Introduction
Pelvic fracture urethral distraction defect (PFUDD)
seriously affects voiding function and the quality of life of
patients (1). Urethral defects
are healed by fibroblasts, but overexpression of extracellular
matrix (ECM) components in the resultant scar tissue may cause a
reduction or complete blockage in the urethral caliber and
impairment of the flow of urine (2). Open urethroplasty of urethral
strictures is performed worldwide and has become the gold standard
for the treatment of urethral strictures. However, this surgery is
often performed in high-volume urethral referral centers, and only
surgeons with multiple years of training are able to obtain high
(~95%) success rates (3).
Overall, the success rate of urethral reconstructive surgeries
remains relatively low globally (short-term 79–88%, long term
42–86%) (4,5). Reconstructive surgery may also
result in serious morbidity at the defect location and the donor
site, including infection, nerve injury and difficulties in opening
the mouth (2,6). To minimize side-effects and
complications, micro invasive surgeries [for example, direct visual
internal urethrotomy (DVIU)] has been used to treat posterior
urethral stricture caused by PFUDD. DVIU is commonly performed in
patients with urethral stricture and remains the first choice in
secondary centers with limited equipment or those that lack
experienced surgeons. However, high recurrence and poor long-term
curative rates have been associated with this technique, and its
effectiveness remains questionable among urologists (7,8).
Thus, there is an urgent need for novel techniques and
complementary strategies to prevent fibrosis and scar formation in
patients following surgery. Although the local injection and
dilution of steroids and other anti-fibrosis reagents were
introduced as an adjuvant treatment for DVIU, a previous systematic
review and meta-analysis by our group has demonstrated that these
drugs are not efficient or effective enough to decrease the
recurrence rate (7).
Gene therapy has emerged as a promising treatment
option for a number of diseases. Accumulating evidence indicates
that microRNAs (miRNAs/miRs) are involved in multiple disease
processes, including the pathogenesis of fibrosis in the skin and
liver (9). Our goal is to add to
the information regarding miRNA dysregulation in urethral fibrosis
to broaden our understanding of the underlying pathological
mechanisms and discover novel treatment targets. By conducting
miRNA sequencing, temporal miRNA expression profiles were generated
in urethral scar and normal urethral tissues, and potential gene
targets and pathways involved in fibrosis were identified in
patients suffering from PFUDD. This established miRNA profile may
provide further insight into the involvement of miRNAs and their
functions in the development and progression of pathological
conditions.
Materials and methods
Urethral scar samples
All procedures performed in studies involving human
participants were in accordance with the ethical standards of the
institutional research committee and in accordance with the 1964
Declaration of Helsinki and its later amendments or comparable
ethical standards. The present study was approved by the Ethics
Committee of Shanghai Sixth People's Hospital (Shanghai, China).
Informed consent was obtained from all patients. Scar and normal
urethral tissues were collected from patients with urethral
stricture undergoing urethroplasty (n=5). All subjects were male
with an age range of 16–59 years. Baseline patient information is
summarized in Table I.
 | Table IBaseline patient characteristics. |
Table I
Baseline patient characteristics.
| Patient number | Age | Sex | Health status |
|---|
| 1 | 59 | Male | PFUDD |
| 2 | 50 | Male | PFUDD |
| 3 | 16 | Male | PFUDD |
| 4 | 43 | Male | PFUDD |
| 5 | 44 | Male | PFUDD |
A total of 3 pairs of samples were used for miRNA
sequencing, and 3 pairs were used for quantitative polymerase chain
reaction (qPCR) verification (one patient's tissue was used in both
miRNA sequencing and qPCR). The etiology of patients with urethral
stricture was PFUDD. All the participating patients underwent
primary surgery. The mean length of the stricture was 1.5 cm and
the locations were all at the membranous segment of the urethra.
Samples were harvested following surgery, sectioned and stored at
−80°C until the process of RNA extraction.
RNA extraction and miRNA sequencing
analysis
Total RNA from each sample (Table II) was used to prepare the miRNA
sequencing library, which included the following steps: i)
3′-adaptor ligation; ii) 5′-adaptor ligation; iii) cDNA synthesis;
iv) PCR amplification; v) size selection of ~135–155 bp PCR
amplified fragments (corresponding to ~15–35 nt small RNAs).
 | Table IIRNA quality control characteristics
for microRNA sequencing samples. |
Table II
RNA quality control characteristics
for microRNA sequencing samples.
| Sample IDa | OD260/280
ratio | OD260/230
ratio | Conc. (ng/µl) | Volume (µl) | Quantity (ng) | QC |
|---|
| Normal 1 | 1.87 | 2.40 | 316.68 | 30 | 9500.40 | Pass |
| Scar 1 | 1.85 | 2.38 | 282.52 | 40 | 11300.80 | Pass |
| Normal 2 | 1.86 | 2.41 | 182.11 | 20 | 3642.20 | Pass |
| Scar 2 | 1.88 | 2.39 | 237.14 | 40 | 9485.60 | Pass |
| Normal 3 | 1.81 | 2.40 | 144.77 | 10 | 1447.70 | Pass |
| Scar 3 | 1.88 | 2.30 | 267.73 | 40 | 10709.20 | Pass |
The TruSeq SR Cluster kit (cat. no. GD-402-4001;
Illumina, Inc., San Diego, CA, USA) was used for cluster generation
in an Illumina cBOT instrument, following the manufacturer's
protocol. Sequencing was performed on an Illumina HiSeq®
2000 instrument (Illumina, Inc.) using a TruSeq Rapid SBS kit (cat.
no. FC-402-4002; Illumina, Inc.).
Following sequencing, the Solexa CHASTITY quality
filtered reads were harvested as clean reads. Adaptor sequences
were trimmed, and the retained reads (≥15 nt) were aligned to
merged pre-miRNA databases (known pre-miRNAs from miRBase v21
(10–14) plus the newly predicted pre-miRNAs)
using Novoalign software v2.07.11 [(Novocraft Technologies Sdn Bhd,
Selangor, Malaysia) maximum mismatch was 1, and low reads (total
reads <2) were discarded]. To characterize the isomiR
variability, sequences matching each miRNA precursor(s) in the
mature miRNA region ±4 nt (≤1 mismatch) were accepted as mature
miRNA isomiRs and grouped according to the 5-prime (5p) or 3-prime
(3p) arm of the precursor hairpin. The number of mapped tags was
defined as the raw expression level of the specific miRNAs.
Transcripts per million values were generated based on the total
number of aligned tags for normalization. The most abundant isomiR,
mature miRNA (annotated in miRBase) and all isoforms of miRNA (5p
or 3p) were used for miRNA expression quantification. For
comparison of miRNA expression profiles between two groups (scar
tissue vs. normal tissue), fold change and P-value were calculated
to identify miRNAs with differentiated expression.
Reverse transcription (RT)-qPCR for the
validation of miRNAs
A total of 3 pairs of urethral samples were used for
qPCR validation of miRNA expression. The primers used for PCR were
designed with Primer Premier software (version 5.0; Premier Biosoft
International, Palo Alto, CA, USA) (primer sequence details are
summarized in Table III). cDNA
synthesis was performed on a GeneAmp PCR System 9700 (Applied
Biosystems; Thermo Fisher Scientific, Inc., Waltham, MA, USA) using
a RevertAid First Strand cDNA Synthesis kit, cat. no. K1622 (Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocols.
qPCR was performed on a ViiA 7 Real-time PCR System (Applied
Biosystems; Thermo Fisher Scientific, Inc.) using a PowerUp SYBR
Green Master Mix, (cat. no. A25778, Thermo Fisher Scientific,
Inc.). qPCR was conducted at: i) 95°C for 10 min, ii) 95°C for 15
sec and iii) 60 sec at >60°C for 40 cycles. The fold change for
each miRNA was calculated using the 2−∆∆Cq method
(15).
 | Table IIIMicroRNA primers for quantitative
real-time PCR. |
Table III
MicroRNA primers for quantitative
real-time PCR.
