Introduction
Leukemia is a heterogeneous group of hematologic
malignancies characterized by the clonal expansion and accumulation
of neoplastic white blood cells. Most leukemias are classified into
myeloid or lymphoid lineages based on the differentiation of
leukemia stem cells and also may be further classified as either
acute or chronic according to blast cells count (1). Acute leukemias (ALs), as a
hematological malignancy with different morphological
characteristics, include acute promyelocytic leukemia (APL), acute
myeloid leukemia (AML, non-APL) and acute lymphocytic leukemia
(ALL), account for ~42.7% of all leukemia cases and 54.8%
leukemia-related mortality (1,2). These
leukemias are characterized by a rapid increase of abnormally
immature white blood cells, which inhibit normal blood cell
production in the bone marrow (1).
Despite the availability of advanced techniques available,
including cytogenetics, immunophenotypes and increasing molecular
genetics, the initial analysis of cell morphology continues to hold
significance in numerous intramedullary and extramedullary
pathological diagnoses (3).
However, morphological analysis of ALs poses challenges due to its
high subjectivity and shortage of skilled technicians specializing
in hematopathology (4,5). Consequently, there is a critical
demand for an automated and intelligent diagnostic assistance
platform to enhance efficiency, alleviate examiner burden and
advance medical research.
Artificial intelligence (AI), which emerged in the
mid-20th century, focuses on developing computer algorithms capable
of performing human-like tasks (6,7). AI is
increasingly utilized across multiple domains within the medical
field, offering potential benefits such as aiding radiologists in
swiftly and precisely diagnosing tumors and assisting pathologists
in improving the overall accuracy of cancer diagnosis (8,9). Key
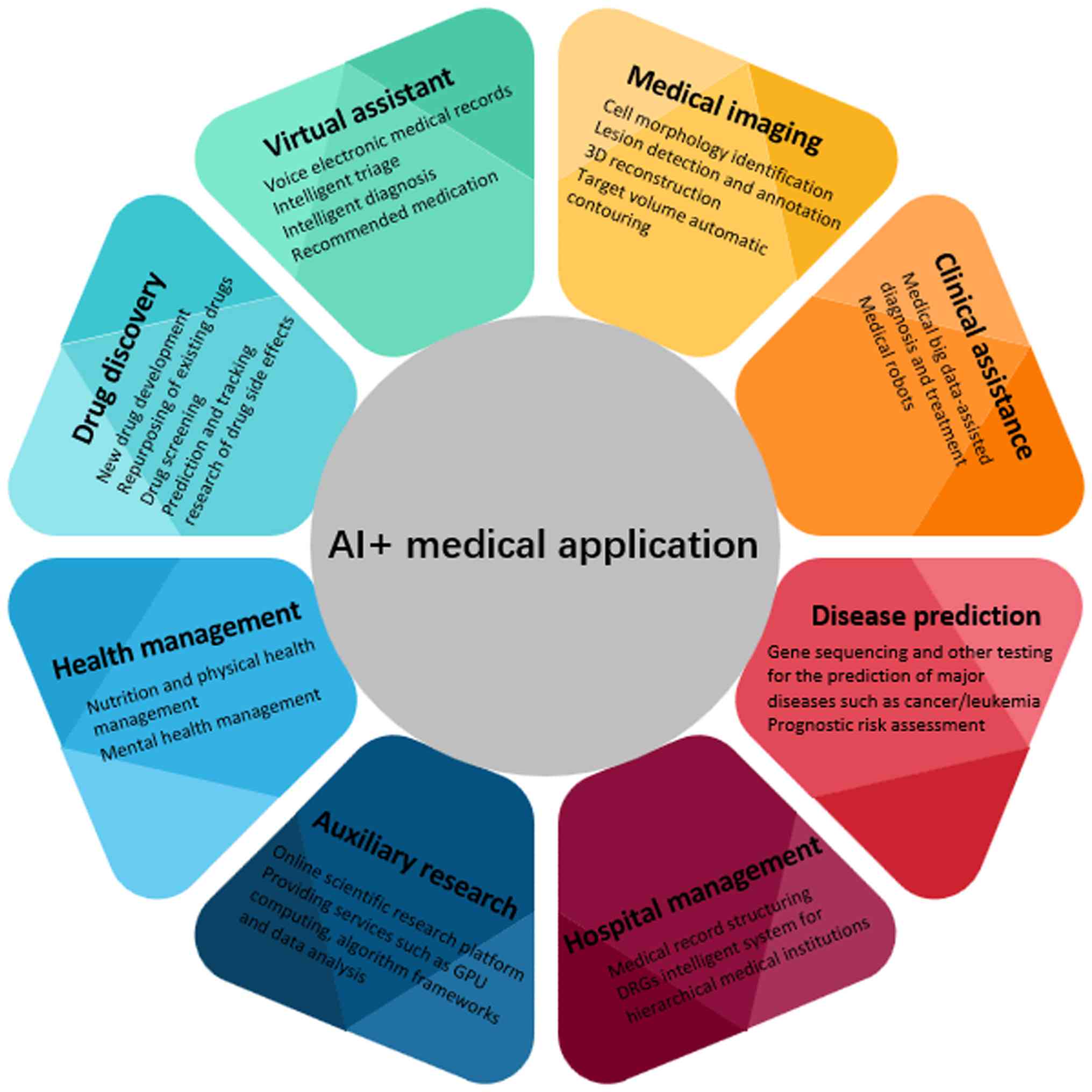
applications of AI are summarized in Fig. 1. Expertise in analyzing bone marrow
cell morphology is vital for diagnosing and tracking the
effectiveness of hematological disorders (10–12).
Currently, numerous studies have attempted AI-assisted
morphological examinations based on bone marrow smears and have
acquired notable results (3,10,13–16).
Early computer-assisted systems for hematological image analysis
relied on rule-based algorithms and handcrafted feature extraction
to classify blood cells, often constrained by limited computational
power and dataset scale. With advancements in computational
capabilities and the availability of large, annotated image
datasets, these traditional approaches gradually evolved into
data-driven machine learning (ML) methods, and subsequently into
deep learning (DL) models capable of automatic feature learning and
superior classification performance (17).
The present review first elaborated on the
application of AI in image recognition, followed by a summary of
previous and recent studies on bone marrow and/or peripheral blood
cell morphology for the detection of various ALs, with particular
emphasis on the latest progress and contributions in morphological
image analysis performance for APL, AML (non-APL) and ALL. Finally,
the current limitations and outline future directions for
integrating AI into routine hematopathology practice are
discussed.
AI for image recognition
In clinical practice, the diagnostic accuracy of
cancer and/or numerous other diseases is largely dependent on the
expertise of radiologists and pathologists, but notable
inter-observer variability exists in reading and interpreting
medical images (18). To address
this, numerous computer-aided detection and diagnosis systems have
been developed to help clinicians interpret medical images more
effectively and support diagnostic decisions (3,19–22).
ML, a type of AI, refers to computer algorithms that
use training data to acquire the ability to perform tasks. ML
models can be classified into supervised, unsupervised and
semi-supervised learning based on the intended output (23). During supervised learning, each
trained image is labeled, and the model is optimized using these
labels to predict categories for new images (24). Unsupervised learning examines
unlabeled data to discover underlying patterns, while
semi-supervised learning utilizes a small amount of labeled data
along with unlabeled data to improve model performance (25). Regardless of the aforementioned
learning methods described, they are all based on convolutional
neural network (CNN) algorithms.
CNNs are widely used in computer vision to recognize
and capture relevant details from visual data. Inspired by
biological visual systems, CNNs process images through multiple
layers, from detecting basic edges and textures in initial layers
to identifying complex objects in deeper layers (26). They have been extensively applied in
medical fields: In radiology for interpreting chest X-rays and
classifying pulmonary nodules (27,28)
and in pathology for classifying tumors, detecting lymph node
metastases and predicting PD-L1 status directly from images
(29–32). A notable milestone was reached in
2018, when the US Food and Drug Administration (US FDA) granted
approval for a retinopathy detection system that relied on CNNs
technology and fulfilled the necessary criteria for clinical
application (33).
