Introduction
RNA-binding proteins (RBPs) are known to play a
crucial role in post-transcriptional regulation in gene expression,
and regulate all aspects of RNA metabolism and function, such as
polyadenylation, RNA splicing, transport, stability and
translation; thus, they represent critical mechanisms for gene
regulation in mammalian cells (1,2).
They contain one or more RNA-binding motifs, such as the RNA
recognition motif (RRM), the human heterogeneous nuclear
ribonucleoprotein (hnRNP) K homology motif, the RGG box and the
double-stranded RNA binding domain (dsRBD) motif. RRM is the most
prevalent type of eukaryotic RNA-binding motif (3), which is composed of two submotifs,
RNP1 and RNP2 (3). RBPs are
involved in the expression of various genes responsible for
regulating biological processes and cellular functions, and thus
expected mutations or the aberrant production of RBPs can cause
cancer progression (4,5).
The RNA binding motif protein 38 (RBM38, also known
as RNPC1) gene is located on chromosome 20q13 and is expressed in a
variety of tissues. It belongs to the RRM family of RBPs, is
expressed as RNPC1a with 239 amino acids and as RNPC1b with 121
amino acids (6). RNPC1 plays
pivotal roles in regulating a wide range of biological processes,
ranging from cell proliferation and cell cycle arrest to cell
myogenic differentiation (7,8).
It is capable of regulating these biological processes by binding
and stabilizing the mRNA of p21, p73, Hu antigen R (HuR) and
macrophage inhibitory cytokine-1 (MIC-1) (6,7,9,10),
or by binding to the mRNAs of p63, murine double minute-2 (MDM2)
and p53 and mediating the decrease in the mRNA levels and the
attenuation of the translation of these proteins (11–13).
RNPC1 was originally recognized as an oncogene, and
was frequently found to be amplified in prostate (14,15), ovarian cancer (16), colorectal cancer (17,18), chronic lymphocytic leukemia
(19), colon carcinoma (20), esophageal adenocarcinoma (21), dog lymphomas (13) and breast cancer (22–24). Recently, new evidence suggests
that RNPC1 acts as a tumor suppressor. It has been reported that
RNPC1 is part of a negative feedback loop, which restricts E2F
transcription factor 1 (E2F1) activity by limiting cell cycle
progression at the G1-S boundary (25). The expression of RNPC1 has been
shown to highly correlate with increased survival in patients with
ovarian cancer (25). In breast
cancer, RNPC1 functions as a tumor repressor, possibly through
promoter hypermethylation silencing (26). In the present study, we identified
RNPC1 genes from mammalian genomes using comparative genomic
analyses. We then searched for conserved transcription
factor-binding sites within the promoter regions of the human RNPC1
gene. Analysis of the expression data and functionally relevant
single nucleotide polymorphisms (SNPs), and comparative proteomic
analyses were conducted. Furthermore, a meta-analysis of the
prognostic value of the RNPC1 gene in various types of cancer was
also performed.
Materials and methods
Identification of the complete RNPC1 gene
in vertebrate genomes and integrative genomic analyses
The RNPC1 gene and amino acid sequences were
selected from the Ensembl database (http://www.ensembl.org/index.html), based on
orthologous and paralogous associations. The selected RNPC1
sequences were applied as queries in order to search for the RNPC1
gene using the BLAST tool at the National Center for Biotechnology
Information (NCBI), in order to confirm whether their best hit was
an RNPC1 gene (27–33). The number, length and structure of
the exons and introns in the RNPC1 gene in all species were
collected from Ensembl. The number and length of the RNPC1 exons
and introns in all sequences were then subjected to exon-intron
conservation analyses. Conserved transcription factor-binding sites
within the promoter region of the human RNPC1 gene were obtained
from the SABiosciences’ proprietary database, which combines Text
Mining Application and data from the UCSC Genome Browser
(http://genome.ucsc.edu/) (27–33).
Comparative proteomic analyses of RNPC1
protein
The protein-coding sequences of RNPC1 were aligned
using the ClustalW program in MEGA 5.05. We constructed a maximum
likelihood (ML) tree of RNPC1 amino acid sequences using MEGA 5.05
with the optimal model (Kimura 2-parameter). Relative support of
the internal node was performed by bootstrap analyses with 1,000
replications for ML reconstructions (34). The CodeML program, implemented in
the PAML 4.7 software package, was used to investigate whether the
RNPC1 protein is under positive selection (35). The site-specific model was
developed using the likelihood ratio test (LRT) to compare the M7
(null model) with the M8 model. M7 is a null model that does not
allow for any codons with ω>1, whereas the M8 model allows for
positively selected sites (ω>1). When the M8 model fits the data
significantly (P-value <0.05) better than the null model (M7),
the presence of sites with ω>1 is suggested. On the contrary,
the results of P-value >0.05 are proof the absence of sites with
ω>1. Twice the log likelihood difference between the two
compared models (2Δl) is compared against χ2 with
critical values being 5.99 and 9.21 at the 0.05 and 0.01
significance levels, respectively (36).
Identification of functionally relevant
SNPs in the human RNPC1 gene and somatic mutations in human
cancer
Functionally relevant SNPs of the human RNPC1 gene
were identified as previously described (27–33). The SNPs were extracted from
Ensembl (http://www.ensembl.org) and NCBI’s SNPdb
(http://www.ncbi.nlm.nih.gov). The SNPs
that disrupted exonic splicing enhancer (ESE)/exonic splicing
silencer (ESS) motifs and caused missence mutations were also
identified. The identification of somatic mutations of the human
RNPC1 gene in human cancer was conducted using COSMIC, a database
for mining complete cancer genomes in the catalogue of somatic
mutations in cancer (37).
Analysis of the expression of the human
RNPC1 gene
The expression profiles of RNPC1 in normal human
tissues were obtained from ArrayExpress (38). Virtual northern blot analysis of
NCBI’s UniGene dataset was also performed, as previously described
(31–33).
Meta-analysis of the prognostic value of
the RNPC1 gene in cancer
For meta-analysis, the PrognoScan database was used
(39). This includes: i) a large
collection of publicly available cancer microarray datasets with
clinical annotation, and ii) a tool for assessing the biological
association between gene expression and prognosis. PrognoScan
employs the minimum P-value approach to group patients for survival
analysis. PrognoScan provides a powerful platform for evaluating
potential tumor markers and therapeutic targets, and is publicly
accessible at http://www.prognoscan.org/. The human RNPC1 gene was
inputted as a query, and the data were collected for analysis.
PrognoScan displays a summary in table format of tests for RNPC1
with columns for dataset, cancer type, subtype, endpoint, cohort,
contributor, array type, probe ID, number of patients, optimal
cutpoint, Pmin and Pcor.
