Introduction
Coiled-coil domain containing 134 (CCDC134) was
first identified through high-throughput functional screening
systems using an Elk1 trans-reporting system in the Peking
University Center for Human Disease Genomics (Beijing, China). Our
earlier study demonstrated that CCDC134 is a classical secreted
protein that inhibited Elk1 transcriptional regulation and
mitogen-activated protein kinase (MAPK) signal transduction through
the Raf-1/MEK/extracellular signal-regulated kinase and c-Jun
N-terminal kinase/stress-activated protein kinase pathways
(1). A previous study also
identified a role for CCDC134 in tumor development; CCDC134 was
identified as a candidate biomarker of malignant transformation
with decreased expression in gastric cancer, and targeted small
interfering RNA knockdown of CCDC134 promoted tumor migration and
invasion via the MAPK pathway (2).
CCDC134 was proposed to have immune cytokine
function, and directly promoted CD8+ T-cell activation,
proliferation and cytotoxicity (3). Additionally, CCDC134 demonstrated
its potent antitumor effects by augmenting CD8+
T-cell-mediated immunity (3).
Mechanistically, exposure to CCDC134 promoted CD8+
T-cell proliferation through the Janus kinase 3-signal transducer
and activator of transcription 5 pathway, and two members of the
γc cytokine family could effectively block CCDC134
binding to activated CD8+ T-cells, which provided
evidence that CCDC134 may serve as a potential member of the
γc cytokine family (3).
On the basis of CCDC134 molecular structure and
transcription regulatory capacity, it is suggested that CCDC134 is
also a nuclear protein that acts as a critical regulator of human
alteration/deficiency in activation 2a (hADA2a) to enhance the
stability of hADA2a and inhibit its proteasome-dependent
degradation (4). Additionally,
CCDC134 participated in the p300-CBP-associated factor (PCAF)
complex via hADA2a to affect its histone acetyltransferase (HAT)
activity, which primarily acetylated lysine 14 of H3, but also less
efficiently acetylated lysine 8 of H4 (5). Also, CCDC134 was involved in the
repair of ultraviolet-induced DNA damage (4). The above evidence indicates that
CCDC134 may function as a cytokine that mediates immune responses
and a nuclear protein, similar to high mobility group box
chromosomal protein 1, interleukin-1α (IL-1α) and IL-33 (6,7).
Thus, CCDC134 may serve as a multi-faceted
adaptor/scaffolding protein to relay cellular signals to the
cytoplasm and the nucleus. To determine if any of these cellular
mechanisms for CCDC134 may be biologically relevant and significant
in vivo, the present study generated a murine model carrying
a conditional allele for Ccdc134. The present study reports
the generation and first characterization of a germline
Ccdc134 null mutant allele in mice, which, when homozygous,
is embryonic lethal and may impair embryonic angiogenesis.
Materials and methods
Targeting the Ccdc134 allele
The Ccdc134 targeting vector was generated
with two flippase recognition target (FRT) sites flanked by a
neomycin (neo)-resistant cassette upstream of the existing 3′ loxP
site, and the 5′ loxP site was inserted into upstream exon 3, by
Nanjing Biomedical Research Institute of Nanjing University
(Nanjing, China), according to well-described principles and
methods (8). The targeting region
of the recombination vector and its relation with the
Ccdc134 locus are demonstrated in Fig. 1A. The targeting vector was
linearized with NotI (New England BioLabs, Inc., Ipswich,
MA, USA) and electroporated into B6/BLU embryonic stem (ES) cells.
G418-resistant clones were screened for homologous recombination by
long range polymerase chain reaction (PCR). Out of 64 clones, 4
were further identified with correct targeting by Southern blot
analysis.
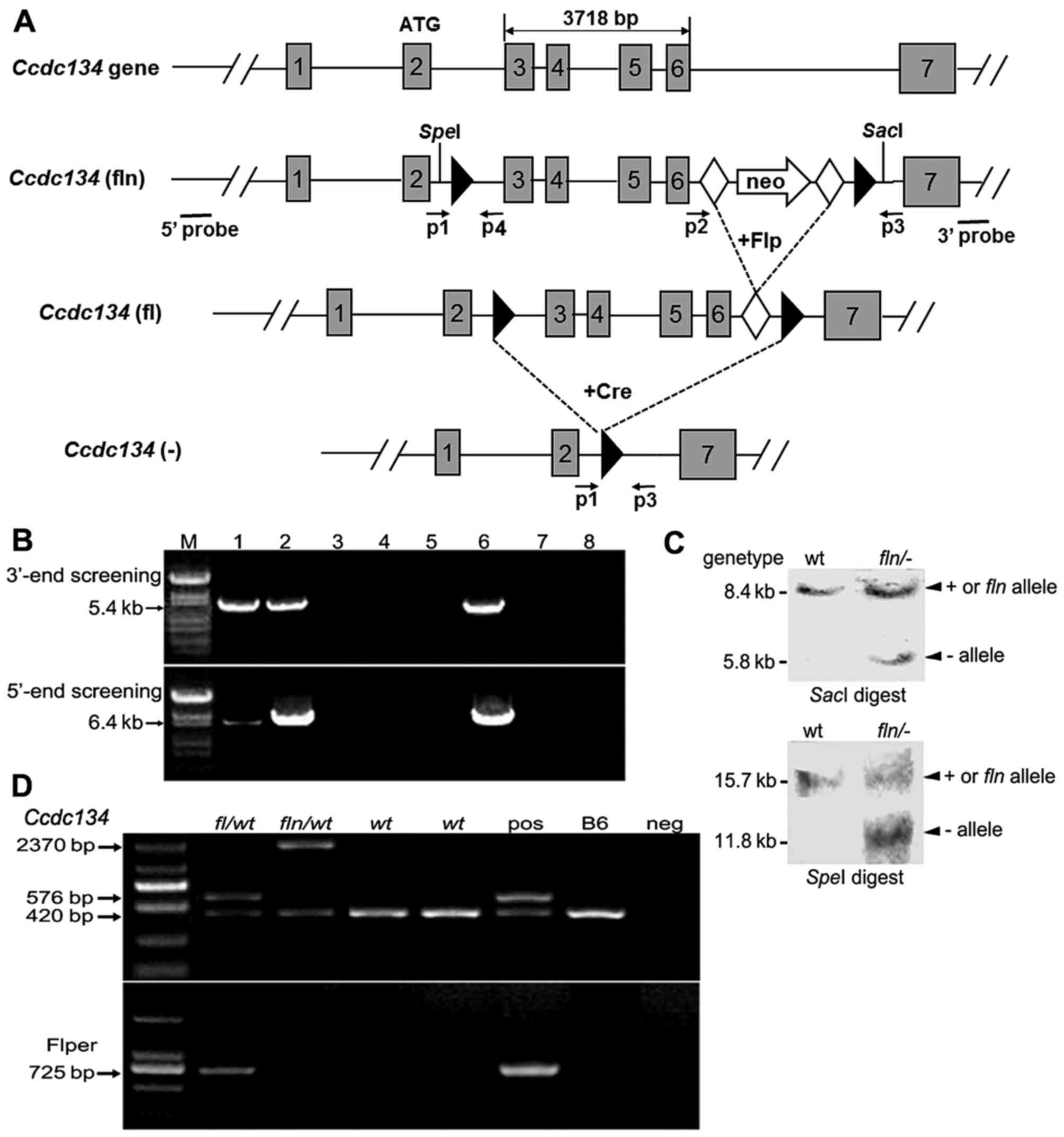 | Figure 1Conditional targeting strategy and
screening for homologous recombination of the Ccdc134 gene.
