Introduction
Nuclear receptors (NRs) are one of the essential
classes of transcriptional factors. NRs play a critical role in all
aspects of human development, metabolism and physiology. Since they
generally act as ligand-activated transcription factors, they are
an essential component of cell signaling (1). Glucocorticoid receptor (GR) is part
of the NRs protein superfamily that is clustered in the family of
the steroid hormone. GR has been shown to interact with a variety
of proteins and is a transcriptional factor that regulates multiple
genes, while is simultaneously expressed in almost every cell in
the human body. Glucocorticoids activate GR (2). Glucocorticoids greatly contribute to
the maintenance of homeostasis and partake in the regulation of the
immune system (3). They exert an
impressively diverse and tissue-specific effect. In the absence of
glucocorticoids, the majority of the GR resides in the cytoplasm,
forming a complex with heat shock proteins. Upon ligand binding,
GRs are released from the complex and translocate to the nucleus,
where they influence the transcription of a number of
glucocorticoid-responsive genes (2). GR is the product of a single gene,
NR3C1, which was first cloned in 1985(4). NR3C1 is located on chromosome
5q31-32 in humans and undergoes alternative processing, leading to
functionally distinct subtypes of GR. The human GR gene (NR3C1)
consists of 9 exons. The predominant isoforms of GR in humans are
hGRα and hGRβ (5). These isoforms
are identical through amino acid 727, but then diverge. hGRα has an
additional 50 amino acids, while hGRβ has an additional 15
non-homologous amino acids. The most well-studied isoform is hGRα,
which is comprised of 777 amino acids, while hGRβ has a dominant
negative effect on hGRα (6). As
far as GRs taxonomy is concerned, it specifically belongs to the
steroid hormone receptor (SHR) subfamily (7). GR belongs to the 3-ketosteroid
receptor group, which also includes receptors for
mineralocorticoids (MRs), progesterone (PR) and androgens (ARs).
Another steroid hormone receptor related to the previous group,
with an essential role in sexual maturation, is the estrogen
receptor (ER). The majority of NRs can be considered an assembly of
smaller protein modules. They share a common modular structure
composed of a highly conserved DNA-binding domain (DBD; C), a less
conserved ligand-binding domain/LBD (E) and some less extensively
studied and highly variable N-terminal (A-B) and C-terminal domains
(F). The N-terminal regions of NRs sometimes harbor potent
transcriptional activation functions, known as activation
function-1 (AF-1), which is independent of the LBD-ligand
interaction. LBD is the site on which ligands bind, and where the
main interaction with coregulators takes place. In some cases, the
LBD can form the hetero- or homodimerization surfaces in NRs.
Within the LBD also lies the activation function-2 (AF-2), with it
referring to the recruitment of transcriptional activators in a
ligand-dependent manner (8). A
flexible hinge region/HR connects the DBD and LBD (D), playing a
crucial role in the selection of the repertoire of DNA-binding
sites. The hinge region contains a series of residues that interact
with the DNA minor groove towards the C-terminal of the DBD
[C-terminal extensions (CTEs)]. NRs bind to the regulatory regions
of target genes as homodimers, heterodimers and monomers. The major
steroid hormone receptors (GR, MR, PR, AR and ER) bind mainly, as
homodimers to response elements, configured as palindromes composed
of two hexad nucleotide sequences separated by 3 base pairs
(9). Apart from homodimerization,
monomeric binding seems to also play a significant role in
tissue-specific target gene expression in GRs (10). In contrast to steroid receptors,
non-steroid receptors seem to bind DNA as part of a heterodimer
with retinoid X receptor (RXR). They mainly bind to response
elements composed of 2 hexad half-sites arranged as tandem repeats
(9). The classical mode of action
of NRs implies that in the absence of their ligands, they behave as
transcriptional repressors through the recruitment of specific
corepressors. Ligand binding in the specific ligand binding pocket
induces a conformational change of the receptor, leading to the
release of the corepressors, the recruitment of co-activators and
the subsequent transactivation of genes (11). A proportion of unligated NRs, and
more specifically, several steroid hormones, reside in the nucleus
bound to DNA (12). GR and MR are
exceptions since they reside in the cytoplasm in association with a
variety of proteins in the absence of ligand.
NRs form an ancient and conserved family that arose
early in the metazoan lineage (11). NR molecular evolution is
characterized by major events of gene duplication and gene losses.
During the evolution of species, gene duplication and loss are not
regularly distributed in the NR superfamily evolution (13). NR gene duplication seems to be
frequent, while gene loss in the superfamily is rare. In some
cases, gene loss is paralleled by duplication of a specific set of
genes, which gives rise to a greater diversity of NRs. This
observation suggests that the lineage-specific expansion of one
gene can more than compensate for an overall trend to loss. A great
difficulty emerges in studying NR evolution, since NR ligands are
distinctive. NR ligands are small molecules not encoded by genes;
hence, the ligand is not the product of a single gene, but the
product of a metabolic pathway which interacts with other pathways.
This attribute is an important facet of their role in signaling.
Therefore, evolution does not occur only on a single gene by the
slow accumulation of mutations, but on an entire network of genes
(11). Thus, predicting how the
specificity of a receptor for its ligand evolves is not an easy
task.
Data on the origins of NRs suggest that they were
not a hormonal receptor with high affinity for a particular ligand
and that feature was acquired later during evolution. It is
considered that the first NR had the ability to bind different
molecules with small affinity (14). This is not surprising, since data
suggest that a single NR can bind several ligands, with different
biological activities, and those ligands can act as modulators that
can selectively activate an NR in a tissue or target specific
manner (11). Consequently,
investigating the mechanisms through which selectivity functions in
NRs is an important step in understanding their evolutionary
history. An analysis of the mechanisms through which mutations
alter the selectivity and function of receptors can be considered a
valid method in decrypting NR receptor evolution.
Data and methods
Dataset collection and filtering
A search was conducted on the RSCB Protein Data Bank
(PDB) database (15) for amino
acid sequences that are related to the NR LBD, followed by the
acquisition of resulting data. Specific sequences that responded to
the query but did not include the LBD were removed from the
dataset, by using regular expressions techniques and local
alignments with reference sequences. In total, 420 NR ligand
binding domain protein structures were downloaded from several
species.
Sequence alignment
Multiple sequence alignment was performed using the
MATLAB Bioinformatics Toolbox, utilizing the progressive multiple
alignment method and a guide tree (16,17). Pairwise distances between protein
sequences were computed following pairwise alignment with the
Gonnet scoring matrix (18) and
followed by counting the proportion of sites at which each pair of
sequences are different. The guide tree was calculated by the
neighbor-joining method assuming equal variance and independence of
evolutionary distance estimates. The visualization of the consensus
sequence was performed using the JalView platform (19) and based on the multiple sequences
alignment results and parameters such as amino acids quality and
conservation. A more specific alignment with representative members
of the Steroid hormone receptors was performed using the Molecular
Operating Environment (MOE) (20).
Mutation analysis
A collection of currently known natural occurring NR
LBD mutations was assembled following a literature review. The
selected mutations harmed ligand binding. The resulting dataset
includes natural occurring LBD mutations in the following
receptors: GR, MR, AR, ERα, peroxisome proliferator-activated
receptor (PPAR)γ, retinoid acid receptor (RAR)α, RARβ, thyroid
hormone receptor (THR)α, THRβ, liver X receptor (LXR)α, vitamin D
receptor (VDR), hepatocyte nuclear factor 4 (HNF4A), steroidogenic
factor 1 (SF1), RAR-related orphan receptor (ROR)α, RORβ and RORγ
(Table SI). Based on the
generated list, all mutations were detected and marked on the
consensus sequence from the multiple sequence alignment, and in the
alignment of the representative structures of the SHR.
Additionally, the mutation rate was examined in all NRs (Table SII). Specific mutation positions
were highlighted by different colors in order to showcase the
mutation rate within the NRs (Fig.
S1).
Structural and functional
analysis
A thorough analysis of NR LBD structural comparisons
was achieved by superimposing the structures and by calculating a
matrix of root mean square deviation (RMSD). The structural
comparisons between the 420 structures were performed utilizing the
structural superposition method, as described in the MATLAB
Bioinformatics Toolbox (21).
