Introduction
Head and neck squamous cell carcinoma (HNSCC)
remains a highly aggressive malignancy characterized by substantial
heterogeneity in clinical behavior and treatment response (1). Despite advances in surgical
techniques, radiotherapy, chemotherapy and immunotherapy, a
substantial proportion of patients still experience poor
progression, manifested as local recurrence, distant metastasis,
treatment resistance and ultimately reduced survival (2,3).
Identifying and validating biomarkers associated with poor
progression is therefore of particular importance (4). Such markers not only provide
mechanistic insights into tumor aggressiveness and disease
evolution but also serve as prognostic tools to stratify patients
according to risk. Moreover, survival-related markers can inform
personalized treatment strategies, guide therapeutic escalation or
de-escalation and support the development of novel targeted
interventions (5). The established
markers associated with poor progression in HNSCC include human
papillomavirus, TP53 mutations, EGFR overexpression, elevated
programmed cell death-ligand 1 expression and the upregulation of
epithelial-mesenchymal transition (EMT)/stemness-related genes
(6–8). These markers are associated with tumor
invasion, metastasis, therapeutic resistance and immune evasion. A
comprehensive analysis of poor progression markers thus holds
considerable promise for improving prognostication, optimizing
patient management and advancing precision oncology in HNSCC.
Leucine zipper protein 1 (LUZP1) is a highly
conserved cytoskeletal- and signaling-associated protein that plays
key roles in the maturation of contractile actomyosin bundle,
cytoskeletal organization and the regulation of cell division
(9,10). LUZP1 is initially identified in the
brain, where it participates in embryonic neural development
(11). Emerging evidence further
suggests that LUZP1 exerts vital functions in tumor biology
(12–16). Aberrant expression of LUZP1 has been
shown to enhance tumor cell migration, invasion and EMT in
osteosarcoma and triple-negative breast cancer (12,13).
Previous proteomic analyses have also identified LUZP1 as a
component of oncogenic signaling networks that contribute to
therapeutic resistance (14).
Moreover, LUZP1 expression has been associated with poor prognosis
in several malignancies, including osteosarcoma, renal papillary
cell carcinoma and ovarian cancer (12,15,16).
However, the oncogenic role and clinical importance of LUZP1 in
HNSCC remain to be elucidated.
The present study first performed a proteomic
analysis comparing tumor specimens from patients with HNSCC with
favorable long-term survival and those with poor short-term
survival. Bioinformatic pathway enrichment was then conducted to
identify candidate survival-associated regulators. Based on this
discovery-driven approach, the present study aimed to functionally
characterize selected candidates in vitro and in clinical
specimens, and to determine their mechanistic contribution to
metastasis, chemoresistance and patient prognosis in HNSCC.
Materials and methods
Pathological tissue specimens
The present study protocol was approved by the
Ethics Committee of China Medical University Hospital (approval no.
CMUH108-REC1-127). Tumor mass and tumor slice specimens from
patients with HNSCC were obtained from the Human Biobank of China
Medical University Hospital (Taichung, Taiwan). For proteomic
analysis and protein expression validation, three tumor masses from
the short-term survival group and three from the long-term survival
group were included. Patients with an overall survival of <2
years were classified as the short-term survival group, whereas
those with an overall survival of ≥5 years were defined as the
long-term survival group. For immunohistochemical analysis, tumor
tissue samples from 130 patients enrolled between June 2014 and
September 2016 were retrospectively analyzed. The inclusion
criteria included i) newly diagnosed squamous cell carcinoma; ii)
advanced-stage disease (stage III–IV); and iii) receipt of both
radiotherapy and surgical treatment. Specimens were excluded if
patients i) had documented HIV infection or ii) declined to provide
informed consent for tissue use.
The demographic and clinicopathological
characteristics of the HNSCC cohort are summarized in Table I. Pathological tissue specimens were
primarily obtained from tumors of the oral cavity, tongue and
oropharynx. In addition, one paired set of tumor slices derived
from the primary tumor and lymph node metastasis of the same
patient was also included in the analysis.
 | Table I.Demographic characteristics of
patients with head and neck squamous cell carcinoma. |
Table I.
Demographic characteristics of
patients with head and neck squamous cell carcinoma.
|
Characteristics | n | % |
|---|
| Sex |
|
|
|
Male | 121 | 93.1 |
|
Female | 9 | 6.9 |
| Age |
|
|
|
<40 | 28 | 26.5 |
|
40-60 | 79 | 60.8 |
|
>60 | 23 | 17.7 |
| Site |
|
|
| Buccal
mucosa | 49 | 37.7 |
|
Tongue | 49 | 37.7 |
|
others | 32 | 24.6 |
| T |
|
|
| 1 | 14 | 10.8 |
| 2 | 45 | 34.6 |
| 3 | 25 | 19.2 |
| 4 | 46 | 35.4 |
| NS |
|
|
| 0 | 44 | 33.8 |
| 1 | 15 | 11.5 |
| 2 | 47 | 36.2 |
| 3 | 24 | 18.5 |
|
Differentiation |
|
|
|
Well | 54 | 41.5 |
|
Moderate | 58 | 44.6 |
|
Poor | 18 | 13.8 |
| PNI |
|
|
| Y | 67 | 51.5 |
| N | 63 | 48.5 |
| LVI |
|
|
| Y | 48 | 36.9 |
| N | 82 | 63.1 |
| ENE |
|
|
| Y | 36 | 27.7 |
| N | 50 | 38.5 |
| Nodal negative | 44 | 33.8 |
In silico analyses
LUZP1 gene expression levels between normal tissue
and primary tumor, as well as across different nodal metastasis
statuses, were analyzed using the UALCAN web platform based on The
Cancer Genome Atlas (TCGA) database (17). Pairwise gene expression correlation
analyses were conducted using the Gene Expression Profiling
Interaction Analysis (GEPIA) web server, (http://gepia.cancer-pku.cn/), which integrates TCGA
and Genotype Tissue Expression data processed through a
standardized analytical pipeline (18). The monotonic associations between
LUZP1 and NFKB1 or NFKB2 expression levels were evaluated using
Spearman's rank correlation coefficient. ChIP-seq bioinformation of
cell lines was obtained from the Encyclopedia of DNA Elements
(ENCODE) project database (https://www.encodeproject.org/). ENCODE cell lines are
well-characterized and publicly available cell models that have
undergone extensive validation, including authentication and
quality control assessments. Data were accessed through the ENCODE
portal and used in accordance with the project's data usage
guidelines.
Cell lines and cell culture
The human pharyngeal squamous cell carcinoma cell
line FaDu (cat. no. HTB-43) was obtained from the American Type
Culture Collection. The human oral squamous carcinoma cell line
OECM-1 (cat. no. SCC180) was purchased from Merck KGaA and the
human tongue squamous cell carcinoma cell line SAS (cat. no.
JCRB0260) was obtained from Japanese Collection Research
Bioresources Cell Bank. FaDu cells were maintained in Dulbecco's
Modified Eagle Medium (DMEM; cat. no. 12100046; Gibco; Thermo
Fisher Scientific, Inc.); OECM-1 cells were cultured in RPMI-1640
medium (cat. no. 31800022; Gibco; Thermo Fisher Scientific, Inc.)
supplemented with 2 mM l-glutamine (cat. no. 5-10K00-H; BioConcept
Ltd.), while SAS cells were maintained in DMEM supplemented with 2
mM l-glutamine. Docetaxel-resistant (DTXr) and cisplatin-resistant
(CISr) OECM-1 cell lines were established by continuous exposure to
DTX (0.2 nM; cat. no. BML-T129; Enzo Life Sciences) and CIS (0.5
µM; cat. no. 232120 Merck KGAa) for 3 months, which gradually
enhanced their chemoresistance. All cells were maintained in a
humidified incubator with 5% CO2 and 95% air at
37°C.
Chemical reagents and treatments
HNSCC cell lines (FaDu, OECM-1 and SAS) were treated
with pharmacological inhibitors and cytokines to evaluate their
effects on LUZP1 expression. The pharmacological inhibitors used in
the present study included SB431542 (5 and 10 µM; cat. no. 616464;
Merck KGaA), LY294002 (5 and 10 µM; cat. no. HY-10108;
MedChemExpress), rapamycin (5 and 10 µM; cat. no. R0395; Merck
KGaA), BAY 11-7085 (5 and 10 µM; cat. no. HY-10257;
MedChemExpress), and YC-1 (10 and 30 µM; cat. no. sc-202856; Santa
Cruz Biotechnology, Inc.). Cytokine treatments included IL-1β (3
ng/ml; cat. no. C01002-100UG; Croyez Bioscience Co., Ltd.) and TNFα
(10 ng/ml; cat. no. C01047-100UG; Croyez Bioscience Co., Ltd.).
Cells were incubated with the indicated inhibitors or cytokines at
37°C for 24 h unless otherwise specified. For time-course analysis
of LUZP1 induction, cells were treated with IL-1β (3 ng/ml) for 0,
1, 4 and 24 h or with TNFα (10 ng/ml) for 0, 6 and 24 h. In
addition, TNFα (10 ng/ml) was applied to OECM-1 cells to assess its
effects on tumor cell migration. Cell migratory capacity was
assessed using a Transwell migration assay, as described in the
Transwell migration and invasion assays section.
Transwell migration and invasion
assays
For the in vitro migration assay, tumor cells
(2×104 cells in 200 µl of media) were seeded into the
upper chamber of a polyethylene terephthalate membrane Transwell
insert (cat. no. 353097; Corning, Inc.) placed on a 24-well plate.
For the in vitro invasion assay, tumor cells
(4×104 cells in 200 µl of media) were seeded into the
upper chamber of Transwell inserts precoated (at 37°C for 24 h)
with Matrigel (1 g/l; cat. no. 356234; BD Biosciences). Serum-free
media was added to the upper chamber, while medium supplemented
with 10% FBS (Corning, Inc.) was added into the lower chamber as a
chemoattractant. After incubation at 37°C for 24 h (for migration
assay) or 48 h (for invasion assay), the cells that migrated or
invaded through the membrane were fixed with 3.7% formalin (10 min;
room temperature; cat. no. 15512; Merk KGAa) and stained with 0.1%
crystal violet (30 min; room temperature; cat. no. C0775; Merck
KGAa). The stained cells were destained with 50% ethanol (30 min;
room temperature; cat. no. 459836; Merck KGAa) and 0.1% acetic acid
(cat. no. A6283; Merck KGAa), and then the absorbance of the
extracted dye was measured spectrophotometrically at 570 nm for
quantification and cell images were acquired using a bright-field
light microscope.