| Primer name | Primer sequence
(5′-3′) |
|---|
| U6-S |
CTCGCTTCGGCAGCACA |
| U6-A |
AACGCTTCACGAATTTGCGT |
| general control
primer-A |
TGGTGTCGTGGAGTCG |
|
hsa-miR-129-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGCAAGCCC |
|
hsa-miR-129-5p-S |
ACACTCCAGCTGGGCTTTTTGCGGTCTGG |
|
hsa-miR-9-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCATACAG |
| hsa-miR-9-5p-S |
ACACTCCAGCTGGGTCTTTGGTTATCTAGCT |
|
hsa-miR-9-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGACTTTCGG |
| hsa-miR-9-3p-S |
ACACTCCAGCTGGGATAAAGCTAGATAACC |
|
hsa-miR-183-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGTGAATT |
|
hsa-miR-183-5p-S |
ACACTCCAGCTGGGTATGGCACTGGTAGAA |
|
hsa-miR-6720-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCTACCAG |
|
hsa-miR-6720-3p-S |
ACACTCCAGCTGGGCGCGCCTGCAGGAACT |
|
hsa-miR-96-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGCAAAAA |
|
hsa-miR-96-5p-S |
ACACTCCAGCTGGGTTTGGCACTAGCACATT |
|
hsa-miR-486-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCTCGGGGC |
|
hsa-miR-486-5p-S |
ACACTCCAGCTGGGTCCTGTACTGAGCTGC |
|
hsa-miR-363-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTACAGATG |
|
hsa-miR-363-3p-S |
ACACTCCAGCTGGGAATTGCACGGTATCCA |
|
hsa-miR-135a-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCACATAG |
|
hsa-miR-135a-5p-S |
ACACTCCAGCTGGGTATGGCTTTTTATTCCT |
|
hsa-miR-374a-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAATTACAA |
|
hsa-miR-374a-3p-S |
ACACTCCAGCTGGGCTTATCAGATTGTATT |
|
hsa-miR-374b-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAATGATAA |
|
hsa-miR-374b-3p-S |
ACACTCCAGCTGGGCTTAGCAGGTTGTATT |
| hsa-miR-941-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGCACATGT |
| hsa-miR-941-S |
ACACTCCAGCTGGGCACCCGGCTGTGTGCAC |
|
hsa-miR-3158-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGTCCTGCA |
|
hsa-miR-3158-3p-S |
ACACTCCAGCTGGGAAGGGCTTCCTCTCTG |
|
hsa-miR-29c-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTAACCGAT |
|
hsa-miR-29c-3p-S |
ACACTCCAGCTGGGTAGCACCATTTGAAAT |
|
hsa-miR-10a-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTATTCCCC |
|
hsa-miR-10a-3p-S |
ACACTCCAGCTGGGCAAATTCGTATCTAGG |
|
hsa-miR-182-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGTGTGAG |
|
hsa-miR-182-5p-S |
ACACTCCAGCTGGGTTTGGCAATGGTAGAACT |
|
hsa-miR-106a-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCTACCTGC |
|
hsa-miR-106a-5p-S |
ACACTCCAGCTGGGAAAAGTGCTTACAGTGC |
|
hsa-miR-4284-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGATGGGGTG |
| hsa-miR-4284-S |
ACACTCCAGCTGGGGGGCTCACATCA |
|
has-mir192-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGGCTGTCA |
|
has-mir192-5p-S |
ACACTCCAGCTGGGCTGACCTATGAATTG |
|
hsa-miR-26a-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGCCTATC |
|
hsa-miR-26a-5p-S |
ACACTCCAGCTGGGTTCAAGTAATCCAGGA |
|
hsa-miR-223-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTGGGGTAT |
|
hsa-miR-223-3p-S |
ACACTCCAGCTGGGTGTCAGTTTGTCAAAT |
|
hsa-miR-92a-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGACAGGCCG |
|
hsa-miR-92a-3p-S |
ACACTCCAGCTGGGTATTGCACTTGTCCCG |
|
hsa-miR-29b-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAACACTGA |
|
hsa-miR-29b-3p-S |
ACACTCCAGCTGGGTAGCACCATTTGAAATC |
|
hsa-miR-125b-5p |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCACAAGT |
|
hsa-miR-125b-5p |
ACACTCCAGCTGGGTCCCTGAGACCCTAAC |
|
hsa-miR-194-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTCCACATG |
|
hsa-miR-194-5p-S |
ACACTCCAGCTGGGTGTAACAGCAACTCCA |
|
hsa-miR-20a-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGCTACCTGC |
|
hsa-miR-20a-5p-S |
ACACTCCAGCTGGGTAAAGTGCTTATAGTGC |
|
hsa-miR-128-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAAAGAGAC |
|
hsa-miR-128-3p-S |
ACACTCCAGCTGGGTCACAGTGAACCGGT |
|
hsa-miR-654-5p-S |
ACACTCCAGCTGGGTGGTGGGCCGCAGAAC |
|
hsa-miR-654-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGCACATGT |
|
hsa-miR-31-3p-S |
ACACTCCAGCTGGGTGCTATGCCAACATAT |
|
hsa-miR-31-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGATGGCAAT |
|
hsa-miR-329-3p-S |
ACACTCCAGCTGGGAACACACCTGGTTAAC |
|
hsa-miR-329-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAAAGAGGT |
|
hsa-let-7b-3p-S |
ACACTCCAGCTGGGCTATACAACCTACTGC |
|
hsa-let-7b-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGGAAGGC |
|
hsa-miR-433-3p-S |
ACACTCCAGCTGGGATCATGATGGGCTCCT |
|
hsa-miR-433-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGACACCGAG |
|
hsa-miR-379-3p-S |
ACACTCCAGCTGGGTATGTAACATGGTCCA |
|
hsa-miR-379-3p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGAGTTAGTG |
|
hsa-miR-744-5p-S |
ACACTCCAGCTGGGTGCGGGGCTAGGGCTA |
|
hsa-miR-744-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGTGCTGTTA |
|
hsa-miR-29c-5p-S |
ACACTCCAGCTGGGTGACCGATTTCTCCTG |
|
hsa-miR-29c-5p-RT |
CTCAACTGGTGTCGTGGAGTCGGCAATTCAGTTGAGGAACACCA |
Bioinformatics analysis
Hierarchical clustering was performed. miRNA target
prediction was performed by TargetScan, miRanda, and microcosm with
DAVID Bioinformatics Resources 6.7 (https://david-d.ncifcrf.gov/home) (16–22). Enrichment analysis for Gene
Ontology (GO) and Kyoto Encylopedia of Genes and Genomes (KEGG)
pathways was performed using the topGO package in R (https://www.bioconductor.org/packages/3.7/bioc/html/topGO.html)
(20–22).
Statistical analysis
Comparisons between average fold changes of
different miRNAs were performed using Student's t-tests (using the
T.TEST function in Microsoft Excel; Microsoft Corporation, Redmond,
WA, USA). For analysis based on miRNA sequencing only, the t-tests
used a two-tailed distribution and the comparison was paired for
expression in normal and scar tissues (n=3; Table IV). For analysis based on both
miRNA sequencing and qPCR validation (Table V), the t-tests used a two-tailed
distribution and the comparison was paired for expression evaluated
by the two methods [miRNA sequencing and qPCR, n=7 (for upregulated
miRNAs) and n=15 (for downregulated miRNAs)].
 | Table IVmiRs with significantly different
expression, as identified by miR sequencing. |
Table IV
miRs with significantly different
expression, as identified by miR sequencing.
A, Upregulated
genes
|
|---|
| miR | Fold change
(scar/normal) | P-valuea |
|---|
| hsa-miR-129-5p | 3.75 | 0.026 |
| hsa-miR-9-5p | 2.08 | 0.003 |
|
hsa-miR-374a-3p | 1.92 | 0.039 |
|
hsa-miR-374b-3p | 1.42 | 0.008 |
| hsa-miR-128-3p | 1.19 | 0.014 |
|
| B, Downregulated
genes |
|
| miR | Fold change
(scar/normal) | P-valuea |
|
| hsa-miR-26a-5p | 0.91 | 0.004 |
| hsa-miR-941 | 0.90 | 0.019 |
| hsa-miR-223-3p | 0.83 | 0.043 |
|
hsa-miR-3158-3p | 0.78 | 0.025 |
| hsa-miR-20a-5p | 0.72 | 0.035 |
| hsa-miR-92a-3p | 0.70 | 0.039 |
| hsa-miR-29c-3p | 0.69 | 0.002 |
| hsa-miR-194-5p | 0.67 | 0.000 |
|
hsa-miR-125b-5p | 0.67 | 0.005 |
| hsa-miR-29b-3p | 0.64 | 0.008 |
| hsa-miR-192-5p | 0.63 | 0.041 |
| hsa-miR-10a-3p | 0.62 | 0.023 |
| hsa-miR-182-5p | 0.51 | 0.016 |
|
hsa-miR-106a-5p | 0.46 | 0.038 |
| hsa-miR-183-5p | 0.45 | 0.013 |
|
hsa-miR-6720-3p | 0.38 | 0.038 |
| hsa-miR-96-5p | 0.33 | 0.037 |
| hsa-miR-486-5p | 0.27 | 0.035 |
| hsa-miR-363-3p | 0.26 | 0.014 |
|
hsa-miR-135a-5p | 0.19 | 0.004 |
| hsa-miR-4284 | 0.17 | 0.038 |
 | Table VComparison between miR sequencing and
qPCR sequencing. |
Table V
Comparison between miR sequencing and
qPCR sequencing.
| miR | Average change by
miR sequencing (scar/normal) | Average change by
qPCR (scar/normal) |
|---|
| has-miR-129-5p | 3.75 | 2.13 |
| hsa-miR-654-5p | 2.06 | 1.57 |
| hsa-miR-31-3p | 1.55 | 1.08 |
| hsa-miR-433-3p | 1.42 | 2.36 |
| hsa-miR-329-3p | 1.32 | 2.46 |
| hsa-let-7b-3p | 1.17 | 2.71 |
| hsa-miR-379-3p | 1.17 | 2.23 |
| hsa-miR-26a-5p | 0.91 | 1.13 |
| hsa-miR-744-5p | 0.84 | 2.52 |
| hsa-miR-223-3p | 0.83 | 0.54 |
| hsa-miR-20a-5p | 0.72 | 0.98 |
| hsa-miR-92a-3p | 0.70 | 1.12 |
| hsa-miR-29c-3p | 0.69 | 1.74 |
|
hsa-miR-125b-5p | 0.67 | 1.70 |
| hsa-miR-29c-5p | 0.67 | 3.01 |
| hsa-miR-192-5p | 0.63 | 2.35 |
|
hsa-miR-106a-5p | 0.46 | 1.19 |
| hsa-miR-183-5p | 0.45 | 2.22 |
| hsa-miR-96-5p | 0.33 | 1.77 |
| hsa-miR-486-5p | 0.27 | 1.84 |
| hsa-miR-4284 | 0.17 | 2.03 |
Histological evaluation
Freshly harvested scar and normal tissue were fixed
in 4% paraformaldehyde at 4°C for over 24 h, dehydrated and
embedded in paraffin following standard protocols by Servicebio,
Inc. (Wuhan Servicebio Technology Co., Ltd., Wuhan, Hubei, China).