AI based on ML can utilize existing data more
effectively than traditional analysis. As summarized by Rodellar
et al (34), the AI for
analyzing hematopathology data involves cell identification and
segmentation, algorithm-based feature extraction, and subsequent
classification based on these features (Fig. 2). ML models, particularly CNNs, have
advanced to allow direct feature extraction and classification with
minimal manual intervention. While CNNs have demonstrated high
accuracy (>95%) in classifying numerous nucleated cell images,
challenges remain in distinguishing morphologically similar
lineages, such as lymphocytes and reactive lymphocytes (35,36).
The present review subsequently explores how these AI methods are
specifically applied to the morphological diagnosis of ALs.
Morphology diagnosis of ALs by AI
The diagnosis and classification of ALs are
conventionally achieved through microscopic observation of cell
morphology from bone marrow smear by hematologists according to
5th World Health Organization classification criteria
(37). This process is heavily
reliant on the examiner's expertise, underscoring the need for
automated systems to enhance diagnostic consistency and efficiency.
AI offers transformative potential in this field by enabling
high-throughput image analysis and facilitating classifications
that may elude human observers. Conventional image-based AI systems
typically follow a three-stage pipeline: Object segmentation,
feature extraction and disease classification (38). Boldú et al (39) proposed a ML model for AL lineage
diagnosis using peripheral blood smears, which demonstrated high
diagnostic performance by achieving 100% sensitivity, 92.3%
specificity and 93.7% precision for myeloid leukemia, and 89%
sensitivity with 100% specificity and precision for lymphocytic
leukemia, suggesting a potential screening role in in routine
hematology. Alcazer et al (40) developed an AI-based prediction
model, termed AI-PAL, for diagnosing APL, AML (non-APL) and ALL.
The model was stratified into confidence and overall variants based
on prediction probability scores. In internal and external
validation, the confidence model achieved accuracies of 99.7, 98.8
and 99.5% for APL, AML (non-APL) and ALL, while the overall model
yielded corresponding accuracies of 87.9, 86.3 and 96.1%,
respectively. However, real-world performance was more moderate,
with area under the receiver operating characteristic curve (AUROC)
values of 0.67 for ALL and 0.71 for AML (41).
Recent advances in CNNs have led to models with
near-perfect metrics, such as 98.37% accuracy using an Orthogonal
SoftMax Layer-based model (42),
99.37% accuracy with the BSNEU-net framework (43) and 96.15% accuracy with LeuFeatx, a
VGG16-adapted feature extractor (44). Despite these promising results,
barriers to clinical deployment remain, due to high computational
demands, sensitivity to hyperparameters and variability in image
quality. The present review synthesized the performance and
limitations of representative AI models in diagnosing APL, AML
(non-APL) and ALL (Fig. 3), with
emphasis on their clinical feasibility and intended roles.
APL
APL is a medical emergency associated with an early
mortality rate of up to 30%, necessitating rapid and accurate
diagnosis (45). Morphologically,
APL is characterized by abnormal promyelocytes with heavy
granulation and Auer rods (46).
While cytomorphology remains the fastest diagnostic modality,
definitive confirmation often requires cytogenetic and molecular
testing (47–49).
AI models, particularly CNNs, have been developed to
assist in rapid morphological screening. For instance, Qiao et
al (50) proposed a compact CNN
that distinguished promyelocytes from normal nucleated cells with
accuracies of 96.53 and 99.20% on two public datasets
(APL-Cytomorphology-LMU and -JHH; The Cancer Imaging Archive).
Eckardt et al (14)
developed a multi-stage DL platform for automated bone marrow smear
analysis, reporting an average precision and recall of 0.97 for
cell segmentation and an AUROC of 0.8575 and 0.9585 for
distinguishing APL from non-APL AML and healthy donors,
respectively, enabling timely diagnosis of APL and early
intervention for patients. Manescu et al (51) introduced a multi-instance learning
(MILLIE) method, which detected APL with an area under curve (AUC)
of 0.94±0.04 in peripheral blood smears and 0.99±0.01 in bone
marrow smears without requiring cell-level annotations. Integrating
MILLIE into clinical workflows may markedly improve detection
throughput and reduce cognitive human errors. Recently, our
previous study exhibited a CELLSEE model for APL screening, which
demonstrated a joint diagnostic performance achieving 93.00%
accuracy and 100% recall at magnifications of ×10 and ×100 in batch
samples, thereby ensuring no APL cases were missed (52). The aforementioned models for rapid
diagnosis of APL demonstrated satisfactory morphological
recognition performance (Table
I).
 | Table I.Studies utilizing AI for APL
morphological diagnosis. |
Table I.
Studies utilizing AI for APL
morphological diagnosis.
| First author,
year | Purpose | AI model | Validation
strategy/dataset | Performance
results | Highlight | (Refs.) |
|---|
| Qiao et al,
2021 | Timely APL
diagnosis | End-to-end pipeline
based on CNN model | Public
database | Precision, 0.9920;
AUC, 0.9977 | Distinguishing
promyelocytes from normal leukocytes | (50) |
| Eckardt et
al, 2022 | APL prediction and
diagnosis | Deep learning based
on CNN | Clinical
dataset | Precision and
recall 0.97; ROC, 0.8575 (APL vs. AML); ROC, 0.9585 (APL vs.
HD) | Earlier diagnosis
vs. genetics/molecular biology | (14) |
| Manescu et
al, 2023 | Improving APL
diagnostic efficiency and reducing human error | Multiple instance
learning for leukocyte identification | Public
database | AUC, 0.94 (blood
films); AUC, 0.99 (bone marrow) | Augmented
throughput for blood film assessment | (51) |
| Xiao et al,
2024 | Rapid APL
screening | CELLSEE model based
on CNN | Clinical
dataset | Accuracy, 0.93;
precision, 0.85; recall, 1.00 | Rapid, batch-sample
screening | (52) |
AML (non-APL)
AML represents the most common type of AL, with an
incidence rate of ~33.5% among all leukemias (2). The morphological characteristics of
AML blasts are as follows: i) The cell size is moderate; ii) the
nucleus is generally round or ovoid, and may exhibit indentation
and folding; iii) the nuclear chromatin is finely granular and lacy
with distinct nucleoli; iv) the amount of cytoplasm varies, is
basophilic, and frequently contains variable numbers of azurophilic
granules; and v) a small proportion of blasts may show Auer rods
(37).
Diagnosis is primarily based on blast cell counts in
bone marrow or peripheral blood (53), with subclassification into M0-M7
according to French-American-British classification criteria
(54). However, cellular
morphological examination is an efficient but highly demanding
task, relying on well-trained and experienced medical
professionals. Therefore, there is a need for an automated AML
classification system that overcomes these issues.
