Introduction
High mobility group (HMG) proteins, isolated
initially from bovine thymus in 1973, are named according to their
high electrophoretic mobility in polyacrylamide gels (1). The HMG family constitutes the second
most abundant family of chromatin-associated proteins in the cell
nucleus after histones. Based on structural characteristics, it
consists of three major subfamilies: HMGA, HMGB and HMGN. HMGB1,
HMGB2, HMGB3 and HMGB4 are the four major members of the HMGB
subfamily. The former three members share high structural homology,
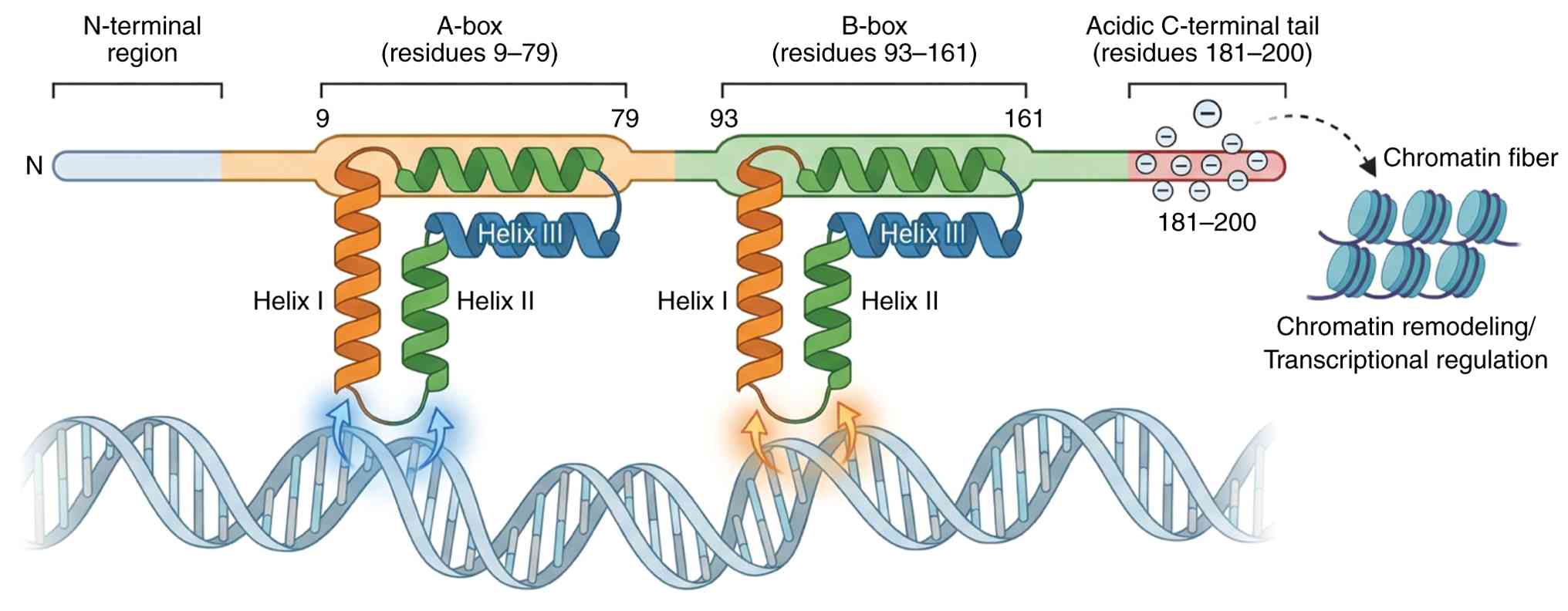
each containing two tandem HMG-box domains (A- and B-box) and an
acidic C-terminal tail, as illustrated in Fig. 1. These proteins can bind and bend
DNA, in turn being involved in diverse cellular processes such as
DNA repair, replication, recombination and transcription (2–4).
Critically, HMGB3 exhibits distinct spatiotemporal
specificity, unlike the widely expressed HMGB1 and the more
restricted HMGB2. Specifically, it may exhibit typical expression
patterns, manifesting as high expression during embryonic
development that is critical for maintaining stem cell pluripotency
and self-renewal, while it is nearly silenced in most
differentiated adult tissues (5–7).
Therefore, the reactivation of HMGB3 may have an intimate
association with cellular dedifferentiation and malignant
transformation. Indeed, HMGB3 is aberrantly overexpressed in
multiple cancers, such as breast cancer (8), ovarian cancer (9), lung cancer (10), colorectal cancer (CRC) (11) and bladder cancer (12), indicating a certain correlation with
poor prognosis simultaneously. In addition to driving malignant
progression, this overexpression may be linked to therapy
resistance, particularly to chemotherapy and tumor recurrence. For
instance, HMGB3 is considered a key indicator in relapse and
treatment failure of acute myeloid leukemia (AML) (13). Its knockdown in gastric cancer cells
may inhibit cell proliferation and migration, and affect
chemosensitivity (14). HMGB3 may
also promote the repair of interstrand DNA crosslinks and
double-strand breaks (DSBs), suggesting its potential role in
resistance to DNA-damaging chemotherapeutic agents (3). In addition, this member may also be
implicated in non-neoplastic diseases, including ulcerative colitis
(15), silica-induced pulmonary
inflammation (16) and myocardial
infarction (17).
With respect to the above, the present work
systematically reviews the multifaceted roles of HMGB3 in driving
resistance to cancer therapy and explores its underlying molecular
mechanisms. Based on the examination of its functions in cancer
cell proliferation, invasion, metastasis and cancer stem cell (CSC)
maintenance, this study continues to integrate its roles in DNA
damage repair and autophagy, with the purpose to uncover its
mechanism in synergistically promoting multidrug resistance through
multiple pathways (14,18). Additionally, the clinical potential
of HMGB3 as a diagnostic biomarker and therapeutic target was
discussed, thereby offering novel strategies for overcoming
resistance to cancer therapy. Overall, HMGB3 exhibits complicated
expression and function across different cancers, and tumor
heterogeneity may exert an impact on treatment response (19,20).
Therefore, in the future, our understanding of the specific
mechanisms of HMGB3 within a distinct tumor microenvironment (TME)
should be refined to facilitate the development of targeted
inhibitors or regulatory strategies, thereby advancing precision
oncology.
Transcriptional and epigenetic regulation of
HMGB3
Multi-level molecular mechanisms, covering
transcription factors and epigenetic modifications, are involved in
the co-regulation of the dysregulated HMGB3 expression in cancer,
eventually forming an interconnected regulatory network (21). The expression of HMGB3, acting as a
chromatin-associated protein, is directly regulated by the binding
of multiple transcription factors. In gastric cancer, nine
transcription factors, including GATA1/2/3 and MZF1, can directly
bind to the promoter or enhancer regions of HMGB3, developing a
‘HMGB3-target gene’ axis that may further promote tumor progression
(22). In AML, its expression may
also be modulated by disease-specific transcription factors
(13). Epigenetically, HMGB3
expression is shaped by histone modifications, non-coding RNAs
(ncRNAs) and DNA methylation. For instance, the histone H3 lysine
27 trimethylation modification can be catalyzed by enhancer of
zeste homolog 2 in glioma, further suppressing microRNA
(miR)-142-3p, thereby relieving its inhibition of the long (l)ncRNA
KCNQ1OT1, consequently enhancing the expression of HMGB3 by binding
to LIN28B (23). Additional
investigations on DNA methylation analysis support a negative
correlation between DNA methylation and HMGB3 expression, as
hypomethylation of the HMGB3 gene is frequent in tumors with HMGB3
upregulation, while its hypermethylation is associated with its
downregulation (24–26).
Notably, HMGB3 also functionally synergizes with
other transcription factors. In prostate cancer, the HMGB3 A-box
domain can interact with SOX9, resulting in a co-activation of
NANOG homeobox (NANOG), thereby enhancing tumor stemness and
progression (27). Collectively,
HMGB3 is critical within a complex regulatory network in cancer.
Its expression can be dynamically balanced by direct
transcriptional regulation and epigenetic modifications, ultimately
determining its functional output in tumorigenesis and
progression.
Post-transcriptional regulation
HMGB3 expression is finely regulated at the
post-transcriptional level by a network of ncRNAs [miRNAs, lncRNAs
and circular (circ)RNAs] through competitive endogenous RNA (ceRNA)
mechanisms. It may eventually affect tumor progression and therapy
resistance significantly, as summarized in Table I.
 | Table I.ncRNA-mediated regulation of HMGB3 in
therapy resistance and apoptosis-associated tumor adaptation. |
Table I.
ncRNA-mediated regulation of HMGB3 in
therapy resistance and apoptosis-associated tumor adaptation.