Three-dimensional (3D) tumor spheroid
invasion assay
Cells (3,000 cells per well) were seeded into
ultra-low attachment (ULA) 96-well round-bottom plates and cultured
for 3 days to allow spheroid formation. After spheroids were
established, a 3D invasion assay was performed. Growth
factor-reduced Matrigel (cat. no. 354230; Corning, Inc.) was
diluted in serum-free DMEM to obtain a 2× working concentration.
The Matrigel solution was added to pre-chilled ULA 96-well
round-bottom plates containing the spheroids at a final volume of
100 µl per well, resulting in a final Matrigel concentration of
3.75 mg/ml. Plates were then incubated at 37°C for 1 h to allow
Matrigel polymerization. 3D tumor spheroid invasion was quantified
using a radial outgrowth distance approach. Phase-contrast images
were acquired at 0, 24, 48 and 96 h following spheroid embedding
using a Nikon TS100 inverted bright-field light microscope. For
each spheroid, the invasion front radius (RX) at each time point
was determined by measuring the radial distance from the spheroid
center to the outer boundary of the contiguous invading cell
population. To minimize the influence of isolated cells and image
noise, the invasion front was defined as the 95th percentile (p95)
of radial distance measurements obtained along multiple angular
axes. The baseline spheroid radius at 0 h (R0) was defined using
the same p95 criterion. Invasion distance at each time point was
calculated as D=RX−R0. Spatial measurements were converted from
pixels to micrometers using the scale bar provided in the 0 h
images. All image analyses were performed using ImageJ software
(version, 1.54 g; National Institutes of Health) and individual
spheroids were analyzed independently as biological replicates for
statistical analysis.
Colony formation assay
For the colony formation assay, tumor cells with or
without LUZP1 knockdown (1,000 cells per well) were seeded into
6-well plates and cultured for 10 days to allow colony formation. A
colony was defined as a cluster containing ≥50 cells. The resulting
colonies were fixed with 3.7% formalin (10 min; room temperature)
and stained with 0.1% crystal violet (30 min; room temperature).
For quantitative analysis, the bound dye was solubilized using 50%
ethanol containing 0.1% acetic acid, and the absorbance was
measured spectrophotometrically at 570 nm.
Cell viability assay
The effects of chemotherapeutic agents, DTX and CIS,
on cell viability were assessed using the methylthiazol MTT method.
Tumor cells were seeded in 24-well plates at a density of
2×104 cells per well and treated with DTX (0, 1, 5 and
10 nM) for 48 h or CIS (0, 5, 10 and 20 µM) for 24 h at 37°C.
Following treatment, 200 µl of MTT solution
[3-(4,5-dimethylthiazol-2-yl)-2,5-diphenyltetrazolium bromide (cat.
no. M6494; Invitrogen; Thermo Fisher Scientific, Inc.); 1 g/l in
PBS] was added to each well and incubated at 37°C for 4 h. The
resulting formazan crystals were solubilized in DMSO (cat. no.
C6164, Merck KGAa), and absorbance was measured at 570 nm using an
enzyme-linked immunosorbent assay reader (DYNEX®
Technologies). Cell viability (%) was calculated as (T/U) × 100,
where T represents the absorbance of treated cells and U represents
absorbance of untreated cells.
Western blot assay
Total protein lysates from homogenized tumor tissues
or cultured cells were extracted using RIPA lysis and Extraction
buffer (cat. no. 89901; Thermo Fisher Scientific, Inc.)
supplemented with a proteinase inhibitor cocktail (cat. no.
11836170001; Roche Diagnostics GmbH). Protein concentrations were
quantified using the Protein Assay kit (cat. no. 5000006EDU;
Bio-Rad Laboratories, Inc.) by measuring absorbance at 595 nm.
Equal amounts of protein (20 µg) were resolved by 9.5% SDS-PAGE
gels and transferred onto PVDF membranes (cat. no. 88518; Thermo
Scientific, Inc.) at 400 mA for 3 h. Membranes were blocked with 5%
skimmed milk in TBS-0.05% Tween (TBST) for 1 h at room temperature,
and incubated with primary antibodies at 4°C for 16–24 h. The
primary antibodies used were as follows: Anti-LUZP1 (1:1,000; cat.
no. 17483-1-AP; Proteintech Group, Inc.), anti-E-cadherin (2,000;
cat. no. 3195; Cell Signaling Technology, Inc.), anti-vimentin
(2,000; cat. no. 3932; Cell Signaling Technology Inc.) and
anti-β-actin (10,000; cat. no. 4970; Cell Signaling Technology,
Inc.). After three washes with TBST (15 min each), membranes were
incubated with horseradish peroxidase-conjugated secondary
antibody, peroxidase AffiniPure Goat Anti-Rabbit IgG (H+L) (10,000;
cat. no. 111-035-144; Jackson ImmunoResearch Laboratories, Inc.) at
room temperature for 1 h. Immunoreactive bands were visualized
using an enhanced chemiluminescence substrate (Western Lighting
Plus ECL; cat. no. 0RT2655, PerkinElmer, Inc.) and captured with an
ImageQuant LAS 4000 luminescence image system (Cytiva). Original
western blot images are provided in Table SI.
Chromatin immunoprecipitation followed
by quantitative PCR (ChIP-qPCR)
ChIP-qPCR was performed to assess NF-κB p65 (RelA)
occupancy at selected genomic loci in OECM-1 cells using the
SimpleChIP Enzymatic Chromatin IP Kit (Magnetic Beads) (cat. no.
9003; Cell Signaling Technology, Inc.). OECM-1 cells were cultured
under standard conditions, stimulated with TNF-α (10 ng/ml; cat.
no. C01047-100UG; Croyez Bioscience Co., Ltd.) for 24 h. Cells were
cross-linked with 1% formaldehyde at room temperature for 10 min
and quenched with glycine. Nuclei were isolated, and chromatin was
enzymatically digested with micrococcal nuclease at 37°C for 25
min, followed by sonication (40 kHz on ice) using eight cycles of
20 s ON and 30 s OFF to generate chromatin fragments ranging from
150 to 900 bp, as confirmed by 1% agarose gel electrophoresis. A 2%
aliquot of sheared chromatin was processed in parallel as an input
control. For immunoprecipitation, 10 µg of chromatin was incubated
overnight at 4°C with 5 µl of ChIP-grade anti-NF-κB p65 antibody
(cat. no. 8242; Cell Signaling Technology, Inc.).
Immunoprecipitation with an anti-histone H3 antibody (cat. no.
4620; Cell Signaling Technology, Inc.) was included as a positive
control. Immune complexes were captured using ChIP-grade Protein G
magnetic beads (cat. no. 9006; Cell Signaling Technology, Inc.) and
subjected to sequential low-salt and high-salt washes. Bound
chromatin was eluted and reverse cross-linked at 65°C for 2 h in
the presence of proteinase K, followed by DNA purification using
spin columns and elution in DNA elution buffer. qPCR was performed
using a QuantStudio 5 Real-Time PCR System (Applied Biosystems,
Thermo Fisher Scientific, Inc.) with KAPA SYBR FAST qPCR Master Mix
(cat. no. KM4106; Roche Diagnostics). Each reaction contained 1 µl
of ChIP or input DNA in a total volume of 10 µl. Technical
triplicates were averaged and enrichment was calculated as a
percentage of input using ΔCq normalization (19) with adjustment for the 2% input
dilution. Information of qPCR primers is provided in Table SII.
Gene knockdown by shRNA
Lentiviral plasmids pCMV-ΔR8.91 and pMD.G, along
with specific short hairpin RNA (shRNA) constructs in the pLKO.1
vector, were obtained from the National RNAi Core Facility,
Academia Sinica. A GFP-targeting shRNA (cat. no. 30323; pLKO.1 GFP
shRNA) was used as a control and was purchased from Addgene, Inc.
The shRNA sequences used in this study are listed in Table SIII. For lentivirus production,
HEK293T cells were transiently co-transfected with the shRNA
expression plasmid and the packaging plasmids pCMV-ΔR8.91 and pMD.G
at a plasmid ratio of 4 µg:3.6 µg:0.4 µg (shRNA:pCMV-ΔR8.91:pMD.G).
Plasmid DNA was mixed with Lipofectamine® 2000
(Invitrogen; Thermo Fisher Scientific, Inc.) and incubated at RT
for 20 min to allow complex formation, followed by transfection,
according to the manufacturer's instructions. A total of 48 h after
transfection at 37°C, the viral supernatants were collected,
filtered through a 0.22 µm membrane filter, and used for subsequent
infection. HNSCC cells were infected with shRNA-containing viral
supernatant in the presence of polybrene (8 µg/ml). After 24 h of
incubation at 37°C, the medium was replaced with fresh complete
medium containing puromycin (2 µg/ml) for 2 weeks for selection.
Cells were maintained under puromycin selection and harvested for
downstream analyses as indicated in each experiment.
Immunofluorescent analysis
Tumor cells cultured on cover glasses or in 6-well
plates were rinsed three times with PBS and fixed in 4%
formaldehyde (cat. no. 1.04003; Merck KGAa) at room temperature for
7 min. Following fixation, cells were washed three times with PBS
(5 min each). Permeabilization was performed with 0.1% Triton X-100
(cat. no. T8532; Merck KGAa) in PBS for 7 min, at room temperature,
followed by three additional PBS washes. Non-specific binding was
blocked with 1% BSA in PBS at room temperature for 1 h or at 4°C
overnight. Tumor cells were then incubated with LUZP1 antibodies
(1:100 dilution in 1% BSA/PBS; cat. no. 17483-1-AP, Proteintech
Group Inc.) at room temperature for 1 h, washed and subsequently
incubated with fluorophore-conjugated secondary antibodies (1:400
dilution in 1% BSA/PBS; cat. no. A-11008, Thermo Fisher Scientific,
Inc.) at room temperature in the dark for 1 h. F-actin was
visualized by staining with fluorescent phalloidin (1:200 dilution
in 1% PBS; cat. no. R415; Invitrogen; Thermo Fisher Scientific,
Inc.) at room temperature in the dark for 30 min. After three final
PBS washes, cover glasses were mounted onto slides using
DAPI-containing mounting medium (cat. no. 20004; AAT Bioquest,
Inc.), sealed with nail polish and stored at 4°C until imaging.