Results
Comparative proteomic analysis of the
RNPC1 protein identified in vertebrate genomes
All the RNPC1 nucleotide and protein sequences were
collected from ENSEMBL and checked using BLAST at NCBI. The
complete RNPC1 gene was identified in the human, bushbaby,
chimpanzee, macaque, gorilla, olive baboon, vervet-AGM (vervet
monkey), guinea pig, mouse, rat, cow, dog, ferret, hedgehog,
armadillo, elephant, lesser hedgehog tenrec, anole lizard, chicken,
Chinese softshell turtle, duck, Amazon molly, flycatcher, cave
fish, Fugu, medaka, platyfish, spotted gar, stickleback, tilapia,
Tetraodon and zebrafish genomes. The sequences and
structural alignment of RNPC1 in these genomes are shown in
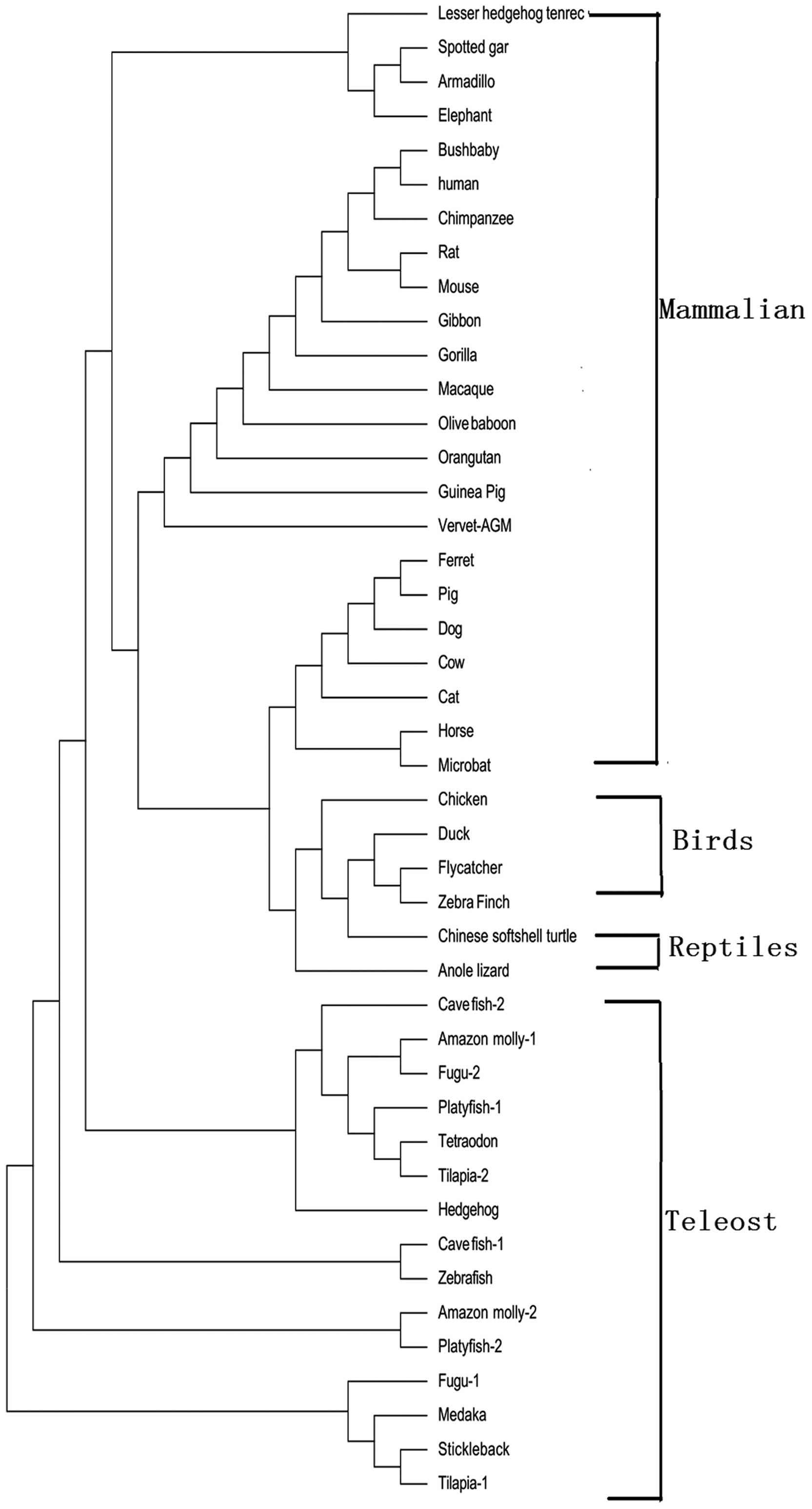
Fig. 1. The phylogenetic tree was
constructed according to the protein-coding sequences of RNPC1,
using the maximum likelihood method; the RNPC1 gene from the
mammalian, bird, reptile and teleost lineages formed
species-specific clusters (Fig.
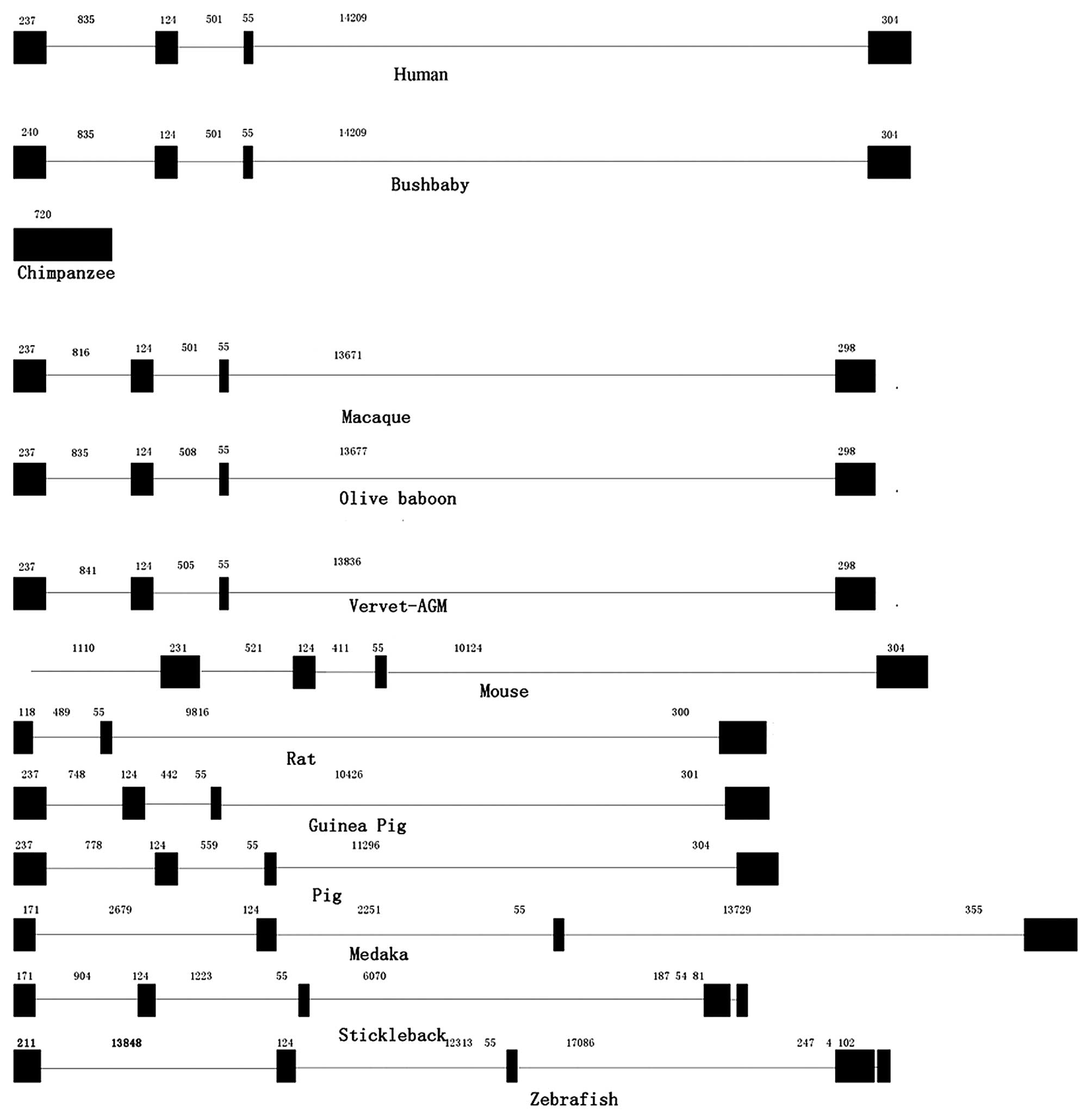
2). The exon-intron data collected from the ENSEMBL database
are shown in Table I and Fig. 3. In the majority of vertebrates,
the RNPC1 gene exhibited exon-intron conservation, with 4 exons and
3 introns, with similar sizes for each exon and intron (Table I). However, there were 5 exons and
4 introns in the RNPC1 gene in the mouse and 3 fish species
(stickleback, tetraodon and zebrafish). Thus, the intron
deletions in the RNPC1 gene may occur during the evolutionary
process of these 3 species of fish. Furthermore, site-specific
tests for positive selection were performed for the vertebrate,
mammalian, primate, and mammalian excluding primate, rodent and
teleost lineages. We were unable to identify any site which was
under positive selection by the M7 and M8 models in the RNPC1
protein. It seemed that RNPC1 in vertebrates was under purifying
selection (data not shown).