| Tumor/cancer | CircRNA/lncRNA | Regulatory
axis | Function and
mechanism | (Refs.) |
|---|
| Non-small cell lung
cancer | - | miR-758/HMGB3 | Inhibits the
proliferation, migration and invasion of cancer cells and promotes
their apoptosis. | (10) |
|
| Circ_0060937 |
miR-195-5p/HMGB3 | Inhibits cancer
cell proliferation, migration, invasion and glycolysis and triggers
apoptosis. | (33) |
|
| Circ_0020123 | miR-1299/HMGB3 | Inhibits the
proliferation, migration and invasion of cancer cells and promotes
their apoptosis. | (39) |
|
| - | miR-513b/HMGB3 | Activates the mTOR
signaling pathway and promotes cancer cell proliferation, invasion,
migration and apoptosis. | (40) |
| Breast cancer | - |
miR-142-3p/HMGB3 | Modulates autophagy
and induces apoptosis through reactive oxygen species accumulation
and mitochondrial dysfunction. | (29) |
|
| - | miR-27b/HMGB3 | Enhances cancer
cell sensitivity to tamoxifen, inhibits invasion and reverses
epithelial to mesenchymal transition-like phenotypes. | (41) |
|
| - |
miR-381-3p/HMGB3 | Exosome-delivered
OIP5-AS1 confers resistance to trastuzumab in cancer cells via its
competing endogenous RNA mechanism. | (42) |
| CRC | - |
miR-200b-3p/HMGB3 | The loss of
exosomal miR-200b-3p in hypoxic cancer-associated fibroblasts
reduces sensitivity to 5-fluorouracil in CRC by targeting
HMGB3. | (38) |
|
| - | miR-429/HMGB3 | Inhibits cancer
cell proliferation and induces apoptosis. | (43) |
|
| - | miR-93/HMGB3 | Inhibits the
activation of the PI3K/AKT pathway and promotes apoptosis in CRC
cells. | (44) |
| Gastric cancer | - | miR-513b/HMGB3 | Inhibits cancer
cell proliferation and migration, and promotes their
apoptosis. | (45) |
| Prostate
cancer | LncRNA SOX2-OT |
miR-452-5p/HMGB3 | Inactivates the
Wnt/β-catenin pathway and inhibits the proliferation and metastasis
of prostate cancer cells. | (46) |
| Ovarian cancer | - |
miRNA-374b-5p/HMGB3 | Downregulates the
Wnt/β-catenin pathway axis and inhibits the malignant progression
of epithelial ovarian cancer. | (47) |
| Cervical
cancer | LncRNA BCYRN1 |
miR-330-5p/HMGB3 | Disrupts the
cisplatin resistance of cancer cells. |
|
|
| - | miR-758/HMGB3 | Downregulates the
Wnt/β-catenin signaling pathway and inhibits cancer cell
proliferation and metastasis. | (48,49) |
| Esophageal
cancer | - | miR-216a/HMGB3 | Activates the
Wnt/β-catenin pathway and promotes the progression of esophageal
cancer. | (50) |
Multiple miRNAs (e.g., miR-101-5p, miR-142-3p,
miR-205 and miR-758) may inhibit HMGB3 expression by directly
targeting the corresponding 3′UTR. In this regard, it may
effectively suppress cancer cell proliferation, migration and
invasion in various tumor types, such as breast cancer, cervical
cancer and non-small cell lung cancer (NSCLC) (12,28–30).
In particular, beyond targeting HMGB3 directly, miR-142-3p can also
induce mitochondrial dysfunction and promote apoptosis by
inactivating the mTOR/STAT3 pathway (29).
In addition to direct regulation, lncRNAs (e.g.,
SNHG5, HOTTIP) and circRNAs (e.g., circFOXO3, circIGF1R,
circN4BP2L2) can sequester corresponding miRNAs to upregulate HMGB3
expression directly, which may further promote the progression of
malignancies such as nasopharyngeal carcinoma (NPC), NSCLC and CRC
(31–37). In CRC, circIGF1R and circN4BP2L2 can
remarkably stimulate tumor cell proliferation, invasion and
glycolysis while inhibiting apoptosis through the activation of the
HMGB3/Wnt/β-catenin signaling pathway (36,37).
Under hypoxic conditions, downregulation of miR-200b-3p in
cancer-associated fibroblast-derived exosomes can attenuate the
suppression of HMGB3, in turn weakening the sensitivity of CRC
cells to 5-fluorouracil (5-FU)-based chemotherapy (38).
Altogether, the multi-level regulation of HMGB3
expression by ncRNAs through ceRNA networks is essential in
malignant progression and therapy resistance. Additional
representative ncRNA-mediated regulatory axes involving HMGB3
across different cancers, including NSCLC, breast cancer,
colorectal cancer, gastric cancer, and cervical cancer, are
summarized in Table I (10,29,33,38–50).
Post-translational modifications
(PTMs) of HMGB3 protein
Critically, despite less advanced research compared
to that on HMGB1 and HMGB2, the function of HMGB3 from the HMG
family may be dynamically regulated by PTMs (2,51,52).
The current understanding of the PTMs of HMGB3 relies largely on
homology-based inference. In particular, these modifications may
play a role in cancer therapy resistance by affecting subcellular
localization, DNA damage response (DDR) and extracellular release
under stress conditions.
In HMGB1, acetylation and phosphorylation act as
crucial regulators of nuclear-cytoplasmic trafficking and
participation in DNA repair. Following these modifications, there
may be reduced chromatin-binding stability, which may further boost
the exposure of DNA damage sites and recruitment of repair factors,
while driving nuclear export and extracellular release under
conditions of severe stress, thereby influencing inflammatory
signaling and cell survival (53–63).
Similarly, the PTM status of HMGB3 may modulate DNA damage
tolerance, replication stress response and post-chemotherapy or
post-radiotherapy cell survival, considering that it retains highly
conserved DNA-binding domains and regulatory regions (2). Furthermore, HMGB proteins may have a
relationship with therapy resistance, given the redox regulation.
To be specific, the oxidation state of cysteine residues in HMGB1
can determine the functional switch from a nuclear DNA chaperone to
an extracellular signaling molecule, a process that is highly
relevant under radiotherapy- or chemotherapy-induced oxidative
stress (53,64–67).
With the presence of conserved cysteine residues as well, HMGB3 may
also have similar stress-induced functional reprogramming,
potentially affecting DNA repair efficiency and TME-associated
inflammatory responses, both of which are recognized to be key
components of acquired therapy resistance (68). Additionally and significantly, there
is so far limited research on the methylation and glycosylation of
HMGB3. Furthermore, existing evidence indicates a negative
correlation of HMGB3 DNA methylation with its expression in tumors
(24). Despite the scarcity of
direct experimental evidence, at the protein level, HMGB3 activity
may be further modulated by lysine methylation and potential
glycosylation sites within the B-box domain (69–72).
Collectively, even with limited direct experimental
evidence for HMGB3 PTMs, PTMs may modulate HMGB3 activity during
DDR and therapy-induced stress, as evidenced by homology- and
structure-based functional insights. Consequently, it may
participate in the modulation of tumor cell tolerance to
radiotherapy, chemotherapy and targeted therapies potentially
(2,51–53).
Core functional mechanisms of HMGB3:
Bridging cancer stemness and treatment resistance
By coordinating multiple critical biological
processes, HMGB3 can drive malignant progression and therapeutic
resistance, thus forming an integrated regulatory network. These
may include maintenance of CSC properties, modulation of the DDR,
regulation of apoptosis-autophagy balance and remodeling of the
TME. Therefore, these mechanisms may constitute the molecular
foundation for resistance across diverse therapeutic modalities, as
depicted in Fig. 2.
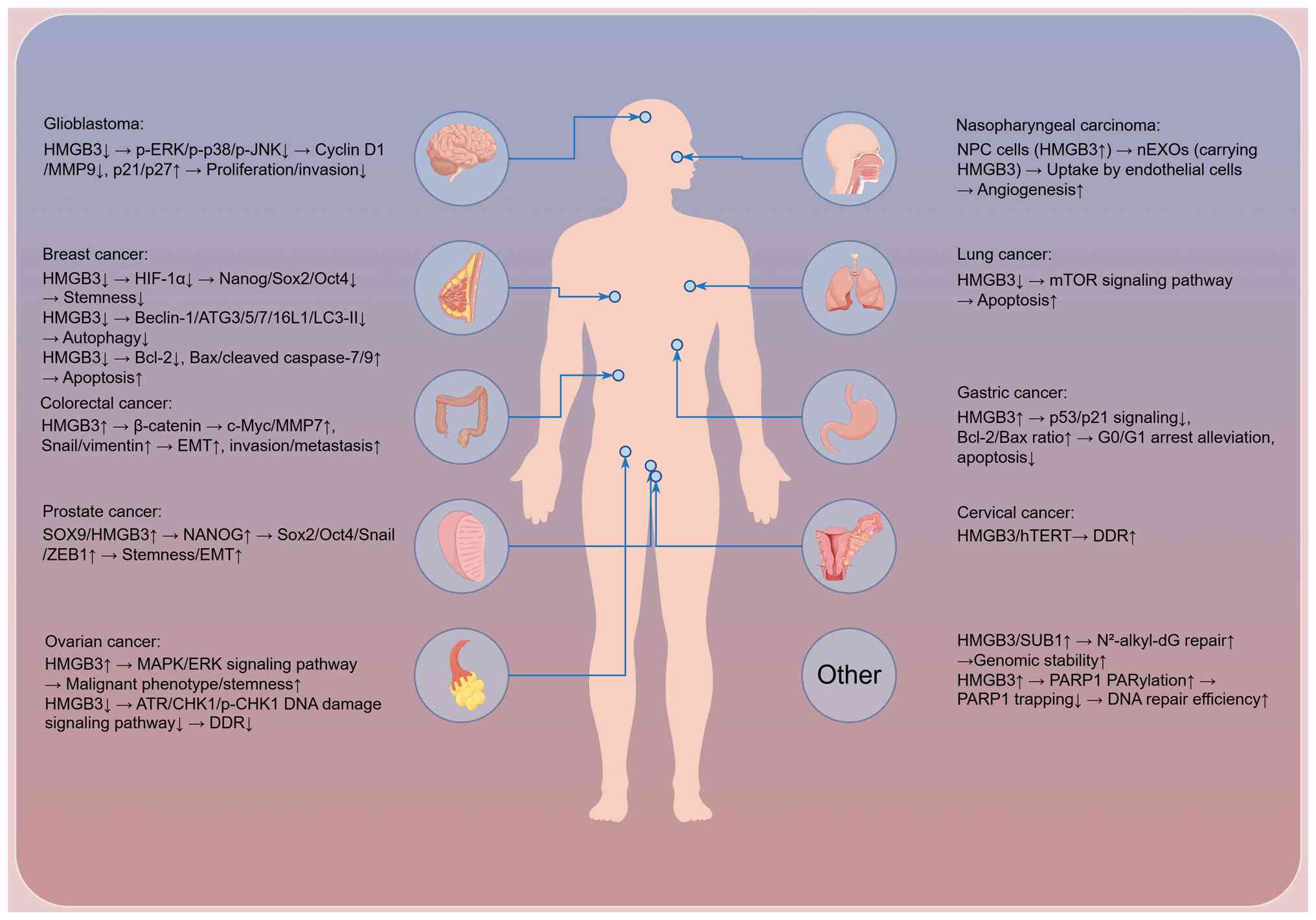 | Figure 2.HMGB3 serving as a central driver of
tumor malignancy. This schematic illustrates the mechanisms of
HMGB3 in coordinating a multi-dimensional regulatory network,
driving tumor progression and therapy resistance by sustaining
cancer stemness, modulating DDR, inhibiting apoptosis, inducing
autophagy and remodeling the tumor microenvironment. HMGB, high
mobility group box; EMT, epithelial to mesenchymal transition; HIF,
hypoxia-inducible factor; DDR, DNA damage response; p-CHK,
phosphorylated checkpoint kinase; Nanog, Nanog homeobox; Oct4,
octamer-binding transcription factor 4; ZEB1, Zinc finger E-box
binding homeobox 1; ATR, Ataxia telangiectasia and Rad3-related;
hTERT, human telomerase reverse transcriptase; SUB1, SUB1 regulator
of transcription; PARP1, poly(ADP-ribose) polymerase 1. |
Maintenance of CSC properties
HMGB3 is a chromatin-associated protein
overexpressed in multiple cancers, establishing an intimate
association with CSC characteristics. CSCs represent a
subpopulation of tumor cells that can undergo self-renewal and
differentiation, exhibiting close relationships with tumor
recurrence, metastasis and intrinsic therapeutic resistance
(73–75).