This method computes and applies a linear transformation to
superpose the coordinates of the atoms of the first structure to
the coordinates of the atoms of the second structure. Alpha carbon
atom coordinates of single chains for each structure are considered
for computing the linear transformation. The structural similarity
matrix was displayed using MATLAB in 5 different colors (blue for
the range 0 to 1, light blue for the range 1.1 to 2, light gray for
the range 2.1 to 3, orange for the range 3.1 to 4.9 and red for the
values ≥5).
A more in-depth look at steroid hormone receptor
structures was gained using MOE (20,22-24),
where all the extracted mutations and interactions sites with
several proteins and ligands were identified and studied.
Specifically, each PDB entry was examined for ligand interaction
using the ligand interaction function. MOE showcased the LBD
amino-acids that interacted with said ligands or co-activators.
Finally, beneficial information from the NCBI conserved domain
database was extracted and assigned with the MOE results in the
consensus sequence from the multiple sequences alignment.
Phylogenetic analysis
A specialized phylogenetic analysis was performed
using the MATLAB Bioinformatics Toolbox (20) utilizing the Unweighted Pair-Group
Method (UPGMA) (25-27)
and a specific hybrid matrix of pairwise distances (28). This matrix combines both
information from the distance matrix of the multiple sequence
alignment, and the RMSD matrix of the structural analysis. The
combined specialized matrix is calculated through element by
element matrices proliferation (29,30). This technique allows the
clustering of less similar proteins in sequence level that are more
conserved in the structural level. Finally, the constructed
phylogenetic tree was visualized using the MEGA (31) radiation option, and the final
clusters were separated by different colors.
Ligand analysis
The notion of chemical similarity plays an important
role in predicting the properties of chemical compounds, clustering
chemicals, and in particular, in conducting functional analysis
studies. The calculation of the similarity of any two molecules is
achieved by comparing their molecular fingerprints (32). These fingerprints are comprised of
structural information about the molecule which has been encoded as
a series of bits. The most popular similarity measure for comparing
chemical structures represented by means of fingerprints is the
Tanimoto coefficient (33).
First, a list with all extract ligands that
co-crystallized in the corresponding SHR LBD structures was created
(Table SIII). We set the same
order in the list as the structures' order from the phylogenetic
analysis. The structural comparisons between the 179 SHR ligands
were performed using the Tanimoto coefficient algorithm (34). The Tanimoto coefficient ranges
from 0, when the fingerprints have no bits in common, to 1, when
the fingerprints are identical. In the end, all the similarities
were saved in a chemical-specific similarity matrix. Finally, the
chemical-specific similarity matrix was displayed using MATLAB in 4
different colors (blue for the range 1 to 0.9, light blue for the
range 0.89 to 0.7, purple for the range 0.69 to 0.6 and black for
the range 0.59 to 0).
Results
The 4 major clusters of the NR
LBD
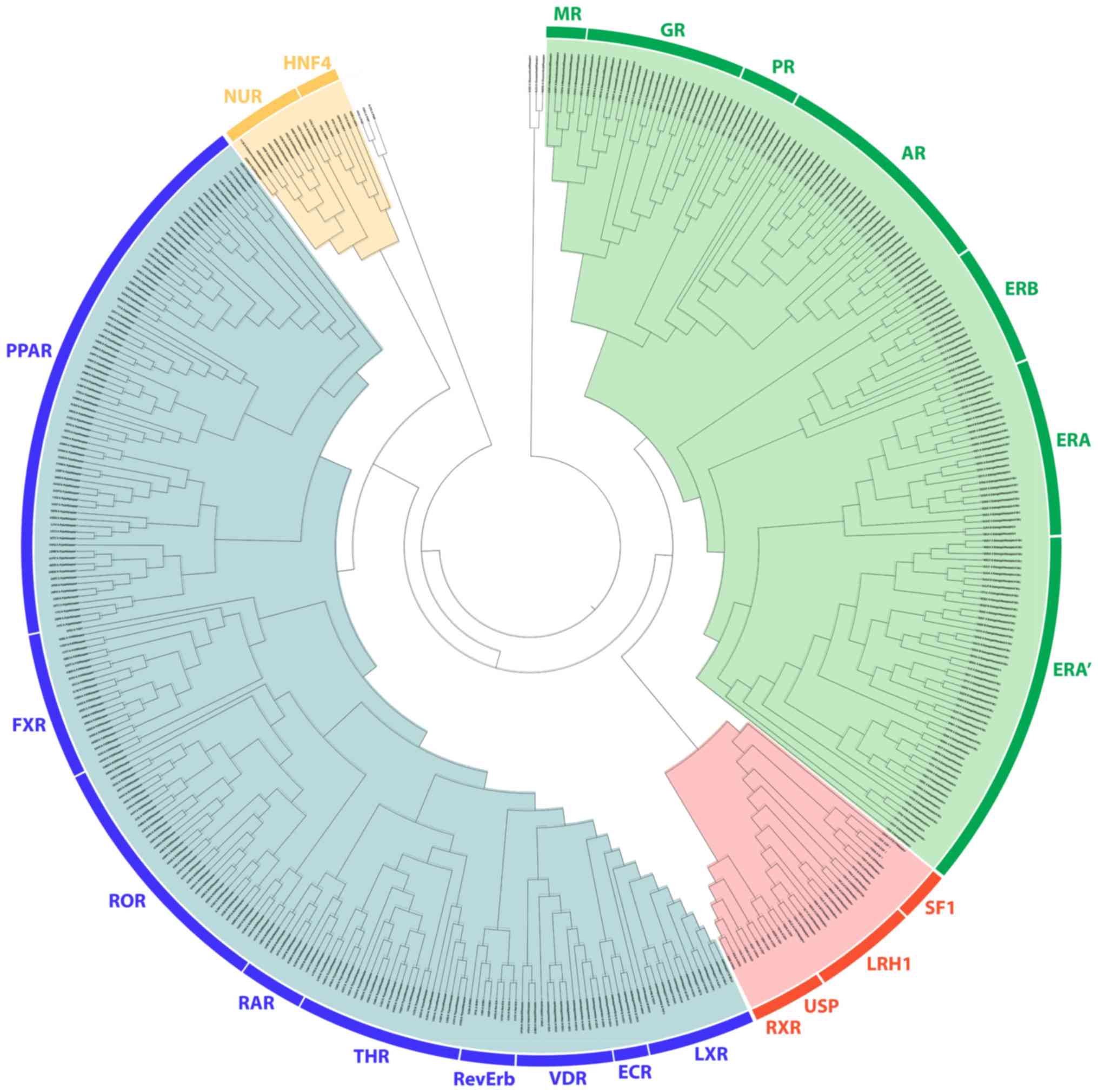
Phylogenetic analysis revealed a distinct separation
of NR LBDs into 4 monophyletic branches, the steroid hormone
receptor-like cluster, the thyroid hormone-like receptors cluster,
the retinoid X-like and steroidogenic factor-like receptor cluster
and the nerve growth factor-like/HNF4 receptor cluster (Fig. 1). All known steroid hormone
receptors (GRs, MRs, PRs, ARs and ERs) have been grouped in
separated sub-clusters in the steroid hormone-like receptor
cluster. As expected, the LBD of GRs, MRs, PRs and Ars was found to
be well-related to the ERs. The ER receptors also separate into two
different subclasses, the ERα and the ERβ, from which there is a
substantial separation of the ERα in 2 groups. A more detailed
structural analysis was performed in order to understand this
strange distribution of the ERs, and the results are described
below. Moreover, the SRH-like cluster was found to be directly
related with the second cluster of the retinoid X-like and
steroidogenic factor-like receptor cluster. The linkage of this
protein family with the steroid hormone receptor family was noted
for the first time, to the best of our knowledge. Within the second
cluster are contained the RXR, the liver receptor homolog 1 (LRH1),
the SF1 and the ultraspiracle protein (USP) subunit of the ecdysone
receptor. The SF1 is a protein that controls many aspects of
adrenal and reproductive function. It is encoded by the NR5A1 gene,
which is a member of the NR protein family (35). Along with the LRH1 encoded by the
NR5A1 gene, they belong to the steroidogenic factor-like subfamily
of NRs. They both hold a critical role in steroidogenesis (36). Another thoroughly different
observation on the SHR branch is its inclusion of the RXR and its
Drosophila homolog USP, which is a subunit of the ecdysone
receptor (37,38).