Initial applications of ML in this domain relied on
traditional algorithms. For instance, techniques such as K-means
and fuzzy C-means clustering were primarily employed for the
segmentation of nuclei and cells in bone marrow smears, serving as
foundational tools for the screening and identification of AML
(55,56). The advent of DL has markedly
improved diagnostic performance. Huang et al (57) combined preprocessing algorithms with
a CNN to achieve 99% accuracy in AML classification, demonstrating
the viability of transfer learning in this context. Building on
these preprocessing techniques, several integrated DL frameworks
have been developed for comprehensive AML recognition and
classification. Wang et al (58) constructed an ImageNet-pretrained CNN
for recognizing aplastic anemia (AA), myelodysplastic syndromes
(MDS) and AML, achieving an AUC of 0.968, accuracy of 0.929 and
sensitivity of 0.857 for AML recognition in the testing set,
providing a convenient tool for clinical doctors to distinguish AA,
MDS and AML. More advanced DL pipelines have since been developed
to address increasingly complex diagnostic tasks. Eckardt et
al (59) extended DL
applications to predict nucleophosmin 1 mutation status directly
from morphological images, attaining an AUROC of 0.92. Yu et
al (60) developed AMLnet, a DL
pipeline capable of AML subtype discrimination with image- and
patient-level AUCs of 0.885 and 0.921, respectively, offering a
rapid prescreening and decision support tool for morphological
pathologists. Other notable models include a hybrid deep
convolutional autoencoder-CNN model by Elhassan et al
(61), which reported 97% accuracy,
97% sensitivity and 98% precision for classifying atypical AML
cells. In the realm of image segmentation, Zhang et al
(62) employed an AML conditional
generative adversarial network (AMLcGAN) for blast segmentation,
achieving a precision of 0.8496 and a recall of 0.8831. The
application of DL has also extended to peripheral blood smear
analysis. For instance, an auxiliary classification GAN evaluated
by Zhang et al (63)
achieved 97.1% accuracy for AML screening. More recently, Aby et
al (64) conducted a
comparative study of activation functions within deep neural
networks, specifically InceptionV3, ResNet50v2 and VGG16, for AML
subtype classification. The findings indicated that the Gaussian
Error Linear Unit function yielded the highest accuracy of 94.02%
when implemented in InceptionV3. Cheng et al (65) developed an AI model using 65,039
myeloblast images from 205 patients with AML to identify
RUNX1::RUNX1T1 fusion gene abnormalities via cell
morphology, which achieved a maximum sensitivity of 95.65% and
specificity of 92.68% under different thresholds, enabling
effective recognition of this genetic alteration. Collectively,
these studies underscore the strong capability of DL-based systems
in enhancing the accuracy and efficiency of AML diagnosis and
classification (Table II).
 | Table II.Studies utilizing AI for AML
morphological diagnosis. |
Table II.
Studies utilizing AI for AML
morphological diagnosis.
| First author,
year | Purpose | AI model | Validation
strategy/dataset | Performance
results | Highlight | (Refs.) |
|---|
| Huang et al,
2020 | Establishing an
objective, rapid and accurate leukemia classification method | The perfect
reflection algorithm and a self-adaptive filter algorithm based on
CNN | Clinical
dataset | Accuracy, 0.90 | Faster, more
accurate and more objective compared with traditional manual
microscopes. | (57) |
| Wang et al,
2022 | Automatically
distinguishing AA, MDS, and AML | CNN model | ASH image bank | AUC, 0.968;
accuracy, 0.929; sensitivity, 0.857 | Distinguishing AA,
MDS, AML with high accuracy | (58) |
| Liu et al,
2022 | Automatically
classifying AML-M1 and M2 subtypes | Random forest
method and broad learning system | TCIA open
database | Accuracy 0.998;
AUC, 0.998; F1-score, 0.998; recall, 0.996 | Enabling
classification of AML-M1 and M2 subtypes | (13) |
| Eckardt et
al, 2022 | Detecting AML and
predicting NPM1 mutation status | Multi-step DL | Clinical
dataset | AUROC, 0.9699 (AML
vs. HD); AUROC, 0.92 (prediction for NPM1 mutation) | Predicting NPM1
mutation status from myeloblast morphology | (59) |
| Yu et al,
2023 | Diagnosing and
classifying AML | AMLnet, a
deep-learning pipeline based on bone marrow images | Clinical
dataset | AUC, 0.885 (image
level); AUC, 0.921 (patient level) | Rapid prescreening
and decision-making aid | (60) |
| Elhassan et
al, 2023 | Building an AML
classification model | GT-DCAE WBC
augmentation model |
AML-Cytomorphology-LMU | Accuracy, 0.97,
sensitivity, 0.97; precision, 0.98 | Introducing a
hybrid data augmentation model | (61) |
| Zhang et al,
2024 | Accurately
differentiating myeloblasts | AMLcGAN | Clinical
dataset | Precision, 0.8496;
recall, 0.8831 | Aiding pathologists
in accurate AML diagnosis | (62) |
| Zhang et al,
2024 | Assessing ACGAN
applicability | ACGAN model | TCIA dataset | Precision, 0.975;
recall, 0.973; F1 scores, 0.974 | High performance
compared with other advanced methods | (63) |
| Aby et al,
2025 | Exploring the
effectiveness of various CNN architectures for AML subtyping | VGG16, InceptionV3
and ResNet50v2 | ASH image bank | Accuracy, 0.9283
(VGG16); accuracy, 0.9402; (InceptionV3) accuracy, 0.9253
(ResNet50v2) | Demonstrating
potential of CNN for automated AML subtyping and identifying GELU
as the optimal activation function | (64) |
ALL
ALL accounts for roughly 9.2% of all leukemia cases
and primarily affects children, featuring excessive proliferation
of lymphocytes in the bone marrow. Despite a favorable cure rate in
the pediatric population, ALL still accounts for 6.7% of
cancer-related deaths among all age groups (2,66).
Diagnosis relies on cytomorphology, immunology and genetics, but
resource limitations often restrict access to advanced assays,
making morphological assessment indispensable. The morphological
features of ALL blasts are as follows: i) The cell size is
moderately small to small; ii) the nuclei are mostly round with
coarse granular or blocky nuclear chromatin, which is coarser than
that of AML blasts; iii) nucleoli are generally small and distinct,
though inconspicuous in some cases; iv) the cytoplasm is extremely
scanty and basophilic, occasionally containing a few azurophilic
granules; and v) basket cells and mitotic figures are commonly
observed (37). With the continuous
development of AI, there have been numerous studies on
morphological computer-aided diagnosis for ALL (Table III).
 | Table III.Included studies utilizing AI for ALL
morphological diagnosis. |
Table III.
Included studies utilizing AI for ALL
morphological diagnosis.