Initially, HMGB3 was found to be preferentially
expressed in hematopoietic stem cells, where it balances
self-renewal and differentiation. For example, there are reduced
common lymphoid progenitors and common myeloid progenitors in
HMGB3-deficient mice, suggesting its potential role in maintaining
primitive stem cell properties (76). Importantly, the proposed
stemness-maintaining function appears to be conserved in
malignancies. In solid tumors, HMGB3 can enhance spheroid
formation, colony formation and the expression of
pluripotency-associated transcription factors (e.g., NANOG, SOX2
and OCT4) to promote the formation of CSC-like phenotypes (27,77–79).
In ovarian and breast cancer models, HMGB3 overexpression may
indicate increased expression of stemness markers and unfavorable
prognosis (77–80). Conversely, HMGB3 silencing can
reduce self-renewal capacity and impair tumorigenicity. Its
mechanisms may be related to the regulation of multiple
stemness-associated signaling pathways. It can improve the
expression of downstream targets [e.g., c-Myc and MMP7] by boosting
β-catenin nuclear accumulation and transcriptional activity. Its
deficiency may also increase the level of E-cadherin while reducing
that of mesenchymal markers (e.g., Snail and vimentin), indicating
its role in driving epithelial-mesenchymal transition (EMT) and
reinforcing invasive phenotypes (21,27,81,82).
In addition, HMGB3 may also activate MAPK/ERK signaling and
cooperate with transcriptional regulators (e.g., SOX9) to enhance
the transcription of NANOG, thereby stabilizing the pluripotency
network (80,83–85).
HMGB3 can activate Wnt/β-catenin, MAPK/ERK and
related regulatory axes collaboratively, thereby sustaining the CSC
pool and enhancing tumor adaptability under therapeutic stress.
Regulation of the DDR pathway
Genomic instability is a hallmark of cancer.
Sustained activation of DDR pathways may enable tumor cells to
survive under endogenous and exogenous genotoxic stress. HMGB3 is a
chromatin-associated architectural protein that may contribute to
DDR regulation at multiple functional levels, including damage
sensing, chromatin remodeling, transcriptional control of repair
genes and coordination of complex DNA lesion repair.
At the early stage of the DDR, rapid lesion
recognition and chromatin relaxation constitute the premise of
efficient repair to permit the recruitment of repair complexes.
Depending on its HMG box domains, HMGB3 can bind distorted DNA
structures and modulate chromatin conformation, thereby
facilitating access of repair machinery to damaged sites. HMGB3 can
recognize specific DNA adducts, such as
N2-alkyl-2′-deoxyguanosine (dG) lesions induced by
benzo[a]pyrene-7,8-dihydrodiol-9,10-epoxide (BPDE), and, together
with SUB1 regulator of transcription, functions as a damage sensor
exhibiting stereoselective repair activity toward
trans-N2-BPDE-dG adducts (86). Thus, HMGB3 exerts potential roles in
early lesion recognition and damaged DNA stabilization. Following
the recognition of damage, HMGB3 can further promote repair
progression by interacting with key repair enzymes. Through a
direct interaction with poly(ADP-ribose) polymerase 1 (PARP1), it
can enhance its PARylation activity and potentially influence its
retention at DNA damage sites (9,87). By
modulating PARP1 activity, HMGB3 can support efficient repair of
single-strand breaks and alkylation-induced lesions. Beyond its
structural role, HMGB3 can transcriptionally regulate major DDR
components. By binding to its promoter, it can activate human
telomerase reverse transcriptase (hTERT), resulting in telomere
maintenance and genomic stability under stress (88,89).
HMGB3 can also activate the transcription of ATR checkpoint kinase
(ATR) and checkpoint kinase 1 (CHK1) (90–92),
which may further strengthen checkpoint signaling, replication fork
stabilization and homologous recombination repair. Consistently,
the depletion of HMGB3 may attenuate ATR/CHK1 signaling and impair
the efficiency of interstrand crosslink repair (8). These coordinated functions converge on
the repair of interstrand crosslinks (ICLs) and DSBs, two complex
DNA lesions. However, its deficiency may impair the deficiency of
ICL repair (3,93) and DSB repair (3,94,95).
Accordingly, HMGB3 can facilitate the recruitment or stabilization
of core repair factors involved in homologous recombination and
non-homologous end joining.
Collectively, HMGB3 works to maintain genomic
stability by integrating chromatin remodeling, enzymatic activation
and transcriptional amplification within the DDR network.
Inhibition of apoptosis and induction
of autophagy
HMGB3 can modulate the balance between apoptosis and
autophagy, serving as a critical indicator in tumor cell survival.
There is growing evidence that HMGB3 can suppress apoptotic
signaling while sustaining pro-survival autophagic activity,
thereby promoting cellular adaptation under stress.
In breast cancer, there is an increased expression
of Beclin-1, ATG family proteins and light chain 3-II, supporting
the role of HMGB3 in maintaining protective autophagy (29). Suppression of HMGB3 by miR-142-3p
can disrupt this autophagic program, leading to mitochondrial
dysfunction, featured by the accumulation of reactive oxygen
species (ROS) and the loss of mitochondrial membrane potential.
These alterations may further activate the caspase cascade and
shift cellular fate toward apoptosis, suggesting that HMGB3 can
regulate autophagy through the preservation of mitochondrial
integrity and prevention of cell death. Consistently,
downregulation of HMGB3 can promote apoptotic signaling in multiple
tumor types. For instance, in gastric cancer, reduced HMGB3
expression can induce G0/G1 cell cycle arrest and modulate the
p53/p21 pathway, while decreasing the Bcl-2/Bax ratio to favor
apoptosis (1). Similar HMGB3
silencing-induced pro-apoptotic effects have been reported in NSCLC
(10,40), cervical cancer (96), ovarian cancer (47), CRC (37,44),
thyroid cancer (97) and esophageal
cancer (50), highlighting its
broad role in sustaining tumor cell viability. Despite the scarcity
of direct evidence linking HMGB3 to autophagy regulation, its
structural similarity to HMGB1 suggests potential mechanistic
parallels. HMGB1 can regulate autophagy by interacting with
Beclin-1 (98), highlighting
potential roles of HMGB3 in autophagosome formation or autophagy
flux via related pathways. Of note, context-dependent effects were
also noted. HMGB3 overexpression combined with selinexor can
enhance apoptosis in myelodysplastic syndromes, possibly through
the activation of cytoplasmic DNA-sensing pathways and
interferon-related innate immune signaling (99). It underlines the complexity of
HMGB3-mediated regulation of cell fate across different cellular
contexts.
HMGB3-driven remodeling of the
TME
In addition to intrinsic genetic alterations, tumor
progression may also be determined by dynamic interactions between
malignant cells and their surrounding microenvironment. Therefore,
HMGB3 functions as a critical regulator linking tumor cell
plasticity to microenvironmental remodeling. HMGB3 can regulate
cellular behavior, intercellular communication and oncogenic
signaling networks coordinately, thereby benefiting the reshaping
of the structural and functional landscape of the TME.
Tumor cell plasticity as a driver of
microenvironmental change
HMGB3 can enhance malignant cell plasticity, serving
as a primary force in TME remodeling. Its overexpression may
promote the proliferation and cell cycle progression across
multiple tumor types. In leukemia, HMGB3 may enhance cell
proliferation by activating MAPK/ERK signaling (13). In ovarian cancer, its overexpression
may accelerate cell growth, whereas its silencing can induce G2/M
arrest (80). In prostate cancer,
depletion of HMGB3 may cause G0/G1 arrest by regulating cyclin D1,
p21 and p27 (46). More
importantly, HMGB3 may trigger EMT, a process that fundamentally
alters tumor-stroma interactions. Its upregulation can escalate the
expression of mesenchymal markers (e.g., N-cadherin, vimentin,
β-catenin, snail and slug) (80),
while its silencing may hinder cell migration and invasion in
gastric cancer (14) and CRC
(24). HMGB3 can strengthen EMT to
disrupt tissue architecture, thereby facilitating dynamic cellular
redistribution within the TME. Additionally, HMGB3 can also support
CSC phenotypes. In breast cancer, it can increase mammosphere
formation, while upregulating the expression of Nanog, SOX2 and
OCT-4 (100), with similar effects
observed in ovarian cancer (80).
Maintenance of stem-like subpopulations can contribute to
intratumoral heterogeneity and continuous microenvironmental
adaptation. With respect to the above, HMGB3-driven cellular
plasticity may offer a possible biological foundation for TME
restructuring.
