Introduction
Cervical cancer is a major global health burden,
with an estimated 342,000 patients dying from the disease annually
(1). The overall prognosis of
patients with cervical cancer is poor due to recurrence and
metastasis (2). Cervical cancer is
a complex process that involves multiple links, stages and genes
and the molecular mechanisms of cervical cancer are unclear
(3). Thus, a deeper understanding
of the underlying molecular mechanisms governing cervical cancer
development and progression is key to devising novel preventive
strategies and targeted therapeutic interventions.
Protein arginine methyltransferases (PRMTs) catalyze
the methylation of guanidinium nitrogen atoms of arginine residues,
using S-adenosylmethionine as the methyl donor (4). In eukaryotic organisms, methylated
arginine exists in three primary forms: Monomethylarginine,
asymmetric dimethylarginine and symmetric dimethylarginine (SDMA)
(5). To date, nine PRMT family
members have been identified, which are classified into three
different types based on their catalytic activity. Type I PRMTs
mediate asymmetric dimethylation of the two ω-nitrogen atoms on the
arginine side chain, while type II PRMTs catalyze symmetric
dimethylation of the same residues. A third subgroup comprises
monomethyltransferases, which introduce a single methyl group onto
arginine residues (6). Notably, an
additional type IV PRMT class has been characterized in fungi,
which generates δ-Ng-methylarginine (7). Accumulating evidence in recent years
has underscored the key role of PRMT-mediated arginine methylation
in multiple hallmarks of tumorigenesis, including cancer
development, metastasis and chemoresistance (8,9).
As a key type II protein arginine methyltransferase,
protein arginine methyltransferase 5 (PRMT5) catalyzes the
symmetric dimethylation of histone residues including H2R3, H4R3,
H3R2 and H2R8, thereby participating in the epigenetic regulation
of gene transcription (7). PRMT5
also mediates the methylation of non-histone proteins substrates
such as the transcription factors P21, P53 and Enhancer of Zeste
Homolog 2 (10). In the fetal
period, PRMT5 is highly expressed in the ovary, while it is lowly
expressed in other tissue and organs, and shows low expression in
all organs in adults, suggesting high PRMT5 expression is closely
related to the development of a number of tumors (11). Literature reports that high PRMT5
expression is associated with poor prognosis in triple-negative
breast cancer (12). PRMT5 is
upregulated in colorectal cancer and promotes tumor formation
(13,14). PRMT5 may serve as a potential target
for the treatment of cervical cancer and the development of
PRMT5-specific inhibitors or degraders in combination with
chemotherapeutic agents as therapeutic agents for cervical cancer
is becoming a popular study (15,16).
Studies have confirmed that multiple signaling
pathways are involved in tumorigenesis, including AKT/PI3K,
RAS/ERK, mTORC and NF-κB signaling pathways (17–19).
Wnt/β-catenin signaling controls key embryonic and somatic cellular
processes, such as determining cell fate, organogenesis and tissue
homeostasis. It contributes to pathological conditions including
inflammatory disorder and metabolic diseases, and is critically
involved in cancer development. Abnormal activation of
Wnt/β-catenin signaling is associated with many aspects of cancer
progression, malignant transformation and poor prognosis (20). The Wnt/β-catenin signaling pathway
serves an important role in the pathogenesis of cervical cancer
(21). Activation of Wnt signaling
promotes cervical cancer cell proliferation, migration, invasion,
cell cycle progression and resistance to apoptosis (22). Therefore, the Wnt/β-catenin
signaling pathway is considered an important target for the
treatment of cervical cancer.
MS4322 is a first-in-class, highly selective
proteolysis targeting chimera specifically targeting PRMT5, which
was developed via protein hydrolysis-targeted chimeric technology
by conjugating the PRMT5 inhibitor EPZ015666 with Von Hippel-Lindau
(VHL) E3 ligase ligand (S,R,S)-AHPC-Me (VHL-2) (23). Notably, MS4322 decreases PRMT5
expression in breast cancer MCF7 cells in a concentration-, time-,
VHL- and proteasome-dependent manner and exhibits favorable plasma
exposure in mouse pharmacokinetic studies (23). Further studies have revealed that
MS4322 treatment also leads to a significant decrease in PRMT5
protein expression and robust inhibition of cell proliferation in
multiple other cancer cell lines, including A549 (lung
adenocarcinoma), A172 (glioblastoma), and Jurkat (leukemia) cells
(24–26).
In summary, PRMT5 is highly expressed in a variety
of malignant tumors, and its high expression is associated with
malignant tumor progression and patient prognosis. To the best of
our knowledge, it has not been reported whether MS4322 has
antitumor effects on cervical cancer cells. Therefore, the present
study focused on whether PRMT5 is highly expressed in cervical
cancer tissue and has a tumor-promoting effect, as well as the
in vitro antitumor effect of MS4322.
Materials and methods
Chemicals and cell lines
MS4322 (purity, >95%) was synthesized by Tianjin
Gao Tan Technology Co. MS4322 was prepared as 90 mM master mix with
DMSO and stored at −20°C. Human cervical cancer HeLa cells were
obtained from Shanghai Institute of Cell Biology, Chinese Academy
of Sciences. Cells were incubated at 37°C in DMEM with 10%
(vol/vol) FBS and antibiotics (penicillin 100 U/ml and streptomycin
10 µg/ml; all Gibco (Thermo Fisher Scientific, Inc.).
Clinical samples
A total of three female patients diagnosed with
cervical cancer were enrolled at Linyi People's Hospital (Linyi,
China). Clinical samples, including cancerous and adjacent
non-cancerous tissues (>2 cm from the tumor margin) were
collected. Sample collection was conducted from June 2023 to August
2023. The patient ages ranged from 52 to 68 years, with a median
age of 60 years. A total of one patient was classified as
well-differentiated (G1), one as moderately differentiated (G2) and
one as poorly differentiated (G3); based on the International
Federation of Gynecology and Obstetrics (FIGO) 2021 staging system,
one patient was at stage IB2, one at stage IIA1 and one at stage
IIA2 (27). All patients had no
history of other malignant tumors or severe chronic disease before
enrollment, and their preoperative vital signs were stable. The
inclusion criteria were as follows: Histopathologically confirmed
cervical cancer with complete pathological data; and complete
clinical medical records containing key information. The exclusion
criteria were as follows: Concurrent in situ malignant
tumors at other sites; severe dysfunction of vital organs including
the heart, liver or kidney; uncontrolled infection; previous
exposure to any anti-tumor therapies; and incomplete medical
records or data that preclude data analysis. The present study was
approved by the Ethics Committee of Linyi People's Hospital, Linyi,
China; approval no. 202306-H-093) and conducted in accordance with
the Declaration of Helsinki. Written informed consent was obtained
from all participants prior to sample collection.
Cell Counting Kit (CCK)-8 assay
Cells (3×103/well) were seeded in 96-well
plate and incubated at 37°C overnight. The culture medium was
replaced with fresh complete medium (Gibco; Thermo Fisher
Scientific), and cells were treated with MS4322 (20, 40, 60, 80,
100, 120, 140 and 160 µM) at 37°C for 24, 48 and 72 h followed by
addition of 20 µl 10 µM CCK-8 solution (Beyotime Biotechnology) at
37°C for 4 h. Cell viability was calculated as follows:
[(Absorbance of experimental well)-(absorbance of blank
well)]/[(absorbance of control well)-(absorbance of blank well)]
×100%. Absorbance at 450 nm was measured using a microplate reader
(EPOCH-SN; Bio Tek Instruments, Inc.).
Colony formation assay
HeLa cells were plated in 6-well plates at 350
cells/well and treated with 120 µM MS4322 at 37°C, and the medium
was changed every 3 days. After 14 days at 37°C, cells were washed
twice with PBS at room temperature, fixed in 75% ethanol on ice for
30 min and then stained with 0.5% crystal violet at room
temperature for 30 min. A colony was defined as a cell cluster
consisting of >50 cells. Colonies were counted manually under an
inverted light microscope. Clone formation rate was calculated as
follows: Number of clones/number of inoculated cells ×100%. Each
experiment was repeated three times.
Wound healing assay
Hela cells were cultured to 90–100% confluence in
6-well plates. The cell monolayer was scraped with a 10 µl pipette
tip to produce a wound, washed with PBS to remove detached cells
and the cells were incubated in serum-free DMEM (Gibco; Thermo
Fisher Scientific, Inc.). Cells were incubated at 37°C for 24 h.
Photographs were taken at 0 and 24 h using a Nikon inverted light
microscope (Nikon Instruments). Wound width analysis was performed
with Image-Pro Plus 6.0 software (Media Cybernetics, Inc.). Data
are shown as the mean of three independent experiments.
Migration and invasion assay
For cell migration, 2.5×104 cells in 100
µl serum-free DMEM (Gibco, Thermo Fisher Scientific) were seed into
the upper compartment of Transwell inserts in 6-well plates. The
upper chamber was filled with serum-free DMEM. The lower chamber
was filled with DMEM containing 10% FBS (Gibco; Thermo Fisher
Scientific). Cells were incubated at 37°C for 72 h, then fixed with
4% paraformaldehyde for 5 min at room temperature. Following
washing with PBS three times, cells were stained with 0.5% crystal
violet blue at room temperature for 5 min, then washed with
double-distilled water. Cells on the upper surface of the insert
were removed with a cotton swab. The positively stained cells were
examined under an inverted light microscope. For the cell invasion
assay, Transwell inserts were precoated with Matrigel (Corning) at
37°C for 4 h to allow gelation.
Cell transfection
The PRMT5-small interfering RNA: Forward:
5′-GCCCAGUUUGAGAUGCCUU-3′ and reverse, 5′-CGGUCAAACUCUACGGAA-3′ and
negative control: Forward: 5′-UUCUCCGAACGUGUCACGU-3′ and reverse,
5′-ACGUGACACGUUCGGAGAA-3′ (HANBIO Biotechnology). The PRMT5
overexpression plasmid (OE-PRMT5) and PRMT5-small interfering RNA
(siRNA) were purchased from HANBIO Biotechnology Co., Ltd. HeLa
cells were seeded in culture plates at a density of
1.85×104 cells/cm2 and transfected at 70–80%
confluence. The PRMT5 overexpression plasmid, empty vector, PRMT5
or negative control siRNA were transfected into HeLa cells using
Lipofectamine 3000 (Thermo Fisher Scientific, Inc.) according to
the manufacturer's instructions. For plasmid transfection, 4 µg of
plasmid DNA was used per well of a 6-well plate. For siRNA
transfection, 1 µg of siRNA was used per well of a 6-well plate.