(A) Schematic diagram of the Ccdc134 gene targeting strategy
and gene deletion in mutant mice. The targeting construct was made
by inserting a neo resistance gene flanked by flippase recognition
target sites (closed diamond) between exon 6 and 7. LoxP sites
(closed triangles) were placed flanking exon 3–6. The neo cassette
was excised by crossing with FLP-expressing transgenic mice to
generate the floxed allele. 5′ and 3′ probes outside the homologous
region were used for Southern blot analysis. (B) Examples of PCR
screening assays of neo-resistant clones (lanes 1–5) following
transfection of linearized targeting construct and drug selection.
Each clone was amplified separately for the recombinant allele (5.4
and 6.4 kb, as predicted) using specific primers for 3′-and 5′-end
screening by long range PCR. Here, clones 1 and 2 (lanes 1 and 2)
were identified as positive. In the agarose gel: Lane 1–5,
amplification using DNA template from clone 1–5; lane 6, positive
control; lane 7, control mouse DNA; lane 8, control mix, no DNA.
(C) Representative Southern blot analysis used to confirm correct
targeting. DNA was digested with either SpeI or SacI
restriction enzymes and the 5′ or 3′ probe was used to detect wt,
fln and - alleles at the expected size: wt allele, 15.7
(SpeI) or 8.4 kb (SacI); and fln, 11.8 (SpeI)
or 5.8 kb (SacI). (D) A representative gel image of the PCR
products. The wt and targeted alleles without neo produce products
of 420 and 576 bp, respectively. Also, fl/wt was found to
lack neo but contain FLP to produce a 725-bp fragment, whereas the
PCR product of fln/wt should be 2,370 bp. Ccdc134,
coiled-coil domain containing 134; FLP, flippase in the recombinant
embryonic stem cells; neo, neomycin; PCR, polymerase chain
reaction; wt, wild-type; fl, floxed; fln, floxed allele with
neomycin cassette; +, wild-type; −, null; pos, positive control;
neg, negative control; M, marker. |
Subsequently, injection of two targeted ES cells
into B6 recipient blastocysts (obtained from C57BL/6 female mice)
produced 2 male and 2 female chimeras that, on crossing with B6
mice (6–8-week-old male or female mice; weight, 18–20 g) purchased
from Beijing Vital River Laboratory Animal Technology Co., Ltd.
(Beijing, China), showed germline transmission. Male chimeras were
mated with C57BL/6 females in the ratio of 1:3 to generate the
Ccdc134fln/+ mouse line. Neo cassette was removed
by crossing with FLP-expressing transgenic mice to generate a
Ccdc134fl/+ mouse line.
Ccdc134fl/+ mice were intercrossed to generate a
Ccdc134fl/fl conditional allele line, and
maintained as a homozygous stock. Ccdc134fl/fl
females were bred with human β-actin (ACTB)-Cre or zona pellucida 3
(ZP3)-Cre mice to generate a Ccdc134+/−
heterozygous knockout (KO) mouse line. Finally,
Ccdc134+/− mice were intercrossed to generate
Ccdc134+/+, Ccdc134+/− and
Ccdc134−/−embryos used for experiments.
The mice were housed and bred under pathogen-free
conditions at the Laboratory Animal Research Facility of Peking
University Health Science Center (Beijing, China). A total of 5
mice/cage were maintained under laboratory conditions at 25°C,
under a normal 12-h light/dark cycle with a humidity of 55% and
access to food and water ad libitum. Experimental procedures
were approved by the Institutional Animal Care and Use Committee of
Peking University Health Science Center (Beijing, China), following
the guidelines of the Care and Use of Laboratory Animals.
Genotypic analyses
DNA was extracted from ES cells using a genomic DNA
extraction kit (Qiagen China Co., Ltd., Shanghai, China) following
the manufacturer's protocol. Screening of ES cells was performed by
long range PCR analysis for 3′-end screening using a targeting
vector-specific forward primer (5′-GCATCGCATTGTCTGAGTAGGTG-3′) and
a reverse primer (5′-TCTTGCAGAGCAAGAGCGAG-3′) inside the targeted
region. In addition, a second pair of primers specific for 5′-end
screening outside of the target region was used forward,
5′-AACCTCACCCACTCTCTCACCG-3′ and reverse,
5′-AAGGGTTATTGAATATGATCGGA-3′. PCR analysis was performed using a
Takara LA Taq long PCR system (Takara Biomedical Technology Co.,
Ltd., Beijing, China) using conditions as follows: Denaturation at
94°C for 5 min; followed by 30 cycles of amplification at 94°C for
1 min, 55°C for 1 min and 65°C for 5 min; and a final extension
step at 72°C for 10 min.
High molecular weight genomic DNA was extracted from
ES cells, digested with either SpeI or SacI (New
England BioLabs, Inc.) and subjected to electrophoresis in 0.9%
agarose gel. For Southern blot analysis, genomic DNA was
transferred to a nylon membrane overnight, and then hybridized
overnight at 42°C using a 5′ external probe with
SpeI-digested DNA or 3′ external probe with
SacI-digested DNA specific to the Ccdc134 sequence
with 32P (GE Healthcare, Chicago, IL, USA) by a random
prime labeling method. Finally, the blot was monitored with
radioautograph to confirm homologous recombination.
Genomic DNA from tails of 3-wcditions as follows:
Denaturation at 94°C for 5 min; 35 cycles of amplification at 94°C
for 30 sec, 55°C for 30 sec and 72°C for 30 sec; and a final
extension step at 72°C for 5 min. Two types of Ccdc134 PCR
primers used for PCR analysis were as follows: p1,
5′-TCCTAACCCTGTCGCTCCCT-3′; p2, 5′-CCAGACAGAGGTGAGCTGCT-3′; p3,
5′-GCACCCTGAGCCAAGTTTAG-3′; and p4, 5′-CCTAACCTATGCCTCCAAAG-3′.
Genomic DNA from the targeted allele yields a 615-bp fragment with
a primer pair p1/p3, and a wild-type allele yields a 420-bp
fragment with primer pair p2/p3, or a 490-bp fragment with
p1/p4.
Quantification of mRNA by reverse
transcription-quantitative PCR (RT-qPCR)
Total RNA was extracted from
Ccdc134+/+, Ccdc134+/− and
Ccdc134−/− embryos at different stages with
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.,
Waltham, MA, USA), following the manufacturer's protocol. cDNA was
synthesized using a Revert Aid First Strand cDNA synthesis kit
(Fermentas; Thermo Fisher Scientific, Inc., Pittsburgh, PA, USA),
following the manufacturer's recommendations. The following PCR
cycling conditions were used: Denaturation at 94°C for 3 min; 35
cycles of amplification at 94°C for 30 sec, 55°C for 30 sec and
72°C for 30 sec; and a final extension step at 72°C for 5 min.
Ccdc134 primer sequences used for RT were as follows F1,
5′-GTTGGCACTGAAGAACCTGG-3′ and R1, 5′-ACGGGTTCCGGAAGTCAGAA-3′. The
qPCR analysis using SYBR-Green master mix was performed using an
ABI 7500 Real-Time PCR system (both from Applied Biosystems; Thermo
Fisher Scientific, Inc., Waltham, MA, USA) according to the
manufacturer's protocol (9). The
following primer sequences were used: Ccdc134 forward,
5′-GCTCCCTTCTCCCTGCAC-3′ and reverse, 5′-AGGCCACAGGAGGACAGA-3′; and
glyceraldehyde 3-phosphate dehydrogenase (GAPDH) forward,
5′-AAGAGGGATGCTGCCCTTAC-3′ and reverse, 5′-CCATTTTGTCTACGGGACGA-3′.
The mRNA expression levels of Ccdc134 were normalized to
GAPDH. All samples were assayed in duplicate, and average
values were used for quantification (9).