Fluorescent images were acquired using a fluorescence microscope
(Nikon Corporation).
Immunohistochemical analysis
Formalin-fixed, paraffin-embedded clinical tissue
specimens were sectioned at 4-µm thickness and baked at 60–65°C for
1 h, followed by a 10 min cooling step. Slides were deparaffinized
twice in xylene (cat. no. 4312, Muto Pure Chemicals Co. Ltd.) 10
min each and rehydrated through a graded ethanol series (cat. no.
459836, Merck KGAa; 100, 95, 75 and 50%; 5 min each), then rinsed
in double distilled H2O for 5 min. Heat-induced epitope
retrieval was performed in either Tris buffer (pH 9.0; cat. no.
920p-06, Merck KGAa) or citrate buffer (pH 6.0; cat. no. CBB500;
ScyTek Laboratories, Inc.) under pressure at 95–100°C for 15 min,
followed by cooling at room temperature for 30 min. Endogenous
peroxidase activity was quenched with 3% hydrogen peroxide (cat.
no. 31642, Thermo Fisher Scientific Inc.) for 10 min and sections
were washed three times with PBS (10 min each). After blocking with
5% normal goat serum (cat. no. 005-000-121; Jackson ImmunoResearch
Laboratories, Inc.) for 1 h at room temperature, slides were
incubated overnight at 4°C with anti-LUZP1 primary antibody (1:15
dilution; cat. no. 17483-1-AP, Proteintech Group Inc.). Following
three PBS washes, sections were incubated with biotinylated goat
anti-rabbit IgG (1:400; cat. no. BA-1000; Vector Laboratories,
Inc.) at room temperature for 1 h, and signal amplification was
performed using the ABC-HRP kit (cat. no. K-6100; Vector
Laboratories, Inc.). Visualization was achieved with Gill's
hematoxylin (cat. no. 3801520, Leica Biosystems) for nuclear
counterstaining. Finally, slides were dehydrated through graded
ethanol (cat. no. 459836; Merck KGAa; 75, 95 and 100%; 3 min each),
cleared in xylene (cat. no. 4312, Muto Pure Chemicals Co. Ltd.) 10
min twice and mounted with coverslips. Immunohistochemical images
were acquired using a Zeiss Axio Imager Z2 upright fluorescence
microscope (Duke) and quantitative analysis was conducted using
Tissue FAXS cell analysis software (version, 7.1; Tissuegnostics).
The immunohistochemical signals were scored as 0, 1, 2 and 3, and a
score ≥2 indicated positive detection.
Proteomic analysis
Total protein extracts (20 µg per sample) were
separated on a 9.5% SDS-PAGE gel and divided into four gel
fractions for in-gel tryptic digestion as previously described
(20). Peptide mixtures were
analyzed using an Orbitrap Exploris 480 mass spectrometer (Thermo
Scientific, Inc.) coupled with an Ultimate 3000 RSLC nano-liquid
chromatography system (Thermo Scientific, Inc.) equipped with a
nanospray ionization source (Thermo Scientific, Inc.). Mass
spectrometric analyses were performed in positive ion mode under
data-dependent acquisition settings. Full mass spectrometry survey
scans were acquired in the m/z range of 375-1,500 with an automatic
gain control (AGC) target of 4×105 and a resolution of
120,000 at m/z 200. The twenty most abundant multiple-charged
precursor ions were sequentially fragmented by collision-induced
dissociation and the tandem mass was analyzed in the Orbitrap at a
resolution of 30,000. Protein identification and label-free
quantification were conducted using Proteome Discovery software
(version, 2.4; Thermo Scientific, Inc.), with the identification
threshold set at P<0.05. Label-free quantification was conducted
using Proteome Discoverer version 2.4 (Thermo Fisher Scientific,
Inc.) incorporating the Minora feature detection algorithm.
Quantitative analysis was based on the peak abundances of precursor
ions corresponding to the identified peptides. Proteins exhibiting
significant upregulation in the experimental group were subjected
to identify significantly enriched biological pathways by Kyoto
Encyclopedia of Genes and Genomes (KEGG) pathway enrichment
analysis in the DAVID website (https://davidbioinformatics.nih.gov/).
Statistical analysis
All data are presented as the means ± SD.
Statistical significance between two independent groups was
assessed using a two-tailed unpaired Student's t-test, whereas
comparisons among multiple groups were performed using one-way
analysis of variance (ANOVA) followed by Tukey's post hoc test.
One-to-many comparisons were conducted using one-way ANOVA followed
by Dunnett's post hoc test. A factorial two-way ANOVA was performed
to assess the effects of two independent variables, followed by
Tukey's honestly significant difference post hoc test for multiple
comparisons among groups. Correlations between gene expression
levels were assessed using Spearman's rank correlation coefficient.
Overall survival (OS) and progression-free survival (PFS) were
analyzed using the Kaplan-Meier method, and differences between
survival curves were evaluated by the log-rank test. All
statistical analyses were conducted using SPSS Statistics software
(version, 22.0; IBM, Corp.). P<0.05 was considered to indicate a
statistically significant difference.
Results
LUZP1 is highly expressed in HNSCC
tumor tissues with short-term survival
To identify protein signatures potentially
associated with poor prognosis, a comparative proteomic analysis of
tumor tissues from patients with HNSCC was performed with long- and
short-term survival. Patients with an overall survival of <3
years were classified as the short-term survival group, whereas
those with an overall survival of ≥5 years were defined as the
long-term survival group. Among the 4,246 proteins identified, 187
exhibited a ≥4-fold increase in abundance in short-term survival
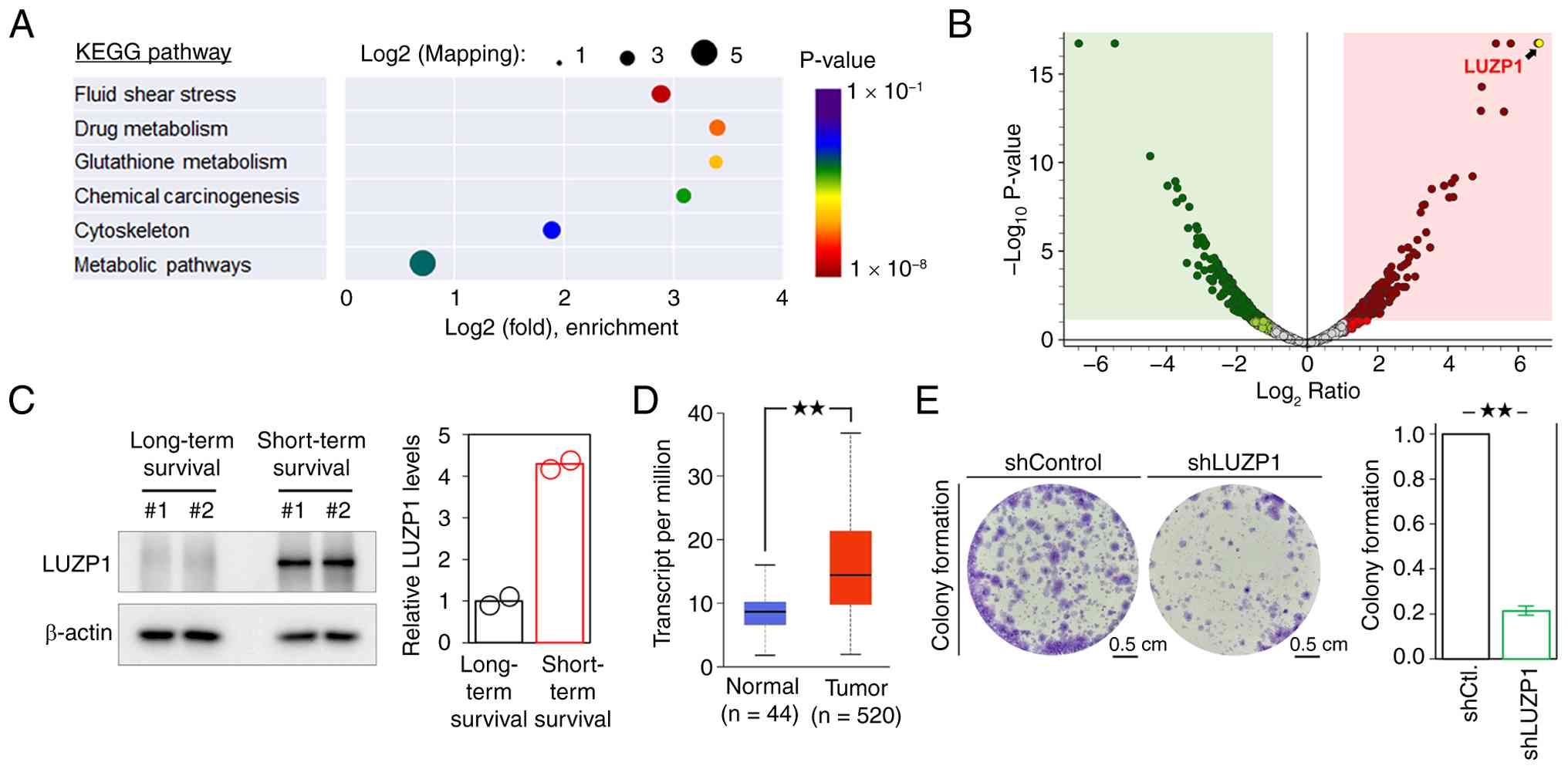
compared with long-term survival tissues (Table SIV). KEGG pathway enrichment
analysis revealed that these upregulated proteins were primarily
associated with pathways related to fluid shear stress, drug
metabolism, glutathione metabolism and chemical carcinogenesis
(Fig. 1A). Fluid shear stress is
associated with the metastatic dissemination of tumor cells within
the circulatory system (21),
whereas drug metabolism is associated with increased therapeutic
resistance (22). These findings
suggest that tumor metastasis and treatment resistance may
represent key determinants underlying poor prognosis and short-term
survival in patients with HNSCC. Differential protein expression
between short- and long-term survival tissues was visualized using
a proteomic volcano plot (Fig. 1B),
where the x-axis represents the log2 fold change in
protein abundance and the y-axis represents the -log10
p-value, indicating statistical significance. LUZP1 emerged as one
of the most upregulated proteins in tumor tissues of patients with
HNSCC with short-term survival (Fig.
