Introduction
Breast cancer-specific gene 1 (BCSG1) is a group of
specific expression genes in human breast cancers (1–5). The
high expression of BCSG1 can promote the survival of tumor cells in
adverse environment and reduce the effect of chemotherapeutics
(6–9), thus making BCSG1 a potential target
for breast cancer treatment (10–12).
RNA interference (RNAi), as a gene-blocking technology widely used
in the regulation of gene expression, has provided a new gene
therapy for cancer and other serious diseases. However, RNAi is
degradable in vivo and off-target distribution, which limits
its application in cancer therapy. Therefore, the encapsulation and
release technique for RNAi molecules has become one of the key
issues in this field.
The recent decades have witnessed the emergence of
some packaging materials (13–18),
among which, chitosan, as a basic polysaccharide, is biocompatible,
biodegradable, nontoxic, showing good tolerance in the human body
(19–27). In addition, the positive charged
chitosan nanoparticles are compatible with the RNAi molecules.
Moreover, mesoporous silica nanoparticle has exhibited its high
drug loading capability and low-toxicity degradation (28,29).
Arginine-glycine-aspartate (RGD) peptide modification significantly
increases the selectivity between cancer and normal cells, through
receptor-mediated endocytosis (30,31).
Taken together, RGD labeled chitosan-silicon dioxide nanoparticles
are particularly prominent material for RNAi packing and
releasing.
In this study, through comprehensive assessment on
BCSG1 genes, the BCSG1-small interference RNA (siRNA) plasmid has
been designed and synthesized for breast cancer treatment.
Moreover, chitosan-silicon dioxide is selected as targeted carrier
to encapsulate BCSG1-siRNA. Finally, the feasibility of
chitosan-silicon dioxide as gene carrier will be analyzed from the
perspectives of its encapsulation and releasing efficiency.
Materials and methods
Materials
Chitosan (MW: 100–150 kDa, DD, 85%;
Amresco, Inc., Solon, OH, USA). Tetraethylorthosilicate (TEOS) and
RGD was supplied by Shanghai Biological Technology Co. (Shanghai,
China). MCF-7 cells were provided by Shanghai Cell Library
(Shanghai, China). Fetal bovine serum, penicillin, streptomycin and
trypsin were purchased from Chongqing Biological Pharmaceutical
Co., Ltd. (Chongqing, China), pGPU6 carrier and RNAi turned dye
reagents were obtained from Shanghai Zimmer Pharmaceutical
Technology Co., Ltd. (Shanghai, China). PCR product recycling
reagents box and BCSG1 probe were provided by Beijing Dingguo
Changsheng Biotechnology Co., Ltd. (Beijing, China).
The design of BCSG1-siRNA
According to the BCSG1 cDNA sequences [provided by
Ambion; Thermo Fisher Scientific, Inc. (Waltham, MA, USA) and
Takara Bio, Inc. (Otsu, Japan) website design and analysis
software], four siRNA sequences and one control sequences of siRNA
have been designed as follows: BCSG1-siRNA-1 sense strand,
5′-TGGTGAGCAGCGTCAACACTGT-3 and antisense strand,
5′-UUUGUGCAGCCAACCCUCCTT-3′; BCSGI-siRNA-2 sense strand,
5′-CCAAGGAGAATGTTGTACAGATT-3′ and antisense strand,
5′-UUUGUGCAGCCAACCCUCCTT-3′; BCSG1-siRNA-3 sense strand,
5′-CAAGACCAAGGAGAATGTTGTTT-3′ and antisense strand,
5′-UUUGUGCAGCCAACCCUCCTT-3′; BCSG1-siRNA-4 sense strand,
5′-GCCAAGACCAAGGAGAAIGTTTT-3′ and antisense strand,
5′-UUUGUGCAGCCAACCCUCCTT-3′; negative control siRNA sense strand,
5′-GTTCTCCGAACGTGTCACGTTT-3′ and antisense strand,
5′-ACGUGACACGUUCGGAGAATT-3′
Preparation of chitosan-silicon
dioxide/BCSG1-siRNA nanoparticles
RGD-silica nanoparticles were prepared by sol-gel
method. Briefly, cyclohexane (7.5 ml) was mixed with hexyl alcohol
(1.6 ml) uniformly, then RGD peptide was added as core material
under constant stirring for 30 min at room temperature. Then
ammonia (100 µl) and TEOS (100 µl) were added drop wise to the
mixture over a period of 24 h with stirring moderately. The
products were collected by centrifugation, then washed by ethanol
and water successively to remove the template cyclohexane.
To obtain chitosan-silicon dioxide/BCSG1-siRNA
nanoparticles, 1% chitosan (100 µl) was added into the RGD
peptide-silicon dioxide under stirring for 30 min at 180 r/min.
Then preheated the equal volume of chitosan/silicon dioxide and
BCSG1-siRNA plasmid at 55°C, and mixed rapidly under stirring for
30 min at room temperature. Then a certain concentration of sodium
tripolyphosphate solution (5 ml) was added into the mixture under
stirring for 3.5 h at room temperature, and set over night.
Finally, the nanoparticles were collected by centrifugation, then
filtered by distilled water and lyophilized about 24 h.
Characterization of chitosan-silicon
dioxide/BCSG1-siRNA
The nanoparticles were characterized by laser
particle size analyzer transmission electron microscope (TEM),
scanning electron microscope (SEM), UV-visible spectroscopy and
fluorescence spectroscopy.
The encapsulation efficiency of BCSG1-siRNA was
measured in accordance with equation [1] (32–36):
Encapsulation efficiency = (C0 –
Ct)/C0 × 100% [1].