Western blot analysis
Whole embryos were homogenized with a Dounce glass
homogenizer (Kimble Glass, Inc., Deerfield, IL, USA) in
radioimmunoprecipitation assay lysis buffer containing fresh
protease inhibitor cocktail (Roche Applied Science, Penzberg,
Germany). Protein concentrations were determined using a BCA
Protein assay kit (Pierce; Thermo Fisher Scientific, Inc.,
Rockford, IL, USA). The cell lysates (100 μg) were separated
by 12.5% SDS-PAGE, transferred to a nitrocellulose membrane and
blocked with 5% non-fat milk for 1 h at room temperature. The
membrane was probed with rabbit anti-CCDC134 polyclonal antibody
(1:500; sc-86363; Santa Cruz Biotechnology, Inc., Dallas, TX, USA)
and mouse anti-β-actin monoclonal antibody (1:1,000; TA09; OriGene
Technologies, Inc., Beijing, China) at 4°C overnight. Subsequently,
the membrane was incubated at room temperature for 1 h using
DyLight 780-conjugated secondary antibodies (1:5,000; 53064; Thermo
Fisher Scientific, Inc., Waltham, MA, USA). The protein bands were
visualized and an infrared fluorescence image was obtained using an
Odyssey infrared imaging system (LI-COR Biosciences, Lincoln, NE,
USA).
Histology and immunohistochemistry
analysis
Whole embryos at 13.5 days post coitus (E13.5) were
fixed in 10% formaldehyde solution at room temperature for over 24
h, embedded in paraffin, sectioned at 5-μm thickness,
stained with hematoxylin for 4 min, and stained with eosin for 2
min. All procedures were performed at room temperature for
histologic examination. Images were captured on a BX-53 inverted
fluorescence microscope (Olympus Corp., Tokyo, Japan) at different
magnifications (×2, ×10 and ×20), and processed with Adobe
Photoshop CS 5.0 (Adobe, San Jose, CA, USA).
For immunohistochemistry staining, slides were
deparaffinized, hydrated, and boiled in a steamer for 3 min in 0.01
M sodium citrate buffer for antigen retrieval. Sections were first
treated with 3% H2O2 at room temperature to
quench endogenous peroxidase, washed several times with
phosphate-buffered saline (PBS) (pH 7.2), blocked with 10% normal
goat or rabbit serum (OriGene Technologies, Inc.) at 37°C for 1 h,
and then incubated with primary antibodies at 4°C overnight. The
following antibodies were used: Rabbit anti-CCDC134 (1:100;
sc-86363; Santa Cruz Biotechnology, Inc.), rabbit anti-vascular
endothelial growth factor receptor-2 (Vegfr-2; 1:100; ab2349;
Abcam, Cambridge, UK), biotin-labeled anti-Ter119 (1:100;
13-5921-81; eBioscience; Thermo Fisher Scientific, Inc., Waltham,
MA, USA), and anti-Ki-67 (1:50; ab16667). After thorough washing, a
GTVision™ III detection system/Mo&Rb HRP (GK500705; GeneTech
Co., Ltd., Shanghai, China) was applied directly for 30 min at room
temperature. After rinsing in PBS, all sections were visualized
with 0.05% 3,3′-diaminobenzidine. The sections were then
counterstained with hematoxylin at room temperature for 4 min. To
quantify proliferating cells, total numbers of Ki-67+
nuclei were counted and data presented as Ki-67+
nuclei/mm2. Fetal brains were stained for Vegfr-2
expression to quantify the vascular density. Images were captured
on a BX-53 inverted fluorescence microscope (Olympus Corp.) at
different magnifications (×2, ×10 and ×20), and processed with
Adobe Photoshop CS 5.0 (Adobe).
Statistical analysis
All data were expressed as the mean ± standard
deviation. The differences among groups were analyzed using one-way
analysis of variance followed by Bonferroni correction. Statistical
analyses were performed using SPSS 11.0 (SPSS, Inc., Chicago, IL,
USA). P<0.05 was considered to indicate a statistically
significant difference.
Results
Generation of floxed Ccdc134 allele and
mice
To investigate further the biological role of
Ccdc134 in vivo, the present study attempted to generate
conditional Ccdc134-null mice using a Cre/loxP strategy. A
homologous targeting construct was prepared with the two loxP sites
flanking Ccdc134 exon 3–6, as well as a neo resistance
cassette (a positive selection marker) within intron 6, flanked by
FRT sites (Fig. 1A). Upon
transfection of ES cells with the linearized targeting vector and
G418 selection, 64 independent drug-resistant clones were selected
and screened for homologous recombination by long range PCR
analyses. The 5.4-kb fragment was amplified from the targeted
allelic genomic DNA with a primer pair for 3′-end screening,
whereas the 6.4-kb fragment was amplified with a primer pair for
5′-end screening (Fig. 1B). A
total of 17 clones were identified as potentially homologous
targeted lines. Additionally, 4 correctly targeted clones were
identified by Southern blot analysis (Fig. 1C).
To generate chimeras, two ES cell lines were
injected into blastocysts of C57BL/6J mice. Male offspring with a
high degree of chimerism were crossed with C57BL/6J females to
generate floxed Ccdc134 mice
(Ccdc134fln/wt). The neo locus was then removed
by crossing with FLP-expressing transgenic mice (10,11). Ccdc134 conditional allele
mice (Ccdc134fl/fl) were then generated by
intercrossing Ccdc134fl/wt mice, and their
genotyping was performed by PCR analyses using tail-derived DNA
(Fig. 1D). The transmission of
Ccdc134wt/wt, Ccdc134fl/wt and
Ccdc134fl/fl followed a Mendelian ratio, and
homozygous flox mice exhibited wild-type characteristics with
normal CCDC134 expression, reproductive capability and lifespan
(data not shown), suggesting that the flox alleles do not influence
Ccdc134 gene activity.
Ccdc134 deficiency is embryonically
lethal
The Ccdc134fl/fl mice were crossed
with ACTB-Cre transgenics, which expressed Cre recombinase under
the control of the ACTB gene promoter in all cells of the embryo by
the blastocyst stage of development, to generate Ccdc134
constitutive KOs, referred to as
ACTB-Cre-Ccdc134−/− (Fig. 1A) (12). Genotyping of progeny from
intercrossed Ccdc134 hetero-zygote
(ACTB-Cre-Ccdc134+/−) demonstrated that, among
284 pups born, 185 (65%) were heterozygous for a null allele and 99
(35%) were wild-type (Ccdc134+/+). No homozygous
mice were born, while heterozygous mice were present at the
expected Mendelian ratio (Table
I). Subsequently, another Cre line, the Zp3-Cre transgenic
line, was used in an attempt to delete Ccdc134 at an earlier
stage in embryonic development. Unlike ACTB, Zp3 is expressed in
the growing oocyte prior to the completion of the first meiotic
division (13). The
Ccdc134 heterozygotes (Zp3-Cre-Ccdc134+/−)
were generated by crossing female
Zp3-Cre-Ccdc134fl/wt mice with male wild-type.