On the third monophyletic branch, the LBD of the
THR-like cluster, all known members are grouped, including THR,
PPAR, LXR, VDR, RAR, FXR, ROR and orphan nuclear hormone receptor
(NR1D1; RevErb). The EcR subunit of the ecdysone receptor also
belongs in this cluster. The EcR subunit of the ecdysone receptor
is closely related to the FXR, based on their similarities in the
DNA binding domain (39). The
EcR/USP heterodimer seems to be the corresponding arthropod complex
to the FXR/RXR heterodimer. This analysis provides the insight that
this heterodimer appears to be composed of 2 subunits with each one
belonging to a different monophyletic branch. Moreover, the LBD of
the THR-like cluster was found to be directly related to the fourth
cluster of the LBDs of the nerve growth factor (Nurr77) and HNF4.
HFN4α and Nurr77 are suggested to be in the same subfamily. Some
other interesting observations are the existence of a number of
distinct GRs showcasing marked differences with regard to other
NRs.
The conserved signaling motifs of the
NR LBD
An in-depth review of the consensus sequence and NR
LBD mutations provides a basis for 7 highly conserved signaling
motifs (Fig. 2A). A more targeted
approach to steroid hormone receptors can provide data on their
importance for NR function (Fig.
2B). Motif A occupies consensus sequence positions 378 to 385
(Fig. 2A). This LLxxL motif is an
inverse NR-box (LLxxL) and still contains the ability to interact
with several NRs (40). Based on
NCBI's conserved domain database (CDD) (41), this position is one of the main
ligand interaction sites of the family and is of utmost importance
to NRs. It also coincides with the SHR interaction Motif A
(Fig. 2B). Motif B occupies
consensus sequence positions 391 to 401 (Fig. 2A). A cross-check with CCD NR
action sites (Fig. 2B) reveals
that Motif B resides in a region critical to co-activator function.
The GR mutation V575G does support this hypothesis (42). V575 is part of the AF-2 surface,
and it contributes directly to the attraction of LxxLL (NR-box
sequence motif). A mutation on that specific region hinders the
ability of co-activators to interact with the NR. The 1L2I PDB
entry for ERα also implies the region's high impact role in
interacting with co-activators, and more specifically, the NR-box
sequence motif. Motif C occupies positions 404 to 413, a region
that is also critical to co-activator function (Fig. 2A). Motif C could be described as
LxxDDQ. Along with the R (402 alignment position) residue, this
motif creates a structure specific to GRs, ARs and PRs. The study
by Bledsoe et al was the first one that recognized this
specific region's critical role in LBD function (7). Motif C partakes in the creation of
GR's second charge clamp. This distinct structure is vital to the
binding of the third NR-box of co-activators. The residues
responsible for the second charge clamp are not present in the
remaining SHRs (ERs and MRs). Motif D occupies positions 512 to
516. This region is part of the highly conserved C-terminal end of
the helix eight domain in SHRs, whose function is critical to
ligand binding. Mutations in this motif, including GR's L672P and
AR's L813P lead to a complete lack of ligand binding (43,44). More experiments also showcase that
the mutant proteins are possibly prone to higher degradation
(44). Motif E is an inverse
NR-box and occupies positions 546 to 550. Motif F occupies
positions 568 to 575. This LxxLL motif (NR-box) is found in all
SHRs, with the ERα LxxLL comprising positions 568 to 572. Motif G
occupies alignment positions 601-613. It contains an ERα LxxLL
motif on positions 601 to 609, a PPAR LxxLL motif on positions 605
to 609, and an LLxxL ERα motif on position 609 to 611. It is quite
intriguing that ERα exhibits a motif of LxxLLLxxL, which contains
both an NR-box and its inverse. A prime example of a malfunctioning
mutation on motif G is the Y537S, which is present on ERαs
appearing in breast cancer cells (45).
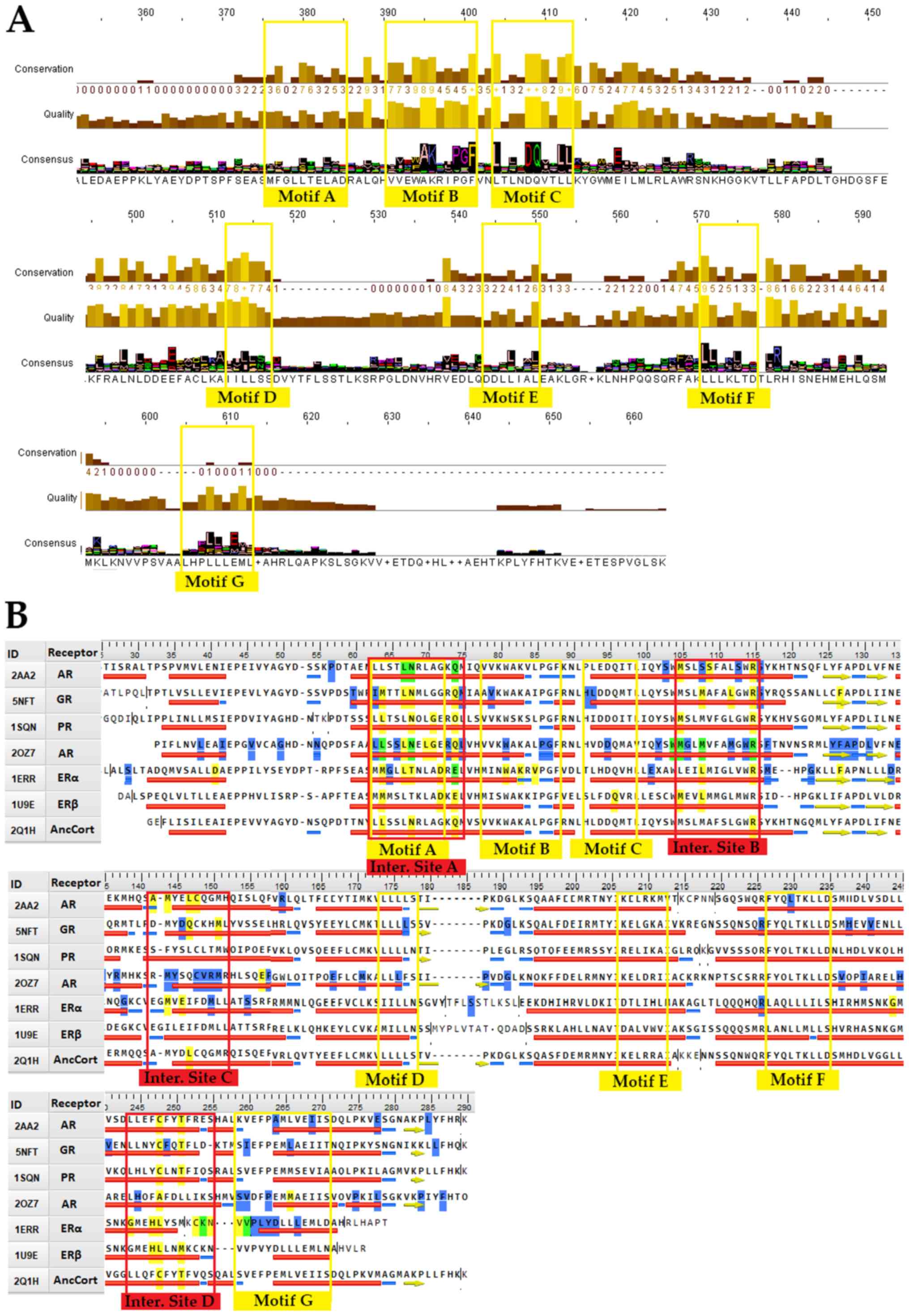 | Figure 2.Conserved signaling motifs and
interaction sites of the NR LBD. (A) Consensus sequence based on
the 420 NR LBD multiple sequences alignment results and parameters,
such as amino acids quality and conservation. The 7 major conserved
signaling motifs of the NR LBD have been marked in the consensus
sequence (colored yellow). (B) Sequence alignment of SHRs with the
7 conserved signaling motifs (colored yellow) and the 4 interaction
sites (colored red). The figure features representative sequences
from each SHR group plus an ancestral corticoid receptor. The
reference sequences used are PDB: 2AA2 for mineralocorticoid
receptor, PDB: 5NFT for glucocorticoid receptor, PDB: 1SQN for
progesterone receptor, PDB: 2OZ7 for androgen receptor, PDB: 1ERR
for estrogen receptor α, 1U9E for estrogen receptor β and PDB: 2Q1H
for the ancestral corticoid receptor. Specific amino acid residues
have been colored to showcase specific attributes. Yellow-colored
residues are known interaction points, blue-colored residues are
prone to mutation, while green-colored residues are both
interaction sites and prone to mutation. NR, nuclear receptor; LBD,
ligand-binding domain; PDB, Protein Data Bank; SHR, steroid hormone
receptor. |
Mutations in the NR LBD
A list of all known NR LBD natural mutations was
composed in this study from previous publications (Tables SI and SII). Studying both the common natural
mutations and their results gives way to a few observations.