| First author,
year | Purpose | AI model | Validation
strategy/dataset | Performance
results, % | Highlight | (Refs.) |
|---|
| Shafique et
al, 2018 | Automating ALL
detection and classification | AlexNet, a deep
convolutional neural network | ALL-IDB | Accuracy, 99.50;
specificity, 98.11; sensitivity, 100.00 | Providing
directions for diagnosing other leukemias | (67) |
| Rehman et
al, 2018 | Improving ALL
diagnosis | Computer vision
toolbox and Alexnet model on GaPU based on CNN | Amreek Clinical
Laboratory | Accuracy,
97.78 | Novel image
segmentation technique | (68) |
| Rezayi et
al, 2021 | Facilitating rapid
ALL identification and classification | A convolutional
network with ten convolutional layers and 2×2 max-pooling
layers-with strides 2 | ALL-IDB and ASH
image bank | Test accuracy,
85.79; validation accuracy, 82.10 | Improved accuracy
with increased image volume | (69) |
| Pałczyński et
al, 2021 | Autonomously
classifying ALL | Hybrid artificial
intelligence systems | ALL-IDB
Database | Accuracy,
97.50 | Demonstrating
advantages of transfer learning | (70) |
| Jiang et al,
2021 | Timely and accurate
ALL diagnosis | ViT-CNN ensemble
model | VGG-16 | Accuracy,
99.03 | Combining vision
transformer and CNN architectures | (71) |
| Mirmohammadi et
al, 2021 | Classifying ALL,
normal, reactive and atypical cells | RF classifier | Images data from
the author's own laboratory samples | Accuracy,
98.00 | Superior
performance vs. other common classifiers | (72) |
| Musleh et
al, 2022 | Realizing early and
rapid ALL diagnosis | ALL Detector, a
deep learning-based network to distinguish patients with ALL | ALL-IDB2
database | Accuracy,
98.00 | Best performance in
distinguishing ALL from healthy groups | (73) |
| Jawahar et
al, 2022 | Realizing early ALL
diagnosis | ALNett, a deep
neural network | ResNet50, VGG-16,
AlexNet, GoogleNet | Training accuracy
99.73; test accuracy, 91.13; F1-score, 0.96 | Improved tool to
aid clinical decision-making | (74) |
| Almadhor et
al, 2022 | Establishing
automated ALL prediction techniques | KNN, SVM, RF,
NB | C-NMC leukemia
dataset | Accuracy, 90 (SVM
vs. other algorithms) | Practical method
for determining ALL diagnosis | (75) |
| Atteia et
al, 2022 | Autonomously
identifying ALL in blood images | Bayesian-based
optimized CNN | ALL-IDB1,
ALL-IDB2 | Test accuracy,
100.00; test specificity, 100.00; test sensitivity, 100.00 | Bayesian-optimized
CNN outperforming other models | (76) |
| Chen et al,
2022 | Rapidly and
accurately detecting ALL | Resnet101-9
ensemble model | C-NMC leukemia
dataset | Accuracy, 85.11;
F1-score, 88.94 | Ensemble model
outperforming individual ResNet-101 | (77) |
| Abunadi et
al, 2022 | Developing
diagnostic systems for early ALL detection | ANN, FFNN, SVM,
AlexNet, GoogleNet and ResNet-18 consist of hybrid systems | ALL-IDB1,
ALL-IDB2 | Accuracy,
100.00 | Improved accuracy
via hybrid model combinations | (78) |
| Albeeshi et
al, 2023 | Finding an
effective and fast method for ALL diagnosis | VGG16 pre-trained,
and using SVM and MLP classification algorithms | VGG16, SVM | Best training
accuracy, 92.27; best validation accuracy, 85.62 | Enhanced
performance via learning rate adjustment | (79) |
| Ahmed et al,
2023 | Assisting in early
ALL detection | DenseNet121,
ResNet50, and MobileNet consist of hybrid systems | C-NMC 2019 and
ALL-IDB2 | Accuracy, 100.00;
precision, 100.00; specificity, 100.00; sensitivity, 100.00 | Significantly
improved detection via hybrid systems | (80) |
| Kaur et al,
2023 | Early and accurate
ALL detection | Deep Skip
Connections-Based Dense Network | Kaggle dataset | Accuracy, 97.89;
sensitivity, 99.13; specificity, 96.76 | Efficiently
handling challenges in existing diagnosis models. | (81) |
| Huang et al,
2024 | Timely ALL
diagnosis | Ensemble-ALL
model | C-NMC leukemia
dataset | Accuracy, 96.26;
precision, 96.26; specificity, 96.26 | Valuable
contributions to ALL diagnosis | (82) |
| Jawahar et
al, 2024 | Detecting ALL | Deep Dilated
Residual Convolutional Neural Network | Kaggle dataset | Training accuracy,
99.86; test accuracy, 91.98 | Leveraging
optimized algorithms and parallel processing for high
workloads | (83) |
| MoradiAmin et
al, 2024 | Developing an
accurate and efficient system for identifying ALL cells | A novel CNN | Images data from
the author's own laboratory samples | Accuracy,
97.00 | Eliminating manual
feature extraction in classification | (84) |
| Elrefaie et
al, 2024 | Improving criterion
for classifying ALL microscopic images | Neural network
classifier based on Bayesian regularization method | ALL-IDB2
dataset | Accuracy, 98.70;
sensitivity, 99.30; specificity, 98.10 | Combining Bayesian
regularization with neural networks | (85) |
| Alsaykhan et
al, 2024 | Exploring the
automated detection of ALL | Hybrid SVM-PSO
model | ALL-IDB1 and
ALL-IDB2 dataset | Accuracy,
97.40 | Novel hybrid
algorithm improving detection performance | (86) |
| Mei et al,
2025 | Achieving early ALL
diagnosis | Progressive
shrinking convolutional neural network model | Images data from
the author's own laboratory samples | Accuracy,
92.51 | Enhancing
diagnostic efficiency and accuracy for experts | (87) |
| Mei et al,
2025 | Enabling rapid
whole slide analysis of ALL | SGLNet
framework | Images data from
the author's own laboratory samples | Average precision,
95.90 | Facilitating
large-scale clinical diagnosis | (88) |
| Anand et al,
2025 | Early diagnosis and
detection of ALL. | Deep optimized
CNN | Images data from
the author's own laboratory samples | Accuracy, 96.00;
precision, 95.00 | Achieving superior
performance by adjusting hyperparameters | (89) |
Early applications of DL demonstrated excellent
performance in the diagnosis of ALL. Shafique et al used a
DL CNN for ALL diagnosis, achieving 100% sensitivity, 98.11%
specificity and 99.50% accuracy, enabling high performance without
the need for microscopic image segmentation (67). Rehman et al (68) also employed a CNN-based model for
ALL-training, with a reported accuracy of 97.78% for the diagnosis
of ALL and subtype classification. These foundational studies
catalyzed rapid growth in AI-based ALL diagnosis using cell image
recognition.
Subsequent studies in 2021 explored diverse
architectural approaches. Rezayi et al (69) compared ResNet-50, VGG-16 and a
proposed convolutional network, achieving accuracies ranging from
81.63 to 84.62%. The proposed network is simpler than the two
pretrained networks and can be employed by pathologists to
recognize ALL. Furthermore, an optimized IoT-friendly neural
network achieved 97.5% accuracy for ALL classification (70), while a ViT-CNN ensemble model
reached 99.03% accuracy in distinguishing blasts from normal cell
images to support diagnosis (71).
Additionally, a Random Forest classifier integrated with image
enhancement yielded 98% accuracy for recognition of ALL (72).
In 2022, several innovations were introduced. The
ALL Detector achieved 98% accuracy in distinguishing patients with
ALL from healthy individuals, serving as a screening tool (73), and a deep neural network-based model
that employs depth-wise convolution with different dilation rates
to classify nucleated cell images and yielded a classification
accuracy of 91.13%, which is useful for ALL categorization
(74). Among traditional ML
algorithms, such as K-nearest neighbor, support vector machine
(SVM), random forest and naive bayes, SVM performed best with 90.0%
accuracy for predicting ALL (75).
A Bayesian-optimized CNN achieved 100% across all evaluation
metrics, showing superiority over existing DL systems developed for
ALL diagnosis (76). A ResNet101-9
ensemble model attained 85.11% accuracy, which performed well in
classifying ALL (77), and a hybrid
artificial neural network-feedforward neural network-SVM model
achieved 100% accuracy for diagnosis of ALL (78).
In 2023, model development continued with varied
architectures. A VGG16 model combined with SVM and multilayer
perceptron (MLP) classifiers yielded a training accuracy of 92.27%
and a validation accuracy of 85.62%, achieving fast and accurate
diagnosis (79). The hybrid CNN-RF
and CNN-XGBoost systems exhibited 100% AUC, accuracy and
sensitivity, thereby supporting effective early diagnosis of ALL
(80). Furthermore, a Deep Skip
Connections-Based Dense Network tailored for ALL diagnosis obtained
99.37% accuracy, 99.71% sensitivity, 99.03% specificity and an AUC
of 99.37%, demonstrating the potential of an effective tool for
early and accurate diagnosis of ALL (81).
In 2024, the focus shifted toward model refinement
and clinical applicability. An ensemble-ALL model achieved notable
performance, with accuracy, precision, recall, F1-score and kappa
scores of 96.26, 96.26, 96.26, 96.25 and 91.36%, respectively; this
model has made valuable contributions to medical image recognition,
particularly for the diagnosis of ALL (82). A Deep Dilated Residual CNN
demonstrated minimal computational complexity and improved the
discrimination of crucial features for accurate multi-class blood
cell image recognition, with an accuracy rate of 91.98% (83). Another deep neural network-based
model has a classification accuracy of ~97% for ALL and lymphocyte
subtypes (84). A method leveraging
Hilbert Huang supervision achieved 98.7% accuracy, 99.3%
sensitivity, and 98.1% specificity for classifying normal and blast
ALL cell images, which could assist in the screening of ALL
(85). Additionally, a hybrid
detection model combining SVM with particle swarm optimization
reported an accuracy of 93%, which is superior to stand-alone
algorithms and exhibiting increased overall performance, an
improved confusion matrix and a higher detection rate of ALL
(86).