C0 was the original amount of RNA before
encapsulation, Ct was the remanent amount of RNA in
supernatants after encapsulation.
The BCSG1-siRNA chitosan nanoparticles and
phosphate-buffered saline (PBS) (pH 7.4) were put in cell
incubators at 37°C, respectively. Then the supernatant was taken to
calculate the released BCSG1-siRNA concentration by using
UV-spectrophotometer after 12, 24, 36, 48, 60 and 72 h. The release
efficiency of BCSG1-siRNA was measured by equation [2]:
Release rate = C2/C1 × 100%
[2].
C1 and C2 were the quantity of
BCSG1-siRNA in PBS before and after incubation, respectively.
BCSG1 protein expression by
immunocytochemistry detection
According to SP immunohistochemistry promega, PBS
was used as a negative control, and the positive signal of BCSG1
protein expression as a positive control.
Results showed the positive signal of BCSG1 is a
brown granular substance, located within the cytoplasm. Ten
microscopic fields were randomly selected under high magnification
microscope (the amount of the cells are no less than 100). The
results were caculated by the percentage of positive cells and the
color intensity (37).
Semi-quantitative RT-PCR
DNase-treated RNA was extracted and used as a
template, following the reverse transcription kit for cDNA
synthesis.
The loop parameter of PCR was shown in Table I, it was about 35 cycles. The
agarose electrophoresis of PCR product: PCR product (3–5 µl) was
mixed with loading buffer (1 µl), then the point sample was added
to 2% agarose gel. And voltage was adjusted to 5 V/cm about 30 min
in the 0.5 X TBE electrophoresis fluids. Finally, PCR products
electrophoresis strips were observed by gel imaging system. The
melting curve was obtained by quantitative analysis software of PCR
instrument (38–40).
 | Table I.The loop parameter of polymerase
chain reaction. |
Table I.
The loop parameter of polymerase
chain reaction.
| Reaction
conditions | Temperature
(°C) | Reaction time
(min) |
|---|
| Initial
denaturation | 94 | 2 |
| Denaturation | 94 | 0.5 |
| Annealing | 56 | 0.5 |
| Extension | 72 | 0.5 |
| Preservation | 72 | 10 |
In vitro cellular uptake assays
The BCSG1-siRNA chitosan nanoparticles were diluted
to 10, 5, 1 and 0.5 µg/ml. MCF-7 breast cancer cells were divided
into 6 groups and treated with the indicated concentrations of
