Introduction
Bacterial biofilms consist of organized populations
of microorganisms embedded within a self-produced extracellular
polymeric matrix (EPS), enabling surface attachment and conferring
substantial resistance to antimicrobial agents and host immune
defense (1). As biofilm-embedded
cells are markedly more tolerant than their planktonic
counterparts, they play a central role in persistent, recurrent and
device-associated infections, rendering effective biofilm control a
major clinical priority (1,2).
Among these pathogens, Gram-negative bacteria characterized by
their lipopolysaccharide rich outer membrane and intrinsic
resistance mechanisms are particularly challenging (3). Notably, Escherichia coli
(E. coli) and Klebsiella pneumoniae (K.
pneumoniae) are key Gram-negative species commonly associated
with infections of the urinary tract, respiratory system,
bloodstream and indwelling medical devices, particularly among
hospitalized or immunocompromised individuals. Their strong ability
to adhere to host tissues and medical devices, combined with robust
biofilm formation, enables persistent colonization and
substantially hinders eradication with conventional antibiotic
therapies (4-6).
In the present study, E. coli and K.
pneumoniae were selected as representative Gram-negative
pathogens due to their clinical prevalence, strong biofilm-forming
capacity and reliance on well-characterized virulence and quorum
sensing (QS) regulated adhesion mechanisms. E. coli
encompasses diverse pathogenic pathotypes that employ a variety of
virulence factors, including fimbrial adhesins (such as type 1
fimbriae), curli fibers (CsgA fibrils), toxins, iron acquisition
systems and secretion systems, to adhere to host tissues and evade
immune defenses. Curli fibers, composed primarily of CsgA subunits,
are major components of the extracellular matrix and show a
critical role in surface adhesion, cell-cell aggregation, and
biofilm stability. Similarly, the E. coli common pilus
(ECP), assembled with the aid of the chaperone EcpB, contributes to
initial attachment and biofilm maturation on both biotic and
abiotic surfaces (7-10).
Biofilm development in E. coli progresses through stages of
initial adhesion, microcolony formation, maturation and dispersion,
and is tightly regulated by global transcriptional regulators and
QS systems. The QS network, involving signaling molecules, such as
autoinducer-2 and indole, coordinates the expression of genes
controlling curli and pili production, biofilm formation, motility
and antibiotic tolerance in a cell-density-dependent manner
(8).
K. pneumoniae is a capsulated Gram-negative
pathogen of major clinical concern, selected in the present study
due to its potent biofilm-forming ability, extensive virulence
repertoire and global association with multidrug resistance (MDR)
(11). Among these, the type 3
fimbrial subunit MrkA serves as a critical structural component of
type 3 fimbriae, mediating attachment to host tissues and medical
devices and playing a central role in biofilm initiation,
maturation and stability. QS systems in K. pneumonia can
control the production of fimbriae, EPS and other
virulence-associated factors in response to AI accumulation. QS
regulators, such as SdiA coordinate fimbriae expression, biofilm
development, and signal molecule production, emphasizing the
importance of cell-cell communication in pathogenicity (12,13).
In addition, K. pneumoniae is a major reservoir of
antibiotic resistance determinants, most notably New Delhi
metallo-β-lactamase-1, which confers resistance to carbapenems and
other β-lactam antibiotics. The coexistence of robust biofilm
formation and MDR significantly complicates treatment, rendering
K. pneumoniae a critical threat in healthcare settings and a
driver of the global spread of MDR infections (14).
In the current clinical landscape, conventional
antibiotics frequently fail to eradicate biofilm-associated
infections due to restricted penetration through the EPS matrix,
the diminished metabolic activity of sessile cells, and the
persistence of highly tolerant subpopulations. In addition,
biofilm-embedded persister cells exhibit metabolic dormancy and
upregulated efflux mechanisms, further compromising therapeutic
efficacy in clinical conditions, such as catheter-associated
urinary tract infections (CAUTIs) and ventilator-associated
pneumonia (VAP) (7,15,16).
Collectively, the well-characterized biofilm architectures,
QS-regulated virulence systems, and MDR profiles of E. coli
and K. pneumoniae render them highly relevant models for
evaluating antibiofilm and anti-virulence strategies. These
challenges underscore an escalating clinical crisis and highlight
the urgent need for alternative therapeutic approaches that move
beyond conventional bactericidal paradigms, focusing instead on
disrupting biofilm formation and virulence regulation while
minimizing selective pressure for resistance development.
Medicinal plants represent a valuable source of
structurally diverse natural compounds that exhibit well-documented
antimicrobial, antibiofilm and anti-virulence properties. Notably,
these plant-derived molecules often exhibit fewer side-effects
compared to conventional antibiotics, rendering them promising
alternatives for managing resistant infections (17-19).
Plant-derived phytochemicals, including flavonoids, phenolic acids,
terpenoids, alkaloids and essential oils, have been widely reported
to interfere with QS signaling, inhibit biofilm formation and
suppress the expression of virulence factors in Gram-negative
pathogens, frequently at sub-minimum inhibitory concentration (MIC)
concentrations (20-22).
Among these, Ruellia tuberosa L. (R. tuberosa L) is a
medicinal plant traditionally recognized for its diverse
pharmacological properties, including anti-inflammatory,
antioxidant and antimicrobial activities. Previous antimicrobial
studies on R. tuberosa L. have largely reported modest
antibacterial activity, often requiring high extract concentrations
to achieve growth inhibition (23-26).
Amajida et al (26)
observed antibacterial effects against E. coli only at
concentrations as high as 500 mg/ml, indicating bactericidal
potency under conventional MIC-based screening conditions. The
discrepancy in potency may be attributed to differences in extract
preparation, bacterial strains tested, and the use of growth
inhibition endpoints rather than biofilm-specific or anti-virulence
assays (23-26).
In addition, the present study demonstrates significant antibiofilm
inhibition at substantially lower, sub-MIC levels, suggesting a
mechanism that selectively attenuates virulence without affecting
bacterial growth. This mechanistic divergence from conventional
bactericidal approaches reinforces the novelty of the present
findings.
Although several medicinal plants have been
evaluated for antimicrobial properties (20), studies that systematically
integrate sub-MIC antibiofilm evaluation, QS-related virulence
targeting, phytochemical profiling, and in silico
mechanistic prediction against clinically relevant E. coli
and K. pneumoniae biofilm-forming pathogens remain limited.
The novelty of the present study lies in demonstrating the
growth-neutral antibiofilm activity of R. tuberosa extract
at sub-MIC level against MDR E. coli and K.
pneumoniae, combined with gas chromatography-mass spectrometry
(GC-MS)-guided molecular docking to predict interactions with key
adhesion and virulence-associated proteins. This integrated
experimental computational approach provides mechanistic insight
into anti-virulence activity beyond conventional bactericidal
screening.
Materials and methods
Isolates and growth conditions
The present study included two MDR clinical
isolates: One E. coli isolate obtained from a urinary tract
infection and one K. pneumoniae isolate obtained from a
respiratory tract specimen. The bacterial isolates used in the
present study were provided as a laboratory gift from the
Department of Microbiology, Saveetha Dental College and Hospitals,
Chennai, India, at an earlier time and were subsequently
transferred to Kannur University, Kerala, for research purposes.
The isolates were obtained from the hospital and were transferred
by the laboratory authorities to the Kannur University laboratory
for research use. The present study did not involve direct patient
interaction, access to patient records, or collection of clinical
specimens. The isolates were handled strictly as microbiological
strains for in vitro experimental purposes. Upon receipt,
the isolates were reconfirmed in the laboratory based on standard
morphological characteristics prior to experimental use. As the
study involved only de-identified bacterial isolates and did not
include human participants or identifiable patient data, ethics
approval and informed patient consent were waived. The cultures
were maintained under aerobic conditions at 37˚C in brain heart
infusion (BHI) broth (HiMedia Laboratories). Prior to the
experimental assays, the isolates were sub-cultured to the
logarithmic phase to ensure optimal growth and viability. Isolates
identities were confirmed using the automated VITEK® 2
system (BioMérieux, Inc.) for precise microbial identification. The
results were further validated using the standardized
identification keys described in Bergey's Manual of Determinative
Bacteriology (27), along with the
assessment of characteristic colony morphology and growth patterns
on MacConkey agar.
Collection of plant material
Fresh leaves of R. tuberosa L. were collected
from the Velappanchavadi area, Chennai, Tamil Nadu, India. The
plant material (leaves) was authenticated by a qualified botanist
prior to further solvent extract processing.