1B; Table SIV). The elevated
expression of LUZP1 was further confirmed in additional tumor
specimens by western blot analysis (Fig. 1C). The LUZP1 mRNA expression level
of HNSCC tissues was significantly higher when compared with that
in the normal tissues via the UALCAN database (Fig. 1D), which provides an in-depth
analysis of gene expression data from TCGA (17). Additionally, LUZP1 gene knockdown by
shRNA significantly impaired the colony formation of OECM-1 cells
(Fig. 1E), suggesting that LUZP1
could enhance the capacity for unlimited proliferation of HNSCC
cells (23).
LUZP1 enhances the metastatic
abilities of HNSCC cells
Since LUZP1 may contribute to HNSCC metastasis, its
effects on the metastatic properties of HNSCC cells was examined. A
one-way ANOVA analysis of the Transwell assays revealed a
significant difference among Control, shLUZP1#1 and shLUZP1#2
groups (migration: F(2,6)=142.71, P=9.0×10−6; invasion:
F(2,6)=193.09, P=4.0×10−6). Post hoc comparisons using
Dunnett's test demonstrated that both shLUZP1#1 and shLUZP1#2 were
significantly different from the Control group (shLUZP1#1 vs.
Control, P=1.14×10−4; shLUZP1#2 vs. Control,
P=1.82×10−4), indicating that knockdown markedly reduced
the migration and invasion abilities of HNSCC cells (Fig. 2A and B). Transwell assays
demonstrated that LUZP1 knockdown markedly attenuated cell
migration and invasion (Fig. 2A and
B). 3D tumor spheroid invasion assay was conducted to validate
the role of LUZP1 in regulating tumor metastatic abilities. A
one-way ANOVA revealed a significant difference among Control,
shLUZP1#1 and shLUZP1#2 groups (F(2,6)=74.81,
P=5.7×10−5). Dunnett's post hoc test demonstrated that
both shLUZP1#1 and shLUZP1#2 were significantly reduced compared
with the Control group (shLUZP1#1 vs. Control,
P=2.6×10−4 and shLUZP1#2 vs. Control,
P=1.1×10−4), indicating that LUZP1 knockdown
significantly attenuated the invasive capacity of tumor spheroids
(Fig. 2C), consistent with the
previous two-dimensional migration and invasion assays (Fig. 2A and B). In addition, LUZP1
knockdown reduced the formation of invadopodia-like protrusive
structures (Fig. 2D), visualized by
phalloidin staining of the key cytoskeletal component F-actin
(24). Furthermore, LUZP1 knockdown
increased the expression of the epithelial marker E-cadherin while
decreasing the expression of the mesenchymal marker vimentin
(Fig. 2E), suggesting a role for
LUZP1 in promoting EMT. Moreover, analysis of paired primary tumors
and metastatic sites from the same patients with HNSCC revealed
markedly increased LUZP1 expression in metastatic lesions compared
with their matched primary tumors (Fig.
2F). This observation further supports the notion that LUZP1
enhances the metastatic potential of HNSCC cells.
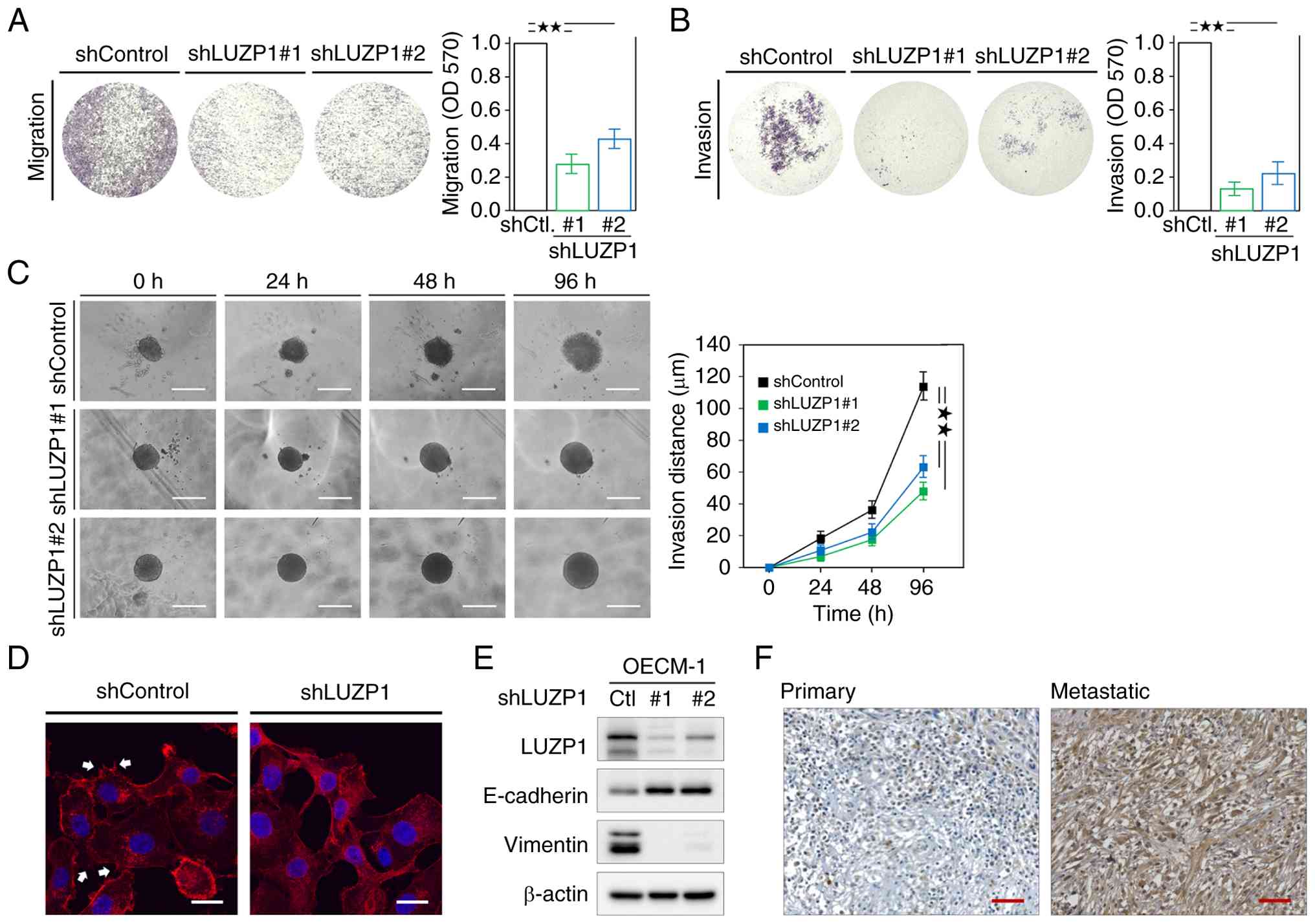 | Figure 2.LUZP1 enhances the metastatic
abilities of HNSCC cells. (A) cell migration and (B) invasion assay
in OECM-1 cells with or without LUZP1 knockdown. Signal
quantification using crystal violet extract was measured by
colorimetric analysis at 570 nm. The relative signal intensities
were normalized to the shControl (n=3). (C) The representative
images for 3D invasion assay for tumor spheroids in SAS cells with
or without LUZP1 knockdown. A radial outgrowth distance approach is
used for quantification of 3D tumor spheroid invasion. Scale bar,
500 µm. (D) Invadopodia-like protrusive structures were visualized
by F-actin staining using phalloidin (cat. no. R415; Invitrogen;
Thermo Fisher Scientific, Inc.), which are indicated by the arrow.
Scale bar, 20 µm. (E) The expression of LUZP1, E-cadherin and
vimentin in OECM-1 cells with or without LUZP1 knockdown was
determined by western blot assay. β-actin, loading control. (F) The
expression of LUZP1 in both primary and metastatic tumors of a
patient with HNSCC was examined using immunohistochemical analysis.
Scale bar, 50 µm. Data are shown as the means ± SD. For statistical
analyses, (A and B) a 2-tailed unpaired Student's t-test; (C)
one-way ANOVA with Tukey's post hoc test. **P<0.01. LUZP1,
leucine zipper protein 1; HNSCC, head and neck squamous cell
carcinoma; sh, short hairpin; Ctl., control. |
LUZP1 increases chemoresistance of
HNSCC cells
KEGG pathway analysis of upregulated proteins
identified in tumor tissues from short-term survival patients with
HNSCC revealed significant enrichment of pathways associated with
increased therapeutic resistance (Fig.