Crosses between these heterozygous mice also delivered only
heterozygous (128) or wild-type (64) live pups in the ratio of 2:1,
consistent with prenatal lethality of
ACTB-Cre-Ccdc134−/− embryos (Table I). A representative genotype is
demonstrated in Fig. 2A. The
complete absence of Ccdc134 homozygous KO mice is a
statistically significant deviation from the expected ratio
(P<0.0001; data not shown), suggesting that a homozygous
Ccdc134-null genotype is embryonically lethal. Timed mating
indicated that mortality of Ccdc134−/− embryos
began between E11.5 and E12.5. The rate of mortality of
Ccdc134−/− embryos increased after E11.5 (0%
until E11.5, 16.67% at E12.5, 42.86% at E13.5 and 100% at E14.5)
and mortality was observed in all Ccdc134−/− mice
at E14.5 (Table I). These data
indicate that Ccdc134 deficiency causes embryonic lethality,
supporting a crucial role for Ccdc134 during
embryogenesis.
 | Table IGenotypes of embryos from timed
pregnancies. |
Table I
Genotypes of embryos from timed
pregnancies.
| Stage | Total | Embryos in each
genotype, n
|
|---|
|
Ccdc134+/+ |
Ccdc134+/− |
Ccdc134−/− |
|---|
| Weaning (human
β-actin-Cre) | 284 | 99 | 185 | 0 |
| Weaning (zona
pellucida 3-Cre) | 192 | 64 | 128 | 0 |
| E14.5 | 33 | 10 | 18 | 5 (5) |
| E13.5 | 54 | 13 | 27 | 14 (6) |
| E12.5 | 39 | 8 | 25 | 6 (1) |
| E11.5 | 32 | 10 | 17 | 5 |
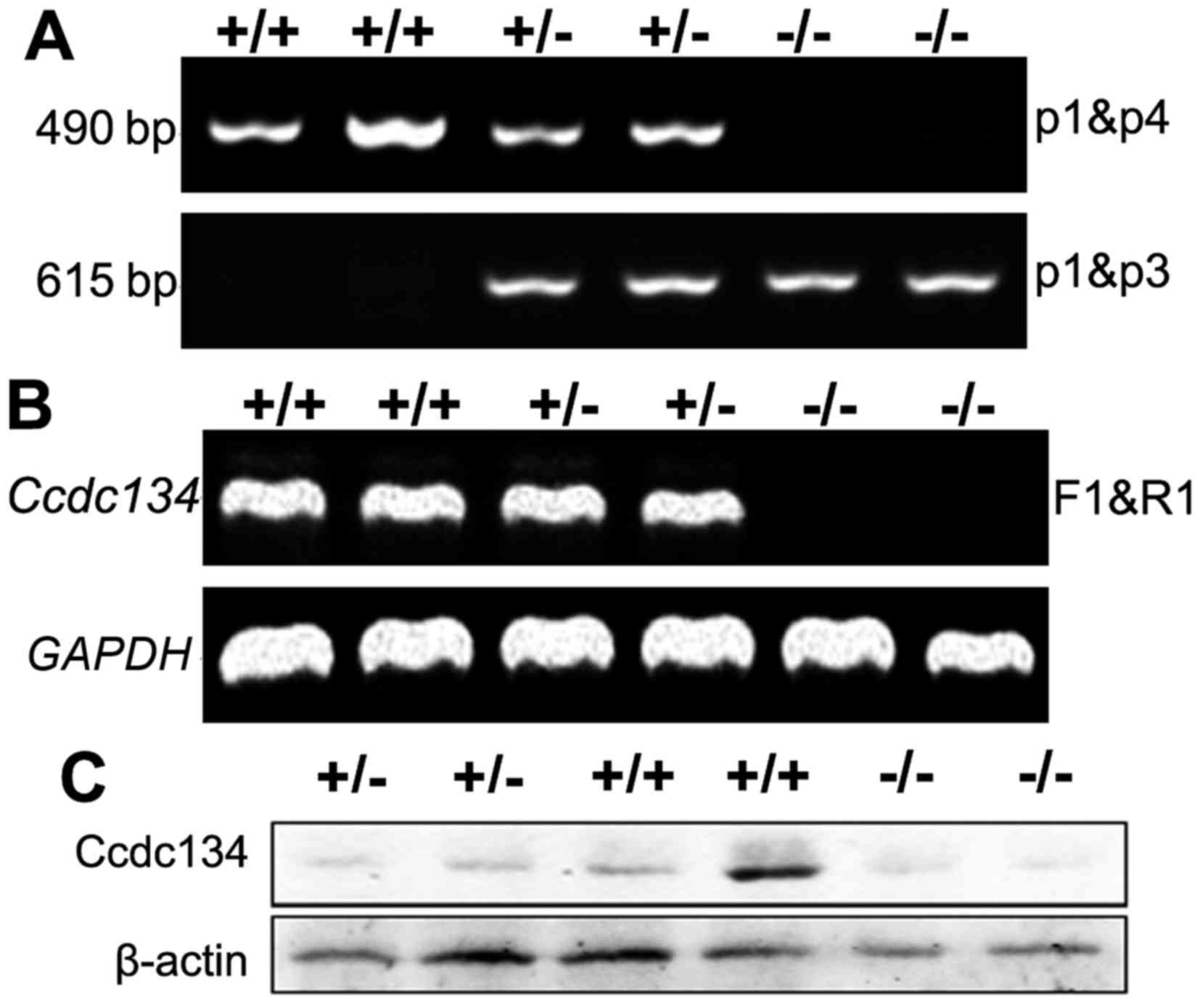
To confirm the inability of the targeted
Ccdc134 allele to support CCDC134 expression, the wild-type,
heterozygous and KO mice embryos at E13.5 were prepared and
analyzed by RT-PCR and western blot analysis. Ccdc134 mRNA
were not detected in Ccdc134−/− mice (Fig. 2B), and no CCDC134 protein was
detected with the use of a specific antibody to CCDC134 (Fig. 2C).
However, Zp3-Cre-Ccdc134+/− mice
appeared to be grossly normal. These heterozygotes bred without
difficulty and delivered normal sized pups. No histologic deficits
were observed. No apparent anatomic or microscopic features could
reliably discriminate heterozygotes from their wild-type
littermates. Previous study revealed that CCDC134 illustrated
potent antitumor effects by augmenting CD8+
T-cell-mediated immunity (3),
therefore different immune cell populations were analyzed in the
spleen and thymus. The results indicated no obvious difference
between heterozygotes and their wild-type littermates.
Additionally, B16 graft tumors were established in
Ccdc134+/− and Ccdc134+/+
controls, and it was demonstrated that Ccdc134+/−
mice may slightly accelerate tumor growth compared with wild-type
controls; however, no significant difference was observed (data not
shown). These data prompted us to restrict our efforts to the
comparison of the wild-type and Ccdc134−/−
populations, to define rigorously the mutant phenotype without
consideration of subtle or dose-dependent deficits.
Dynamic expression analyses of Ccdc134 in
whole embryos during embryogenesis
To determine whether Ccdc134 expression was
altered at various developmental stages of the embryo, the mRNA
expression level of Ccdc134 at four developmental stages was
compared. The expression level of Ccdc134 decreased from
E6.5 to E9.5, followed by an increasing trend from E9.5 to E12.5,
and then a decrease at E14.5. Thus, Ccdc134 mRNA showed the
highest expression level at E12.5 (Fig. 3A).
To gain further insight into the spatial and
temporal expression patterns of CCDC134 during embryonic
development, whole mount embryo sections were analyzed by
immunohistochemistry using specific rabbit anti-CCDC134 antibody.
This revealed histological details of CCDC134 expression in the
developing mouse embryo. At E6.5, the most prominent expression was
observed in the Reichert's membrane and endoderm. Furthermore,
CCDC134 expression was detected in the neural tube and
trophoblastic giant cells at E9.5. With the development of the
mouse embryo, prominent expression of CCDC134 was detected in the
somite and major organs, including the liver and lung, from E12.5
to E14.5 (Fig. 3B). At these
stages, CCDC134 was highly expressed in the somite and liver,
suggesting that CCDC134 participated in the development and
formation of major organs.