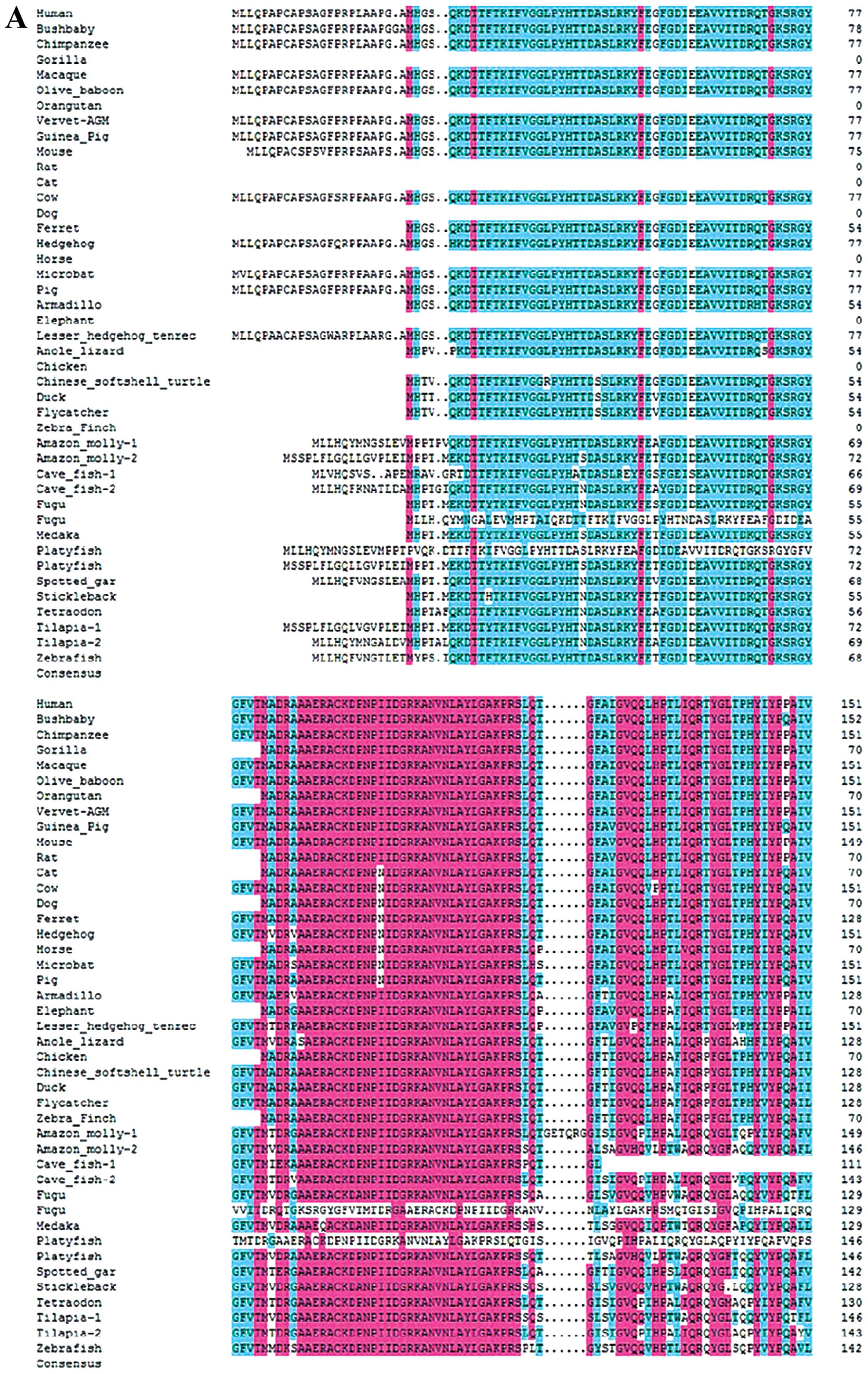 | Figure 1Sequence and structural alignment of
RNPC1 in vertebrates. (A) Alignment of RNPC1 in vertebrates from
position 1–151. (B) Alignment of RNPC1 in vertebrates from position
152–239. All the RNPC1 gene and protein sequences were collected
from Ensembl and checked using the BLAST tool at NCBI. The complete
RNPC1 gene was identified in the human, bushbaby, chimpanzee,
macaque, gorilla, olive baboon, vervet-AGM, guinea pig, mouse, rat,
cow, dog, ferret, hedgehog, armadillo, elephant, lesser hedgehog
tenrec, anole lizard, chicken, Chinese softshell turtle, duck,
Amazon molly, flycatcher, cave fish, fugu, medaka, platyfish,
spotted gar, stickleback, tilapia, tetraodon and zebrafish
genomes. |
 | Table IExon and intron lengths of RNPC1. |
Table I
Exon and intron lengths of RNPC1.
| Species | Exon 1 | Intron 1 | Exon 2 | Intron 2 | Exon 3 | Intron 3 | Exon 4 | Intron 4 | Exon 5 | Total exons |
|---|
| Human | 237 | 835 | 124 | 501 | 55 | 14209 | 304 | | | 720 |
| Bushbaby | 240 | 842 | 124 | 430 | 55 | 11713 | 304 | | | 723 |
| Chimpanzee | 720 | | | | | | | | | 720 |
| Macaque | 237 | 816 | 124 | 503 | 55 | 13671 | 298 | | | 714 |
| Olive baboon | 237 | 835 | 124 | 508 | 55 | 13677 | 298 | | | 714 |
| Vervet-AGM | 237 | 841 | 124 | 505 | 55 | 13836 | 298 | | | 714 |
| Mouse | 0 | 1110 | 231 | 521 | 124 | 411 | 55 | 10124 | 304 | 714 |
| Rat | 118 | 489 | 55 | 9816 | 300 | | | | | 473 |
| Guinea Pig | 237 | 748 | 124 | 442 | 55 | 10426 | 301 | | | 717 |
| Cow | 237 | 753 | 124 | 483 | 55 | 12228 | 304 | | | 720 |
| Pig | 237 | 778 | 124 | 559 | 55 | 11296 | 304 | | | 720 |
| Armadillo | 168 | 747 | 124 | 399 | 55 | 12317 | 283 | | | 630 |
| Amazon molly-1 | 213 | 4109 | 141 | 1707 | 56 | 6429 | 349 | | | 759 |
| Amazon molly-2 | 222 | 4039 | 124 | 2832 | 55 | 13455 | 364 | | | 765 |
| Cave fish-2 | 213 | 2209 | 124 | 151 | 55 | 4745 | 316 | | | 708 |
| Fugu-1 | 171 | 789 | 124 | 1405 | 55 | 5561 | 340 | | | 690 |
| Medaka | 171 | 2679 | 124 | 2251 | 55 | 13729 | 355 | | | 705 |
| Platyfish-2 | 222 | 4066 | 124 | 2973 | 55 | 15548 | 352 | | | 753 |
| Spotted gar | 210 | 2827 | 124 | 2302 | 55 | 6210 | 301 | | | 690 |
| Stickleback | 171 | 904 | 124 | 1223 | 55 | 6070 | 187 | 54 | 81 | 618 |
|
Tetraodon | 174 | 1629 | 124 | 269 | 55 | 3386 | 214 | 27 | 99 | 666 |
| Zebrafish | 211 | 13848 | 124 | 12313 | 55 | 17086 | 247 | 4 | 102 | 739 |
Expression profile of the human RNPC1
gene
The investigation of the available microarray data
and virtual northern blot analysis, we revealed the predominant
expression of RNPC1 in bone marrow, whole blood, lymph node,
thymus, brain, cerebellum, retina, spinal cord, heart, smooth
muscle, skeletal muscle, small intestine, colon, adipocyte,
kidneys, liver, lungs, pancreas, thyroid, salivary gland, skin,
breast, ovaries, uterus, placenta, prostate and testes. When we
searched the PrognoScan database, we found that human RNPC1 was
also expressed in bladder, blood, brain, breast, colorectal, eye,
head and neck, lung, ovarian, skin and soft tissue cancer.