Preparation and preliminary screening
of R. tuberosa L
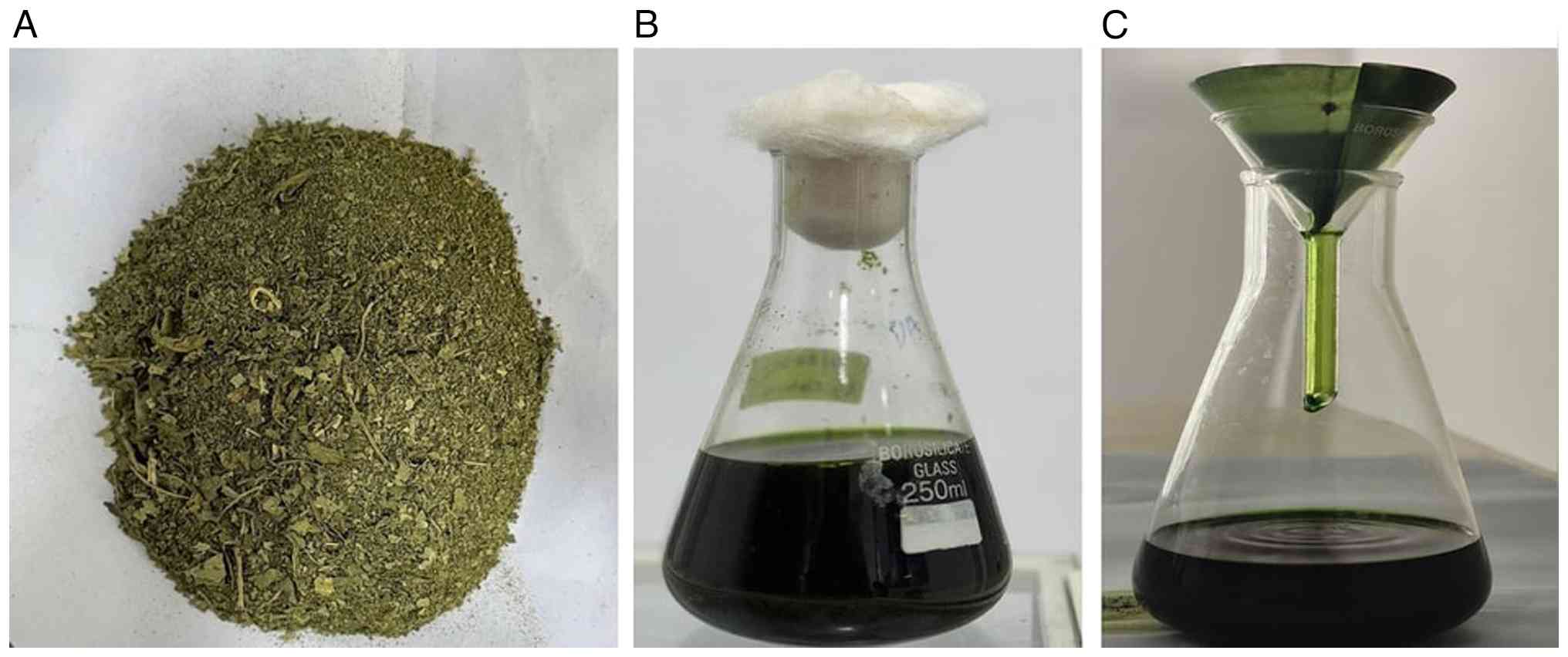
The harvested leaves were rinsed three times with
distilled water to eliminate adhering debris and surface
impurities. Following surface sterilization, the leaves were
shade-dried at room temperature and was then pulverized into a
coarse powder using a mechanical grinder. For extraction, 50 g of
the powdered leaves were soaked in 150 ml ethanol (Rankem
Laboratories, LLC, India) and incubated in a shaking incubator at
150 rpm and 37˚C for 4 days. The resulting extract was filtered
using Whatman filter paper (HiMedia Laboratories) after 4 days to
obtain a clear filtrate (Fig. 1).
The dried material was weighed and stored at 4˚C for subsequent
use.
Antibiogram profiling and
antimicrobial screening of R. tuberosa L
Antibiogram profiling was followed by the disk
diffusion method as per the Clinical and Laboratory Standards
Institute (CLSI) 2022 guidelines (28). A 1.5x108 CFU/ml (0.5
McFarland standard) inoculum was uniformly spread across
Mueller-Hinton agar (MHA) plates (HiMedia Laboratories) with a
sterile swab, antibiotic discs placed following brief drying, and
plates incubated at 37˚C for a day. The antibiotics evaluated
included ampicillin, ceftriaxone, ciprofloxacin, gentamicin,
imipenem and colistin (HiMedia Laboratories Private Limited). Zone
diameters were measured post-incubation and categorized as
susceptible or resistant using CLSI breakpoints.
Additionally, the antibacterial potential of R.
tuberosa L. extract was assessed via the agar well diffusion
assay, following established protocols (29). Standardized suspensions of E.
coli and K. pneumoniae, adjusted to a final density of
1.5x108 CFU/ml (0.5 McFarland standard), were uniformly
swabbed onto MHA plates. An 8-mm well was aseptically created in
the agar using a sterile cork borer, into which 40 µl of the
extract (dissolved in DMSO at 50 mg/ml) was pipetted. Pure DMSO
(Rankem Laboratories, LLC) served as the negative control. The
plates were incubated at 37˚C for 24 h, following which the
diameters of the inhibition zones were accurately measured in mm
using a vernier calliper to assess antibacterial activity.
Broth microdilution method
The MIC of R. tuberosa L. extract was
determined using a 2-fold broth microdilution method, as previously
described (30). Briefly,
concentrations ranging from 40-0.078 mg/ml were tested by
inoculating standardized suspensions of E. coli and K.
pneumoniae (1.5x108 CFU/ml) into BHI broth
containing R. tuberosa L. extract. The mixtures were
incubated at 37˚C for 24 h. Post-incubation,
2,3,5-triphenyltetrazolium chloride (TTC) (HiMedia Laboratories
Private Limited) was added to each well to detect viable cells
through color development. The lowest concentration exhibiting no
color change was recorded as the MIC. Based on the MIC values
obtained, subsequent biofilm inhibition assays were conducted using
sub-MIC concentrations derived from these preliminary
determinations to specifically assess growth-independent
antibiofilm activity.
Biofilm quantification assay
The impact of R. tuberosa L. extract on
biofilm formation by E. coli and K. pneumoniae was
assessed using a previously described crystal violet (CV) staining
method with slight modifications (31). Briefly, microtiter plates
containing 180 µl fresh BHI medium were inoculated with 20 µl of
overnight culture of the test organisms. Subsequently, R.
tuberosa L. extract was added at sub-MIC concentrations (E.
coli: 10 to 0.019 mg/ml; K. pneumoniae: 20 to 0.039
mg/ml), and the plates were incubated at 37˚C for 48 h. Adherent
biofilms were then stained with 0.1% CV solution (HiMedia
Laboratories) for 15 min at room temperature, followed by washing
with phosphate-buffered saline (PBS) (HiMedia Laboratories Private
Limited) to remove planktonic cells. Bound CV was eluted with 200
µl of 70% ethanol following a 5-min incubation at 37˚C, and the
absorbance was measured at 525 nm using a UV-Vis spectrophotometer
(JASCO).
Growth kinetics analysis of E. coli
and K. pneumoniae
The growth kinetics of E. coli and K.
pneumoniae were evaluated in the presence of R. tuberosa
L. extract at sub-MIC concentration. For E. coli, bacterial
growth was assessed at a sub-MIC of 10 mg/ml, whereas for K.
pneumoniae, growth kinetics were evaluated at a sub-MIC of 20
mg/ml. Overnight cultures were supplemented with the extract and
incubated at 37˚C for 24 h. Bacterial cell density was monitored
hourly by measuring the optical density (OD600) using a
spectrophotometer (JASCO).
GC-MS profiling of R. tuberosa L.
extract
For GC-MS analysis, the dried ethanolic extract of
R. tuberosa L. was reconstituted in n-hexane at a final
concentration of 1 mg/ml. The analysis was performed according to a
previously described protocol, with minor modifications adapted for
the present study (32). An
aliquot (1 µl) was injected in split mode (100:1) using an AOC-5000
autosampler (Shimadzu Corporation). Analysis was performed on a
Shimadzu GCMS-QP2010 Plus system equipped with an RTx-5MS capillary
column (cross-bond diphenyl dimethyl polysiloxane-fused silica)
(Shimadzu Corporation). Ultra-high-purity helium (≥99.99%) served
as the carrier gas at a constant flow rate of 0.8 ml/min. Compound
identification was achieved by comparison of mass spectra with the
NIST/WILEY7 library (John Wiley & Sons), and only compounds
with a match quality ≥80% were accepted. GC-MS was conducted for
qualitative phytochemical profiling only; individual compounds were
not isolated for in-vitro testing. Biological assays were
performed using the crude extract, and GC-MS results were used
solely to guide compound selection for in silico docking
analysis.
Computational analysis of compounds
from R. tuberosa L. extract
In silico molecular docking analysis was
conducted on compounds identified from the R. tuberosa L.
ethanolic extract. GC-MS profiling revealed multiple
phytochemicals, with diethyl phthalate [DEP, 42.56%; relative
abundance, retention time (RT) 14.98 min] as the most prominent
constituent. DEP was selected for docking analysis based on its
relative abundance in the extract, serving as a representative
compound for preliminary assessment of potential protein-ligand
interactions.
Docking-based interaction
analysis
DEP (C12H14O4;
PubChem CID: 6781) has a molecular weight of 222.24 g/mol and
consists of 12 carbon atoms, 14 hydrogen atoms and 4 oxygen atoms.
DEP was detected in the GC-MS chromatogram through its prominent
high-intensity peak, exhibiting the highest relative abundance
based on peak area. Its chemical structure was retrieved and
verified using PubChem database information (NCBI, NIH).
Structure-based molecular docking and
binding interaction analysis
For molecular docking analysis, only the
biologically relevant monomeric chain (Chain A) of each protein was
retained, while all other chains, crystallographic water molecules,
and co-crystallized ligands were removed to prevent potential
interference during binding simulations. The present study targeted
key proteins implicated in biofilm formation and virulence in E.
coli and K. pneumoniae. The selected structures included
8ENQ (cryo-EM structure of E. coli CsgA fibril), 5DFK
(crystal structure of E. coli common pilus chaperone EcpB),
8FFK (apo structure of the K. pneumoniae AcrB multidrug
efflux pump), and 9HW9 (solution NMR structure of the type 3
fimbrial subunit MrkA from K. pneumoniae). Three-dimensional
coordinates of these proteins were retrieved from the RCSB Protein
Data Bank (PDB). Protein preparation involved the removal of
H2O molecules, non-essential chains, and bound ligands,
followed by the addition of polar hydrogens and assignment of
appropriate atomic charges using BIOVIA Discovery Studio Visualizer
2024 (v24.1.0.23298, Dassault Systèmes Biovia Corp.).
Computational docking analyses were conducted to
evaluate the interaction of DEP with the selected protein targets.
Docking simulations were performed using PyRx-Python Prescription
0.8 integrated with AutoDock Vina. The grid box center coordinates
and dimensions were defined to encompass the predicted active or
binding regions of each protein, and the detailed parameters used
for each target are provided in Table
SI to ensure computational reproducibility. Multiple ligand
conformations were generated, and the most favorable docking pose
for each protein-ligand complex was selected based on the lowest
(most negative) binding energy values. The final binding modes and
three-dimensional interaction patterns were visualized and analyzed
using BIOVIA Discovery Studio Visualizer 2024.
Statistical analysis
All assays were performed in triplicate, and data
are presented as the mean ± SD. Statistical analysis was performed
using one-way ANOVA followed by Tukey's post hoc test in GraphPad
Prism (v5.03; Dotmatics) and was used to analyze biofilm and growth
curve data. A value of P<0.05 was considered to indicate a
statistically significant difference compared to the controls.