1A). This prompted the investigation of whether elevated LUZP1
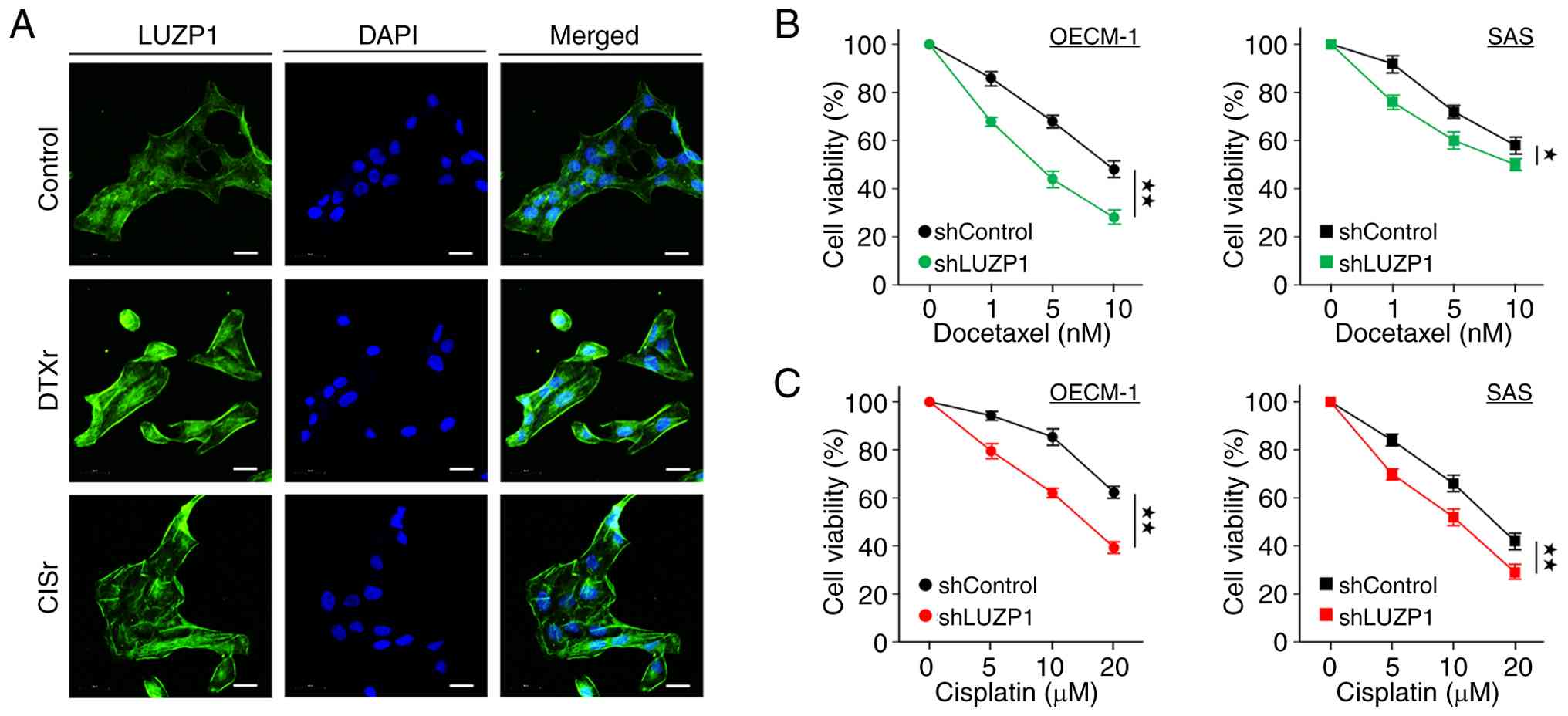
expression contributes to enhanced chemoresistance in HNSCC. To
this end, DTXr and CISr OECM-1 cell lines were established by
continuous exposure to DTX or CIS, two chemotherapeutic agents
commonly used in HNSCC treatment (25). Immunofluorescence analysis
demonstrated that LUZP1 expression was markedly elevated in both
OECM-1/DTXr and OECM-1/CISr cells compared with their parental
counterparts (Fig. 3A).
Furthermore, MTT assays revealed that LUZP1 knockdown increased the
sensitivity of OECM-1 and SAS cells to DTX and CIS (Fig. 3B and C), indicating that LUZP1 may
play a functional role in mediating chemoresistance in HNSCC.
NF-κB signaling activation promotes
LUZP1 expression in HNSCC cells
Based on the aforementioned findings, the signaling
pathways associated with metastasis and chemoresistance that may
regulate LUZP1 expression were investigated using pathway-specific
inhibitors. A total of five inhibitors were employed in the present
study, including SB431542 for the TGF-β/Smad pathway, LY294002 for
PI3K/AKT/mTOR signaling, Rapamycin for mTOR, BAY 11-7085 for NF-κB
signaling and YC-1 for the HIF-1 pathway. Semi-quantitative western
blot analysis demonstrated that BAY 11-7085 exerted the most
pronounced inhibitory effect on LUZP1 expression across all three
HNSCC cells (FaDu, OECM-1 and SAS; Fig.
4A), and repeated treatments with BAY 11-7085 consistently
reproduced this inhibitory effect on LUZP1 protein levels (Fig. 4B). These results suggest that LUZP1
expression may be regulated by NF-κB signaling activation.
Consistently, in silico analysis using the GEPIA database
revealed a significant positive correlation between LUZP1
expression and that of NF-κB p105 (NFKB1) and NF-κB p100 (NFKB2)
subunits (Fig. 4C). Based on the
aforementioned findings that LUZP1 knockdown increased the
sensitivity of HNSCC cells to DTX and CIS (Fig. 3B and C), validation of the
contribution of NF-κB-LUZP1 signaling to chemoresistance was next
sought. To this end, NF-κB p65 was silenced using gene knockdown by
shRNA. Western blot analysis showed that NF-κB knockdown resulted
in a marked reduction of LUZP1 expression (Fig. 4D). Consistently, MTT assays
demonstrated that NF-κB knockdown significantly enhanced the
sensitivity of both OECM-1 and SAS cells to DTX and CIS, leading to
a pronounced decrease in drug IC50 values (Fig. 4E and F). These results support that
NF-κB-LUZP1 signaling may play a key role in mediating
chemoresistance in HNSCC cells. To further validate the regulation
of LUZP1 by NF-κB signaling, HNSCC cells were stimulated with the
cytokines IL-1β and TNF-α, both known activators of NF-κB signaling
(26). Western blot analysis showed
that IL-1β and TNF-α both induced LUZP1 expression in a
time-dependent manner (Fig. 4G and
H). Moreover, to examine the functional consequences of
NF-κB-mediated regulation of LUZP1, cell migration following LUZP1
knockdown and/or TNF-α stimulation was assessed. Using a two-way
ANOVA to assess the effects of shLUZP1 and TNFα treatment,
significant main effects of shLUZP1 (F(1,8)=68.87,
P=3.35×10−5) and TNFα (F(1,8)=216.30,
P=4.49×10−7) were observed, whereas the interaction term
did not reach significant (F(1,8)=0.026, P=0.877) (Fig. 4I). Tukey's multiple comparisons post
hoc test confirmed significant differences among all four groups
(adjusted P≤0.01). The results indicated that TNF-α treatment
enhanced the migratory ability of parental HNSCC cells and
effectively rescued the migration defect caused by LUZP1 knockdown
(Fig. 4I).
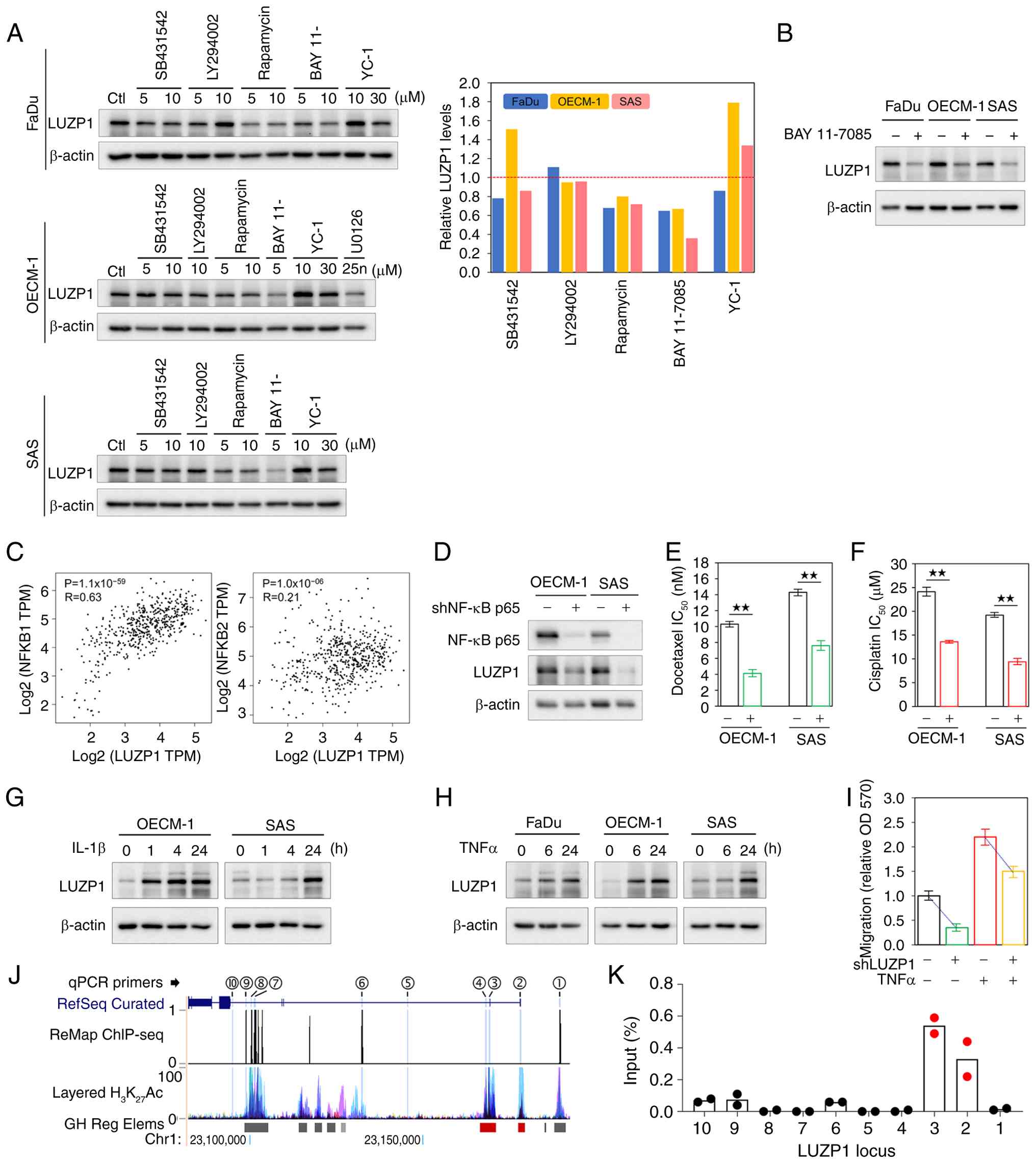 | Figure 4.NF-κB signaling activation promotes
LUZP1 expression in HNSCC cells. (A) The expression of LUZP1 in
FaDu, OECM-1 and SAS cells treated with SB431542 (10 µM), LY294002
(10 µM), Rapamycin (10 µM), BAY 11–7085 (5 µM) or YC-1 (30 µM) was
determined by western blot assay. Signal quantification was
measured using ImageJ 1.54 g software (National Institutes of
Health) and the relative intensity was normalized to untreated
control. The red dashed line represents the normalized value as 1.
