Introduction
Gastric cancer is a prevalent malignancy within the
digestive system. According to data from a global burden of disease
study (1), compared with in 1990,
there were an estimated 1.23 million new cases of gastric cancer
and 954,000 gastric cancer-related deaths worldwide in 2021.
According to data from the American Cancer Society (2), there were an estimated 1.23 million
new gastric cancer cases and 954,000 gastric cancer-related deaths
in the United States in 2024. This demonstrates that gastric cancer
remains a notable global health concern.
One of the major challenges in the treatment of
gastric cancer is cisplatin resistance (3). The mechanisms that lead to drug
resistance are complex and diverse, involving multiple aspects,
such as increased drug efflux and the upregulation of DNA damage
repair capabilities. Enhanced drug efflux is mainly mediated by
upregulation or increased activity of ABC transporters, leading to
a decrease in intracellular drug concentration (4,5).
Notably, experimental findings by Chen et al (6) demonstrated that the upregulation of
NBS1 can increase the DNA damage repair capacity, consequently
enhancing chemotherapy resistance (6). Moreover, the targeting and
downregulation of checkpoint kinase 1 has been demonstrated to be
an efficacious method of reversing drug resistance in gastric
cancer cells (7). SIVA-1, a
pro-apoptotic protein that can be activated by the P53 tumor
suppressor protein (8), has been
demonstrated to be associated with the induction of apoptosis in a
variety of tumor cells, such as breast cancer (9), ovarian cancer (10) and glioblastoma (11). Previous experimental results have
demonstrated that SIVA-1 serves an important role in reversing the
sensitivity of gastric cancer-resistant cells to vincristine (VCR)
(12,13). However, the effects of SIVA-1
silencing on the drug sensitivity and biological function of
cisplatin (DDP)-resistant human gastric cancer cells (AGS/DDP)
remain to be elucidated, and the mechanisms underlying altered drug
sensitivity must be explored. In the present study, AGS/DDP cells,
in which SIVA-1 was silenced, were constructed, and the changes in
the sensitivity of the cells to drugs, and alterations in their
proliferation, migration and invasion were observed, to assess the
role of SIVA-1 in gastric cancer chemoresistance.
Materials and methods
Bioinformatics analysis
The stomach adenocarcinoma (STAD) transcriptome
dataset in The Cancer Genome Atlas (TCGA; http://portal.gdc.cancer.gov/) was acquired, which
contains data from 379 gastric cancer tissue and 34 normal gastric
tissue. The GSE186205 (14) dataset
was acquired from the Gene Expression Omnibus (https://www.ncbi.nlm.nih.gov/geo). The GSE186205
dataset contains one immortalized gastric cancer cell line
(KATOIII) and one immortalized cisplatin-resistant gastric cancer
cell line (KATO/DDP), with the experiment repeated three times. The
R 4.2.3 software (https://cran.r-project.org/bin/windows/base/old/4.2.3/)
was used for analysis. The ‘DESeq2’ package (version 1.40.2)
(15) was used to analyze the
differential genes between the tumor group and the drug-resistant
group in the GSE186205 dataset. P<0.05 and absolute
log2(FoldChange)>2 were considered to indicate differentially
expressed genes. The inter-group risk scoring of the SIVA-1 gene in
TCGA-STAD was performed using the ‘survival’ package (https://github.com/therneau/survival),
and the impact of SIVA-1 expression levels on overall patient
survival was assessed using a Cox proportional hazards regression
model. Data visualization was performed using the ‘ggrisk’ package
(https://github.com/yikeshu0611/ggrisk). The gene-drug
correlation of SIVA-1 in the GSE186205 dataset was analyzed using
the oncoPredict package (16) and a
scatter plot of the results of the Pearson correlation analysis was
generated. A comprehensive search was conducted in the STRING 12.0
database (https://cn.string-db.org/) for
SIVA-1, Bcl-2, BAX, X-linked inhibitor of apoptosis protein (XIAP),
MAPK8 and baculoviral inhibitor of apoptosis repeat-containing 5
(BIRC5), with a minimum required interaction score of 0.40. The
output of this analysis showed the six protein interactions using
Cytoscape 3.10.3 (https://cytoscape.org/), and protein-protein
interaction (PPI) networks were plotted. The MSigDB database
(http://www.gsea-msigdb.org/gsea/msigdb/index.jsp)
was used for Gene Set Enrichment Analysis (GSEA). Gene Ontology
(GO) and Kyoto Encyclopedia of Genes and Genomes (KEGG) enrichment
analyses were performed using the ‘clusterProfiler’ package
(17), with GO enrichment analysis
including biological process (BP), cellular component (CC) and
molecular function (MF) terms. The GSEA, and KEGG and GO enrichment
analyses were used to analyze the GSE186205 dataset. Functional
pathways with filter-corrected P<0.05 and q<0.05 were
considered to be differential. Subsequently, the GO enrichment data
were visualized using the ‘ggSankey’ package (https://github.com/davidsjoberg/ggsankey).
Reagents
DDP (cat. no. H20073653) and Adriamycin (ADM) were
procured from Qilu Pharmaceutical Co., Ltd. Trypsin, streptomycin,
penicillin and VCR were procured from Sigma-Aldrich; Merck KGaA.
The following antibodies were procured from Cell Signaling
Technology, Inc.: SIVA-1 (cat. no. 12532S), Bcl-2 (cat. no. 4223S),
BAX (cat. no. 2772S), MAPK8 (cat. no. 3708S), BIRC5 (cat. no.
8756S), XIAP (cat. no. 2042S) and GAPDH (cat. no. 5174). The SIVA-1
RNA interference (RNAi) recombinant lentiviral vector
[pGV358-GFP-SIVA-1-short hairpin RNA (shRNA)] and the null vector
[pGV358-GFP-negative control (NC)] were obtained from Shanghai
Jikai Gene Medical Technology Co., Ltd.
Cell culture
AGS/DDP cells (cat. no. KGG3506-1) were procured
from Nanjing KeyGen Biotech Co., Ltd., and cultured in RPMI 1640
medium (Thermo Fisher Scientific, Inc.), which was supplemented
with 10% FBS (Thermo Fisher Scientific, Inc.) and antibiotics (100
U/ml penicillin + 100 mg/ml streptomycin). The cells were cultured
at 37°C under 5% CO2 gas conditions. To maintain the
drug-resistant phenotype, AGS/DDP cells were cultivated in medium
augmented with 0.6 µg/ml DDP. The 293T cells were obtained from the
Xiangya Center Laboratory at Central South University (Changsha,
China).
Infection with the shRNA-SIVA-1
lentiviral vector and formation of stably silenced expression cell
lines
The pGV358-GFP-SIVA-1-shRNA and pGV358-GFP-NC were
separately co-transfected with pHelper 1.0 and pHelper2.0 into 293T
cells using a second-generation lentiviral vector system. The total
amount of lentiviral vector, and pHelper 1.0 and pHelper 2.0
plasmid DNA was 45 µg, with a mass ratio of 2:1.5:1. The mixture
(lentiviral vector, pHelper 1.0 and pHelper 2.0),
Lipofectamine® 3000 Transfection Reagent (cat. no.
L3000015; Invitrogen; Thermo Fisher Scientific, Inc.) and Opti-MEM™
medium (cat. no. 31985062; Gibco; Thermo Fisher Scientific, Inc.)
were added to a 6-well plate containing 293T cells, and the plate
were incubated at 37°C in a 5% CO2 incubator. A total of
6 h post-transfection, the medium was replaced with fresh DMEM
(Hyclone™; Cytiva), and the cells were cultured for an additional 4
days before viral supernatant collection. Lentiviral particles were
harvested by ultracentrifugation (53,780 × g, 2 h, 4°C). The viral
titer (Lv-SIVA-1-shRNA and Lv-NC) was measured by fluorescence
microscopy (Keyence Corporation). The titer of the concentrated
lentiviral particles (Lv-SIVA-1-shRNA and Lv-NC) was determined
using a gradient dilution method and was calculated to be
2×109 TU/ml. AGS/DDP cells were then transduced with
Lv-SIVA-1-shRNA and Lv-NC recombinant lentiviruses particles at a
multiplicity of infection of 30 for a period of 48 h at 37°C.
Stable cell line selection commenced 72 h after transduction of
AGS/DDP cells with lentiviral particles. After being treated with
0.6 µg/ml DDP, the cell lines exhibiting stable silencing of SIVA-1
were then subjected to a 4-week screening process using 5 µg/ml
puromycin for initial selection, followed by maintenance at a
concentration of 3 µg/ml puromycin. Following the completion of
this screening phase, the AGS/DDP cells were divided into the
following three distinct groups based on the outcomes obtained: i)
shRNA-SIVA-1 group; ii) shRNA-NC group; and iii) control group
(uninfected).
Cell viability assay
Following cell recovery via thawing in a 37°C water
bath and centrifugation at 100 × g for 5 min, cell digestion was
performed using 0.5% trypsin-EDTA (Thermo Fisher Scientific, Inc.)
at 37°C and was terminated with medium containing 10% FBS. The
resulting cell pellets were then resuspended in RPMI 1640 medium
and the cell concentration was adjusted to 1.5×104
cells/ml. The cells in the shRNA-SIVA-1, shRNA-NC and control
groups were inoculated in 96-well plates (~5,000 cells/well) and
were placed in a cell culture incubator set to 37°C with 5%
CO2. Following a 24-h incubation period, cell viability
was assessed after treatment with DDP (0–500 µg/ml), ADM (0–1,000
µg/ml) and VCR (0–850 µg/ml) at 37°C for 48 h. Briefly, 10 µl Cell
Counting Kit-8 reagent (Dojindo Laboratories, Inc.) was added and
the cells were incubated at 37°C for 2 h. Subsequently, the
IC50 of each group was analyzed and calculated at 450 nm
using a microplate reader (PR 3100 TSC; Bio-Rad Laboratories,
Inc.).
Colony formation assay
The colony formation assay is a reliable method for
assessing the proliferative capacity of cells. Following the
completion of cell recovery and cell digestion, cells from the
shRNA-SIVA-1, shRNA-NC and control groups were inoculated in 6-well
plates (~500 cells/well). The cells were cultured at 37°C in the
presence of DDP (0.6 µg/ml), under humidified 5% CO2 gas
conditions for 2 weeks, during which the complete medium was
changed every 3 days. Subsequently, the cells were fixed with 500
µl 4% paraformaldehyde/well at 37°C for 30 min, and stained with
500 µl 0.5% crystal violet staining solution/well at 37°C for 5 min
after washing twice with PBS. Colony formation was manually counted
under an inverted optical microscope (CKX53; Olympus Corporation)
at ×40 magnification. Colonies were defined as clusters containing
≥50 cells. The colony formation rate (%) was calculated using the
formula: (number of colonies formed/total number of cells
inoculated) ×100.