Firstly, highly conserved sequences are not prone to mutations
(Fig. S1). They seem critical to
normal protein function, which leads to their evolutionary
conservation. Indeed, natural mutations on highly conserved
positions lead to deliberating effects. Some positions exhibit no
mutation at all. The lack of mutations may be attributed to their
quintessential role in protein function, since mutations on said
positions could be lethal, so no surviving phenotypes exist. The
majority of mutations on steroid receptors are associated with
hormone level changes, which are linked to their respective ligand.
It should be mentioned that phenotypes that could be attributed to
mutations on NRs may not exhibit said mutations on the final
protein product. A number of factors could be responsible for this
observation, such as epigenetic actions on NR genes, mutations in
non-coding regions affecting enhancers, or mutations on NR
cofactors (46). An in-depth look
into GR mutations confirmed the aforementioned. No mutations were
found on highly conserved NR amino acids. Mutations on GR LBD have
a severe effect on adrenocortical function. Mutations of GR LBD can
lead to Chrousos syndrome, a condition characterized by tissue
insensitivity to glucocorticoids. An interesting fact is that some
LBD mutations have a dominant negative effect. These kinds of
mutations on the LBD can be considered more severe than DBD
mutations since they affect normal functioning proteins as
well.
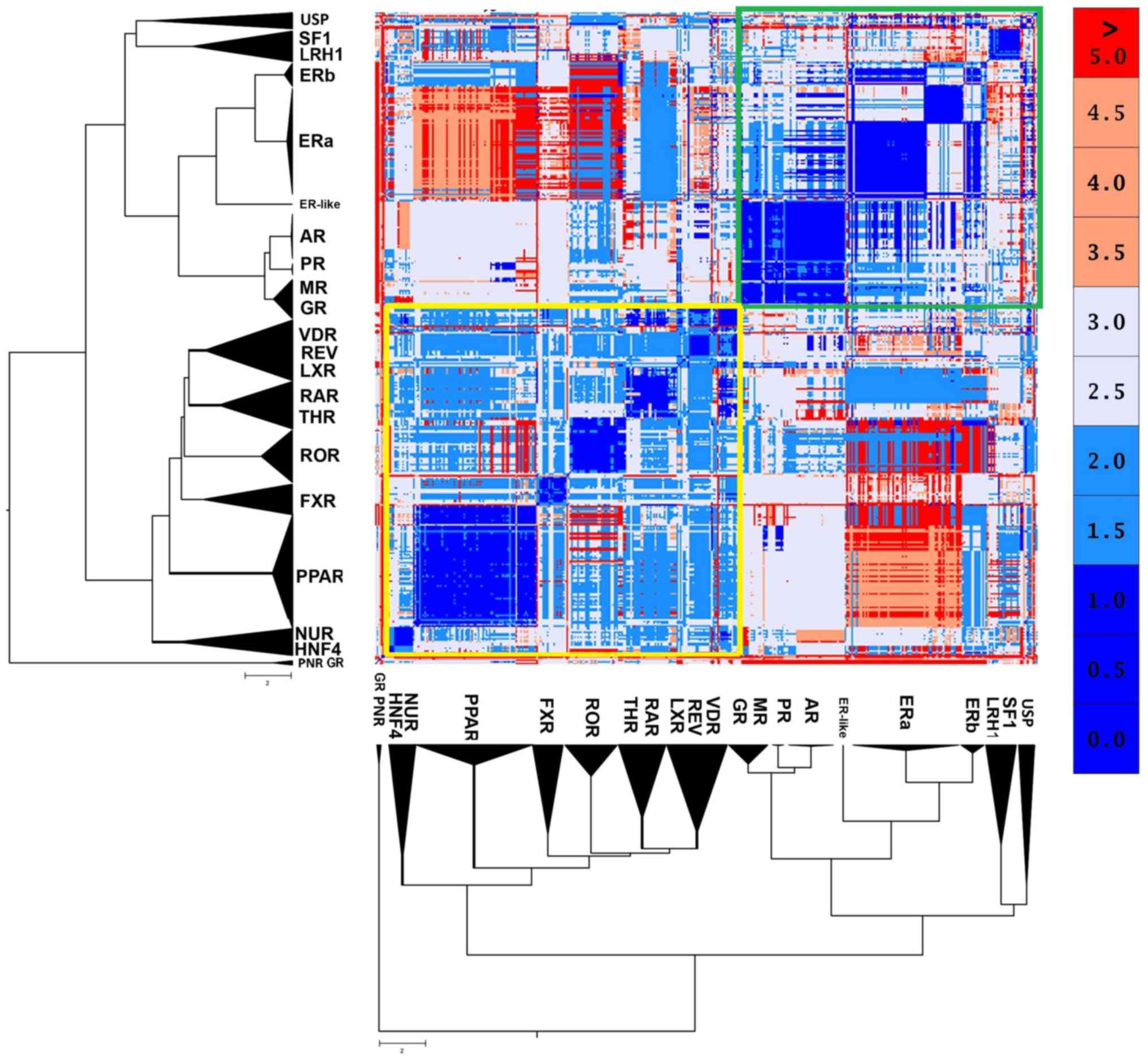
The 2 major canonical forms
NRs are structurally quite conserved. However, the
structural analysis results of NR LDB displayed 2 distinct
canonical forms (Fig. 3). The
first one seems to be prevalent in the SHR-like, while the second
one is prevailing in the THR-like receptors. The well-defined
retinoid X receptor-like/steroidogenic factor-like cluster of the
USP, SF1 and LRH1 receptors may appear phylogenetically close,
although they have a small structural correlation with the two
major forms. Based on these findings, the SHR-like LBD canonical
form exhibits highly conserved structural domains, albeit it
validates the notion above, that estrogen receptors are pretty
different from the rest of SHRs. A peculiar observation emerges,
one in which ERβ found to be more structurally similar to the rest
of SHRs than ERα (Fig. 3).
Another observation is the number of β-strands that compose SHRs is
4, while the number of α-helixes is not constant in each specific
entry. Based on crystallographic observations, a steroid hormone
receptor is formed either with 11 or 12 α-helixes. A specific
examination in an alignment featuring only GRs also showcases the
different activation states of NRs, based on the position of the
AF-2 containing helix (7). The
activation state is dependent on the ligand featured and the
presence or absence of co-activators/corepressors. On the other
hand, 3 GR LBD structures (PDB: 3H52, 4LSJ and 4MDD) and 3
photoreceptor-specific NRs (PNR) LBD structures (PDB: 4LOG, 4XAJ
and 4XAI) appear to distance themselves from the entirety of all NR
LBD structures. An in-depth look proposes that those GR LBD
structures correspond to a specific antagonist form of the
receptor, in which the dislocation of the 12th helix leads to
complete disruption of the receptor's function (21). The second major canonical form of
the thyroid hormone-like receptor LBD is highly conserved, with
small differences amongst receptors. These differences create
somewhat distinct subclasses, the PPAR-like, the ROR/THR, such as
VDR-like and the HNF4/Nur77-like.