Intercellular communication and
stromal remodeling
Beyond tumor cell-intrinsic changes, HMGB3 may also
play a role in non-malignant components of the TME, among which
angiogenesis represents a well-defined mechanism. In NPC, HMGB3,
secreted via nuclear exosomes, may be internalized by endothelial
cells to promote proliferation and tube formation (101). HMGB3-containing exosomes may
increase microvascular density to facilitate the expansion of the
vascular network within the TME. Furthermore, HMGB3 can modulate
the immune microenvironment. Even with an insufficiency of the
direct mechanistic data, its structural homology to HMGB1-an
established damage-associated molecular pattern-may suggest
potential immunoregulatory functions (18,102,103). In glioblastoma, the overexpression
of HMGB3 may be related to reduced immune cell infiltration and an
immunosuppressive microenvironment (104). Dysregulation of HMGB3 expression
can also amplify ROS generation and activate the NF-κB signaling to
mediate the production of cytokines such as VEGF and IL-6, thereby
shaping the status of inflammation within the TME (1). The remodeling of extracellular matrix
can further induce microenvironmental restructuring. HMGB3 can
regulate MMPs, as evidenced by reduced MMP2 expression following
HMGB3 knockdown in bladder cancer and decreased MMP2/MMP9 activity
in gastric cancer, ultimately altering the physical and biochemical
properties of the TME (1,105).
Integrated signaling networks
underlying TME remodeling
Diverse effects of HMGB3 on TME architecture are
mediated through interconnected oncogenic signaling pathways. HMGB3
can activate the MAPK/ERK signaling in leukemia and ovarian cancer
(13,80), promote β-catenin nuclear
accumulation and downstream target (e.g., MMP7 and c-Myc)
transcription (44,46,80,105),
and modulate PI3K/AKT signaling, as revealed by miR-93-mediated
suppression of HMGB3 in CRC (44).
The interaction between HMGB3 and hypoxia-inducible factor (HIF)-1α
in breast cancer also establish a relationship of HMGB3 to
hypoxia-associated microenvironmental adaptation (100).
By integrating these signaling cascades, HMGB3 can
coordinate malignant cell plasticity, stromal reprogramming and
microenvironmental adaptation, thereby reshaping the organization
and function of TME.
HMGB3-mediated tumor therapy resistance
HMGB3 may induce resistance across multiple
therapeutic modalities. As described above, the present study has
systematically elucidated its roles in DDR, apoptosis regulation
and microenvironmental remodeling. This chapter continues to unveil
the manifestations of these biological functions in specific
therapeutic contexts. Notably, HMGB3-mediated resistance appears to
be treatment-dependent, highlighting the presence of mechanistic
heterogeneity across varied anticancer therapies.
Chemotherapy resistance
Chemotherapeutic agents may produce cytotoxic
effects through diverse mechanisms and HMGB3 has been reported to
modulate resistance in a drug-specific manner. Platinum compounds,
such as cisplatin, can induce cytotoxicity primarily through the
formation of DNA adducts that distort the DNA helix and activate
DNA damage signaling pathways (1,4). HMGB3
is predominantly involved in DNA damage processing, supporting its
implication in platinum resistance. Through potential binding to
cisplatin-DNA adducts, it may facilitate lesion recognition and
subsequent repair (1). In
cisplatin-resistant ovarian cancer models, the inhibition of HMGB3
may enhance drug sensitivity and attenuate the activation of the
ATR/CHK1/p-CHK1 axis, suggesting its role in supporting sustained
DDR signaling under platinum-induced genotoxic stress (8). HMGB3 may also influence the clearance
rate of cisplatin-DNA adducts by interacting with the cisplatin
resistance-associated protein (also known as LUC7L3). HMGB3
knockdown may reduce the efficiency of adduct removal, thereby
altering cellular responses to cisplatin exposure (93). Of note, in gastric cancer cells,
HMGB3 silencing may amplify the sensitivity to cisplatin and
paclitaxel but reduce the sensitivity to oxaliplatin (14,106),
indicating the possible compound- and cellular context-dependent
HMGB3-mediated modulation of platinum response.
Microtubule-targeting agents represent another major class of
chemotherapeutics. For example, paclitaxel can stabilize
microtubules and prevent their depolymerization (107), while vincristine can inhibit
tubulin polymerization and disrupt spindle formation (108). Alterations in cell cycle
regulation and apoptotic signaling thresholds have been proven to
be strongly associated with resistance to these agents. In gastric
cancer, HMGB3 knockdown can suppress cell proliferation, induce
G0/G1 arrest and enhance paclitaxel sensitivity by modulating p53,
p21 and the Bcl-2/Bax ratio (14).
Similarly, in cervical cancer, given reduced IC50 values, its
depletion may enhance the sensitivity to both paclitaxel and
vincristine (88). Collectively,
HMGB3 enables the modulation of cell cycle progression and
apoptosis-related pathways to promote cell survival under mitotic
stress eventually. HMGB3 has been proposed to be associated with
resistance to antimetabolites and endocrine therapies, in addition
to the aforementioned agents. In CRC, by activating Wnt/β-catenin
signaling and EMT-associated transcriptional programs, HMGB3 may
mediate the resistance to 5-FU (109). In breast cancer, HMGB3 expression
is negatively regulated by miR-27b, and its overexpression is
associated with resistance to tamoxifen (41). Currently, although there is still an
incomplete definition of the mechanistic details, there is reason
to believe that HMGB3 exerts a broader role in shaping
chemotherapeutic responsiveness through transcriptional and
signaling reprogramming.
Radiotherapy resistance
Radiotherapy induces cytotoxicity primarily through
the induction of DSBs. Therefore, the efficiency of DNA repair and
the propensity to undergo post-damage apoptosis remain the major
determinants for cellular radiosensitivity. Current evidence
suggests that HMGB3 may induce radioresistance by regulating the
HMGB3/hTERT axis. HMGB3, a transcriptional regulator, can bind to
the hTERT promoter and enhance its expression. Elevated hTERT
levels may be related to increased DNA repair capacity and reduced
radiation-induced apoptosis. Conversely, HMGB3 knockdown may
contribute to accumulated γH2AX foci, impaired DSB repair, and
enhanced radiosensitivity in both in vitro and in
vivo models (88). As a result,
HMGB3 may possibly promote repair competence and restrict apoptotic
execution to sustain radiation tolerance.
Targeted therapy resistance
Targeted therapies frequently encounter resistance
owing to pathway reactivation, compensatory signaling or DNA repair
dynamic alterations. Via both protein-protein interactions and
oncogenic signaling network modulation, HMGB3 appears to
participate in these adaptive processes. Notably, HMGB3 can
interact with PARP1 to affect its functional activity. Loss of
HMGB3 can induce PARP1 ‘trapping’ at sites of DNA damage and weaken
the activity of PARylation, highlighting the regulatory role of
HMGB3 in PARP1 DNA-binding kinetics. Such modulation can
potentially attenuate the cytotoxic effect of olaparib and other
PARP inhibitors (PARPi) (9,110). Besides, given the common function
of the MAPK/ERK pathway as a bypass pathway in targeted therapy
failure, HMGB3-mediated activation of the pathway-reported in
multiple tumor types-may induce adaptive resistance (80,83–85).
However, there is still an insufficiency of direct causal evidence
linking HMGB3 to resistance against specific kinase inhibitors,
necessitating further mechanistic studies to clarify this
relationship.
Immunotherapy resistance
Immune checkpoint inhibitors (ICIs) may disrupt
inhibitory signaling pathways that restrain T-cell activity to
restore antitumor immunity. Emerging evidence indicates that, on
the basis of both tumor-intrinsic signaling alterations and
microenvironmental modulation, HMGB3 may boost immune evasion and
reduce responsiveness to ICIs. In triple-negative breast cancer,
HMGB3 can suppress interferon (IFN)-γ-induced STAT1 phosphorylation
and IFN regulatory factor 1 expression while enhancing STAT3
activation. Concurrently, it can also upregulate
ferroptosis-inhibitory proteins [e.g., solute carrier family 7
member 11 (SLC7A11), glutathione peroxidase 4 and SLC3A2] to
accumulate lipid ROS and restrict IFN-γ-mediated ferroptotic cell
death. These molecular changes are associated with the resistance
to anti-programmed cell death 1 (PD-1) therapy (111). In glioblastoma, elevated
expression of HMGB3 is associated with a non-inflammatory,
immune-excluded microenvironment characterized by reduced immune
cell infiltration (104). Such an
immunologically ‘cold’ tumor phenotype may compromise the efficacy
of ICIs, further implicating HMGB3 in immunotherapy resistance.
Overall, HMGB3 can mediate the resistance to
chemotherapy, radiotherapy, targeted therapy and immunotherapy in a
context-dependent manner. Instead of functioning through a single
dominant mechanism, HMGB3 may enhance tumor survival under
therapeutic pressure by integrating DNA repair regulation, cell
cycle control, apoptotic modulation, signaling pathway activation
and immune adaptation. Nonetheless, in order to determine whether
HMGB3 can serve as a predictive biomarker or actionable therapeutic
target for overcoming treatment resistance, further mechanistic
validation and clinical correlation are required. Representative
HMGB3-mediated mechanisms contributing to resistance to
chemotherapy, radiotherapy, targeted therapy and immunotherapy are
summarized in Table II.
 | Table II.HMGB3-mediated mechanisms of therapy
resistance in cancer. |
Table II.
