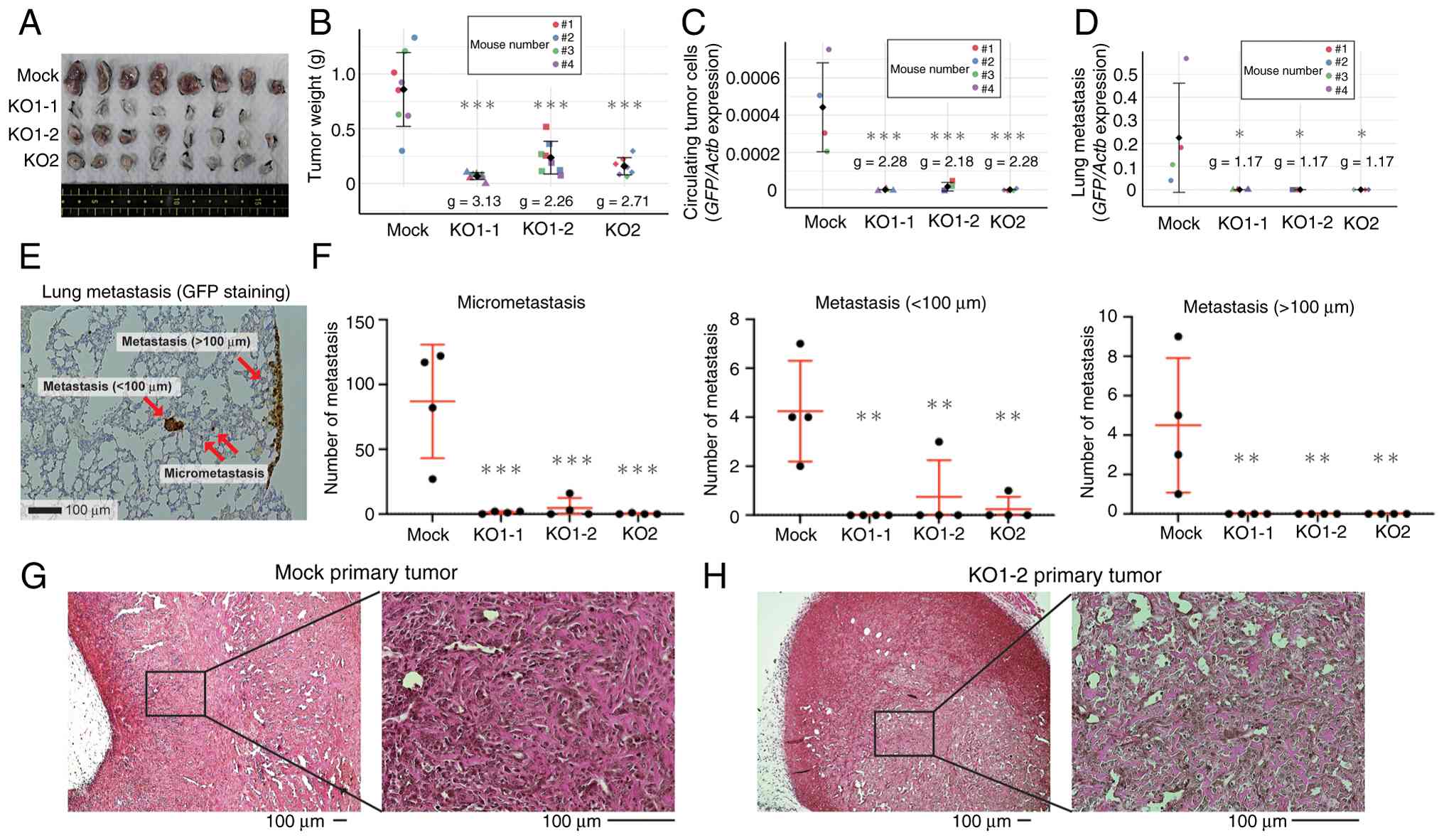
Depleting HIF-1α decreases tumorigenic
activity in vivo
AXT cells form osteoid-rich tumors, a characteristic
essential for the definitive diagnosis of osteosarcoma, as well as
lung metastasis in C57BL/6 mice (4–6). In
the present study, HIF-KO cells were transplanted into mice to
evaluate tumorigenicity. The weight of the primary KO cell tumors
21 days after subcutaneous implantation of 1×106 cells
was significantly lower when compared with that of Mock tumors
(Fig. 3A and B). No tumors
developed at the transplantation site in one mouse inoculated with
KO1-1, suggesting that the suppression of tumorigenic activity
in vivo is stronger when compared with that of cell growth
in vitro.
After forming primary tumors, AXT cells
spontaneously develop hematogenous metastatic lesions. Because AXT
cells express GFP, the amount of lung metastases and CTCs in the
blood can be quantified by measuring the expression level of GFP
(4,5,35).
Such measurements revealed that the number of lung metastases and
CTCs was reduced significantly after KO of HIF-1α (Fig. 3C and D). Further evaluation of lung
metastases by immunohistochemical staining for GFP also showed a
significant reduction of lung metastasis; in particular, there were
no lesions with a diameter >100 µm in HIF-KO cell-inoculated
mice (Fig. 3E and F). Importantly,
tumors derived from HIF-KO cells also contained osteoids, and there
were no histological changes suggestive of osteosarcoma after
knockout of HIF-1α (Figs. 3G and H,
S3A and B).