Results
Identification of E. coli and K.
pneumoniae
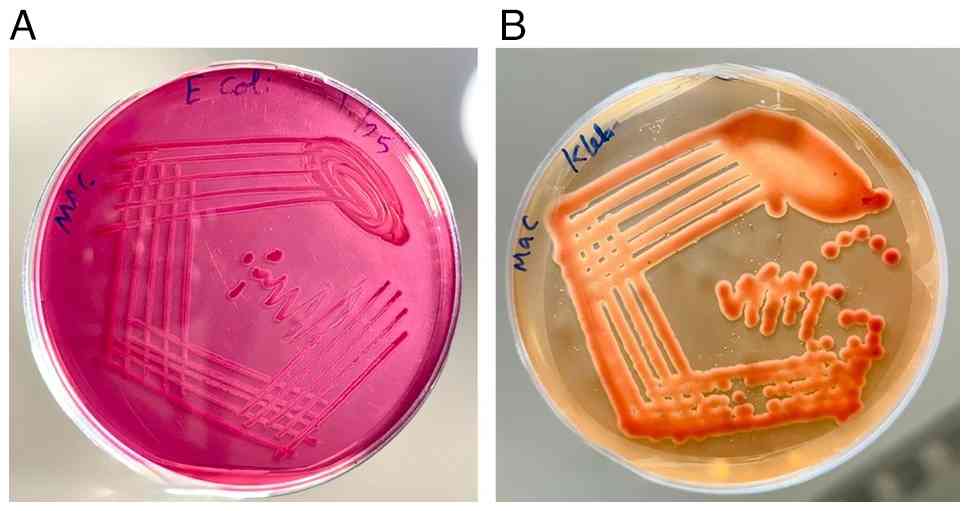
E. coli and K. pneumoniae were
accurately confirmed via the automated VITEK® 2 system
(https://www.biomerieux.com).
Complementary validation was achieved through distinctive colony
morphology and growth characteristics on MacConkey agar (Fig. 2).
Antibiogram and antimicrobial
analysis
Antibiogram analysis adhered to CLSI guidelines.
Both E. coli and K. pneumonia isolates exhibited
substantial resistance to most tested antibiotics including
ampicillin, ceftriaxone, ciprofloxacin and gentamicin.
Susceptibility persisted primarily to last-line options imipenem
and colistin (Table I). The
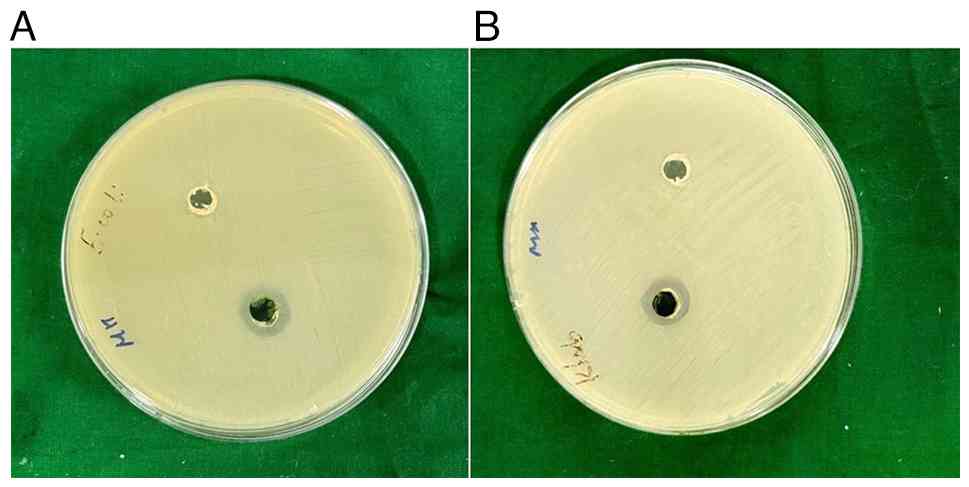
antimicrobial potential of R. tuberosa L. extract was
evaluated using the agar well diffusion method against E.
coli and K. pneumoniae. The results revealed a distinct
zone of inhibition measuring 14 mm and 13 mm, indicating a
significant level of antimicrobial efficacy against this pathogen
(Fig. 3). This suggests that R.
tuberosa L. extract possesses the capability to inhibit the
growth of E. coli and K. pneumoniae, which is
particularly relevant given the MDR nature of the pathogen.
 | Table IComparative antibiotic susceptibility
profile of clinical E. coli and K. pneumoniae
isolates determined by the Kirby-Bauer disk diffusion method
according to CLSI guidelines. |
Table I
Comparative antibiotic susceptibility
profile of clinical E. coli and K. pneumoniae
isolates determined by the Kirby-Bauer disk diffusion method
according to CLSI guidelines.
| Antibiotic | E. coli
(mean ± SD, mm) | Interpretation | K.
pneumoniae (mean ± SD, mm) | Interpretation |
|---|
| Ampicillin | No zone | Resistant | No zone | Resistant |
| Ceftriaxone | No zone | Resistant | 12±1.3 | Resistant |
| Ciprofloxacin | 14±1.2 | Resistant | 14±1.2 | Resistant |
| Gentamicin | 14±1.1 | Resistant | 14±0.5 | Resistant |
| Imipenem | 20±1.3 | Sensitive | 20±1.4 | Sensitive |
| Colistin | 21±1.2 | Sensitive | 22±1.3 | Sensitive |
MIC analysis of R. tuberosa L.
extract
Using a 2-fold serial broth dilution method, the MIC
of R. tuberosa L. extract was determined to be 20 mg/ml for
E. coli and 40 mg/ml for K. pneumoniae (Table II). Based on these results,
sub-MIC concentrations were subsequently selected and employed for
anti-biofilm assays.
 | Table IIMinimum inhibitory concentration of
R. tuberosa L. against E. coli and K. pneumoniae. |
Table II
Minimum inhibitory concentration of
R. tuberosa L. against E. coli and K. pneumoniae.
| | | Growth
measured |
|---|
| Serial no. | Two-fold dilution
concentration (mg/ml) | E. coli | K.
pneumoniae |
|---|
| Sl.1 | 40 | - | - |
| 2 | 20 | - | + |
| 3 | 10 | + | + |
| 4 | 5 | + | + |
| 5 | 2.5 | + | + |
| 6 | 1.25 | + | + |
| 7 | 0.625 | + | + |
| 8 | 0.312 | + | + |
| 9 | 0.156 | + | + |
| 10 | 0.078 | + | + |
Inhibition of biofilm formation by E.
coli and K. pneumoniae
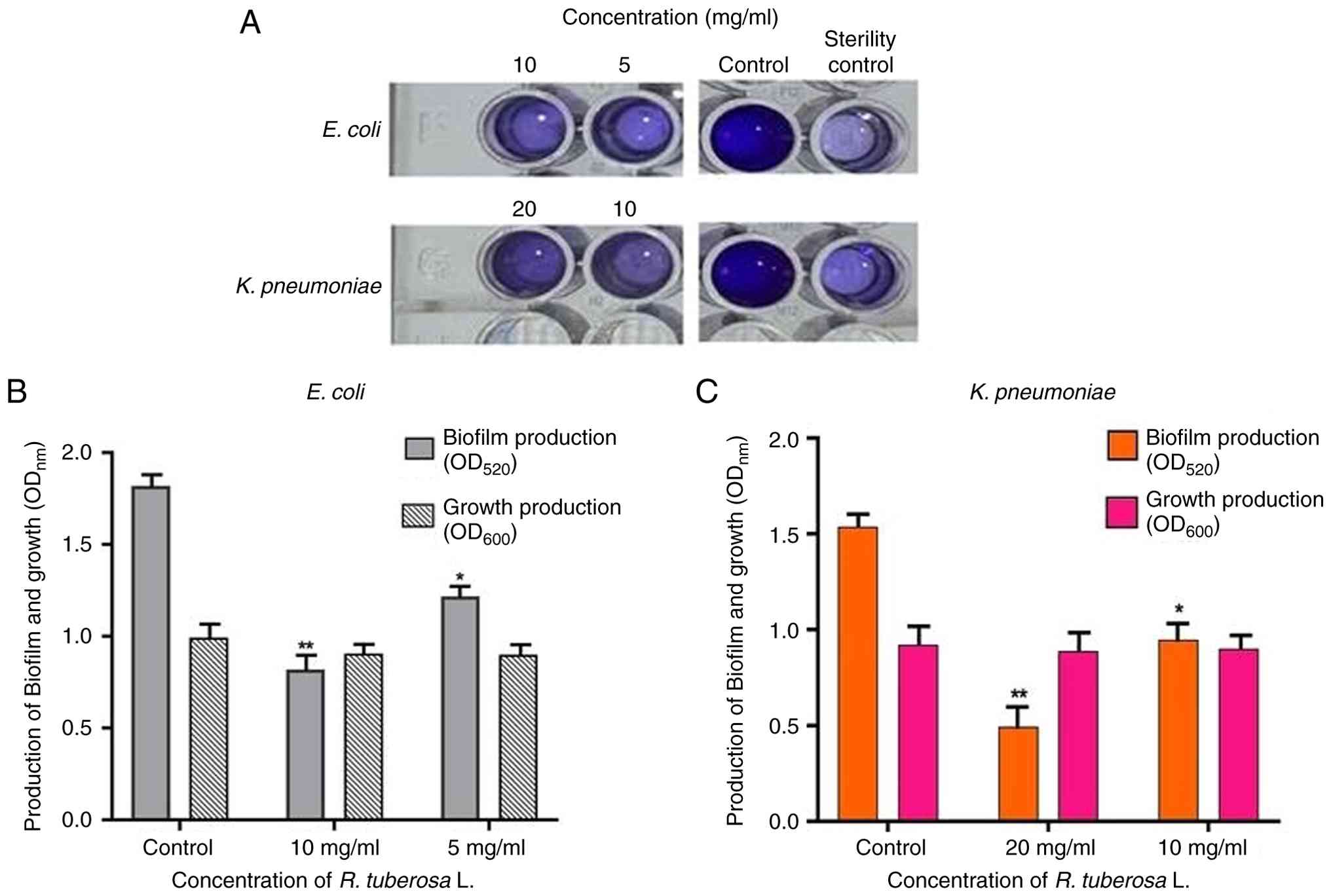
The CV staining biofilm assay demonstrated
significant inhibition by R. tuberosa L. extract (Fig. 4A). Spectrophotometric analysis at
520 nm (OD520) revealed 55 and 33% biofilm reduction in
E. coli at 10 and 5 mg/ml, respectively (Fig. 4B). For K. pneumoniae,
biofilm formation decreased by 67.81% at 20 mg/ml and 38.83% at 10
mg/ml (Fig. 4C). All observed
reductions were statistically significant compared to the control
(P<0.05). The statistical analysis of OD600 growth
measurements revealed no significant difference between the treated
and untreated groups at the tested sub-MIC concentrations,
confirming that the observed reductions in biofilm biomass were
independent of bacterial growth inhibition.