Morphological and histological pathology
of Ccdc134−/− embryos
To assess the role of CCDC134 throughout
development, Ccdc134+/+,
Ccdc134+/− and Ccdc134−/−
embryos were euthanized at E10.5–E16.5. Necropsies through this
interval demonstrated that Ccdc134−/− embryos
appeared similar to Ccdc134+/+ and
Ccdc134+/− littermates at E10.5, while
Ccdc134−/− embryos exhibited severe hemorrhaging
(100% penetrance) in the brain ventricular space and neural tube at
E13.5. Additionally, Ccdc134−/− embryos later
than E13.5 looked anemic, and their size was slightly smaller than
those of wild-type embryos (Fig.
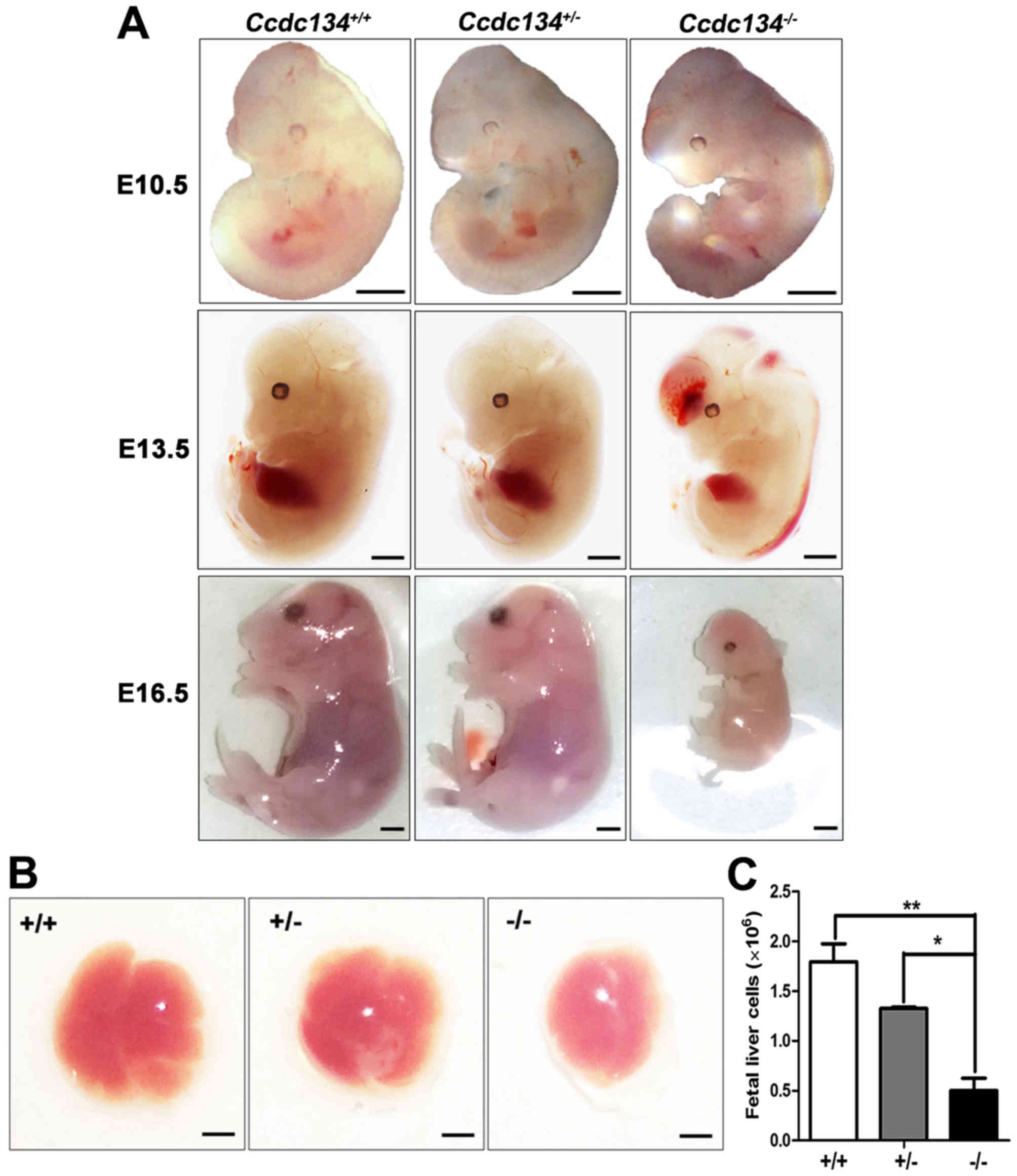
4A). Subsequently, embryos at E16.5 were isolated, and the
absorbed embryos with genotypes Ccdc134−/− were
noted. Thus, these data further suggest that Ccdc134
deletion causes embryonic lethality. In addition, in contrast to
Ccdc134+/+ and Ccdc134+/−
embryos whose red-brown livers invariably filled a large portion of
the abdomen, the livers of Ccdc134−/− embryos
were considerably smaller (Fig.
4B). The total cell number per fetal liver was
~2×106 in Ccdc134+/+ livers and
~5×105 in Ccdc134−/− livers,
suggesting that Ccdc134 deficiency affects the number of
fetal liver cells and may lead to a defect in hematopoiesis
(Fig. 4C).
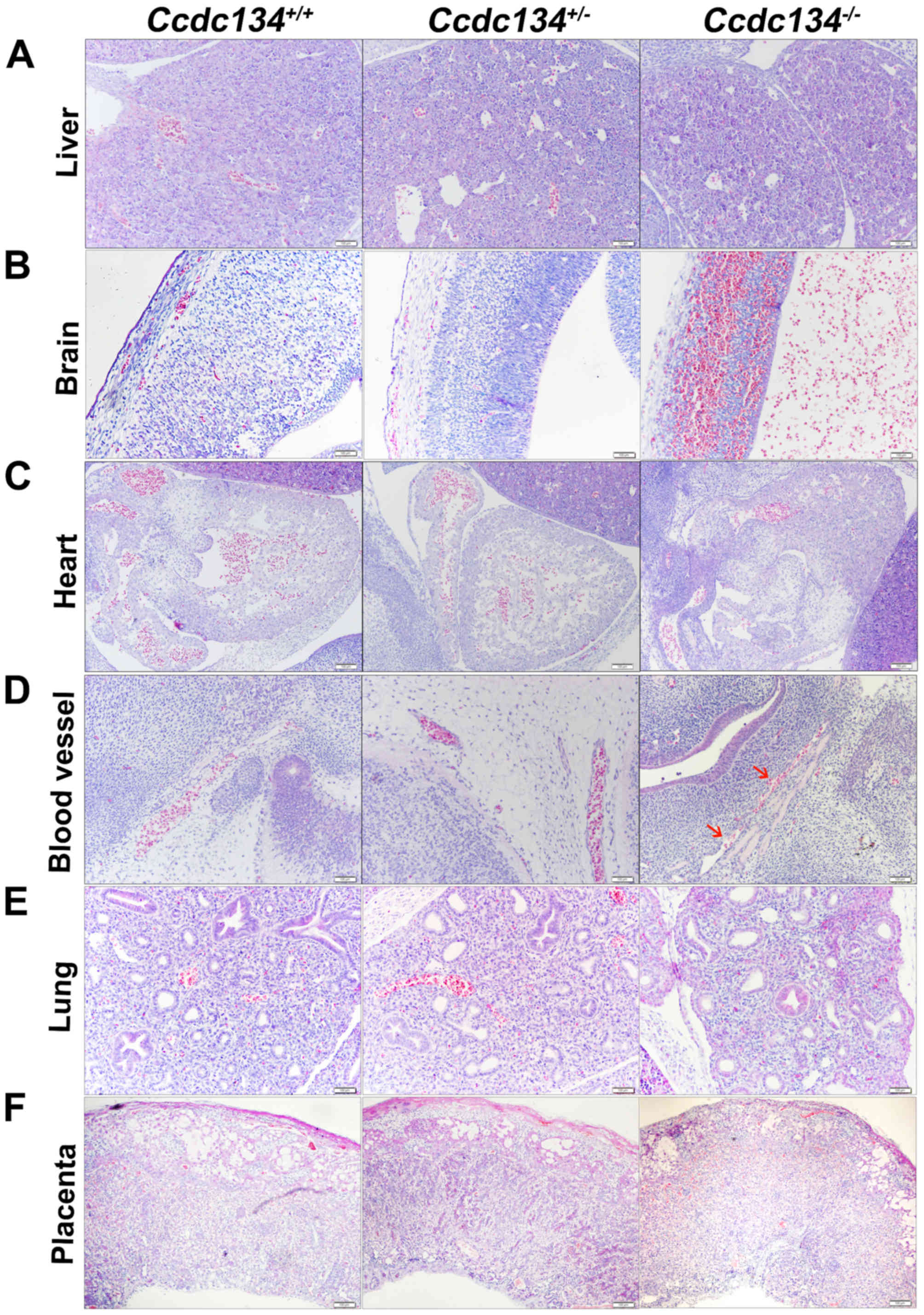
Whole-mount histologic sections of
Ccdc134−/− embryos demonstrated liver
abnormalities in later stage embryos. The fetal livers were
markedly smaller and hypoplastic (Fig. 5A). Additionally, compared with
their littermates, severe hemorrhage and overall brain
disorganization was detected in the Ccdc134−/−
embryo brain ventricular space (Fig.