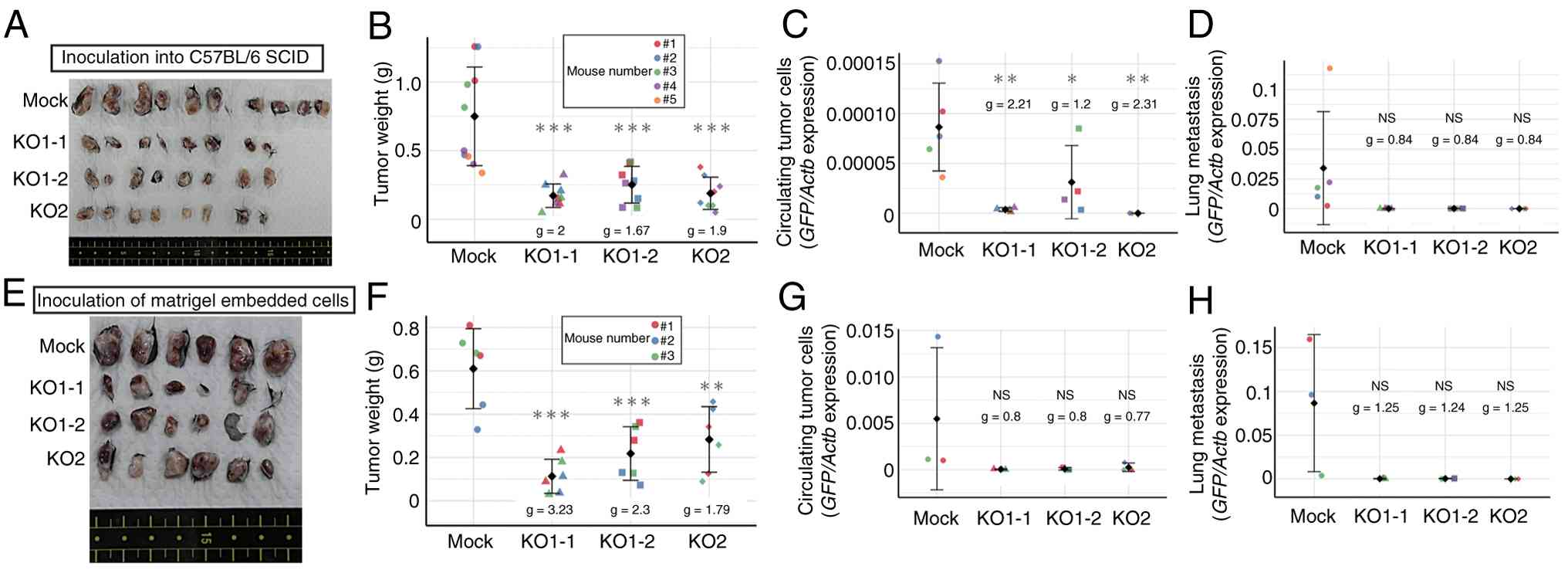
Altering inoculation conditions does
not rescue the reduced tumorigenicity induced by HIF-1α KO
Accumulating evidence indicates that hypoxic
environments affect tumor immunity (38,39),
and that suppression of HIF-1α in tumor cells activates tumor
immunity (40,41). Therefore, tumor cells were
transplanted into SCID mice to examine whether tumor immunity is
involved in the regression of HIF-1α KO tumors. Similar to
transplantation into C57BL/6 mice, transplantation into SCID mice
did not restore tumorigenic and metastatic activity completely
(Fig. 4A-D). Thus, it seems that
tumor immunity mediated by T cells and B cells is not the main
suppressor of tumor formation by HIF-KO cells. Next, whether
further improvements in the transplant environment increased tumor
cell engraftment was tested. For this purpose, Matrigel was used to
provide a basement membrane to prepare the microenvironment from
the time of transplantation (42).
Studies show that co-injection with Matrigel increases tumor
formation by cancer cell lines and enhances engraftment of primary
human epithelial cancer cells in immunocompromised mice (43,44);
however, in the present study, KO cells failed to form tumors and
metastases as efficiently as Mock cells (Fig. 4E-H).
The small group sizes of in vivo experiments
is a limitation in the present study. However, the knockout groups
demonstrated reduced tumor growth compared with the Mock controls.
The effect size estimates consistently indicate reduced tumor
growth in all knockout groups relative to the Mock control,
suggesting that HIF-1α plays an important role in tumorigenesis and
progression of osteosarcoma in vivo.
Osteosarcoma develops a rich
vasculature even in the absence of HIF-1α
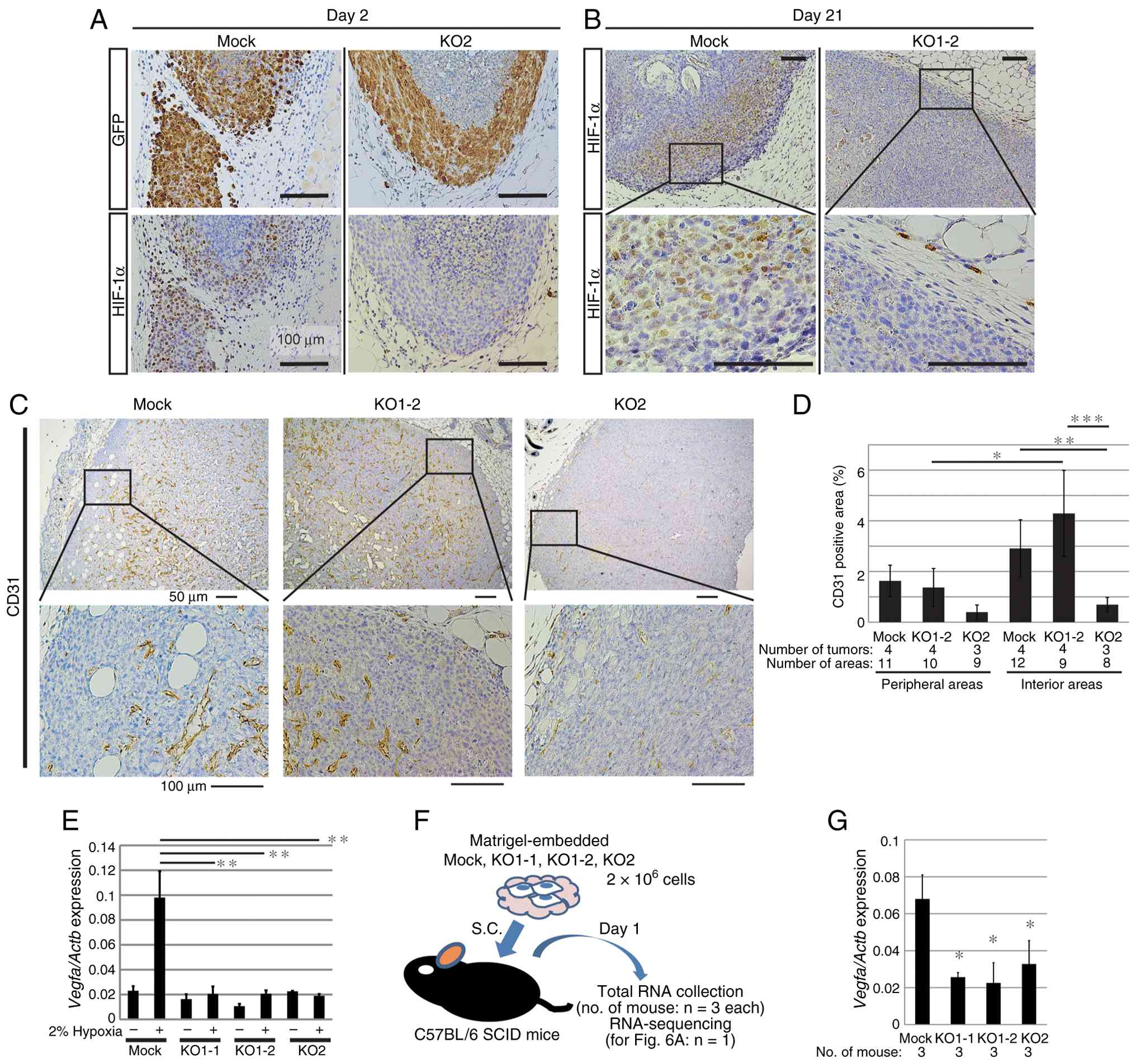
To clarify the functions of HIF-1α, its expression
in tumors immunohistochemically was evaluated over time. At 2 days
after subcutaneous implantation of Mock-derived tumors, high
nuclear expression of HIF-1α was observed throughout areas of dense
viable cells (except the interior, which harbored GFP-negative dead
cells), whereas KO tumors did not express HIF-1α (Fig. 5A). At 21 days post-implantation, the
interior of the tumor was filled with viable cells, with notable
bone formation and HIF-1α was highly expressed in the peripheral
areas in which cells were proliferating rapidly (Fig. 5B). Again, no expression of HIF-1α
was observed in KO cell tumors. Analysis with pimonidazole
suggested that areas with high HIF-1α expression were hypoxic
(Fig. 1D and E). The pattern of
hypoxic areas appearing mainly around the tumor periphery was the
same in Mock- and HIF-KO-derived tumors (Fig. S4A), suggesting that oxygen supply
inside the tumor was not disrupted by HIF-1α loss. Therefore, tumor
angiogenesis was evaluated. Staining for CD31, a marker of vascular
endothelial cells, suggested that the inside of the tumor was
vascularized (Figs. 5C and S4A). Quantification of CD31 positive