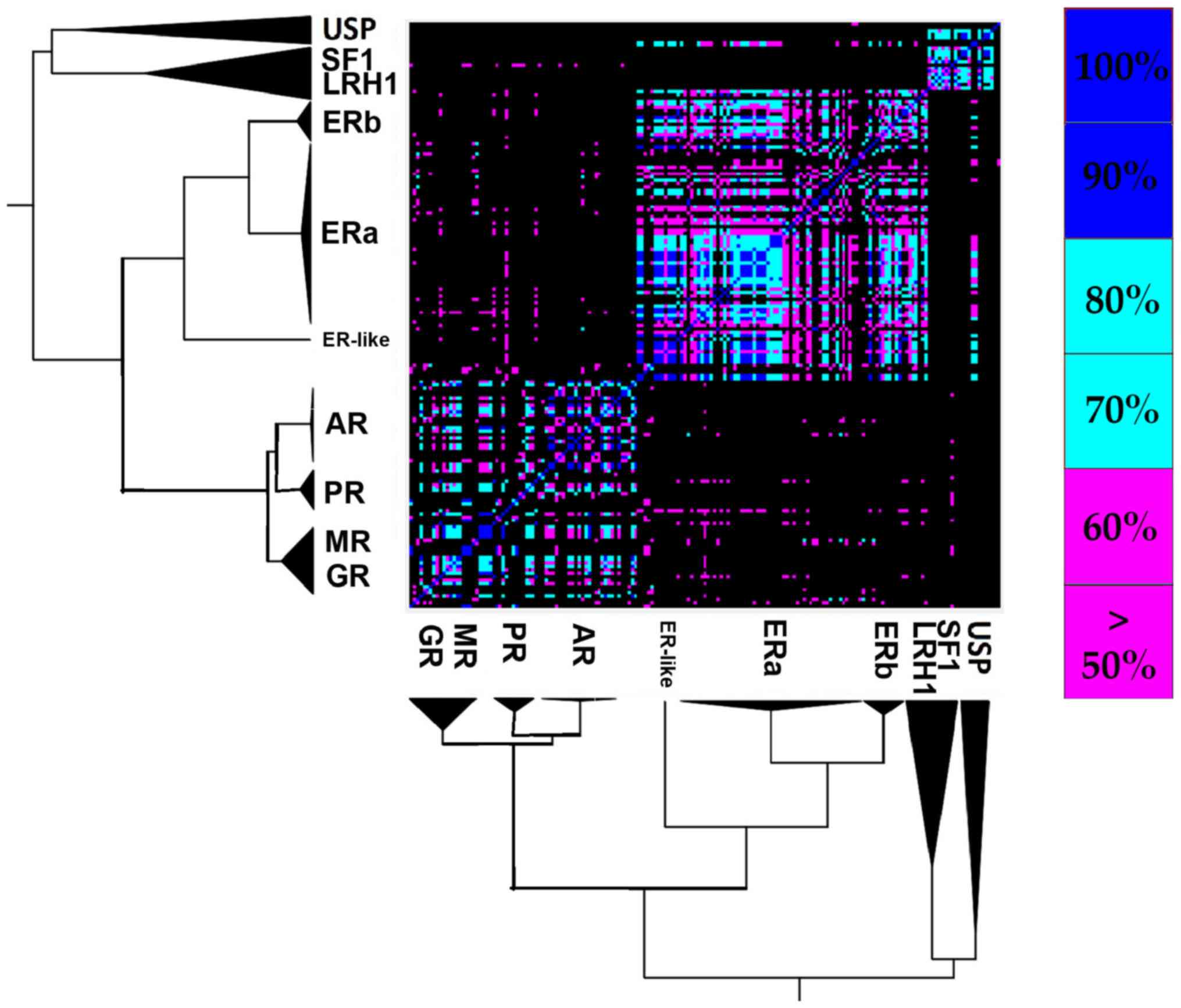
The 3 ligand specificity clusters of
the SHR LBD
Ligand specificity is one of the major and most
important characteristics of NRs (47). NR ligands are small hydrophobic
molecules that bind their corresponding NR LBD's hydrophobic
pocket. A list featuring all SHR ligands that were co-crystallized
in the corresponding SHR structures was created (Table SIII). The majority of ligands
seem to be receptor-specific, with the exceptions of MOF, which
binds both GR and PR and R18, which binds both PR and AR. This
observation is quite interesting since, as mentioned above, AR, MR,
PR and GR appear to create a steroid receptor sub-class of their
own, different from ERs. The PDB entry 1GS4 sheds light onto the
specific association between GR, PR and AR (48). Mutation T877A on the AR adds the
ability to bind progesterone 17b-estradiol and some anti-androgens.
The threonine at position 877 on AR is unique, though its
corresponding alignment position (251, Fig. 2B) partakes in ligand interactions
in all of the steroid receptors. This mutation gives the AR
abilities that have more in common with PR and ER. Mutation L701H
substantially impairs AR's own ability to bind androgen, but allows
it to bind cortisol (48). The
leucine at the 701 position is present in MR, PR and AR, while its
corresponding alignment position (67, Fig. 2B) partakes in ligand interaction
in GRs, PRs, ARs and ERs.
A comparative analysis of the ligands that have been
co-crystallized with their corresponding receptors was conducted,
in order to create a more clear-cut idea for the interactions that
are characteristic of the steroid hormone receptors' ligand binding
pocket (Fig. 4). A total of 94
unique steroid hormone receptors ligands were collected and were
compared in order to search for similarities and identify new
relations. Based on the results, there is a clear separation in the
main clusters, as shown in Fig.
4. Those are the USP/SF1/LRH1 ligand-specific cluster, the ER
ligand-specific cluster and the AR/PR/ MR/ GR ligand-specific
cluster. It is quite interesting that the USP/SF1/LRH1
ligand-specific cluster contains ligands that are similar to the ER
ligand-specific cluster. Moreover, focusing on the ER
ligand-specific cluster, there is a clear separation of ERα in 2
sub-clusters.
A sub-cluster of ERas forms and acts
as estrogen β, and the risk of breast cancer
As expected, the majority of NRs in the same
subfamily display high structural similarity, with a great example
being the ARs, PRs and MRs (Fig.
3). A review of the SHR-like branch adds some new information.