Cell apoptosis assays
To detect the effects of silencing SIVA-1 on the
apoptosis of AGS/DDP cells, each group of cells was treated with 20
µg/ml DDP. To illustrate more clearly the changes in apoptosis
caused by variations in cisplatin sensitivity among the groups, a
concentration of 20 µg/ml (close to the IC50
concentration) was selected for this assay. The Annexin V-PE/7-AAD
Apoptosis Kit (cat. no. AP-105; Hangzhou Lianke Biotechnology Co.,
Ltd.) was then utilized in accordance with the manufacturer's
guidelines. The cells were incubated with 1X Binding Buffer
containing Annexin V-PE and 7-AAD for 15 min at 25°C in the dark.
Subsequently, the stained cells were detected by flow cytometry
using a FACScan instrument (Agilent Technologies, Inc.) and data
analysis was performed using MultiCycle for Windows (version AV;
Phoenix Flow Systems).
Cell scratch assay
The cell scratch assay is a well-established in
vitro experiment that models the process of cell migration.
Following completion of the cell recovery and cell digestion
procedures, cell growth was observed continuously on a daily basis.
The cells were seeded in 12-well plates until they reached 90–95%
confluence, after which, a sterile pipette tip was used to create a
linear wound in the center of the confluent cells, and the cells
were rinsed twice with PBS to remove cell debris. Subsequently, the
cells were cultured in serum-free medium. The healing process was
observed under ×40 magnification using an inverted optical
microscope (CKX53; Olympus Corporation) at 0, 24 and 48 h,
respectively. The area of the scratches was measured using the
microscope companion software (Confluency Checker Software; version
1.2; Olympus Corporation). Wound healing (%) was calculated as
follows: (0-h scratch area-48-h scratch area)/0-h scratch area
×100.
Transwell assay
The capacity of cells to migrate or invade was
evaluated by employing a 24-well Transwell system (8.0 µm pore
polycarbonate membrane; cat. no. 3422; Corning Inc.), which served
to replicate the physiological conditions under which cells
traverse in vivo tissues. Initially, 500 µl complete medium
containing 10% FBS was added to the lower chamber of a 24-well
plate, and a Transwell chamber was placed above it. For the
invasion assay, 100 µl diluted Matrigel was added and cured at 37°C
for 1 h to form a gel matrix. Following the completion of cell
recovery and cell digestion, the cells in the shRNA-SIVA-1,
shRNA-NC and control groups were inoculated in the upper chamber of
the 24-well plate (~5×104 cells/well), and the plate was
returned to the incubator to continue cultivation. After 24 h, the
Transwell chamber was fixed in 400 µl 4% paraformaldehyde at room
temperature for 10 min, washed once with PBS and stained with 400
µl crystal violet at room temperature for 5 min. Subsequently, the
chambers were washed three times, both internally and externally,
with PBS. The number of visible cells that had migrated/invaded the
membrane was subsequently observed under an inverted optical
microscope (CKX53; Olympus Corporation) at ×40 magnification. For
the cell migration assay, the aforementioned protocol was repeated
in the absence of Matrigel.
Reverse transcription-quantitative
PCR
After completion of cell recovery and cell
digestion, the cell concentration was adjusted to
1.5×104 cells/ml. Total RNA was extracted from cells in
all groups (shRNA-SIVA-1, shRNA-NC and control) using Triquick
Reagent (cat. no. R1100; Beijing Solarbio Science & Technology
Co., Ltd.) according to the manufacturer's instructions.
Specifically, RNA Extraction Reagent (cat. no. P1011; Beijing
Solarbio Science & Technology Co., Ltd.) was added during the
phase separation step. The extracted RNA was then reverse
transcribed into cDNA using the Reverse Transcription Kit (cat. no.
MR05101M; Monad Biotech Co., Ltd.) according to the manufacturer's
protocol. qPCR was performed using the 2X S6 Universal SYBR qPCR
Mix [cat. no. Q204-01; XinBei Biotechnology (Suzhou) Co., Ltd.] on
a Real-Time PCR System (MA-6000; Molarray Research, Inc.) under the
following thermocycling conditions: Initial denaturation at 95°C
for 30 sec, followed by 40 cycles of denaturation at 95°C for 3–10
sec and annealing/extension at 60°C for 10–30 sec. The primers for
PCR are shown in Table I. Relative
gene expression levels were calculated using the 2−ΔΔCq
method (18) with GAPDH as the
internal reference gene. The experiment was repeated three times,
with the mean value taken as the final result.
 | Table I.Primer sequences used for reverse
transcription-quantitative PCR. |
Table I.
Primer sequences used for reverse
transcription-quantitative PCR.
| Gene | Base sequence,
5′-3′ |
|---|
| SIVA-1 | F:
GTCGTTCACCTGCCAGAGTC |
|
| R:
CCGTCTGGTCCAATCAGCAT |
| MAPK8 | F:
CTTGGCATGGGCTACAAGGA |
|
| R:
GGTGGGGACATAAGGTGGTG |
| Bcl-2 | F:
GTGTGTGGAGAGCGTCAACC |
|
| R:
ACAGTTCCACAAAGGCATCC |
|
| CAG |
| BAX | F:
AGTAACATGGAGCTGCAGAGGA |
|
| TGA |
|
| R:
TGGAGACAGGGACATCAGTCG |
| XIAP | F:
ATTTCCAGATTGGGGCTCGG |
|
| R:
CTTGTCCACCTTTTCGCGCC |
| BIRC5 | F:
CCACTGAGAACGAGCCAGACTT |
|
| R:
GTATTACAGGCGTAAGCCACCG |
| GAPDH | F:
TGACGTGGACATCCGCAAAG |
|
| R:
CTGGAAGGTGGACAGCGAGG |
Western blotting
After cell recovery and digestion, total proteins
were extracted from whole cell lysates using RIPA Lysis and
Extraction Buffer (cat. no. 89901; Thermo Fisher Scientific, Inc.).
The concentration of the extracted proteins was measured using the
BCA Protein Quantification Kit (Thermo Fisher Scientific, Inc.).
The absorbance value of each sample was measured at 562 nm using a
microplate reader and the concentration was calculated from the
standard curve. Total proteins (20 µg) were collected for SDS-PAGE
on 10% gels and were transferred onto a PVDF membrane, before being
incubated with QuickBlock™ Blocking Buffer (cat. no. P0235;
Beyotime Biotechnology) for 1 h at room temperature with agitation.
The following primary antibodies were then added and incubated
overnight at 4°C with agitation: Anti-SIVA-1 (1:500), anti-Bcl-2
(1:500), anti-BAX (1:500), anti-MAPK8 (1:1,000), anti-BIRC5
(1:500), anti-XIAP (1:1,000) and anti-GAPDH (1:1,000).
Subsequently, Goat Anti-Rabbit IgG H&L (HRP) (cat. no. ab6721;
Abcam) was added and incubated for 1 h at room temperature in the
dark, and washed three times with 0.1% TBS Tween-20 Buffer (cat.
no. 28360; Thermo Fisher Scientific, Inc.). A gel imager (iBright
CL750; Invitrogen; Thermo Fisher Scientific, Inc.) was used to scan
the protein blotting membrane, and the gray scale values of the
bands were analyzed and recorded using ImageJ software (bundled
with 64-bit Java 8; National Institutes of Health). The relative
expression levels of the target proteins were expressed as the
ratio of the integrated optical densities of the target protein
bands and the GAPDH bands.
Establishment of a subcutaneous tumor
xenograft model in nude mice
Ethics approval for the animal experiments in the
present study was obtained from The Medical Ethics Committee of
People's Hospital of Guangxi Zhuang Autonomous Region (approval no.
KY-GZR-2016-416; Nanning, China). A total of 30 5-week-old male
BALB/c nude mice (weight, 16–18 g), were purchased from Beijing
Vital River Laboratory Animal Technology Co., Ltd., and maintained
at the Laboratory Animal Center of Guangxi Medical University
(Nanning, China). The housing environment was maintained at a
temperature of 22±2°C and a relative humidity of 55±10%, with a
12-h light/dark cycle. Autoclaved standard rodent diet and sterile
drinking water were provided ad libitum. The nude mice were
subcutaneously inoculated with AGS/DDP cells (2×106
cells in 100 µl suspension/mouse) into the left anterior abdomen.
The health status and behavior of the nude mice were monitored
daily. According to the animal operational procedures established
by the Animal Ethics Committee of the People's Hospital of Guangxi
Zhuang Autonomous Region, a standardized management plan was put in
place to deal with any in-fighting as followed: i) The immediate
separation of involved animals and tiered intervention based on the
severity of the wounds; ii) superficial wounds were treated with
topical cleansing and antisepsis; iii) severe trauma cases were
assessed by a veterinarian to determine the appropriateness of
euthanasia; and iv) animals were humanely euthanized once they
reached predefined humane endpoints (19–22).
The presence of a tumor measuring ≥1.5 cm in any direction,
accompanied by body weight loss of >20% of the initial weight,
or 10% weight loss accompanied by severe clinical signs of distress
(such as hypotrichosis, loss of appetite or reduced vitality), were
considered humane endpoints. If mice reached these endpoints they
were euthanized (23). When the
resulting tumor diameter was measured to be 5 mm, the mice were
randomly divided into the following three groups (n=10/group): i)
shRNA-SIVA-1 group; ii) shRNA-NC group; and iii) control group.
Nude mice in the shRNA-SIVA-1 group or the shRNA-NC group were
injected intratumorally with 100 µl Lv-SIVA-1-shRNA or Lv-NC
recombinant lentivirus particles every 2 days. Each injection
consisted of 100 µl lentiviral suspension with a titer of
2×109 TU/ml. When the subcutaneous xenografts were
visible, the length and width of the tumor were measured every 2
days. Tumor volume was calculated using the following formula:
Length × width2 ×0.5 cm3. The relative tumor
volume (RTV) was calculated on day 13 using the following formula:
Tumor volume on day 13/tumor volume before administration. DDP was
injected intraperitoneally at a dose of 3 mg/kg every 2 days. A
total of 13 days after tumor injections, the mice were sacrificed
by cervical dislocation. Death was confirmed by the absence of
vital signs (respiratory arrest, loss of corneal reflex and no
response to a toe pinch), the xenografts were dissected and
harvested for further analysis. Only mice that survived until the
end of the experiment were included in the tumor growth and
parameter analysis.
Hematoxylin and eosin (H&E)
staining and immunohistochemical (IHC) staining
The obtained pathological specimens were immersed in
10% neutral buffered formalin solution for fixation at room
temperature (~22°C) for 48 h, dehydrated through increasing ethanol
series (75, 85, 95, 100 and 100%) and embedded in paraffin. After
sectioning, deparaffinization and rehydration, the specimens were
stained with 0.5% hematoxylin for 3–6 min and then with 0.5% eosin
Y for 2–3 min, both at room temperature. Subsequently, the stained
sections were dehydrated through a graded ethanol series, cleared
in xylene and mounted with a neutral resin.