(B) The expression of LUZP1 in FaDu, OECM-1 and SAS cells with or
without BAY 11–7085 was validated by western blot assay. (C)
Spearman's monotonic correlation between LUZP1 and NFKB1 or NFKB2
expression in HNSCC was analyzed using The Cancer Genome Atlas
RNA-Sequencing database on the GEPIA server. (D) Protein expression
of NF-κB p65 and LUZP1 in OECM-1 and SAS cells with NF-κB p65
knockdown (+) or control shRNA -), as determined by western blot
analysis. β-actin, loading control. (E) IC50 values of
docetaxel in OECM-1 and SAS cells with or without NF-κB p65
knockdown. (F) IC50 values of cisplatin in OECM-1 and
SAS cells with or without NF-κB p65 knockdown. The expression of
LUZP1 in different HNSCC cells treated with (G) IL-1β (3 ng/ml) or
(H) TNFα (10 ng/ml) for the indicated times was determined by
western blot assay. β-actin, loading control. (I) Transwell cell
migration assay was conducted using OECM-1 cells with or without
LUZP1 knockdown in the presence or absence of TNFα (10 ng/ml)
treatment. Signal quantification using crystal violet extract was
measured by colorimetric analysis at 570 nm, and the relative
signal intensities were normalized to untreated shControl (shLUZP1,
-; TNFα, -) (n=3). (J) Genomic visualization of the human LUZP1
locus (GRCh38/hg38) showing RefSeq-curated exon annotations, NF-κB
RelA ChIP-seq binding signals in FaDu cells (ReMap), layered
H3K27ac ChIP-seq profiles from ENCODE cell lines, and GeneHancer
regulatory element annotations. Red boxes denote promoter regions
and gray boxes indicate putative enhancers. Blue vertical bars mark
the locations of ChIP-qPCR primer sets designed for experimental
validation. (K) ChIP-qPCR analysis showing increased NF-κB (RelA)
occupancy at the LUZP1 promoter in response to TNF-α treatment. For
statistical analyses, (E and F) a 2-tailed unpaired Student's
t-test; (I) a factorial two-way ANOVA, followed by Tukey's
Honestly Significant Difference post hoc test. **P<0.01. TPM,
transcripts per million; ChIP-seq, chromatin immunoprecipitation
sequencing; LUZP1, leucine zipper protein 1; sh, short hairpin. |
To examine whether LUZP1 is transcriptionally
regulated by NF-κB. An in silico analysis was first carried
out to evaluate potential NF-κB-mediated regulation of the LUZP1
locus. Genomic snapshots of the human genome (GRCh38/hg38 assembly)
spanning the LUZP1 region (Chr 1:23,082,260-23,191,800) were
obtained from the UCSC Genome Browser (27). The resulting genomic visualization
(Fig. 4J) integrates multiple
regulatory features, including RefSeq-curated exon annotations
(28) of LUZP1, NF-κB RelA ChIP-seq
binding signals in FaDu cells from the ReMap database (29), and layered H3K27ac ChIP-seq profiles
from several ENCODE cell lines (30), which mark active regulatory regions
and are associated with transcriptional activation. Additionally,
GeneHancer regulatory element annotations (31) were included, with red boxes denoting
promoter regions and gray boxes indicating putative enhancers.
Together, this analysis revealed potential NF-κB binding sites
within the promoter and enhancer regions of LUZP1, suggesting a
regulatory relationship between NF-κB and LUZP1 expression. Based
on these sequence features, 10 sets of ChIP-qPCR primers were
designed targeting candidate NF-κB-enriched regions (indicated by
blue vertical bars in Fig. 4J) for
experimental validation. ChIP-qPCR analysis demonstrated that TNF-α
stimulation significantly enhanced NF-κB binding at the LUZP1
promoter (Fig. 4K), indicating that
NF-κB occupancy at the LUZP1 promoter is inducible and context
dependent. Collectively, these results demonstrate that the
oncogenic roles of LUZP1 could be mediated through NF-κB signaling
activation.
LUZP1 is associated with poor survival
of patients with HNSCC
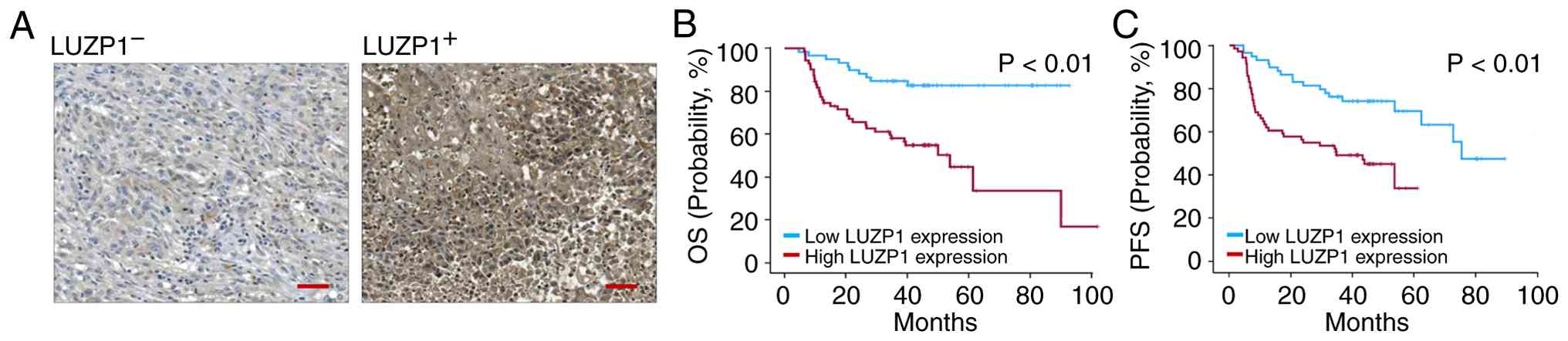
To assess the impact of LUZP1 on clinical outcomes,
IHC assays were conducted in tissue specimens to determine the
correlation between LUZP1 expression levels and OS and PFS in
patients with HNSCC. A total of 130 pathologic specimens were
collected from patients with HNSCC, with demographic information
shown in Table I. Representative
IHC staining images are shown in Fig.
5A. IHC signals were semi-quantitatively scored as 0, 1, 2 or
3, and a score ≥2 was defined as positive (LUZP1+), as
described in the Materials and methods section. The Kaplan-Meier
plot analysis revealed that LUZP1+ was significantly
associated with shorter OS and PFS in the present study cohort
(Fig. 5B and C).
To determine whether LUZP1 expression independently
predicts patient outcomes, multivariate Cox proportional hazards
regression analyses were conducted with adjustment for age, T
stage, nodal status, lymphovascular invasion (LVI), tumor
differentiation, perineural invasion (PNI) and surgical margin
status. High LUZP1 expression emerged as an independent prognostic
factor for both OS and PFS (Tables
II and III). Specifically,
patients with high LUZP1 expression exhibited a significantly
increased risk of mortality (OS; HR, 1.789–6.006; P<0.001) and
disease progression or recurrence (PFS; HR, 2.202–9.924;
P<0.001), independent of established clinicopathological
variables. By contrast, tumor differentiation, LVI, PNI and
surgical margin status did not retain independent prognostic
significance for either OS or PFS in the multivariate models. In
multivariate Cox regression analyses (Tables II and III), advanced nodal disease (N3)
remained significantly associated with worse survival outcomes.
Specifically, N3 was independently associated with reduced overall
survival compared with N0 (Table
II, P=0.029; Table III,
P=0.004). Importantly, the prognostic impact of LUZP1 expression
persisted after adjustment for nodal status and tumor stage,
indicating that LUZP1 provides prognostic information beyond
conventional staging parameters.
 | Table II.Multivariate Cox regression analysis
of overall survival in patients with head and neck squamous cell
carcinoma. |
Table II.
Multivariate Cox regression analysis
of overall survival in patients with head and neck squamous cell
carcinoma.
|
|
| 95.0% CI |
|---|
|
|
|
|
|---|
|
Characteristics | P-value | Low | Upper |
|---|
| Age |
|
|
|
| 40-60
vs. <40 | 0.364 | 0.281 | 1.594 |
| 40-60
vs. >60 | 0.108 | 0.166 | 1.195 |
| T | 0.095 |
|
|
| T4 vs.
T1 | 0.039 | 0.062 | 0.933 |
| T4 vs.
T2 | 0.045 | 0.186 | 0.981 |
| T4 vs.
T3 | 0.486 | 0.273 | 1.854 |
| Nodal | 0.012 |
|
|
| N3 vs.
N0 | 0.029 | 0.135 | 0.896 |
| N3 vs.
N1 | 0.092 | 0.056 | 1.243 |
| N3 vs.
N2 | 0.433 | 0.601 | 3.279 |
| LVI |
|
|
|
| No vs.
yes | 0.138 | 0.841 | 3.471 |
|
Differentiation | 0.210 |
|
|
| Well
vs. moderate | 0.918 | 0.495 | 1.884 |
| Well
vs. poor | 0.101 | 0.166 | 1.172 |
| PNI |
|
|
|
| No vs.
yes | 0.099 | 0.895 | 3.602 |
| Margin |
|
|
|
| Free
vs. positive | 0.244 | 0.357 | 1.300 |
| LUZP1 |
|
|
|
| Low vs.
High | <0.001 | 2.202 | 9.924 |
 | Table III.Multivariate Cox regression analysis
of progression-free survival in patients with head and neck
squamous cell carcinoma. |
Table III.
Multivariate Cox regression analysis
of progression-free survival in patients with head and neck
squamous cell carcinoma.
|
|
| 95.0% CI |
|---|
|
|
|
|
|---|
|
Characteristics | P value | Low | Upper |
|---|
| Age |
|
|
|
| 40-60
vs. <40 | 0.395 | 0.672 | 2.740 |
| 40-60
vs. >60 | 0.160 | 0.244 | 1.263 |
| T |
|
|
|
| T4 vs.
T1 | 0.424 | 0.250 | 1.791 |
| T4 vs.
T2 | 0.817 | 0.295 | 2.617 |
| T4 vs.