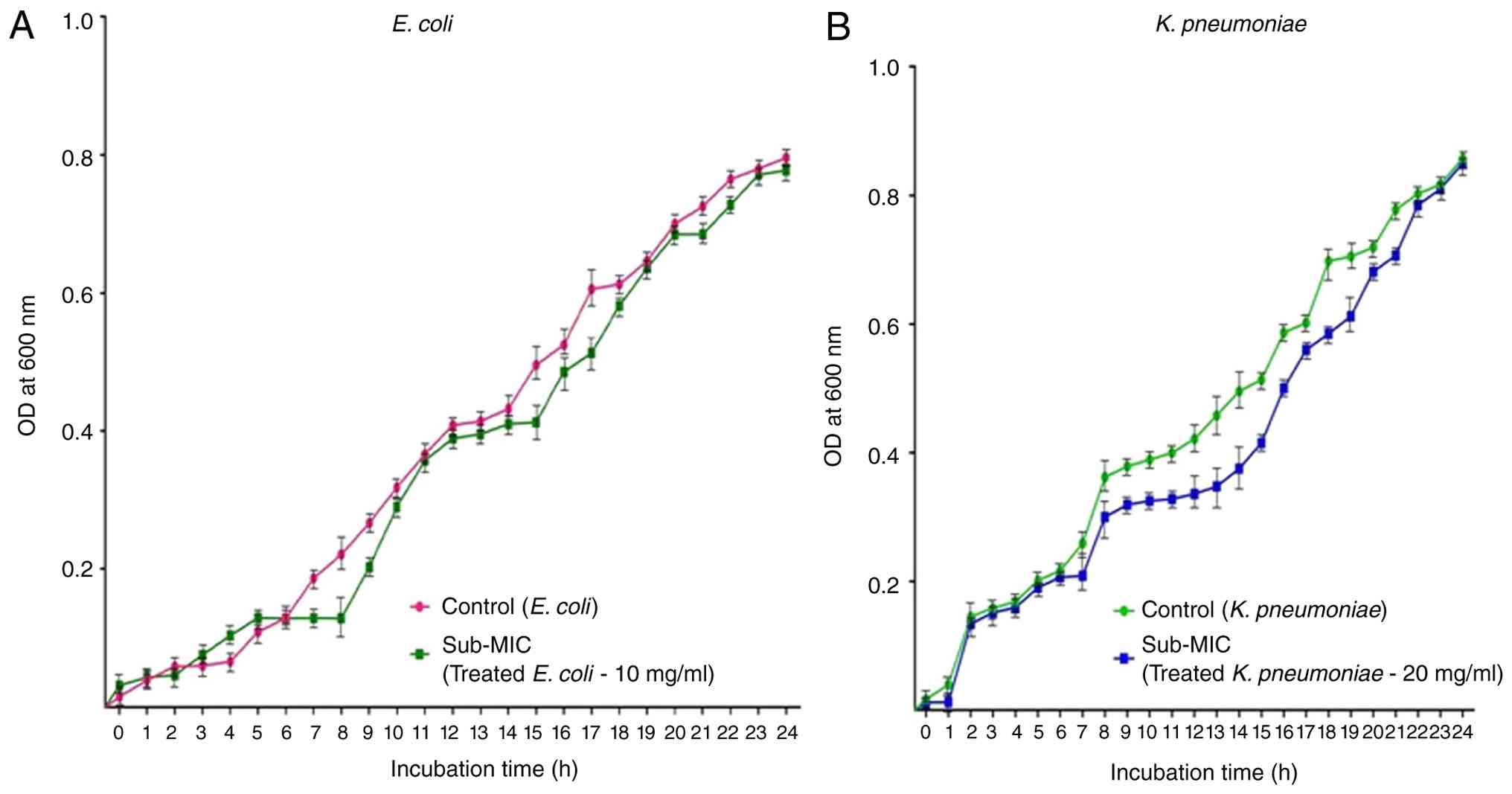
Growth kinetics analysis
The growth kinetics of E. coli and K.
pneumoniae exhibited comparable patterns upon exposure to the
R. tuberosa L. extract at sub-MIC levels. E. coli was
evaluated at a sub-MIC concentration of 10 mg/ml (Fig. 5A), whereas K. pneumoniae was
assessed at 20 mg/ml. Regardless of the observed differences in
concentrations, no marked difference in growth was observed between
the two species relative to the control (Fig. 5B).
GC-MS analysis of R. tuberosa L
GC-MS profiling of the ethanolic extract of R.
tuberosa L. identified >30 distinct chemical constituents
through matching with the NIST and Wiley spectral libraries
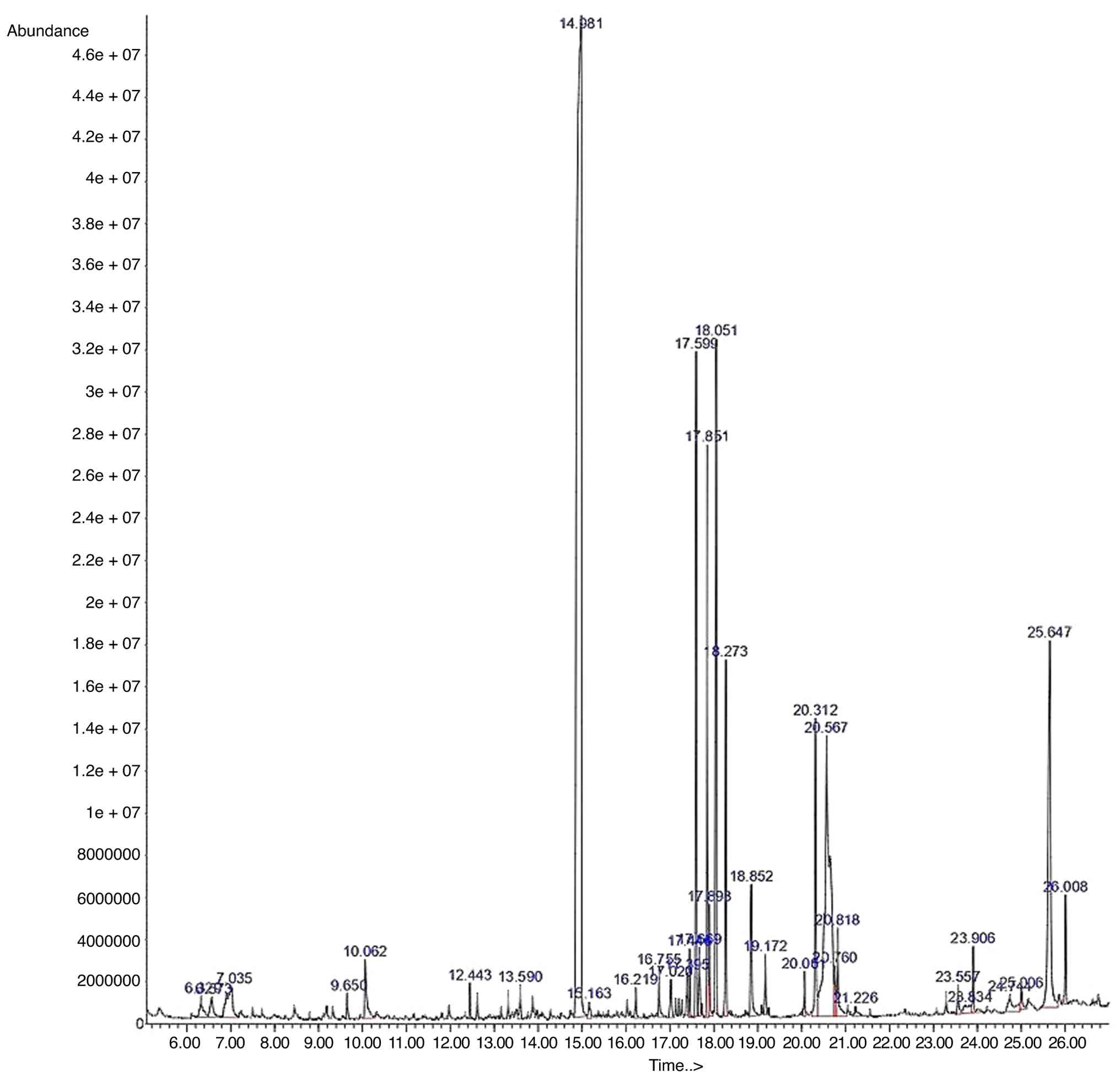
(Table III). The chromatograms
displayed multiple compounds with distinct RT, featuring diethyl
phthalate (DEP; 42.56%; RT, 14.98 min) as the predominant
components (Fig. 6). This rich
phytochemical diversity underscores the therapeutic promise of
R. tuberosa L., with DEP selected for further molecular
docking analysis due to its highest abundance. The minor baseline
drift observed is due to solvent background and column
characteristics and did not affect compound identification, as no
well-defined peaks in this region satisfied the library match
quality criteria.
 | Table IIIGC-MS identification of compounds in
Ruellia tuberosa L. with corresponding retention times and
area percentage. |
Table III
GC-MS identification of compounds in
Ruellia tuberosa L. with corresponding retention times and
area percentage.