The samples were stored and sectioned at room temperature; 4 µm
sections were incubated in a 65°C oven for 2 h and de-paraffinized
by washing with xylene and a descending alcohol gradient (100, 100,
85 and 75% for immunohistochemistry staining; 100, 95, 90, 80 and
70% for Masson trichrome staining) prior to staining.
Immunohistochemistry and Masson trichrome staining
were performed using mouse-anti-human monoclonal transforming
growth factor (TGF)-β1 antibodies (1:200, cat. no. GB11179,
Servicebio, Inc., Wuhan Servicebio Technology Co., Ltd.) and Masson
trichrome (cat. no. PT003; Shanghai Bogoo Biotechnology. Co., Ltd.,
Shanghai, China) following the manufacturer's protocols. Secondary
antibody staining involved the DAKO REAL EnVision Detection System,
Peroxidase/DAB, Rabbit/Mouse (cat. no. K5007; Agilent Technologies,
Inc., Santa Clara, CA, USA) and a Masson staining kit (cat. no.
G1006; Servicebio, Inc., Wuhan Servicebio Technology Co., Ltd.)
according to the manufacturer's protocols.
Results
miRNA expression results based on
sequencing data
All sequencing samples were evaluated for RNA
quality (Table II) and an miRNA
expression profile was generated to identify differentially
expressed miRNAs in scar tissues of patients with urethral
stricture. Out of the 1,620 human miRNAs that were sequenced, 55
miRNAs were upregulated (fold change >2) and 39 were
downregulated (fold change <0.5).
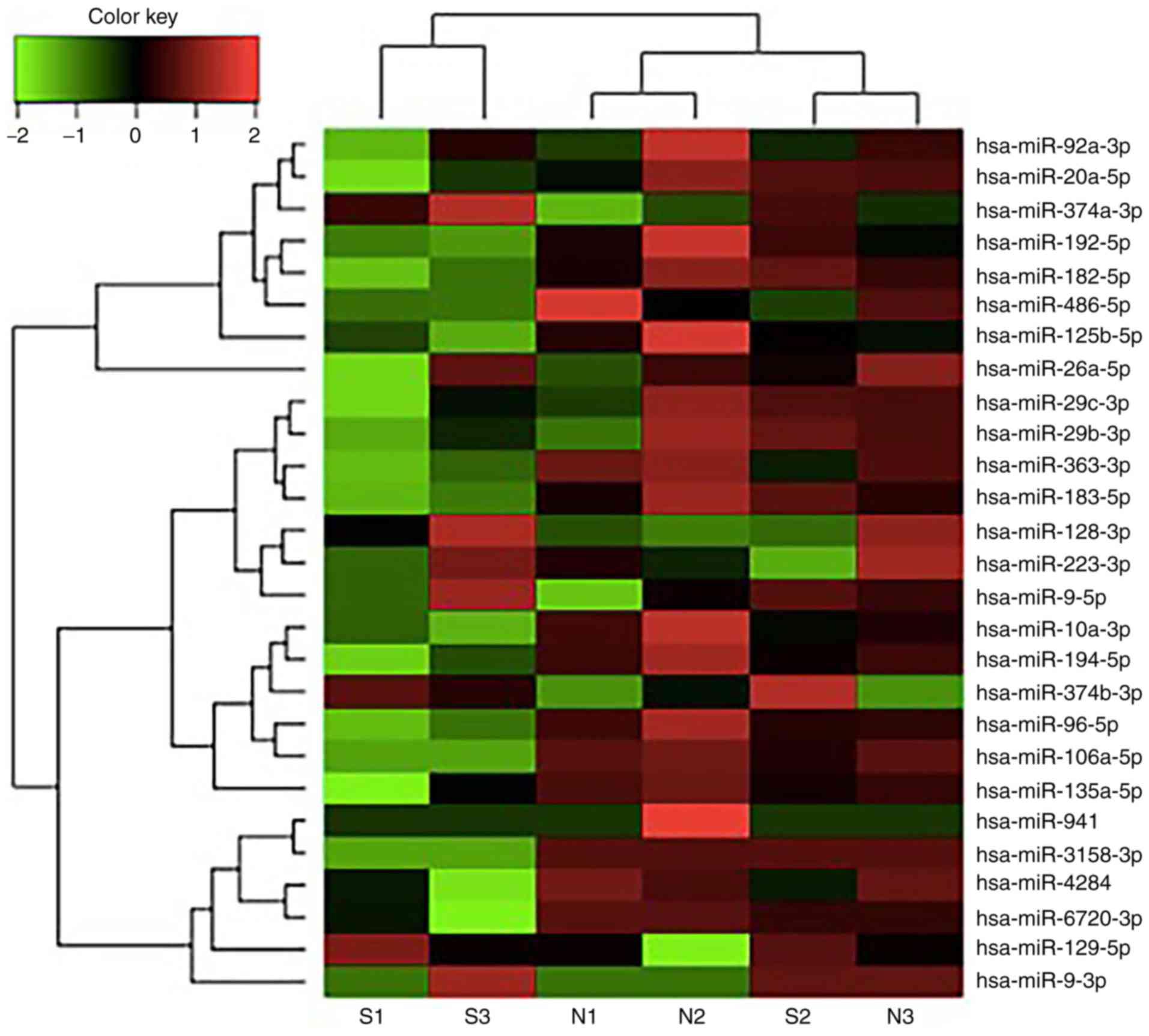
A total of 26 differentially expressed miRNAs were
identified, with 5 upregulated and 21 downregulated (P<0.05;
Fig. 1). The fold changes and
P-values were summarized in Table
IV.
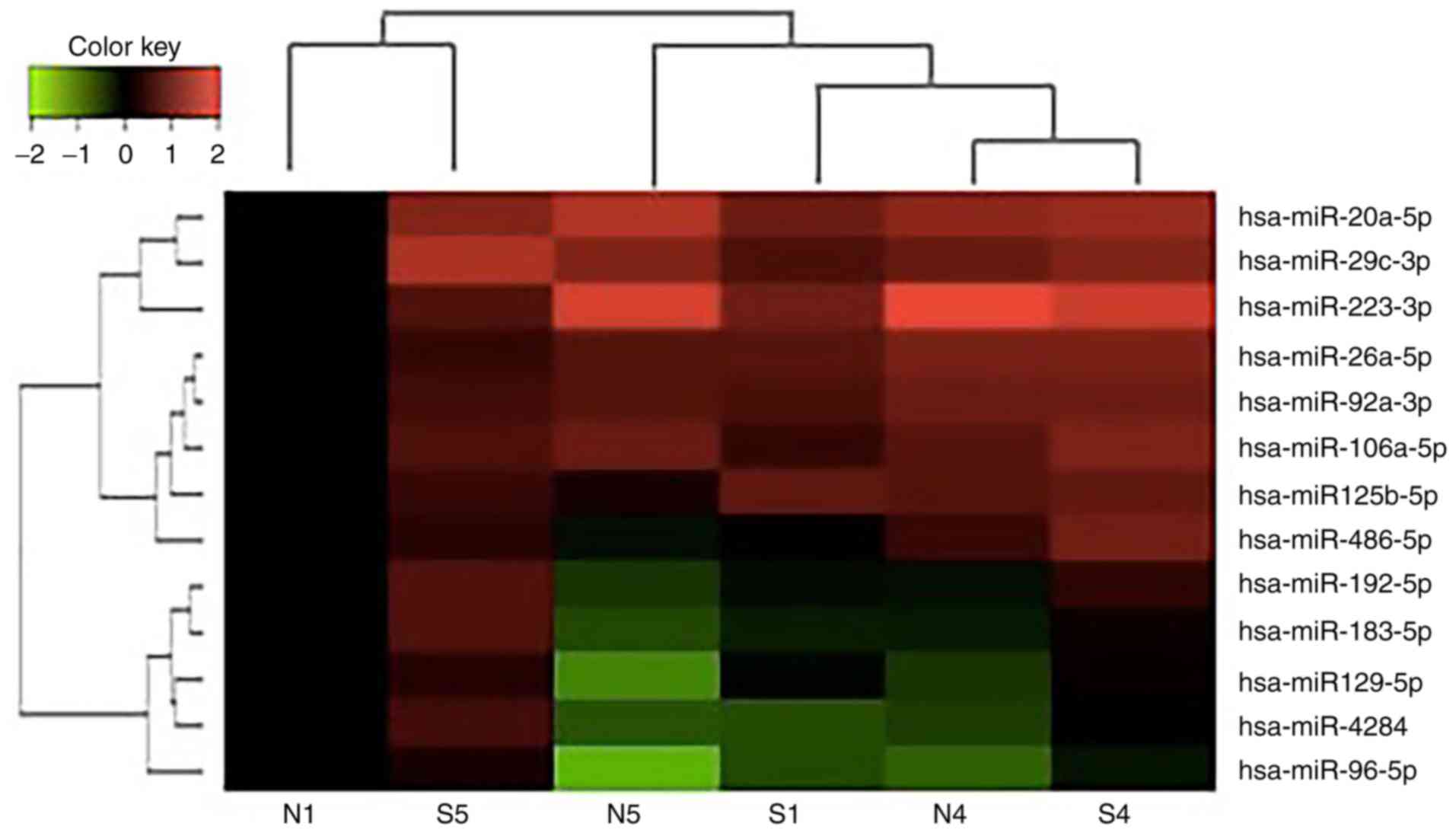
Valitation of miRNAs with qPCR
qPCR was performed for miRNAs with significant
changes based on the miRNA sequencing data (Fig. 2). miRNA upregulation identified by
miRNA sequencing was validated by qPCR (P=0.51; paired t-test
comparisons; n=7), while miRNA downregulation identified by miRNA
sequencing was quite different from the data generated by qPCR
(P<0.01; paired t-test comparisons; n=15; Table V). Among these, hsa-miR-129-5p was
overexpressed in scar tissue compared with normal tissue, although
the difference was not significant. Hsa-miR-223-3p and
hsa-miR-20a-5p expression was slightly decreased in scar tissues
compared with normal tissues. Hsa-miR-26a-5p, hsa-miR-92a-3p,
hsa-miR-29c-3p and hsa-miR-125b-5p demonstrated slight changes in
expression, but showed opposite regulation patterns as validated by
qPCR. Given the fact that sample size was small and the materials
harvested were limited in the present study, it was not possible to
generate certain conclusions or rule out the possibility of sample
variety in this case. Further functional validation will be
required to confirm the involvement of these candidate miRNAs in
fibrosis, scar formation and PFUDD.