T3 | 0.731 | 0.436 | 3.264 |
| Nodal |
|
|
|
| N3 vs.
N0 | 0.004 | 0.122 | 0.665 |
| N3 vs.
N1 | 0.120 | 0.164 | 1.231 |
| N3 vs.
N2 | 0.251 | 0.289 | 1.383 |
| LVI |
|
|
|
| No vs.
Yes | 0.133 | 0.870 | 2.856 |
|
Differentiation |
|
|
|
| Well vs
Moderate | 0.924 | 0.562 | 1.886 |
| Well
vs. Poor | 0.089 | 0.190 | 1.125 |
| PNI |
|
|
|
| No vs.
Yes | 0.639 | 0.639 | 2.074 |
| Margin |
|
|
|
| Free
vs. Positive | 0.835 | 0.591 | 1.917 |
| LUZP1 |
|
|
|
| Low vs.
High | <.001 | 1.789 | 6.006 |
Discussion
HNSCC is a biologically and clinically heterogeneous
malignancy and its poor prognosis is frequently driven by local
recurrence, distant metastasis and therapeutic resistance despite
the availability of multimodal treatment approaches. Identifying
molecular determinants of poor survival is therefore essential for
risk stratification and therapeutic guidance. The present study
suggests that LUZP1 functions as a key oncogenic regulator in
HNSCC, associating its expression with metastatic potential and
chemoresistance through NF-κB signaling activation. Notably, IHC
analysis of a patient cohort revealed that high LUZP1 expression
was significantly associated with shorter OS and PFS, suggesting
its potential utility as a prognostic biomarker.
LUZP1 was initially identified as an essential
regulator of embryonic development, as Luzp1-null mice
display neural tube closure defects and cardiovascular
abnormalities that result in perinatal lethality (11). Subsequent studies have shown that
LUZP1 functions as an actin-stabilizing protein and a negative
regulator of ciliogenesis (32),
and it has also been implicated in cell division, cell migration
and epithelial apical constriction (9). In Townes-Brocks syndrome, a
mechanistic study demonstrated that truncated SALL1 promotes
proteasomal degradation of LUZP1, thereby associating LUZP1
deficiency with impaired cilia regulation and providing a direct
pathway by which LUZP1 abnormality contributes to disease (33). Consistently, diseases associated
with LUZP1 deficiency predominantly affect organs that depend on
proper cytoskeletal dynamics and ciliary function, aligning with
its established molecular roles in actin stabilization, inhibition
of ciliogenesis and regulation of cell architecture and
motility.
Accumulating evidence indicates that LUZP1 exerts
context-dependent functions in cancer, yet its effects consistently
converge on the regulation of cytoskeletal dynamics, cell migration
and EMT-associated phenotypes (9,10). In
glioma, deletions or loss of LUZP1 function have been associated
with enhanced migration and invasion, likely through disruption of
actin organization, cell division and polarity control,
underscoring its role in maintaining cytoskeletal integrity
(34). By contrast, a study in
triple-negative breast cancer demonstrate that LUZP1 can
heterodimerize with DAPK3, protecting it from proteasomal
degradation and thereby indirectly promote EMT, desmosome
suppression and invasive behavior (13). Similarly, in osteosarcoma, LUZP1
acts as a downstream effector of circFIRRE-miRNA regulatory
circuits, facilitating tumor progression, angiogenesis and
metastatic dissemination (12).
Consistent with these reports, the functional analyses of the
present study show that LUZP1 knockdown markedly suppresses
migration and invasion of HNSCC cells, reduces invadopodia-like
protrusive structures and prevents EMT-associated marker
expression, as evidenced by E-cadherin upregulation and vimentin
downregulation. Moreover, the observation that LUZP1 expression is
elevated in metastatic lesions compared with matched primary tumors
further supports its involvement in metastatic dissemination.
Collectively, these findings suggest that although LUZP1 is not a
canonical EMT transcription factor, it may function as an
EMT-enabling factor by modulating actin cytoskeleton organization,
actomyosin contractility and cellular plasticity.
Importantly, the EMT-associated plasticity driven by
LUZP1 also provides a mechanistic basis for its contribution to
drug resistance. KEGG pathway analysis of proteins upregulated in
short-term survival tissues revealed significant enrichment of drug
metabolism and glutathione-associated pathways, both of which are
associated with chemoresistance. In line with these findings, the
present study experimentally confirmed that LUZP1 expression is
markedly elevated in DTZr and CISr OECM-1 cell lines. Conversely,
LUZP1 knockdown restored chemosensitivity in both OECM-1 and SAS
cells, indicating a functional role for LUZP1 in sustaining
resistant phenotypes. Chemoresistance in HNSCC is widely recognized
as a multifactorial process involving drug sequestration, efflux,
detoxification and activation of pro-survival signaling pathways
(35). Data of the present study
suggests that LUZP1 represents an integral component of this
resistance network. Rather than directly regulating drug metabolism
or efflux, LUZP1 may promote therapy tolerance by coordinating
cytoskeletal remodeling with cellular stress-adaptive programs.
Such remodeling could influence intracellular drug uptake or
trafficking or functionally cooperate with antioxidant and
metabolic pathways to buffer cytotoxic stress (36–38).
In addition, the functional convergence of LUZP1 with inflammatory
and stress-responsive signaling pathways, including
NF-κB-associated programs, may further reinforce anti-apoptotic
responses during treatment (39).
Together, these observations support a model in which LUZP1
integrates EMT-associated plasticity, cytoskeletal adaptation and
metabolic stress buffering to promote both metastatic progression
and therapeutic resistance in aggressive HNSCC, highlighting its
potential value as a predictive biomarker and therapeutic target
for chemoresistant disease.
NF-κB is a central regulator of inflammatory
signaling, tumor progression and therapeutic resistance in HNSCC
(26). It controls the
transcription of cytokines, survival factors and invasion-promoting
proteins, thereby shaping a tumor microenvironment that favors
malignant progression (40). The
present study identified NF-κB signaling as a key upstream
regulator of LUZP1 expression. Pharmacologic inhibition with BAY
11–7085 consistently produced the strongest suppression of LUZP1
across multiple HNSCC cell lines, whereas stimulation with IL-1β
and TNF-α, two potent NF-κB activators, induced LUZP1 expression in
a time-dependent manner and rescued the migration defect caused by
LUZP1 knockdown. These findings suggest that LUZP1 is a novel NF-κB
effector contributing to pro-metastatic and chemoresistant
phenotypes. The importance of this axis is reinforced by research
that shows IL-1β and TNF-α enhance EMT, stemness, migration,
invasion and metastatic outgrowth through NF-κB-driven programs
across multiple types of cancer (25,41–43).
IL-1β acts via PI3K/AKT and ZEB1-dependent pathways, while TNF-α
stabilizes Snail to accelerate epithelial-mesenchymal plasticity,
both converging on NF-κB activation. Collectively, the results of
the present study indicate that the NF-κB-LUZP1 axis integrates
inflammatory cytokine signaling with EMT and therapy resistance,
representing a potential therapeutic vulnerability in HNSCC.
The present study has several limitations that
should be acknowledged. Although the promotion of metastasis and
chemoresistance by LUPZ1 is demonstrated, the precise molecular
mechanisms remain to be clarified. Future investigations should
delineate how LUZP1 interacts with cytoskeletal and metabolic
networks at the molecular level. Furthermore, while NF-κB was
identified as an upstream regulator of LUZP1, it remains uncertain
whether additional signaling pathways also converge on its
regulation. Given the complexity of HNSCC signaling, it is
plausible that LUZP1 functions as an integrator of multiple
oncogenic inputs. Finally, although BAY 11–7085 effectively
suppressed LUZP1 expression in vitro, its clinical
translation is limited by issues of toxicity and specificity. The
development of more selective inhibitors or RNA-based therapeutic
approaches may therefore be required to target LUZP1 effectively in
the clinical setting.
In summary, the present study identifies LUZP1 as a
key oncogenic regulator in HNSCC that associates poor survival with
enhanced metastatic potential and chemoresistance. The present
study demonstrated that elevated LUZP1 expression promotes EMT,
invasion and drug resistance, and is significantly associated with
shorter OS and PFS in patient cohorts. Mechanistically, LUZP1
expression is driven, at least in part, by NF-κB signaling and can
be induced by pro-inflammatory cytokines such as IL-1β and TNF-α,
thereby integrating inflammatory cues with tumor aggressiveness.
These findings highlight LUZP1 not only as a potential prognostic
biomarker but also as a candidate therapeutic target in HNSCC.
Further studies are warranted to delineate the detailed molecular
mechanisms of LUZP1 regulation and to explore therapeutic
strategies for its clinical translation.
Supplementary Material
Supporting Data
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
All MS raw datasets generated in the present study
have been deposited in the jPOST (44) and ProteomeXchange repositories. The
accession numbers are JPST004371 for jPOST and PXD073924 for
ProteomeXchange. The data will be publicly accessible starting on 2
Feb 2026 and can be previewed via the jPOST repository using the
following link and access key: https://repository.jpostdb.org/preview/1096459080698079a08d66e
(access key: 6669).
Authors' contributions
CYL, CYH, YAT and WCC conceived the present study.
CYL, CYH, HCL, CCL, TTC and WCT performed the experiments. HCL, WCT
and YAT analyzed the data. TTC and WCT confirm the authenticity of
all the raw data. CYL, CYH, HCL, YAT and WCC wrote the manuscript.
WCC revised the manuscript. All authors read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study protocol was approved by the
Ethics Committee of China Medical University Hospital (Taichung,
Taiwan; approval no. CMUH108-REC1-127).
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Johnson DE, Burtness B, Leemans CR, Lui
VWY, Bauman JE and Grandis JR: Head and neck squamous cell
carcinoma. Nat Rev Dis Primers. 6:922020. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Liu YP, Zheng CC, Huang YN, He ML, Xu WW
and Li B: Molecular mechanisms of chemo- and radiotherapy
resistance and the potential implications for cancer treatment.