| Serial no. and
peak | Name of the
compounds | Molecular
formula | Retention time | Percentage |
|---|
| 1 | Benzyl
chloride |
C7H7Cl | 6.9116 | 0.56 |
| 2 |
2,5-Hexanedione |
C6H10O2 | 7.0354 | 1.08 |
| 3 | Dodecane |
C12H26 | 9.6500 | 0.33 |
| 4 |
5-Hydroxymethylfurfural |
C6H6O3 | 10.0615 | 1.00 |
| 5 | Tetradecane |
C14H30 | 12.4430 | 0.32 |
| 6 |
1,1,6-Trimethyl-1,2-dihydronaphthalene |
C13H16 | 12.6142 | 0.24 |
| 7 |
5-Methylene-4,5,6,6a-tetrahydro-3ah-pentalen-1-one |
C9H10O | 13.3170 | 0.25 |
| 8 | Phorone |
C9H14O | 13.5901 | 0.33 |
| 9 |
1,4-Dihydrothujopsene-(I1) |
C15H26 | 13.8705 | 0.25 |
| 10 | Diethyl
phthalate |
C12H14O4 | 14.9811 | 42.56 |
| 11 | Terpinyl
propionate |
C13H22O2 | 16.2193 | 0.32 |
| 12 | Tetradecanoic
acid |
C14H28O2 | 16.7546 | 0.51 |
| 13 | Loliolide |
C11H16O3 | 17.0204 | 0.54 |
| 14 |
Beta-homocyclocitral |
C11H18O | 17.3955 | 0.52 |
| 15 | Isopropyl
myristate |
C17H34O2 | 17.4464 | 0.56 |
| 16 | Erucic acid |
C22H42O2 | 17.5994 | 6.29 |
| 17 |
3,7,11,15-Tetramethyl-2-hexadecen-1-ol |
C20H40O | 17.6686 | 0.84 |
| 18 | Linolenic acid |
C18H30O2 | 17.8560 | 5.22 |
| 19 | Isooctanol |
C8H180 | 17.8980 | 0.92 |
| 20 | 2-Hexadecen-1-ol,
3,7,11,15-tetramethyl-, acetate, [R [R*,R*-(E)]] | C22H42O2 | 18.0510 | 7.77 |
| 21 | Cholest-5-ene, 3
bromo-, (3á)- | C27H45Br | 18.2730 | 2.98 |
| 22 | n-Hexadecanoic
acid |
C16H32O2 | 18.8521 | 1.79 |
| 23 | Hexadecanoic acid,
ethyl ester |
C18H36O2 | 19.1725 | 0.65 |
| 24 | 2H-Pyran-2-one |
C5H4O2 | 20.0610 | 0.43 |
| 25 | Phytol |
C20H40O | 20.3123 | 3.16 |
| 26 |
13Z,16Z-docosadienoic acid |
C22H40O2 | 20.5672 | 5.96 |
| 27 | Octacosanol |
C28H58O | 20.6437 | 3.59 |
| 28 | Linoleic acid ethyl
ester |
C20H36O2 | 20.7602 | 0.34 |
| 29 |
Dihomo-gamma-linolenic acid |
C20H34O2 | 20.8185 | 0.64 |
| 30 |
1-O-hexadecyl-2-(9Z-octadecenoyl)-sn-glycerol |
C37H72O4 | 23.2874 | 0.23 |
| 31 | Glycerol
1-palmitate |
C19H38O4 | 23.5569 | 0.31 |
| 32 | Bis(2-ethylhexyl)
phthalate |
C24H38O4 | 23.9064 | 0.53 |
| 33 | Octacosanol |
C28H58O | 25.6471 | 8.04 |
| 34 | Squalene |
C30H50 | 26.0076 | 0.94 |
In silico analysis of DEP interactions
with E. coli and K. pneumoniae regulators
DEP was selected for molecular docking based on its
highest relative abundance in the GC-MS profile, allowing
assessment of one of the major constituents of the extract.
Although other compounds such as erucic acid, phytol and
octacosanol were also identified and may possess biological
activity, the present analysis focused on DEP as a representative
constituent for preliminary mechanistic evaluation. Docking
simulations were performed to evaluate the binding interactions of
DEP with key regulatory and virulence-associated proteins of E.
coli and K. pneumoniae. The selected targets included
8ENQ (cryo-EM structure of E. coli CsgA fibril), 5DFK
(crystal structure of E. coli common pilus chaperone EcpB),
8FFK (apo form of the K. pneumoniae AcrB multidrug efflux
pump), and 9HW9 (solution NMR structure of the type 3 fimbrial
subunit MrkA from K. pneumoniae). DEP exhibited binding
affinities of -4.00 kcal/mol and -5.2 kcal/mol toward the E.
coli targets 8ENQ and 5DFK (Table
IV), respectively. In K. pneumoniae, DEP showed binding
energies of -5.4 kcal/mol for 8FFK and -4.6 kcal/mol for 9HW9
(Table V).
 | Table IVMolecular docking results of DEP with
E. coli targets 8ENQ and 5DFK. |
Table IV
Molecular docking results of DEP with
E. coli targets 8ENQ and 5DFK.
| Binding of DEP with
E. coli targets 8ENQ and 5DFK | Binding
affinity | rmsd/ub | rmsd/lb |
|---|
| 8ENQ_6781_
E=211.67 | -4 | 0 | 0 |
|
8ENQ_6781_E=211.67 | -3.9 | 31.488 | 28.751 |
|
8ENQ_6781_E=211.67 | -3.9 | 34.752 | 32.937 |
| 8ENQ_6781_
E=211.67 | -3.9 | 31.964 | 30.064 |
| 8ENQ_6781_
E=211.67 | -3.8 | 32.266 | 29.807 |
| 8ENQ_6781_
E=211.67 | -3.8 | 31.754 | 29.339 |
| 8ENQ_6781_
E=211.67 | -3.8 | 30.881 | 29.36 |
| 8ENQ_6781_
E=211.67 | -3.7 | 34.065 | 32.006 |
| 8ENQ_6781_
E=211.67 | -3.7 | 34.363 | 31.683 |
|
5DFK_6781_E=211.67 | -5.2 | 0 | 0 |
| 5DFK_6781_
E=211.67 | -5.2 | 3.542 | 1.13 |
|
5DFK_6781_E=211.67 | -5.1 | 11.338 | 9.815 |
|
5DFK_6781_E=211.67 | -5.1 | 3.63 | 1.379 |
|
5DFK_6781_E=211.67 | -5.1 | 11.271 | 9.789 |
| 5DFK_6781_
E=211.67 | -4.9 | 2.342 | 1.534 |
| 5DFK_6781_
E=211.67 | -4.8 | 14.807 | 12.768 |
| 5DFK_6781_
E=211.67 | -4.7 | 12.057 | 10.719 |
| 5DFK_6781
_E=211.67 | -4.6 | 11.588 | 10.441 |
 | Table VMolecular docking analysis of DEP
with K. pneumoniae targets 8FFK and 9HW9. |
Table V
Molecular docking analysis of DEP
with K. pneumoniae targets 8FFK and 9HW9.
| Binding of DEP with
K. pneumoniae targets 8FFK and 9HW9 | Binding
affinity | rmsd/ub | rmsd/lb |
|---|
|
8FFK_6781_E=211.67 | -5.4 | 0 | 0 |
|
8FFK_6781_E=211.67 | -5.4 | 4.241 | 0.071 |
|
8FFK_6781_E=211.67 | -5.4 | 19.269 | 17.333 |
|
8FFK_6781_E=211.67 | -5.3 | 19.879 | 17.537 |
|
8FFK_6781_E=211.67 | -5.3 | 3.664 | 1.197 |
|
8FFK_6781_E=211.67 | -5.3 | 2.101 | 1.282 |
|
8FFK_6781_E=211.67 | -5.2 | 19.48 | 17.171 |
|
8FFK_6781_E=211.67 | -5.2 | 20.076 | 17.843 |
|
8FFK_6781_E=211.67 | -5.1 | 30.794 | 28.043 |
| 9HW9_
6781_E=211.67 | -4.6 | 0 | 0 |
| 9HW9_
6781_E=211.67 | -4.5 | 4.513 | 2.415 |
| 9HW9_
6781_E=211.67 | -4.4 | 21.575 | 19.773 |
| 9HW9_
6781_E=211.67 | -4.3 | 8.499 | 6.76 |
| 9HW9_
6781_E=211.67 | -4.3 | 20.282 | 18.199 |
| 9HW9_
6781_E=211.67 | -4.3 | 20.033 | 18.223 |
| 9HW9_
6781_E=211.67 | -4.3 | 21.633 | 19.821 |
| 9HW9_
6781_E=211.67 | -4.3 | 21.658 | 19.916 |
| 9HW9_
6781_E=211.67 | -4.1 | 15.602 | 13.318 |
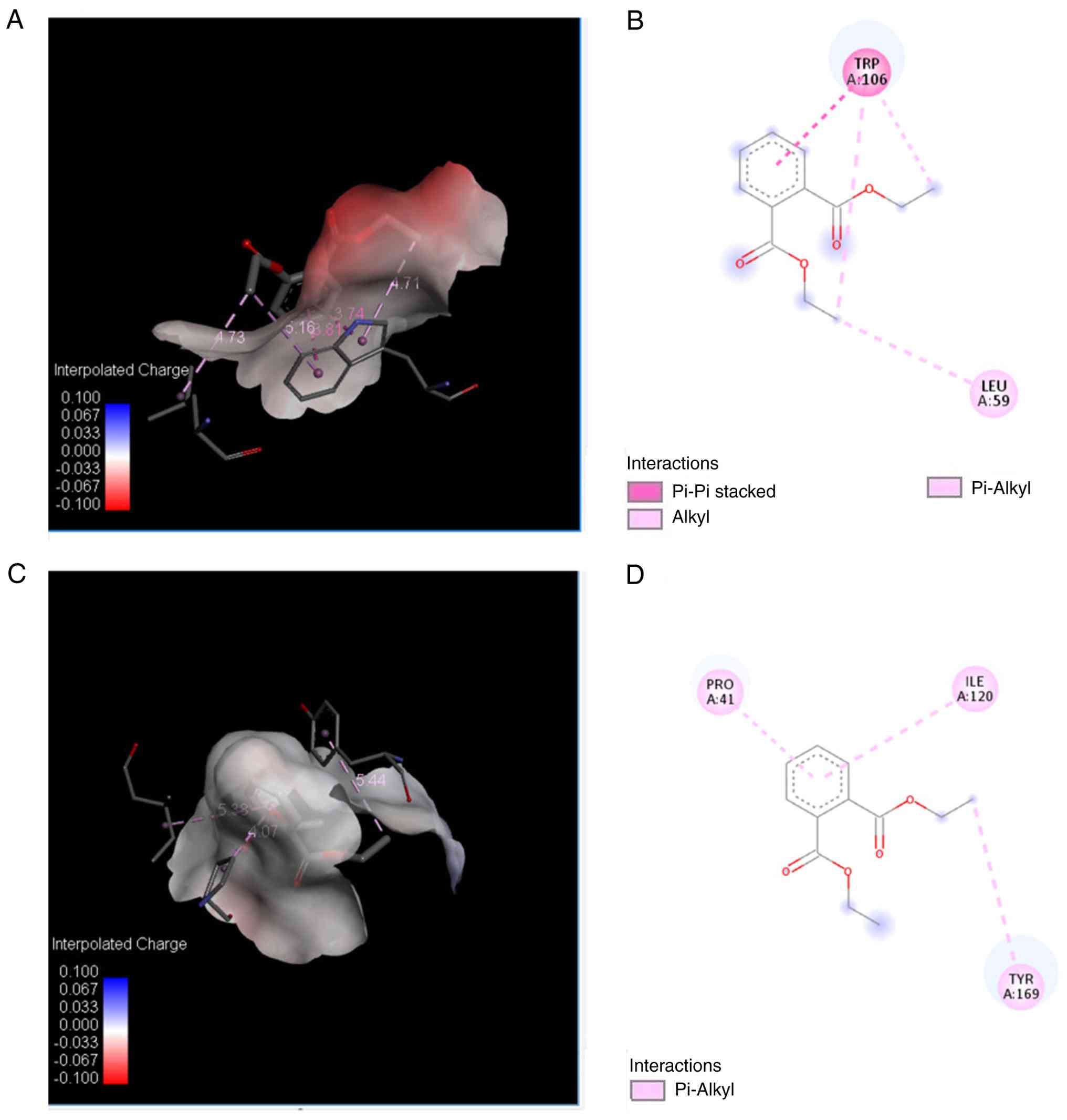
Interaction analysis revealed that DEP binding to
8ENQ involved π-π stacking with TRP A:106, along with alkyl and
π-alkyl interactions with LEU A:59 and TRP A:106. For the 5DFK
protein, DEP formed π-alkyl interactions with PRO A:41, ILE A:120
and TYR A:169 (Fig. 7). In the
case of 8FFK, DEP exhibited π-π stacking with PHE A:616 and
additional alkyl and π-alkyl interactions with LEU A:572, PHE
A:665, ILE A:625, and MET A:574. Notably, DEP formed three
conventional hydrogen bonds with 9HW9 involving GLN A:107 (3.01 Å),
SER A:154 (2.39 Å) and GLN A:104 (2.62 Å), with bond lengths
falling within the typical hydrogen bond distance range.