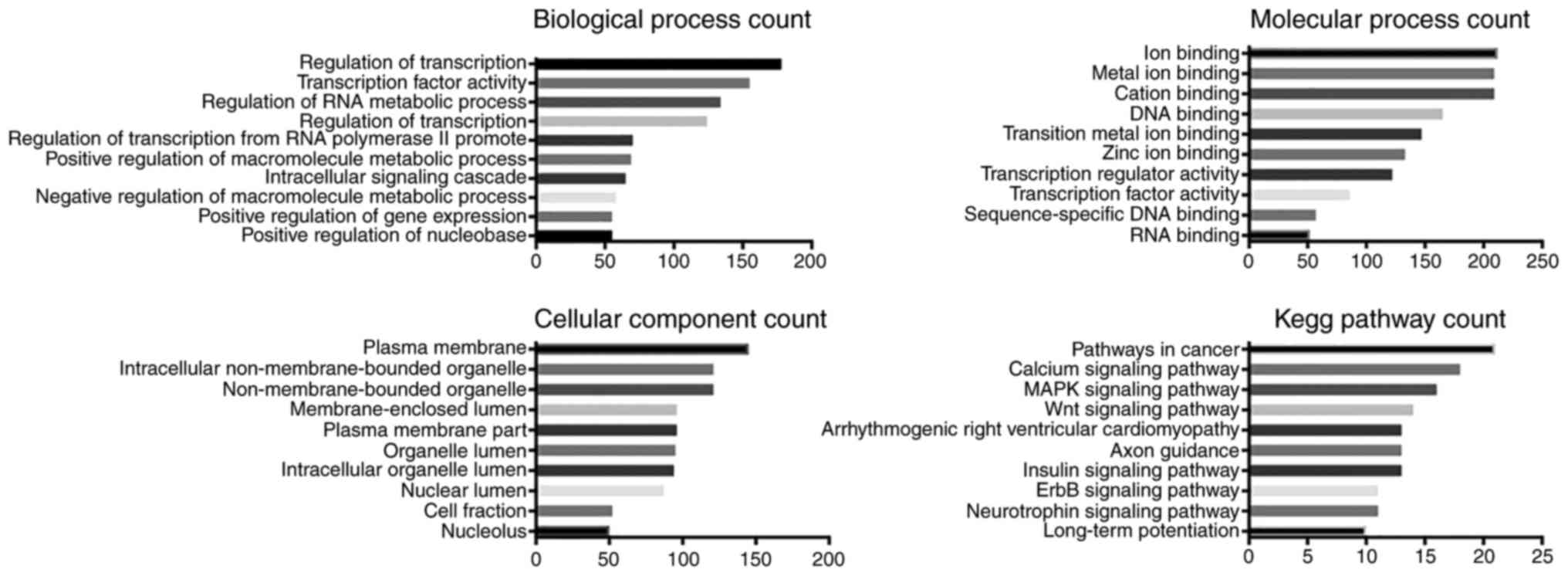
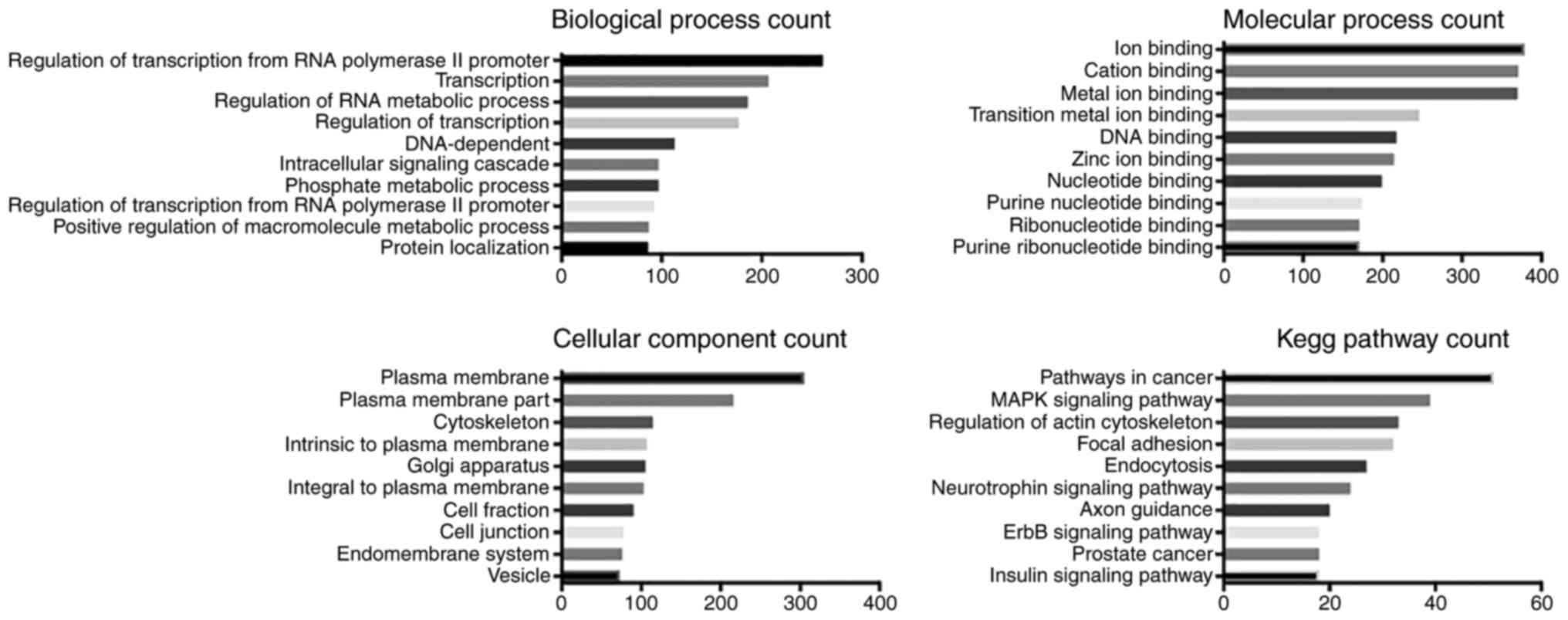
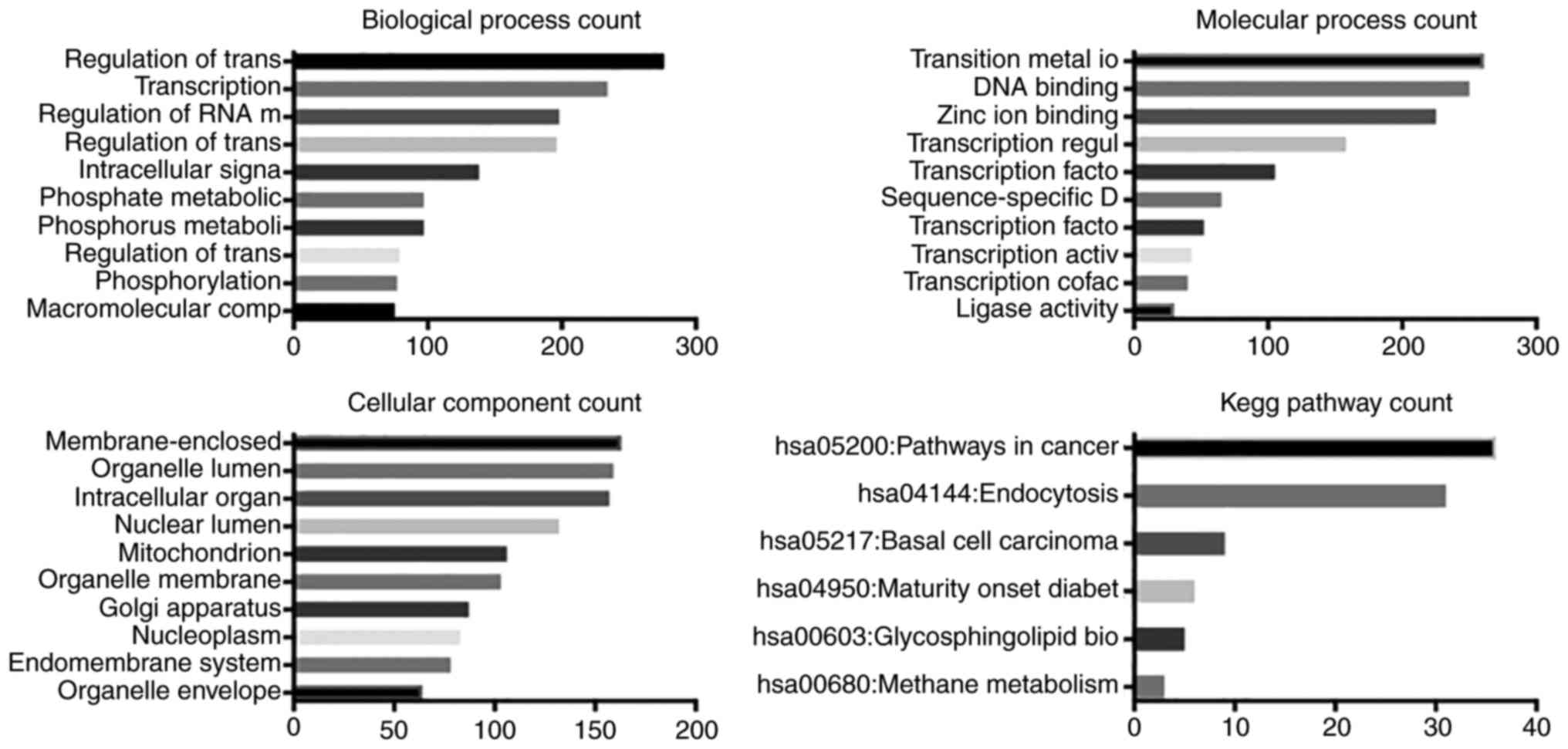
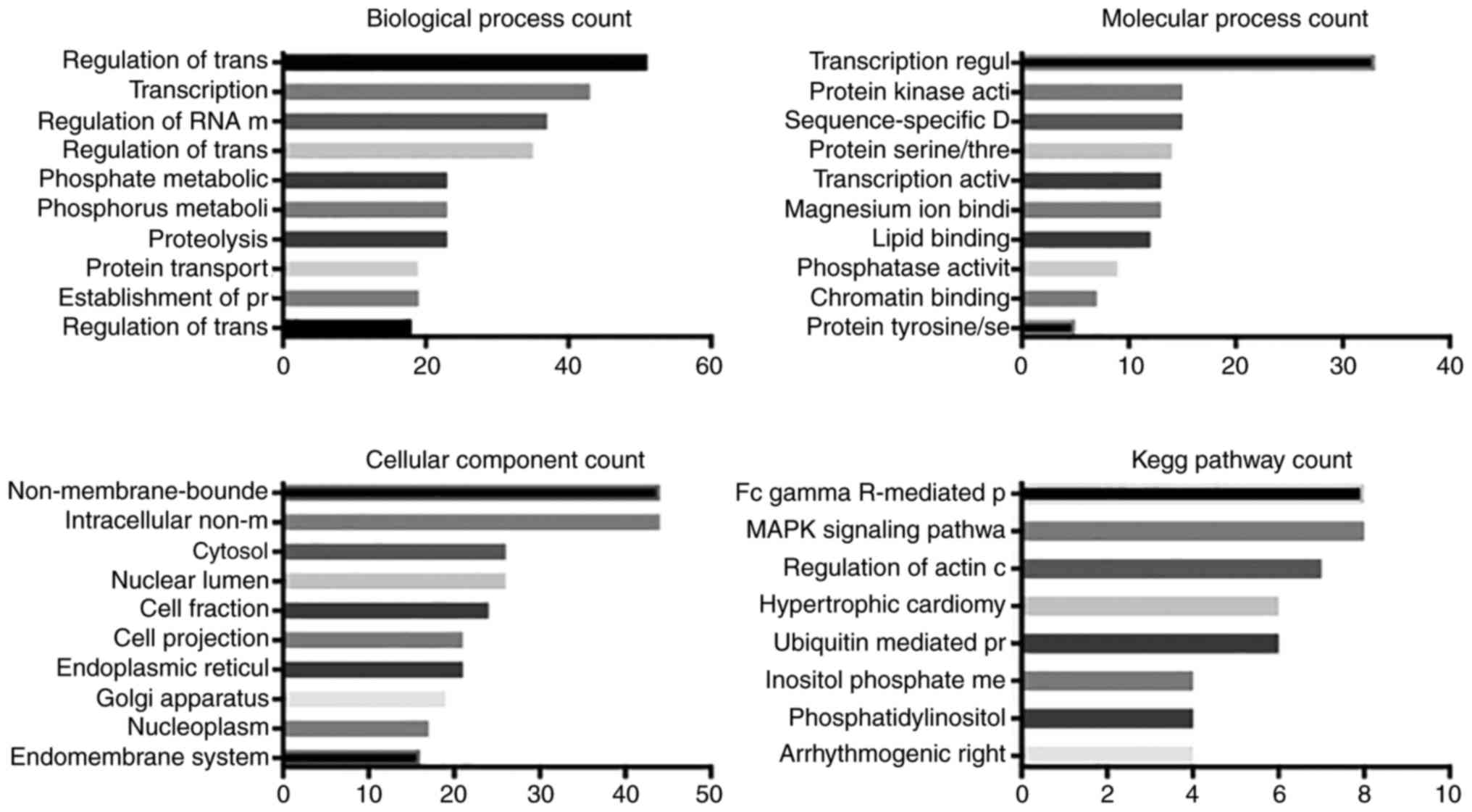
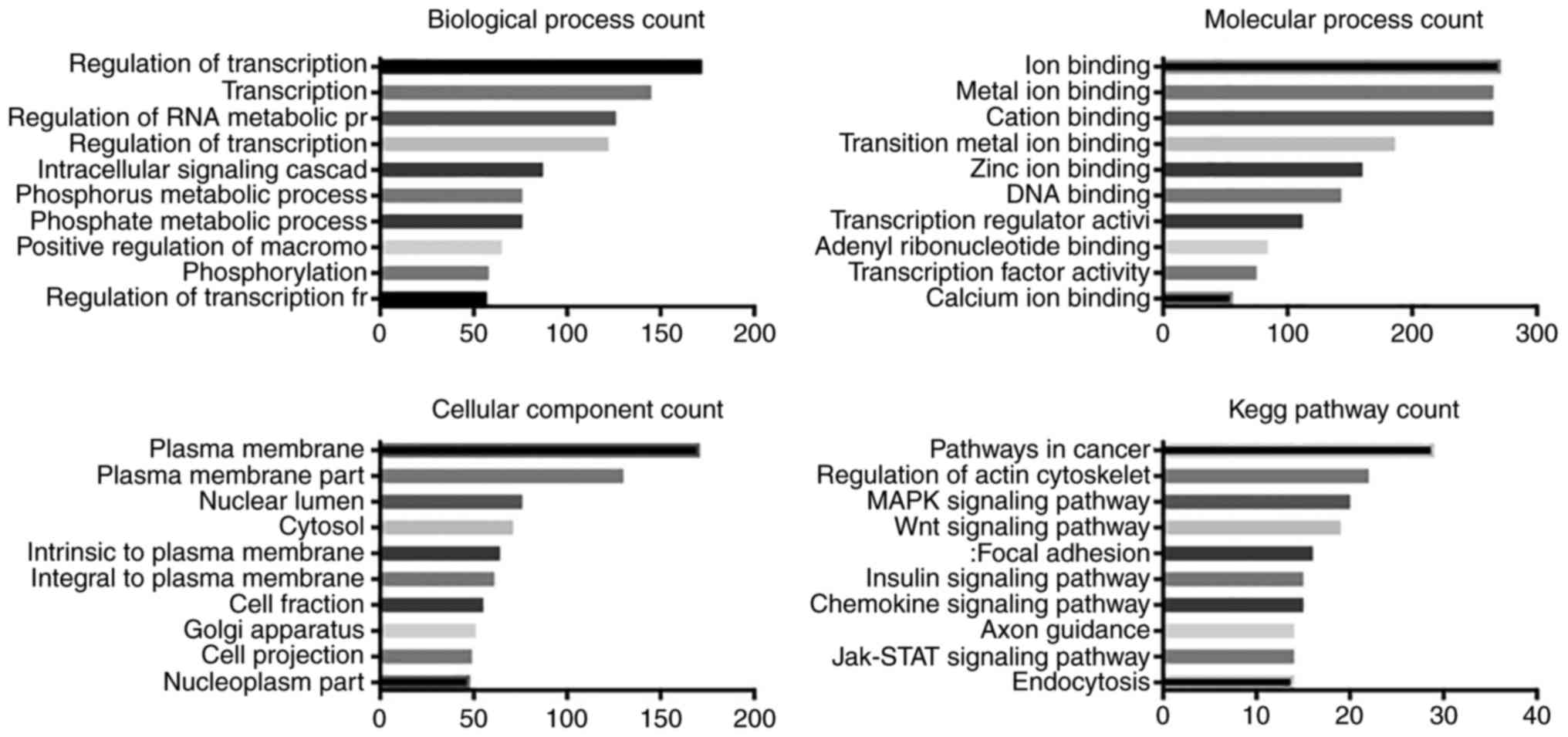
GO and KEGG pathway analysis
The predicted biological processes, molecular
function, cellular components and KEGG pathways associated with
miR-129-5p, miR-9-5p, miR-6720-3p, miR-363-3p and miR-135a-5p were
analyzed (Figs. 3Figure 4Figure 5Figure 6–7). As expected, these miRNAs were
predicted to be most frequently involved in 'regulation of
transcription' and 'RNA metabolic process'. miR-129-5p, miR-9-5p
and miR-135a-5p were predicted to function in ion binding, while
miR-6720-3p in both ion binding and DNA binding. miR-363-3p was the
only miRNA predicted to molecularly regulate transcription and
protein kinase activity. Cellular component analysis suggested that
miR-129-5p, miR-9-5p and miR-135a-5p were associated with 'plasma
membrane', miR-6720-3p in 'membrane-enclosed lumen', and miR-363-3p
in 'non-membrane bounded organelles'. KEGG pathway analysis
indicated that four miRNAs (miR-129-5p, miR-9-5p, miR-6720-3p and
miR-135a-5p) may regulate pathways in cancer.