In 2025, Mei et al (87) carried out two related studies on
lymphoblastic leukemia detection: in one study, they proposed a
lightweight model focusing on throughput (111 slides per second)
with a maintained accuracy of 92.51%, reflecting the trend of
developing clinically applicable efficient models. Furthermore, Mei
et al (88) built a
high-quality dataset containing 1,794 microscopic images and
developed the spatially-guided learning framework, which solved key
challenges in whole-slide leukemia detection and achieved a mean
average precision of 95.9% for ALL. In addition, Anand et al
(89) proposed a deep optimized CNN
model for early ALL diagnosis, which, optimized with the Adam
optimizer among other methods, attained an accuracy of 0.96 and a
precision of 0.95 after fine-tuning the corresponding
hyperparameters.
Despite these advances, to the best of our
knowledge, no model has undergone rigorous external validation in
real-world cohorts, and challenges related to model
interpretability, data heterogeneity and computational efficiency
still persist.
A comparison of AI methodologies reveals distinct
trade-offs among data requirements, interpretability, computational
efficiency and clinical roles (Table
IV). Traditional ML models offer high interpretability and low
computational cost but require manual feature engineering and
plateau in performance. Classic deep CNNs achieve state-of-the-art
accuracy through automated feature learning but suffer from low
inherent interpretability and high computational demands.
Lightweight architectures balance efficiency and accuracy, making
them suitable for point-of-care applications. Hybrid models combine
the strengths of DL and ML, offering moderate interpretability and
robust performance on smaller datasets. Ensemble methods, while
often achieving top benchmark performance, entail high complexity
and risk of overfitting. Generative models are valuable for data
augmentation and unsupervised segmentation but may produce
artifacts. Clinically, lightweight CNNs are preferable for rapid
screening in resource-limited settings, whereas hybrid or ensemble
models may serve as high-precision diagnostic aids in well-equipped
laboratories.
 | Table IV.Comparison of AI techniques in
medical image analysis. |
Table IV.
Comparison of AI techniques in
medical image analysis.
| Technique | Representative
architectures | Data and annotation
requirements |
Interpretability | Strengths | Weaknesses | Typical clinical
role | (Refs.) |
|---|
| Traditional ML
(feature-based) | RF, SVM, k-means,
fuzzy C-means | Small to medium
datasets; handcrafted features; annotation for seg. and labels | High (features
human-defined) | Fast training;
transparent decisions; works with limited data; easy to deploy | Performance
saturates; poor generalization; manual feature engineering is
labor-intensive | Screening tools,
educational aids, legacy systems | (13,55,56,72,84,85) |
| Classic deep
CNNs | VGG16, ResNet50,
InceptionV3 | Large datasets
(thousands/class); pixel/image-level labels | Low (black box);
post-hoc explanations (Grad-CAM) | SOTA accuracy;
automatic feature learning; transfer learning mitigates data
scarcity | Computationally
expensive; overfitting risk; sensitive to hyperparameters; low
inherent interpretability | High-performance
classifiers, research prototypes | (2,10,39,42,44,51,52,57,58,59,68,69,73,79,89) |
| Lightweight or
efficient CNNs | MobileNet,
EfficientNet, compact CNN | Moderate datasets;
benefits from pretraining | Low to
moderate | Real-time
inference; point-of-care feasible; low power | Slightly lower
accuracy than large models; still lacks built-in
interpretability | Bedside decision
support, large-scale screening | (43,50,60,65,87,88) |
| Hybrid models | CNN + RF, CNN +
SVM, CNN + XGBoost | Moderate datasets;
leverages pretrained CNN features | Moderate (CNN
features visualizable) | Balances accuracy
and interpretability; often outperforms pure DL on small data | Two-stage pipeline;
not fully end-to-end optimized | Generalizable tools
when training data limited | (9,15,40,70,74,76,78,80,81,83,86) |
| Ensemble
models | Voting/averaging of
multiple CNNs, CNN + ViT | Large datasets;
multiple models required | Very low | Robustness; often
top benchmark scores | High computational
cost; complex deployment; risk of over-ensembling | Competition
solutions, seldom deployed clinically | (14,71,77,82) |
| Generative
models | AMLcGAN,
ACGANs | Unlabeled/paired
data for synthesis/segmentation | Low | Data augmentation;
segmentation without extensive labels | Training
instability; mode collapse; synthetic images may miss diagnostic
details | Enhancing training
sets, unsupervised segmentation | (62,63) |
| Vision
transformers | ViT, Swin
Transformer | Very large datasets
(>1M) or strong pretraining | Very low | Captures global
context; can outperform CNNs on massive data | Extreme data
hunger; extensive regularization needed; limited validation in
cytology | Cutting-edge
research, potential future tools | (61,67,75) |
Conclusions and perspectives
ALs are life-threatening diseases that can lead to
death if left untreated. Currently, microscopic bone marrow
examination remains the primary diagnostic procedure. However,
given the dynamic nature of hematological diseases and the frequent
presence of atypical cell morphology, there is a growing need for
an automated, efficient and highly accurate diagnostic system,
which would help eliminate intra- and inter-observer variability
and improve diagnostic consistency. Previous studies have
demonstrated that AI significantly outperformed manual microscopy
in diagnosing and subtyping AML based on bone marrow cell
morphology, as well as in predicting the RUNX1:RUNX1T1 AML
subtype. This advantage is particularly evident in metrics such as
sensitivity and positive predictive value (60,65).
In recent years, AI has begun integrating into
clinical hematology. By June 2025, the US FDA had already approved
20 AI/ML-based devices to assist hematologists or medical
laboratory technicians in extracting information from specimens
(https://www.fda.gov/medical-devices/software-medical-device-samd/artificial-intelligence-and-machine-learning-aiml-enabled-medical-devices).
To date, none of the specific AI models in this review have
received FDA approval for standalone diagnosis of ALs. However,
several AI-based digital pathology and hematology systems have been
cleared for broader clinical use in supporting cell counting,
segmentation or pre-classification tasks. This trend not only
underscores the growing recognition of the potential of
AI-augmented diagnostic workflows but also expands the broad
application of AI in leukemia, encompassing minimal residual
disease monitoring, prediction of genetic abnormalities, prognostic
stratification and treatment management.
AI adoption in this field faces several significant
challenges, and it cannot yet fully replace human expertise. First,
in data quality and standardization: Variations in staining
protocols, smear preparation, imaging parameters and scanner types
introduce technical artifacts. Future efforts must prioritize
standardized protocols or AI-based normalization techniques to
minimize these inconsistencies. Second, model generalization and
real-world performance: While several models achieve high accuracy
(>95%) on internal or curated datasets, their performance often
declines in external, real-world validation. Prospective,
multi-center clinical trials using consecutive patient samples are
essential to assess true generalizability. Third, interpretability
and trust: The ‘black-box’ nature of numerous DL models hinders
clinical trust. Integrating explainability tools, such as attention
maps or saliency maps that highlight decision-relevant cellular
regions, is crucial for transparency. Fourth, integration into
clinical workflows: Successful adoption depends on seamlessly
embedding AI into existing diagnostic pathways. Finally, ethics and
application: Data privacy, algorithm transparency and clinician
acceptance present critical hurdles that must be addressed to
ensure ethical, trustworthy and effective deployment. Key
considerations include defining its role, as a triage tool, a
diagnostic assistant or an independent system, each requiring
different levels of sensitivity, specificity and regulatory
approval. Additionally, regulatory and legal frameworks for
AI-aided diagnoses must be established alongside technological
development.