5B). Concentric hypertrophy of the cardiac ventricular wall was
also present in Ccdc134−/− embryos, suggesting
increased vascular resistance (Fig.
5C). Furthermore, vasculature malformation was also noted,
although blood cells present in Ccdc134−/−
embryos appeared morphologically normal (Fig. 5D). However, the lungs of
Ccdc134−/− embryos and littermate controls
presented normal branching of the bronchioalveolar tree, with
progressive thinning of the alveolocapillary membrane and
flattening of the terminal sac epithelium (Fig. 5E). Structural changes in the
placenta, leading to altered hemodynamics or surface area available
for nutrient exchange, have been demonstrated to result in
reductions in growth, heart defects and perinatal morbidity
(14) (Fig. 5F). Given this, whether loss of
Ccdc134 altered the morphology of the placenta was
investigated. The placenta of Ccdc134−/− embryos
was thinner and poorly developed, and demonstrated a prominence of
labyrinth trophoblasts, which were arranged into poorly formed
maternal vascular spaces; however, the thin-walled capillary bed of
fetal circulation was ill defined (Fig. 5F).
Lethality due to reduced cell
proliferation and vascular defects in Ccdc134−/−
embryos
To clarify the causes of the liver abnormality in
Ccdc134−/− embryos, immunohistochemical analysis
was performed in transverse sections from
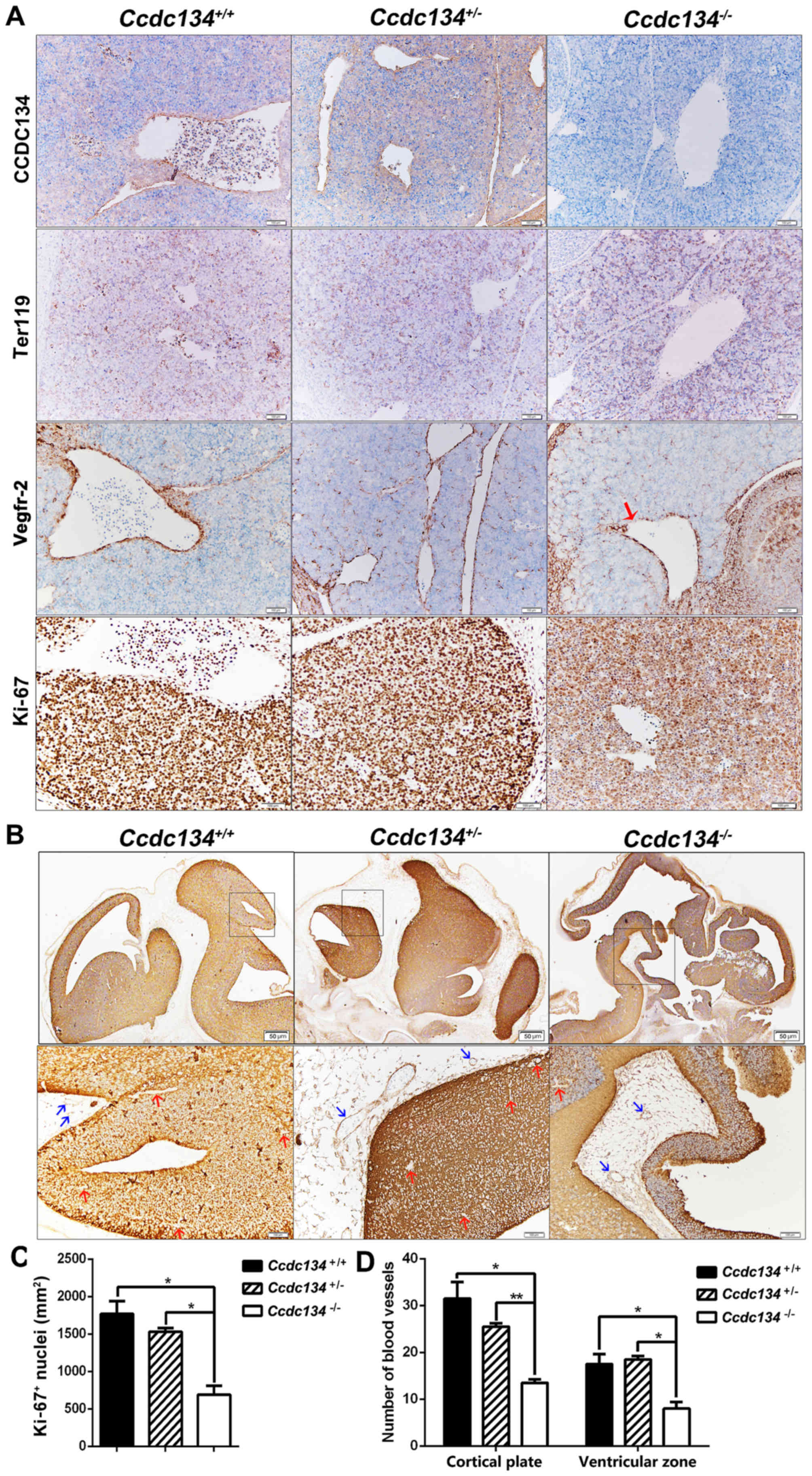
Ccdc134−/− embryos at E13.5. As demonstrated in
Fig. 6, CCDC134 expression was
initially examined in vascular endothelial cells and some
differentiated erythroid cells of fetal livers; however,
Ccdc134−/− embryos were identified by
immunostaining for the absence of CCDC134 protein (Fig. 6A). As the embryonic liver is the
main site of hematopoiesis in the second-half of murine gestation,
it appeared that poor oxygenation secondary to anemia may
contribute to embryonic demise (15). Erythroid cells of fetal livers
were detected using the antibody against Ter119, which is a surface
marker for differentiated erythroid cells (16). As demonstrated in Fig. 6A, no obvious difference was
observed between the fetal liver of Ccdc134−/−
embryos and wild-type embryos.