Comparative genomic analysis of the human
RNPC1 gene
The sex determining region Y (SRY)-box 5 (Sox5),
runt-related transcription factor 3 (RUNX3), CCAAT displacement
protein 1 (CUTL1), v-rel avian reticuloendotheliosis viral oncogene
homolog (Rel)A, peroxisome proliferator-activated receptor γ
isoform 2 (PPARγ2) and activating transcription factor 6 (ATF6)
regulatory transcription factor binding sites were identified in
the upstream (promoter) region of the RNPC1 gene.
Identification of functionally relevant
SNPs in the human RNPC1 gene and somatic mutations in human
cancer
A total of 429 SNPs were identified in the human
RNPC1 gene. Of these, 34 SNPs were functionally relevant, including
14 SNPs causing missense mutations, 8 exonic splicing enhancer SNPs
and 12 SNPs causing nonsense mutations (Table II). By searching the COSMIC
database, we identified 30 somatic mutations of RNPC1 in 10,148
cancer samples (Table III).
 | Table IIFunctionally relevant SNPs in the
human RNPC1 gene. |
Table II
Functionally relevant SNPs in the
human RNPC1 gene.
| SNP ID | Chr 20 position
sequence | Sequence | Type | Amino acid
change |
|---|
| rs150246007 | 55982673(+) | CCCGTC/TGCTGT | mis | S L |
| rs201278266 | 55982720(+) | CCTACA/GCCCAG | mis | T A |
| rs201875738 | 55982772(+) | GCCTGC/TCACGG | mis | A V |
| rs199521379 | 55982775(+) | TGCCAC/TGGCTG | mis | T M |
| rs201066490 | 55982813(+) | CCGCCA/GTGCCC | mis | M V |
| rs201744631 | 55982843(+) | CACCCA/GCGGGC | mis | T A |
| rs16980970 | 55982858(+) | CTTTCG/CTGCAG | mis | L V |
| rs1065289 | 55982781(+) | GGCTGC/ACAGCT | mis | D A |
| rs369246420 | 55982844(+) | ACCCGC/TGGGCA | mis | A V |
| rs373452137 | 55982615(+) | ACTACA/GTCTAC | mis | I V |
| rs377081682 | 55982714(+) | GCCCGG/TCCTAC | mis | A S |
| rs368322258 | 55982789(+) | GCTTCA/GTGGGC | mis | M V |
| rs10652881 | 55982715(+) | CCCGGC/TCTACG | mis | A V |
| rs1065290 | 55982817(+) | CGTGCC/ACCAGG | mis | H P |
| rs11546710 | 55983073(+) | AGAGAC/TGGCTT | ese | |
| rs6128022 | 55983476(+) | TCCCAG/AGCGCA | ese | |
| rs8126441 | 55983505(+) | GGGGCC/AGCCGG | ese | |
| rs3829703 | 55983509(−) | TTGGCC/TGGCGG | ese | |
| rs11546713 | 55984212(−) | CCCCCA/GCCCTC | ese | |
| rs1065292 | 55983556(+) | TTTTTC/TTTGTA | ese | |
| rs1052752 | 55983512(+) | CCGGCC/AAAAGG | ese | |
| rs3207621 | 55983521(+) | GGCCCC/TTTTCC | ese | |
| rs141028132 | 55982671(+) | GTCCCG/ATCGCT | syn | P |
| rs201839752 | 55982683(+) | TCCTCA/GCCCTA | syn | S |
| rs115516069 | 55982695(+) | ATTGAG/ATACAC | syn | E |
| rs200910302 | 55982719(+) | GCCTAC/TGCCCA | syn | Y |
| rs199953546 | 55982758(+) | CCATAC/TGCCGC | syn | Y |
| rs143107197 | 55982812(+) | GCCGCC/TGTGCC | syn | A |
| rs373297597 | 55982875(+) | GCGCCA/GCAGCT | syn | P |
| rs373492567 | 55982887(+) | CAGCCA/TGACAG | syn | P |
| rs202004284 | 55982704(+) | ACGCCG/AGCCAG | syn | P |
| rs376442730 | 55982872(+) | CAGGCA/GCCGCA | syn | A |
| rs377524807 | 55982644(+) | CCCAGC/TGTGGT | syn | S |
| rs374582705 | 55982713(+) | AGCCCA/GGCCTA | syn | P |
 | Table IIISomatic mutations of RNPC1 in cancer
tissue. |
Table III
Somatic mutations of RNPC1 in cancer
tissue.