BCSG1-siRNA chitosan nanoparticles, further incubated for a week.
At 2 days interval, culture medium and BCSG1-siRNA chitosan
nanoparticles were replenished. Afterward, the amount of MCF-7
cells in the five groups was measured by inverted microscope. Cell
suspension (20 µl) was dropped into the counting slide and abserved
by the 10X objective lens. The experiments were carried out in
triplicate and were independently repeated at least 3 times. Then
the proliferation rate and apoptosis rate (equations 3 and 4) were
determined to evaluate the effect of BCSG1-siRNA chitosan
nanoparticles to MCF-7 cells.
Proliferation rate = (N0 –
N1)/N0 × 100% [3].
N1 and was N0 were the number
of cells in experimental group and control group.
Apoptosis rate = N2/(N2 + N) ×
100% [4].
N2 and N were the number of apoptotic
cells apoptosis and normal cells.
Statistical analysis
All the quantitative data have been presented as the
mean ± standard deviation. Analysis of variance with Tukey post hoc
test was used to evaluate statistical significance among different
groups using SPSS software version 20 (SPSS, Inc., Chicago, IL,
USA). Significance was considered when P<0.05.
Results
The immunocytochemistry analysis
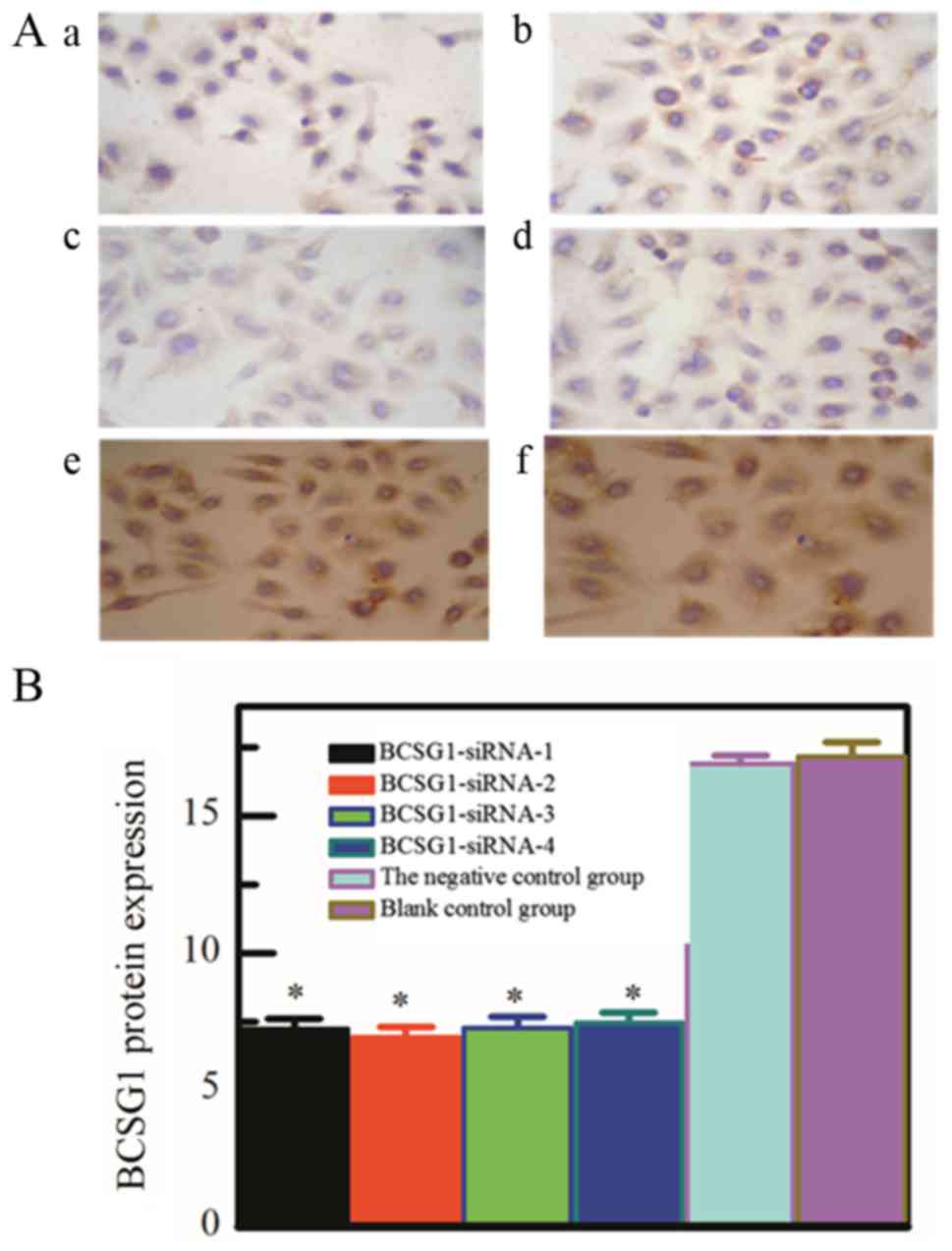
The immunocytochemistry result (Fig. 1) has shown that the BCSG1 protein
expression of the four specific siRNA transfection groups, the
negative control group and blank control group was 7.22, 6.93,
7.26, 7.46, 16.89 and 17.17. The differences among the four
specific siRNA transfection groups were not significant
(P>0.05), while the differences between the former four groups
and control groups had statistical significance (P<0.05).
Compared with negative control group and blank
control group, the protein expression of BCSG1 in the four specific
siRNA transfection groups is significantly lower. The results
showed that the designed siRNA transfection groups could
specifically downregulated the protein expression of BCSG1.
The PCR analysis of mRNA
expression
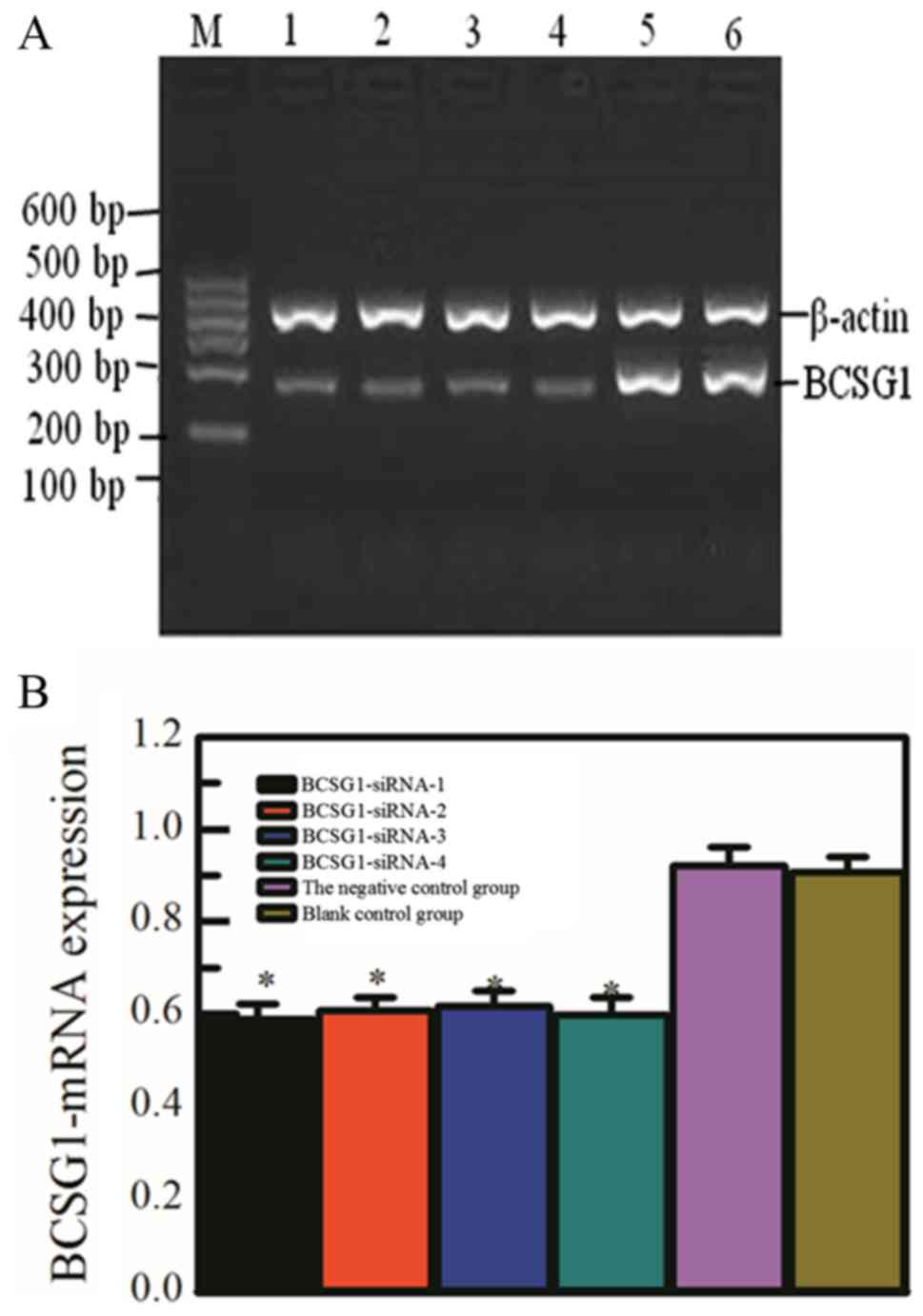
RNA for reverse transcription, PCR amplification and
electrophoresis in gel image system photo were experimented after
transfection (Fig. 2), including
the four specific siRNA transfection groups, the negative control
group and blank control group.