For IHC staining, the paraffin-embedded tumor
sections were first deparaffinized in xylene (cat. no. 10023418;
Sinopharm Chemical Reagent Co., Ltd.) and then rehydrated in a
descending ethanol series (100, 100, 95, 85 and 75%) for 5 min
each. Antigen retrieval was performed using heat-induced epitope
retrieval at a pressure of 103 kPa and a temperature of 125°C for
23 min. After a 10-min treatment with 3% hydrogen peroxide (cat.
no. 10011218; Sinopharm Chemical Reagent Co., Ltd.) at room
temperature, the cells were incubated in a humidified chamber for
30 min to eliminate endogenous peroxidase activity. The cells were
then blocked with 10% goat serum (cat. no. SL038-10 ml; Beijing
Solarbio Science & Technology Co., Ltd.) at room temperature
for 30 min. The sections were incubated overnight at 4°C with
anti-SIVA-1 antibody (1:200; cat. no. PA5-100737; Thermo Fisher
Scientific, Inc.). After washing with PBS, the slides were
incubated with the ready-to-use HRP-Polymer Anti-Mouse/Rabbit
reagent (cat. no. Kit-5020; MaxVision Biosciences) at 37°C for 30
min according to the manufacturer's instructions. Detection was
performed using a DAB substrate kit (cat. no. DA1010-2*3ml; Beijing
Solarbio Science & Technology Co., Ltd.), after which the
sections were counterstained with hematoxylin (cat. no. G1140;
Beijing Solarbio Science & Technology Co., Ltd.). Subsequently,
the sections were dehydrated in an increasing ethanol series (75,
85, 95, 100 and 100%) for 5 min each, and xylene was added for
permeabilization (repeated twice, 3 min each time). The slides were
then mounted using neutral resin and imaging was carried out using
an optical microscope. The immunostaining scoring of each
pathological section was performed by two independent pathologists,
and IHC staining results were semi-quantified using the QuickScore
scoring method (24). The IHC
staining score consisted of the intensity of cell staining and the
percentage of positively stained cells, where the intensity of cell
staining was scored as follows: 0 (no staining), 1+ (weak
staining), 2+ (moderate staining) and 3+ (strong staining). The
percentage of positively stained cells was evaluated according to
the following criteria: 0 (0%), 1 (<10%), 2 (10–25%), 3
(26–50%), 4 (51–75%) and 5 (>75%). The IHC staining score was
calculated as the intensity of cell staining × the percentage of
positively stained cells [score of 0–3 (−), score of 4–6 (+), score
of 7–9 (++) and score of 10–12 (++++)].
Statistical analysis
R 4.2.3 (https://cran.r-project.org/bin/windows/base/old/4.2.3/)
and SPSS 23.0 (IBM, Corp.) statistical software were used for
analysis. Each experiment was independently repeated three times.
The data from the IHC staining scores are presented as the median
with the interquartile range, whereas continuous data are presented
as the mean ± standard deviation. Differences between the groups
were compared using one-way ANOVA, followed by Tukey's post hoc
test to facilitate multiple comparisons. As the data from the IHC
staining scores did not meet the key assumptions required for
parametric tests (normal distribution and homogeneity of variance),
the Kruskal-Wallis test was performed, followed by Dunn post hoc
test for pairwise comparisons. P<0.05 was considered to indicate
a statistically significant difference.
Results
Investigation of the expression
profile of SIVA-1 and its relationship with drug sensitivity
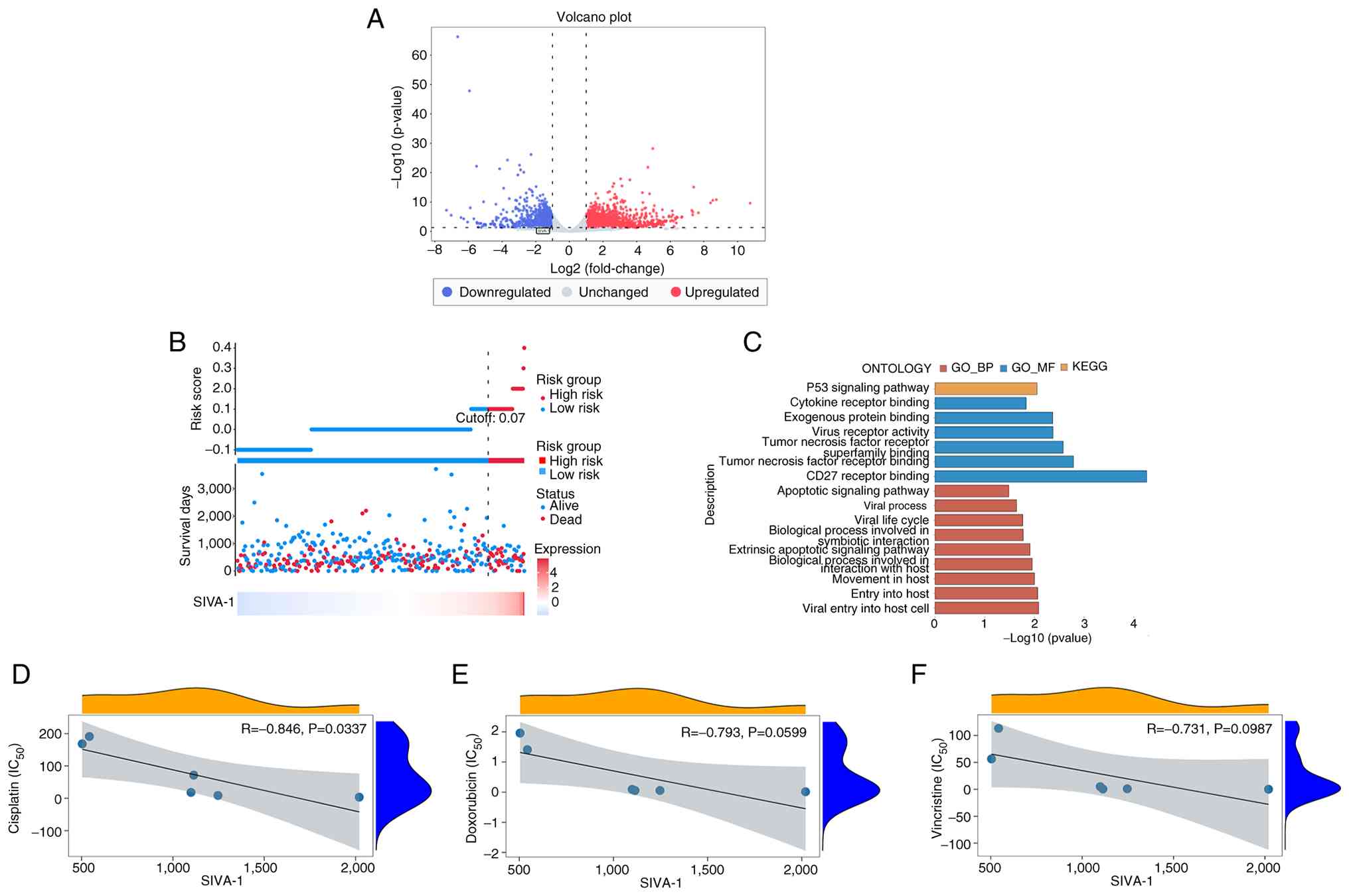
First, differential gene analysis was performed
using the GSE186205 dataset, and the results are shown in a volcano
plot (Fig. 1A). SIVA-1 was
differentially expressed in cisplatin-resistant gastric cancer cell
lines, where its expression was lower compared with that in
cisplatin-sensitive gastric cancer cell lines. The association
between SIVA-1 and survival status was explored, and data
visualization was performed by plotting risk factor associations
using TCGA-STAD data. The results showed that when SIVA-1
expression was high, the risk score for a positive event (death)
was high (risk score cut-off, >0.07). Specifically, the risk of
death was 1.01 times higher in the high-expression group than in
the low-expression group. (Fig.
1B). Therefore, high SIVA-1 expression could be considered a
high-risk factor. KEGG enrichment analysis revealed that SIVA-1 was
associated with the ‘p53 signaling pathway’ (Fig. 1C). In addition, GO enrichment
analysis revealed that SIVA-1 was involved in the BP terms of
‘tumor necrosis factor receptor binding’ and ‘CD27 receptor
binding’, among others, the MF terms of ‘apoptotic signaling
pathway’ and ‘extrinsic apoptotic signaling pathway’, as well as
other functions. The results of the drug sensitivity analysis using
the GSE186205 dataset are shown in Fig.
1D-F. SIVA-1 expression was negatively correlated with the
IC50 value of DDP (R=−0.846, P<0.05), whereas
statistically significant correlations were not observed with ADM
or VCR (P>0.05). This indicated that when SIVA-1 expression was
lower, the resistant group was less sensitive to DDP.
Obtaining and measuring the titers of
shRNA-SIVA-1 and shRNA-NC lentiviral particles
The titer of shRNA-SIVA-1 and shRNA-NC lentiviral
particles was measured by gradient dilution method to be
2×109 TU/ml. Fluorescence-labeled lentiviruses were also
observed by fluorescence microscopy and the viral titer was
calculated using the following formula: Viral titer=number of
fluorescent cells/volume of viral stock solution (Fig. 2A).