| Position (AA) | Mutation (CDS) | Mutation (amino
acid) | Mutation ID
(COSM) | Count | Mutation type |
|---|
| 19 | c.55G>C | p.A19P | COSM1412673 | 2 | Substitution -
missense |
| 20 | c.56_57insC |
p.A20fs*70 | COSM3724433 | 2 | Insertion -
frameshift |
| 49 | c.146C>G | p.S49W | COSM3963694 | 1 | Substitution -
missense |
| 55 | c.163G>C | p.E55Q | COSM397137 | 1 | Substitution -
missense |
| 78 | c.234C>T | p.G78G | COSM724075 | 1 | Substitution -
coding silent |
| 83 | c.249C>T | p.A83A | COSM1165242 | 1 | Substitution -
coding silent |
| 89 | c.265G>A | p.E89K | COSM724074 | 1 | Substitution -
missense |
| 89 | c.266A>G | p.E89G | COSM117504 | 1 | Substitution -
missense |
| 97 | c.290C>T | p.P97L | COSM224597 | 1 | Substitution -
missense |
| 109 | c.325G>A | p.A109T | COSM1412674 | 1 | Substitution -
missense |
| 112 | c.336C>T | p.G112G | COSM192074 | 1 | Substitution -
coding silent |
| 116 | c.346C>T | p.R116W | COSM1579588 | 1 | Substitution -
missense |
| 116 | c.347G>A | p.R116Q | COSM724073 | 1 | Substitution -
missense |
| 120 | c.359C>T | p.T120M | COSM1028363 | 1 | Substitution -
missense |
| 131 | c.392C>G | p.P131R | COSM3405224 | 1 | Substitution -
missense |
| 132 | c.395C>G | p.T132S | COSM3363328 | 1 | Substitution -
missense |
| 139 | c.415G>A | p.G139R | COSM125790 | 1 | Substitution -
missense |
| 147 | c.441A>C | p.P147P | COSM4134683 | 1 | Substitution -
coding silent |
| 150 | c.450C>T | p.I150I | COSM1565701 | 1 | Substitution -
coding silent |
| 160 | c.478G>A | p.A160T | COSM1412675 | 1 | Substitution -
missense |
| 163 | c.487C>T | p.P163S | COSM3548089 | 1 | Substitution -
missense |
| 168 | c.502C>T | p.P168S | COSM3548090 | 1 | Substitution -
missense |
| 172 | c.515A>G | p.Y172C | COSM4099709 | 1 | Substitution -
missense |
| 174 | c.521C>T | p.P174L | COSM1412676 | 1 | Substitution -
missense |
| 176 | c.527G>A | p.S176N | COSM270012 | 1 | Substitution -
missense |
| 193 | c.577G>A | p.A193T | COSM3770849 | 1 | Substitution -
missense |
| 193 | c.579C>T | p.A193A | COSM3548091 | 1 | Substitution -
coding silent |
| 210 | c.628G>A | p.A210T | COSM4099710 | 1 | Substitution -
missense |
| 220 | c.659C>T | p.P220L | COSM3911642 | 1 | Substitution -
missense |
| 225 | c.675C>T | p.F225F | COSM263280 | 1 | Substitution -
coding silent |
Meta-analysis of the prognostic value of
the human RNPC1 gene in cancer
When provided with the specific gene, PrognoScan
displays a summary (in table format) of tests for the gene, with
columns for the dataset, cancer type, subtype, endpoint, cohort,
contributor, array type, probe ID, number of patients, optimal
cut-point, Pmin and Pcor. Among the databases which detected the
expression of the RNPC1 gene, an association between the expression
of the RNPC1 gene and cancer prognosis was noted in 14 of the 94
tests (blood cancer 2/9, brain cancer 1/5, breast cancer 3/30,
colorectal cancer 1/9, eye 1/1, head and neck cancer 0/1, lung
cancer 5/24, ovarian cancer 1/10, skin cancer 0/1 and soft tissue
cancer 0/1), with a 5% significance level (Table IV). As regards blood, colorectal
and eye cancer, a correlation between the decreased expressino of
the RNPC1 gene and poor survival was observed. However, a higher
expression of the RNPC1 gene was found to correlated with a poor
survival in patients with brain and ovarian cancer. Of the 3 breast
cancer cases, a lower expression of the RNPC1 gene, which
correlated with poor survival, was observed in 2 cases (E-TABM-158
and GSE7849), while a higher expression of the RNPC1 gene
correlated with a poor survival in the case of GSE11121. Of the
lung cancer cases, a lower expression of the RNPC1 gene, which
correlated with poor survival, was noted in 2 cases (GSE31210 and
GSE31211), while a higher expression of the RNPC1 gene correlated
with poor survival in 3 cases (HARVARD-LC, GSE4716-GPL3694 and
Jacob-00182-CANDF) cases.
 | Table IVDataset content from PrognoScan
demonstrating an association between the expression of the RNPC1
gene and cancer prognosis. |
Table IV
Dataset content from PrognoScan
demonstrating an association between the expression of the RNPC1
gene and cancer prognosis.
| Database | Case type | Subsyte | No. of
patients | Endpoint | Cut-point | P-value | Prognosis | Refs. |
|---|
|
GSE12417-GPL570 | Blood cancer | AML | 79 | Overall
survival | 0.15 | 0.001223 | 1 | (57) |
| GSE12417-GPL96 | Blood cancer | AML | 163 | Overall
survival | 0.39 | 0.035113 | 1 | (57) |
| GSE4271-GPL96 | Brain cancer | Astrocytoma | 77 | Overall
survival | 0.82 | 0.003856 | 2 | (58) |
| GSE7849 | Breast cancer | | 76 | Disease-free
survival | 0.3 | 0.030908 | 1 | (59) |
| GSE11121 | Breast cancer | | 200 | Distant
metastasis-free survival | 0.82 | 0.008577 | 2 | (60) |
| E-TABM-158 | Breast cancer | | 117 | Disease-specific
survival | 0.7 | 0.035878 | 1 | (61) |
| GSE17537 | Colorectal
cancer | | 55 | Overall
survival | 0.11 | 0.02856 | 1 | (62) |
| GSE22138 | Eye cancer | Uveal melanoma | 63 | Distant
metastasis-free survival | 0.33 | 0.018941 | 1 | (63) |
|
jacob-00182-CANDF | Lung cancer | Adenocarcinoma | 82 | Overall
survival | 0.82 | 0.020135 | 2 | (64) |
| HARVARD-LC | Lung cancer | Adenocarcinoma | 84 | Overall
survival | 0.69 | 0.004177 | 2 | (65) |
| GSE31210 | Lung cancer | Adenocarcinoma | 204 | Relapse-free
survival | 0.51 | 0.001557 | 1 | (66) |
| GSE31211 | Lung cancer | Adenocarcinoma | 204 | Overall
survival | 0.34 | 0.000939 | 1 | (66) |
|
GSE4716-GPL3694 | Lung cancer | NSCLC | 50 | Overall
survival | 0.82 | 0.044441 | 2 | (67) |
| DUKE-OC | Ovarian cancer | | 133 | Overall
survival | 0.85 | 0.010709 | 2 | (68) |
Discussion
RNPC1 (also known as RBM38), is an RBP that contains
one RRM domain. It is expressed as two isoforms, RNPC1a and RNPC1b
(6). RNPC1 is a direct target of
p53 and can interact with other members of the p53 family; it can
stabilize p21 and p73 transcripts and destabilize p63 transcripts.
It can also bind and stabilize the mRNA of the CDK inhibitor, p21,
thereby inducing cell cycle arrest in the G1 phase (7,8).
RNPC1 also binds and stabilizes the mRNA of another RBP HuR, which
in turn facilitates RNPC1-mediated growth arrest (7). In the present study, the complete
RNPC1 gene was identified in the human, bushbaby, chimpanzee,
macaque, gorilla, olive baboon, vervet-AGM, guinea pig, mouse, rat,
cow, dog, ferret, hedgehog, armadillo, elephant, lesser hedgehog
tenrec, anole lizard, chicken, Chinese softshell turtle, duck,
Amazon molly, flycatcher, cave fish, Fugu, medaka, platyfish,
spotted gar, stickleback, tilapia, tetraodon and zebrafish
genomes, suggesting that RNPC1 exists in all types of vertebrates,
including fish, amphibians, birds and mammals. In the different
genomes, the gene had a similar organization, namely 4 exons/3
introns, and all the genetic loci were syntenically conserved. The
phylogenetic tree revealed that the RNPC1 gene from the mammalian,
bird, reptile and teleost lineage formed species-specific clusters.
As observed from the alignment and phylogenetic tree, RNPC1 in
mammals is conserved among vertebrate genomes, suggesting that the
function of RNPC1 plays an important physiological role in all
vertebrates during the evolution process.
The investigation of available microarray data and
virtual northern blot analysis confirmed the predominant expression
of RNPC1 in the bone marrow, whole blood, the lymph node, thymus,
brain, cerebellum, retina, spinal cord, heart, smooth muscle,
skeletal muscle, small intestine, colon, adipocyte, kidneys, liver,
lungs, pancreas, thyroid, salivary gland, skin, breast, ovaries,
uterus, placenta, prostate and testes. Thus, RNPC1 is widely
expressed in a number of tissues and organs. A total of 429 SNPs
were identified in the human RNPC1 gene. Of these, 34 SNPs were
functionally relevant, including 14 SNPs causing missense
mutations, 8 exonic splicing enhancer SNPs and 12 SNPs causing
nonsense mutations, which may affect the multiple functions of
RNPC1. However, the effects of these SNPs on RNPC1 physiological
and pathological functions require further investigation.