Transfection was performed at 37 for 6 h, after which the cells
were cultured for 24–48 h at 37°C in a 5% CO2 incubator
and then harvested for subsequent experiments.
Cell apoptosis HeLa cells were harvested and
co-stained with Annexin V-fluorescein isothiocyanate and propidium
iodide using the Annexin V apoptosis detection kit (cat. no.
556547; BD Biosciences), according to the manufacturer's
instructions. The apoptotic cells were analyzed by flow cytometry
(Accuri C6; BD Biosciences) using CellQuest Pro software, v3.3
(Becton, Dickinson and Company). Apoptosis rate was calculated as
follows: (early apoptotic cells + late apoptotic cells)/total cells
×100%
Cell cycle assay
The concentration of HeLa cells was adjusted to
2.5×105 cells/ml using DMEM and seeded into 6-well
plates. HeLa cells were treated with 120 µM MS4322 at 37°C for 3
days. Cells were collected and washed twice with PBS and 700 µl
pre-cooled 80% ethanol was added to give a final ethanol
concentration of 70%. Cells were fixed at 4°C for >4 h, then
washed twice again with PBS. A total of 100 µl RNase (50 µg/ml) and
50 µg/ml propidium iodide (50 µg/ml) were added in the dark at room
temperature for 30 min. Cell cycle distribution was analyzed by
flow cytometry using a BD FACSCalibur (BD Biosciences). Data were
analyzed using FlowJo software (version 10.6.2, BD Biosciences)
Untreated cells were used as the control group. Each experiment was
repeated three times.
RNA extraction and reverse
transcription-quantitative (RT-q)PCR
RNA was extracted using TRIzol (Thermo Fisher
Scientific, Inc.) and RT was performed using a PrimeScript RT
Reagent kit (Takara Bio, Inc.) at 37°C for 15 min, followed by heat
inactivation at 85°C for 5 sec. The qPCR reaction system consisted
of 10 µl SYBR Green Master Mix (Thermo Fisher Scientific, Inc.),
0.4 µl forward primer, 0.4 µl reverse primer, 0.4 µl 50X ROX
Reference Dye 2 (Thermo Fisher Scientific, Inc.), and 4.8 µl
double-distilled water. Amplification was performed using a
real-time PCR system under the following conditions: initial
denaturation at 95°C for 30 sec, followed by 40 cycles of
denaturation at 95°C for 5 sec and annealing at 60°C 30 sec. The
relative expression of target genes was calculated by
2−ΔΔCq method (28)
using GAPDH as the internal reference gene. Each experiment was
repeated three times. The primer sequences were as follows: PRMT5:
Forward, 5′-TGACCAATAAGAAGGGAT-3′ and reverse,
5′-GGCATTAGGTGGAGGAC-3′ and GAPDH: Forward,
5′-TCAAGAAGGTGGTGAAGCAGG-3′ and reverse,
5′-TCAAAGGTGGAGGAGTGGGT-3′.
Protein extraction and western
blotting
Cells were collected, washed twice with PBS, lysed
with PARP buffer (Beyotime Biotechnology) and centrifuged at 14,800
× g at 4°C for 5 min. The protein concentration was determined via
a BCA Protein Assay kit (Beyotime Biotechnology). Proteins (20
µg/lane) were separated using 10% SDS-PAGE and transferred to a
PVDF membrane, which was blocked with 5% non-fat milk in TBST (0.1%
Tween-20) for 1 h at room temperature. Membranes were then
incubated with primary antibodies at 4°C overnight. The membrane
antibodies included PRMT5 (cat. no. ab109451; 1:1,000; Abcam),
E-cadherin (cat. no. ab231303; 1:1,000; Abcam), snail (cat. no.
ab167609; 1:1,000; Abcam), Vimentin (cat. no. ab92547; 1:1,000;
Abcam), Bax (cat. no. ab32503; 1:1,000; Abcam), Bcl-2 (cat. no.
ab194583; 1:1,000; Abcam), GSK-3β (cat. no. ab32391; 1:1,000;
Abcam), C-myc (cat. no. ab32072; 1:1,000; Abcam), Wnt-3a (cat. no.
Ab219412; 1:1,000; Abcam) and GAPDH (cat. no. ab181602; 1:5,000;
all Abcam). Subsequent incubation with horseradish peroxidase
(HRP)-conjugated secondary antibodies (cat. no. SA00001-2; 1:3,000;
Wuhan Sanying) at room temperature for 1 h. Immunoreactive bands
were visualized using an enhanced chemiluminescence kit (Beyotime
Biotechnology). Images were scanned using a chemiluminescence and
fluorescence imaging system (Hangzhou Shenhua Technology Co., Ltd.)
and analyzed using Image-Pro plus software (Image-Pro plus 5.1,
Media Cybernetics, Inc.). GAPDH was used as an internal control to
regulate protein loading.
Immunohistochemistry (IHC)
The samples were fixed with 10% formalin at room
temperature for 24 h and embedded in paraffin. The
paraffin-embedded samples were cut into 4-µm-thick sections. The
prepared tissue sections were subjected to IHC staining for the
detection of PRMT5 protein. The samples were baked at 60°C for 3 h
to enhance tissue adherence. Next, the sections were deparaffinized
in xylene and rehydrated through a graded series of ethanol
solution. Endogenous peroxidase activity was quenched with 3%
hydrogen peroxide at room temperature for 10 min. Antigen retrieval
was performed by incubating the sections in EDTA buffer (cat. no.
ZLI-9072; OriGene) and heating in a microwave oven for 13–15 min.
After blocking with 5% goat serum (cat. no. DD13; OriGene) at room
temperature for 30 min, the sections were incubated with primary
antibody against PRMT5 (cat. no. ab109451; 1:200; Abcam) overnight
at 4°C. Following four washes with PBS, the sections were incubated
with horseradish peroxidase-conjugated goat anti-rabbit/mouse
secondary antibody (cat. no. PV-9000; 1:1,000; ZSGB Biotechnology)
at 37°C for 30 min. The sections were washed four times with PBS
again. Finally, the sections were developed with DAB at room
temperature for 3–5 min, followed by counterstaining with Mayer's
hematoxylin at room temperature for 1–2 min. All staining and
morphological observations were performed using a light microscope.
Image acquisition and quantitative analysis of IHC were performed
using ImageJ software (Version 1.53k, National Institutes of
Health).
Immunofluorescence detection
Following incubation with 120 µM MS4322 for 72 h at
37°C in a humidified incubator with 5% CO2, HeLa cells
were fixed with 70% methanol at room temperature for 10 min and
blocked with PBS containing 10% BSA (Thermo Fisher Scientific,
Inc.) at room temperature for 30 min. Cells were incubated with
primary antibodies overnight at 4°C. The primary antibodies were as
follows: E-cadherin (cat. no. ab231303; 1:200; Abcam), Snail (cat.
no. ab167609; 1:200; Abcam), and Vimentin (cat. no. ab92547; 1:200;
Abcam). After primary antibody incubation, cells were washed three
with PBS. Subsequently, cells were incubated with Cy3-labeled
anti-rabbit secondary antibody (cat. no. ab6939; 1:400; Abcam) at
room temperature for 1 h and stained with DAPI (2.0 µg/ml) at room
temperature for 5 min. Fluorescence signals were detected using an
inverted fluorescence microscope (cat. no. DM505, Nikon Co.,
Ltd.).
Statistical analysis
All quantitative data were obtained from at least
three independent biological replicates and are presented as mean ±
standard deviation (SD). Comparisons between two independent groups
were performed using the unpaired Student's t-test, while
comparisons among >2 groups were conducted via one-way ANOVA
followed by Tukey's post hoc test; for data that violated variance
homogeneity, Dunnett's T3 test was employed instead. All
statistical analyses were carried out using SPSS 20.0 (IBM Corp.)
and GraphPad Prism 9.0 software (GraphPad Software Inc.;
Dotmatics). P<0.05 was considered to indicate a statistically
significant difference.
Results
PRMT5 is highly expressed in cervical
cancer tissue
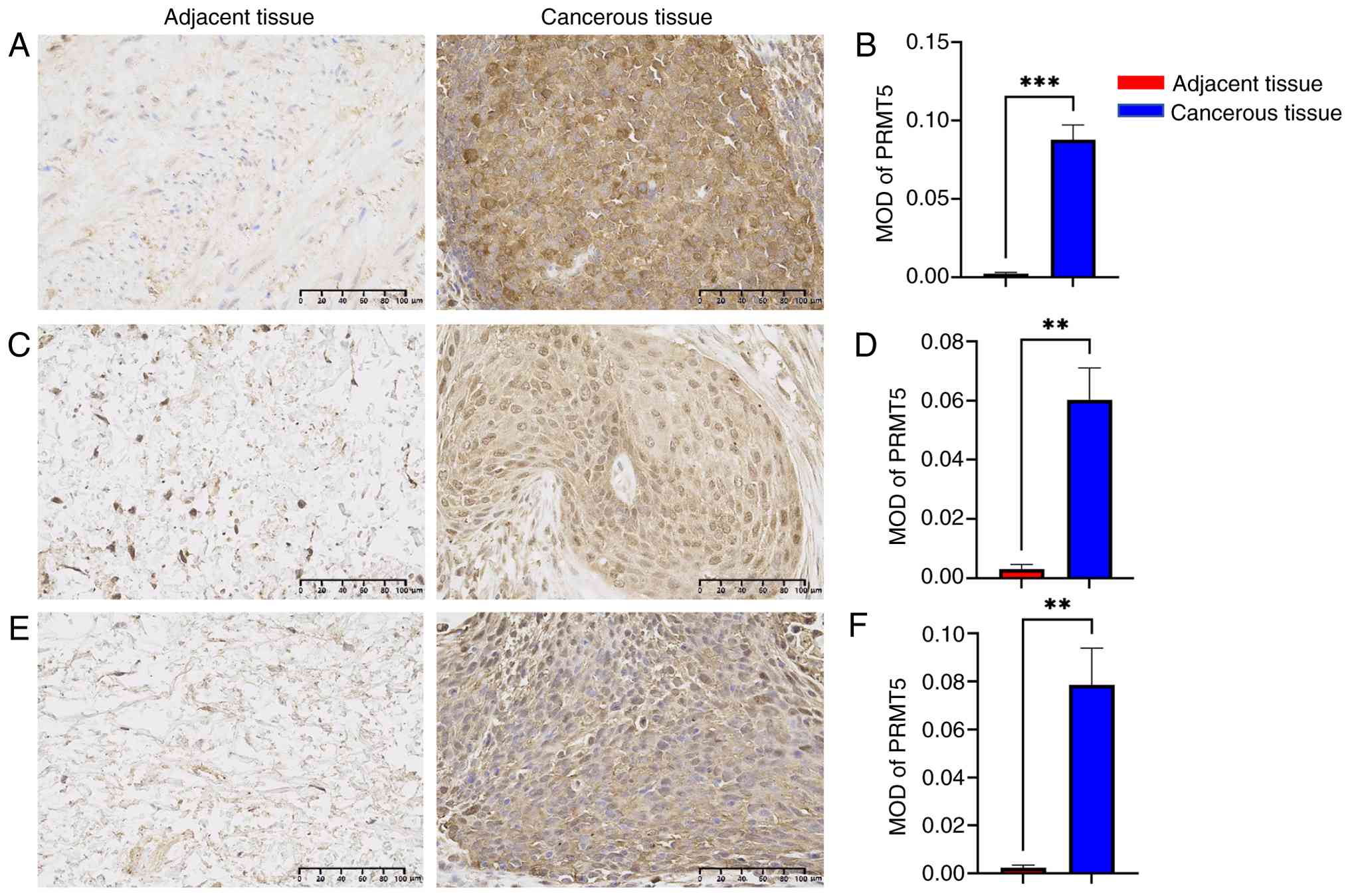
PRMT5 has been documented to be overexpressed in
multiple types of malignancy, including breast (12) and colorectal cancer (13). To investigate the expression profile
of PRMT5 in cervical cancer and their matched adjacent tissue, IHC
was performed to detect PRMT5 expression in paired tissue samples
derived from three patients. PRMT5 exhibited significantly elevated
expression in cervical cancer tissues compared with adjacent tissue
(Fig. 1).