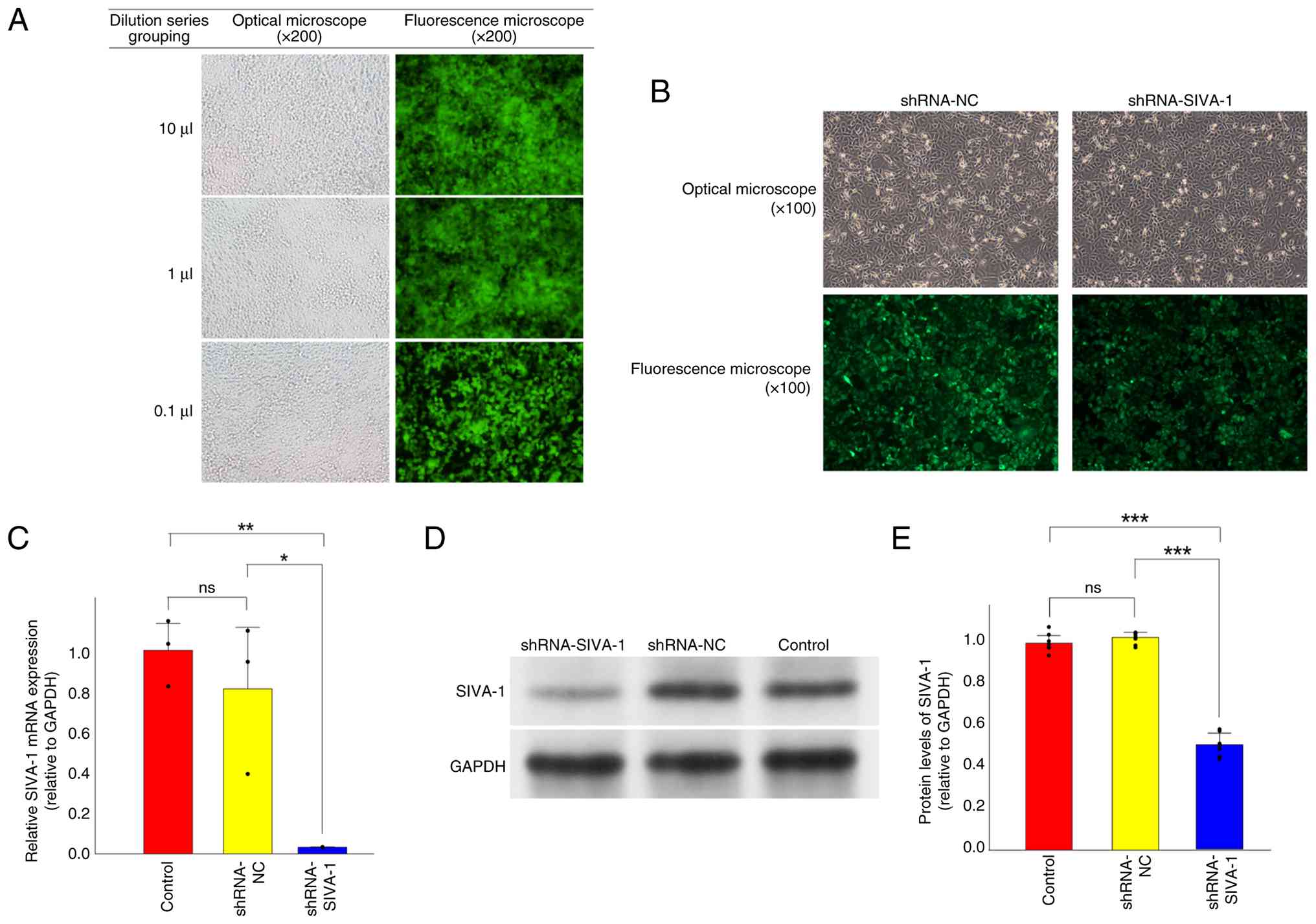 | Figure 2.Generation and validation of stable
SIVA-1-silenced AGS/DDP cells. (A) Infection efficiency of
lentiviral plasmids at gradient dilution concentrations of 10, 1
and 0.1 µl is represented by GFP fluorescence intensity (original
magnification, ×200). (B) Effects of lentiviral infection on
AGS/DDP cells (original magnification, ×100). (C) mRNA expression
levels of SIVA-1 in cells from each group, with GAPDH used as an
internal reference. (D) Western blot analysis was performed to
measure the protein expression levels of SIVA-1 in each group of
cells. (E) Semi-quantification of SIVA-1 protein expression levels
in cells from each group, with GAPDH used as an internal reference.
The results indicate the ratio of the optical density of SIVA-1
bands to the optical density of GAPDH bands. *P<0.05,
**P<0.01, ***P<0.001, as determined by ANOVA tests. DDP,
cisplatin; NC, negative control; ns, not significant; shRNA, short
hairpin RNA. |
Infection of drug-resistant gastric
cancer cells with shRNAs and screening of cell lines with stable
SIVA-1 knockdown
The shRNA-SIVA-1 and shRNA-NC lentiviral particles
contain the GFP reporter gene. They were infected into AGS/DDP
cells to establish the shRNA-SIVA-1, shRNA-NC and control groups.
The stable SIVA-1 knockdown cell line was then established from the
GFP-positive cells from the shRNA-SIVA-1 group (Fig. 2B).
Expression of SIVA-1 in AGS/DDP
cells
To assess the effects of the shRNA-SIVA-1 lentivirus
on SIVA-1 expression following infection into AGS/DDP cells, the
relative mRNA expression levels of SIVA-1 in each group of cells
were measured by reverse transcription-quantitative PCR. The
results showed that the mRNA levels of SIVA-1 were significantly
decreased in the shRNA-SIVA-1 group compared with those in the
shRNA-NC and control groups (P<0.05; Fig. 2C). These findings indicated that the
shRNA-SIVA-1 lentivirus downregulated the mRNA expression levels of
SIVA-1 in AGS/DDP cells. In addition, the protein expression levels
of SIVA-1 were determined by western blotting. The protein levels
of SIVA-1 were also significantly decreased in the shRNA-SIVA-1
group compared with in the shRNA-NC and control groups (P<0.05;
Fig. 2D and E).
Silencing of SIVA-1 results in
increased resistance to DDP in AGS/DDP cells
To investigate the effect of silencing SIVA-1 on
drug resistance in resistant gastric cancer cells, the
IC50 values of ADM, DDP and VCR in each group of cells
was measured by performing at least three independent cell
viability assays. The results showed that the IC50 value
(µg/ml) of DDP in the shRNA-SIVA-1 group (21.43±0.52) was
significantly increased compared with that in the control
(17.49±0.30) and shRNA-NC (17.33±0.39) groups (P<0.05) (Table II). By contrast, the differences in
the IC50 values of ADM and VCR were not significant
between the cells in each group (P>0.05). These findings
indicated that SIVA-1 knockdown could enhance the resistance
AGS/DDP gastric cancer cells to DDP.
 | Table II.Effect of SIVA-1 silencing on DDP,
ADM and VCR resistance in AGS/DDP cells. |
Table II.
Effect of SIVA-1 silencing on DDP,
ADM and VCR resistance in AGS/DDP cells.
|
| IC50
values |
|---|
|
|
|
|---|
| Drug | shRNA-SIVA-1 | shRNA-NC | Control |
|---|
| DDP, µg/ml |
21.43±0.52a |
17.33±0.39a |
17.49±0.30a |
| VCR, µg/ml | 318.89±9.12 | 315.52±6.49 | 321.67±9.10 |
| ADM, µg/ml | 4.49±1.12 | 4.68±0.75 | 3.27±0.63 |
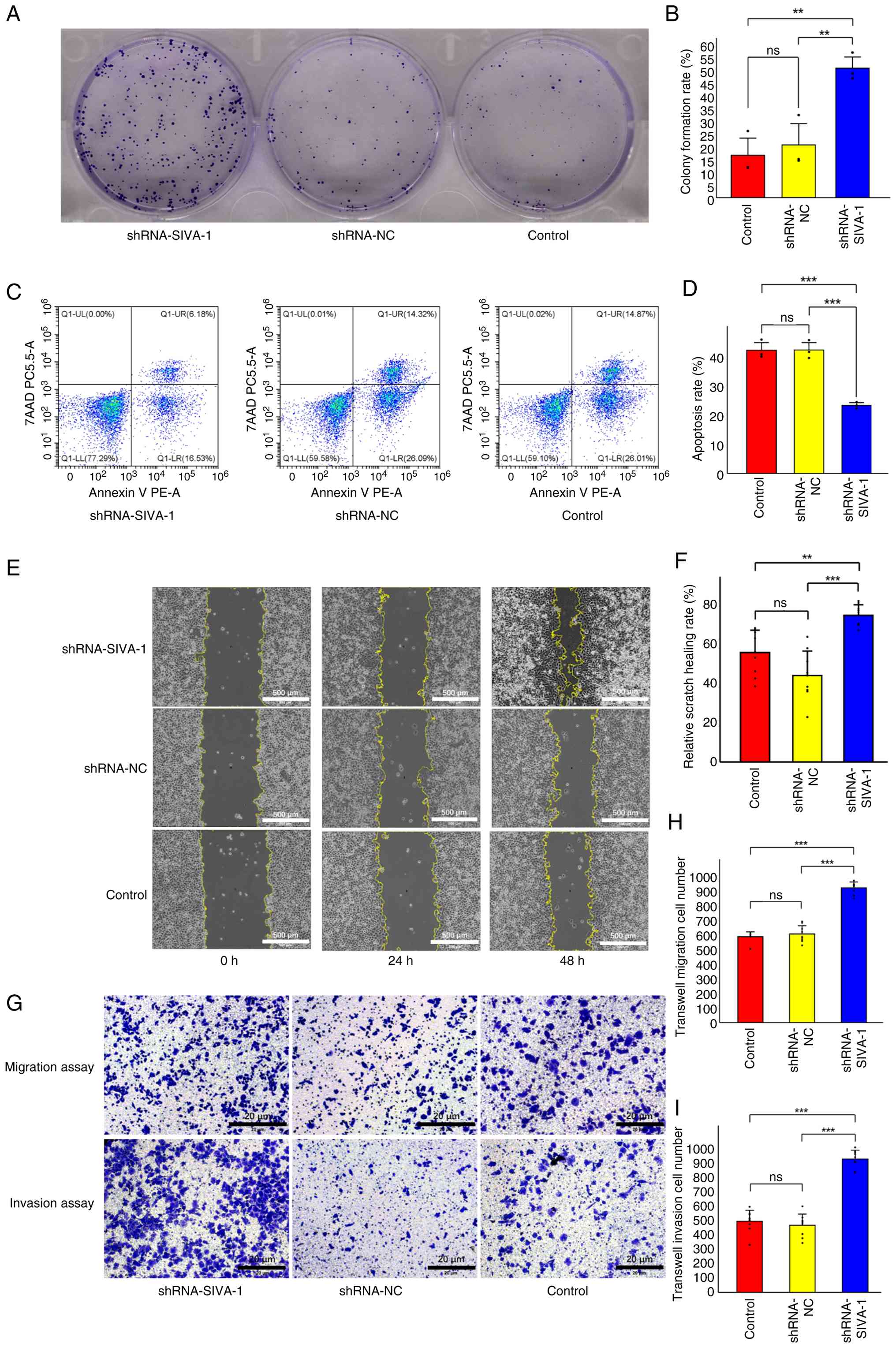
Silencing of SIVA-1 promotes the
proliferation of AGS/DDP cells
To investigate the role of SIVA-1 in the
proliferation of AGS/DDP gastric cancer cells, a colony formation
assay was performed (Fig. 3A and
B). Compared with in the control (16.96±6.87%) and shRNA-NC
(21.09±8.47%) groups, the colony formation rate was significantly
higher in the shRNA-SIVA-1 group (51.84±4.51%) (P<0.05). These
results indicated that SIVA-1 silencing could promote the
proliferative function of DDP-resistant gastric cancer cells.