Contrary to the observations, some receptors indicate significant
structural differences even among themselves (Fig. 3). This observation points out that
the structures within the same category share more than one
canonical form due to a critical mutation, disease, or cancer. The
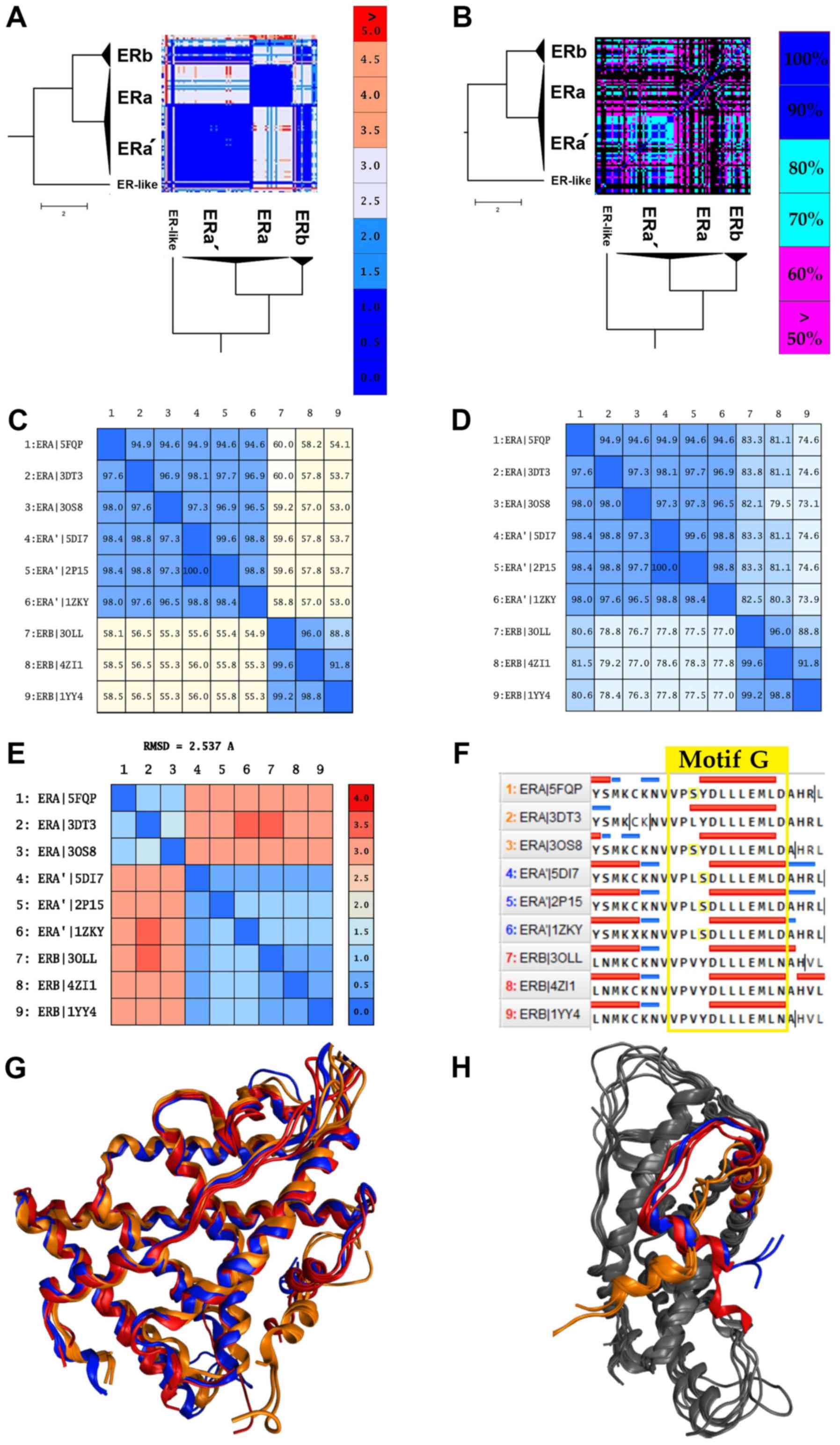
ERα data are provided in the same direction. Despite ERα showing
high sequence similarity (Fig.
5D), it is structurally separated into 2 different canonical
forms (Fig. 5A). An in-depth look
at the ERα structures comprising of the 2 canonical forms yielded
some interesting results. The 2 canonical forms of ERα correspond
to 2 different types of ERα.
The first canonical form of ERα contains various
mutant ERαs and a small number of wild-type receptors. The most
common mutants in this sub-cluster are located at positions C381,
C417, C530 and L536. The second canonical form of ERα, which will
be referred to as ERα', is mainly characterized by the Y537S
mutation. This mutation is commonly found in breast cancer patients
and is associated with resistance to several endocrine therapies
(49). More specifically, this
mutation is located in the AF-2 helix of the ERα' LBD and shifts
the receptor equilibrium towards the agonist conformation, even in
the absence of a ligand (23).
Moreover, the ERα' canonical form appears to be identical in the
structural level with the ERβ (RMSD <2) rather than the ERα,
while in the sequence level, all ERαs and ERαs' exhibit minimal
differences (Fig. 5A, C, E
and G). Based on these findings,
the Y537S mutation induces a critical conformational change in the
ERα structure. In particular, it changes the angle of the AF-2
helix of the ERα' LBD, in a position which is identified coequally
on the ERβ AF-2 helix (Fig. 5H).
The displacement and the 90˚ turn of the AF-2 helix plays a key
role in the action of the ERα' since it contains the signaling
‘Motif G’, one of the most important motifs of the NR LBD (Fig. 5F). This hypothesis seems to be
confirmed with the functional analysis we had performed by
analyzing all the available ligands and chemicals that have been
co-crystallized in the cavity of the ERα' and generally in all ERs.
Based on these results, ERα' can interact without any specificity
with all the identified ligands on ERs, including ERβ (Fig. 5B). This particularity has also
been described by several studies (48-50)
that are found to refer to a number of ERα' structures. In the
study by Nettles et al, a member of the ERα' (PDB: 2P15) was
crystallized, and it was concluded that this ERα could interact
with a wider array of pharmacophores than was previously thought
(50). In summary, it is possible
that the Y537S mutation induces a LBD domain with higher plasticity
in the ERα'.
Discussion
Hybrid phylogenetic analysis is more eloquent than a
common phylogenetic analysis. It combines both sequence and
structural information from the NRs' LBD to propose clusters with
higher confidence. It has been observed that protein structure is
more conserved than sequence (51), something visible on the current
analysis. Differences that are not detected on sequence analysis
alone are visible on this analysis. The phylogenetic analysis
performed depicts the distinct separation of NR LBDs into the
monophyletic branches of the steroid hormone receptor-like cluster,
the thyroid hormone-like receptors cluster, the retinoid X-like and
steroidogenic factor-like receptors cluster, and the nerve growth
factor-like/HNF4 receptors cluster and the LBD of GRs, MRs, PRs and
ARs found to be well-related to ERs. The linkage of the SRH-like
cluster with the second cluster of the retinoid X-like and
steroidogenic factor-like receptors cluster is noted for the first
time, at least to the best of our knowledge.
Researching motifs in the NR ligand binding domain
is of utmost importance. As mentioned above in the ‘Introduction’,
the ligand binding domain is a somewhat conserved domain. Finding
motifs that have been conserved through the evolutionary process
should highlight regions of critical importance in ligand binding.
Those regions can provide amino acid patterns that are a unique
signature to the NR LBD. Indeed, a previous study published in 1996
found signature sequences that are essential in maintaining the
ligand binding pocket structure (52). Those sequences seem to agree with
the proposed motifs that are described below, and some new ‘key’
motifs are introduced. The existence of NR-boxes or inverse
NR-boxes on SHRs may seem peculiar on first glance, but it should
not be a surprise. NRs have the ability of both hetero- and
homo-dimerization. Since NR-boxes can bind to specific NR regions,
they may have a role in the interaction between NRs, specifically
hetero and homo-dimerization. The results showcase moderate
conservation of specific amino acid sequences. Length-wise all NR
ligand binding domains are pretty similar. Looking through the
subcategory of steroid hormone receptors also provides some
interesting insights. There are many more sequence similarities, as
expected, though it is interesting, that estrogen receptors seem to
create a subcategory of their own since they exhibit amino acid
variations not present in the rest steroid hormone receptors.
Steroid hormone receptors share 4 main interaction
sites from where directly interact with several ligands and
co-activators (Fig. 2B). The
interaction sites A and B of the SHR LBD are well characterized and
found across all NR members. The interaction sites C and D should
be highly variable among NRs and somewhat conserved on SHRs. The
incorporation of both mutations and ligand interaction points can
specify the importance of these action regions. Fig. 2B provides information on both
interaction points and mutations. It shows that all interaction
sites are prone to mutations. This fact also seems logical, since
the activation sites are the ones responsible for the selectivity
of NRs. Mutations in those regions possibly are at the forefront of
NR evolution. The consensus sequence also adds to this speculation,
since the main interaction regions are characterized by relatively
small sequence conservation (Fig.
2B). It should also be highlighted that the interaction sites A
and B, which are conserved on all NRs, seem to be bridged by 3
regions of highly conserved motifs A, B and C.
Based on the comparative analysis of the
co-crystallized ligands to their receptors, there is a clear
separation in the main clusters, the USP/SF1/LRH1 ligand specific
cluster, the ER ligand specific cluster, and the AR/PR/MR/GR ligand
specific cluster. The USP/SF1/LRH1 ligand specific cluster contains
ligands similar to the ERs ligands specific cluster, where there is
evident separation of ERα in two sub-clusters. Moreover, ERα' can
interact without any specificity with all the identified ligands on
ERs, including ERβ, and there is a possible evidence that the Y537S
mutation induces a LBD domain with higher plasticity in the
ERα'.