PRMT5 overexpression promotes HeLa
cell migration and invasion, while knockdown of PRMT5 inhibits HeLa
cell migration and invasion
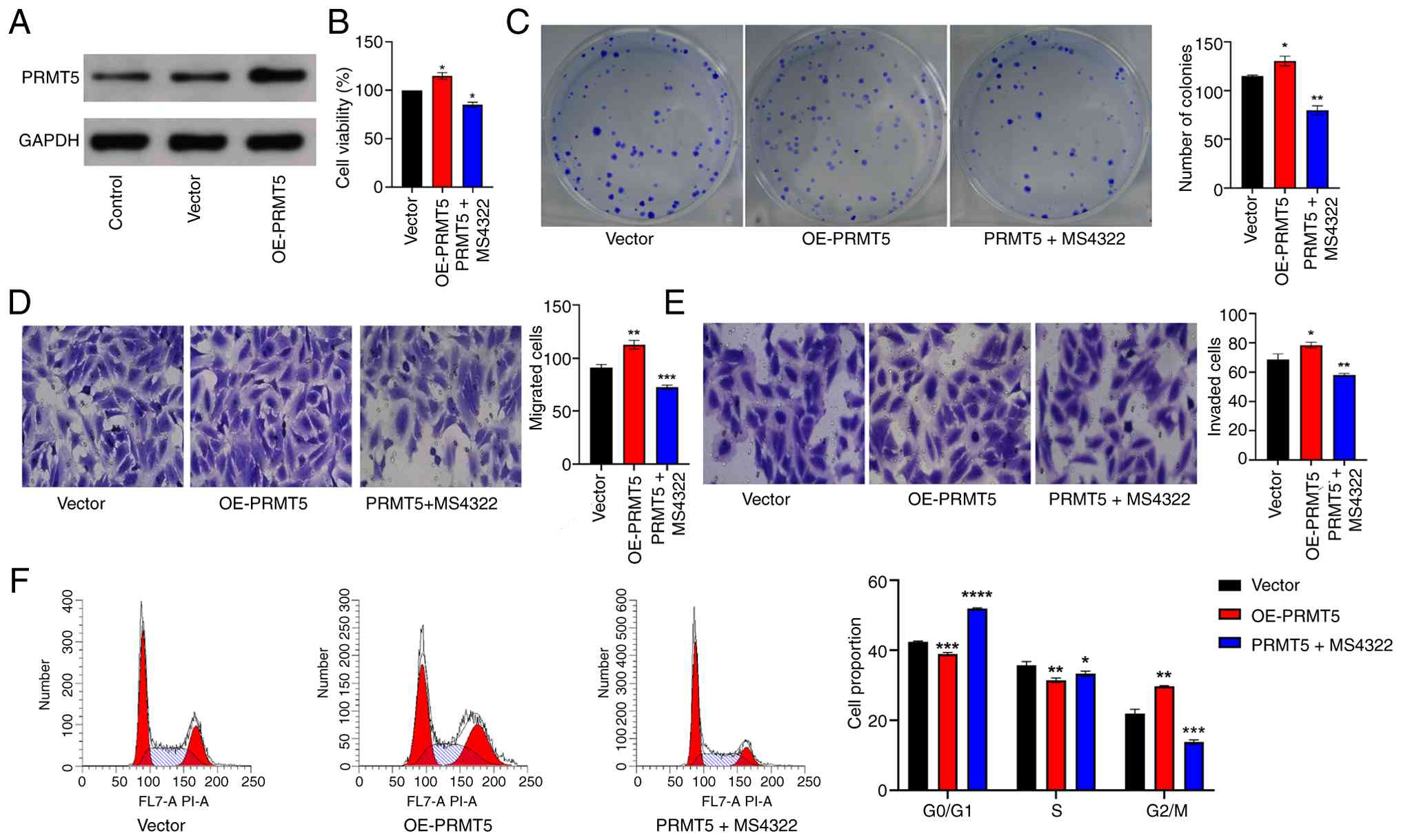
To elucidate the functional role of PRMT5 in
cervical cancer cells, the present study generated stable HeLa cell
lines with PRMT5 overexpression (Fig.
2A) and knockdown (Fig.
3A).
CCK-8 and colony formation assay revealed that
stable PRMT5 overexpression significantly enhanced cell
proliferation and colony-forming capacity relative to control cells
(Fig. 2B and C). Transwell
migration and invasion assays further indicated that
PRMT5-overexpressing HeLa cells exhibited a significant increase in
migratory and invasive potential (Fig.
2D and E). In addition, flow cytometry cell cycle analysis
showed that compared with control cells, PRMT5-overexpressing HeLa
cells displayed a decreased proportion of cells in the G0/G1 and S
phases, accompanied by an increase in the G2/M phase fraction
(Fig. 2F). These PRMT5-mediated
pro-tumorigenic effects were abrogated following treatment with
MS4322.
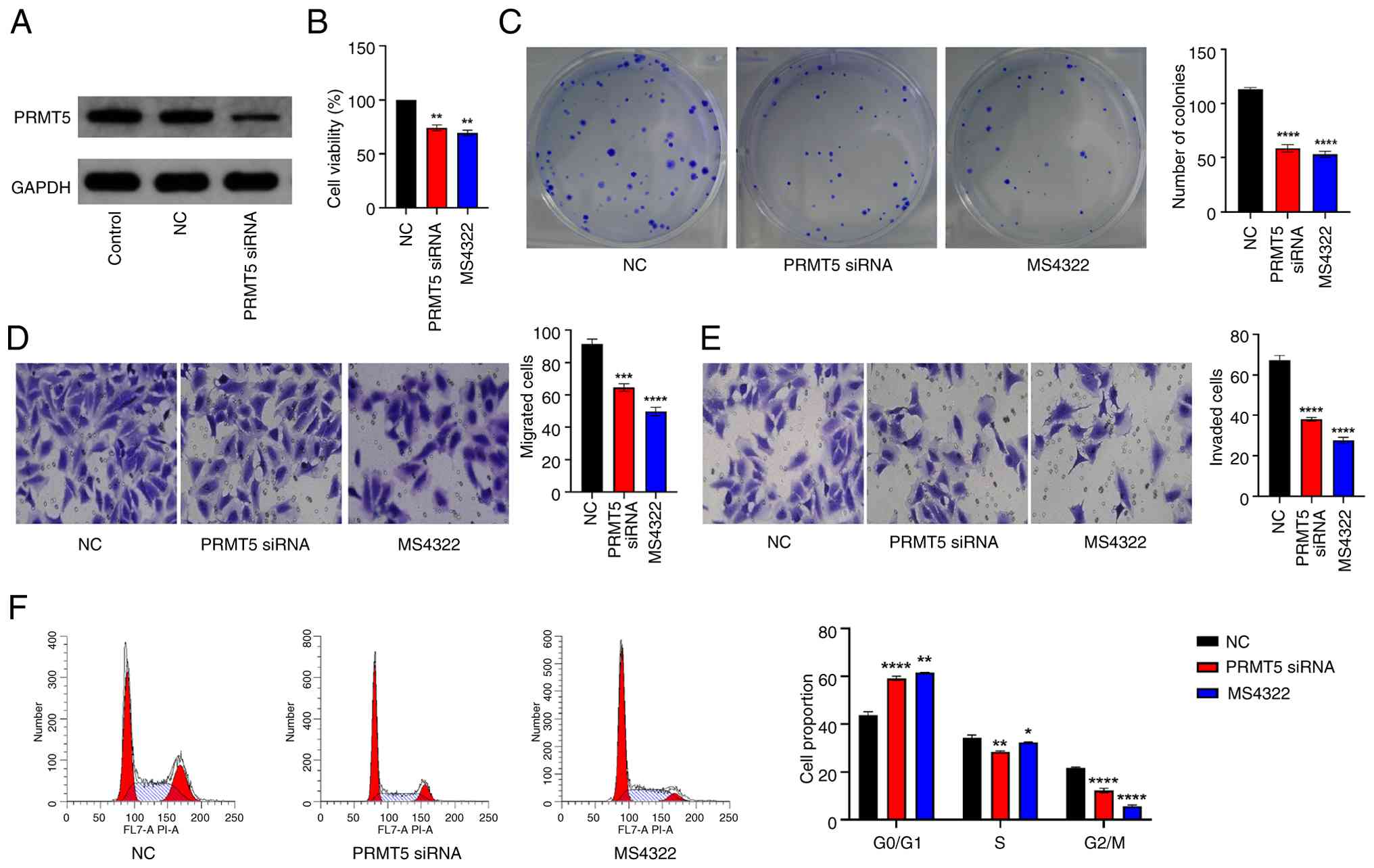
Conversely, PRMT5 knockdown in HeLa cells resulted
in a significant inhibition of cell proliferation and
colony-forming ability (Fig. 3B and
C). Consistently, Transwell assay demonstrated that migratory
and invasive capacity were significantly impaired in
PRMT5-knockdown HeLa cells (Fig. 3D and
E). Flow cytometric analysis further confirmed that PRMT5
knockdown induced a cell cycle arrest characterized by an increased
proportion of cells in the G0/G1 phase and a decrease in the S and
G2/M phases (Fig. 3F).