RNPC1 was originally recognized as an oncogene, and
was frequently found to be amplified in prostate (14,15), ovarian (16) and colorectal cancer (17,18), chronic lymphocytic leukemia
(19), colon carcinoma (20), esophageal cancer (21), dog lymphomas (13) and breast cancer (22–24). In our previous study, we found
that RNPC1 played a tumor suppressor role role in breast cancer
(40). In the present study, we
first noted that RNPC1 was indeed expressed in bladder, blood,
brain, breast, colorectal, eye, head and neck, lung, ovarian, skin
and soft tissue cancer. Out of 94 tests, 14 revealed an association
between RNPC1 gene expression and cancer prognosis (blood 2/9,
brain 1/5, breast 3/30, colorectal 1/9, eye 1/1, head and neck 0/1,
lung 5/24, ovarian 1/10, skin 0/1 and soft tissue cancer 0/1). It
is important to note that the association between the expression of
RNPC1 and prognosis varied in different types of cancer, and even
in the same type of cancer from different databases. This suggests
that the function of RNPC1 in these tumors may be multidimensional,
and that RNPC1 is not just a tumor suppressor or promoter.
Moreover, we identified 30 somatic mutations of RNPC1 in cancer
tissues in the present study. Further investigation is required to
elucidate the mechanisms through which these mutations affect tumor
formation. The mechanisms underlying the role of RNPC1 in the
process of these tumors may be involve the mRNA stabilizion of
oncogenes or anti-oncogenes, such as p53 (13), p63 (11), MDM2 (12), p73 (9), HuR (7) and p21 (6). However, the mechanisms underlying
the role of RNPC1 in the developmental process of these tumors
require further investigation.
The Sox5, RUNX3, CUTL1, RelA, CCAAT-enhancer-binding
protein (C/EBP)α, c-Ets-1, PPARγ2 and ATF6 regulatory transcription
factor binding sites were identified in the upstream (promoter)
region of the RNPC1 gene. Sox5 plays a role in the regulation of
embryonic development and in the determination of cell fate. It can
function as a transcriptional regulator after forming a protein
complex with other proteins. It has a negative effect on cell
proliferation in some cell types and functions as a target of
microRNAs (41,42). RUNX3 encodes a member of the runt
domain-containing family of transcription factors. A heterodimer of
this protein and a β subunit forms a complex that binds to the core
DNA sequence 5′-PYGPYGGT-3′ found in a number of enhancers and
promoters, and can either activate or suppress transcription. It
functions as a tumor suppressor and is frequently deleted or
transcriptionally silenced in cancer (43–46). CUTL1 is a transcription factor
which plays a role in development and multiple physiological
processes. Emerging evidence indicates that CUTL1 is not only
involved in developmental events, but also in pathological
processes, such as tumorigenesis and multiple signal transduction
pathways of cancer (47,48). RelA is a subunit of the nuclear
factor (NF-κB) p65. NF-κB is an ubiquitous transcription factor
which plays a role in several biological processes. NF-κB is
composed of NFKB1 or NFKB2 bound to either REL, RELA or RELB. NF-κB
is a pleiotropic transcription factor present in almost all cell
types and is the endpoint of a series of signal transduction events
that are initiated by a vast array of stimuli related to a number
of biological processes, such as inflammation, immunity,
differentiation, cell growth, tumorigenesis and apoptosis (49–52). C/EBPα is required for the proper
control of adipogenesis, glucose metabolism, granulocytic
differentiation, lung development and the development of various
types of cancer (53,54). c-Ets-1 is known to play an
important role in various biological processes, such as
development, differentiation, proliferation, apoptosis, migration,
tissue remodeling, invasion and angiogenesis in a variety of cell
types, including B cells, endothelial cells, fibroblasts and
neoplastic cells (55,56). These tumor-related transcriptional
factors may be involved in the effects of RNPC1 in tumors (14–24).
In conclusion, integrative genomic analyses of RNPC1
and its role in cancer prediction provide a powerful tool for the
evaluation of RNPC1 as a potential tumor markers and therapeutic
targets in cancer research.
Acknowledgments
The present study was supported by grants from the
National Natural Science Foundation of China (nos. 81272916 and
81202077), the Key Project of Jiangsu Provincial Health (H201110 to
Q.D.), the Project of Jiangsu Province Traditional Chinese Medicine
Bureau (LZ11084), the ̔Six Talents Peak̓ projects of Jiangsu
Province (to T.-S.X.), the Qinglan project of Jiangsu Province (to
T.-S.X.) and a project funded by the Priority Academic Program
Development of Jiangsu higher Education Institutions (PAPD).
References
|
1
|
Kim MY, Hur J and Jeong S: Emerging roles
of RNA and RNA-binding protein network in cancer cells. BMB Rep.
42:125–130. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Krecic AM and Swanson MS: hnRNP complexes:
composition, structure, and function. Curr Opin Cell Biol.
11:363–371. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Dreyfuss G, Matunis MJ, Piñol-Roma S and
Burd CG: hnRNP proteins and the biogenesis of mRNA. Annu Rev
Biochem. 62:289–321. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Audic Y and Hartley RS:
Post-transcriptional regulation in cancer. Biol Cell. 96:479–498.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yisraeli JK: VICKZ proteins: a
multi-talented family of regulatory RNA-binding proteins. Biol
Cell. 97:87–96. 2005. View Article : Google Scholar
|
|
6
|
Shu L, Yan W and Chen X: RNPC1, an
RNA-binding protein and a target of the p53 family, is required for
maintaining the stability of the basal and stress-induced p21
transcript. Genes Dev. 20:2961–2972. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cho SJ, Zhang J and Chen X: RNPC1
modulates the RNA-binding activity of, and cooperates with, HuR to
regulate p21 mRNA stability. Nucleic Acids Res. 38:2256–2267. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Miyamoto S, Hidaka K, Jin D and Morisaki
T: RNA-binding proteins Rbm38 and Rbm24 regulate myogenic
differentiation via p21-dependent and -independent regulatory
pathways. Genes Cells. 14:1241–1252. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yan W, Zhang J, Zhang Y, Jung YS and Chen
X: p73 expression is regulated by RNPC1, a target of the p53
family, via mRNA stability. Mol Cell Biol. 32:2336–2348. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yin T, Cho SJ and Chen X: RNPC1, an
RNA-binding protein and a p53 target, regulates macrophage
inhibitory cytokine-1 (MIC-1) expression through mRNA stability. J
Biol Chem. 288:23680–23686. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Zhang J, Jun Cho S and Chen X: RNPC1, an
RNA-binding protein and a target of the p53 family, regulates p63
expression through mRNA stability. Proc Natl Acad Sci USA.
107:9614–9619. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Xu E, Zhang J and Chen X: MDM2 expression
is repressed by the RNA-binding protein RNPC1 via mRNA stability.
Oncogene. 32:2169–2178. 2013. View Article : Google Scholar
|
|
13
|
Zhang J, Cho SJ, Shu L, Yan W, Guerrero T,
Kent M, Skorupski K, Chen H and Chen X: Translational repression of
p53 by RNPC1, a p53 target overexpressed in lymphomas. Genes Dev.
25:1528–1543. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zheng SL, Xu J, Isaacs SD, Wiley K, Chang
B, Bleecker ER, Walsh PC, Trent JM, Meyers DA and Isaacs WB:
Evidence for a prostate cancer linkage to chromosome 20 in 159
hereditary prostate cancer families. Hum Genet. 108:430–435. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Bar-Shira A, Pinthus JH, Rozovsky U,
Goldstein M, Sellers WR, Yaron Y, Eshhar Z and Orr-Urtreger A:
Multiple genes in human 20q13 chromosomal region are involved in an
advanced prostate cancer xenograft. Cancer Res. 62:6803–6807.