HMGB3-mediated mechanisms of therapy
resistance in cancer.
| Treatment
modality | Classes of
resistance mechanisms | Core mechanism | Tumor type | (Refs.) |
|---|
| Targeted
therapy | PARPi
resistance | HMGB3/PARP1 | Ovarian cancer | (9) |
| Radiotherapy |
Radioresistance | HMGB3/hTERT | Cervical
cancer | (88) |
| Chemotherapy | Paclitaxel
resistance |
HMGB3/Bcl-2/Bax | Gastric cancer | (14) |
|
|
|
HMGB3/MMP2/MMP9 |
|
|
|
| Tamoxifen
resistance | miR-27b/HMGB3 | Breast cancer | (41) |
|
| Cisplatin/platinum
resistance |
ATR/CHK1/p-CHK1 | Ovarian cancer | (8,93) |
|
|
| CROP/LUC7L3 | Gastric cancer | (106) |
|
|
| miR-200b/HMGB3 |
|
|
|
| 5-FU
resistance |
miR-200b-3p/HMGB3 | Colorectal
cancer | (38) |
|
|
| Wnt/β-catenin | Breast cancer | (109) |
| Immunotherapy | Anti-PD-1/PD-L1
therapy resistance |
HMGB3/IFNγ/STAT1/ferroptosis | Triple-negative
breast cancer | (111) |
HMGB3-targeted therapeutic strategies and
clinical prospects
HMGB3 features high embryonic expression, limited
expression in normal adult tissues and aberrant overexpression in
multiple malignancies (e.g., leukemia, breast cancer, CRC, lung
cancer, glioma), underscoring its potential as an attractive
therapeutic target.
Direct targeting strategies
Targeting HMGB3 directly is the best approach for
precision therapy. Through antisense oligonucleotides (ASOs) or
small interfering RNA (siRNA), the specific silencing of HMGB3 has
been documented to possess significant antitumor activity in
multiple cancer models, which can hinder the proliferation and
invasion while enhancing chemosensitivity (24,80,88).
Such findings provide proof-of-concept for developing ASO- or
siRNA-based HMGB3-targeted therapeutics.
The function of HMGB3, a DNA-binding protein and
transcription factor, depends on specific domains and protein
interactions. Small-molecular compounds developed by targeting its
key functional sites (e.g., the HMGB domain or protein-protein
interaction interfaces) may block its pro-cancer activities. For
example, HMGB3can promote PARPi resistance by interacting with
PARP1 in ovarian cancer (9),
enhance radioresistance by binding the hTERT promoter in cervical
cancer (88) and promote breast
cancer growth by interacting with HIF-1α (100). Developing inhibitors that
specifically block these interactions can precisely suppress
specific oncogenic functions of HMGB3.
HMGB3 may exist extracellularly, although it is
primarily nuclear, offering antibody targets. In NPC, HMGB3 is
secreted via nuclear exosomes, and circulating nuclear exosomes
HMGB3 may link to angiogenesis and metastasis (101), suggesting the potential of
neutralizing antibodies in impeding extracellular HMGB3 functions.
The high expression of HMGB3, even without extracellular activity,
in specific cancers warrants the development of antibody-drug
conjugates targeting cell surface markers associated with HMGB3
expression, thereby enabling precise drug delivery.
Indirect targeting strategies
miRNAs are recognized as post-transcriptional
regulators that may participate in HMGB3 regulation, which can be
modulated to indirectly influence HMGB3 levels. In prostate cancer,
HMGB3 can be negatively regulated by miR-205-5p and its elevated
expression is associated with poor outcomes (112). In breast cancer, by targeting
HMGB3, miR-27b is linked to tamoxifen resistance. Thus, targeting
miR-27b or HMGB3 may reverse the resistance to tamoxifen (109), indicating the miR/HMGB3 axis
serving as a potential therapeutic target (113).
HMGB3 may activate or participate in multiple
oncogenic signaling pathways. In ovarian cancer (92) and leukemia (13), HMGB3 can promote malignancy and
stemness via MAPK/ERK. Significantly, MEK/ERK inhibitors (e.g.,
AZD6244, PD0325901) can effectively reverse HMGB3-induced
pro-cancer effects (80). It may
also promote disease progression by activating pathways such as
Wnt/β-catenin, PI3K/AKT, hypoxia/HIF-1α (44,100,114,115). Inhibiting these pathways
represents an effective indirect strategy targeting HMGB3.
Combination therapy
Given the central role in resistance and stemness,
HMGB3 may be an ideal combination therapy target. Targeted HMGB3
inhibition combined with PARPi can restore sensitivity to PARPi,
thereby overcoming resistance (9).
When combined with chemotherapy, targeting HMGB3 can enhance the
sensitivity to multiple agents (e.g., paclitaxel, cisplatin)
(9,109). While combined with radiotherapy,
it can enhance radiation response by blocking the HMGB3/hTERT axis.
When used jointly with immunotherapy, it may overcome
HMGB3-mediated anti-PD-1 resistance by inhibiting IFN-γ-driven
ferroptosis in triple-negative breast cancer. In addition, HMGB3
can promote the stemness (80) and
CSC-associated EMT in ovarian cancer via modulating the
Wnt/β-catenin pathway (24).
Accordingly, targeting HMGB3 may suppress or eliminate CSCs,
reducing the risks of recurrence and metastasis.
Potential of HMGB3 as a prognostic
biomarker and efficacy predictor
HMGB3 can also be regarded as a promising biomarker
for diagnosis, prognosis and treatment response prediction, given
its abnormal expression in multiple cancers and strong association
with clinical outcomes. HMGB3 is overexpressed in various tumors
(e.g., breast cancer, NSCLC, CRC, bladder cancer, prostate cancer),
with elevated levels being associated with poor prognosis.
Furthermore, its expression also exhibits a positive association
with advanced clinicopathological features, such as tumor grade,
size, stage and lymph node metastasis (105,109,114,116). In breast cancer, HMGB3 shows
excellent diagnostic potential, with an area under the receiver
operating characteristic curve of 0.932 (109). In NPC and thyroid cancer,
circulating HMGB3 in exosomes or serum may imply metastasis,
suggesting the potential of liquid biopsy (101,115). Elevated HMGB3 may predict
resistance to radiotherapy, chemotherapy, targeted therapy and
ICIs, as well as unfavorable survival, underscoring its role in
treatment response prediction and prognostic assessment.
Conclusions and future perspectives
This review systematically summarizes the central
roles of HMGB3 in resistance to cancer therapy and cancer stemness.
HMGB3 is highly and specifically expressed during embryonic
development, but remains largely silent in adult tissues. It may
function significantly in malignant transformation, given its
aberrant reactivation in various malignancies. HMGB3 can regulate
tumor cell functions through a multi-level network, maintaining CSC
properties, promoting DNA damage repair and cellular survival and
further supporting therapy resistance via TME remodeling.
Mechanistically, these functions are finely modulated by ncRNAs and
PTMs, highlighting that HMGB3 may be a core molecular node linking
cancer stemness, genomic instability, cellular survival and
microenvironmental regulation.
Despite notable progress, there are significant
deficiencies in existing research on HMGB3. Specifically, most
functional validations are limited to in vitro models, with
a lack of support from complex in vivo systems. Meanwhile,
there is a poor interpretation of its roles and underlying
molecular mechanisms within the tumor immune microenvironment.
Additionally, its clinical translation is blocked owing to the
absence of highly specific HMGB3 small-molecule inhibitors or
neutralizing antibodies. In the future, there is a need to develop
highly specific HMGB3-targeted tools and validate their therapeutic
potential using patient-derived organoids and humanized models, and
to conduct multicenter clinical cohort studies to systematically
assess the correlation between HMGB3 expression and treatment
response or prognosis. Besides, it is important to elucidate its
dual roles within the tumor immune microenvironment and their
molecular bases to provide a rationale for combination
immunotherapy strategies. Overall, HMGB3 serves as a pivotal node
connecting multiple oncogenic processes that exhibits substantial
translational potential and may be a novel precision therapeutic
target for overcoming the resistance of cancer therapy.
Acknowledgements
Not applicable.
Funding
This study was supported by Wuxi Taihu Talent Plan (grant no.
THRCJH20200406) and the Major Scientific Research Projects of Wuxi
City (grant no. Z202325).
Availability of data and materials
Not applicable.
Authors' contributions
NHF and JFS designed this study and provided
clinical guidance as well as data interpretation. JZ and YFS
prepared the figures for this study. JZ and LYW drafted the
manuscript. JYG and YH supervised the study and revised the
manuscript. Data authentication is not applicable. All authors
reviewed the manuscript, provided comments and approved the final
version.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Wen B, Wei YT and Zhao K: The role of high
mobility group protein B3 (HMGB3) in tumor proliferation and drug
resistance. Mol Cell Biochem. 476:1729–1739. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Chikhirzhina E, Tsimokha A, Tomilin AN and
Polyanichko A: Structure and functions of HMGB3 protein. Int J Mol
Sci. 25:76562024. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dangerfield J, Mukherjee A, Reh W,
Battenhouse A and Vasquez KM: High-Mobility group box protein 3
(HMGB3) facilitates DNA interstrand crosslink processing and
Double-strand break repair in human cells. Genes (Basel).
16:10442025. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Niu L, Yang W, Duan L, Wang X, Li Y, Xu C,
Liu C, Zhang Y, Zhou W, Liu J, et al: Biological functions and
theranostic potential of HMGB family members in human cancers. Ther
Adv Med Oncol. 12:17588359209708502020. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Vaccari T, Beltrame M, Ferrari S and
Bianchi ME: Hmg4, a new member of the Hmg1/2 gene family. Genomics.
49:247–252. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Lv Y, Lv M, Ji X, Xue L, Rui C, Yin L,
Ding H and Miao Z: Down-regulated expressed protein HMGB3 inhibits
proliferation and migration, promotes apoptosis in the placentas of
fetal growth restriction. Int J Biochem Cell Biol. 107:69–76. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tang D, Kang R, Zeh HJ and Lotze MT: The
multifunctional protein HMGB1: 50 years of discovery. Nat Rev
Immunol. 23:824–841. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mukherjee A, Huynh V, Gaines K, Reh WA and
Vasquez KM: Targeting the High-mobility group box 3 protein
sensitizes chemoresistant ovarian cancer cells to cisplatin. Cancer
Res. 79:3185–3191. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Ma H, Qi G, Han F, Lu W, Peng J, Li R, Yan
S, Yuan C and Kong B: HMGB3 promotes PARP inhibitor resistance
through interacting with PARP1 in ovarian cancer. Cell Death Dis.