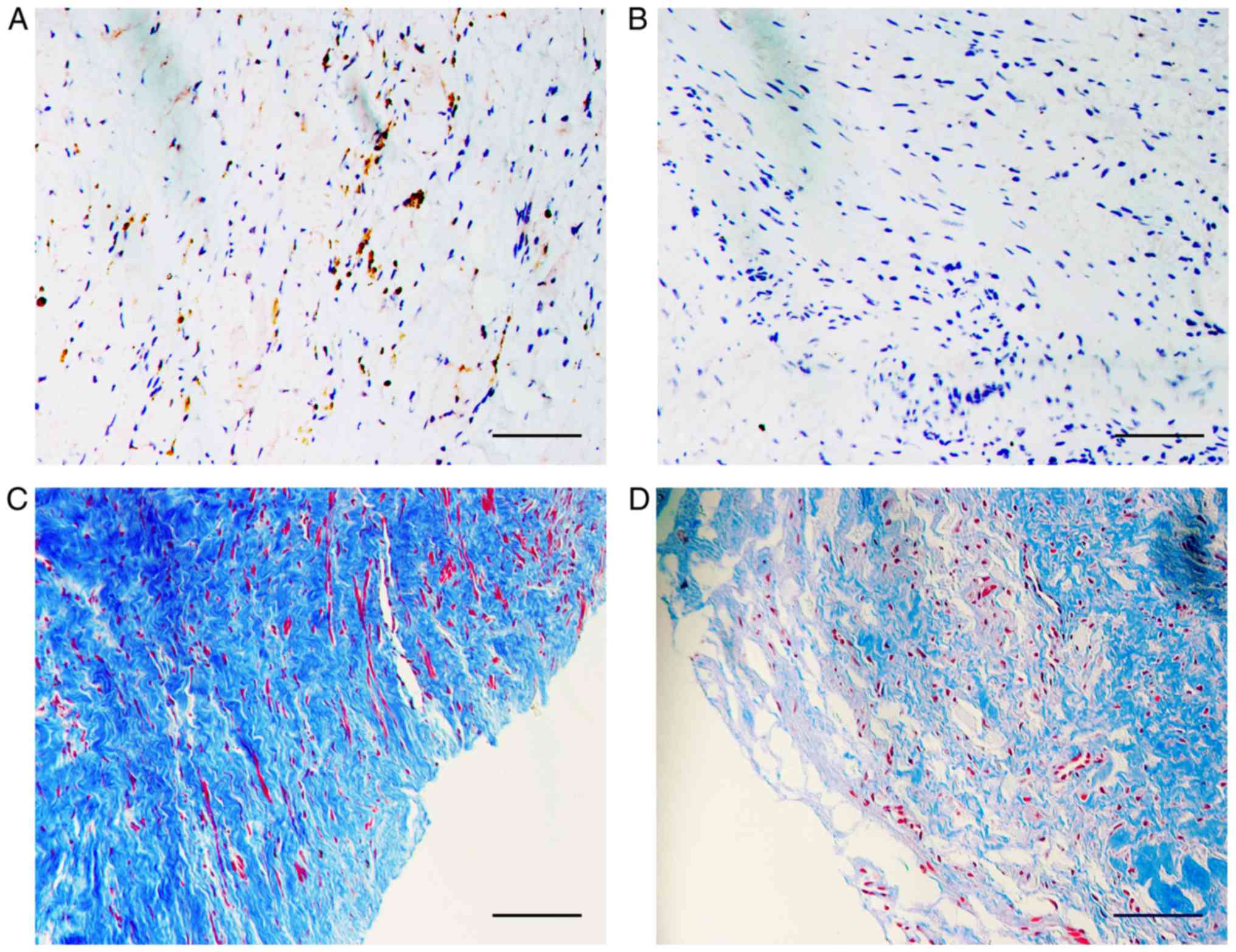
TGF-β and collagen are upregulated in
urethral scar tissue, but not normal tissue
TGF-β signaling has been confirmed to be involved in
fibrosis. To investigate whether TGF-β signaling was affected in
scar tissues, immunostaining for TGF-β1 protein was performed. As
expected, TGF-β1 expression was increased in urethral scar tissues
compared with normal tissues from the same patient (Fig. 8A and B). Similarly, dense collagen
distribution and irregular fiber alignment was observed in urethral
scar tissues, but not in normal tissue (Fig. 8C and D).
Discussion
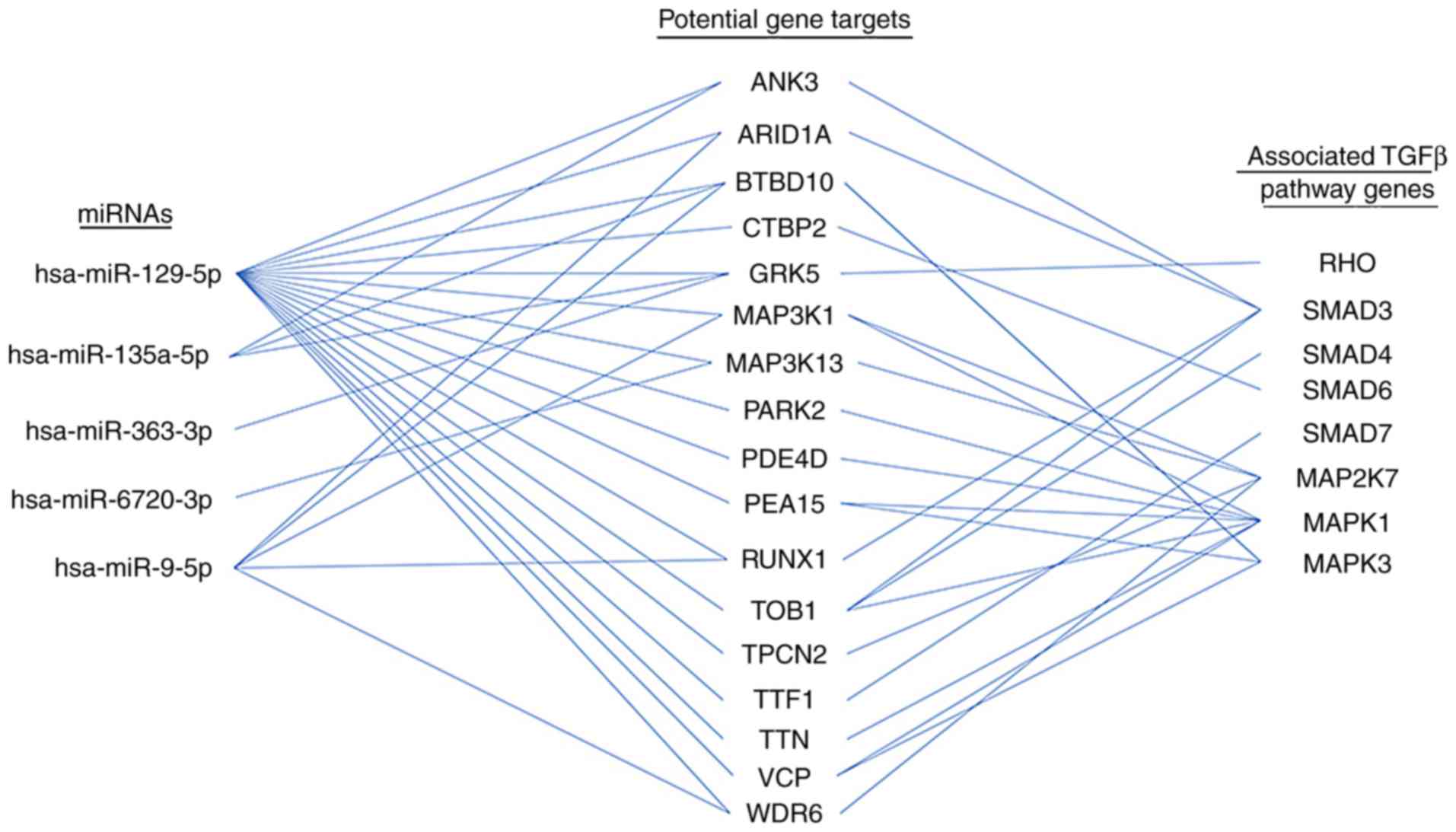
miR-129-5p potentially regulates 17 genes that are
directly associated with TGF-β pathway members (Fig. 9), indicating that it may serve an
important function in fibrosis through the regulation of TGF-β
signalling. miR-129-5p has previously been reported to be
significantly downregulated following TGF-β1 treatment in kidney
proximal tubular cells (23). In
addition, miR-129-5p is downregulated in fibroblasts in systemic
sclerosis (or scleroderma, an acquired disorder that typically
results in fibrosis of the skin and internal organs), and
upregulated miR-129-5p (induced by interleukin-17A) suppresses α1
collagen protein expression levels in normal fibroblasts (24). The present study suggested that
miR-129-5p is upregulated in scar tissues from patients with PFUDD
compared with normal tissues, and further analysis is required to
assess whether the downstream targets in TGF-β pathway are
functionally affected, and whether miR-129-5p regulation is driving
the fibrosis or results from fibrosis as a feedback effect.