areas indicated that the KO1-2 cell-derived tumors had abundant
blood vessels at a level comparable to that of the mock-derived
tumors, while tumors derived from KO2 are less vascularized when
compared with those from Mock cells (Figs. 5D and S4B). Consistent with the emerging pattern
of hypoxic areas, vascularization tended to be more prevalent
within the tumor when compared with its periphery. These findings
suggest that global tumor oxygen supply is not severely disrupted
by HIF-1α deletion. Expression of vascular endothelial growth
factor A (VEGF-A), a transcriptional target of HIF-1α, increased in
Mock cells under hypoxic conditions in vitro, but not in KO
cells (Fig. 5E); however, unlike
Car9 (Fig. 2C), expression
of Vegfa was not negligible in KO cells (Fig. 5E). To analyze gene expression in
tumor cells in vivo, each cell type (embedded in Matrigel)
was implanted subcutaneously into mice, harvested on the next day
and total RNA was extracted (Fig.
5F). Expression of Vegfa in Mock cells was ~2-fold
higher when compared with in KO cells, but robust expression was
still detected in KO cells (as observed in vitro; Fig. 5G). Regarding an alternative driver
of VEGF-A regulation, HIF-2α was expressed at significantly lower
levels when compared with HIF-1α in Mock and HIF-KO cells as
assessed by RT-qPCR analysis (Fig.
S5A). Furthermore, under hypoxic conditions, the addition of
HIF-2α inhibitors, PT2385 or TC-S7009, had little effect on
Vegfa expression (Fig.
S5B). STAT3 activation differed between HIF-KO clones, making
it unlikely to be a common molecular mechanism for increasing
Vegfa in HIF-KO cells (Fig.
S5C). Notably, treatment of mTOR inhibitors, temsirolimus or
everolimus, under hypoxia reduced the expression level of
Vegfa in all HIF-KO cells, although temsirolimus did not
significantly affect that in KO1-2 cells (Fig. S5D). Therefore, mTOR pathway is
partially responsible for maintaining the basal Vegfa
expression in the absence of HIF-1α. Importantly, it was suggested
that non-tumor cells may be involved in angiogenesis in
vivo. Immunostaining of α-smooth muscle actin (αSMA), a useful
marker for a part of fibroblasts including cancer-associated
fibroblasts (45), revealed
αSMA-positive cells comprising various cells including tumor cells,
fibroblastic cells and CD31-positive cells, likely representing
mature vascular endothelial cells (Fig. S5E). In addition, F4/80-positive
macrophages were abundantly accumulated at the tumor periphery and
within the tumor (Fig. S5F). These
stromal cells may support the angiogenesis of HIF-KO-derived tumors
in vivo.
These results indicate that HIF-1α-independent
expression of VEGF-A in osteosarcoma cells may be sufficient to
form vasculature to support tumor growth, even though the HIF-KO
tumors are smaller than those formed by Mock cells.
HIF-KO cells show differing dependence
on glycolysis for growth
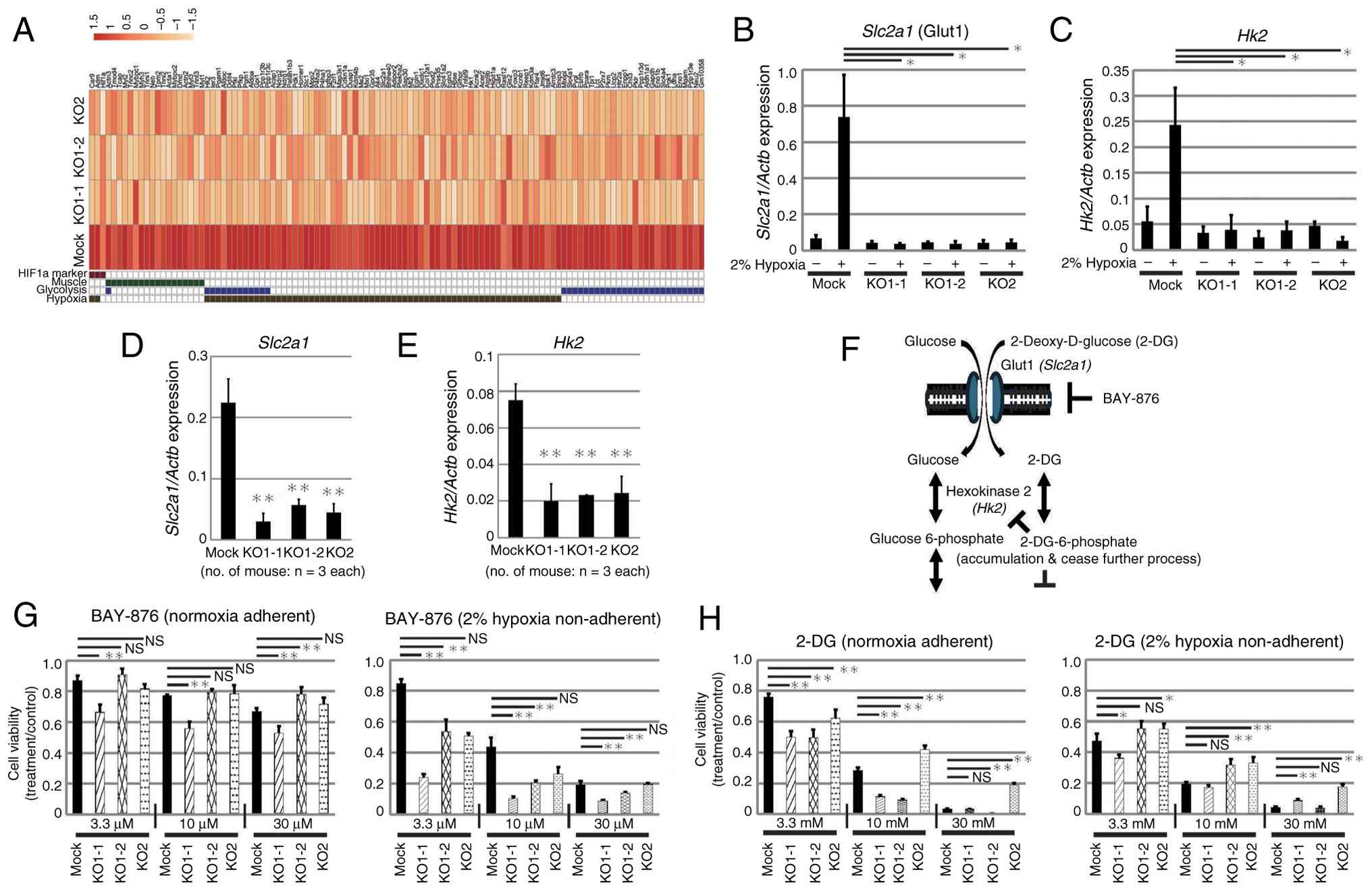
To explore changes in gene expression upon depletion