13:2632022. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhou GH, Lu YY, Xie JL, Gao ZK, Wu XB, Yao
WS and Gu WG: Overexpression of miR-758 inhibited proliferation,
migration, invasion, and promoted apoptosis of non-small cell lung
cancer cells by negatively regulating HMGB. Biosci Rep.
39:BSR201808552019. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhou D, He S, Zhang D, Lv Z, Yu J, Li Q,
Li M, Guo W and Qi F: LINC00857 promotes colorectal cancer
progression by sponging miR-150-5p and upregulating HMGB3 (high
mobility group box 3) expression. Bioengineered. 12:12107–12122.
2021. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xie X, Pan J, Han X and Chen W:
Downregulation of microRNA-532-5p promotes the proliferation and
invasion of bladder cancer cells through promotion of
HMGB3/Wnt/β-catenin signaling. Chem Biol Interact. 300:73–81. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Swenson SA, Dhir A, Weber IS, Dobish KK,
Xiao P, Buckley SM and Hyde KT: HMGB3: A novel regulator of
leukemia proliferation. Blood. 144:4112–4113. 2024. View Article : Google Scholar
|
|
14
|
Guo S, Wang Y, Gao Y, Zhang Y, Chen M, Xu
M, Hu L, Jing Y, Jing F, Li C, et al: Knockdown of high mobility
Group-Box 3 (HMGB3) expression inhibits proliferation, reduces
migration, and affects chemosensitivity in gastric cancer cells.
Med Sci Monit. 22:3951–3960. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Qiu C, Chen Y, Xia H, Duan J and Zhang L,
Zhang Y, Chen Z and Zhang L: Hsa_circ_0004662 accelerates the
progression of ulcerative colitis via the microRNA-532/HMGB3
signalling axis. J Cell Mol Med. 29:e704302025. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Qin X, Niu Z, Chen H and Hu Y:
Macrophage-derived exosomal HMGB3 regulates silica-induced
pulmonary inflammation by promoting M1 macrophage polarization and
recruitment. Part Fibre Toxicol. 21:122024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Manhas A, Tripathi D and Jagavelu K:
Involvement of HIF1α/Reg protein in the regulation of HMGB3 in
myocardial infarction. Vascul Pharmacol. 152:1071972023. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Chen R, Zou J, Zhong X, Li J, Kang R and
Tang D: HMGB1 in the interplay between autophagy and apoptosis in
cancer. Cancer Lett. 581:2164942024. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sethi N and Kang Y: Unravelling the
complexity of metastasis-molecular understanding and targeted
therapies. Nat Rev Cancer. 11:735–748. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Vemula S, Bonala S, Vadde NK, Natu JZ,
Basha R, Vadde R and Ahmad S: Drug resistance and immunotherapy in
gynecologic cancers. Life Sci. 332:1221042023. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Mirzaee F, Abbaszade-CheragheAli A and
Khamoushi A: Overview of high mobility group box 3 (HMGB3] protein.
Mol Genet Genomics. 300:592025. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Gong Y, Cao Y, Song L, Zhou J, Wang C and
Wu B: HMGB3 characterization in gastric cancer. Genet Mol Res.
12:6032–6039. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zhang Y, Yu Y, Yuan L and Zhang B: EZH2
promotes glioma cell proliferation, invasion, and migration via
Mir-142-3p/KCNQ1OT1/HMGB3 Axis: Running Title: EZH2 promotes glioma
cell malignant behaviors. Mol Neurobiol. 61:8668–8687. 2024.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gong W, Guo Y, Yuan H, Hu X, Chai R, Zheng
B, Wan Z and Tu S: HMGB3 is a potential therapeutic target by
affecting the migration and proliferation of colorectal cancer.
Front Cell Dev Biol. 10:8914822022. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zheng M, Wu L, Xiao R, Zhou Y, Cai J, Chen
W, Chen C, Sun K and Shen S: Integrated analysis of coexpression
and a tumor-specific ceRNA network revealed a potential prognostic
biomarker in breast cancer. Transl Cancer Res. 12:949–964. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Moore LD, Le T and Fan G: DNA methylation
and its basic function. Neuropsychopharmacology. 38:23–38. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Xu Y, Xu M, Li X, Weng X, Su Z, Zhang M,
Tan J, Zeng H, Li X, Nie L, et al: SOX9 and HMGB3 co-operatively
transactivate NANOG and promote prostate cancer progression.
Prostate. 83:440–453. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Toda H, Seki N, Kurozumi S, Shinden Y,
Yamada Y, Nohata N, Moriya S, Idichi T, Maemura K, Fujii T, et al:
RNA-sequence-based microRNA expression signature in breast cancer:
Tumor-suppressive miR-101-5p regulates molecular pathogenesis. Mol
Oncol. 14:426–446. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Sharma P, Yadav P, Sundaram S, Venkatraman
G, Bera AK and Karunagaran D: HMGB3 inhibition by miR-142-3p/sh-RNA
modulates autophagy and induces apoptosis via ROS accumulation and
mitochondrial dysfunction and reduces the tumorigenic potential of
human breast cancer cells. Life Sci. 304:1207272022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Elgamal OA, Park JK, Gusev Y,
Azevedo-Pouly AC, Jiang J, Roopra A and Schmittgen TD: Tumor
suppressive function of mir-205 in breast cancer is linked to HMGB3
regulation. PLoS One. 8:e764022013. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Liu D, Wang Y, Zhao Y and Gu X: LncRNA
SNHG5 promotes nasopharyngeal carcinoma progression by regulating
miR-1179/HMGB3 axis. BMC Cancer. 20:1782020. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Shi J, Wang H, Feng W, Huang S, An J, Qiu
Y and Wu K: Long non-coding RNA HOTTIP promotes hypoxia-induced
glycolysis through targeting miR-615-3p/HMGB3 axis in non-small
cell lung cancer cells. Eur J Pharmacol. 862:1726152019. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Xi J, Xi Y, Zhang Z, Hao Y, Wu F, Bian B,
Hao G, Li W and Zhang S: Hsa_circ_0060937 accelerates non-small
cell lung cancer progression via modulating miR-195-5p/HMGB3
pathway. Cell Cycle. 20:2040–2052. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Zhou ZF, Wei Z, Yao JC, Liu SY, Wang F,
Wang Z, Chen XF, Lin H, Ye Y and Zheng QF: CircRNA_102179 promotes
the proliferation, migration and invasion in non-small cell lung
cancer cells by regulating miR-330-5p/HMGB3 axis. Pathol Res Pract.
216:1531442020. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yang M, Zheng E, Ni J, Xu X, Jiang X and
Zhao G: Circular RNA circFOXO3 facilitate non-small cell lung
cancer progression through upregulating HMGB3 via sponging
miR-545-3p/miR-506-3p. Tissue Cell. 75:1017022022. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Gao S, Zhang X, Bai W, Wang J and Jiang B:
Circ-IGF1R affects the progression of colorectal cancer by
activating the miR-362-5p/HMGB3-Mediated Wnt/β-Catenin signal
pathway. Biochem Genet. 61:1210–1229. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yang K, Zhang F, Luo B and Qu Z:
CAFs-derived small extracellular vesicles circN4BP2L2 promotes
proliferation and metastasis of colorectal cancer via
miR-664b-3p/HMGB3 pathway. Cancer Biol Ther. 23:404–416. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yuan H, Chen B, Chai R, Gong W, Wan Z,
Zheng B, Hu X, Guo Y, Gao S, Dai Q, et al: Loss of exosomal
micro-RNA-200b-3p from hypoxia cancer-associated fibroblasts
reduces sensitivity to 5-flourouracil in colorectal cancer through
targeting high-mobility group box 3. Front Oncol. 12:9201312022.
View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Sun F, Yang X, Song W, Yu N and Lin Q:
Tanshinone IIA (TSIIA) represses the progression of non-small cell
lung cancer by the circ_0020123/miR-1299/HMGB3 pathway. Mol Cell
Biochem. 478:1973–1986. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wang J, Sheng Z and Cai Y: Effects of
microRNA-513b on cell proliferation, apoptosis, invasion, and
migration by targeting HMGB3 through regulation of mTOR signaling
pathway in non-small-cell lung cancer. J Cell Physiol.
234:10934–10941. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Li X, Wu Y, Liu A and Tang X: MiR-27b is
epigenetically downregulated in tamoxifen resistant breast cancer
cells due to promoter methylation and regulates tamoxifen
sensitivity by targeting HMGB3. Biochem Biophys Res Commun.
477:768–773. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Yu Q, Li Y, Peng S, Li J and Qin X:
Exosomal-mediated transfer of OIP5-AS1 enhanced cell
chemoresistance to trastuzumab in breast cancer via up-regulating
HMGB3 by sponging miR-381-3p. Open Med (Wars). 16:512–525. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Tian X, Chang J, Zhang N, Wu S, Liu H and
Yu J: MicroRNA-429 acts as a tumor suppressor in colorectal cancer
by targeting high mobility group box 3. Oncol Lett. 21:2502021.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Gu M, Jiang Z, Li H, Peng J, Chen X and
Tang M: MiR-93/HMGB3 regulatory axis exerts tumor suppressive
effects in colorectal carcinoma cells. Exp Mol Pathol.
120:1046352021. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Chen X, Zhao G, Wang F, Gao F, Luo H, Wang
Y, Du Y, Chen X, Xue C, Dong Z, et al: Upregulation of miR-513b
inhibits cell proliferation, migration, and promotes apoptosis by
targeting high mobility group-box 3 protein in gastric cancer.
Tumour Biol. 35:11081–11089. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Song X, Wang H, Wu J and Sun Y: Long
noncoding RNA SOX2-OT knockdown inhibits proliferation and
metastasis of prostate cancer cells through modulating the
miR-452-5p/HMGB3 axis and inactivating Wnt/β-Catenin pathway.