The first four groups were significantly darkened
strip, showed the lower expression of BCSG1-mRNA. Compared with the
gray value of β-actin mRNA, the PCR result indicated that the
relative transcript level of mRNA in each group was 0.589±0.033,
0.608±0.029, 0.617±0.033, 0.598±0.037, 0.922±0.040, 0.908±0.031, it
can be seen that there were no significant differences among the
former four groups (P>0.05), but the difference between the
former four groups and control groups had statistical significance
(P<0.05).
Compared with negative control group and blank
control group, the mRNA expression of BCSG1 in the four specific
siRNA transfection groups is significantly lower. The results
showed that the designed siRNA transfection groups could
specifically downregulated the mRNA expression of BCSG1.
The characteristic of chitosan
nanoparticle
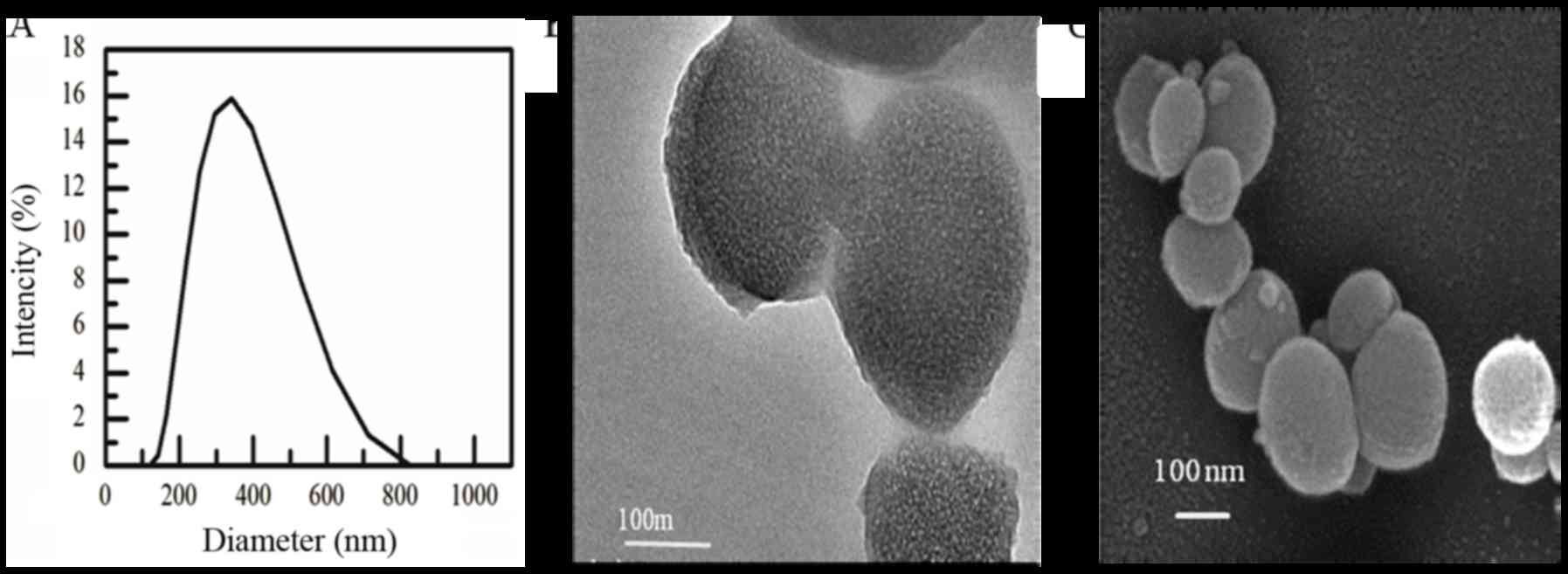
According to the laser particle size Analyzer test
(Fig. 3A), results showed that the
particle sizes was between 200–600 nm, average out at 303±51 nm,
the polydispersity index was 0.18. However, the shapes and sizes of
nanoparticles after freeze drying were irregular sphere, whose size
was slightly smaller than 200 nm from the TEM and SEM (Fig. 3B and C).