Additionally, a carbon-hydrogen bond was observed with ASN A:105,
along with π-alkyl and alkyl interactions involving LEU A:84 and
TYR A:94 (Fig. 8). Although the
binding energies obtained for DEP (-4.0 to -5.4 kcal/mol) are
modest compared to high-affinity drug-target interactions, they
fall within the range commonly reported for small, non-drug-like
natural compounds involved in anti-virulence and antibiofilm
modulation. Similar docking scores have been reported for
phytochemicals that exert biologically meaningful effects through
multi-target, low-affinity interactions rather than single
high-affinity inhibition (22,30).
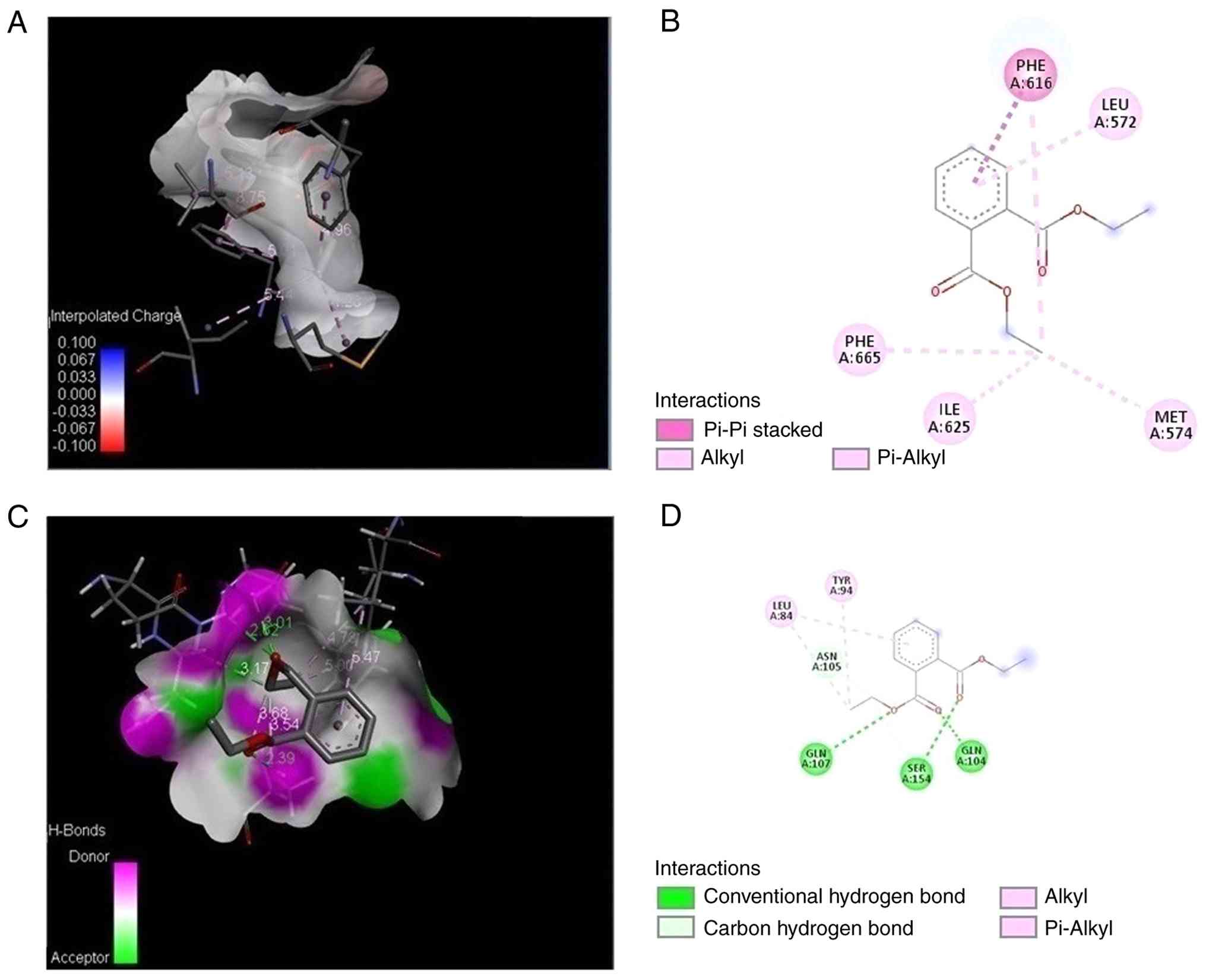 | Figure 8Molecular docking and interaction
analysis of K. pneumoniae proteins. (A and C) 3D binding
conformations of DEP within the active sites of 8FFK and 9HW9,
shown as surface representations; (B and D) 2D interaction diagrams
highlighting key stabilizing interactions. In the 2D maps, green
dashed lines represent conventional hydrogen bonds, light green
dashed lines indicate carbon-hydrogen bonds, pink lines denote
alkyl interactions, light pink dashed lines indicate π-alkyl
interactions, and dark pink dashed lines represent π-π stacking
interactions. DEP binding to 8FFK is characterized by π-π stacking
with PHE A:616 and alkyl/π-alkyl interactions involving LEU A:572,
PHE A:665, ILE A:625, and MET A:574. By contrast, DEP interaction
with 9HW9 involves three conventional hydrogen bonds with GLN
A:107, SER A:154, and GLN A:104, a carbon-hydrogen bond with ASN
A:105, and additional alkyl/π-alkyl interactions with LEU A:84 and
TYR A:94. |
Discussion
The ability of E. coli and K.
pneumoniae to form robust biofilms is a major contributor to
their persistence, virulence and MDR in clinical settings.
Biofilm-associated infections are notoriously difficult to
eradicate, as embedded cells exhibit enhanced tolerance to
antibiotics and host immune responses (33). Therefore, identifying natural
agents that disrupt biofilm formation without exerting strong
bactericidal pressure is a critical strategy to mitigate resistance
development.
In the present study, preliminary antibiogram
analysis revealed that the clinical isolate of E. coli and
K. pneumoniae exhibited resistance to the majority of the
tested antibiotics. Owing to this MDR profile, the present study
subsequently evaluated the antimicrobial potential of the ethanolic
leaf extract of R. tuberosa L. The extract demonstrated
appreciable antibacterial activity against E. coli and K.
pneumoniae, producing zones of inhibition of 14 and 13 mm,
respectively, at a concentration of 50 mg/ml in the agar well
diffusion assay. These findings align closely with values
documented in prior studies for R. tuberosa ethanolic or
n-hexane extracts, which typically produced inhibition zones of
7-16 mm only at the highest tested concentrations (26); notably, in the present study,
similar inhibitory effects were achieved at substantially lower
concentrations against E. coli and K. pneumoniae.
Similarly, Tinospora cordifolia has been reported as an
effective source for the biogenic synthesis of ZnO nanoparticles
exhibiting notable antimicrobial activity (34). These findings are consistent with
the results of the present study and further support the potential
of plant-derived systems for antimicrobial applications at low
concentrations.
MIC analysis further highlighted the potency of the
extract, with MIC values of 20 mg/ml for E. coli and 40
mg/ml for K. pneumoniae. These concentrations are markedly
lower than those reported in earlier studies, such as in the study
by Amajida et al (26), who
observed inhibition of E. coli and Bacillus subtilis
only at 500 mg/ml. The lower MIC values observed in the present
study may reflect differences in phytochemical composition,
extraction efficiency and strain-specific susceptibility. In
addition, methodological variations including solvent type,
extraction duration, plant source variability, and growth
conditions employed during antimicrobial assays may also contribute
to the discrepancies observed when compared with previous reports,
including the study by Amajida et al (26).