of HIF-1α in vivo, RNA-seq analysis was carried out. GSEA
revealed that genes included in the QI_HYPOXIA gene set were
enriched among those downregulated in HIF-KO samples relative to
Mock (Fig. S6) which is consistent
with the GSEA findings from the comparison between non-adherent and
adherent cells (Fig. 1B). In
addition to hypoxia-related genes, several gene sets associated
with muscle function or anion transport and carbohydrate catabolism
were downregulated in HIF-1α KO samples compared with the Mock
control (Figs. 6A and S6). By contrast, few gene sets contained
genes with high upregulation ratios in KO cells, and no gene set
with significantly increased expression was extracted. In addition,
there were no common genes whose expression levels in KO cells were
>2 fold when compared with those in Mock cells. Although
depleting HIF-1α suppressed tumor formation significantly in
vivo, it failed to eliminate tumors completely (Figs. 3 and 4). Thus, treatments in addition to HIF-1α
depletion would be required to overcome osteosarcoma, and
elucidation of the survival pathways on which KO cells depend in
the absence of HIF-1α was attempted.
Expression of glucose transporter 1 (Glut1 coded by
Slc2a1), which is involved in glucose uptake and hexokinase
2 (Hk2), an enzyme involved in the early stage of
glycolysis, increased significantly in Mock cells under hypoxic
culture, but not in KO cells (Fig. 6B
and C). In addition, in vivo expression levels of these
genes in cells collected as described in Fig. 5F, was significantly increased in
Mock cells when compared with that in KO cells (Fig. 6D and E). Therefore, the functions of
these molecules were inhibited and the effects on proliferation
under normoxia and hypoxia were analyzed (Fig. 6F). The sensitivity of all cells to
BAY-876, a specific inhibitor of Glut1, was higher under hypoxic
conditions, and proliferation was suppressed in a dose-dependent
manner, suggesting that hypoxic cells are more dependent on
glycolysis when compared with normoxic cells (Fig. 6G). KO cells tended to be more
sensitive when compared with Mock cells to low concentrations of
BAY-876. KO1-1 cells were the most sensitive, suggesting a
dependency on glycolysis. 2-Deoxy-D-glucose (2-DG) is taken up into
cells (as is glucose) and phosphorylated by hexokinase 2; however,
the subsequent reactions do not proceed, resulting in accumulation
within the cell and inhibition of glycolysis (46). Dose-dependent inhibition of
proliferation was observed for all cells under both normoxic and
hypoxic conditions (Fig. 6H).
Notably, KO1-1 cells tended to be more sensitive to 2-DG compared
with Mock, KO1-2 and KO2 cells while KO2 cells were less sensitive
(as in the case of BAY-876). These results indicate that Glut1 and
HK2 function during proliferation of HIF-KO cells, and that
sensitivity to inhibition varies among clones, suggesting that the
clones have differing levels of dependence on glycolysis for
survival.