In conclusion, NRs are vital transcription factors,
and the aforementioned information acquired can be used in a
variety of ways. The interaction sites and conserved motifs can be
used as selected targeted regions for novel drugs. The development
of new drugs can be achieved through specific in silico techniques
that can compose ligands, which can interact with those specific
regions and force new alterations in the protein's dynamics
(53). The phylogenetic analysis
also provided new insights for NR clustering and identified several
key regions that exist through evolution in the NR LBD such as the
amino acid repeating motif ‘LxxLL’ or ‘LLxxL’. The mutation
analysis highlighted mutational hotspots, while also providing
insights on their effects in structure and function, especially
when they are localized in NR conserved signaling motifs.
Structural and functional analysis of the NR LBD display two major
canonical forms and identify 3 ligand specific clusters within the
steroid hormone receptor family. Last but not least, a new
sub-cluster of ERα with a very specific canonical form has been
identified and related to breast cancer through a well-known
mutation in ERs. This new information may be of high importance in
order to understand the signaling mechanism underlying NRs and
cancer.
Supplementary Material
Mutation rates on the consensus
sequence from the multiple sequence alignment. Specific mutation
positions are highlighted by different colors in order to showcase
frequencies. Blue is used to mark positions which showcase
mutations on two different NRs, and green is used to mark positions
which showcase mutations on 3 different NRs; red is used to mark
positions which showcase mutations on four different NRs, while
purple is used to mark positions which showcase mutations on 5
different NRs. NRs, nuclear receptors.
List of various nuclear receptor
mutations located in the ligand binding domain.
Mutations rates on different nuclear
receptors based on the multiple alignment position.
List featuring all SHR ligands and
their corresponding receptors.
Acknowledgements
Not applicable.
Funding
DV would like to acknowledge funding from: i)
Microsoft Azure for Genomics Research Grant (CRM:0740983); ii)
FrailSafe Project (H2020-PHC-21-2015-690140) ‘Sensing and
predictive treatment of frailty and associated co-morbidities using
advanced personalized models and advanced interventions’, co-funded
by the European Commission under the Horizon 2020 research and
innovation program; iii) Amazon Web Services Cloud for Genomics
Research Grant (309211522729); iv) AdjustEBOVGP-Dx
(RIA2018EF-2081): Biochemical Adjustments of native EBOV
Glycoprotein in Patient Sample to Unmask target Epitopes for Rapid
Diagnostic Testing. A European and Developing Countries Clinical
Trials Partnership (EDCTP2) under the Horizon 2020 ‘Research and
Innovation Actions’ DESCA. EE would like to acknowledge funding by
the project ‘INSPIRED-The National Research Infrastructures on
Integrated Structural Biology, Drug Screening Efforts and Drug
Target Functional Characterization’ (Grant MIS 5002550) and by the
project: ‘OPENSCREEN-GR An Open-Access Research Infrastructure of
Chemical Biology and Target-Based Screening Technologies for Human
and Animal Health, Agriculture and the Environment’ (Grant MIS
5002691), which are implemented under the Action ‘Reinforcement of
the Research and Innovation Infrastructure’, funded by the
Operational Programme ‘Competitiveness, Entrepreneurship and
Innovation’ (NSRF 2014-2020) and co-financed by Greece and the
European Union (European Regional Development Fund).
Availability of data and materials
All data generated or analyzed during this study are
included in this published article or are available from the
corresponding author on reasonable request.
Authors' contributions
TM, LP, AE, FB, DV, GPC, EE have all equally
contributed to the writing, drafting, revising, editing, reviewing,
and the conception and design of the study. All authors have read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Robinson-Rechavi M, Escriva Garcia H and
Laudet V: The nuclear receptor superfamily. J Cell Sci.
116:585–586. 2003. View Article : Google Scholar
|
|
2
|
Chrousos GP: The glucocorticoid receptor
gene, longevity, and the complex disorders of Western societies. Am
J Med. 117:204–207. 2004.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Bereshchenko O, Migliorati G, Bruscoli S
and Riccardi C: Glucocorticoid-Induced Leucine Zipper: A Novel
Anti-inflammatory Molecule. Front Pharmacol. 10(308)2019.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Hollenberg SM, Weinberger C, Ong ES,
Cerelli G, Oro A, Lebo R, Thompson EB, Rosenfeld MG and Evans RM:
Primary structure and expression of a functional human
glucocorticoid receptor cDNA. Nature. 318:635–641. 1985.PubMed/NCBI View
Article : Google Scholar
|
|
5
|
Nicolaides NC, Galata Z, Kino T, Chrousos
GP and Charmandari E: The human glucocorticoid receptor: Molecular
basis of biologic function. Steroids. 75:1–12. 2010.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Kadmiel M and Cidlowski JA: Glucocorticoid
receptor signaling in health and disease. Trends Pharmacol Sci.
34:518–530. 2013.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Bledsoe RK, Montana VG, Stanley TB, Delves
CJ, Apolito CJ, McKee DD, Consler TG, Parks DJ, Stewart EL, Willson
TM, et al: Crystal structure of the glucocorticoid receptor ligand
binding domain reveals a novel mode of receptor dimerization and
coactivator recognition. Cell. 110:93–105. 2002.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Huang P, Chandra V and Rastinejad F:
Structural overview of the nuclear receptor superfamily: Insights
into physiology and therapeutics. Annu Rev Physiol. 72:247–272.
2010.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Evans RM and Mangelsdorf DJ: Nuclear
Receptors, RXR, and the Big Bang. Cell. 157:255–266.
2014.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Lim HW, Uhlenhaut NH, Rauch A, Weiner J,
Hübner S, Hübner N, Won KJ, Lazar MA, Tuckermann J and Steger DJ:
Genomic redistribution of GR monomers and dimers mediates
transcriptional response to exogenous glucocorticoid in vivo.
Genome Res. 25:836–844. 2015.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Holzer G, Markov GV and Laudet V:
Evolution of Nuclear Receptors and Ligand Signaling: Toward a Soft
Key-Lock Model? Curr Top Dev Biol. 125:1–38. 2017.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Klinge CM: Steroid Hormone Receptors and
Signal Transduction Processes. In: Principles of Endocrinology and
Hormone Action. Belfiore A and LeRoith D (eds). Springer
International Publishing, Cham, pp187-232, 2018.
|
|
13
|
Bertrand S, Brunet FG, Escriva H,
Parmentier G, Laudet V and Robinson-Rechavi M: Evolutionary
genomics of nuclear receptors: From twenty-five ancestral genes to
derived endocrine systems. Mol Biol Evol. 21:1923–1937.
2004.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Markov GV and Laudet V: Origin and
evolution of the ligand-binding ability of nuclear receptors. Mol
Cell Endocrinol. 334:21–30. 2011.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Berman HM, Westbrook J, Feng Z, Gilliland
G, Bhat TN, Weissig H, Shindyalov IN and Bourne PE: The Protein
Data Bank. Nucleic Acids Res. 28:235–242. 2000.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Sobie EA: An introduction to MATLAB. Sci
Signal. 4(tr7)2011.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Papageorgiou L, Loukatou S, Sofia K,
Maroulis D and Vlachakis D: An updated evolutionary study of
Flaviviridae NS3 helicase and NS5 RNA-dependent RNA polymerase
reveals novel invariable motifs as potential pharmacological
targets. Mol Biosyst. 12:2080–2093. 2016.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Pearson WR: Selecting the Right
Similarity-Scoring Matrix. Curr Protoc Bioinformatics.
43:3.5.1–3.5.9. 2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Waterhouse AM, Procter JB, Martin DM,
Clamp M and Barton GJ: Jalview Version 2 - a multiple sequence
alignment editor and analysis workbench. Bioinformatics.
25:1189–1191. 2009.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Vilar S, Cozza G and Moro S: Medicinal
chemistry and the molecular operating environment (MOE):
Application of QSAR and molecular docking to drug discovery. Curr
Top Med Chem. 8:1555–1572. 2008.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Kufareva I and Abagyan R: Methods of
protein structure comparison. Methods Mol Biol. 857:231–257.