MS4322 inhibits cervical cancer HeLa
cell viability in vitro
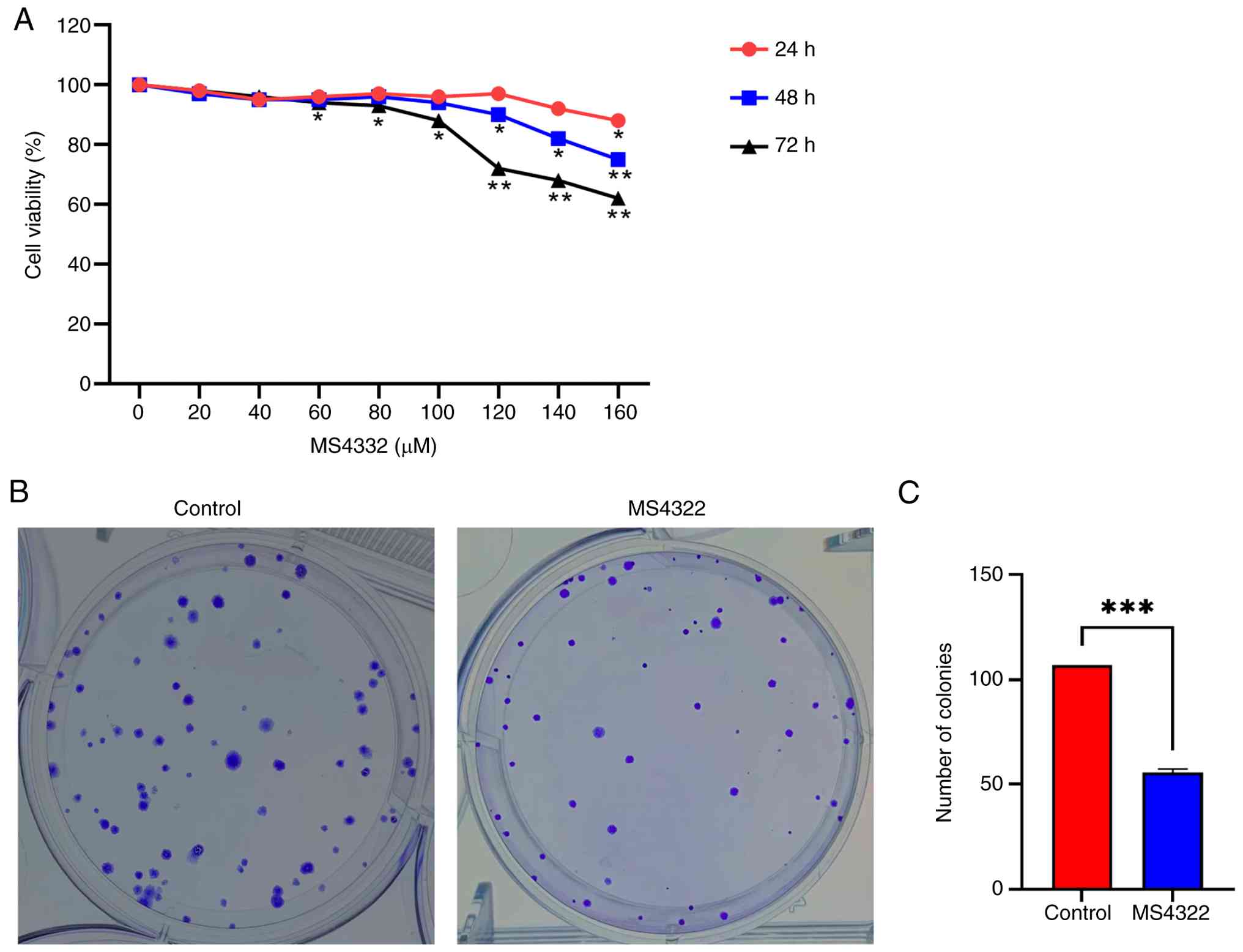
HeLa cells were treated with MS4322 and the cell
viability was detected by the CCK-8 assay (Fig. 4A). MS4322 significantly inhibited
the viability of HeLa cells in vitro in a concentration- and
time-dependent manner. The inhibitory effect was not obvious when
the concentration was <120 µM and at 24 and 48 h. Therefore,
MS4322 at 120 µM and 72 h were selected for the subsequent
experiments. The half-maximal inhibitory concentration of MS4322 on
HeLa cells was 170.408 µM. Compared with the control group, the
number of colonies formed by HeLa cells was reduced after 72 h
MS4322 treatment (Fig. 4B),
indicating that MS4322 had a significant inhibitory effect on the
colony formation ability of HeLa tumor cells (Fig. 4C).
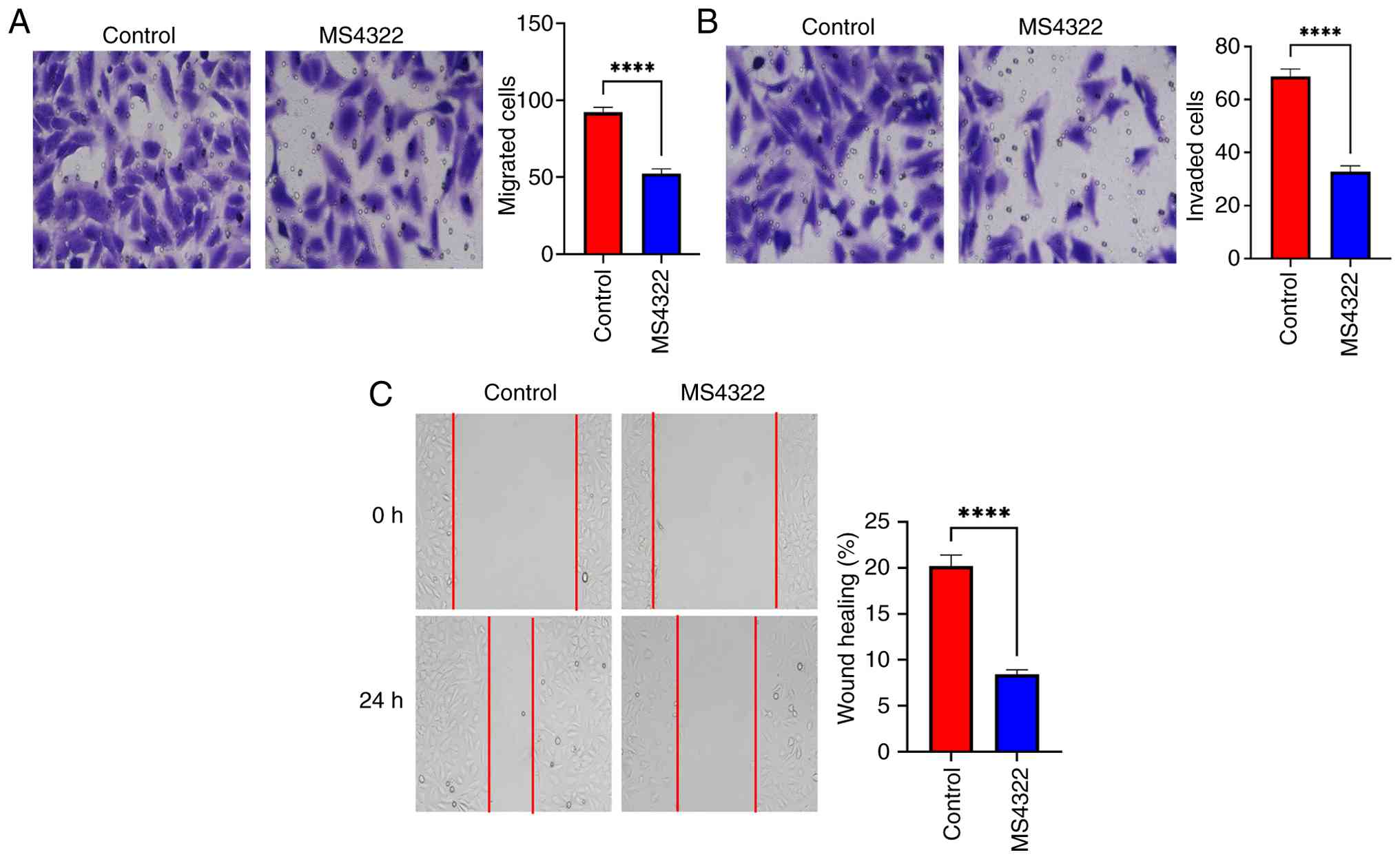
MS4322 inhibits the migration and
invasion of HeLa cells
Cell migration plays a key role in a variety of
cellular physiopathological processes, including tumor metastasis
(29). The present study examined
the effect of MS4322 on HeLa cell migration and invasion. The
number of migrating HeLa cells was significantly decreased when
treated with MS4322 compared with the control (Fig. 5A). In the wound healing assay,
untreated HeLa cells migrated to fill the scratched area within 24
h, but MS4322 significantly prevented migration of HeLa cells
(Fig. 5C). Taken together, these
experiments confirmed that MS4322 inhibited the migration of HeLa
cells. To investigate the effect of MS4322 on HeLa cell invasion,
Matrigel assay was performed. Compared with the control group, the
number of invaded HeLa cells was significantly decreased in the
MS4322-treated group (Fig. 5B).
These results indicated that MS4322 significantly inhibited the
migration and invasion ability of HeLa cells in vitro.