Moreover, a key strength of the present study lies
in the sub-MIC antibiofilm evaluation, which revealed substantial
inhibition of biofilm formation without affecting planktonic
growth. At the sub-MIC level, the extract inhibited biofilm
formation by 55% in E. coli (10 mg/ml) and 67.81% in K.
pneumoniae (20 mg/ml), while growth kinetics (OD600)
remained indistinguishable from the untreated controls. This growth
neutrality strongly indicates a true antibiofilm mechanism. Similar
antibiofilm effects have been reported in previous research for
several natural phenolic compounds, including epigallocatechin
gallate, octyl gallate, scutellarein, wedelolactone and
resveratrol, which reduce E. coli biofilm formation by
50-80% at sub-MIC concentrations through the suppression of curli
expression, motility and chemotactic responses (35). Additionally, a number of essential
oils and plant-derived agents (e.g., cinnamaldehyde and eugenol)
also impair membrane integrity or significantly alter growth and
motility, making it difficult to distinguish antibiofilm-specific
effects from general antibacterial stress (36-37).
By contrast, R. tuberosa extract achieved comparable biofilm
inhibition, while fully sparing planktonic growth, suggesting that
it may represent a promising anti-virulence candidate capable of
inhibiting biofilm formation without affecting planktonic growth.
Similar antibiofilm activities have been observed with other plant
extracts in different Gram-negative species, such as Rhamnus
frangula against Acinetobacter baumannii at 5 mg/ml
(38). Additionally, R.
officinalis and P. paniculata are known to impede
biofilm formation in K. pneumoniae by up to 58.41 and
55.71%, respectively (39). In
comparison with these reports, in the present study, R.
tuberosa extract exhibited comparable antibiofilm activity
against E. coli and K. pneumoniae, highlighting its
potential as an effective antibiofilm agent.
Several natural products, including Coffea
arabica L. extracts, promysalin, and specialized microbial
metabolites, are known to inhibit biofilm formation at sub-MIC or
nanomolar concentrations through defined molecular targets
(40-42).
While these compounds are highly potent, they are often
structurally complex (e.g., promysalin requiring multi-step total
synthesis), difficult to source (microbial fermentation-dependent),
or associated with specific metabolic targets (e.g., succinate
dehydrogenase) that may limit broader applicability. In contrast,
the activity of R. tuberosa L. extract appears to arise from
a multi-target, phytochemical-driven mechanism, which may reduce
the likelihood of resistance development, while enhancing
compatibility with existing antibiotics.
In the present study, the GC-MS profiling of the
R. tuberosa extract identified >30 compounds, with DEP
(42.56% relative abundance) as the predominant constituent. To
explore potential molecular interactions underlying the observed
antibiofilm activity, in silico docking analyses were
performed. For clarity, only the principal binding interactions and
key stabilizing residues are summarized. DEP exhibited stable
interactions with major biofilm and resistance-associated targets,
including the E. coli CsgA curli fibril (-4.0 kcal/mol,
π-stacking with TRP A:106), EcpB chaperone (-5.2 kcal/mol), K.
pneumoniae AcrB efflux pump (-5.4 kcal/mol, π-stacking with PHE
A:616) and MrkA fimbrial protein (-4.6 kcal/mol, hydrogen bonding
with GLN and SER residues). These findings suggest that DEP may
contribute to the observed bioactivity; however, the potential role
of other phytoconstituents and synergistic interactions cannot be
excluded. DEP may affect bacterial virulence and biofilm formation
through multiple mechanisms, including disruption of cell membrane
integrity, interference with QS, inhibition of adhesion and
fimbriae-associated proteins, and modulation of efflux-related
pathways. Previous studies indicate that DEP can exert antibiofilm
and anti-virulence effects (43,44).
Additionally, DEP has been reported to alter membrane permeability
and lipid organization, leading to impaired surface attachment and
early biofilm establishment (45,46).
In the present study, the observed docking interactions with
adhesion and fimbrial-related proteins further support a possible
role for DEP in disrupting bacterial attachment and biofilm
maturation. Moreover, interactions with efflux-associated proteins
may also contribute to reduced persistence within the biofilm
matrix. While these mechanisms are supported by in silico
predictions and previous reports, experimental validation will be
required to confirm the relative contribution of each pathway.
Further validation using advanced experimental approaches,
including in vitro and in vivo assays, is required to
substantiate the antibiofilm potential of DEP. It is important to
emphasize that molecular docking provides predictive insights into
potential binding interactions and does not constitute functional
evidence of target inhibition; therefore, the proposed mechanisms
remain hypothetical until validated through targeted biochemical,
genetic, or phenotypic assays.
Taken together, the present study advances beyond
descriptive antibiofilm observations by integrating quantitative
biofilm assays, growth-neutral validation, GC-MS chemical
profiling, and predictive molecular docking to propose a
mechanistic framework for anti-virulence activity against clinical
MDR pathogens. The findings indicate that R. tuberosa L.
extract, particularly its major constituent DEP, represents a
promising antibiofilm strategy that may enhance antibiotic efficacy
and reduce resistance selection when used in combination therapies,
particularly for biofilm-associated infections, such as CAUTIs and
VAP. However, in vitro bioactivity likely reflects the
synergistic effects of multiple constituents within the crude
extract rather than the action of individual compounds.
Accordingly, molecular docking served as a predictive tool to
prioritize candidate bioactives, and further validation using
purified fractions will be necessary to confirm compound-specific
activity.
Nevertheless, the present study has certain
limitations. The antibiofilm activity was evaluated using a single
clinical isolate each of E. coli and K. pneumoniae,
which limits the generalizability of the findings. These strains
were clinical isolates obtained from Saveetha Dental College and
Hospitals and identified using the VITEK® 2 system;
however, no reference strains were included for comparative
analysis. In addition, detailed genotypic characterization,
including sequence typing and resistance gene profiling, was not
performed. Future studies incorporating a larger and genetically
diverse panel of clinical and reference isolates will be necessary
to validate and extend the applicability of these findings.
Additionally, GC-MS identified the major phytochemical
constituents, the individual compounds were not fractionated or
tested separately; therefore, the specific bioactive component(s)
could not be conclusively determined. Furthermore, in vivo
validation using appropriate infection models will be essential to
establish the therapeutic potential, safety profile, and
translational relevance of R. tuberosa L. for possible
clinical application.
In conclusion, the present study demonstrates that
the ethanolic leaf extract of R. tuberosa L. exerts
significant, growth-neutral antibiofilm effects against MDR E.
coli and K. pneumoniae, with sub-MIC concentrations
reducing biofilm biomass, while leaving planktonic growth largely
unaffected, consistent with an anti-virulence rather than
bactericidal mode of action. GC-MS profiling coupled with molecular
docking identified DEP as a major constituent and predicted its
potential interactions with key biofilm- and virulence-associated
proteins, including curli fibrils and pilus assembly factors in
E. coli and efflux pump and fimbrial components in K.
pneumoniae, providing a plausible mechanistic hypothesis rather
than direct functional evidence for the observed inhibition of
adhesion and biofilm maturation. Altogether, these findings suggest
that R. tuberosa L. represents a promising phytochemical
resource for the development of anti-virulence adjuncts to
conventional antibiotics in the management of biofilm-associated
infections caused by MDR Gram-negative pathogens, warranting
further validation through expanded strain panels, detailed
omics-based pathway analysis, and in vivo infection
models.
Supplementary Material
Docking grid parameters used for
molecular docking analysis.
Acknowledgements
Not applicable.
Funding
Funding: No funding was received.
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
AC contributed to the study design, data collection
and manuscript preparation. AB was involved in data collection,
data analysis and in the drafting of the manuscript. GRV supervised
the study, contributed to data interpretation, and critically
revised the manuscript. All authors confirm the authenticity of all
the raw data. All authors have read, and approved the final version
of the manuscript.
Ethics approval and consent to
participate
The bacterial isolates used in the present study
were provided as a laboratory gift from the Department of
Microbiology, Saveetha Dental College and Hospitals, Chennai,
India, at an earlier time and were subsequently transferred to
Kannur University, Kerala, for research purposes. The isolates were
obtained from the hospital and were transferred by the laboratory
authorities to the Kannur University laboratory for research use.
The present study did not involve direct patient interaction,
access to patient records, or collection of clinical specimens. The
isolates were handled strictly as microbiological strains for in
vitro experimental purposes. Upon receipt, the isolates were
reconfirmed in the laboratory based on standard morphological
characteristics prior to experimental use. As the study involved
only de-identified bacterial isolates and did not include human
participants or identifiable patient data, ethics approval and
informed patient consent were waived.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Liu HY, Prentice EL and Webber MA:
Mechanisms of antimicrobial resistance in biofilms. NPJ Antimicrob
Resist. 2(27)2024.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Luo Y, Yang Q, Zhang D and Yan W:
Mechanisms and control strategies of antibiotic resistance in
pathological biofilms. J Microbiol Biotechnol. 31:1–7.
2021.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Maldonado RF, Sá-Correia I and Valvano MA:
Lipopolysaccharide modification in Gram-negative bacteria during
chronic infection. FEMS Microbiol Rev. 40:480–493. 2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Vuotto C, Longo F, Balice M, Donelli G and
Varaldo P: Antibiotic resistance related to biofilm formation in
Klebsiella pneumoniae. Pathogens. 3:743–758. 2014.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Stahlhut SG, Struve C, Krogfelt KA and
Reisner A: Biofilm formation of Klebsiella pneumoniae on
urethral catheters requires either type 1 or type 3 fimbriae. FEMS
Immunol Med Microbiol. 65:350–359. 2012.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Surgers L, Boyd A, Girard PM, Arlet G and
Decré D: Biofilm formation by ESBL-producing strains of
Escherichia coli and Klebsiella pneumoniae. Int J Med
Microbiol. 309:13–18. 2019.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Lüthje P and Brauner A: Virulence factors
of Uropathogenic E. coli and their interaction with the
Host. Adv Microb Physiol. 65:337–372. 2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Subashchandrabose S and Mobley HLT:
Virulence and fitness determinants of uropathogenic Escherichia
coli. Microbiol Spectr 3: 10.1128/microbiolspec.UTI-0015-2012,
2015.
|
|
9
|
de Pace F, Nakazato G, Pacheco A, Boldrin
de Paiva J, Sperandio V and Dias da Silveira W: The type VI
Secretion system plays a role in type 1 fimbria expression and
pathogenesis of an avian pathogenic Escherichia coli strain.