Future studies should not only focus on improving
model performance but also emphasize practical clinical
application. Based on the findings of the aforementioned studies,
key directions for improvement include: i) Reducing the impact of
staining and imaging variations; ii) enriching and diversifying
databases; iii) establishing standardization; iv) preventing
systemic errors and biases; v) validating models through clinical
trials and regulatory processes; and vi) integrating multimodal
data. Notably, education on AI-assisted diagnostics should also be
incorporated into medical curricula to prepare future
practitioners.
In summary, addressing these challenges requires a
concerted interdisciplinary effort combining hematology, pathology,
computer science and biomedical engineering. Through collaborative
innovation and rigorous validation, AI could evolve into a reliable
adjunct in the morphological diagnosis of ALs.
Acknowledgements
Not applicable.
Funding
The present work was supported by Clinical Research Physician
Capability Enhancement Project (grant no. 2024LYC04).
Availability of data and materials
Not applicable.
Authors' contributions
HC and GZ designed and wrote original draft. YY and
CX reviewed and edited the manuscript. CH and GT corrected and
revised the paper. Data authentication is not applicable. All
authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Chennamadhavuni A, Iyengar V, Mukkamalla
SKR and Shimanovsky A: Leukemia. StatPearls. (Treasure Island, FL).
2026.Available from:. https://www.ncbi.nlm.nih.gov/books/NBK560490/
|
|
2
|
Siegel RL, Kratzer TB, Wagle NS, Sung H
and Jemal A: Cancer statistics, 2026. CA Cancer J Clin.
76:e700432026.PubMed/NCBI
|
|
3
|
Matek C, Krappe S, Münzenmayer C,
Haferlach T and Marr C: Highly accurate differentiation of bone
marrow cell morphologies using deep neural networks on a large
image data set. Blood. 138:1917–1927. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zini G: How I investigate difficult cells
at the optical microscope. Int J Lab Hematol. 43:346–353. 2021.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Varotto E, Munaretto E, Stefanachi F,
Della Torre F and Buldini B: Diagnostic challenges in acute
Monoblastic/monocytic leukemia in children. Front Pediatr.
10:9110932022. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Turing AM: Computing machinery and
intelligence. Mind. 49:433–460. 2001.
|
|
7
|
Schinkel M, Paranjape K, Nannan Panday RS,
Skyttberg N and Nanayakkara PWB: Clinical applications of
artificial intelligence in sepsis: A narrative review. Comput Biol
Med. 115:1034882019. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mazurowski MA, Buda M, Saha A and Bashir
MR: Deep learning in radiology: An overview of the concepts and a
survey of the state of the art with focus on MRI. J Magn Reson
Imaging. 49:939–954. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Shinde S, Kalbhor M and Wajire P:
DeepCyto: A hybrid framework for cervical cancer classification by
using deep feature fusion of cytology images. Math Biosci Eng.
19:6415–6434. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Chandradevan R, Aljudi AA, Drumheller BR,
Kunananthaseelan N, Amgad M, Gutman DA, Cooper LAD and Jaye DL:
Machine-based detection and classification for bone marrow aspirate
differential counts: Initial development focusing on nonneoplastic
cells. Lab Invest. 100:98–109. 2020. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Fuentes-Arderiu X and Dot-Bach D:
Measurement uncertainty in manual differential leukocyte counting.
Clin Chem Lab Med. 47:112–115. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Shah A, Khan A, Azam F, Rauf M, Basit A,
Shah J, Saleem Y and Khan SU: A machine learning-based approach for
the segmentation and classification of acute lymphoblastic leukemia
cells in microscopic blood images. Computers Materials Continua.
69:767–784. 2021.
|
|
13
|
Liu K and Hu J: Classification of acute
myeloid leukemia M1 and M2 subtypes using machine learning. Comput
Biol Med. 147:1057412022. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Eckardt JN, Schmittmann T, Riechert S,
Kramer M, Sulaiman AS, Sockel K, Kroschinsky F, Schetelig J,
Wagenführ L, Schuler U, et al: Deep learning identified acute
promyelocytic leukemia in bone marrow smears. BMC Cancer.
22:2012022. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Wang W, Luo M, Guo P, Wei Y, Tan Y and Shi
H: Artificial intelligence-assisted diagnosis of hematologic
diseases based on bone marrow smears using deep neural networks.
Comput Methods Programs Biomed. 231:1073432023. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ghete T, Kock F, Pontones M, Pfrang D,
Westphal M, Höfener H and Metzler M: Models for the marrow: A
comprehensive review of AI-based cell classification methods and
malignancy detection in bone marrow aspirate smears. Hemasphere.
8:e700482024. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Obeagu EI: Revolutionizing hematological
disorder diagnosis: Unraveling the role of artificial intelligence.
Ann Med Surg (Lond). 87:3445–3457. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Kruger RP, Townes JR, Hall DL, Dwyer SJ
III and Lodwick GS: Automated radiographic diagnosis via feature
extraction and classification of cardiac size and shape
descriptors. IEEE Trans Biomed Eng. 19:174–186. 1972. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhou Y, Chia MA, Wagner SK, Ayhan MS,
Williamson DJ, Struyven RR, Liu T, Xu M, Lozano MG, Woodward-Court
P, et al: A foundation model for generalizable disease detection
from retinal images. Nature. 622:156–163. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kermany DS, Goldbaum M, Cai W, Valentim
CCS, Liang H, Baxter SL, McKeown A, Yang G, Wu X, Yan F, et al:
Identifying medical diagnoses and treatable diseases by image-based
deep learning. Cell. 172:1122–1131.e9. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Dehkharghanian T, Mu Y, Tizhoosh HR and
Campbell CJV: Applied machine learning in hematopathology. Int J
Lab Hematol. 45 (Suppl 2):S87–S94. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Fan BE, Yong BSJ, Li R, Wang SSY, Aw MYN,
Chia MF, Chen DTY, Neo YS, Occhipinti B, Ling RR, et al: From
microscope to micropixels: A rapid review of artificial
intelligence for the peripheral blood film. Blood Rev.
64:1011442024. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Deo RC: Machine learning in medicine.
Circulation. 132:1920–1930. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
LeCun Y, Bengio Y and Hinton G: Deep
learning. Nature. 521:436–444. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chen Y, Mancini M, Zhu X and Akata Z:
Semi-supervised and unsupervised deep visual learning: A survey.
IEEE Trans Pattern Anal Mach Intell. 46:1327–1347. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Esteva A, Robicquet A, Ramsundar B,
Kuleshov V, DePristo M, Chou K, Cui C, Corrado G, Thrun S and Dean
J: A guide to deep learning in healthcare. Nat Med. 25:24–29. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Lee SM, Seo JB, Yun J, Cho YH,
Vogel-Claussen J, Schiebler ML, Gefter WB, van Beek EJR, Goo JM,
Lee KS, et al: Deep learning applications in chest radiography and
computed tomography: Current state of the art. J Thorac Imaging.
34:75–85. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Trebeschi S, van Griethuysen JJM,
Lambregts DMJ, Lahaye MJ, Parmar C, Bakers FCH, Peters NHGM,
Beets-Tan RGH and Aerts HJWL: Deep learning for fully-automated
localization and segmentation of rectal cancer on multiparametric
MR. Sci Rep. 7:53012017. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Ehteshami Bejnordi B, Veta M, Johannes van
Diest P, van Ginneken B, Karssemeijer N, Litjens G, van der Laak
JAWM; the CAMELYON16 Consortium, ; Hermsen M, Manson QF, et al:
Diagnostic assessment of deep learning algorithms for detection of
lymph node metastases in women with breast cancer. JAMA.
318:2199–2210. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Komura D and Ishikawa S: Machine learning
approaches for pathologic diagnosis. Virchows Arch. 475:131–138.