Silencing SIVA-1 induces apoptosis in
AGS/DDP cells
The effects of silencing SIVA-1 on apoptosis in
AGS/DDP cells were explored through at least three independent
replicate experiments (Fig. 3C and
D). The results demonstrated that the apoptosis rate in the
shRNA-SIVA-1 group (23.84±0.94%) was significantly lower than that
in the shRNA-NC (43.21±2.60%) and control (43.15±2.65%) groups
(P<0.05). By contrast, no significant differences were observed
between the shRNA-NC and control groups.
Silencing of SIVA-1 enhances the
migration and invasion of AGS/DDP cells
Firstly, the effect of SIVA-1 silencing on the
migration of AGS/DDP cells was assessed by scratch assay (Fig. 3E and F). According to the healing
rate results at 48 h, the scratch healing rate of the shRNA-SIVA-1
group (74.54±5.08%) was significantly increased compared with that
in the control (55.61±10.70%) and shRNA-NC (43.94±11.69%) groups
(P<0.05). These findings suggested that silencing SIVA-1 could
improve the mobility of DDP-resistant gastric cancer cells. In
addition, using the Transwell assay, the effect of SIVA-1 silencing
on the migration and invasion of AGS/DDP cells was investigated. As
shown in Fig. 3G and H, the number
of migrating cells was significantly increased in the shRNA-SIVA-1
group (931.78±38.28) relative to the shRNA-NC (612.56±53.92) and
control (594.22±30.98) groups (P<0.05). The results of the
invasion assay, shown in Fig. 3G and
I, showed that the number of invasive cells in the shRNA-SIVA-1
group (928.78±57.93) was significantly higher compared with that in
the shRNA-NC (467.00±74.15) and control (495.56±71.28) groups
(P<0.05).
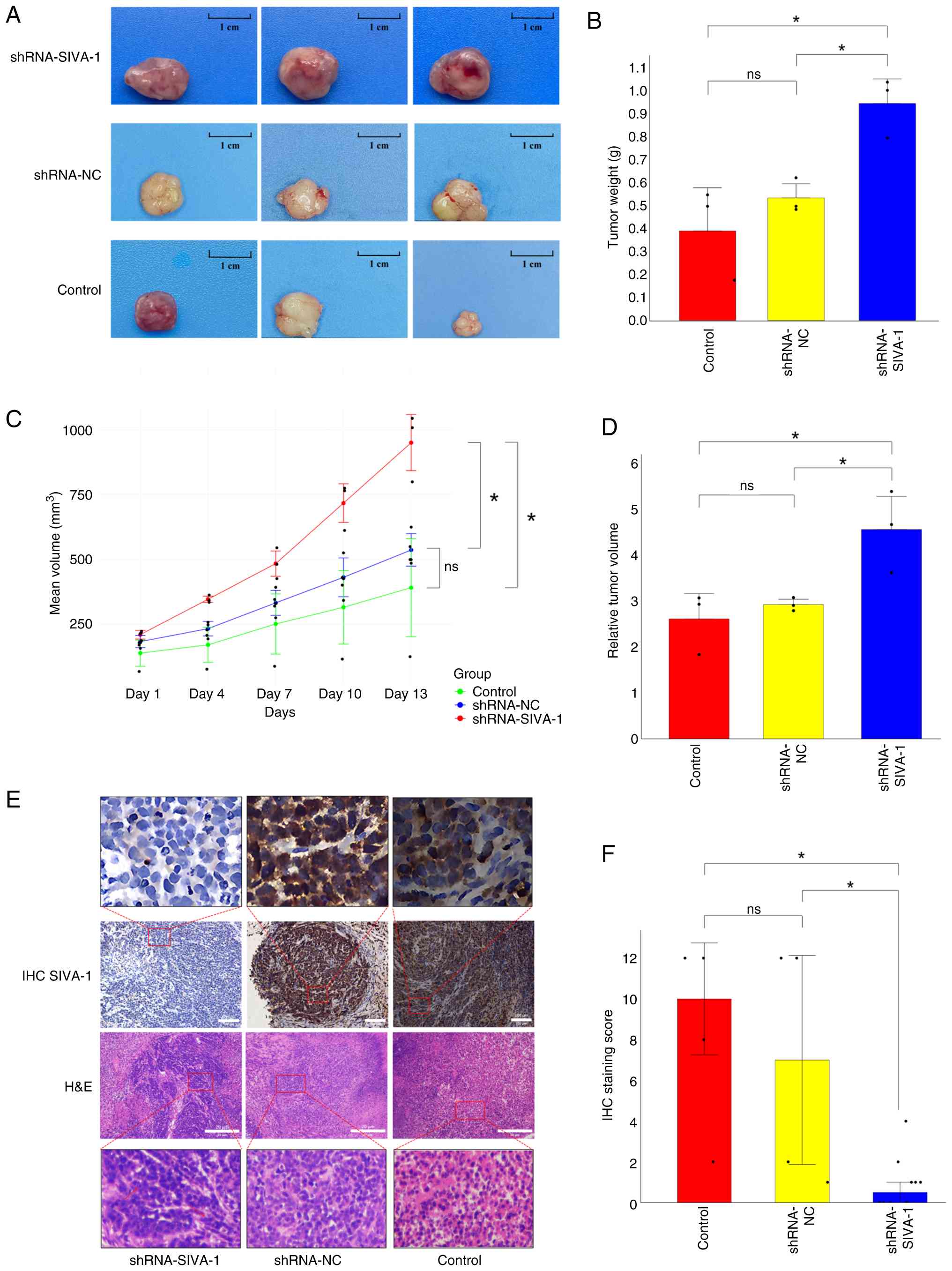
SIVA-1 silencing increases tumor
burden in vivo
To further validate the biological role of SIVA-1
silencing in the gastric cancer environment in vivo,
subcutaneous tumor loading experiments were performed in nude mice.
The changes in the weight of the individual nude mice and the
volume of the subcutaneous tumors in each group were observed. In
the final analysis, only three nude mice survived in each group,
with some mice reaching pre-defined humane endpoints (due to severe
injuries from fighting or complications of DDP chemotherapy) and
were therefore euthanized. Throughout the entire experimental
period, a total of four nude mice died as a result of in-fighting;
among these, one was from the shRNA-SIVA-1 group, one from the
shRNA-NC group and two from the control group. The results showed
that the weight of tumors in the shRNA-SIVA-1 group (0.95±0.11 g)
was significantly increased compared with that in the shRNA-NC
(0.54±0.06 g) and control (0.39±0.19 g) groups (P<0.05; Fig. 4A and B). The growth rate of
subcutaneous xenograft tumors in the shRNA-SIVA-1 group was
significantly faster than that in the shRNA-NC and control groups
(P<0.05; Fig. 4C). Fig. 4D shows the RTV on day 13. The RTV
value of the shRNA-SIVA-1 group (4.55±0.73) was significantly
higher than that in the shRNA-NC (2.92±0.12) and control
(2.61±0.55) groups (P<0.05). Cell staining results are shown in
Fig. 4E. Observing the
paraffin-embedded sections of subcutaneous xenograft tumors in nude
mice after H&E staining, the shRNA-SIVA-1 group showed abnormal
cellular morphology with uneven nuclei sizes, and nuclear
pleomorphism was observed in some areas. In addition, the results
of the IHC staining showed that the expression of SIVA-1 in the
subcutaneous metastases of nude mice in the shRNA-SIVA-1 group was
lower than that in the shRNA-NC and control groups (P<0.05;
Fig. 4F), indicating that SIVA-1
was silenced in the subcutaneous metastases of the shRNA-SIVA-1
group.
Screening of Bcl-2, BAX, XIAP, MAPK8
and BIRC5 as interaction genes of SIVA-1
To explore the genes associated with SIVA-1, GSEA
was performed on the GSE186205 dataset. The two signaling pathways
‘MicroRNAs in cancer’ and ‘Chemical carcinogenesis-receptor
activation’ were differentially expressed in cisplatin-resistant
gastric cancer cell lines compared with those in
cisplatin-sensitive gastric cancer cell lines in KEGG enrichment
analyses (Fig. 5A). In GO
enrichment analyses, ‘mitochondrial protein-containing complex’,
‘protein localization to mitochondrion’, ‘apoptotic mitochondrial
changes’, ‘regulation of release of cytochrome c from
mitochondria’, ‘metaphase/anaphase transition of cell cycle’ and
‘regulation of endopeptidase activity’ were differentially
expressed in drug-resistant tissues (Fig. 5B). BAX was enriched in the following
terms: ‘mitochondrial protein-containing complex’, ‘protein
localization to mitochondrion’, ‘apoptotic mitochondrial changes’
and ‘regulation of release of cytochrome c from mitochondria’. In
addition, BIRC5 has the biological function of ‘metaphase/anaphase
transition of cell cycle’ and ‘regulation of endopeptidase
activity’. In a previous study, experimental results showed that
SIVA-1 affects the expression of Bcl-2 (25). Furthermore, the data in the previous
study suggested that SIVA-1 influences apoptosis by interacting
with XIAP and concomitantly regulating MAPK8 (26). Therefore, based on the GSEA, a GO
enrichment analysis of Bcl-2, BAX, XIAP, MAPK8 and BIRC5 was
performed to investigate the functions of these five genes. The
results are shown in Fig. 5C; the
GO enrichment analysis revealed that Bcl-2, BAX, XIAP, MAPK8 and
BIRC5 were primarily enriched in the BP terms ‘regulation of
mitochondrial outer membrane permeabilization involved in apoptotic
signaling pathway’, ‘regulation of intrinsic apoptotic signaling
pathway in response to DNA damage by p53 class mediator’,
‘regulation of protein insertion into mitochondrial membrane
involved in apoptotic signaling pathway’ and ‘intrinsic apoptotic
signaling pathway in response to DNA damage’. The CC term ‘Bcl-2
family protein complex’ and ‘mitochondrial outer membrane’
indicated that Bcl-2 may form a protein complex with BAX and is
located on the outer membrane of the mitochondria. The MF terms,
particularly those related to protein binding (‘BH3 domain binding’
and ‘death domain binding’), receptor binding (‘CD27 receptor
binding’ and ‘tumor necrosis factor receptor binding’) and enzyme
regulator activity (‘cysteine-type endopeptidase inhibitor activity
involved in apoptotic processes’), showed that the aforementioned
five genes had the ability to regulate enzyme activities or bind to
receptors. The gene expression correlation heatmap showed that
SIVA-1 was correlated with MAPK8 (r=0.1, P<0.05), MAPK8 was
correlated with Bcl-2 (r=0.2, P<0.05), Bcl-2 was correlated with
XIAP (r=0.2, P<0.05) and with BIRC5 (r=−0.2, P<0.05), and BAX
was correlated with XIAP (r=0.1, P<0.05) and BIRC5 (r=0.3,
P<0.05), although these correlations were very weak (Fig. 5D). PPI networks (Fig. 5E), along with its visual legend
(Fig. 5F), showed that SIVA1 acts
as an upstream gene linked to MAPK8 by regulating MDM2, which
indirectly regulates Bcl-2 and BAX, and ultimately affects XIAP and
BIRC5.