To further evaluate the existence of the vascular
integrity defect, immunohistochemical analysis with specific
antibody against Vegfr-2 was conducted, which is a type V receptor
tyrosine kinase, predominantly known to be expressed in vascular
endothelial cells (17). The
representative example of Fig. 6A
illustrates a discontinuous Vegfr-2-positive endothelial cell
barrier in fetal livers in Ccdc134−/− embryos
compared with Ccdc134+/+ and
Ccdc134+/− embryos, which may contribute to fetal
liver abnormality. In order to further investigate the influence of
Ccdc134 deficiency on cell proliferation in fetal livers, a
proliferation study was performed at E13.5. Ki-67 immunostaining
detection of cycling cells revealed a significant decrease of
Ki-67+ nuclei in fetal livers of
Ccdc134−/− embryos compared with wild-type
embryos (Fig. 6A and C).
Observations of Ccdc134−/− embryos support that
the phenotype of an abnormal fetal liver described at E13.5 above
is due to a significant decrease in proliferation and impaired
blood vessels.
Gross morphology suggested severe hemorrhage in the
brain ventricular space of Ccdc134−/− embryos at
E13.5. Vascular integrity defect was further analyzed with
anti-Vegfr-2 antibody. As demonstrated in Fig. 6B, Vegfr-2 was expressed by both
cerebral tissue and vessels in the forebrain, midbrain and
hindbrain of embryos at E13.5; however, compared with the
Ccdc134+/+ and Ccdc134+/−
embryos, overall brain disorganization in the
Ccdc134−/− embryo was observed. Additionally, a
significant reduction of vascular density was observed in the
Ccdc134−/− embryos compared with their
littermates (Fig. 6D). Taken
together, these findings suggest that Ccdc134 may be
associated with angiogenesis.
Discussion
The present study reported the generation of
complete Ccdc134 null mice by deleting the four coding exons
of Ccdc134 using a Cre-loxP system, which abolishes the
expression of CCDC134 protein. The present study also demonstrated
the CCDC134 expression pattern during embryogenesis. Furthermore, a
large number of postnatal mice were examined and no homozygous
Ccdc134 KO mice were identified, suggesting that complete
loss of Ccdc134 resulted in embryonic lethality.
The main processes involved in mouse embryonic
development include regional specification, morphogenesis, cell
differentiation, cell growth and the overall control of timing
(18). We speculate that the main
cause of mortality in Ccdc134−/− embryos at E13.5
was anemia, as severe hemorrhage in the brain and neural tube was
identified. Notably, mortality at this age corresponds to a
developmental period during which defects of the heart or placenta
often lead to embryonic lethality (19). Additionally, the placentas of
Ccdc134−/− embryos were thinner and poorly
developed and showed a prominence of labyrinth trophoblasts. The
cell numbers in Ccdc134−/− fetal livers were
significantly reduced compared with the number in wild-type livers.
Also, fewer Vegfr-2-positive endothelial cells and reduced cell
proliferation were demonstrated in fetal livers of
Ccdc134−/− embryos compared to that of wild-type
embryos. It is also interesting to note that
Ccdc134−/− embryos demonstrated broad areas of
extravasated blood associated with discontinuous and defective
blood vessels. Angiogenesis is important in embryonic development,
in which endothelial progenitor cells (EPCs) serve critical roles
(20). Furthermore, angiogenesis
is driven by newly formed EPCs migrating from the sites of
hematopoietic stem cell (HSC) development (21). Bone marrow begins to function as a
source of HSCs just before birth, whereas in embryogenesis,
multi-lineage hematopoietic progenitors exist in the extraembryonic
yolk sac at E8.25, and in the placenta and embryonic
aorta-gonad-mesonephros region at E10 (22). From E12 to birth, the fetal liver
is the main site for definitive HSC formation (23). These findings implied that
Ccdc134 may have a role in hematopoiesis and angiogenesis
during embryonic development. Therefore, conditional deletion of
Ccdc134 in hematopoietic stem cells (Vav-Cre) and
endothelial cells (Tie2-Cre) may be utilized to further investigate
the function of CCDC134 in the differentiation of HSCs and
angiogenesis.
In conjunction with our previous results (4), CCDC134 was identified to be a novel
partner of hADA2a protein, which was a core component of the yeast
alteration/deficiency in activation (ADA)/GCN5 HAT complexes to
facilitate the acetylation of nucleosomal histones (24). Additionally, CCDC134 may act as a
critical regulator of hADA2a stability and activity, and
participate in the PCAF complex via hADA2a to affect its HAT
activity, which acetylates H3K14 and H4K8 (24). Deletion of GCN5 and PCAF resulted
in embryo lethality between E9.5 and E11.5, indicating that PCAF
and GCN5 served important roles in embryogenesis (25). Also, GCN5 and PCAF had redundant
functions in mouse embryonic fibroblasts (26). Additionally, HAT PCAF/lysine
acetyltransferase 2B was an important factor of the Hedgehog
signaling pathway that served an important role in embryonic
patterning and development of various tissues and organs, as well
as in maintaining and repairing mature tissues in adults (27). During embryogenesis, the early
mammalian embryo was characterized by large-scale chromatin
remodeling, including changes in histone variant incorporation,
global changes in DNA, and histone tail modification (28,29). Histone H3 acetylation in the
nucleosome core occurred with different temporal kinetics during
mouse pre-implantation development, and further affected embryo
development (30). In addition,
various studies have demonstrated that angiogenesis is precisely
regulated by soluble growth factors and receptor-mediated signals.
Vegfr-2 is a key regulator of angiogenesis, and its expression and
function are regulated by acetylation under dynamic control of the
acetyltransferase p300 (31,32). Furthermore, PCAF acts as a master
switch in the inflammatory processes required for effective
arteriogenesis (33). Therefore,
the cause of embryonic mortality in the present study may be due to
defective hematopoiesis and angiogenesis, which likely cause
mortality due to failure of acetylation modification of key
regulators during embryo development.
In conclusion, the present results support the
conclusion that disruption of Ccdc134 expression in mice
leads to embryonic lethality. In addition, the present preliminary
studies suggested that Ccdc134 may be related to
hematopoiesis and angiogenesis, and further studies will be
performed. Thus, the Ccdc134 null line provides a critical
tool for determining the physiological roles of Ccdc134.
Furthermore, Ccdc134fl/fl mice will be important
for the analysis of Ccdc134 function in specific cell types
and the extension into analysis of Ccdc134 loss of function
in adult animals.
Acknowledgments
We thank Mrs. Weiyan Xv (Peking University Human
Disease Genomics Center, Beijing, China) for technical support. The
present study was supported by the National Natural Sciences
Foundation of China (grant no. 81372254), the National Basic
Research Program of China (grant no. 2013CB837201), Beijing Natural
Sciences Foundation (grant no. 7142082) and the National Science
and Technology Major Projects of New Drugs (grant no.
2012ZX09103301-032).