MedComm (2020). 2:315–340. 2021. View
Article : Google Scholar : PubMed/NCBI
|
|
3
|
Lee DY, Abraham J, Ross E, Ridge JA, Lango
MN, Liu JC, Bauman JR, Avkshtol V and Galloway TJ: Rapid recurrence
in head and neck cancer: Underappreciated problem with poor
outcome. Head Neck. 43:212–222. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Shirima CA, Bleotu C, Spandidos DA,
El-Naggar AK, Gradisteanu Pircalabioru G and Michalopoulos I:
Epithelial-derived head and neck squamous tumourigenesis (Review).
Oncol Rep. 52:1412024. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Duan XP, Qin BD, Jiao XD, Liu K, Wang Z
and Zang YS: New clinical trial design in precision medicine:
Discovery, development and direction. Signal Transduct Target Ther.
9:572024. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Park R and Chung CH: Advanced human
papillomavirus-negative head and neck squamous cell carcinoma:
unmet need and emerging therapies. Mol Cancer Ther. 23:1717–1730.
2024. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Tathineni P, Joshi N and Jelinek MJ:
Current state and future directions of EGFR-directed therapy in
head and neck cancer. Curr Treat Options Oncol. 24:680–692. 2023.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Wang Y, Han J, Zhu Y, Huang N and Qu N:
New advances in the therapeutic strategy of head and neck squamous
cell carcinoma: A review of latest therapies and cutting-edge
research. Biochim Biophys Acta Rev Cancer. 1880:1892302025.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Bozal-Basterra L, Gonzalez-Santamarta M,
Muratore V, Martín-Martín N, Ercilla A, Rodríguez JA, Carracedo A,
Sutherland JD and Barrio R: LUZP1 controls cell division, migration
and invasion through regulation of the actin cytoskeleton. Front
Cell Dev Biol. 9:6240892021. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang L, Tsang HY, Yan Z, Tojkander S,
Ciuba K, Kogan K, Liu X and Zhao H: LUZP1 regulates the maturation
of contractile actomyosin bundles. Cell Mol Life Sci. 81:2482024.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hsu CY, Chang NC, Lee MW, Lee KH, Sun DS,
Lai C and Chang AC: LUZP deficiency affects neural tube closure
during brain development. Biochem Biophys Res Commun. 376:466–471.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Yu L, Zhu H, Wang Z, Huang J, Zhu Y, Fan
G, Wang Y, Chen X and Zhou G: Circular RNA circFIRRE drives
osteosarcoma progression and metastasis through
tumorigenic-angiogenic coupling. Mol Cancer. 21:1672022. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang J, Tran-Huynh AM, Kim BJ, Chan DW,
Holt MV, Fandino D, Yu X, Qi X, Wang J, Zhang W, et al:
Death-associated protein kinase 3 modulates migration and invasion
of triple-negative breast cancer cells. PNAS Nexus. 3:pgae4012024.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Poel D, Boyd LNC, Beekhof R, Schelfhorst
T, Pham TV, Piersma SR, Knol JC, Jimenez CR, Verheul HMW and
Buffart TE: Proteomic analysis of miR-195 and miR-497 replacement
reveals potential candidates that increase sensitivity to
oxaliplatin in MSI/P53wt colorectal cancer cells. Cells.
8:11112019. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Chen M, Wang J and Xiao Y: Leucine zipper
protein 1 (LUZP1) serves as a prognostic biomarker for patients
with renal papillary cell carcinoma. Asian J Surg. 46:4011–4013.
2023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Yu D, Luo Y, Guo R, Ma F, Chang Y and Dang
J: Comprehensive profiling of endocrine metabolism identifies a
novel signature with robust predictive value in ovarian cancer. J
Gene Med. 26:e36862024. View
Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chandrashekar DS, Karthikeyan SK, Korla
PK, Patel H, Shovon AR, Athar M, Netto GJ, Qin ZS, Kumar S, Manne
U, et al: UALCAN: An update to the integrated cancer data analysis
platform. Neoplasia. 25:18–27. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Li C, Tang Z, Zhang W, Ye Z and Liu F:
GEPIA2021: integrating multiple deconvolution-based analysis into
GEPIA. Nucleic Acids Res. 49((W1)): W242–W246. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Schmittgen TD and Livak KJ: Analyzing
real-time PCR data by the comparative C(T) method. Nat Protoc.
3:1101–1108. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Tung JC, Huang WC, Yang JC, Chen GY, Fan
CC, Chien YC, Lin PS, Candice Lung SC and Chang WC: Auramine O, an
incense smoke ingredient, promotes lung cancer malignancy. Environ
Toxicol. 32:2379–2391. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Espina JA, Cordeiro MH, Milivojevic M,
Pajić-Lijaković I and Barriga EH: Response of cells and tissues to
shear stress. J Cell Sci. 136:jcs2609852023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hsieh CY, Lin CC and Chang WC: Taxanes in
the treatment of head and neck squamous cell carcinoma.
Biomedicines. 11:28872023. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Franken NA, Rodermond HM, Stap J, Haveman
J and van Bree C: Clonogenic assay of cells in vitro. Nat Protoc.
1:2315–2319. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fan CC, Cheng WC, Huang YC, Sher YP, Liou
NJ, Chien YC, Lin PS, Lin PS, Chen CH and Chang WC: EFHD2 promotes
epithelial-to-mesenchymal transition and correlates with
postsurgical recurrence of stage I lung adenocarcinoma. Sci Rep.
7:146172017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hsieh CY, Lin CC, Huang YW, Chen JH, Tsou
YA, Chang LC, Fan CC, Lin CY and Chang WC: Macrophage secretory
IL-1β promotes docetaxel resistance in head and neck squamous
carcinoma via SOD2/CAT-ICAM1 signaling. JCI Insight. 7:e1572852022.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Liu T, Zhang L, Joo D and Sun SC: NF-κB
signaling in inflammation. Signal Transduct Target Ther.
2:170232017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Casper J, Speir ML, Raney BJ, Perez G,
Nassar LR, Lee CM, Hinrichs AS, Gonzalez JN, Fischer C, Diekhans M,
et al: The UCSC genome browser database: 2026 update. Nucleic Acids
Res. 54:D1331–D1335. 2026. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Pruitt KD, Brown GR, Hiatt SM,
Thibaud-Nissen F, Astashyn A, Ermolaeva O, Farrell CM, Hart J,
Landrum MJ, McGarvey KM, et al: RefSeq: An update on mammalian
reference sequences. Nucleic Acids Res. 42:D756–D763. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Hammal F, de Langen P, Bergon A, Lopez F
and Ballester B: ReMap 2022: A database of Human, Mouse, Drosophila
and Arabidopsis regulatory regions from an integrative analysis of
DNA-binding sequencing experiments. Nucleic Acids Res.
50:D316–D325. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Moore JE, Pratt HE, Fan K, Phalke N,
Fisher J, Elhajjajy SI, Andrews G, Gao M, Shedd N, Fu Y, et al: An
expanded registry of candidate cis-regulatory elements. Nature. Jan
7–2026.(Epub ahead of print). View Article : Google Scholar
|
|
31
|
Fishilevich S, Nudel R, Rappaport N, Hadar
R, Plaschkes I, Iny Stein T, Rosen N, Kohn A, Twik M, Safran M, et
al: GeneHancer: Genome-wide integration of enhancers and target
genes in GeneCards. Database (Oxford). 2017:bax0282017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Gonçalves J, Sharma A, Coyaud É, Laurent
EMN, Raught B and Pelletier L: LUZP1 and the tumor suppressor EPLIN
modulate actin stability to restrict primary cilia formation. J
Cell Biol. 219:e2019081322020. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bozal-Basterra L, Gonzalez-Santamarta M,
Muratore V, Bermejo-Arteagabeitia A, Da Fonseca C, Barroso-Gomila
O, Azkargorta M, Iloro I, Pampliega O, Andrade R, et al: LUZP1, a
novel regulator of primary cilia and the actin cytoskeleton, is a
contributing factor in Townes-Brocks Syndrome. Elife. 9:e559572020.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Dong X, Zhang P, Liu L, Li H, Cheng S, Li
S, Wang Y, Zheng C, Dong J and Zhang L: The
Circ_0001367/miR-545-3p/LUZP1 axis regulates cell proliferation,
migration and invasion in glioma cells. Front Oncol. 11:7814712021.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Khera N, Rajkumar AS, Abdulkader M Alkurdi
K, Liu Z, Ma H, Waseem A and The MT: Identification of multidrug
chemoresistant genes in head and neck squamous cell carcinoma
cells. Mol Cancer. 22:1462023. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
DeWane G, Salvi AM and DeMali KA: Fueling
the cytoskeleton-links between cell metabolism and actin
remodeling. J Cell Sci. 134:jcs2483852021. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Aseervatham J: Cytoskeletal Remodeling in
Cancer. Biology (Basel). 9:3852020.PubMed/NCBI
|
|
38
|
Song Z, Cui Y, Xin L, Xiao R, Feng J, Li
C, Yin Z, Wang H, Li Q, Wang M, et al: Mechano-oncogenic
cytoskeletal remodeling drives leukemic transformation with
mitochondrial vesicle-mediated STING activation. Cell Stem Cell.
32:581–597.e11. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Mao H, Zhao X and Sun SC: NF-κB in
inflammation and cancer. Cell Mol Immunol. 22:811–839. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Cao Y, Yi Y, Han C and Shi B: NF-κB
signaling pathway in tumor microenvironment. Front Immunol.
15:14760302024. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Liu C, Wu K, Li C, Zhang Z, Zhai P, Guo H
and Zhang J: SPP1+ macrophages promote head and neck squamous cell
carcinoma progression by secreting TNF-α and IL-1β. J Exp Clin
Cancer Res. 43:3322024. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Rébé C and Ghiringhelli F: Interleukin-1β
and cancer. Cancers (Basel). 12:17912020. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Chen S, Yang Y, Zheng Z, Zhang M, Chen X,
Xiao N and Liu H: IL-1β promotes esophageal squamous cell carcinoma
growth and metastasis through FOXO3A by activating the PI3K/AKT
pathway. Cell Death Discov. 10:2382024. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Okuda S, Yoshizawa AC, Kobayashi D,
Takahashi Y, Watanabe Y, Moriya Y, Hatano A, Takami T, Matsumoto M,
Araki N, et al: jPOST environment accelerates the reuse and
reanalysis of public proteome mass spectrometry data. Nucleic Acids
Res. 53:D462–D467. 2025. View Article : Google Scholar : PubMed/NCBI
|