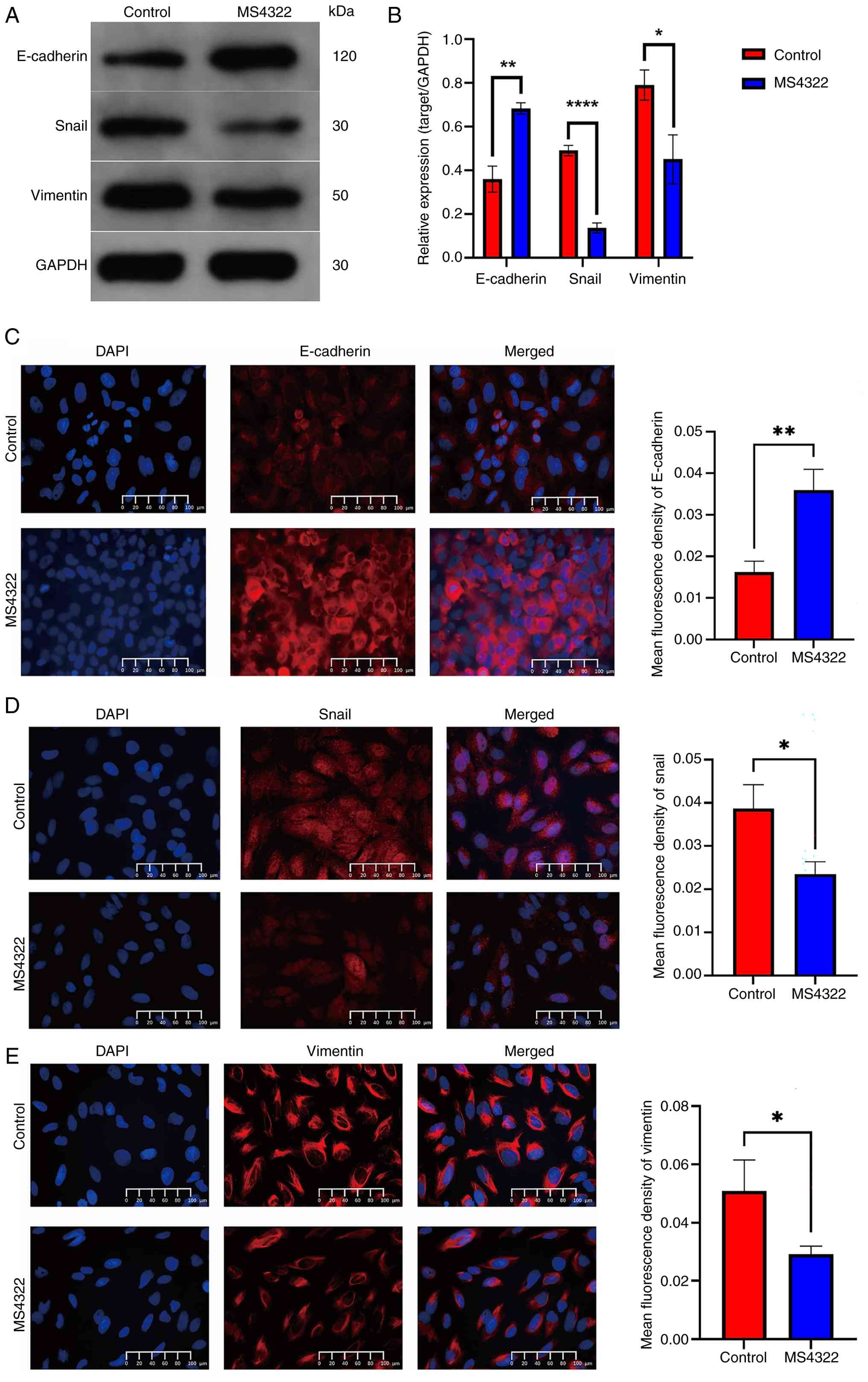
MS4322 inhibits epithelial-mesenchymal
transition (EMT) in HeLa cells
EMT is hypothesized to play an important function in
tumor cell migration and invasion (30). Therefore, the present study examined
the effect of MS4322 on the expression of EMT-associated proteins
in HeLa cells. E-cadherin protein expression was significantly
upregulated, while Snail and vimentin protein expression was
significantly downregulated in HeLa cells treated with MS4322
(Fig. 5A and B). The fluorescence
intensity of Snail and Vimentin was significantly weakened
(Fig. 6D and E) and the
fluorescence intensity of E-cadherin was significantly enhanced
(Fig. 6C) after MS4322 treatment
compared with the control group. These results suggested that the
inhibitory effect of MS4322 on HeLa cell migration and invasion may
be related to EMT.
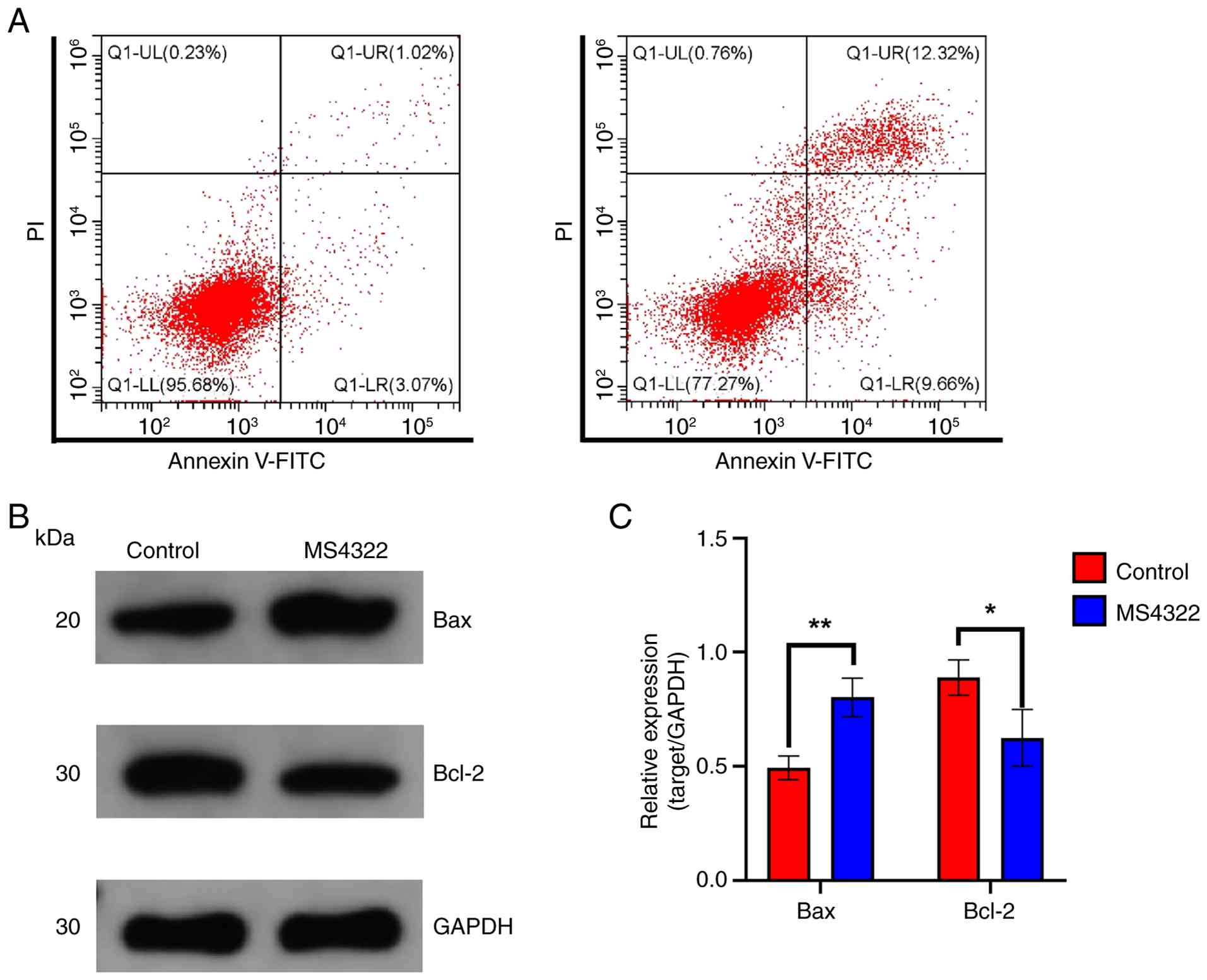
MS4322 induces apoptosis in HeLa
cells
Apoptosis serves an important role in both
tumorigenesis and therapy (31).
The number of Annexin V-positive cells significantly increased from
4.09 to 21.98% after 72 h MS4322 treatment, suggesting that MS4322
promoted apoptosis in HeLa cells (Fig.
7A). In addition, the expression of apoptosis-associated
proteins was detected by western blotting following MS4322
treatment of HeLa cells. MS4322 significantly inhibited the
expression of Bcl-2 protein and elevated the expression of Bax
protein (Fig. 7B and C). These
results suggested that MS4322 influenced the degree of apoptosis in
cervical cancer HeLa cells.
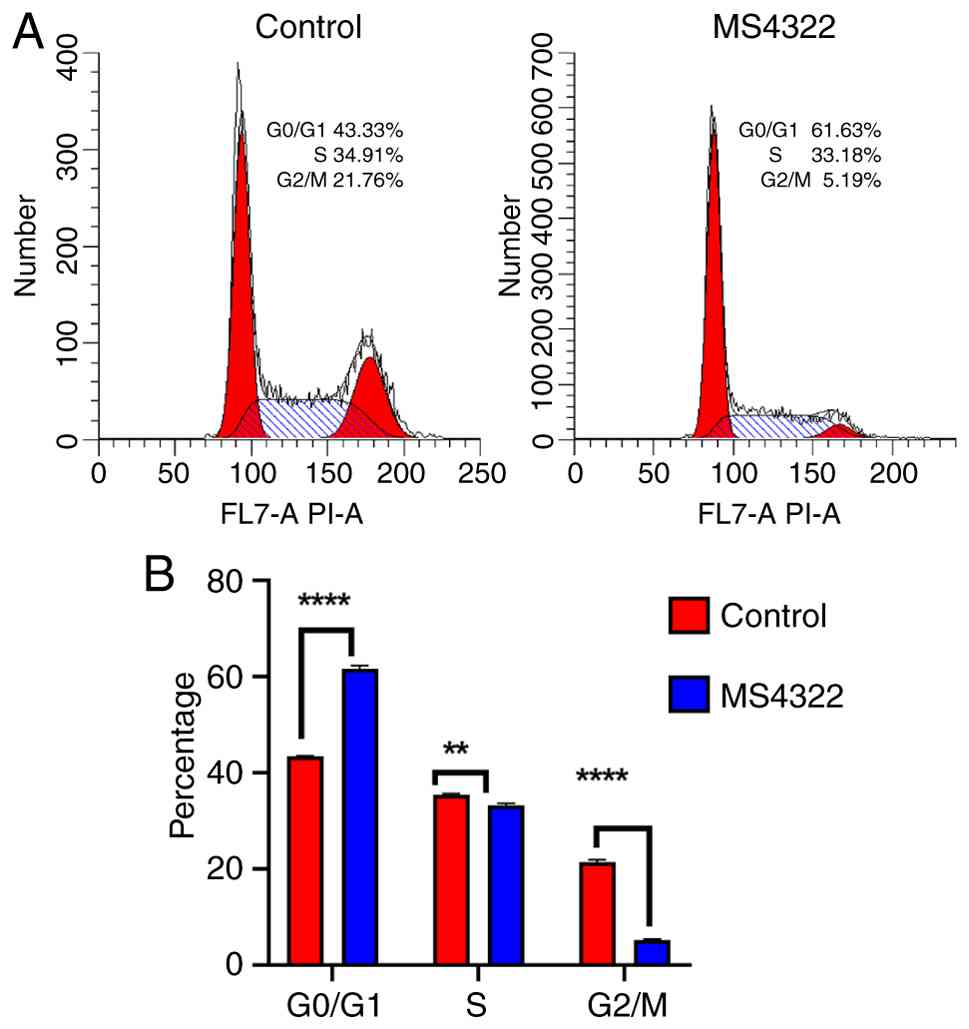
MS4322 induces G0/G1 phase arrest in
HeLa cells
MS4322 induced apoptosis in HeLa cells, and
inhibition of cancer cell proliferation is usually associated with
cell cycle arrest. To investigate the potential mechanism by which
MS4322 inhibited the proliferation of HeLa cells, the present study
analyzed the effect of MS4322 on the cell cycle by flow cytometry.