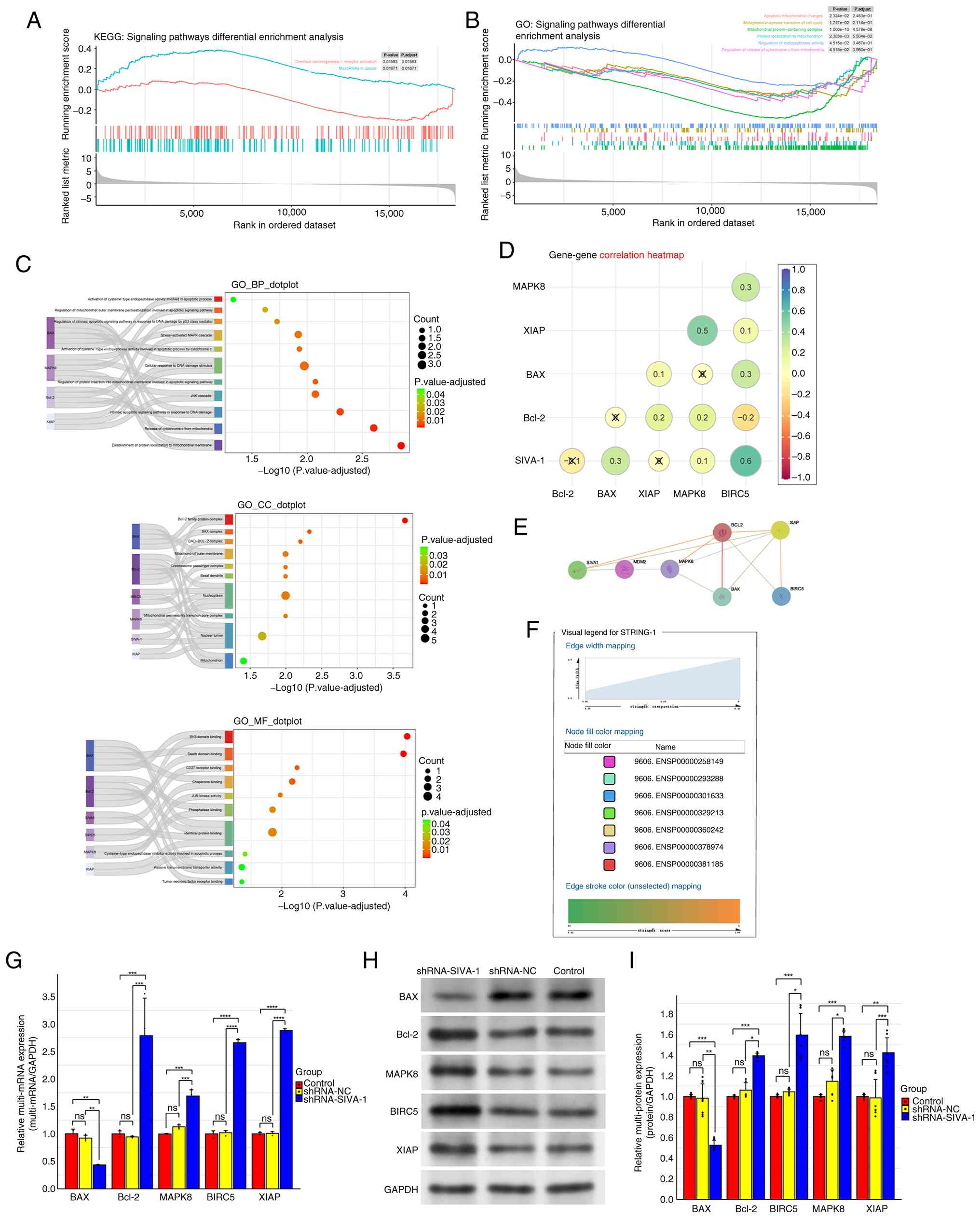 | Figure 5.Analysis and validation of SIVA-1
interaction with Bcl-2, BAX, XIAP, MAPK8 and BIRC5, and related
signaling pathways. (A) GSEA of KEGG pathways in the GSE186205
dataset. (B) GSEA of GO terms in the GSE186205 dataset. (C) BP, CC
and MF terms associated with SIVA-1, Bcl-2, BAX, XIAP, MAPK8 and
BIRC5, as determined by GO enrichment analysis in the GSE186205
dataset. (D) Gene expression correlation heatmap of SIVA-1, Bcl-2,
BAX, XIAP, MAPK8 and BIRC5 in The Cancer Genome Atlas-stomach
adenocarcinoma dataset. ‘X’ indicates P≥0.05. (E) Interactions
between SIVA-1, and Bcl-2, BAX, XIAP, MAPK8 and BIRC5 proteins in
the STRING 12.0 database. (F) Visual legend of the protein-protein
interaction network. (G) mRNA expression levels of Bcl-2, BAX,
XIAP, MAPK8 and BIRC5 in each group of cells, as detected by
reverse transcription-quantitative PCR after SIVA-1 silencing. (H)
Protein expression levels of Bcl-2, BAX, XIAP, MAPK8 and BIRC5, as
detected by western blotting after SIVA-1 silencing in each group
of cells. (I) Semi-quantification of the protein expression levels
of Bcl-2, BAX, XIAP, MAPK8 and BIRC5 in different groups of cells
(using GAPDH as an internal reference). *P<0.05, **P<0.01,
***P<0.001, ****P<0.0001, as determined by ANOVA tests. BP,
biological process; CC, cellular component; BIRC5, baculoviral
inhibitor of apoptosis repeat-containing 5; GO, Gene Ontology;
GSEA, Gene Set-Enrichment Analysis; KEGG, Kyoto Encyclopedia of
Genes and Genomes; MF, molecular function; ns, not significant;
XIAP, X-linked inhibitor of apoptosis protein. |
Silencing of SIVA-1 upregulates the
expression of Bcl-2, BIRC5, XIAP and MAPK8, and downregulates the
expression of BAX
Combined with the protein interaction results of
bioinformatics analysis, the mechanism by which SIVA-1 knockdown
led to enhanced drug resistance in AGS/DDP cells was investigated
by detecting the expression levels of apoptosis-related genes and
genes of apoptosis-related signaling pathways (Bcl-2, BAX, XIAP,
MAPK8 and BIRC5). The mRNA expression levels of these genes were
detected by reverse transcription-quantitative PCR. The results
showed that compared with those in the control and shRNA-NC groups,
the mRNA expression levels of Bcl-2, BIRC5, XIAP and MAPK8 were
increased, whereas the mRNA expression levels of BAX were decreased
in the shRNA-SIVA-1 group (P<0.05; Fig. 5G). The expression levels of the
aforementioned apoptosis-related proteins were detected by western
blotting. The results were similar to those of the reverse
transcription-quantitative PCR, with the expression levels of
Bcl-2, BIRC5, XIAP and MAPK8 proteins increased, and BAX protein
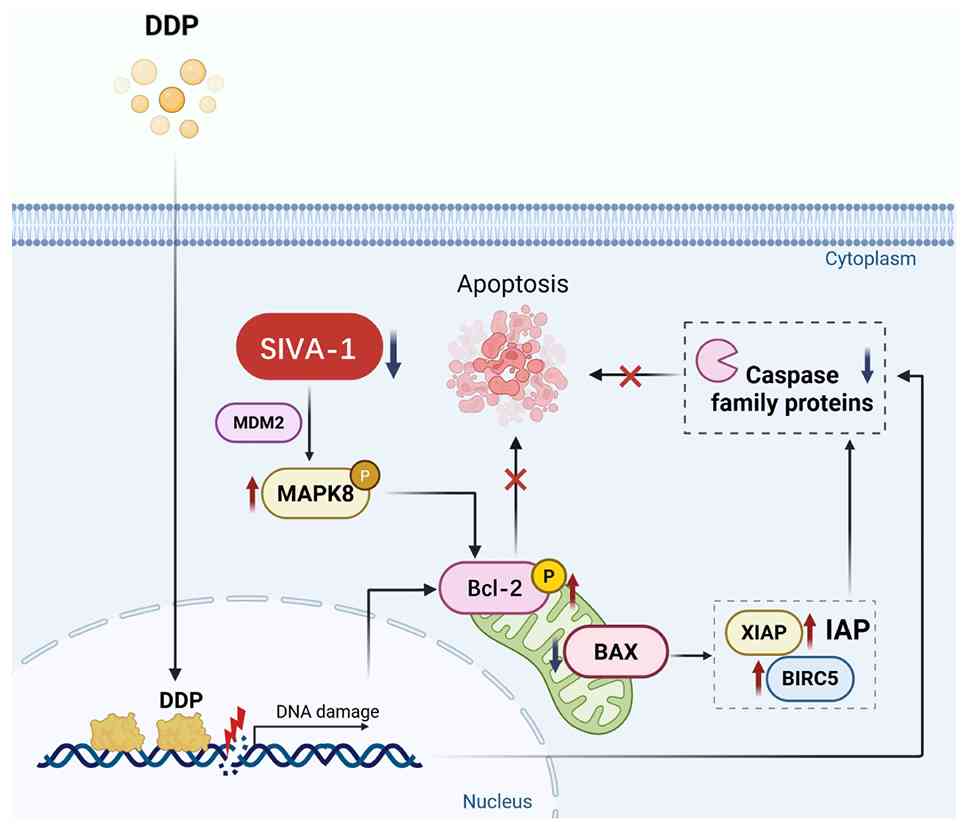
decreased in the shRNA-SIVA-1 group (P<0.05; Fig. 5H and I). Fig. 6 demonstrates that silencing SIVA-1
indirectly increases the expression of MAPK8 through MDM2, thereby
triggering the mitochondrial-dependent apoptosis pathway, which is
mediated by the Bcl-2/BAX signaling pathway. It is specifically
manifested as the upregulation of Bcl-2 and the downregulation of
BAX, and XIAP and BIRC5 are downstream regulatory genes of BAX.
Their simultaneous upregulation ultimately affects apoptosis
mediated by the caspase family of proteins. Therefore, silencing
SIVA-1 was shown to block the apoptotic pathway caused by DDP
through the MAPK8/Bcl-2/BAX signaling pathway.
Discussion
SIVA-1 is a gene closely related to apoptosis. Under
the environmental conditions of hyperbaric oxygen, apoptosis can be
promoted in glioma cells by downregulating SIVA-1 expression
(11). Existing evidence has
suggested that SIVA-1 has the potential to influence drug
resistance in various types of cancer, including gastric cancer,
colorectal cancer (27), cervical
cancer (28) and hepatocellular
carcinoma (29).
Gastric cancer is one of the most common malignant
tumors of the digestive system, and the leading treatment is
radical surgical resection of the lesion. As the signs or clinical
symptoms of patients with early gastric cancer are not obvious,
according to Tan et al (30), ~70% of patients with gastric cancer
are in the advanced stages of the disease at the time of diagnosis.