Cancer Biother Radiopharm. 35:682–695. 2020.PubMed/NCBI
|
|
47
|
Chen F, Sun F, Liu X, Shao J and Zhang B:
Glaucocalyxin A inhibits the malignant progression of epithelial
ovarian cancer by affecting the
MicroRNA-374b-5p/HMGB3/Wnt-β-catenin pathway axis. Front Oncol.
12:9558302022. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Sun D, Cao R, Han L, Yu X, Wang H, Wang X
and Chen X: Long noncoding RNA brain cytoplasmic RNA 1 induces
Cisplatin-resistance of cervical cancer cells by sponging
MicroRNA-330-5p and upregulating High-Mobility group Box 3. Gynecol
Obstet Invest. 87:200–210. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Song T, Hou X and Lin B: MicroRNA-758
inhibits cervical cancer cell proliferation and metastasis by
targeting HMGB3 through the WNT/β-catenin signaling pathway. Oncol
Lett. 18:1786–1792. 2019.PubMed/NCBI
|
|
50
|
Sun CX, Zhu F and Qi L: Demethylated
miR-216a regulates high mobility group box 3 promoting growth of
esophageal cancer cells through Wnt/β-Catenin pathway. Front Oncol.
11:6220732021. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Chen R, Kang R and Tang D: The mechanism
of HMGB1 secretion and release. Exp Mol Med. 54:91–102. 2022.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Starkova T, Polyanichko A, Tomilin AN and
Chikhirzhina E: Structure and functions of HMGB2 protein. Int J Mol
Sci. 24:83342023. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Kwak MS, Kim HS, Lkhamsuren K, Kim YH, Han
MG, Shin JM, Park IH, Rhee WJ, Lee SK, Rhee SG and Shin JS:
Peroxiredoxin-mediated disulfide bond formation is required for
nucleocytoplasmic translocation and secretion of HMGB1 in response
to inflammatory stimuli. Redox Biol. 24:1012032019. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Wei T, Liu J, Li C, Tan Y, Wei R, Wang J,
Wu H, Li Q, Liu H, Tang Y and Li X: Revealing the extracellular
function of HMGB1 N-terminal region acetylation assisted by a
protein semi-synthesis approach. Chem Sci. 14:10297–10307. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Elenkov I, Pelovsky P, Ugrinova I,
Takahashi M and Pasheva E: The DNA binding and bending activities
of truncated tail-less HMGB1 protein are differentially affected by
Lys-2 and Lys-81 residues and their acetylation. Int J Biol Sci.
7:691–699. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Pasheva E, Sarov M, Bidjekov K, Ugrinova
I, Sarg B, Lindner H and Pashev IG: In vitro acetylation of HMGB-1
and −2 proteins by CBP: The role of the acidic tail. Biochemistry.
43:2935–2940. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
He Y, Ding Y, Wang D, Zhang W, Chen W, Liu
X, Qin W, Qian X, Chen H and Guo Z: HMGB1 bound to cisplatin-DNA
adducts undergoes extensive acetylation and phosphorylation in
vivo. Chem Sci. 6:2074–2078. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Zhou S, Lu H, Chen R, Tian Y, Jiang Y,
Zhang S, Ni D, Su Z and Shao X: Angiotensin II enhances the
acetylation and release of HMGB1 in RAW264.7 macrophage. Cell Biol
Int. 42:1160–1169. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Pelovsky P, Pashev IG and Pasheva E:
Interplay between in vitro acetylation and phosphorylation of
tailless HMGB1 protein. Biochem Biophys Res Commun. 380:138–142.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Zhang X, Wheeler D, Tang Y, Guo L, Shapiro
RA, Ribar TJ, Means AR, Billiar TR, Angus DC and Rosengart MR:
Calcium/calmodulin-dependent protein kinase (CaMK) IV mediates
nucleocytoplasmic shuttling and release of HMGB1 during
lipopolysaccharide stimulation of macrophages. J Immunol.
181:5015–5023. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Youn JH and Shin JS: Nucleocytoplasmic
shuttling of HMGB1 is regulated by phosphorylation that redirects
it toward secretion. J Immunol. 177:7889–7897. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Lv WL, Arnesano F, Carloni P, Natile G and
Rossetti G: Effect of in vivo post-translational modifications of
the HMGB1 protein upon binding to platinated DNA: A molecular
simulation study. Nucleic Acids Res. 46:11687–11697. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Liu Y, Song D, Li S, Guo Z and Zheng P:
Click Chemistry-based force spectroscopy revealed enhanced binding
dynamics of phosphorylated HMGB1 to cisplatin-DNA. J Am Chem Soc.
146:13126–13132. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Venereau E, Casalgrandi M, Schiraldi M,
Antoine DJ, Cattaneo A, De Marchis F, Liu J, Antonelli A, Preti A,
Raeli L, et al: Mutually exclusive redox forms of HMGB1 promote
cell recruitment or proinflammatory cytokine release. J Exp Med.
209:1519–1528. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Taverna S, Tonacci A, Ferraro M, Cammarata
G, Cuttitta G, Bucchieri S, Pace E and Gangemi S: High mobility
group box 1: Biological functions and relevance in oxidative stress
related chronic diseases. Cells. 11:8492022. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Kazama H, Ricci JE, Herndon JM, Hoppe G,
Green DR and Ferguson TA: Induction of immunological tolerance by
apoptotic cells requires caspase-dependent oxidation of
high-mobility group box-1 protein. Immunity. 29:21–32. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Yang H, Hreggvidsdottir HS, Palmblad K,
Wang H, Ochani M, Li J, Lu B, Chavan S, Rosas-Ballina M, Al-Abed Y,
et al: A critical cysteine is required for HMGB1 binding to
Toll-like receptor 4 and activation of macrophage cytokine release.
Proc Natl Acad Sci USA. 107:11942–11947. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Raucci A, Di Maggio S, Scavello F,
D'Ambrosio A, Bianchi ME and Capogrossi MC: The Janus face of HMGB1
in heart disease: A necessary update. Cell Mol Life Sci.
76:211–229. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Ito I, Fukazawa J and Yoshida M:
Post-translational methylation of high mobility group box 1 (HMGB1)
causes its cytoplasmic localization in neutrophils. J Biol Chem.
282:16336–16344. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Wu F, Zhao ZH, Ding ST, Wu HH and Lu JJ:
High mobility group box 1 protein is methylated and transported to
cytoplasm in clear cell renal cell carcinoma. Asian Pac J Cancer
Prev. 14:5789–5795. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Vergoten G and Bailly C: N-glycosylation
of high mobility group box 1 protein (HMGB1) modulates the
interaction with glycyrrhizin: A molecular modeling study. Comput
Biol Chem. 88:1073122020. View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Kim YH, Kwak MS, Park JB, Lee SA, Choi JE,
Cho HS and Shin JS: N-linked glycosylation plays a crucial role in
the secretion of HMGB1. J Cell Sci. 129:29–38. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Bhavnagari H, Raval A and Shah F:
Deciphering potential role of hippo signaling pathway in breast
cancer: A comprehensive review. Curr Pharm Des. 29:3505–3518. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
74
|
Zhao Q, Zong H, Zhu P, Su C, Tang W, Chen
Z and Jin S: Crosstalk between colorectal CSCs and immune cells in
tumorigenesis, and strategies for targeting colorectal CSCs. Exp
Hematol Oncol. 13:62024. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Lian JW, Li SY, Clarke RB, Howell SJ and
Meng QJ: Can we utilise the circadian clock to target cancer stem
cells? Cancer Lett. 611:2173602024. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Nemeth MJ, Kirby MR and Bodine DM: Hmgb3
regulates the balance between hematopoietic stem cell self-renewal
and differentiation. Proc Natl Acad Sci USA. 103:13783–13788. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Gu J, Xu T, Zhang CM, Chen HY, Huang QH
and Zhang Q: HMGB3 small interfere RNA suppresses mammosphere
formation of MDA-MB-231 cells by down-regulating expression of
HIF1α. Eur Rev Med Pharmacol Sci. 23:9506–9516. 2019.PubMed/NCBI
|
|
78
|
Leis O, Eguiara A, Lopez-Arribillaga E,
Alberdi MJ, Hernandez-Garcia S, Elorriaga K, Pandiella A, Rezola R
and Martin AG: Sox2 expression in breast tumours and activation in
breast cancer stem cells. Oncogene. 31:1354–1365. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Varisli L, Zoumpourlis P, Spandidos DA,
Zoumpourlis V and Vlahopoulos S: ALDH1A1 in breast cancer: A
prospective target to overcome therapy resistance (review). Oncol
Lett. 29:2132025. View Article : Google Scholar : PubMed/NCBI
|
|
80
|
Ma H, Qi G, Han F, Gai P, Peng J and Kong
B: HMGB3 promotes the malignant phenotypes and stemness of
epithelial ovarian cancer through the MAPK/ERK signaling pathway.
Cell Commun Signal. 21:1442023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Wang LK, Xie XN, Song XH, Su T, Chang XL,
Xu M, Liang B and Huang DY: Upregulation of miR-200b inhibits
hepatocellular carcinoma cell proliferation and migration by
targeting HMGB3 protein. Technol Cancer Res Treat.
17:15330338188064752018. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Chen X and Zeng L: Ginkgo biloba extract
761 enhances 5-fluorouracil chemosensitivity in colorectal cancer
cells through regulation of high mobility group-box 3 expression.
Am J Transl Res. 10:1773–1783. 2018.PubMed/NCBI
|
|
83
|
Zhang S, Liu J, Yuan T, Liu H, Wan C and
Le Y: Circular RNA 0001313 knockdown suppresses non-small cell lung
cancer cell proliferation and invasion via the
microRNA-452/HMGB3/ERK/MAPK Axis. Int J Gen Med. 13:1495–1507.
2020. View Article : Google Scholar : PubMed/NCBI
|
|
84
|
Liu J, Wang L and Li X: HMGB3 promotes the
proliferation and metastasis of glioblastoma and is negatively
regulated by miR-200b-3p and miR-200c-3p. Cell Biochem Funct.