The proportions of untreated HeLa cells in G0/G1, G2/M and S phase
were 43.33, 34.91 and 21.76%, respectively; however, the
proportions of HeLa cells in G0/G1, G2/M and S phases were 61.63,
33.18 and 5.19%, respectively, when treated with MS4322 (Fig. 8A). The proportion of cells in G0/G1
phase was significantly increased, and the proportion of cells in
G2/M and S phases was significantly decreased following treatment,
which indicated that MS4322 arrested cells in G0/G1 phase (Fig. 8B).
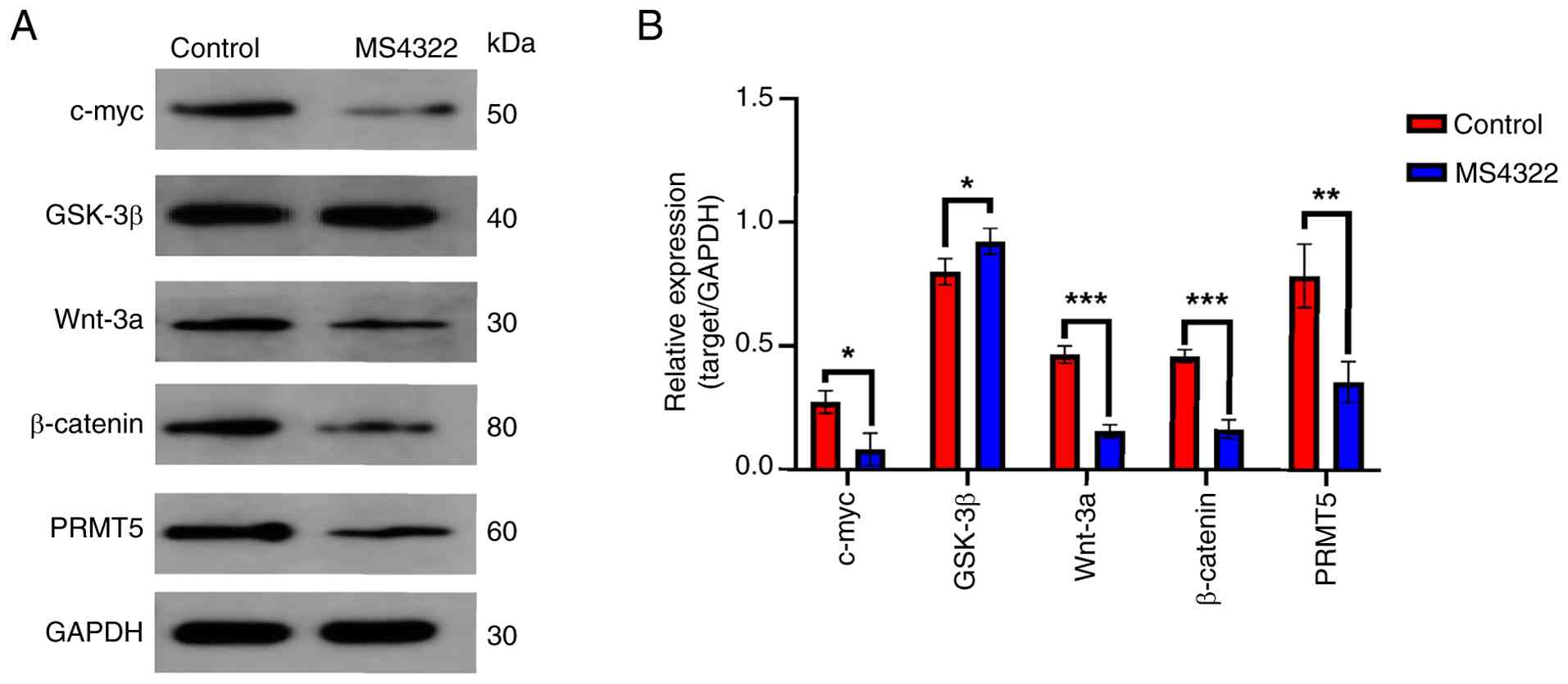
MS4322 inhibits Wnt/β-catenin pathway
activity in cervical cancer cells
The Wnt/β-catenin pathway serves a key role in
several biological processes, including cell proliferation,
differentiation, migration and apoptosis (32). β-catenin is the key molecule of the
pathway, and its stability and nuclear translocation are key steps
in the activation of the pathway. Wnt-3a, a member of the Wnt
family, activates the pathway, while glycogen synthase kinase 3β
(GSK-3β), a negative regulator of the pathway, inhibits pathway
activation by phosphorylating β-catenin and promoting its
degradation (20,21).
Decreased expression of β-catenin and Wnt-3a
protein, as well as increased expression of GSK-3β following MS4322
treatment suggested that MS4322 may inhibit the activation of the
Wnt/β-catenin pathway. In addition, the expression of c-myc, a
target protein downstream of the Wnt/β-catenin pathway, was also
significantly inhibited (Fig.
9).
Discussion
Cervical cancer is the fourth most common female
malignant tumor in the world, with ~660,000 new cases and 350,000
deaths globally in 2022 (33). It
is the second most prevalent cancer in developing countries,
accounting for 94% of global cervical cancer deaths and with an
incidence 2–3 times higher than that in developed countries
(34). Notably, its incidence is on
the rise globally, with a 50.1% increase in cases, and the age of
onset tends to be younger (35).
Epidemiological studies have shown that HeLa cells are associated
with human papillomavirus (HPV) infection and defective immune
function (36,37). Early cervical cancer has a high cure
rate, with a 5-year survival rate >90% for FIGO Stage I, while
advanced and recurrent cervical cancer has a poor prognosis, with a
5-year survival rate of only 10–20% (38). Cervical cancer cell proliferation,
metastasis and drug resistance are notable problems in clinical
treatment. Therefore, clarifying the specific mechanisms underlying
proliferation, recurrence and metastasis of cervical cancer and
searching for effective therapeutic targets are key to improve
treatment prospects.
Arginine methylation is one of the most common types
of post-translational modification of proteins in mammalian cells.
PRMTs are notable regulators of epigenetically mediated gene
expression, which modulates a variety of processes, including mRNA
splicing, DNA damage response, stem cell function, TGF-β and EMT
signaling pathways and immune response (39). Dysregulation of the expression of
PRMTs leads to cancer recurrence, decreases patient survival and
induces drug resistance by modulating the TGF-β, PI3K/Akt and
MAPK/ERK signaling pathways (40–42).
Numerous PRMTs are aberrantly expressed in cancer cells and promote
cancer stem cell generation, EMT and tumor cell proliferation. The
development of small molecules targeting PRMTs has led to the
generation of chemical probes for the modulation of most PRMTs and
treatment based on targeting PRMT1 and PRMT5 (43). PRMT5 is a key member of the type II
PRMT family. It contains an inter-terminal domain that binds
methylated epitope protein 50, which is essential for its full
methyltransferase activity. Its C-terminal domain harbors the
methyltransferase catalytic motifs required for plasma membrane
binding (44). PRMT5 serves a key
role in normal cellular processes by catalyzing monomethylation and
symmetric dimethylation of a range of histone and non-histone
substrates, and is also an oncoprotein that epigenetically
regulates the expression of certain oncogenes (45). For example, PRMT5 mediates the
symmetric dimethylation of histone substrates such as H4R3 and H3R8
to epigenetically silence tumor suppressor genes (46). It also targets multiple non-histone
substrates, including KEAP1 (9,12),
SMAD4 (42), and AKT (46), thereby modulating the NRF2/HMOX1,
TGF-β (42), and PI3K/AKT (46) signaling pathways to drive cancer
cell proliferation, metastasis and immunotherapy resistance.
Clinical studies (4,11) have shown that PRMT5 is highly
expressed in numerous types of solid tumor and hematological
malignancies and is associated with cancer development and
progression. In addition, increased PRMT5 expression is associated
with poorer overall survival and tumor metastasis (46,47).
In ovarian cancer cells, PRMT5 gene expression is significantly
upregulated, and this high expression is associated with poor
prognosis. Mechanistically, PRMT5 enhances glycolytic flux and
promotes the proliferation of ovarian cancer cells by regulating
SDMA-mediated activity of enolase 1 activity (48). PRMT5 is highly expressed in lung
cancer tissue and is associated with the progression of lung tumors
and patient survival; PRMT5 affects lung cancer growth by promoting
EMT and metastasis via hypoxia-inducible factor
1α/VEGFR/Akt/endothelial nitric oxide synthase (eNOS) signaling and
promoting angiogenesis (49).
Therefore, PRMT5 is becoming a promising anticancer target and has
received great attention from the pharmaceutical industry and
academia (50,51).
The present study demonstrated that PRMT5 was
significantly upregulated in human cervical cancer compared with
adjacent normal tissue. Functional experiments in HeLa cells
revealed that PRMT5 overexpression markedly promoted cell
proliferation, migration, invasion and colony formation, whereas
PRMT5 knockdown exerted the opposite effects. The molecular
mechanism by which PRMT5 induces HeLa cell proliferation and
migration primarily involves the Wnt/β-catenin signaling pathway.
PRMT5 upregulates Wnt-3a and β-catenin expression, thereby
activating the downstream target gene c-myc to promote cell cycle
progression and EMT (52). PRMT5
also suppresses apoptosis by upregulating the anti-apoptotic
protein Bcl-2 and downregulating the pro-apoptotic protein Bax
(53,54). At the phenotypic level, PRMT5
decreases the epithelial marker E-cadherin and increases the
mesenchymal markers vimentin and Snail, thereby facilitating
migration and invasion (52).