According to the UV-Spectrophotometric analysis, the
encapsulation rate of chitosan nanoparticles was 94.23±0.43%,
showing that most of the DNA are encapsulated in chitosan
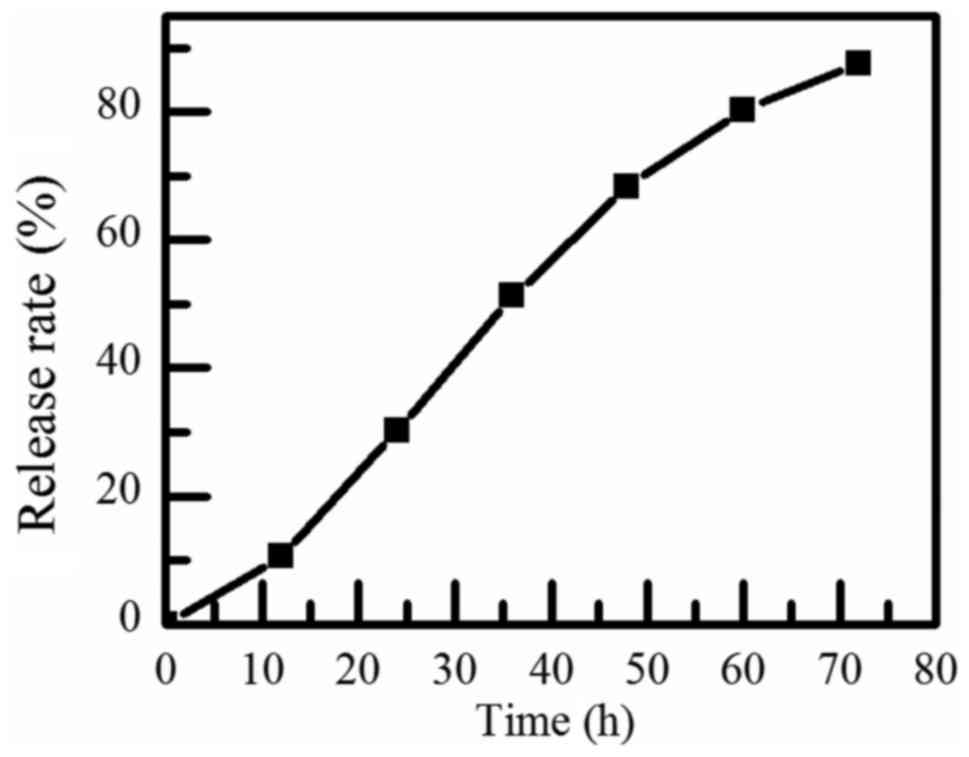
nanoparticles, the encapsulation effect was good. Similarly, the
release efficiency of BCSG1-siRNA was measured and drawn into the
release curve, shown in Fig. 4.
The release rate was almost 30% after 24 h, and the releasing
effect of BCSG1-siRNA nanoparticles was sustained. It has a good
application in drug controlled release.
The proliferation and apoptosis rate
assay
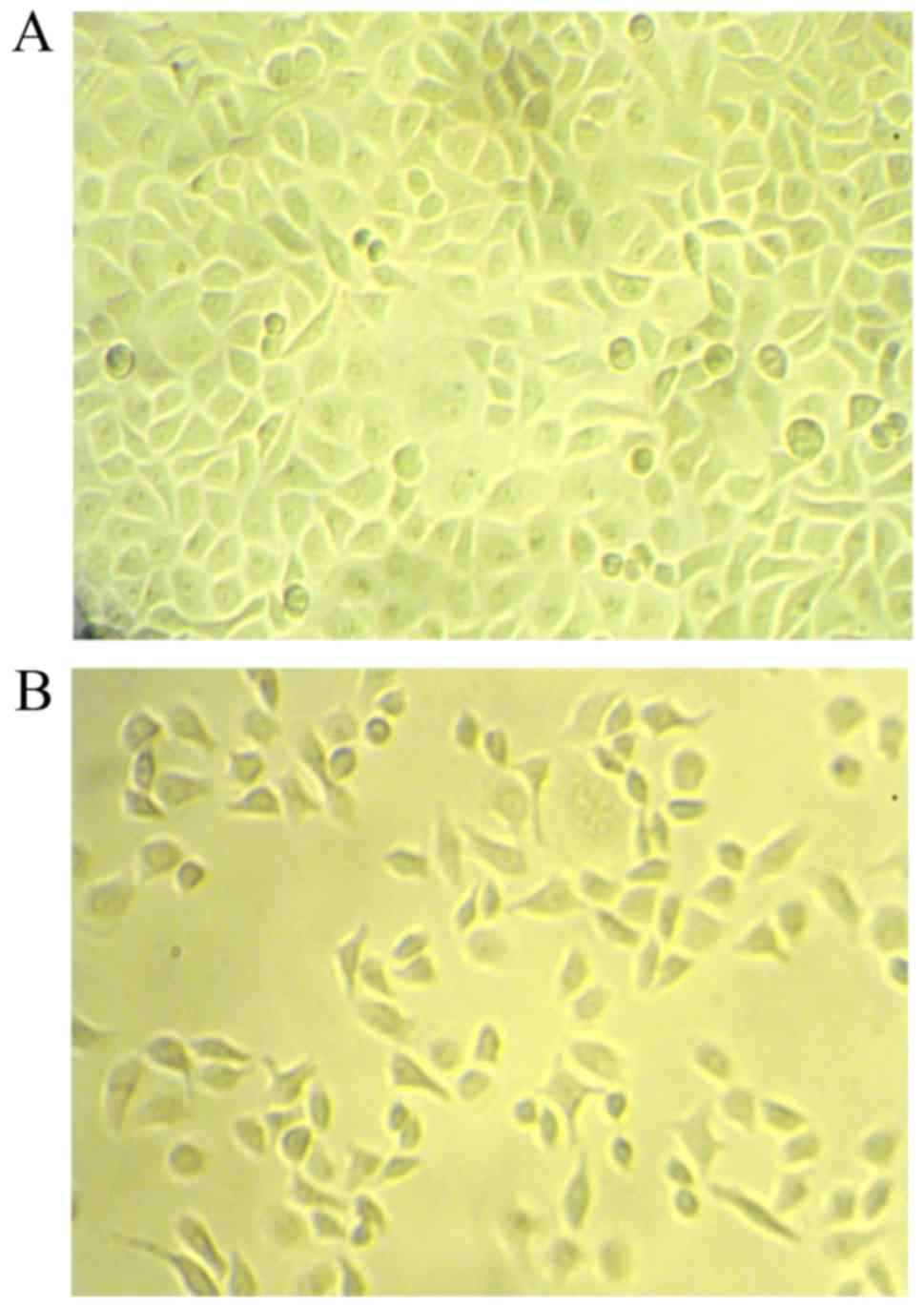
As shown in Fig. 5,
under the inverted microscope, cells of the control group were in
good condition and grew with attaching on the plate; interestingly,
cells of the trancfected group exhibited smaller volume, narrow
shape and loose junctions, more cells floated on the plate.