Besides its fibrosis-associated functions,
miR-129-5p has also been reported as a regulator of
epithelial-to-mesenchymal transition (EMT) in epithelial cells
(23). miR-129-5p is
significantly downregulated in various types of cancer, including
lung cancer, breast cancer, gastric cancer and glioblastoma, and
over-expression of miR-129-5R suppresses cell viability,
proliferation, migration and invasion, while promoting cell
apoptosis (25–28). miR-129-5p has also been
demonstrated to negatively regulate a specific circular RNA
(hsa-circ-0005986) and Notch1, and downregulated hsa-circ-0005986
may promote the proliferation of hepatocellular carcinoma cells
(29). Investigation of
miR-129-5p and its potential targets through TGF-β-independent
regulations in PFUDD may prove interesting.
miR-9-5p expression is increased in lung tissues
from patients with idiopathic pulmonary fibrosis and in the lungs
of mice with bleomycin-induced lung fibrosis (30). Overexpression of miR-9-5p inhibits
TGF-β1-induced lung fibrosis, and inhibition of miR-9-5p amplifies
the transformation of human lung fibroblasts into myofibroblasts
(30). Overexpression of
hsa-miR-9-5p may prevent fibrogenesis in skin fibrosis, mediated by
suppressing TGF-β signaling, and inhibition of miR-9-5p results in
increased expression of fibrosis markers (31). Hsa-miR-9-5p is upregulated in
TGF-β1-treated human dermal fibroblasts (31). Overexpression of miR-9-5p
significantly delays TGF-β-dependent transformation of dermal
fibroblasts, decreases the expression of the ECM proteins collagen,
type I, α1 and fibronectin, the amount of secreted collagen
proteins and the expression of the archetypal myofibroblast marker
α-smooth muscle actin (31). In
contrast to this, specific inhibition of miR-9-5p resulted in
enhanced presence of fibrosis markers (31). The upregulation of hsa-miR-9-5p
observed in the present study may have been an induced
auto-suppression effect of fibrosis, and an analysis with larger
cohort and further validation may be helpful to understand its
function during the process.
Little is known about the involvement of miR-363-3p,
miR-135a-5p or miR-6720-3p in fibrosis regulation. The majority of
the research concerning these three miRNAs focuses on their
function in carcinogenesis. For example, miR-363-3p is
downregulated in tissues from patients with non-small cell lung
cancer and gallbladder cancer, and in cluster of differentiation
133-positive cancer stem-like cells isolated from patients with
larynx cancer (32–34). miR-363-3p significantly inhibits
cell growth in vitro and in vivo in lung cancer
cells, and induces cell cycle arrest and promote cell apoptosis,
potentially through proliferating cell nuclear antigen and its
ability to inhibit the activation of the mechanistic target of
rapamycin and extracellular signal-regulated kinase (ERK) signaling
pathways (33,35). In patients with prostate cancer,
miR-363-3p expression was decreased in serum and its level was
negatively correlated with risk of recurrence and progression,
suggesting that it may serve as a potential diagnostic and
prognostic biomarker for prostate cancer (36). Similarly, in colorectal cancer,
miR-363-3p was downregulated in the tissues of patients, and its
expression was negatively associated with metastatic status
(37). miR-363-3p inhibits
colorectal cancer cell migration and invasion and inhibits EMT,
potentially through targeting SRY-box 4 (37). Inhibition of miR-363-3p induces
cell cycle arrest and suppresses apoptosis in gallbladder cancer
cells, potentially by targeting and competing with the long
non-coding RNA metastasis associated lung adenocarcinoma transcript
1 in the regulation of the MCL1, BCL2 family apoptosis regulator
gene (34).
miR-135a-5p is upregulated in liver biopsy samples
from patients with chronic hepatitis C or hepatitis C virus (HCV),
but is not induced by anti-viral response to HCV infection
(38). By targeting and inducing
silencing of protein tyrosine phosphatase receptor δ, a tumor
suppressor, miR-135a-5p may potentially drive HCV-associated
hepatocarcinogenesis (38).
miR-6720 has been reported to be upregulated in
hepatocellular carcinoma cells following treatment with
alternariol, a mycotoxin (39).
However, not much evidence is available concerning its functions in
TGF-β regulated signaling transduction.
As a shared potential target of hsa-miR-129-5p,
hsa-miR-135a-5p and hsa-miR-363-3p, the G protein-coupled receptor
kinase 5 (GRK5) gene encodes a member of the guanine
nucleotide-binding protein-coupled receptor kinase subfamily of the
Ser/Thr protein kinase family. GRK5 mRNA and protein expression is
increased in the lungs of patients with cystic fibrosis compared
with normal donors (40). GRK5
has been identified as a regulator of aldosterone-mineralocorticoid
receptor/angiotensin II type 1 receptor-mediated cardiomyocyte
hypertrophy (41). Molecular
analysis has revealed that overexpression of N-terminal GRK5
inhibits apoptosis and fibrosis, which are major pathogenic
phenotypes that accompany cardiac hypertrophy, through nuclear
factor-κB signaling (42).
Knockout of GRK5 inhibits cardiac fibrosis induced by metoprolol, a
β1-adrenergic receptor-selective blocker, and this
regulation is dependent on β-arrestins but independent of G-protein
signaling (43).
TGF-β1 and collagen were overexpressed in scar
tissues in the present study (Fig.
8), indicating that the TGF-β pathway was upregulated. TGF-β1
is a key regulatory molecule in the pathogenesis of fibrosis, and
is frequently upregulated during this process (44,45). Inhibition of TGF-β1 or its
receptors, including TGF-β receptor 1, may reverse the unilateral
ureteral occlusion-induced increase of fibrotic biomarkers,
including TGF-β1, collagens and fibronectin, at the mRNA level
(44). The increase of TGF-β1 and
collagen may result in excessive fibrosis in scar tissue, and it is
unclear how the regulatory miRNAs and genes that regulate TGF-β
signaling are involved in this process. Potential crosstalk between
miRNAs and their target genes are listed, together with the
associated signaling molecules involved in TGF-β1 regulated
fibrosis (Fig. 9).
It is worth noting that mitogen-activated protein
kinase 1, proliferation and apoptosis adaptor protein 15,
transducer of ERBB2, 1 and valosin containing protein gene products
are predicted to interact with one or more TGF-β pathway proteins,
as presented in Fig. 9. Among the
associated TGF-β pathway genes, SMAD family member 3,
mitogen-activated protein kinase 7 and ERK1/2 are the most
frequently targeted candidates for potential interactions. Given
the fact that crosstalk between SMADs and the ERK signaling pathway
is essential in fibrosis (46),
further functional analysis focusing on those 'hotspots' in larger
cohorts may prove interesting.
In conclusion, the present study sequenced the
urethral scar and normal tissue from patients with urethral
stricture. miRNA sequencing indicated significantly altered
expression of hsa-miR-129-5p, hsa-miR-135a-5p, hsa-miR-363-3p,
hsa-miR-6720-3p and hsa-miR-9-5p. Bioinformatics analysis revealed
that these miRNAs and their potential target genes are associated
with fibrosis in other diseases. These data may provide miRNA
targets for future precision treatments for urethral stricture.
Acknowledgments
Financial support was obtained from the National
Natural Science Fund of China (grant nos. 81470917 and 81700590),
the Science and Technology Commission of Shanghai (grant no.
17410742800), and the Shanghai JiaoTong University Biomedical
Engineering Cross Research Foundation (grant no. YG2017QN15).
Notes
[1] Competing
interests
The authors declare that they have no competing
interests.
References
|
1
|
Tritschler S, Roosen A, Füllhase C, Stief
CG and Rübben H: Urethral stricture: Etiology, investigation and
treatments. Dtsch Arztebl Int. 110:220–226. 2013.PubMed/NCBI
|
|
2
|
Osman NI, Hillary C, Bullock AJ, MacNeil S
and Chapple CR: Tissue engineered buccal mucosa for urethroplasty:
Progress and future directions. Adv Drug Deliv Rev. 82–83:69–76.
2015. View Article : Google Scholar
|
|
3
|
Hampson LA, McAninch JW and Breyer BN:
Male urethral strictures and their management. Nat Rev Urol.
11:43–50. 2014. View Article : Google Scholar :
|
|
4
|
Andrich DE, Dunglison N, Greenwell TJ and
Mundy AR: The long-term results of urethroplasty. J Urol.
170:90–92. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lumen N, Hoebeke P and Oosterlinck W:
Urethroplasty for urethral strictures: Quality assessment of an
in-home algorithm. Int J Urol. 17:167–174. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Nerli RB, Neelagund SE, Guntaka A, Patil
S, Hiremath SC, Jali SM, Vernekar R and Hiremath MB: Staged buccal
mucosa urethroplasty in reoperative hypospadias. Indian J Urol.
27:196–199. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Zhang K, Qi E, Zhang Y, Sa Y and Fu Q:
Efficacy and safety of local steroids for urethra strictures: A
systematic review and meta-analysis. J Endourol. 28:962–968. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Dubey D: The current role of direct vision
internal urethrotomy and self-catheterization for anterior urethral
strictures. Indian J Urol. 27:392–396. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
O'Reilly S: MicroRNAs in fibrosis:
Opportunities and challenges. Arthritis Res Ther. 18:112016.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Griffiths-Jones S: The microRNA registry.