MS4322 is a highly selective PRMT5 degrader and the
first reported degrader targeting any PRMT family member. It is
constructed by conjugating the PRMT5 inhibitor EPZ015666 to VHL E3
ligase ligand (S,R,S)-AHPC-Me (VHL-2) (44). MS4322 decreases PRMT5 expression in
various cancer cell lines in a concentration-, time-, VHL- and
proteasome-dependent manner and effectively suppresses cancer cell
proliferation (44). The present
study demonstrated that MS4322 significantly suppressed the
viability, migration, invasion and colony formation of HeLa cells.
Mechanistically, MS4322 downregulates PRMT5 at both mRNA and
protein levels, reverses PRMT5-mediated oncogenic signaling,
induces G0/G1 cell cycle arrest, inhibits EMT by upregulating
E-cadherin and downregulating vimentin and snail, and promotes
apoptosis by increasing Bax and decreasing Bcl-2 expression.
Moreover, MS4322 suppresses the Wnt/β-catenin pathway by degrading
PRMT5, downregulating Wnt-3a, β-catenin and c-myc while enhancing
GSK-3β expression. Collectively, MS4322 targets PRMT5 to disrupt
the Wnt-β-catenin-EMT axis, thereby inhibiting proliferation,
migration and invasion and promoting apoptosis in HeLa cells.
EMT is regarded as a key pathological process in
tumor progression. In the malignant evolution of tumors, EMT
confers stronger invasive and metastatic abilities, and may allow
tumor cells to escape apoptosis induced by chemotherapeutic agents,
death receptor ligands and immune surveillance, which increases the
difficulty of treatment and the mortality rate of patients
(55). The present study
demonstrated EMT-associated protein changes following MS4322
treatment: MS4322 upregulated the expression of E-cadherin protein
and downregulated the expression of Snail and vimentin protein in
HeLa cells.
Apoptosis is a tightly regulated and evolutionarily
conserved program of cell death that serves a key role in life
activities. It not only serves a key role in normal physiological
processes such as embryogenesis and adult tissue homeostasis, but
is also known for its role as a tumor suppressor mechanism
(56). The role of apoptosis in
cancer has received widespread attention for its importance in
maintaining body homeostasis and preventing cancer development
(9,15,18).
However, when apoptosis is inhibited or resisted, it confers a
survival advantage to cancer cells, promotes tumor evolution and
growth and may lead to therapeutic failure (31,56,57).
The Bcl-2 family is associated with tumor progression and Bcl-2/Bax
ratio (pro- and anti-apoptotic proteins, respectively) determines
tumor cell survival or death (58).
The present study confirmed that MS4322 inhibited the protein
expression of Bcl-2 by increasing Bax protein expression in
cervical cancer HeLa cells. Consistently, BIIB021, an orally
active, fully synthetic and selective small-molecule inhibitor of
heat shock protein 90, has been reported to decrease Bcl-2 and
elevate Bax levels in human cervical cancer cells (59). Cell cycle abnormalities can cause a
variety of diseases, including cancer, and the rapid multiplication
of tumors depends on uncontrolled cell cycle progression (60). Following 72 h MS4322 treatment, the
number of cells in S and G2/M phase was decreased and the cells
were blocked in G0/G1 phase.
Aberrant activation of the Wnt/β-catenin signaling
pathway occurs in numerous types of malignant tumor and is closely
associated with cancer development and progression, as well as
renewal capacity and multidifferentiation potential of tumor cells
(61). Aberrant activation of
β-catenin contributes to cancer development (62). Dysregulation of the Wnt/β-catenin
pathway is associated with HPV-associated cervical cancer (63). The present study confirmed that
MS4322 downregulated PRMT5 mRNA expression in cervical cancer HeLa
cells and investigated its effect on the expression of proteins in
the Wnt/β-catenin signaling pathway. To the best of our knowledge,
the present study is the first to demonstrate that MS4322 enhanced
GSK-3β protein expression. By contrast, β-catenin, Wnt3a and
downstream proto-oncogene c-myc protein expression was
significantly decreased.
The anticancer activity of MS4322 is not restricted
to cervical cancer cells; prior studies have shown that this
compound effectively decreases PRMT5 protein expression and
inhibits proliferation in multiple cancer cell lines, including
MCF7 (breast cancer), A549 (lung adenocarcinoma), A172
(glioblastoma), and Jurkat (leukemia) cells (23,24).
However, the mechanism of MS4322 underlying inhibiting the activity
of Wnt/β-catenin signaling pathway remains to be investigated. The
present study was conducted using the HeLa cell line. Future
studies should validate the function of PRMT5 and the anticancer
efficacy of MS4322 in other cervical cancer cell models or primary
cells, thereby enhancing the generalizability and clinical
relevance of the findings. Nevertheless, the present study
demonstrated the inhibitory effect of MS4322 on HeLa cells and its
potential mechanisms, providing novel insight into the pathogenesis
of cervical cancer and offering promising new targets and
strategies for its clinical diagnosis and treatment.
In conclusion, the present study found that PRMT5
showed high expression in cervical cancer tissues, PRMT5 promoted
the migration and invasion of HeLa cervical cancer cells, while
knockdown of PRMT5 inhibited these malignant phenotypes and induced
cell cycle arrest. Furthermore, the PRMT5 degrader MS4322 exerted
anti-tumor effects by targeting PRMT5, thereby suppressing the
malignant biological behaviors of cervical cancer HeLa cells in
vitro.
Acknowledgements
Not applicable.
Funding
The present study was funded by the Key Research and Development
Program of Linyi (Medical Category) (grant no. 2026YX0066).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
RL designed the study, performed experiments, and
wrote the manuscript. JC performed experiments and data analysis.
ZC and SW contributed to data analysis and participated in data
discussion. TL interpreted data interpretation, participated in
data discussion, and critically revised the manuscript. YX
conceived the study conception, experimental design, and critical
revision of the manuscript. RL and YX confirm the authenticity of
all the raw data. All authors read and approved the final version
of the manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Linyi People's Hospital, Linyi, China (approval no.
202306-H-093) and performed in accordance with the Declaration of
Helsinki. All participants provided written informed consent prior
to sample collection.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Wilailak S, Kengsakul M and Kehoe S:
Worldwide initiatives to eliminate cervical cancer. Int J Gynaecol
Obstet. 155 (Suppl 1):S102–S106. 2021. View Article : Google Scholar
|
|
2
|
Yadav G, Srinivasan G and Jain A: Cervical
cancer: Novel treatment strategies offer renewed optimism. Pathol
Res Pract. 254:1551362024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Voelker RA: Cervical cancer screening.
JAMA. 330:20302023. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zheng J, Li B, Wu Y, Wu X and Wang Y:
Targeting arginine methyltransferase PRMT5 for cancer therapy:
Updated progress and novel strategies. J Med Chem. 66:8407–8427.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Blanc RS and Richard S: Arginine
methylation: The coming of age. Mol Cell. 65:8–24. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Rasheed S, Bouley RA, Yoder RJ and
Petreaca RC: Protein arginine methyltransferase 5 (PRMT5) mutations
in cancer cells. Int J Mol Sci. 24:60422023. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Stopa N, Krebs JE and Shechter D: The
PRMT5 arginine methyltransferase: Many roles in development, cancer
and beyond. Cell Mol Life Sci. 72:2041–2059. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kaganovski A, Smith-Salzberg B, Shimshon
HK, Draheim A, Spivak M, Sapir T and Shifteh D: Current and
emerging therapies for targeting protein arginine
methyltransferases (PRMTs) in cancer. Int J Mol Sci. 26:79072025.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ozturk H, Seker-Polat F, Abbaszadeh N,
Kingham Y, Orsulic S and Adli M: High PRMT5 levels, maintained by
KEAP1 inhibition, drive chemoresistance in high-grade serous
ovarian cancer. J Clin Invest. 135:e1842832025. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Bray C, Balcells C, McNeish IA and Keun
HC: The potential and challenges of targeting MTAP-negative cancers
beyond synthetic lethality. Front Oncol. 13:12647852023. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Bao X, Zhao S, Liu T, Liu Y, Liu Y and
Yang X: Overexpression of PRMT5 promotes tumor cell growth and is
associated with poor disease prognosis in epithelial ovarian
cancer. J Histochem Cytochem. 61:206–221. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang Z, Li R, Hou N, Zhang J, Wang T, Fan
P, Ji C, Zhang B, Liu L, Wang Y, et al: PRMT5 reduces immunotherapy
efficacy in triple-negative breast cancer by methylating KEAP1 and
inhibiting ferroptosis. J Immunother Cancer. 11:e0068902023.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhou B, Chen N, Chen Z, Chen S, Yang J,
Zheng Y and Shen L: Prmt5 deficient mouse B cells display RNA
processing complexity and slower colorectal tumor progression. Eur
J Immunol. 53:e22502262023. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Qiao Y, Wu Z, Wang P, Jin Y, Bai F, Zhang
F, An Y, Xue M, Feng H, Zhang Y, et al: C6orf223 promotes
colorectal cancer growth and metastasis by facilitating PRMT5-MEP50
multiprotein complex assembling. J Clin Invest. 135:e1860522025.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Gao J, Liu R, Feng D, Huang W, Huo M,
Zhang J, Leng S, Yang Y, Yang T, Yin X, et al: Snail/PRMT5/NuRD
complex contributes to DNA hypermethylation in cervical cancer by
TET1 inhibition. Cell Death Differ. 28:2818–2836. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Soeta T, Sugisawa N, Yamamura A, Tanaka N,
Imoto H, Tsuchiya T, Aizawa T, Okamoto K, Kawamura M, Saijo F, et
al: MRTX1719, an MTA-cooperative PRMT5 inhibitor, induces cell
cycle arrest and synergizes with oxaliplatin and gemcitabine for
enhanced anticancer effects. Anticancer Res. 44:5231–5240. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Debnath J, Gammoh N and Ryan KM: Autophagy
and autophagy-related pathways in cancer. Nat Rev Mol Cell Biol.
24:560–575. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li M, Gao Z, Wang S, Zhao Y and Xie H:
miR-27a-3p upregulation by p65 facilitates cervical tumorigenesis
by increasing TAB3 expression and is involved in the positive
feedback loop of NF-κB signaling. Oncol Rep. 50:1322023. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li Y, Meng F, Sui C, Wang Y and Cheng D:
CircRNA hsa_circ_0001627 aggravates cervical cancer progression
through upregulation of FNDC3B and activating PI3K/mTOR signaling
pathway. J Cell Commun Signal. 17:627–638. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Yu F, Yu C, Li F, Zuo Y, Wang Y, Yao L, Wu
C, Wang C and Ye L: Wnt/β-catenin signaling in cancers and targeted
therapies. Signal Transduct Target Ther. 6:3072021. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Wang B, Li X, Liu L and Wang M: β-Catenin:
Oncogenic role and therapeutic target in cervical cancer. Biol Res.