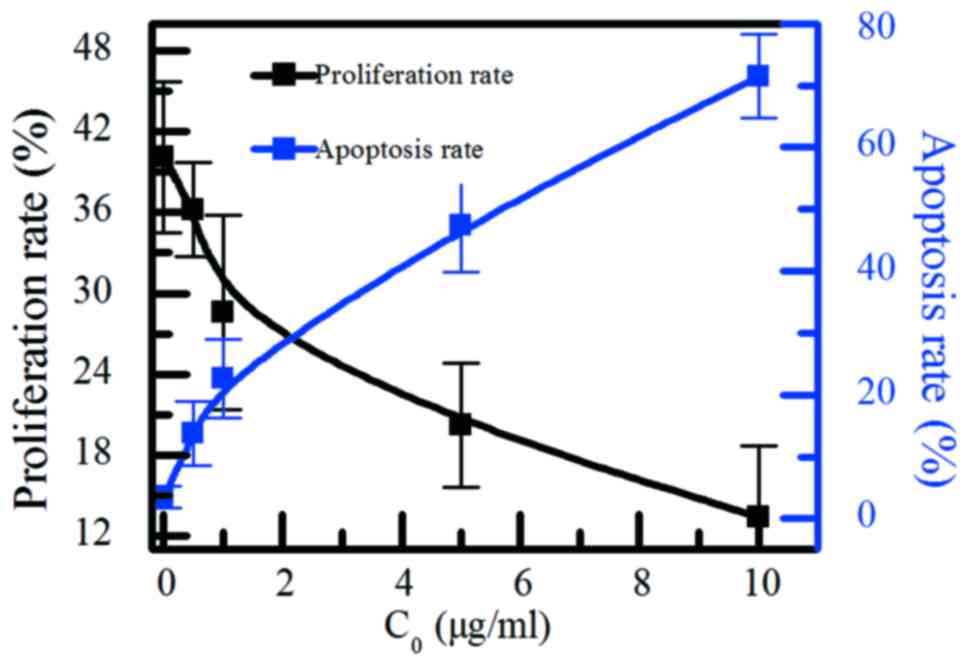
The proliferation rate and the apoptosis rate was
shown in Fig. 6. When the
concentration of the BCSG1-siRNA chitosan nanoparticles up to 10
µg/ml, the proliferation rate reduced from 40.1±5.6 to 13.4±5.3%,
and the apoptosis rate increased from 3.5±1.8 to 71.5±6.8%. When
the concentration of BCSG1-siRNA chitosan nanoparticles increases,
the proliferation effect of MCF-7 cell will decrease, the apoptosis
rate will increase.
The results showed that the designed
BCSG1-siRNA/chitosan-silicon dioxide nanoparticle could
significantly inhibit tumor growth. The BCSG1-siRNA was
encapsulated by chitosan-silicon dioxide, then delivered to the
tumor, avoiding the degradation in the tissue. As improving the
concentration of BCSG1-siRNA, the BCSG1 expression would be well
downregulated, then the tumor growth would be inhibited.
In this study, the BCSG1-siRNA plasmid has been
designed to downregulate the BCSG1 gene expression.
Chitosan-silicon dioxide nanoparticle containing BCSG1-siRNA has
been successfully synthesized, its feasibility and effect as drug
carriers has been analyzed.
From the immunocytochemistry experiment and the PCR
results, the designed BCSG1-siRNA plasmid could significantly
downregulate the BCSG1 gene expression. The encapsulation
efficiency of nanoparticle was almost 90% (compared with the
traditional materials, the encapsulation efficiency was 65–75% in
the purified chitosan nanometer carrier, 65–70% in liposomes or
other loading material).
Chitosan-silicon dioxide nanoparticle has a good
application in drug controlled release, because of the positive
charged chitosan and mesoporous silica nanoparticle. The release
efficiency was enhanced, as increasing the concentration of gene
drug. And the releasing effect of BCSG1-siRNA nanoparticles was
sustained.
As improving the concentration of BCSG1-siRNA, the
BCSG1 expression would be well downregulated, then the tumor growth
would be inhibited. Results reveal the significant selectivity of
BCSG1-siRNA nanoparticles.
Chitosan-silicon dioxide carrier can not only
improve the encapsulation efficiency, but also enhance the release
rate of gene drugs. As increasing the concentration of BCSG1-siRNA
chitosan nanoparticle, its effect for breast cancer cells was
enhanced, while the damage on normal cell is relatively small.
Therefore, it is a better targeted gene delivery system, the
chitosan-silicon dioxide as a targeted carrier for gene therapy is
feasible.