2019. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Korbar B, Olofson AM, Miraflor AP, Nicka
CM, Suriawinata MA, Torresani L, Suriawinata AA and Hassanpour S:
Deep learning for classification of colorectal polyps on
whole-slide images. J Pathol Inform. 8:302017. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sha L, Osinski BL, Ho IY, Tan TL, Willis
C, Weiss H, Beaubier N, Mahon BM, Taxter TJ and Yip SSF:
Multi-field-of-view deep learning model predicts nonsmall cell lung
cancer programmed death-ligand 1 status from whole-slide
hematoxylin and eosin images. J Pathol Inform. 10:242019.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Abràmoff MD, Lavin PT, Birch M, Shah N and
Folk JC: Pivotal trial of an autonomous AI-based diagnostic system
for detection of diabetic retinopathy in primary care offices. NPJ
Digit Med. 1:392018. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Rodellar J, Alférez S, Acevedo A, Molina A
and Merino A: Image processing and machine learning in the
morphological analysis of blood cells. Int J Lab Hematol. 40:46–53.
2018. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang Q, Bi S, Sun M, Wang Y, Wang D and
Yang S: Deep learning approach to peripheral leukocyte recognition.
PLoS One. 14:e02188082019. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Hegde RB, Prasad K, Hebbar H and Singh
BMK: Comparison of traditional image processing and deep learning
approaches for classification of white blood cells in peripheral
blood smear images. Biocybern Biomed Eng. 39:382–392. 2019.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
WHO Classification of Tumours Editorial
Board, . Haematolymphoid tumours. WHO Classification of Tumours
Series. 11. 5th edition. International Agency for Research on
cancer; Lyon: 2024, Available from:. https://publications.iarc.who.int/637October
12–2025
|
|
38
|
Chen X, Wang X, Zhang K, Fung KM, Thai TC,
Moore K, Mannel RS, Liu H, Zheng B and Qiu Y: Recent advances and
clinical applications of deep learning in medical image analysis.
Med Image Anal. 79:1024442022. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Boldú L, Merino A, Acevedo A, Molina A and
Rodellar J: A deep learning model (ALNet) for the diagnosis of
acute leukaemia lineage using peripheral blood cell images. Comput
Methods Programs Biomed. 202:1059992021. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Alcazer V, Le Meur G, Roccon M, Barriere
S, Le Calvez B, Badaoui B, Spaeth A, Kosmider O, Freynet N,
Eveillard M, et al: Evaluation of a machine-learning model based on
laboratory parameters for the prediction of acute leukaemia
subtypes: A multicentre model development and validation study in
France. Lancet Digit Health. 6:e323–e333. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Pucher G, Rostalski T, Nensa F, Kleesiek
J, Reinhardt HC and Sauer CM: Why implementing machine learning
algorithms in the clinic is not a plug-and-play solution: A
simulation study of a machine learning algorithm for acute
leukaemia subtype diagnosis. EBioMedicine. 111:1055262025.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Das PK, Sahoo B and Meher S: An efficient
detection and classification of acute leukemia using transfer
learning and orthogonal softmax layer-based model. IEEE/ACM Trans
Comput Biol Bioinform. 20:1817–1828. 2023. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Saikia R, Deka R, Sarma A and Devi SS:
BSNEU-net: Block feature map distortion and switchable
Normalization-based enhanced Union-net for Acute leukemia detection
on heterogeneous dataset. J Imaging Inform Med. 38:1334–1361. 2025.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Rastogi P, Khanna K and Singh V: LeuFeatx:
Deep learning-based feature extractor for the diagnosis of acute
leukemia from microscopic images of peripheral blood smear. Comput
Biol Med. 142:1052362022. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Infante JB: Predictors of early death in
acute promyelocytic leukemia. Med Sci (Basel).
13:3002025.PubMed/NCBI
|
|
46
|
Bennett JM, Catovsky D, Daniel MT,
Flandrin G, Galton DA, Gralnick HR and Sultan C: Proposals for the
classification of the acute leukaemias. French-American-British
(FAB) co-operative group. Br J Haematol. 33:451–458. 1976.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Bain BJ: Diagnosis from the Blood Smear. N
Engl J Med. 353:498–507. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Döhner H, Wei AH, Appelbaum FR, Craddock
C, DiNardo CD, Dombret H, Ebert BL, Fenaux P, Godley LA, Hasserjian
RP, et al: Diagnosis and management of AML in adults: 2022
recommendations from an international expert panel on behalf of the
ELN. Blood. 140:1345–1377. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Sanz MA, Fenaux P, Tallman MS, Estey EH,
Löwenberg B, Naoe T, Lengfelder E, Döhner H, Burnett AK, Chen SJ,
et al: Management of acute promyelocytic leukemia: Updated
recommendations from an expert panel of the European LeukemiaNet.
Blood. 133:1630–1643. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Qiao Y, Zhang Y, Liu N, Chen P and Liu Y:
An End-to-End pipeline for early diagnosis of acute promyelocytic
leukemia based on a compact CNN model. Diagnostics (Basel).
11:12372021. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Manescu P, Narayanan P, Bendkowski C, Elmi
M, Claveau R, Pawar V, Brown BJ, Shaw M, Rao A and Fernandez-Reyes
D: Detection of acute promyelocytic leukemia in peripheral blood
and bone marrow with annotation-free deep learning. Sci Rep.
13:25622023. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Xiao Y, Huang Z, Wu J, Zhang Y, Yang Y, Xu
C, Guo F, Ni X, Hu X, Yang J, et al: Rapid screening of acute
promyelocytic leukaemia in daily batch specimens: A novel
artificial intelligence-enabled approach to bone marrow morphology.
Clin Transl Med. 14:e17832024. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Short NJ, Rytting ME and Cortes JE: Acute
myeloid leukaemia. Lancet. 392:593–606. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Walter RB, Othus M, Burnett AK, Löwenberg
B, Kantarjian HM, Ossenkoppele GJ, Hills RK, van Montfort KG,
Ravandi F, Evans A, et al: Significance of FAB subclassification of
‘acute myeloid leukemia, NOS’ in the 2008 WHO classification:
Analysis of 5848 newly diagnosed patients. Blood. 121:2424–2431.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Agaian S, Madhukar M and Chronopoulos TA:
Automated screening system for acute myelogenous leukemia detection
in blood microscopic images. IEEE Syst J. 8:995–1004. 2014.
View Article : Google Scholar
|
|
56
|
Bigorra L, Merino A, Alférez S and
Rodellar J: Feature analysis and automatic identification of
leukemic lineage blast cells and reactive lymphoid cells from
peripheral blood cell images. J Clin Lab Anal. 31:e220242017.
View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Huang F, Guang P, Li F, Liu X, Zhang W and
Huang W: AML, ALL, and CML classification and diagnosis based on
bone marrow cell morphology combined with convolutional neural
network: A STARD compliant diagnosis research. Medicine
(Baltimore). 99:e231542020. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Wang M, Dong C, Gao Y, Li J, Han M and
Wang L: A deep learning model for the automatic recognition of
aplastic anemia, myelodysplastic syndromes, and acute myeloid
leukemia based on bone marrow smear. Front Oncol. 12:8449782022.
View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Eckardt JN, Middeke JM, Riechert S,
Schmittmann T, Sulaiman AS, Kramer M, Sockel K, Kroschinsky F,
Schuler U, Schetelig J, et al: Deep learning detects acute myeloid
leukemia and predicts NPM1 mutation status from bone marrow smears.
Leukemia. 36:111–118. 2022. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Yu Z, Li J, Wen X, Han Y, Jiang P, Zhu M,
Wang M, Gao X, Shen D, Zhang T, et al: AMLnet, A deep-learning
pipeline for the differential diagnosis of acute myeloid leukemia
from bone marrow smears. J Hematol Oncol. 16:272023. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Elhassan TA, Mohd Rahim MS, Siti Zaiton
MH, Swee TT, Alhaj TA, Ali A and Aljurf M: Classification of
atypical white blood cells in acute myeloid leukemia using a
two-stage hybrid model based on deep convolutional autoencoder and
deep convolutional neural network. Diagnostics (Basel). 13:1962023.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Zhang Z, Arabyarmohammadi S, Leo P,
Meyerson H, Metheny L, Xu J and Madabhushi A: Automatic myeloblast
segmentation in acute myeloid leukemia images based on adversarial
feature learning. Comput Methods Programs Biomed. 243:1078522024.