For patients diagnosed with advanced gastric cancer, curative
surgical treatment is no longer considered an appropriate option;
instead, a comprehensive treatment plan centered on systemic
chemotherapy is often initiated (31). In order to improve prognosis and
survival, the use of combination chemotherapeutic drugs is
preferred in the clinical treatment of patients with advanced
gastric cancer (32). With the wide
application of various chemical drugs in the treatment of gastric
cancer, an increasing number of patients have experienced reduced
effectiveness of therapeutic drugs (33), ultimately leading to the progression
of advanced gastric cancer and a notable decrease in the survival
of patients. To address chemotherapeutic drug resistance, the
preliminary experimental results of the present study showed that
SIVA-1 expression was positively associated with the inhibition
rate of DDP in gastric cancer cells (34), leading to altered drug sensitivity
by mechanisms which have not yet been elucidated. In the present
study, SIVA-1 was used as an upstream target to study drug
resistance; its effects on drug-resistant gastric cancer cells and
the efficacy of chemotherapeutic drugs were investigated, and the
mechanism of reversing the occurrence of drug resistance was
explored.
Bioinformatics analysis showed that SIVA-1 was
expressed at a low level in DDP-resistant gastric cancer cell lines
and that the low level of expression predicted resistance to DDP.
However, the Cox proportional hazards model revealed that higher
SIVA-1 expression levels in patients with gastric cancer were
associated with an increased risk of death. It may be hypothesized
that SIVA-1 serves a dual role in both ordinary gastric cancer
cells and cisplatin-resistant gastric cancer cells. As demonstrated
in earlier studies, the upregulation of SIVA-1, mediated by the
long non-coding RNA FTX/microRNA-215-3p regulatory axis, promotes
gastric cancer cell proliferation (35). By contrast, a review by Parducci
et al (36) indicated that
SIVA-1 can promote apoptosis in gastric cancer and enhance the
effectiveness of chemotherapy. Nevertheless, the results of the
present study showed that silencing SIVA-1 in AGS/DDP cells
promoted their proliferation, migration and invasion. In addition,
an inextricable association with endogenous apoptosis was
suggested. It was also observed that the resistance of AGS/DDP
cells to DDP was enhanced after silencing SIVA-1, which further
confirmed the effect of SIVA-1 on the drug resistance of gastric
cancer cells.
Preliminary evidence has indicated that SIVA-1
affects gastric cancer DDP resistance by regulating MDM2 expression
(34). A previous study
demonstrated that MAPK8 is a downstream target gene of MDM2
(37). Analysis of the PPI network
diagram in the current study showed that SIVA-1 had an indirect
effect on MAPK8 and was associated with MDM2; this led to the
conclusion that SIVA1 indirectly regulates MAPK8 via MDM2. MAPK8,
also termed c-Jun N-terminal kinase, may be considered a tumor
resistance gene with a positive effect on eliminating tumor cells,
which can be activated by tumor necrosis factor and in turn
triggers the process of apoptosis (38). In addition, MAPK8 has been reported
to affect the biological functions of tumor cells through multiple
pathways, including proliferation, migration and apoptosis
(39). The bioinformatics analysis
results indicated that MAPK8, as a co-interacting protein with
SIVA-1, may be involved in the regulation of endogenous apoptotic
processes, which ultimately leads to a decrease in drug sensitivity
of drug-resistant gastric cancer cells.
According to a previous study, MAPK8 indirectly
inhibits apoptosis in hepatocellular carcinoma through the
phosphorylation of Bcl-2 (40), and
the key factor that influences apoptosis is the Bcl-2/BAX ratio. It
is well established that Bcl-2 is an anti-apoptotic gene and that
Bax is a pro-apoptotic gene (41),
both of which belong to the Bcl-2 family of proteins. The MF term
‘mitochondrial outer membrane’, as determined by GO enrichment
analysis, indicated that both Bcl-2 and BAX are located on the
outer mitochondrial membrane and belong to the same key hub of the
mitochondria-dependent apoptosis pathway. It has been suggested
that an altered Bcl-2/BAX ratio in gastric cancer affects the
mitochondria-dependent apoptotic pathway mediated by caspase family
proteins (42). Low et al
(43) observed that tumor cells
develop drug resistance due to reduced apoptosis as a result of
reduced accumulation of BAX in mitochondria, where the onset of
apoptosis may be related to endogenous apoptosis mediated by
caspase family proteins. Similarly, inhibition of Bcl-2 in acute
myeloid leukemia results in increased drug sensitivity through
activation of the mitochondrial autophagy process mediated by the
BH3 structural domain (44). The
results of the apoptosis assay in the present study demonstrated
that silencing the expression of SIVA-1 in AGS/DDP cells hindered
their progression through apoptosis. This ultimately resulted in an
augmentation of the resistance of AGS/DDP cells to DDP. This result
may be due to the activation of MAPK8/Bcl-2/BAX-mediated
mitochondria-dependent apoptosis by SIVA-1.
XIAP, as a gene downstream of BAX (45), is responsible for regulating caspase
family proteins and indirectly inhibiting the
mitochondria-dependent apoptosis pathway. XIAP is considered to be
an anti-apoptotic gene, similar to Bcl-2. One study showed that
simultaneous inhibition of XIAP with Bcl-2 resulted in a higher
rate of apoptosis in bladder cancer cells than inhibition of XIAP
or Bcl-2 alone (46). Previous
evidence has suggested that XIAP inhibits activation of the caspase
family of proteins, which in turn regulates apoptosis. BIRC5 and
XIAP belong to the IAP proteins family. Similar to XIAP, BIRC5 acts
as a downstream gene of BAX (47),
XIAP and BIRC5 work synergistically to affect cell function
(48). The BIR structural domain is
often considered to be a characteristic structure of the IAP family
of proteins (49), and is
associated with drug resistance (50). In addition, BIRC5 benefits from a
unique BIR structural domain that directly inhibits
mitochondria-dependent apoptosis by interacting with caspase family
proteins (51). Similarly, Zhu
et al (52) reported that
overexpression of BIRC5 can increase drug resistance in lung cancer
and suggested that this was caused by dysregulation of the
apoptotic pathway.
In conclusion, the present study aimed to validate
the effects of silencing SIVA-1 on the sensitivity of AGS/DDP cells
to DDP, and to further investigate the mechanisms of enhancement of
drug resistance and changes in tumor cell biological functions. A
total of five genes associated with mitochondrial apoptotic
pathways, MAPK8, Bcl-2, BAX, XIAP and BIRC5, were identified using
bioinformatics analysis. In addition, using cellular experiments
and constructing in vivo xenograft tumor model experiments,
it was revealed that silencing SIVA-1 expression caused an increase
in the expression levels of MAPK8, Bcl-2, XIAP and BIRC5 proteins,
and a decrease in the expression level of BAX protein. Furthermore,
an increase in resistance to DDP was observed. It was thus
indicated that SIVA-1 acts indirectly on MAPK8, which in turn
activates the Bcl-2/BAX-mediated mitochondria-dependent apoptosis
pathway, with XIAP and BIRC5 acting as downstream target genes of
BAX that synergize with caspase family proteins. Ultimately, it
alters the biological function of gastric cancer-resistant cells
and their sensitivity to chemotherapeutic agents. Thus, it is clear
that SIVA-1 is a central gene affecting drug resistance in gastric
cancer cells.
Acknowledgements
Not applicable.
Funding
The study was funded by the Natural Science Foundation of China
(grant no. 82260468), the Natural Science Foundation of Guangxi
(grant nos. 2023GXNSFAA026341, 2023GXNSFAA026143 and
2025GXNSFAA069965), the National Natural Science Foundation
Cultivation Fund of the People's Hospital of Guangxi Zhuang
Autonomous Region (grant nos. NSFC-HCF2025012 and 2026GPY0209) and
the Guangxi Medical and Health Appropriate Technology Development
and Promotion Project (grant no. S2024006).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
XTW and FBK confirm the authenticity of all the raw
data. ZYS, ZRD, LJX, SX, FBK and XTW contributed to the conception
and design of the study. YQ, YLH, YRL, HBH, MRD, XGZ, XML, LL and
XDD were involved in the acquisition, analysis and interpretation
of the data. ZYS, LJX, ZRD, SX, LL, FBK and XTW drafted the
manuscript. YQ, XW, FBK, XGZ and LL critically revised the
manuscript for important intellectual content. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The present study obtained approval from The Medical
Ethics Committee of Guangxi Zhuang Autonomous Region People's
Hospital (Nanning, China; approval no. KY-GZR-2016-416).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Qin N, Fan Y, Yang T, Yang Z and Fan D:
The burden of Gastric Cancer and possible risk factors from 1990 to
2021, and projections until 2035: Findings from the Global Burden
of Disease Study 2021. Biomark Res. 13:52025. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel RL, Giaquinto AN and Jemal A:
Cancer statistics, 2024. CA Cancer J Clin. 74:12–49.
2024.PubMed/NCBI
|
|
3
|
Gupta J, Ahmed AT, Tayyib NA, Zabibah RS,
Shomurodov Q, Kadheim MN, Alsaikhan F, Ramaiah P, Chinnasamy L and
Samarghandian S: A state-of-art of underlying molecular mechanisms
and pharmacological interventions/nanotherapeutics for cisplatin
resistance in gastric cancer. Biomed Pharmacother. 166:1153372023.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Liu Q, Song Y, Su J, Yang S, Lian Q, Wang
T, Wei H and Fang J: PUF60 promotes chemoresistance through drug
efflux and reducing apoptosis in gastric cancer. Int J Med Sci.
22:269–282. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Ji N, Li H, Zhang Y, Li Y, Wang P, Chen X,
Liu YN, Wang JQ, Yang Y, Chen ZS, et al: Lansoprazole (LPZ)
reverses multidrug resistance (MDR) in cancer through impeding
ATP-binding cassette (ABC) transporter-mediated chemotherapeutic
drug efflux and lysosomal sequestration. Drug Resist Updat.
76:1011002024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chen H, Li Y, Li H, Chen X, Fu H, Mao D,
Chen W, Lan L, Wang C, Hu K, et al: NBS1 lactylation is required
for efficient DNA repair and chemotherapy resistance. Nature.
631:663–669. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Babaei S, Nikbakht M, Majd A and Mousavi
SA: Comparative effects of arsenic trioxide and chemotherapy on
Chk1 and CDC25 gene expression in gastric cancer cells AGS and
MKN45: A potential therapeutic strategy. Mol Biol Rep. 52:1982025.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Xu F, Xu A, Guo Y, Bai Q, Wu X, Ji SP and
Xia RX: PM2.5 exposure induces alveolar epithelial cell apoptosis
and causes emphysema through p53/Siva-1. Eur Rev Med Pharmacol Sci.