Acknowledgements
This study was funded by Health and Family Planning
Commission of Henan Province, China (201302015), and the 5451
Project of Health Planning Commission in Henan Province, 2014
(94-United States). The breast cancer cells in the experiment were
provided from the Fifth Affiliated Hospital of Zhengzhou
University.
References
|
1
|
Anderson BO, Shyyan R, Eniu A, Smith RA,
Yip CH, Bese NS, Chow LW, Masood S, Ramsey SD and Carlson RW:
Breast cancer in limited-resource countries: An overview of the
breast health global initiative 2005 guidelines. Breast J. 12 Suppl
1:S3–S15. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Peng C, Ma W, Xia W and Zheng W:
Integrated analysis of differentially expressed genes and pathways
in triple-negative breast cancer. Mol Med Rep. 15:1087–1094. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Gupta A, Godwin AK, Vanderveer L, Lu A and
Liu J: Hypomethylation of the synuclein gamma gene CpG island
promotes its aberrant expression in breast carcinoma and ovarian
carcinoma. Cancer Res. 63:664–673. 2003.PubMed/NCBI
|
|
4
|
Inaba S, Li C, Shi YE, Song DQ, Jiang JD
and Liu J: Synuclein gamma inhibits the mitotic checkpoint function
and promotes chromosomal instability of breast cancer cells. Breast
Cancer Res Treat. 94:25–35. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Gupta A, Inaba S, Wong OK, Fang G and Liu
J: Breast cancer-specific gene 1 interacts with the mitotic
checkpoint kinase BubR1. Oncogene. 22:7593–7599. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Pan ZZ, Bruening W, Giasson BI, Lee VM and
Godwin AK: Gamma-synuclein promotes cancer cell survival and
inhibits stress and chemotherapy drug-induced apoptosis by
modulating MAPK pathways. J Biol Chem. 277:35050–35060. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Howard EM, Lau SK, Lyles RH, Birdsong GG,
Tadros TS, Umbreit JN and Kochhar R: Correlation and expression of
p53, HER-2, vascular endothelial growth factor (VEGF), and
e-cadherin in a high-risk breast-cancer population. Int J Clin
Oncol. 9:154–160. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Singh VK, Zhou Y, Marsh JA, Uversky VN,
Forman-kay JD, Liu J and Jia Z: Synuclein-gamma targeting peptide
inhibitor that enhances sensitivity of breast cancer cells to
antimicrotubule drugs. Cancer Res. 67:626–633. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Arriagada R, Le Chevalier T, Pignon JP,
Rivière A, Monnet I, Chomy P, Tuchais C, Tarayre M and Ruffié P:
Initial chemotherapeutic doses and survival in patients with
limited small-cell lung cancer. N Engl J Med. 329:1848–1852. 1993.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kester M, Heakal Y, Fox T, Sharma A,
Robertson GP, Morgan TT, Altinoğlu EI, Tabaković A, Parette MR,
Rouse SM, et al: Calcium phosphate nanocomposite particles for in
vitro imaging and encapsulated chemotherapeutic drug delivery to
cancer cells. Nano Lett. 8:4116–4121. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cichewicz RH and Kouzi SA: Chemistry,
biological activity, and chemotherapeutic potential of betulinic
acid for the prevention and treatment of cancer and HIV infection.
Med Res Rev. 24:90–114. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Shree T, Olson OC, Elie BT, Kester JC,
Garfall AL, Simpson K, Bell-McGuinn KM, Zabor EC, Brogi E and Joyce
JA: Macrophages and cathepsin proteases blunt chemotherapeutic
response in breast cancer. Genes Dev. 25:2465–2479. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Davis ME, Zuckerman JE, Choi CH, Seligson
D, Tolcher A, Alabi CA, Yen Y, Heidel JD and Ribas A: Evidence of
RNAi in humans from systemically administered siRNA via targeted
nanoparticles. Nature. 464:1067–1070. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Han HD, Mangala LS, Lee JW, Shahzad MM,
Kim HS, Shen D, Nam EJ, Mora EM, Stone RL, Lu C, et al: Targeted
gene silencing using RGD-labeled chitosan nanoparticles. Clin
Cancer Res. 16:3910–3922. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Varshosaz J and Taymouri S: Hollow
inorganic nanoparticles as efficient carriers for siRNA delivery: A
comprehensive review. Curr Pharm Des. 21:4310–4328. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Chen SJ, Zhang HZ, Wan LC, Jiang SS, Xu
YM, Liu F, Zhang T, Ma D and Xie MQ: Preparation and performance of
a pH-sensitive cisplatin-loaded magnetic nanomedicine that targets
tumor cells via folate receptor mediation. Mol Med Rep.
13:5059–5067. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Kim DH, Villeneuve LM, Morris KV and Rossi
JJ: Argonaute-1 directs siRNA-mediated transcriptional gene
silencing in human cells. Nat Struct Mol Biol. 13:793–797. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Judge AD, Sood V, Shaw JR, Fang D,
McClintock K and MacLachlan I: Sequence-dependent stimulation of
the mammalian innate immune response by synthetic siRNA. Nat
Biotechnol. 23:457–462. 2005. View
Article : Google Scholar : PubMed/NCBI
|
|
19
|
Pillé JY, Li H, Blot E, Bertrand JR,
Pritchard LL, Opolon P, Maksimenko A, Lu H, Vannier JP, Soria J, et
al: Intravenous delivery of anti-RhoA small interfering RNA loaded
in nanoparticles of chitosan in mice: Safety and efficacy in
xenografted aggressive breast cancer. Hum Gene Ther. 17:1019–1026.