36:357–365. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Ji F, Yao Z, Liu C, Fu S, Ren B, Liu Y, Ma
L, Wei J and Sun D: A novel lnc-LAMC2-1:1 SNP promotes colon
adenocarcinoma progression by targeting miR-216a-3p/HMGB3. Heliyon.
8:e123422022. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Zhao T, He X, Liang X, Kellum AH Jr, Tang
F, Yin J, Guo S and Wang Y, Gao Z and Wang Y: HMGB3 and SUB1 bind
to and facilitate the repair of N2-Alkylguanine lesions in DNA. J
Am Chem Soc. 146:22553–22562. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Ray Chaudhuri A and Nussenzweig A: The
multifaceted roles of PARP1 in DNA repair and chromatin
remodelling. Nat Rev Mol Cell Biol. 18:610–621. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Li Z, Zhang Y, Sui S, Hua Y, Zhao A, Tian
X, Wang R, Guo W, Yu W, Zou K, et al: Targeting HMGB3/hTERT axis
for radioresistance in cervical cancer. J Exp Clin Cancer Res.
39:2432020. View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Masutomi K, Possemato R, Wong JM, Currier
JL, Tothova Z, Manola JB, Ganesan S, Lansdorp PM, Collins K and
Hahn WC: The telomerase reverse transcriptase regulates chromatin
state and DNA damage responses. Proc Natl Acad Sci USA.
102:8222–8227. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
90
|
Yang SF, Nelson CB, Wells JK, Fernando M,
Lu R, Allen JAM, Malloy L, Lamm N, Murphy VJ, Mackay JP, et al:
ZNF827 is a single-stranded DNA binding protein that regulates the
ATR-CHK1 DNA damage response pathway. Nat Commun. 15:22102024.
View Article : Google Scholar : PubMed/NCBI
|
|
91
|
Saldivar JC, Hamperl S, Bocek MJ, Chung M,
Bass TE, Cisneros-Soberanis F, Samejima K, Xie L, Paulson JR,
Earnshaw WC, et al: An intrinsic S/G2 checkpoint enforced by ATR.
Science. 361:806–810. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
92
|
Liu Q, Guntuku S, Cui XS, Matsuoka S,
Cortez D, Tamai K, Luo G, Carattini-Rivera S, DeMayo F, Bradley A,
et al: Chk1 is an essential kinase that is regulated by Atr and
required for the G(2)/M DNA damage checkpoint. Genes Dev.
14:1448–1459. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
93
|
Mukherjee A, Benhamou LR and Vasquez KM:
Abstract 1512: Architectural protein HMGB3 interacts with cisplatin
resistance associated overexpressed protein (CROP/LUC7L3) in human
cancer cells and modulates cisplatin-DNA adduct removal. Cancer
Res. 82:15122022. View Article : Google Scholar
|
|
94
|
Kok I, Bayraktar C, Durgun A, Aksu AC,
Kayabolen A, Yedier Bayram O, Sur Erdem I and Bagci Onder T:
P03.10.A identifying DNA damage response (DDR) related factors
Essenti al for radiotherapy response in glioblastoma using a novel
CRISPR/Cas9 library, ddrkol. Neuro Oncol. 25:ii382023. View Article : Google Scholar
|
|
95
|
Li J, Song C, Gu J, Li C, Zang W, Shi L,
Chen L, Zhu L, Zhou M, Wang T, et al: RBBP4 regulates the
expression of the Mre11-Rad50-NBS1 (MRN) complex and promotes DNA
double-strand break repair to mediate glioblastoma
chemoradiotherapy resistance. Cancer Lett. 557:2160782023.
View Article : Google Scholar : PubMed/NCBI
|
|
96
|
Zhuang S, Yu X, Lu M, Li Y, Ding N and
Ding Y: High mobility group box 3 promotes cervical cancer
proliferation by regulating Wnt/β-catenin pathway. J Gynecol Oncol.
31:e912020. View Article : Google Scholar : PubMed/NCBI
|
|
97
|
Dong S, Pan J, Shen YB, Zhu LX, Chen L,
Zhu F, Li H, Shen HX, Xia Q, Wu YJ and Xie XJ: SYT7 plays a role in
promoting thyroid cancer by mediating HMGB3 ubiquitination. Endocr
Relat Cancer. 29:175–189. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
98
|
Wang X, Shao X, Li T, Zhang L, Yang Q, Ye
W, Tong J, Li Z and Fang X: Pingchuanning Formula suppresses airway
inflammation in a rat model of asthmatic cold syndrome by
regulating the HMGB1/Beclin-1 axis-mediated autophagy. Nan Fang Yi
Ke Da Xue Xue Bao. 45:1153–1162. 2025.(In Chinese). PubMed/NCBI
|
|
99
|
Tong HY, Yang WL, Ma LY, Zhang YD, Wang W,
Li KF, Lang W, Wang L, Yang WB and Huang H: HMGB3 as a cargo
protein for XPO1: Implications for myelodysplastic syndromes
prognosis and treatment. Blood. 144:4570–4571. 2024. View Article : Google Scholar
|
|
100
|
Gu J, Xu T, Huang QH, Zhang CM and Chen
HY: HMGB3 silence inhibits breast cancer cell proliferation and
tumor growth by interacting with hypoxia-inducible factor 1α.
Cancer Manag Res. 11:5075–5089. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
101
|
Zhang K, Liu D, Zhao J, Shi S, He X, Da P,
You Y and You B: Nuclear exosome HMGB3 secreted by nasopharyngeal
carcinoma cells promotes tumour metastasis by inducing
angiogenesis. Cell Death Dis. 12:5542021. View Article : Google Scholar : PubMed/NCBI
|
|
102
|
Khambu B, Yan S, Huda N and Yin XM: Role
of High-mobility group Box-1 in liver pathogenesis. Int J Mol Sci.
20:53142019. View Article : Google Scholar : PubMed/NCBI
|
|
103
|
Patra S, Roy PK, Dey A and Mandal M:
Impact of HMGB1 on cancer development and therapeutic insights
focused on CNS malignancy. Biochim Biophys Acta Rev Cancer.
1879:1891052024. View Article : Google Scholar : PubMed/NCBI
|
|
104
|
Wang L, Xu P, Li X and Zhang Q:
Comprehensive bioinformatics analysis identified HMGB3 as a
promising immunotherapy target for glioblastoma multiforme. Discov
Oncol. 16:4782025. View Article : Google Scholar : PubMed/NCBI
|
|
105
|
Li M, Cai Y, Zhao H, Xu Z, Sun Q, Luo M,
Gu L, Meng M, Han X and Sun H: Overexpression of HMGB3 protein
promotes cell proliferation, migration and is associated with poor
prognosis in urinary bladder cancer patients. Tumour Biol.
36:4785–4792. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
106
|
Ke Y, Mai J, Liu Z, Xu Y, Zhao C and Wang
B: Interfering HMGB3 release from cancer-associated fibroblasts by
miR-200b represses chemoresistance and epithelial-mesenchymal
transition of gastric cancer cells. J Gastrointest Oncol.
13:2197–2218. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
107
|
Zhu L and Chen L: Progress in research on
paclitaxel and tumor immunotherapy. Cell Mol Biol Lett. 24:402019.
View Article : Google Scholar : PubMed/NCBI
|
|
108
|
Singh A, Srivastav S, Singh MP, Singh R,
Kumar P and Kush P: Recent advances in phytosomes for the safe
management of cancer. Phytomedicine Plus. 4:1005402019. View Article : Google Scholar
|
|
109
|
Zhou X, Zhang Q, Liang G, Liang X and Luo
B: Overexpression of HMGB3 and its prognostic value in breast
cancer. Front Oncol. 12:10489212022. View Article : Google Scholar : PubMed/NCBI
|
|
110
|
Fu X, Li P, Zhou Q, He R, Wang G, Zhu S,
Bagheri A, Kupfer G, Pei H and Li J: Mechanism of PARP inhibitor
resistance and potential overcoming strategies. Genes Dis.
11:306–320. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
111
|
Luo B, Zheng H, Liang G, Luo Y, Zhang Q
and Li X: HMGB3 contributes to Anti-PD-1 resistance by inhibiting
IFN-γ-Driven ferroptosis in TNBC. Mol Carcinog. 64:490–501. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
112
|
Yamada Y, Nishikawa R, Kato M, Okato A,
Arai T, Kojima S, Yamazaki K, Naya Y, Ichikawa T and Seki N:
Regulation of HMGB3 by antitumor miR-205-5p inhibits cancer cell
aggressiveness and is involved in prostate cancer pathogenesis. J
Hum Genet. 63:195–205. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
113
|
Zhong X, Zhang S, Zhang Y, Jiang Z, Li Y,
Chang J, Niu J and Shi Y: HMGB3 is associated with an unfavorable
prognosis of neuroblastoma and promotes tumor progression by
mediating TPX2. Front Cell Dev Biol. 9:7695472021. View Article : Google Scholar : PubMed/NCBI
|
|
114
|
Zhang Z, Chang Y, Zhang J, Lu Y, Zheng L,
Hu Y, Zhang F and Li X, Zhang W and Li X: HMGB3 promotes growth and
migration in colorectal cancer by regulating WNT/β-catenin pathway.
PLoS One. 12:e01797412017. View Article : Google Scholar : PubMed/NCBI
|
|
115
|
Zhao Y, Lv HJ, Deng XY, Chen P, Garstka
MA, Shi BY and Fu J: Translocated HMGB3 is involved in papillary
thyroid cancer progression by activating cytoplasmic TLR3 and
transmembrane TREM1. Cell Cycle. 22:2584–2601. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
116
|
Song N, Liu B, Wu JL, Zhang RF, Duan L, He
WS and Zhang CM: Prognostic value of HMGB3 expression in patients
with non-small cell lung cancer. Tumour Biol. 34:2599–2603. 2013.
View Article : Google Scholar : PubMed/NCBI
|