2012.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Aertgeerts K, Skene R, Yano J, Sang BC,
Zou H, Snell G, Jennings A, Iwamoto K, Habuka N, Hirokawa A, et al:
Structural analysis of the mechanism of inhibition and allosteric
activation of the kinase domain of HER2 protein. J Biol Chem.
286:18756–18765. 2011.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Papageorgiou L, Loukatou S, Koumandou VL,
Makałowski W, Megalooikonomou V, Vlachakis D and Kossida S:
Structural models for the design of novel antiviral agents against
Greek Goat Encephalitis. PeerJ. 2(e664)2014.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Papageorgiou L, Megalooikonomou V and
Vlachakis D: Genetic and structural study of DNA-directed RNA
polymerase II of Trypanosoma brucei, towards the designing of novel
antiparasitic agents. PeerJ. 5(e3061)2017.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Michener CD and Sokal RR: A quantitative
approach to a problem in classification. Evolution. 11:130–162.
1957. View Article : Google Scholar
|
|
26
|
Sneath PHA and Sokal RR: Unweighted pair
group method with arithmetic mean. In: Numerical Taxonomy. W.H.
Freeman, San Francisco, CA, pp230-234, 1973.
|
|
27
|
Pavlopoulos GA, Soldatos TG, Barbosa-Silva
A and Schneider R: A reference guide for tree analysis and
visualization. BioData Min. 3(1)2010.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Lu J, Xu G, Zhang S and Lu B: An effective
sequence-alignment-free superpositioning of pairwise or multiple
structures with missing data. Algorithms Mol Biol.
11(18)2016.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Leaché AD, Wagner P, Linkem CW, Böhme W,
Papenfuss TJ, Chong RA, Lavin BR, Bauer AM, Nielsen SV, Greenbaum
E, et al: A hybrid phylogenetic-phylogenomic approach for species
tree estimation in African Agama lizards with applications to
biogeography, character evolution, and diversification. Mol
Phylogenet Evol. 79:215–230. 2014.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Fouquier J, Rideout JR, Bolyen E, Chase J,
Shiffer A, McDonald D, Knight R, Caporaso JG and Kelley ST:
Ghost-tree: Creating hybrid-gene phylogenetic trees for diversity
analyses. Microbiome. 4(11)2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Stecher G, Liu L, Sanderford M, Peterson
D, Tamura K and Kumar S: MEGA-MD: Molecular evolutionary genetics
analysis software with mutational diagnosis of amino acid
variation. Bioinformatics. 30:1305–1307. 2014.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Mellor CL, Marchese Robinson RL, Benigni
R, Ebbrell D, Enoch SJ, Firman JW, Madden JC, Pawar G, Yang C and
Cronin MTD: Molecular fingerprint-derived similarity measures for
toxicological read-across: Recommendations for optimal use. Regul
Toxicol Pharmacol. 101:121–134. 2019.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Rácz A, Bajusz D and Héberger K: Life
beyond the Tanimoto coefficient: Similarity measures for
interaction fingerprints. J Cheminform. 10(48)2018.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Bajusz D, Rácz A and Héberger K: Why is
Tanimoto index an appropriate choice for fingerprint-based
similarity calculations? J Cheminform. 7(20)2015.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Ferraz-de-Souza B, Lin L and Achermann JC:
Steroidogenic factor-1 (SF-1, NR5A1) and human disease. Mol Cell
Endocrinol. 336:198–205. 2011.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Fayard E, Auwerx J and Schoonjans K:
LRH-1: An orphan nuclear receptor involved in development,
metabolism and steroidogenesis. Trends Cell Biol. 14:250–260.
2004.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Hall BL and Thummel CS: The RXR homolog
ultraspiracle is an essential component of the Drosophila
ecdysone receptor. Development. 125:4709–4717. 1998.PubMed/NCBI
|
|
38
|
Hill RJ, Billas IM, Bonneton F, Graham LD
and Lawrence MC: Ecdysone receptors: From the Ashburner model to
structural biology. Annu Rev Entomol. 58:251–271. 2013.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Laffitte BA, Kast HR, Nguyen CM, Zavacki
AM, Moore DD and Edwards PA: Identification of the DNA binding
specificity and potential target genes for the farnesoid
X-activated receptor. J Biol Chem. 275:10638–10647. 2000.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Greschik H, Flaig R and Moras D:
Ligand/cofactor complexes of nuclear receptor ligand-binding
domains. https://www.academia.edu/18571843/Ligand_cofactor_complexes_of_nuclear_receptor_ligand-binding_domains.
|
|
41
|
Marchler-Bauer A, Derbyshire MK, Gonzales
NR, Lu S, Chitsaz F, Geer LY, Geer RC, He J, Gwadz M, Hurwitz DI,
et al: CDD: NCBI's conserved domain database. Nucleic Acids Res.
43:D222–D226. 2015.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Nicolaides NC, Roberts ML, Kino T,
Braatvedt G, Hurt DE, Katsantoni E, Sertedaki A, Chrousos GP and
Charmandari E: A novel point mutation of the human glucocorticoid
receptor gene causes primary generalized glucocorticoid resistance
through impaired interaction with the LXXLL motif of the p160
coactivators: Dissociation of the transactivating and
transreppressive activities. J Clin Endocrinol Metab. 99:E902–E907.
2014.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Jääskeläinen J, Mongan NP, Harland S and
Hughes IA: Five novel androgen receptor gene mutations associated
with complete androgen insensitivity syndrome. Hum Mutat.
27(291)2006.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Vitellius G, Fagart J, Delemer B, Amazit
L, Ramos N, Bouligand J, Le Billan F, Castinetti F, Guiochon-Mantel
A, Trabado S, et al: Three Novel Heterozygous Point Mutations of
NR3C1 Causing Glucocorticoid Resistance. Hum Mutat. 37:794–803.
2016.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Harrod A, Fulton J, Nguyen VTM, et al:
Genomic modelling of the ESR1 Y537S mutation for evaluating
function and new therapeutic approaches for metastatic breast
cancer. Oncogene. 36:2286–2296. 2017.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Achermann JC, Schwabe J, Fairall L and
Chatterjee K: Genetic disorders of nuclear receptors. J Clin
Invest. 127:1181–1192. 2017.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Ai N, Krasowski MD, Welsh WJ and Ekins S:
Understanding nuclear receptors using computational methods. Drug
Discov Today. 14:486–494. 2009.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Matias PM, Carrondo MA, Coelho R, Thomaz
M, Zhao XY, Wegg A, Crusius K, Egner U and Donner P: Structural
basis for the glucocorticoid response in a mutant human androgen
receptor (AR(ccr)) derived from an androgen-independent prostate
cancer. J Med Chem. 45:1439–1446. 2002.PubMed/NCBI View Article : Google Scholar
|
|
49
|
Puyang X, Furman C, Zheng GZ, Wu ZJ, Banka
D, Aithal K, Agoulnik S, Bolduc DM, Buonamici S, Caleb B, et al:
Discovery of Selective Estrogen Receptor Covalent Antagonists for
the Treatment of ERαWT and ERαMUT Breast
Cancer. Cancer Discov. 8:1176–1193. 2018.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Nettles KW, Bruning JB, Gil G, O'Neill EE,
Nowak J, Guo Y, Kim Y, DeSombre ER, Dilis R, Hanson RN, et al:
Structural plasticity in the oestrogen receptor ligand-binding
domain. EMBO Rep. 8:563–568. 2007.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Siltberg-Liberles J, Grahnen JA and
Liberles DA: The evolution of protein structures and structural
ensembles under functional constraint. Genes (Basel). 2:748–762.
2011.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Wurtz JM, Bourguet W, Renaud JP, Vivat V,
Chambon P, Moras D and Gronemeyer H: A canonical structure for the
ligand-binding domain of nuclear receptors. Nat Struct Biol.
3:87–94. 1996.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Kandil S, Biondaro S, Vlachakis D, Cummins
AC, Coluccia A, Berry C, Leyssen P, Neyts J and Brancale A:
Discovery of a novel HCV helicase inhibitor by a de novo drug
design approach. Bioorg Med Chem Lett. 19:2935–2937.
2009.PubMed/NCBI View Article : Google Scholar
|