Infect Immun. 78:4990–4998. 2010.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Rendón MA, Saldaña Z, Erdem AL,
Monteiro-Neto V, Vázquez A, Kaper JB, Puente JL and Girón JA:
Commensal and pathogenic Escherichia coli use a common pilus
adherence factor for epithelial cell colonization. Proc Natl Acad
Sci USA. 104:10637–10642. 2007.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Xu L, Li J, Wu W, Wu X and Ren J:
Klebsiella pneumoniae capsular polysaccharide: Mechanism in
regulation of synthesis, virulence, and pathogenicity. Virulence.
15(2439509)2024.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Langstraat J, Bohse M and Clegg S: Type 3
fimbrial shaft (MrkA) of Klebsiella pneumoniae, but not the
Fimbrial adhesin (MrkD), facilitates biofilm formation.
Infect Immun. 69:5805–5812. 2001.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Li Y and Ni M: Regulation of biofilm
formation in Klebsiella pneumoniae. Front Microbiol.
14(1238482)2023.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yan J, Pu S, Jia X, Xu X, Yang S, Shi J,
Sun S and Zhang L: Multidrug resistance mechanisms of carbapenem
resistant Klebsiella pneumoniae strains isolated in
Chongqing, China. Ann Lab Med. 37:398–407. 2017.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Liu X, Sai F, Li L, Zhu C and Huang H:
Clinical characteristics and risk factors of catheter-associated
urinary tract infections caused by Klebsiella pneumoniae.
Ann Palliat Med. 9:2668–2677. 2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Kushnareva М, Markhulia Kh М, Keshishyan E
and Semenov A: Ventilator-associated pneumonia caused by
Klebsiella pneumoniae in preterm newborn infants. Int J
Pediatric Res. 4:2469–5769. 2018.
|
|
17
|
Woo S, Marquez L, Crandall WJ, Risener CJ
and Quave CL: Recent advances in the discovery of plant-derived
antimicrobial natural products to combat antimicrobial resistant
pathogens: Insights from 2018-2022. Nat Prod Rep. 40:1271–1290.
2023.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Lu L, Hu W, Tian Z, Yuan D, Yi G, Zhou Y,
Cheng Q, Zhu J and Li M: Developing natural products as potential
anti-biofilm agents. Chin Med. 14(11)2019.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Shamim A, Ali A, Iqbal Z, Mirza MA, Aqil
M, Kawish SM, Siddiqui A, Kumar V, Naseef PP, Alshadidi AAF and
Saheer Kuruniyan M: Natural medicine a promising candidate in
combating microbial biofilm. Antibiotics (Basel).
12(299)2023.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Bouyahya A, Dakka N, Et-Touys A, Abrini J
and Bakri Y: Medicinal plant products targeting quorum sensing for
combating bacterial infections. Asian Pac J Trop Med. 10:729–743.
2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Asfour HZ: Anti-quorum sensing natural
compounds. J Microsc Ultrastruct. 6:1–10. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Pathoor NN, Ganesh PS, Gopal RK, Anshad
AR, Shankar EM, Mariappan V, Busi S, Salim SA, Kathiresan N,
Kulanthaivel L, et al: Attenuation of biofilm-encoding genes and
virulence attributes in clinical isolates of Acinetobacter
baumannii by essential oil derived from Myroxylon
balsamum. Sci Rep. 16(2861)2026.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Sharma A, Kumar A, Singh AK, Kumar KJ,
Narasimhan B and Kumar P: Ethnomedicinal uses, phytochemistry,
pharmacology, and toxicology of Ruellia tuberosa L.: A
review. Chem Biodivers. 21(e202400292)2024.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Ravi P and Pandey BKS: Phytochemical
studies on Ruellia tuberosa: A review. Int J Pharm Biol Sci.
8:1190–1195. 2018.
|
|
25
|
Guha S, Talukdar D, Mandal GK, Mukherjee
R, Ghosh S, Naskar R, Saha P, Murmu N and Das G: Crude extract of
Ruellia tuberosa L. flower induces intracellular ROS,
promotes DNA damage and apoptosis in triple negative breast cancer
cells. J Ethnopharmacol. 332(118389)2024.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Amajida H, Purwoko T and Susilowati A:
Antibacterial activity of ethanolic and n-hexane extracts of
Ruellia tuberosa leaves against Escherichia coli and
Bacillus subtilis bacteria. Biofarmasi J Natural Product
Biochem. 17:2580–2550. 2019.
|
|
27
|
Williams and Wilkins Publishers. Bergey's
Manual of Systematic Bacteriology. Baltimore, 2: 2001.
|
|
28
|
Clinical and Laboratory Standards
Institute (CLSI). Performance standards for antimicrobial
susceptibility testing. 32nd edition. CLSI supplement M100. Wayne
(PA), Clinical and Laboratory Standards Institute, 2022.
|
|
29
|
Balouiri M, Sadiki M and Ibnsouda SK:
Methods for in vitro evaluating antimicrobial activity: A review. J
Pharm Anal. 6:71–79. 2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
G S, Pathoor NN, Murthykumar K and Ganesh
PS: Targeting Pseudomonas aeruginosa PAO1 pathogenicity: The
role of Glycyrrhiza glabra in inhibiting virulence factors
and biofilms. Diagn Microbiol Infect Dis.
111(116674)2025.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Pathoor NN, Ganesh PS, Anshad AR, Gopal
RK, Ponmalar EM, Suvaithenamudhan S, Rudrapathy P and Shankar EM:
3-Hydroxybenzoic acid inhibits the virulence attributes and
disrupts biofilm production in clinical isolates of
Acinetobacter baumannii. Eur J Clin Microbiol Infect Dis.
44:653–669. 2025.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Girija S, Duraipandiyan V, Kuppusamy PS,
Gajendran H and Rajagopal R: Chromatographic characterization and
GC-MS evaluation of the bioactive constituents with antimicrobial
potential from the pigmented ink of Loligo duvauceli. Int
Sch Res Notices. 2014(820745)2014.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Surgers L, Boyd A, Girard PM, Arlet G and
Decré D: Biofilm formation by ESBL-producing strains of
Escherichia coli and Klebsiella pneumoniae. Int J Med
Microbiol. 309:13–18. 2019.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Kirana P, Prashantkumar CS, Soundarya R,
Shwetha GS, Kalasad MN and Gayathri D: Tinospora
cordifolia's green synthesis approach: Comparative study on
biogenic and chemical synthesis of zinc oxide and zinc sulfide
nanoparticles. Materials Int. 6(40)2024.
|
|
35
|
Buchmann D, Schwabe M, Weiss R, Kuss AW,
Schaufler K, Schlüter R, Rödiger S, Guenther S and Schultze N:
Natural phenolic compounds as biofilm inhibitors of
Multidrug-resistant Escherichia coli- the role of similar
biological processes despite structural diversity. Front Microbiol.
14(1232039)2023.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Kim Y, Kim S, Cho KH, Lee JH and Lee J:
Antibiofilm activities of cinnamaldehyde analogs against
uropathogenic Escherichia coli and Staphylococcus
aureus. Int J Mol Sci. 23(7225)2022.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Olszewska MA, Gędas A and Simões M: The
effects of eugenol, Trans-Cinnamaldehyde, citronellol, and
terpineol on Escherichia coli biofilm control as assessed by
Culture-dependent and -independent methods. Molecules.
25(2641)2020.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Rony Varughese RM, Pathoor NN, Ranganathan
P and Ganesh PS: Efficacy of Rhamnus frangula extract
against Acinetobacter baumannii biofilms: Histopathological
evidence from ex vivo goat models. World Acad Sci J. 7(36)2025.
|
|
39
|
Paula-Ramos L, da Rocha Santos CE, Camargo
Reis Mello D, Nishiama Theodoro L, De Oliveira FE, Back Brito GN,
Junqueira JC, Jorge AOC and de Oliveira LD: Klebsiella
pneumoniae planktonic and biofilm reduction by different plant
extracts: In vitro study. ScientificWorldJournal.
2016(3521413)2016.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Zubair M: Antimicrobial and Anti-biofilm
activities of Coffea Arabica L. against the clinical strains
isolated from diabetic foot ulcers. Cureus.
16(e52539)2024.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Keohane CE, Steele AD, Fetzer C,
Khowsathit J, Van Tyne D, Moynié L, Gilmore MS, Karanicolas J,
Sieber SA and Wuest WM: Promysalin elicits Species-selective
inhibition of Pseudomonas aeruginosa by targeting succinate
dehydrogenase. J Am Chem Soc. 140:1774–1782. 2018.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Mahoney AR, Storek KM and Wuest WM:
Structure-based design of promysalin analogues to overcome
mechanisms of bacterial resistance. ACS Omega. 8:12558–12564.
2023.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Rashiya N, Sangavi J, Padmini N,
Langeswaran K, Alagarsamy A, Selvakumar G and Saravanan M: In
silico and in vitro analysis of diethyl phthalate as a quorum
sensing inhibitor and its antitumor evaluation against MDA-MB-231
cell lines. Mol Divers. 30:761–771. 2026.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Rashiya N, Padmini N, Ajilda AAK,
Prabakaran P, Durgadevi R, Veera Ravi A, Ghosh S, Sivakumar N and
Selvakumar G: Inhibition of biofilm formation and quorum sensing
mediated virulence in Pseudomonas aeruginosa by marine
sponge symbiont Brevibacterium casei strain Alu 1. Microb
Pathog. 150(104693)2021.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Kim YR and Sang MK: Effects of
di-(2-ethylhexyl) phthalate on growth, metabolism, and virulence of
the plant pathogenic bacterium Acidovorax citrulli. Front
Cell Infect Microbiol. 13(1228713)2023.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Louis M, Tahrioui A, Verdon J, David A,
Rodrigues S, Barreau M, Manac'h M, Thiroux A, Luton B, Dupont C, et
al: Effect of phthalates and their substitutes on the physiology of
Pseudomonas aeruginosa. Microorganisms.
10(1788)2022.PubMed/NCBI View Article : Google Scholar
|