The pathways upon which osteosarcoma
cells rely to survive in the absence of HIF-1α are
heterogeneous
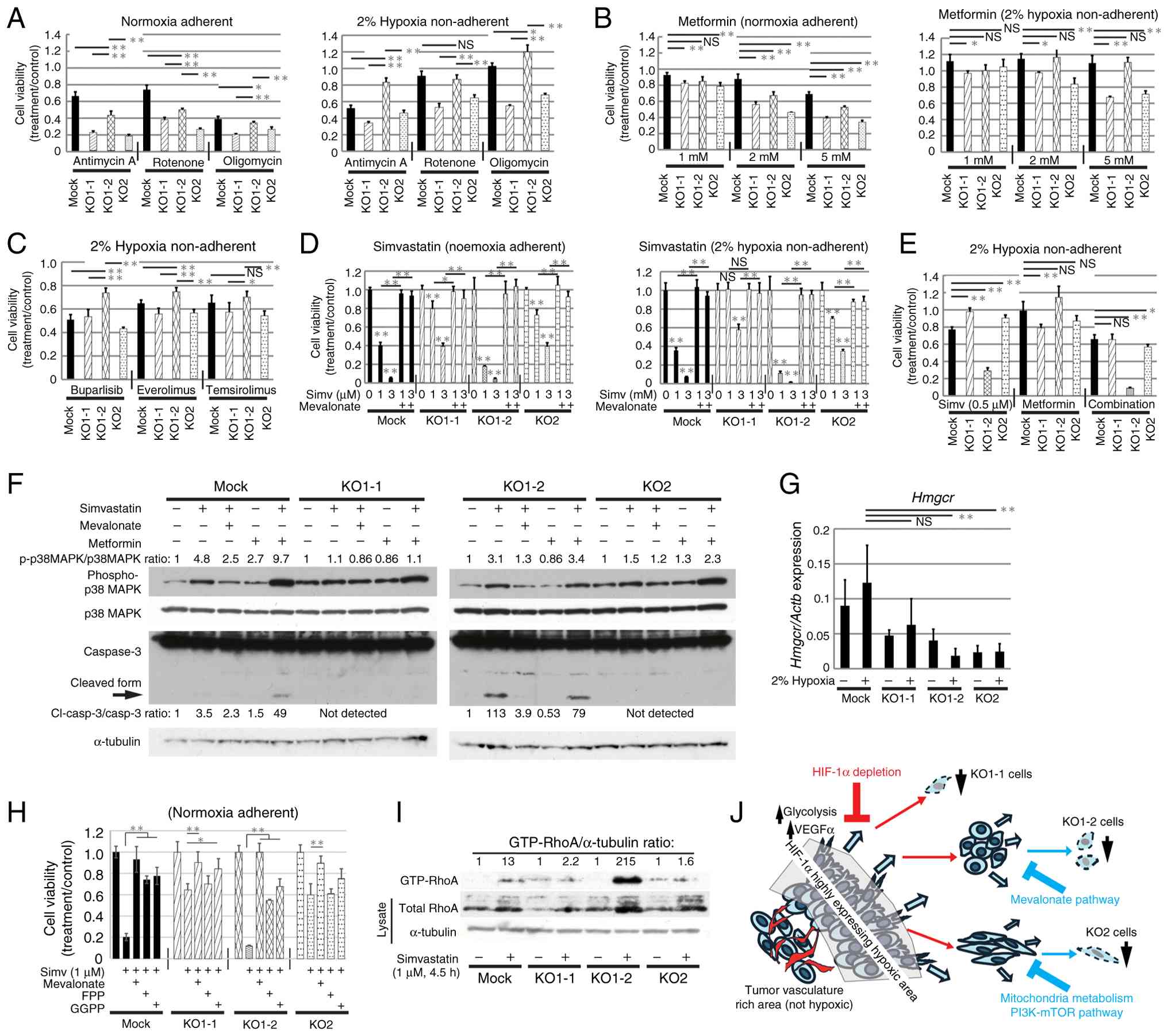
To clarify the pathways upon which HIF-1α KO cells
depend for survival, the effects of various compounds on
proliferation was analyzed. The sensitivity of KO cells to
mitochondrial electron transport inhibitors antimycin A, rotenone
and oligomycin was higher under normoxic conditions when compared
with under hypoxic conditions (Fig.
7A), indicating that the cells were more dependent on
mitochondria for survival under normoxia. Growth inhibition induced
by these inhibitors was weaker in Mock cells and KO1-2 cells when
compared with that in KO1-1 and KO2 cells. A similar trend was
observed when metformin was added. Under hypoxic conditions,
treatment with up to 5 mM metformin did not inhibit growth of Mock
or KO1-2 cells, but did inhibit that of KO1-1 and KO2 cells
(Fig. 7B). These findings suggest
that Mock and KO1-2 cells are less dependent on the mitochondrial
electron transport chain for survival under hypoxia when compared
with KO1-1 and KO2 cells. KO1-2 cells were less sensitive to the
PI3K inhibitor buparlisib and the mTOR inhibitors everolimus and
temsirolimus, when compared with Mock, KO1-1 and KO2 cells
(Fig. 7C). In addition, the
sensitivity of KO1-2 to shikonin, which inhibits Pkm2 (47), was lower when compared with that of
Mock, KO1-1 and KO2 cells (Fig.
S7A), suggesting that survival of KO1-2 cells depends on a
different mechanism.
Inhibition of the mevalonate synthesis pathway
markedly and dose-dependently suppressed the proliferation of KO1-2
cells under both normoxic and hypoxic conditions (Figs. 7D and S7B). Growth inhibition by simvastatin or
atorvastatin was prevented by addition of mevalonate, indicating
that this inhibition is specific to the mevalonate synthesis
pathway. Treatment with statins suppressed growth of all cells, but
the sensitivity of KO1-1 and KO2 was relatively low. Our previous
study demonstrated that combined administration of a statin and
metformin inhibited growth and induced apoptosis of osteosarcoma
cells (6). Therefore, the present
study analyzed the sensitivity of each cell clone to co-treatment
with simvastatin and metformin under hypoxic conditions (Fig. 7E). Co-treatment of Mock and KO1-2
cells, whose proliferation was not suppressed by metformin alone,
with simvastatin decreased proliferation. By contrast, KO1-1 and
KO2 cells, which were less sensitive to simvastatin than Mock and
KO1-2 cells, showed synergistic growth suppression when co-treated
with metformin. Similar synergistic inhibition was observed upon
combined treatment with atorvastatin and metformin (Fig. S7C). Our previous study suggests
that simvastatin induces apoptosis in osteosarcoma cells via
activation of p38MAPK and AMPK (6).
The present study found that simvastatin increased phosphorylation
of p38MAPK in Mock and KO1-2 cells, which was abolished by addition
of mevalonate, indicating that this phosphorylation is
mevalonate-dependent (Fig. 7F).
Activation of AMPK became more pronounced with higher
concentrations and longer treatment of simvastatin, and the
intensity of activation was stronger in KO1-2 and Mock cells
(Fig. S7D and E). In KO1-2 cells
treated with 0.5 µM of simvastatin alone for 17 h, expression of
cleaved Caspase 3 was detected (Fig.
7F), suggesting induction of apoptosis. Co-treatment with
simvastatin and metformin increased phosphorylation of p38MAPK in
all cells. Notably, cleaved Caspase 3 was detected in Mock and
KO1-2 cells after treatment with simvastatin and metformin, which
is consistent with the high sensitivity of both cells to
simvastatin (Fig. 7E and F).
Expression levels of HMG-CoA reductase
(Hmgcr) and synthase (Hmgcs) enzymes, which are key
components to the mevalonate synthesis pathway, did not vary
significantly between cells under normoxic and hypoxic conditions.
The protein level of HMGCR did not change with simvastatin
treatment (Figs. 7G and S7F and G). Furthermore, the present study
found no significant differences in expression between cells after
transplantation (Fig. S7H and I;
samples prepared as in Fig. 5F).
Sensitivity to statins may change with forced expression of HMGCR.