2002.PubMed/NCBI
|
|
16
|
Tanner MM, Grenman S, Koul A, Johannsson
O, Meltzer P, Pejovic T, Borg A and Isola JJ: Frequent
amplification of chromosomal region 20q12-q13 in ovarian cancer.
Clin Cancer Res. 6:1833–1839. 2000.PubMed/NCBI
|
|
17
|
Korn WM, Yasutake T, Kuo WL, Warren RS,
Collins C, Tomita M, Gray J and Waldman FM: Chromosome arm 20q
gains and other genomic alterations in colorectal cancer metastatic
to liver, as analyzed by comparative genomic hybridization and
fluorescence in situ hybridization. Genes Chromosomes Cancer.
25:82–90. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Knösel T, Schlüns K, Stein U, Schwabe H,
Schlag PM, Dietel M and Petersen I: Genetic imbalances with impact
on survival in colorectal cancer patients. Histopathology.
43:323–331. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Krackhardt AM, Witzens M, Harig S, Hodi
FS, Zauls AJ, Chessia M, Barrett P and Gribben JG: Identification
of tumor-associated antigens in chronic lymphocytic leukemia by
SEREX. Blood. 100:2123–2131. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Carvalho B, Postma C, Mongera S, Hopmans
E, Diskin S, van de Wiel MA, van Criekinge W, Thas O, Matthäi A,
Cuesta MA, et al: Multiple putative oncogenes at the chromosome 20q
amplicon contribute to colorectal adenoma to carcinoma progression.
Gut. 58:79–89. 2009. View Article : Google Scholar
|
|
21
|
Hötte GJ, Linam-Lennon N, Reynolds JV and
Maher SG: Radiation sensitivity of esophageal adenocarcinoma: the
contribution of the RNA-binding protein RNPC1 and p21-mediated cell
cycle arrest to radioresistance. Radiat Res. 177:272–279. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Ginestier C, Cervera N, Finetti P,
Esteyries S, Esterni B, Adélaïde J, Xerri L, Viens P, Jacquemier J,
Charafe-Jauffret E, et al: Prognosis and gene expression profiling
of 20q13-amplified breast cancers. Clin Cancer Res. 12:4533–4544.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Letessier A, Sircoulomb F, Ginestier C,
Cervera N, Monville F, Gelsi-Boyer V, Esterni B, Geneix J, Finetti
P, Zemmour C, et al: Frequency, prognostic impact, and subtype
association of 8p12, 8q24, 11q13, 12p13, 17q12, and 20q13
amplifications in breast cancers. BMC Cancer. 6:2452006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Xue JQ, Xia TS, Liang XQ, Zhou W, Cheng L,
Shi L, Wang Y and Ding Q: RNA-binding protein RNPC1: Acting as a
tumor suppressor in breast cancer. BMC Cancer. 14:3222014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Feldstein O, Ben-Hamo R, Bashari D, Efroni
S and Ginsberg D: RBM38 is a direct transcriptional target of E2F1
that limits E2F1-induced proliferation. Mol Cancer Res.
10:1169–1177. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Léveillé N, Elkon R, Davalos V, Manoharan
V, Hollingworth D, Oude Vrielink J, le Sage C, Melo CA, Horlings
HM, Wesseling J, et al: Selective inhibition of microRNA
accessibility by RBM38 is required for p53 activity. Nat Commun.
2:5132011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Yang L, Luo Y and Wei J: Integrative
genomic analyses on Ikaros and its expression related to solid
cancer prognosis. Oncol Rep. 24:571–577. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Yang L, Luo Y, Wei J and He S: Integrative
genomic analyses on IL28RA, the common receptor of
interferon-lambda1, -lambda2 and -lambda3. Int J Mol Med.
25:807–812. 2010.PubMed/NCBI
|
|
29
|
Yang L, Wei J and He S: Integrative
genomic analyses on interferon-lambdas and their roles in cancer
prediction. Int J Mol Med. 25:299–304. 2010.PubMed/NCBI
|
|
30
|
Yu H, Yuan J, Xiao C and Qin Y:
Integrative genomic analyses of recepteur d’origine nantais and its
prognostic value in cancer. Int J Mol Med. 31:1248–1254.
2013.PubMed/NCBI
|
|
31
|
Wang M, Wei X, Shi L, Chen B, Zhao G and
Yang H: Integrative genomic analyses of the histamine H1 receptor
and its role in cancer prediction. Int J Mol Med. 33:1019–1026.
2014.PubMed/NCBI
|
|
32
|
Wang B, Chen K, Xu W, Chen D, Tang W and
Xia TS: Integrative genomic analyses of secreted protein acidic and
rich in cysteine and its role in cancer prediction. Mol Med Rep.
10:1461–1468. 2014.PubMed/NCBI
|
|
33
|
Wang B, Xu W, Tan M, Xiao Y, Yang H and
Xia TS: Integrative genomic analyses of a novel cytokine,
interleukin-34 and its potential role in cancer prediction. Int J
Mol Med. 35:92–102. 2015.
|
|
34
|
Kumar S, Nei M, Dudley J and Tamura K:
MEGA: A biologist-centric software for evolutionary analysis of DNA
and protein sequences. Brief Bioinform. 9:299–306. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Yang Z: PAML: aA program package for
phylogenetic analysis by maximum likelihood. Comput Appl Biosci.
13:555–556. 1997.PubMed/NCBI
|
|
36
|
Yang Z, Nielsen R, Goldman N and Pedersen
AM: Codon-substitution models for heterogeneous selection pressure
at amino acid sites. Genetics. 155:431–449. 2000.PubMed/NCBI
|
|
37
|
Forbes SA, Bindal N, Bamford S, Cole C,
Kok CY, Beare D, Jia M, Shepherd R, Leung K, Menzies A, et al:
COSMIC: Mining complete cancer genomes in the Catalogue of Somatic
Mutations in Cancer. Nucleic Acids Res. 39:D945–D950. 2011.
View Article : Google Scholar :
|
|
38
|
Parkinson H, Sarkans U, Shojatalab M,
Abeygunawardena N, Contrino S, Coulson R, Farne A, Lara GG,
Holloway E, Kapushesky M, et al: ArrayExpress - a public repository
for microarray gene expression data at the EBI. Nucleic Acids Res.
33:D553–D555. 2005. View Article : Google Scholar
|
|
39
|
Mizuno H, Kitada K, Nakai K and Sarai A:
PrognoScan: a new database for meta-analysis of the prognostic
value of genes. BMC Med Genomics. 2:182009. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Xue JQ, Xia TS, Liang XQ, Zhou W, Cheng L,
Shi L, Wang Y and Ding Q: RNA-binding protein RNPC1: Acting as a
tumor suppressor in breast cancer. BMC Cancer. 14:3222014.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Renjie W and Haiqian L: MiR-132, miR-15a
and miR-16 synergistically inhibit pituitary tumor cell
proliferation, invasion and migration by targeting Sox5. Cancer
Lett. 356:568–578. 2015. View Article : Google Scholar
|
|
42
|
Pei XH, Lv XQ and Li HX: Sox5 induces
epithelial to mesenchymal transition by transactivation of Twist1.
Biochem Biophys Res Commun. 446:322–327. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Kang KA, Kim KC, Bae SC and Hyun JW:
Oxidative stress induces proliferation of colorectal cancer cells
by inhibiting RUNX3 and activating the Akt signaling pathway. Int J
Oncol. 43:1511–1516. 2013.PubMed/NCBI
|
|
44
|
Xu HW, Ren F, Yu YM and Cai CZ: Runx3
expression in lymph nodes with metastasis is associated with the
outcome of gastric cancer patients. Oncol Lett. 2:1275–1279.