24:3943–3950. 2020.PubMed/NCBI
|
|
9
|
Li N, Jiang P, Du W, Wu Z, Li C, Qiao M,
Yang X and Wu M: Siva1 suppresses epithelial-mesenchymal transition
and metastasis of tumor cells by inhibiting stathmin and
stabilizing microtubules. Proc Natl Acad Sci USA. 108:12851–1286.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ma Y, Liu T, Song X, Tian Y, Wei Y, Wang
J, Li X and Yang X: Siva 1 inhibits proliferation, migration and
invasion by phosphorylating Stathmin in ovarian cancer cells. Oncol
Lett. 14:1512–1518. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ren ZQ, Wang RD, Wang C, Ren XH, Li DG,
Liu YL and Yu QH: Key genes involved in the beneficial mechanism of
hyperbaric oxygen for glioblastoma and predictive indicators of
hyperbaric oxygen prolonging survival in glioblastoma patients.
Curr Med Sci. 44:1036–1046. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Kong F, Wu K, Pang L, Huang Y, Li L, Xu J,
Li F, Qing Y, Wang Z, Huang X, et al: Inhibition of
apoptosis-regulatory protein Siva-1 reverses multidrug resistance
in gastric cancer by targeting PCBP1. Oncol Res. 30:277–288. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Kong FB, Deng QM, Deng HQ, Dong CC, Li L,
He CG, Wang XT, Xu S and Mai W: Siva-1 regulates multidrug
resistance of gastric cancer by targeting MDR1 and MRP1 via the
NF-κB pathway. Mol Med Rep. 22:1558–1566. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Phoo NLL, Dejkriengkraikul P, Khaw-On P
and Yodkeeree S: Transcriptomic profiling reveals AKR1C1 and AKR1C3
mediate cisplatin resistance in signet ring cell gastric carcinoma
via autophagic cell death. Int J Mol Sci. 22:125122021. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Love MI, Huber W and Anders S: Moderated
estimation of fold change and dispersion for RNA-seq data with
DESeq2. Genome Biol. 15:5502014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Maeser D, Gruener RF and Huang RS:
oncoPredict: An R package for predicting in vivo or cancer patient
drug response and biomarkers from cell line screening data. Brief
Bioinform. 22:bbab2602021. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Yu G, Wang LG, Han Y and He QY:
clusterProfiler: An R package for comparing biological themes among
gene clusters. OMICS. 16:284–287. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Weber EM, Zidar J, Ewaldsson B, Askevik K,
Udén E, Svensk E and Törnqvist E: Aggression in Group-housed male
mice: A systematic review. Animals (Basel). 13:1432022. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lidster K, Owen K, Browne WJ and Prescott
MJ: Cage aggression in group-housed laboratory male mice: An
international data crowdsourcing project. Sci Rep. 9:152112019.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Van Loo PL, Van Zutphen LF and Baumans V:
Male management: Coping with aggression problems in male laboratory
mice. Lab Anim. 37:300–313. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
CCAC Guidelines, . Mice. A4, 118 pages.
Canadian Council on Animal Care. 2019.Available at:. https://ccac.ca/Documents/Standards/Guidelines/CCAC_Guidelines_Mice-Sept2022.pdf
|
|
23
|
Workman P, Aboagye EO, Balkwill F, Balmain
A, Bruder G, Chaplin DJ, Double JA, Everitt J, Farningham DA,
Glennie MJ, et al: Guidelines for the welfare and use of animals in
cancer research. Br J Cancer. 102:1555–1577. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Detre S, Saclani Jotti G and Dowsett M: A
‘quickscore’ method for immunohistochemical semiquantitation:
Validation for oestrogen receptor in breast carcinomas. J Clin
Pathol. 48:876–878. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Kong FB, Shi ZY, Huang YL, Chen HH, Deng
QM, Wu K, Zhu Z, Li L, Xu S, Zhong XG, et al: SIVA-1 interaction
with PCBP1 serves as a predictive biomarker for cisplatin
sensitivity in gastric cancer and its inhibitory effect on tumor
growth in vivo. J Cancer. 15:4301–4312. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Resch U, Schichl YM, Winsauer G, Gudi R,
Prasad K and de Martin R: Siva1 is a XIAP-interacting protein that
balances NFkappaB and JNK signalling to promote apoptosis. J Cell
Sci. 122:2651–2661. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lin Z, Wan AH, Sun L, Liang H, Niu Y, Deng
Y, Yan S, Wang QP, Bu X, Zhang X, et al: N6-methyladenosine
demethylase FTO enhances chemo-resistance in colorectal cancer
through SIVA1-mediated apoptosis. Mol Ther. 31:517–534. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Liu T, Ma Y, Wang Z, Zhang W and Yang X:
Siva 1 inhibits cervical cancer progression and its clinical
prognosis significance. Cancer Manag Res. 12:303–311. 2020.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li T, Lv M, Chen X, Yu Y, Zang G and Tang
Z: Plumbagin inhibits proliferation and induces apoptosis of
hepatocellular carcinoma by downregulating the expression of SIVA.
Drug Des Devel Ther. 13:1289–1300. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Tan Z: Recent advances in the surgical
treatment of advanced gastric cancer: A review. Med Sci Monit.
25:3537–3541. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Guan WL, He Y and Xu RH: Gastric cancer
treatment: Recent progress and future perspectives. J Hematol
Oncol. 16:572023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Takahari D: Second-line chemotherapy for
patients with advanced gastric cancer. Gastric Cancer. 20:395–406.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang W, Chen J, Wei Z, Song J, Zha X,
Wang D and Xu M: Advancements and challenges in immunotherapy for
gastric cancer: Current approaches and future directions. Front
Immunol. 16:15927332025. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang XT, Li L, Kong FB, Zhong XG and Mai
W: Lentivirus-mediated overexpression of SIVA-1 reverses cisplatin
resistance in gastric cancer in vitro. Cell Biochem Biophys.
78:455–463. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhang F, Wang XS, Tang B, Li PA, Wen Y and
Yu PW: Long non-coding RNA FTX promotes gastric cancer progression
by targeting miR-215. Eur Rev Med Pharmacol Sci. 24:3037–3048.
2020.PubMed/NCBI
|
|
36
|
Parducci NS, Garnique ADMB, de Almeida BO
and Machado-Neto JA: Exploring the dual role of SIVA1 in cancer
biology. Gene. 950:1493652025. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Gao X, Wei M, Shan W, Liu Q, Gao J, Liu Y,
Zhu S and Yao H: An oral 2-hydroxypropyl-β-cyclodextrin-loaded
spirooxindole-pyrrolizidine derivative restores p53 activity via
targeting MDM2 and JNK1/2 in hepatocellular carcinoma. Pharmacol
Res. 148:1044002019. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Heaton WL, Senina AV, Pomicter AD, Salama
ME, Clair PM, Yan D, Bell RN, Gililland JM, Prchal JT, O'Hare T and
Deininger MW: Autocrine Tnf signaling favors malignant cells in
myelofibrosis in a Tnfr2-dependent fashion. Leukemia. 32:2399–2411.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Parra E, Gutiérrez L and Ferreira J:
Inhibition of basal JNK activity by small interfering RNAs enhances
cisplatin sensitivity and decreases DNA repair in T98G glioblastoma
cells. Oncol Rep. 33:413–418. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Chen S, Du Y, Xu B, Li Q, Yang L, Jiang Z,
Zeng Z and Chen L: Vaccinia-related kinase 2 blunts sorafenib's
efficacy against hepatocellular carcinoma by disturbing the
apoptosis-autophagy balance. Oncogene. 40:3378–3393. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Adams JM and Cory S: Life-or-death
decisions by the Bcl-2 protein family. Trends Biochem Sci.
26:61–66. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Chen J, Ji Z, Wu D, Wei S, Zhu W, Peng G,
Hu M, Zhao Y and Wu H: MYBL2 promotes cell proliferation and
inhibits cell apoptosis via PI3K/AKT and BCL2/BAX/Cleaved-caspase-3
signaling pathway in gastric cancer cells. Sci Rep. 15:91482025.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Low HB, Wong ZL, Wu B, Kong LR, Png CW,
Cho YL, Li CW, Xiao F, Xin X, Yang H, et al: DUSP16 promotes cancer
chemoresistance through regulation of mitochondria-mediated cell
death. Nat Commun. 12:22842021. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Glytsou C, Chen X, Zacharioudakis E,
Al-Santli W, Zhou H, Nadorp B, Lee S, Lasry A, Sun Z, Papaioannou
D, et al: Mitophagy promotes resistance to BH3 mimetics in acute
myeloid leukemia. Cancer Discov. 13:1656–1677. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
An HG, Shin S, Lee B, Kwon Y, Kwon TU,
Kwon YJ and Chun YJ: Induction of synergistic apoptosis by
tetramethoxystilbene and nutlin-3a in human cervical cancer cells.
Toxicol Res. 38:591–600. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Kunze D, Kraemer K, Erdmann K, Froehner M,
Wirth MP and Fuessel S: Simultaneous siRNA-mediated knockdown of
antiapoptotic BCL2, Bcl-xL, XIAP and survivin in bladder cancer
cells. Int J Oncol. 41:1271–1277. 2012.PubMed/NCBI
|
|
47
|
Al-Astani Tengku Din TA, Shamsuddin SH,
Idris FM, Ariffin Wan Mansor WN, Abdul Jalal MI and Jaafar H:
Rapamycin and PF4 induce apoptosis by upregulating Bax and
down-regulating survivin in MNU-induced breast cancer. Asian Pac J
Cancer Prev. 15:3939–3944. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Hehlgans S, Petraki C, Reichert S, Cordes
N, Rödel C and Rödel F: Double targeting of Survivin and XIAP
radiosensitizes 3D grown human colorectal tumor cells and decreases
migration. Radiother Oncol. 108:32–39. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Berthelet J and Dubrez L: Regulation of
apoptosis by inhibitors of apoptosis (IAPs). Cells. 2:163–187.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Shi L, Lu J, Xia X, Liu X, Li H, Li X, Zhu
J, Li X, Sun H and Yang X: Clinically used drug arsenic trioxide
targets XIAP and overcomes apoptosis resistance in an
organoid-based preclinical cancer model. Chem Sci. 15:8311–8322.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Pavlyukov MS, Antipova NV, Balashova MV,
Vinogradova TV, Kopantzev EP and Shakhparonov MI: Survivin monomer
plays an essential role in apoptosis regulation. J Biol Chem.
286:23296–23307. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhu X, Zhou R, Lu Y, Zhang Y, Chen Q and
Li Y: Identification and validation of afatinib potential drug
resistance gene BIRC5 in Non-Small cell lung cancer. Front Oncol.
11:7630352021. View Article : Google Scholar : PubMed/NCBI
|