The present study attempted to establish HMGCR-highly expressing
KO1-2 cells using lentivirus but this could not be achieved (data
not shown). Subsequently, the present study analyzed the other
molecular changes in each cell type following the treatment of
simvastatin. The amount of cholesterol in the cells tended to be
slightly lower in Mock and KO1-2 cells and decreased similarly in
all cells with the addition of simvastatin (Fig. S7J). Growth inhibition by
simvastatin was significantly prevented by co-treatment of FPP or
GGPP, although the rescue was not as complete as with mevalonate
(Fig. 7H). FPP and GGPP are
required for post-translational prenylation of small GTPases of the
Ras and Rho families (48,49). The highly sensitive KO1-2 cells
showed markedly increased intracellular accumulation of GTP-RhoA
when compared with the Mock, KO1-1 and KO2 cells with 4.5 h of
simvastatin treatment (Fig. 7I).
This finding was consistent with the changes observed in the parent
AXT cell line when simvastatin caused apoptosis (6). Thus, abnormalities in signals involved
in the prenylation pathway such as differences in intracellular
GTP-RhoA accumulation, might be more strongly associated with
simvastatin sensitivity than differences in cholesterol synthesis.
To clarify the mechanisms underlying the emergence of cells with
different sensitivity to simvastatin, the parental AXT cells were
subjected to limiting dilution, single cell cloning was performed
and the sensitivity to simvastatin was evaluated (Fig. S7K). The results showed that the
sensitivity to simvastatin differed between clones, indicating that
heterogeneity pre-existed in the parental AXT cells.
Collectively, these data suggest that
HIF-1α-independent survival pathways differ among KO clones and
that the mevalonate synthesis pathway may support cell growth under
hypoxic conditions in the absence of HIF-1α (Fig. 7J).
Discussion
The findings of the present study suggest that
HIF-1α plays a key role in rapid progression of osteosarcoma in
vivo. The effect of HIF-1α depletion was more pronounced in
vivo than in vitro culture, suggesting that the in
vivo environment is more hostile to cancer cells in terms of
oxygen and nutritional conditions when compared with in
vitro, and that the function of HIF-1α is important in
vivo. Reduction of tumorigenicity persisted even after changing
the transplantation conditions. When transplanted into
immune-competent mice, tumorigenesis of KO1-1 cells was
particularly poor, with one mouse showing no tumor formation after
inoculation. By contrast, when inoculated into SCID mice, KO1-1
cells formed tumors readily, showing a tendency toward increased
tumorigenicity. Reduced growth in vivo after HIF-1α
depletion may be advantageous in that it allows anti-tumor immune
responses to eliminate tumor cells; thus, HIF-1α may be a potential
therapeutic target for osteosarcoma. However, eradication of tumors
that form even in the absence of HIF-1α is necessary for treatment
to be completely successful. Notably, even after HIF-1α depletion,
rich vascularization can occur and the non-negligible basal
expression of Vegfa was observed in the present study. The
findings suggest that HIF-1-independent mechanisms, such as
activation of cell intrinsic signals in cancer cells (for example,
activation of the mTOR pathway) and the tumor microenvironment, are
involved in the regulation of Vegfa expression and vascular
formation. Therefore, although HIF-targeted therapy is expected to
suppress tumor vasculature formation, this strategy may be
ineffective in some cell types.
Previously, drugs that target HIF specifically are
being used clinically to treat cancer types that depend on HIF for
survival (50). In 2023, the
HIF-2α-targeting drug belzutifan was approved by the US Food and
Drug Administration for use in patients with advanced renal cell
carcinoma (51). Notably, a
mutation that interferes with drug binding and precludes HIF-2
complex dissociation has been identified as the cause of resistance
to the HIF-2 inhibitor PT2385 (52). Moreover, even when HIF inhibitors
suppress the function of HIF, tumor progression can still occur, as
shown in the present study in the osteosarcoma model. Analysis of
HIF-1α knockout clones revealed that individual cells have
different properties. The results of compound screening revealed
that each of the three clones depended on a different pathway for
survival. The difficulty is that in the absence of HIF-1α, the
pathway on which survival depends cannot be easily identified.
Furthermore, in clinical settings, these clones may be mixed within
osteosarcoma tumors.
The reasons why such heterogeneity exists, and its
emerging mechanisms are important issues. To adapt to the
environment and survive various stimuli, malignant tumor cells
plastically change their phenotype through the acquisition of
genetic mutations and epigenetic modifications (for example,
drug-tolerant persister cells) (53). Single-cell RNA-seq is a definitive
method providing useful information for solving this problem.
However, due to several constraints, it was difficult to perform
this for the present study. In addition, the present study
acknowledges other limitations including that the group sizes of
in vivo experiments were small owing to the ethical and
feasibility constraints. The transcriptome analysis was conducted
on only one sample (n=1), and statistical analysis was not
possible. Therefore, the graph shows the fold change using lenient
thresholds for the purpose of screening to identify variable genes.
The assessment of metabolic flux using labeled precursors is also
required as a future direction to elucidate the mechanism of
differential mevalonate dependence between HIF-KO clones.
Finally, the data suggested that although targeting
HIF-1 is a potentially attractive therapy for osteosarcoma, to
achieve a complete cure, it may be necessary to identify and
inhibit pathways that tumor cells rely on for survival in the
absence of HIF-1.
Supplementary Material
Supporting Data
Supporting Data
Acknowledgements
We thank Ms. Ikuyo Ishimatsu (Institute of Science
Tokyo) and Ms. Erika Alexandra Meyer (Hoshi University) for
experimental assistance and Dr. Manabu Kawada (Institute of
Microbial Chemistry) and Dr. Hiroyuki Seimiya (The Cancer Institute
of JFCR) of AdAMS project (JSPS KAKENHI JP 22H04922) for technical
assistance of compound screening.
Funding
This work was supported by Japan Society for the Promotion of
Science (grant no. KAKENHI 24K10343).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author. The data generated in the
present study may be found in the Gene Expression Omnibus under
accession number GSE307480 or at the following URL: https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE307480.
Authors' contributions
TS contributed to conceptualization, resources,
data curation, software, formal analysis, supervision, funding
acquisition, validation, investigation, visualization, methodology,
writing-original draft and project administration, AK contributed
to conceptualization, resources, data curation, formal analysis,
validation, investigation, visualization, methodology and
writing-original draft. TT contributed to resources, data curation,
software, formal analysis, validation, investigation,
visualization, methodology and writing-original draft. AS
contributed to data curation, software, formal analysis,
validation, investigation and visualization. HN, HH, SH, YT, HM and
YF contributed to resources, data curation, conducting experiments
and contributing to the design of experimental models. AM and HS
contributed to conceptualization, resources, data curation,
funding, supervision, validation, methodology and project
administration. All authors approved the final version of the
manuscript.