2011.
|
|
45
|
Yu GP, Ji Y, Chen GQ, Huang B, Shen K, Wu
S and Shen ZY: Application of RUNX3 gene promoter methylation in
the diagnosis of non-small cell lung cancer. Oncol Lett. 3:159–162.
2012.PubMed/NCBI
|
|
46
|
Han YX and Liang DY: The role of the tumor
suppressor RUNX3 in giant cell tumor of the bone. Int J Oncol.
40:673–678. 2012.
|
|
47
|
Bian J, Li B, Zeng X, Hu H, Hong Y, Ouyang
H, Zhang X, Wang Z, Zhu H, Lei P, et al: Mutation of TGF-β receptor
II facilitates human bladder cancer progression through altered
TGF-β1 signaling pathway. Int J Oncol. 43:1549–1559.
2013.PubMed/NCBI
|
|
48
|
Liu KC, Lin BS, Zhao M, Wang KY and Lan
XP: Cutl1: A potential target for cancer therapy. Cell Signal.
25:349–354. 2013. View Article : Google Scholar
|
|
49
|
Claudius AK, Kankipati CS, Kilari RS,
Hassan S, Guest K, Russell ST, Perry CJ, Stark LA and Nicholl ID:
Identification of aspirin analogues that repress NF-κB signalling
and demonstrate anti-proliferative activity towards colorectal
cancer in vitro and in vivo. Oncol Rep. 32:1670–1680.
2014.PubMed/NCBI
|
|
50
|
Zhang J, Kou YB, Zhu JS, Chen WX and Li S:
Knockdown of HMGB1 inhibits growth and invasion of gastric cancer
cells through the NF-κB pathway in vitro and in vivo. Int J Oncol.
44:1268–1276. 2014.PubMed/NCBI
|
|
51
|
Guan Z, Ding C, Du Y, Zhang K, Zhu JN,
Zhang T, He D, Xu S, Wang X and Fan J: HAF drives the switch of
HIF-1α to HIF-2α by activating the NF-κB pathway, leading to
malignant behavior of T24 bladder cancer cells. Int J Oncol.
44:393–402. 2014.
|
|
52
|
Yu L, Mu Y, Sa N, Wang H and Xu W: Tumor
necrosis factor α induces epithelial-mesenchymal transition and
promotes metastasis via NF-κB signaling pathway-mediated TWIST
expression in hypopharyngeal cancer. Oncol Rep. 31:321–327.
2014.
|
|
53
|
Xue M, Li X, Wu W, Zhang S, Wu S, Li Z and
Chen W: Upregulation of long non-coding RNA urothelial carcinoma
associated 1 by CCAAT/enhancer binding protein α contributes to
bladder cancer cell growth and reduced apoptosis. Oncol Rep.
31:1993–2000. 2014.PubMed/NCBI
|
|
54
|
Weng W, Wang M, Xie S, Long Y, Li F, Sun
F, Yu Y and Li Z: YY1-C/EBPα-miR34a regulatory circuitry is
involved in renal cell carcinoma progression. Oncol Rep.
31:1921–1927. 2014.PubMed/NCBI
|
|
55
|
Wei W, Hu Z, Fu H, Tie Y, Zhang H, Wu Y
and Zheng X: MicroRNA-1 and microRNA-499 downregulate the
expression of the ets1 proto-oncogene in HepG2 cells. Oncol Rep.
28:701–706. 2012.PubMed/NCBI
|
|
56
|
Shaikhibrahim Z and Wernert N: ETS
transcription factors and prostate cancer: The role of the family
prototype ETS-1 (Review). Int J Oncol. 40:1748–1754.
2012.PubMed/NCBI
|
|
57
|
Metzeler KH, Hummel M, Bloomfield CD,
Spiekermann K, Braess J, Sauerland MC, Heinecke A, Radmacher M,
Marcucci G, Whitman SP, et al Cancer and Leukemia Group B; German
AML Cooperative Group: An 86-probe-set gene-expression signature
predicts survival in cytogenetically normal acute myeloid leukemia.
Blood. 112:4193–4201. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Phillips HS, Kharbanda S, Chen R, Forrest
WF, Soriano RH, Wu TD, Misra A, Nigro JM, Colman H, Soroceanu L, et
al: Molecular subclasses of high-grade glioma predict prognosis,
delineate a pattern of disease progression, and resemble stages in
neurogenesis. Cancer Cell. 9:157–173. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Anders CK, Acharya CR, Hsu DS, Broadwater
G, Garman K, Foekens JA, Zhang Y, Wang Y, Marcom K, Marks JR, et
al: Age-specific differences in oncogenic pathway deregulation seen
in human breast tumors. PLoS One. 3:e13732008. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Schmidt M, Böhm D, von Törne C, Steiner E,
Puhl A, Pilch H, Lehr HA, Hengstler JG, Kölbl H and Gehrmann M: The
humoral immune system has a key prognostic impact in node-negative
breast cancer. Cancer Res. 68:5405–5413. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Jain AN, Chin K, Børresen-Dale AL,
Erikstein BK, Eynstein Lonning P, Kaaresen R and Gray JW:
Quantitative analysis of chromosomal CGH in human breast tumors
associates copy number abnormalities with p53 status and patient
survival. Proc Natl Acad Sci USA. 98:7952–7957. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Smith JJ, Deane NG, Wu F, Merchant NB,
Zhang B, Jiang A, Lu P, Johnson JC, Schmidt C, Bailey CE, et al:
Experimentally derived metastasis gene expression profile predicts
recurrence and death in patients with colon cancer.
Gastroenterology. 138:958–968. 2010. View Article : Google Scholar
|
|
63
|
Laurent C, Valet F, Planque N, Silveri L,
Maacha S, Anezo O, Hupe P, Plancher C, Reyes C, Albaud B, et al:
High PTP4A3 phosphatase expression correlates with metastatic risk
in uveal melanoma patients. Cancer Res. 71:666–674. 2011.
View Article : Google Scholar
|
|
64
|
Director’s Challenge Consortium for the
Molecular Classification of Lung Adenocarcinoma; Shedden K, Taylor
JM, Enkemann SA, Tsao MS, Yeatman TJ, Gerald WL, Eschrich S,
Jurisica I, Giordano TJ, Misek DE, et al: Gene expression-based
survival prediction in lung adenocarcinoma: a multi-site, blinded
validation study. Nat Med. 14:822–827. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Hammerman PS, Lawrence MS, Voet D, Jing R,
Cibulskis K, Sivachenko A, Stojanov P, McKenna A, Lander ES,
Gabriel S, et al Cancer Genome Atlas Research Network:
Comprehensive genomic characterization of squamous cell lung
cancers. Nature. 489:519–525. 2012. View Article : Google Scholar
|
|
66
|
Yamauchi M, Yamaguchi R, Nakata A, Kohno
T, Nagasaki M, Shimamura T, Imoto S, Saito A, Ueno K, Hatanaka Y,
et al: Epidermal growth factor receptor tyrosine kinase defines
critical prognostic genes of stage I lung adenocarcinoma. PLoS One.
7:e439232012. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Tomida S, Koshikawa K, Yatabe Y, Harano T,
Ogura N, Mitsudomi T, Some M, Yanagisawa K and Takahashi T, Osada H
and Takahashi T: Gene expression-based, individualized outcome
prediction for surgically treated lung cancer patients. Oncogene.
23:5360–5370. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Bild AH, Yao G, Chang JT, Wang Q, Potti A,
Chasse D, Joshi MB, Harpole D, Lancaster JM, Berchuck A, et al:
Oncogenic pathway signatures in human cancers as a guide to
targeted therapies. Nature. 439:353–357. 2006. View Article : Google Scholar
|