53:332020. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Adiga D, Bhat S, Chakrabarty S and
Kabekkodu SP: DOC2B is a negative regulator of Wnt/β-catenin
signaling pathway in cervical cancer. Pharmacol Res.
180:1062392022. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shen Y, Gao G, Yu X, Kim H, Wang L, Xie L,
Schwarz M, Chen X, Guccione E, Liu J, et al: Discovery of
first-in-class protein arginine methyltransferase 5 (PRMT5)
degraders. J Med Chem. 63:9977–9989. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Guo Y, Li Y, Zhou Z, Hou L, Liu W, Ren W,
Mi D, Sun J, Dai X, Wu Y, et al: Targeting PRMT5 through PROTAC for
the treatment of triple-negative breast cancer. J Exp Clin Cancer
Res. 43:3142024. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Brown T, Nguyen T, Zhou B and Zheng YG:
Chemical probes and methods for the study of protein arginine
methylation. RSC Chem Biol. 4:647–669. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Abe Y, Sano T and Tanaka N: The Role of
PRMT5 in immuno-oncology. Genes (Basel). 14:6782023. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Koskas M, Amant F, Mirza MR and Creutzberg
CL: Cancer of the corpus uteri: 2021 update. Int J Gynaecol Obstet.
155 (Suppl 1):S45–S60. 2021. View Article : Google Scholar
|
|
28
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Katsaounis D, Chaplain MAJ and Sfakianakis
N: Stochastic differential equation modelling of cancer cell
migration and tissue invasion. J Math Biol. 87:82023. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Li D, Xia L, Huang P, Wang Z, Guo Q, Huang
C, Leng W and Qin S: Heterogeneity and plasticity of
epithelial-mesenchymal transition (EMT) in cancer metastasis:
Focusing on partial EMT and regulatory mechanisms. Cell Prolif.
56:e134232023. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Hänggi K and Ruffell B: Cell death,
therapeutics, and the immune response in cancer. Trends Cancer.
9:381–396. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Tang Y, Dong L, Zhang C, Li X, Li R, Lin
H, Qi Y, Tang M, Peng Y, Liu C, et al: PRMT5 acts as a tumor
suppressor by inhibiting Wnt/β-catenin signaling in murine gastric
tumorigenesis. Int J Biol Sci. 18:4329–4340. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Elzeiny N, Sayed Shafei AE, Wagih S, Saad
M, Sayed D, Salem EY, Wael M, Ellackany R and Matboli M:
Phytochemicals in cervical cancer: An epigenetic overview.
Epigenomics. 15:941–959. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Feng Q, Cui N, Li S, Cao J, Chen Q and
Wang H: Upregulation of SOX9 promotes the self-renewal and
tumorigenicity of cervical cancer through activating the
Wnt/β-catenin signaling pathway. FASEB J. 37:e231742023. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Satari M, Javani Jouni F, Abolmaleki P and
Soleimani H: Influence of static magnetic field on HeLa and Huo2
cells in the presence of aloe vera extract. Asian Pac J Cancer
Prev. 22:9–15. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Mata-Rocha M, Rodríguez-Hernández RM,
Chávez-Olmos P, Garrido E, Robles-Vázquez C, Aguilar-Ruiz S,
Torres-Aguilar H, González-Torres C, Gaytan-Cervantes J,
Mejía-Aranguré JM and Romero-Tlalolini MLA: Presence of HPV DNA in
extracellular vesicles from HeLa cells and cervical samples. Enferm
Infecc Microbiol Clin (Engl Ed). 38:159–165. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Guimarães YM, Godoy LR, Longatto-Filho A
and Reis RD: Management of early-stage cervical cancer: A
literature review. Cancers (Basel). 14:5752022. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Li Q, Sun H, Liu S, Tang J, Liu S, Yin P,
Mi Q, Liu J, Yu L and Bi Y: Ginsenoside Rk1 inhibits HeLa cell
proliferation through an endoplasmic reticulum signaling pathway. J
Ginseng Res. 47:645–653. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Li D, Peng X, Hu Z, Li S, Chen J and Pan
W: Small molecules targeting selected histone methyltransferases
(HMTs) for cancer treatment: Current progress and novel strategies.
Eur J Med Chem. 264:1159822024. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Micallef I, Fenech K and Baron B:
Therapeutic targeting potential of the protein lysine and arginine
methyltransferases to reverse cancer chemoresistance. Front Mol
Biosci. 11:14554152024. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Sapir T, Shifteh D, Pahmer M, Goel S and
Maitra R: Protein arginine methyltransferase 5 (PRMT5) and the
ERK1/2 & PI3K pathways: A case for PRMT5 inhibition and
combination therapies in cancer. Mol Cancer Res. 19:388–394. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Liu A, Yu C, Qiu C, Wu Q, Huang C, Li X,
She X, Wan K, Liu L, Li M, et al: PRMT5 methylating SMAD4 activates
TGF-β signaling and promotes colorectal cancer metastasis.
Oncogene. 42:1572–1584. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Zhu Y, Xia T, Chen DQ, Xiong X, Shi L, Zuo
Y, Xiao H and Liu L: Promising role of protein arginine
methyltransferases in overcoming anti-cancer drug resistance. Drug
Resist Updat. 72:1010162024. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Antonysamy S, Bonday Z, Campbell RM, Doyle
B, Druzina Z, Gheyi T, Han B, Jungheim LN, Qian Y, Rauch C, et al:
Crystal structure of the human PRMT5:MEP50 complex. Proc Natl Acad
Sci USA. 109:17960–17965. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Motolani A, Martin M, Sun M and Lu T: The
structure and functions of PRMT5 in human diseases. Life (Basel).
11:10742021.PubMed/NCBI
|
|
46
|
Huang L, Zhang XO, Rozen EJ, Sun X, Sallis
B, Verdejo-Torres O, Wigglesworth K, Moon D, Huang T, Cavaretta JP,
et al: PRMT5 activates AKT via methylation to promote tumor
metastasis. Nat Commun. 13:39552022. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Barczak W, Jin L, Carr SM, Munro S, Ward
S, Kanapin A, Samsonova A and La Thangue NB: PRMT5 promotes cancer
cell migration and invasion through the E2F pathway. Cell Death
Dis. 11:5722020. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Xie F, Zhang H, Zhu K, Jiang CS, Zhang X,
Chang H, Qiao Y, Sun M, Wang J, Wang M, et al: PRMT5 promotes
ovarian cancer growth through enhancing Warburg effect by
methylating ENO1. MedComm (2020). 4:e2452023. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Zheng Y, Ji H, Yi W, Chen Z, Hu X, Zhou J,
Wang Y and Zheng X: PRMT5 facilitates angiogenesis and EMT via
HIF-1α/VEGFR/Akt signaling axis in lung cancer. Aging (Albany NY).
15:6163–6178. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Lee J, Kim J and Hwang I: Targeting PRMT5
in cancer: Mechanistic insights and clinical progress. Biomed
Pharmacother. 193:1187542025. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Gu Z, Kong W, Liu X, Hu L, Zhou Y, Liang
Z, Zhang M, Chen D, Li F and Chen W: Drug discovery targeting
protein arginine methyltransferase 5 (PRMT5): An update. Bioorg Med
Chem. 128:1182402025. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Brown-Burke F, Zhang Y, Hinterschied C,
Prouty A, Helmig-Mason J, Chung JH, Vaddi K, Chen-Kiang S, Di
Liberto M, Alinari L, et al: PRMT5 inhibition drives therapeutic
vulnerability to BCL-2 inhibition with venetoclax and provides
rationale for combination therapy in mantle cell lymphoma. Blood.
134 (Suppl 1):S3022019. View Article : Google Scholar
|
|
53
|
Wang N, Yan H, Wu D, Zhao Z, Chen X, Long
Q, Zhang C, Wang X, Deng W and Liu X: PRMT5/Wnt4 axis promotes
lymph-node metastasis and proliferation of laryngeal carcinoma.
Cell Death Dis. 11:8642020. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Brown-Burke F, Hwang I, Sloan S,
Hinterschied C, Helmig-Mason J, Long M, Chan WK, Prouty A, Chung
JH, Zhang Y, et al: PRMT5 inhibition drives therapeutic
vulnerability to combination treatment with BCL-2 inhibition in
mantle cell lymphoma. Blood Adv. 7:6211–6224. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Celià-Terrassa T and Kang Y: How important
is EMT for cancer metastasis? PLoS Biol. 22:e30024872024.
View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Morana O, Wood W and Gregory CD: The
apoptosis paradox in cancer. Int J Mol Sci. 23:13282022. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Kulbay M, Paimboeuf A, Ozdemir D and
Bernier J: Review of cancer cell resistance mechanisms to apoptosis
and actual targeted therapies. J Cell Biochem. 123:1736–1761. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Guo A, Chang Y, Lin J, Guo J, He Y, Wang
C, Wu Z, Xing Y, Jin F and Deng Y: Resveratrol enhances anticancer
effects of silybin on HepG2 cells and H22 tumor-bearing mice via
inducing G2/M Phase arrest and increasing Bax/Bcl-2 ratio. Comb
Chem High Throughput Screen. 28:89–98. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Güven CM and Özgür A: BIIB021, an orally
available and small-molecule inhibitor of HSP90, activates
intrinsic apoptotic pathway in human cervical adenocarcinoma cell
line (HeLa). Eur Rev Med Pharmacol Sci. 27:7299–7308.
2023.PubMed/NCBI
|
|
60
|
Liang XR, Liu YF, Chen F, Zhou ZX, Zhang
LJ and Lin ZJ: Cell cycle-related lncRNAs as innovative targets to
advance cancer management. Cancer Manag Res. 15:547–561. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Wu X, Que H, Li Q and Wei X: Wnt/β-catenin
mediated signaling pathways in cancer: Recent advances, and
applications in cancer therapy. Mol Cancer. 24:1712025. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Su M, Shan S, Gao Y, Dai M, Wang H, He C,
Zhao M, Liang Z, Wan S, Yang J and Cai H: 2-Deoxy-D-glucose
simultaneously targets glycolysis and Wnt/β-catenin signaling to
inhibit cervical cancer progression. IUBMB Life. 75:609–623. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Chen X, Jiang M, Zhou S, Chen H, Song G,
Wu Y and Zhu X: PRAME promotes cervical cancer proliferation and
migration via Wnt/β-catenin pathway regulation. Cancers (Basel).
15:18012023. View Article : Google Scholar : PubMed/NCBI
|