Ethics approval and consent to
participate
All animal care and procedures were performed in
accordance with the guidelines of Hoshi University, and the present
study was approved by the Committee on Animal Research of Hoshi
University (approval no. P24-091).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Fletcher CDM, Bridge JA, Hogendoorn PCW
and Mertens F: Tumors of soft tissue and bone. WHO Classification
of tumors. 4th edition. IARC Press; Lyon: 2013
|
|
2
|
Gill J and Gorlick R: Advancing therapy
for osteosarcoma. Nat Rev Clin Oncol. 18:609–624. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kager L, Tamamyan G and Bielack S: Novel
insights and therapeutic interventions for pediatric osteosarcoma.
Future Oncol. 13:357–368. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Shimizu T, Ishikawa T, Sugihara E,
Kuninaka S, Miyamoto T, Mabuchi Y, Matsuzaki Y, Tsunoda T, Miya F,
Morioka H, et al: c-MYC overexpression with loss of Ink4a/Arf
transforms bone marrow stromal cells into osteosarcoma accompanied
by loss of adipogenesis. Oncogene. 29:5687–5699. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shimizu T, Sugihara E, Yamaguchi-Iwai S,
Tamaki S, Koyama Y, Kamel W, Ueki A, Ishikawa T, Chiyoda T, Osuka
S, et al: IGF2 preserves osteosarcoma cell survival by creating an
autophagic state of dormancy that protects cells against
chemotherapeutic stress. Cancer Res. 74:6531–6541. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Kamel WA, Sugihara E, Nobusue H,
Yamaguchi-Iwai S, Onishi N, Maki K, Fukuchi Y, Matsuo K, Muto A,
Saya H and Shimizu T: Simvastatin-induced apoptosis in osteosarcoma
cells: A key role of RhoA-AMPK/p38 MAPK signaling in antitumor
activity. Mol Cancer Ther. 16:182–192. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Paoli P, Giannoni E and Chiarugi P:
Anoikis molecular pathways and its role in cancer progression.
Biochim et Biophys Acta. 1833:3481–3498. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Schafer ZT, Grassian AR, Song L, Jiang Z,
Gerhart-Hines Z, Irie HY, Gao S, Puigserver P and Brugge JS:
Antioxidant and oncogene rescue of metabolic defects caused by loss
of matrix attachment. Nature. 461:109–113. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Semenza GL: Targeting HIF-1 for cancer
therapy. Nat Rev Cancer. 3:721–732. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Pouysségur J, Dayan F and Mazure NM:
Hypoxia signalling in cancer and approaches to enforce tumour
regression. Nature. 441:437–443. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bertout JA, Patel SA and Simon MC: The
impact of O2 availability on human cancer. Nat Rev
Cancer. 8:967–975. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Luo D, Ren H, Zhang W, Xian H, Lian K and
Liu H: Clinicopathological and prognostic value of
hypoxia-inducible factor-1 in patients with bone tumor: A
systematic review and meta-analysis. J Ortho Surg Res. 14:1–12.
2019. View Article : Google Scholar
|
|
13
|
Yang QC, Zeng BF, Dong Y, Shi ZM, Jiang ZM
and Huang J: Overexpression of hypoxia-inducible factor-1α in human
osteosarcoma: Correlation with clinicopathological parameters and
survival outcome. Jpn J Clin Oncol. 37:127–134. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ren H, Zhang YH, Li HY, Xie T, Sun LL, Zhu
T, Wang SD and Ye ZM: Prognostic roles of hypoxia-inducible
factor-1 alpha expression in osteosarcoma: A meta-analysis. Onco
Targets Ther. 9:1477–1487. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen Y, Yang Y, Tuan Z, Wang C and Shi Y:
Predicting chemosensitivity in osteosarcoma prior to chemotherapy:
An investigational study of biomarkers with immunohistochemistry.
Oncol Lett. 3:1011–16. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
El-Naggar AM, Veinotte CJ, Cheng H,
Grunewald TGP, Negri GL, Somasekharan SP, Corkery DP, Tirode F,
Mathers J, Khan D, et al: Translational activation of HIF-1α by
YB-1 promotes sarcoma metastasis. Cancer Cell. 27:682–697. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Zhang Y, Cheng H, Li W, Wu H and Yang Y:
Highly-expressed P2X7 receptor promotes growth and metastasis of
human HOS/MNNG osteosarcoma cells via PI3K/Akt/GSK3b/b-catenin and
mTOR/HIF1a/VEGF signaling. Int J Cancer. 145:1068–1082. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xu WN, Yang RZ, Zheng HL, Jiang LS and
Jiang SD: NDUFA4L2 regulated by HIF-1α promotes metastasis and
epithelial-mesenchymal transition of osteosarcoma cells through
inhibiting ROS production. Front Cell Dev Biol. 8:5150512020.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Luo P, Zhang YD, He F, Tong CJ, Liu K, Liu
H, Zhu SZ, Luo JZ and Yuan B: HIF-1α -mediated augmentation of
miRNA-18b-5p facilitates proliferation and metastasis in
osteosarcoma through attenuation PHF2. Sci Rep. 12:103982022.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Labun K, Montague TG, Krause M, Torres
Cleuren YN, Tjeldnes H and Valen E: CHOPCHOP v3: Expanding the
CRISPR web toolbox beyond genome editing. Nucleic Acids Res.
47:W171–W174. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) Method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Schneider CA, Rasband WS and Eliceiri KW:
NIH image to ImageJ: 25 Years of image analysis. Nat Methods.
9:671–675. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Bligh EG and Dyer WJ: A rapid method for
total lipid extraction and purification. Can J Biochem Physiol.
37:911–917. 1959. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Mootha VK, Lindgren CM, Eriksson KF,
Subramanian A, Sihag S, Lehar J, Puigserver P, Carlsson E,
Ridderstrale M, Laurila E, et al: PGC-1alpha-responsive genes
involved in oxidative phosphorylation are coordinately
downregulated in human diabetes. Nat Genet. 34:267–73. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Subramanian A, Tamayo P, Mootha VK,
Mukherjee S, Ebert BL, Gillette MA, Paulovich A, Pomeroy SL, Golub
TR, Lander ES and Mesirov JP: Gene set enrichment analysis: A
knowledge-based approach for interpreting genome-wide expression
profiles. Proc Natl Acad Sci USA. 102:15545–15550. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Korotkevich G, Sukhov V and Sergushichev
A: Fast gene set enrichment analysis. bioRxiv New York: 2019,
http://biorxiv.org/content/early/2016/06/20/060012
|
|
27
|
Core Team, . R: A language and environment
for statistical computing. R: Foundation for Statistical Computing;
Vienna, Austria: 2023, https://www.R-project.org/
|
|
28
|
Liberzon A, Birger C, Thorvaldsdóttir H,
Ghandi M, Mesirov JP and Tamayo P: The molecular signatures
database (MSigDB) hallmark gene set collection. Cell Syst.