Glossary
Abbreviations
Abbreviations:
|
CCDC134
|
coiled-coil domain containing 134
|
|
PCR
|
polymerase chain reaction
|
|
hADA2a
|
human alteration/deficiency in
activation 2a
|
|
PCAF
|
p300-CBP-associated factor
|
|
ACTB
|
human β-actin
|
|
Zp3
|
zona pellucida 3
|
|
MAPK
|
mitogen-activated protein kinase
|
|
Vegfr-2
|
vascular endothelial growth factor
receptor-2
|
|
EPCs
|
endothelial progenitor cells
|
|
HSCs
|
hematopoietic stem cells
|
|
HAT
|
histone acetyltransferase
|
References
|
1
|
Huang J, Shi T, Ma T, Zhang Y, Ma X, Lu Y,
Song Q, Liu W, Ma D and Qiu X: CCDC134, a novel secretory protein,
inhibits activation of ERK and JNK, but not p38 MAPK. Cell Mol Life
Sci. 65:338–349. 2008. View Article : Google Scholar
|
|
2
|
Zhong J, Zhao M, Luo Q, Ma Y, Liu J, Wang
J, Yang M, Yuan X, Sang J and Huang C: CCDC134 is down-regulated in
gastric cancer and its silencing promotes cell migration and
invasion of GES-1 and AGS cells via the MAPK pathway. Mol Cell
Biochem. 372:1–8. 2013. View Article : Google Scholar
|
|
3
|
Huang J, Xiao L, Gong X, Shao W, Yin Y,
Liao Q, Meng Y, Zhang Y, Ma D and Qiu X: Cytokine-like molecule
CCDC134 contributes to CD8+ T-cell effector functions in
cancer immuno-therapy. Cancer Res. 74:5734–5745. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Huang J, Zhang L, Liu W, Liao Q, Shi T,
Xiao L, Hu F and Qiu X: CCDC134 interacts with hADA2a and functions
as a regulator of hADA2a in acetyltransferase activity, DNA
damage-induced apoptosis and cell cycle arrest. Histochem Cell
Biol. 138:41–55. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Schiltz RL, Mizzen CA, Vassilev A, Cook
RG, Allis CD and Nakatani Y: Overlapping but distinct patterns of
histone acetylation by the human coactivators p300 and PCAF within
nucleosomal substrates. J Biol Chem. 274:1189–1192. 1999.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Garlanda C, Dinarello CA and Mantovani A:
The interleukin-1 family: Back to the future. Immunity.
39:1003–1018. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yang H, Wang H, Chavan SS and Andersson U:
High mobility group box protein 1 (HMGB1): The prototypical
endogenous danger molecule. Mol Med. 21(Suppl 1): S6–S12.
2015.PubMed/NCBI
|
|
8
|
Li G, Xu C, Lin X, Qu L, Xia D, Hongdu B,
Xia Y, Wang X, Lou Y, He Q, et al: Deletion of Pdcd5 in mice led to
the deficiency of placenta development and embryonic lethality.
Cell Death Dis. 8:e28112017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(−ΔΔC(T)) method. Methods. 25:402–408. 2001. View Article : Google Scholar
|
|
10
|
Farley FW, Soriano P, Steffen LS and
Dymecki SM: Widespread recombinase expression using FLPeR (flipper)
mice. Genesis. 28:106–110. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Rodríguez CI, Buchholz F, Galloway J,
Sequerra R, Kasper J, Ayala R, Stewart AF and Dymecki SM:
High-efficiency deleter mice show that FLPe is an alternative to
Cre-loxP. Nat Genet. 25:139–140. 2000. View
Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lewandoski M, Meyers EN and Martin GR:
Analysis of Fgf8 gene function in vertebrate development. Cold
Spring Harb Symp Quant Biol. 62:159–168. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
de Vries WN, Binns LT, Fancher KS, Dean J,
Moore R, Kemler R and Knowles BB: Expression of Cre recombinase in
mouse oocytes: A means to study maternal effect genes. Genesis.
26:110–112. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Renaud SJ, Karim Rumi MA and Soares MJ:
Review: Genetic manipulation of the rodent placenta. Placenta.
32(Suppl 2): S130–S135. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Dzierzak E, Medvinsky A and de Bruijn M:
Qualitative and quantitative aspects of haematopoietic cell
development in the mammalian embryo. Immunol Today. 19:228–236.
1998. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Koulnis M, Pop R, Porpiglia E, Shearstone
JR, Hidalgo D and Socolovsky M: Identification and analysis of
mouse erythroid progenitors using the CD71/TER119 flow-cytometric
assay. J Vis Exp. 54:e28092011.
|
|
17
|
Breier G, Clauss M and Risau W: Coordinate
expression of vascular endothelial growth factor receptor-1 (flt-1)
and its ligand suggests a paracrine regulation of murine vascular
development. Dev Dyn. 204:228–239. 1995. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Slack JM: Essential Developmental Biology.
3rd edition. Wiley-Blackwell; Oxford: 2012
|
|
19
|
Savolainen SM, Foley JF and Elmore SA:
Histology atlas of the developing mouse heart with emphasis on
E11.5 to E18.5. Toxicol Pathol. 37:395–414. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Folkman J: Angiogenesis in cancer,
vascular, rheumatoid and other disease. Nat Med. 1:27–31. 1995.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Smith AG: Embryo-derived stem cells: Of
mice and men. Annu Rev Cell Dev Biol. 17:435–462. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Coşkun S, Chao H, Vasavada H, Heydari K,
Gonzales N, Zhou X, de Crombrugghe B and Hirschi KK: Development of
the fetal bone marrow niche and regulation of HSC quiescence and
homing ability by emerging osteolineage cells. Cell Rep. 9:581–590.
2014. View Article : Google Scholar :
|
|
23
|
Golub R and Cumano A: Embryonic
hematopoiesis. Blood Cells Mol Dis. 51:226–231. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Grant PA, Duggan L, Côté J, Roberts SM,
Brownell JE, Candau R, Ohba R, Owen-Hughes T, Allis CD, Winston F,
et al: Yeast Gcn5 functions in two multisubunit complexes to
acetylate nucleosomal histones: Characterization of an Ada complex
and the SAGA (Spt/Ada) complex. Genes Dev. 11:1640–1650. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Yamauchi T, Yamauchi J, Kuwata T, Tamura
T, Yamashita T, Bae N, Westphal H, Ozato K and Nakatani Y: Distinct
but overlapping roles of histone acetylase PCAF and of the closely
related PCAF-B/GCN5 in mouse embryogenesis. Proc Natl Acad Sci USA.
97:11303–11306. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Jin Q, Yu LR, Wang L, Zhang Z, Kasper LH,
Lee JE, Wang C, Brindle PK, Dent SY and Ge K: Distinct roles of
GCN5/PCAF-mediated H3K9ac and CBP/p300-mediated H3K18/27ac in
nuclear receptor transactivation. EMBO J. 30:249–262. 2011.
View Article : Google Scholar :
|
|
27
|
Malatesta M, Steinhauer C, Mohammad F,
Pandey DP, Squatrito M and Helin K: Histone acetyltransferase PCAF
is required for Hedgehog-Gli-dependent transcription and cancer
cell proliferation. Cancer Res. 73:6323–6333. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Hemberger M, Dean W and Reik W: Epigenetic
dynamics of stem cells and cell lineage commitment: Digging
Waddington's canal. Nat Rev Mol Cell Biol. 10:526–537. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Burton A and Torres-Padilla ME: Chromatin
dynamics in the regulation of cell fate allocation during early
embryogenesis. Nat Rev Mol Cell Biol. 15:723–734. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ziegler-Birling C, Daujat S, Schneider R
and Torres-Padilla ME: Dynamics of histone H3 acetylation in the
nucleosome core during mouse pre-implantation development.
Epigenetics. 11:553–562. 2016. View Article : Google Scholar :
|
|
31
|
Rahimi N and Costello CE: Emerging roles
of post-translational modifications in signal transduction and
angiogenesis. Proteomics. 15:300–309. 2015. View Article : Google Scholar :
|
|
32
|
Zecchin A, Pattarini L, Gutierrez MI, Mano
M, Mai A, Valente S, Myers MP, Pantano S and Giacca M: Reversible
acetylation regulates vascular endothelial growth factor receptor-2
activity. J Mol Cell Biol. 6:116–127. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Bastiaansen AJ, Ewing MM, de Boer HC, van
der Pouw Kraan TC, de Vries MR, Peters EA, Welten SM, Arens R,
Moore SM, Faber JE, et al: Lysine acetyltransferase PCAF is a key
regulator of arteriogenesis. Arterioscler Thromb Vasc Biol.
33:1902–1910. 2013. View Article : Google Scholar : PubMed/NCBI
|