View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Zhang C and Zhu J: AML leukocyte
classification method for small samples based on ACGAN. Biomed Tech
(Berl). 69:491–499. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Aby AE, Salaji S, Anilkumar KK and Rajan
T: Classification of acute myeloid leukemia by pre-trained deep
neural networks: A comparison with different activation functions.
Med Eng Phys. 135:1042772025. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Cheng H, Ding J, Wang J, Xiao Y, Jin X,
Zhang Y, Yang Y, Xu H, Cao X, Guo F, et al: Predicting
RUNX1::RUNX1T1 genetic abnormalities in acute myeloid leukemia from
bone marrow smears: Can artificial intelligence do better?
iScience. 28:1129982025. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Iacobucci I and Mullighan CG: Genetic
basis of acute lymphoblastic leukemia. J Clin Oncol. 35:975–983.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Shafique S and Tehsin S: Acute
lymphoblastic leukemia detection and classification of its subtypes
using pretrained deep convolutional neural networks. Technol Cancer
Res Treat. 17:15330338188027892018. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Rehman A, Abbas N, Saba T, Rahman SIU,
Mehmood Z and Kolivand H: Classification of acute lymphoblastic
leukemia using deep learning. Microsc Res Tech. 81:1310–1317. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
69
|
Rezayi S, Mohammadzadeh N, Bouraghi H,
Saeedi S and Mohammadpour A: Timely diagnosis of acute
lymphoblastic leukemia using artificial intelligence-oriented deep
learning methods. Comput Intell Neurosci. 2021:54781572021.
View Article : Google Scholar : PubMed/NCBI
|
|
70
|
Pałczyński K, Śmigiel S, Gackowska M,
Ledziński D, Bujnowski S and Lutowski Z: IoT Application of
transfer learning in hybrid artificial intelligence systems for
acute lymphoblastic leukemia classification. Sensors (Basel).
21:80252021. View Article : Google Scholar : PubMed/NCBI
|
|
71
|
Jiang Z, Dong Z, Wang L and Jiang W:
Method for diagnosis of acute lymphoblastic leukemia based on
ViT-CNN ensemble model. Comput Intell Neurosci. 2021:75298932021.
View Article : Google Scholar : PubMed/NCBI
|
|
72
|
Mirmohammadi P, Ameri M and Shalbaf A:
Recognition of acute lymphoblastic leukemia and lymphocytes cell
subtypes in microscopic images using random forest classifier. Phys
Eng Sci Med. 44:433–441. 2021. View Article : Google Scholar : PubMed/NCBI
|
|
73
|
Musleh S, Islam MT, Alam MT, Househ M,
Shah Z and Alam T: ALLD: Acute lymphoblastic leukemia detector.
Stud Health Technol Inform. 289:77–80. 2022.PubMed/NCBI
|
|
74
|
Jawahar M, Sharen H, Janianbarasi L and
Gandomi AH: ALNett: A cluster layer deep convolutional neural
network for acute lymphoblastic leukemia classification. Comput
Biol Med. 148:1058942022. View Article : Google Scholar : PubMed/NCBI
|
|
75
|
Almadhor A, Sattar U, Al Hejaili A, Ghulam
Mohammad U, Tariq U and Ben Chikha H: An efficient computer
Vision-based approach for acute lymphoblastic leukemia prediction.
Front Comput Neurosci. 16:10836492022. View Article : Google Scholar : PubMed/NCBI
|
|
76
|
Atteia G, Alhussan AA and Samee NA:
BO-ALLCNN: Bayesian-based optimized CNN for acute lymphoblastic
leukemia detection in microscopic blood smear images. Sensors
(Basel). 22:55202022. View Article : Google Scholar : PubMed/NCBI
|
|
77
|
Chen YM, Chou FI, Ho WH and Tsai JT:
Classifying microscopic images as acute lymphoblastic leukemia by
Resnet ensemble model and Taguchi method. BMC Bioinformatics.
22:6152022. View Article : Google Scholar : PubMed/NCBI
|
|
78
|
Abunadi I and Senan EM: Multi-Method
diagnosis of blood microscopic sample for early detection of acute
lymphoblastic leukemia based on deep learning and hybrid
techniques. Sensors (Basel). 22:16292022. View Article : Google Scholar : PubMed/NCBI
|
|
79
|
Albeeshi AA and Alshanbari HS: Modeling of
the acute lymphoblastic leukemia detection by convolutional neural
networks (CNNs). Curr Med Imaging. 19:734–748. 2023.PubMed/NCBI
|
|
80
|
Ahmed IA, Senan EM, Shatnawi HAS,
Alkhraisha ZM and Al-Azzam MMA: Hybrid techniques for the diagnosis
of acute lymphoblastic leukemia based on fusion of CNN features.
Diagnostics (Basel). 13:10262023. View Article : Google Scholar : PubMed/NCBI
|
|
81
|
Kaur M, AlZubi AA, Jain A, Singh D, Yadav
V and Alkhayyat A: DSCNet: Deep Skip Connections-based dense
network for ALL diagnosis using peripheral blood smear images.
Diagnostics (Basel). 13:27522023. View Article : Google Scholar : PubMed/NCBI
|
|
82
|
Huang ML and Huang ZB: An ensemble-acute
lymphoblastic leukemia model for acute lymphoblastic leukemia image
classification. Math Biosci Eng. 21:1959–1978. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
83
|
Jawahar M, Anbarasi LJ, Narayanan S and
Gandomi AH: An attention-based deep learning for acute
lymphoblastic leukemia classification. Sci Rep. 14:174472024.
View Article : Google Scholar : PubMed/NCBI
|
|
84
|
MoradiAmin M, Yousefpour M,
Samadzadehaghdam N, Ghahari L, Ghorbani M and Mafi M: Automatic
classification of acute lymphoblastic leukemia cells and lymphocyte
subtypes based on a novel convolutional neural network. Microsc Res
Tech. 87:1615–1626. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
85
|
Elrefaie RM, Mohamed MA, Marzouk EA and
Ata MM: A robust classification of acute lymphocytic leukemia-based
microscopic images with supervised Hilbert-Huang transform. Microsc
Res Tech. 87:191–204. 2024. View Article : Google Scholar : PubMed/NCBI
|
|
86
|
Alsaykhan LK and Maashi MS: A hybrid
detection model for acute lymphocytic leukemia using support vector
machine and particle swarm optimization (SVM-PSO). Sci Rep.
14:234832024. View Article : Google Scholar : PubMed/NCBI
|
|
87
|
Mei L, Lian C, Han S, Jin S, He J, Dong L,
Wang H, Shen H, Lei C and Xiong B: High-accuracy and lightweight
image classification network for optimizing lymphoblastic leukemia
diagnosisy. Microsc Res Tech. 88:489–500. 2025. View Article : Google Scholar : PubMed/NCBI
|
|
88
|
Mei L, Lian C, Han S, Ye Z, Hua Y, Sun M,
He J, Ye Z, Mei M, Yalikun Y, et al: High-efficiency spatially
guided learning network for lymphoblastic leukemia detection in
bone marrow microscopy images. Comput Biol Med. 196:1108602025.
View Article : Google Scholar : PubMed/NCBI
|
|
89
|
Anand V, Bachhal P, Koundal D and Dhaka A:
Deep learning model for early acute lymphoblastic leukemia
detection using microscopic images. Sci Rep. 15:291472025.
View Article : Google Scholar : PubMed/NCBI
|