1:417–425. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qi J, Nakayama K, Cardiff RD, Borowsky AD,
Kaul K, Williams R, Krajewski S, Mercola D, Carpenter PM, Bowtell D
and Ronai ZA: Siah2-dependent concerted activity of HIF and FoxA2
regulates formation of neuroendocrine phenotype and neuroendocrine
prostate tumors. Cancer Cell. 18:23–38. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Kim D, Paggi JM, Park C, Bennett C and
Salzberg SL: Graph-based genome alignment and genotyping with
HISAT2 and HISAT-genotype. Nat Biotechnol. 37:907–915. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liao Y, Smyth GK and Shi W: FeatureCounts:
An efficient general purpose program for assigning sequence reads
to genomic features. Bioinformatics. 30:923–930. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Kolde R: pheatmap: Pretty Heatmaps. R
package version 1.0.12. 2019.https://CRAN.R-project.org/package=pheatmap
|
|
33
|
Hedges LV: Distribution theory for glass's
estimator of effect size and related estimators. J Educ Stat.
6:107–128. 1981. View Article : Google Scholar
|
|
34
|
Torchiano M: effsize: Efficient Effect
Size Computation. R package version 0.8.1.
2020.doi:10.5281/zenodo.1480624.
|
|
35
|
Shimizu T, Kimura K, Sugihara E,
Yamaguchi-Iwai S, Nobusue H, Sampetrean O, Otsuki Y, Fukuchi Y,
Saitoh K, Kato K, et al: MEK inhibition preferentially suppresses
anchorage-independent growth in osteosarcoma cells and decreases
tumors in vivo. J Orthop Res. 39:2732–2743. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Cobb LM, Nolan J and Butler SA:
Distribution of pimonidazole and RSU 1069 in tumour and normal
tissues. Br J Cancer. 62:915–918. 1990. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Hoppe G, Yoon S, Gopalan B, Savage AR,
Brown R, Case K, Vasanji A, Chan ER, Silver RB and Sears JE:
Comparative systems pharmacology of HIF stabilization in the
prevention of retinopathy of prematurity. Proc Natl Acad Sci USA.
113:E2516–E2525. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Palazon A, Tyrakis PA, Macias D, Velica P,
Rundqvist H, Fitzpatrick S, Vojnovic N, Phan AT, Loman N, Hedenfalk
I, et al: An HIF-1α/VEGF-A axis in cytotoxic T cells regulates
tumor progression. Cancer Cell. 32:669–683.e5. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Cowman SJ and Koh MY: Revisiting the HIF
switch in the tumor and its immune microenvironment. Trends Cancer.
8:28–42. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Lequeux A, Noman MZ, Xiao M, Van Moer K,
Hasmim M, Benoit A, Bosseler M, Viry E, Arakelian T, Berchem G, et
al: Targeting HIF-1 alpha transcriptional activity drives cytotoxic
immune effector cells into melanoma and improves combination
immunotherapy. Oncogene. 40:4725–4735. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Salman S, Meyers DJ, Wicks EE, Lee SN,
Datan E, Thomas AM, Anders NM, Hwang Y, Lyu Y, Yang Y, et al: HIF
inhibitor 32-134D eradicates murine hepatocellular carcinoma in
combination with anti-PD1 therapy. J Clin Invest. 132:e1567742022.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Passaniti A, Kleinman HK and Martin GR:
Matrigel: History/background, uses, and future applications. J Cell
Commun Signal. 16:621–626. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Quintana E, Shackleton M, Sabel MS, Fullen
DR, Johnson TM and Morrison SJ: Efficient tumor formation by single
human melanoma cells. Nature. 456:593–598. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Pretlow TG, Delmoro CM, Dilley GG,
Spadafora CG and Pretlow TP: Transplantation of human prostatic
carcinoma into nude mice in matrigel. Cancer Res. 51:3814–3817.
1991.PubMed/NCBI
|
|
45
|
Orimo A, Gupta PB, Sgroi DC,
ArenzanaSeisdedos F, Delaunay T, Naeem R, Carey VJ, Richardson AL
and Weinberg RA: Stromal fibroblasts present in invasive human
breast carcinomas promote tumor growth and angiogenesis through
elevated SDF1/CXCL12 secretion. Cell. 121:3353482005. View Article : Google Scholar
|
|
46
|
Dey S, Murmu N, Mondal T, Saha I,
Chatterjee S, Manna R, Haldar S, Dash SK, Sarkar TR and Giri B:
Multifaceted entrancing role of glucose and its analogue,
2-deoxy-D-glucose in cancer cell proliferation, inflammation, and
virus infection. Biomed Pharmacother. 156:1138012022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Chen J, Xie J, Jiang Z, Wang B, Wang Y and
Hu X: Shikonin and its analogs inhibit cancer cell glycolysis by
targeting tumor pyruvate kinase-M2. Oncogene. 30:4297–4306. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Gao J, Liao J and Yang G: CAAX-box
protein, prenylation process and carcinogenesis. Am J Transl Res.
1:312–325. 2009.PubMed/NCBI
|
|
49
|
Hori Y, Kikuchi A, Isomura M, Katayama M,
Miura Y, Fujioka H, Kaibuchi K and Takai Y: Post-translational
modifications of the C-terminal region of the rho protein are
important for its interaction with membranes and the stimulatory
and inhibitory GDP/GTP exchange proteins. Oncogene. 6:515–522.
1991.PubMed/NCBI
|
|
50
|
Jonasch E, Donskov F, Iliopoulos O,
Rathmell WK, Narayan VK, Maughan BL, Oudard S, Else T, Maranchie
JK, Welsh SJ, et al: Belzutifan for renal cell carcinoma in von
Hippel-Lindau disease. N Engl J Med. 385:2036–2046. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Chen AH and Grana AK: Belzutifan's role in
the treatment landscape of clear cell renal cell carcinoma.
Pharmacotherapy. 45:356–366. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Courtney KD, Ma Y, Diaz de Leon A,
Christie A, Xie Z, Woolford L, Singla N, Joyce A, Hill H,
Madhuranthakam AJ, et al: HIF-2 complex dissociation, target
inhibition, and acquired resistance with PT2385, a first-in-class
HIF-2 inhibitor, in patients with clear cell renal cell carcinoma.
Clin Cancer Res. 26:793–803. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
De Conti G, Dias MH and Bernards R:
Fighting drug resistance through the targeting of drug-tolerant
persister cells. Cancers (Basel). 13:11182021. View Article : Google Scholar : PubMed/NCBI
|