Nucleic Acids Res. 32:D109–D111. 2004. View Article : Google Scholar :
|
|
11
|
Griffiths-Jones S, Grocock RJ, van Dongen
S, Bateman A and Enright AJ: miRBase: microRNA sequences, targets
and gene nomenclature. Nucleic Acids Res. 34:D140–D144. 2006.
View Article : Google Scholar :
|
|
12
|
Griffiths-Jones S, Saini HK, van Dongen S
and Enright AJ: miRBase: Tools for microRNA genomics. Nucleic Acids
Res. 36:D154–D158. 2008. View Article : Google Scholar :
|
|
13
|
Kozomara A and Griffiths-Jones S: miRBase:
Annotating high confidence microRNAs using deep sequencing data.
Nucleic Acids Res. 42:D68–D73. 2014. View Article : Google Scholar :
|
|
14
|
Kozomara A and Griffiths-Jones S: miRBase:
Integrating microRNA annotation and deep-sequencing data. Nucleic
Acids Res. 39:D152–D157. 2011. View Article : Google Scholar :
|
|
15
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2−ΔΔCT method. Methods. 25:402–408. 2001. View Article : Google Scholar
|
|
16
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Enright AJ, John B, Gaul U, Tuschl T,
Sander C and Marks DS: MicroRNA targets in Drosophila. Genome Biol.
5:R12003. View Article : Google Scholar
|
|
18
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinfor-matics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar
|
|
19
|
Huang da W, Sherman BT and Lempicki RA:
Bioinformatics enrichment tools: Paths toward the comprehensive
functional analysis of large gene lists. Nucleic Acids Res.
37:1–13. 2009. View Article : Google Scholar
|
|
20
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The Gene
Ontology Consortium. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
The Gene Ontology Consortium: Expansion of
the Gene Ontology knowledgebase and resources. Nucleic Acids Res.
45:D331–D338. 2017. View Article : Google Scholar :
|
|
22
|
Kanehisa M and Goto S: KEGG: Kyoto
encyclopedia of genes and genomes. Nucleic Acids Res. 28:27–30.
2000. View Article : Google Scholar
|
|
23
|
Li Y, An H, Pang J, Huang L, Li J and Liu
L: MicroRNA profiling identifies miR-129-5p as a regulator of EMT
in tubular epithelial cells. Int J Clin Exp Med. 8:20610–20616.
2015.
|
|
24
|
Nakashima T, Jinnin M, Yamane K, Honda N,
Kajihara I, Makino T, Masuguchi S, Fukushima S, Okamoto Y, Hasegawa
M, et al: Impaired IL-17 signaling pathway contributes to the
increased collagen expression in scleroderma fibroblasts. J
Immunol. 188:3573–3583. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang Y, An J, Lv W, Lou T, Liu Y and Kang
W: miRNA-129-5p suppresses cell proliferation and invasion in lung
cancer by targeting microspherule protein 1, E-cadherin and
vimentin. Oncol Lett. 12:5163–5169. 2016. View Article : Google Scholar
|
|
26
|
Xu H, Hu Y and Qiu W: Potential mechanisms
of microRNA-129-5p in inhibiting cell processes including
viability, proliferation, migration and invasiveness of
glioblastoma cells U87 through targeting FNDC3B. Biomed
Pharmacother. 87:405–411. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Luan QX, Zhang BG, Li XJ and Guo MY:
MiR-129-5p is down-regulated in breast cancer cells partly due to
promoter H3K27m3 modification and regulates epithelial-mesenchymal
transition and multi-drug resistance. Eur Rev Med Pharmacol Sci.
20:4257–4265. 2016.PubMed/NCBI
|
|
28
|
Jiang Z, Wang H, Li Y, Hou Z, Ma N, Chen
W, Zong Z and Chen S: MiR-129-5p is down-regulated and involved in
migration and invasion of gastric cancer cells by targeting
interleukin-8. Neoplasma. 63:673–680. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Fu L, Chen Q, Yao T, Li T, Ying S, Hu Y
and Guo J: Hsa_circ_0005986 inhibits carcinogenesis by acting as a
miR-129-5p sponge and is used as a novel biomarker for
hepatocellular carcinoma. Oncotarget. 8:43878–43888.
2017.PubMed/NCBI
|
|
30
|
Fierro-Fernández M, Busnadiego Ó, Sandoval
P, Espinosa-Díez C, Blanco-Ruiz E, Rodríguez M, Pian H, Ramos R,
López-Cabrera M, García-Bermejo ML and Lamas S: miR-9-5p suppresses
pro-fibrogenic transformation of fibroblasts and prevents organ
fibrosis by targeting NOX4 and TGFBR2. EMBO Rep. 16:1358–1377.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Miguel V, Busnadiego O, Fierro-Fernández M
and Lamas S: Protective role for miR-9-5p in the fibrogenic
transformation of human dermal fibroblasts. Fibrogenesis Tissue
Repair. 9:72016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Karatas OF, Suer I, Yuceturk B, Yilmaz M,
Oz B, Guven G, Cansiz H, Creighton CJ, Ittmann M and Ozen M:
Identification of microRNA profile specific to cancer stem-like
cells directly isolated from human larynx cancer specimens. BMC
Cancer. 16:8532016. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Wang Y, Chen J, Lin Z, Cao J, Huang H,
Jiang Y, He H, Yang L, Ren N and Liu G: Role of deregulated
microRNAs in non-small cell lung cancer progression using
fresh-frozen and formalin-fixed, paraffin-embedded samples. Oncol
Lett. 11:801–808. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wang SH, Zhang WJ, Wu XC, Weng MZ, Zhang
MD, Cai Q, Zhou D, Wang JD and Quan ZW: The lncRNA MALAT1 functions
as a competing endogenous RNA to regulate MCL-1 expression by
sponging miR-363-3p in gallbladder cancer. J Cell Mol Med.
20:2299–2308. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wang Y, Chen T, Huang H, Jiang Y, Yang L,
Lin Z, He H, Liu T, Wu B, Chen J, et al: miR-363-3p inhibits tumor
growth by targeting PCNA in lung adenocarcinoma. Oncotarget.
8:20133–20144. 2017.PubMed/NCBI
|
|
36
|
Cochetti G, Poli G, Guelfi G, Boni A,
Egidi MG and Mearini E: Different levels of serum microRNAs in
prostate cancer and benign prostatic hyperplasia: Evaluation of
potential diagnostic and prognostic role. OncoTargets Ther.
9:7545–7553. 2016. View Article : Google Scholar
|
|
37
|
Hu F, Min J, Cao X, Liu L, Ge Z, Hu J and
Li X: MiR-363-3p inhibits the epithelial-to-mesenchymal transition
and suppresses metastasis in colorectal cancer by targeting Sox4.
Biochem Biophys Res Commun. 474:35–42. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Van Renne N, Roca Suarez AA, Duong FH,
Gondeau C, Calabrese D, Fontaine N, Ababsa A, Bandiera S,
Croonenborghs T, Pochet N, et al: miR-135a-5p-mediated
downregulation of protein tyrosine phosphatase receptor delta is a
candidate driver of HCV-associated hepatocarcinogenesis. Gut:
gutjnl. 2016:3122702017.
|
|
39
|
Vejdovszky K, Sack M, Jarolim K, Aichinger
G, Somoza MM and Marko D: In vitro combinatory effects of the
Alternaria mycotoxins alternariol and altertoxin II and potentially
involved miRNAs. Toxicol Lett. 267:45–52. 2017. View Article : Google Scholar
|
|
40
|
Mak JC, Chuang TT, Harris CA and Barnes
PJ: Increased expression of G protein-coupled receptor kinases in
cystic fibrosis lung. Eur J Pharmacol. 436:165–172. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Cannavo A, Liccardo D, Eguchi A, Elliott
KJ, Traynham CJ, Ibetti J, Eguchi S, Leosco D, Ferrara N, Rengo G
and Koch WJ: Myocardial pathology induced by aldosterone is
dependent on non-canonical activities of G protein-coupled receptor
kinases. Nat Commun. 7:108772016. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Sorriento D, Santulli G, Fusco A,
Anastasio A, Trimarco B and Iaccarino G: Intracardiac injection of
AdGRK5-NT reduces left ventricular hypertrophy by inhibiting
NF-κB-dependent hyper-trophic gene expression. Hypertension.
56:696–704. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Nakaya M, Chikura S, Watari K, Mizuno N,
Mochinaga K, Mangmool S, Koyanagi S, Ohdo S, Sato Y, Ide T, et al:
Induction of cardiac fibrosis by β-blocker in G protein-independent
and G protein-coupled receptor kinase 5/β-Arrestin2-dependent
signaling pathways. J Biol Chem. 287:35669–35677. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Richards TL, Minor K and Plato CF: Effects
of transforming growth factor-beta (TGF-β) receptor I inhibition on
renal biomarkers and fibrosis in unilateral ureteral occluded (UUO)
Mice. FASEB J. 31(1030): 10332017.
|
|
45
|
Penn JW, Grobbelaar AO and Rolfe KJ: The
role of the TGF-β family in wound healing, burns and scarring: A
review. Int J Burns Trauma. 2:18–28. 2012.
|
|
46
|
Zhan M and Kanwar YS: Hierarchy of
molecules in TGF-β signaling relevant to myofibroblast activation
and renal fibrosis. Am J Physiol Renal Physiol. 307:F385–F387.
2014. View Article : Google Scholar : PubMed/NCBI
|