2006. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Xu Y and Du Y: Effect of molecular
structure of chitosan on protein delivery properties of chitosan
nanoparticles. Int J Pharm. 250:215–226. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Yin L, Ding J, He C, Cui L, Tang C and Yin
C: Drug permeability and mucoadhesion properties of thiolated
trimethyl chitosan nanoparticles in oral insulin delivery.
Biomaterials. 30:5691–5700. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Shi Z, Neoh KG, Kang ET and Wang W:
Antibacterial and mechanical properties of bone cement impregnated
with chitosan nanoparticles. Biomaterials. 27:2440–2449. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kim JH, Kim YS, Park K, Lee S, Nam HY, Min
KH, Jo HG, Park JH, Choi K, Jeong SY, et al: Antitumor efficacy of
cisplatin-loaded glycol chitosan nanoparticles in tumor-bearing
mice. J Control Release. 127:41–49. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Janes KA, Fresneau MP, Marazuela A, Fabra
A and Alonso MJ: Chitosan nanoparticles as delivery systems for
doxorubicin. J Control Release. 73:255–267. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hwang HY, Kim IS, Kwon IC and Kim YH:
Tumor targetability and antitumor effect of docetaxel-loaded
hydrophobically modified glycol chitosan nanoparticles. J Control
Release. 128:23–31. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wu Y, Yang W, Wang C, Hu J and Fu S:
Chitosan nanoparticles as a novel delivery system for ammonium
glycyrrhizinate. Int J Pharm. 295:235–245. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Kim JH, Kim YS, Park K, Kang E, Lee S, Nam
HY, Kim K, Park JH, Chi DY, Park RW, et al: Self-assembled glycol
chitosan nanoparticles for the sustained and prolonged delivery of
antiangiogenic small peptide drugs in cancer therapy. Biomaterials.
29:1920–1930. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
He L, Huang Y, Zhu H, Pang G, Zheng W,
Wong YS and Chen T: Cancer-targeted monodisperse mesoporous silica
nanoparticles as carrier of ruthenium polypyridyl complexes to
enhance theranostic effects. Adv Func Mat. 24:2754–2763. 2014.
View Article : Google Scholar
|
|
29
|
Zhang Q, Liu F, Nguyen KT, Ma X, Wang X,
Xing B and Zhao Y: Multifunctional mesoporous silica nanoparticles
for cancer-targeted and controlled drug delivery. Adv Func Mat.
22:5144–5156. 2012. View Article : Google Scholar
|
|
30
|
Cai LL, Liu P, Li X, Huang X, Ye YQ, Chen
FY, Yuan H, Hu FQ and Du YZ: RGD peptide-mediated chitosan-based
polymeric micelles targeting delivery for integrin-overexpressing
tumor cells. Int J Nanomedicine. 6:3499–3508. 2011.PubMed/NCBI
|
|
31
|
Mansur AA, de Carvalho SM and Mansur HS:
Bioengineered quantum dot/chitosan-tripeptide nanoconjugates for
targeting the receptors of cancer cells. Int J Biol Macromol.
82:780–789. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Liu TY, Chen SY, Lin YL and Liu DM:
Synthesis and characterization of amphiphatic
carboxymethyl-hexanoyl chitosan hydrogel: Water-retention ability
and drug encapsulation. Langmuir. 22:9740–9745. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Liu KH, Chen SY, Liu DM and Liu TY:
Self-assembled hollow nanocapsule from amphiphatic
carboxymethyl-hexanoyl chitosan as drug carrier. Macromolecules.
41:6511–6516. 2008. View Article : Google Scholar
|
|
34
|
Burdick JA and Anseth KS:
Photoencapsulation of osteoblasts in injectable RGD-modified PEG
hydrogels for bone tissue engineering. Biomaterials. 23:4315–4323.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Li WM, Liu DM and Chen SY:
Amphiphilically-modified gelatin nanoparticles: Self-assembly
behavior, controlled biodegradability, and rapid cellular uptake
for intracellular drug delivery. J Mat Chem. 21:12381–12388. 2011.
View Article : Google Scholar
|
|
36
|
Reynolds AR, Hart IR, Watson AR, Welti JC,
Silva RG, Robinson SD, Da Violante G, Gourlaouen M, Salih M, Jones
MC, et al: Stimulation of tumor growth and angiogenesis by low
concentrations of RGD-mimetic integrin inhibitors. Nat Med.
15:392–400. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
37
|
Krajewska M, Krajewski S, Epstein JI,
Shabaik A, Sauvageot J, Song K, Kitada S and Reed JC:
Immunohistochemical analysis of bcl-2, bax, bcl-X and mcl-1
expression in prostate cancer. Am J Pathol. 148:1567–1576.
1996.PubMed/NCBI
|
|
38
|
Dastjerdi MN, Rarani MZ, Valiani A and
Mahmoudieh M: The effect of adenosine A1 receptor agonist and
antagonist on p53 and caspase 3, 8, and 9 expression and apoptosis
rate in MCF-7 breast cancer cell line. Res Pharm Sci. 11:303–310.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Wu BB, Gong YP, Wu XH, Chen YY, Chen FF,
Jin LT, Cheng BR, Hu F and Xiong B: Fourier transform infrared
spectroscopy for the distinction of MCF-7 cells treated with
different concentrations of 5-fluorouracil. J Transl Med.
13:1082015. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
França EL, Honorio-frança AC, Fernandes
RT, Marins CM, Pereira CC and Fde Varotti P: The effect of
melatonin adsorbed to polyethylene glycol microspheres on the
survival of MCF-7 cells. Neuroimmunomodulation. 23:27–32. 2016.
View Article : Google Scholar : PubMed/NCBI
|