Introduction
Lipoid proteinosis (LP) was first described in 1929
by Wiethe and Urbach as a rare autosomal recessive inheritance
disease (1). The major clinical
features of LP include intracranial calcification, infiltration and
thickening of skin mucosal tissues, hair loss, recurrent parotitis,
papules in the eyelids, dental underdevelopment, hoarseness, and
even suffocation and mortality (2,3).
Certain patients with LP also exhibit mental and psychiatric
symptoms (4). These pathological
features are now standard for clinical LP diagnosis (5).
Although the molecular mechanism of occurrence and
development of LP require further elucidation, it is generally
considered that the gene mutation of extracellular matrix protein 1
(ECM1) forms the pathological basis of LP (6,7).
Hamada et al (6) performed
the first linkage analysis and survey of the ECM1 gene in patients
with LP, and identified the localization of the ECM1 gene at human
chromosome 1q21 locus Further studies indicated the role of an ECM1
gene mutation in LP. Systematic surveys across different countries
and regions demonstrated variable mutation points of the ECM1 gene
in individual patients with LP, suggesting the potential gene
polymorphism of the ECM1 gene in patients with LP (7,8).
The ECM is a group of glycoproteins that are
secreted from animal cells to form the complex matrix between cells
and exert critical functions during the modulation of
intra-cellular transport, signal transduction and energy exchange
(9–11). ECM1 has been demonstrated to
possess important roles in the differentiation of cellular
epidermal cells, and the connection between glycoproteins of the
epidermal layer and collagen (12,13).
Among the 10 exons of ECM1 gene, frequent mutations occur between
exon 6 and exon 7 (7).
Due to its rare incidence, clinical reporting of LP
is sparse in China. The diagnosis of LP in China relies mainly on
clinical features. With the rapid progression of molecular biology
at the clinical level, reverse transcription-quantitative
polymerase chain reaction (RT-qPCR) analysis together with DNA
sequencing have now become important methods to study the existence
of ECM1 gene mutation, thus providing evidence to improve LP
diagnosis (6,7). Such a combination of RT-qPCR and gene
sequencing can also provide important information for genetic
counseling, prenatal diagnosis and prepotency (14).
The present study collected clinical cases for
diagnosing LP based on its pathological features. RT-qPCR was
performed to detect the presence of mutation in the ECM1 gene of
patients. Common gene mutation loci and allele frequencies were
further analyzed in patients with LP. The present study aimed to
investigate the clinical features and inheritance pattern of ECM1
gene mutation in Chinese LP patients, thus providing useful
information for the diagnosis and treatment of LP in China.
Materials and methods
Reagents and drugs
PCR kits (SanTaq PCR Master Mix and Taq Plus DNA
Polymerase) were purchased from Shanghai Shenggong Biology
Engineering Technology Service, Ltd. (Shanghai, China). Primers
were synthesized by Beijing Sanbo Yuanzhi Biotechnology Co., Ltd.
(Beijing, China). Universal DNA Purification Kit for PCR products
was purchased from Tiangen Bio (Beijing, China). Sequencing of PCR
products was performed by Beijing Sanbo Yuanzhi Biotechnology Co.,
Ltd.
Patients
From November 2016 to September 2017, 4 patients (2
females and 2 males with a median age of 49, ranging from 39–65)
with LP in the department of dermatology of Yantai Yuhuangding
Hospital (Yantai, China) were recruited for the present study.
Inclusive criteria were (15):
Intracranial calcification, infiltration and thickening of skin
mucosal tissues, hair loss, recurrent parotitis, papules in the
eyelids, dental underdevelopment and hoarseness. The present study
was approved by the ethical committee of Yantai Yuhuangding
Hospital and obtained written consents from patients or families.
Family surveys were also performed if possible.
Histopathological examination
Pathological examination was performed on patients
with LP, as previously described (16,17).
The keratinization degree of epidermis was examined to see if
atrophy or atypical hyperplasia of the spinous layer existed.
Dermis was then examined for any notable thickening, vessel
dilation, as well as thickening of vascular walls and the existence
of extracellular translucent layer of eosinophilic cells at the
superficial layer of the dermis. Periodic acid-Schiff (PAS)
staining performed for this translucent layer of eosinophilic cells
and observed under an Olympus BH2 light microscope.
Genomic DNA extraction
With the consent from patients and families, 5-ml
peripheral blood samples were collected from veins for extracting
genomic DNA as previously described (18). Equal volumes of phenol were added
to the blood which was then centrifuged (3,000 × g for 8 min) at
room temperature. The upper aqueous phase was collected to a new
tube with the addition of equal volume phenol for another
centrifugation at 3,000 × g for 8 min at 4°C. The upper aqueous
phase was again saved in a new tube with an equal volume of
phenol/chloroform (1:1) mixture. Following centrifugation at 3,000
× g for 8 min at 4°C, the upper phase was saved and mixed with
equal volume of chloroform. The upper phase was then collected
following 3,000 × g centrifugation for 8 min 4°C and mixed with
1/10 volume of sodium acetate (3 M) and absolute ethanol. The white
participation was then carefully removed, followed by washing with
75% ethanol. The DNA pellet was dissolved in sterilized water and
the genomic DNA was then used in the following experiment.
PCR
PCR primers were designed based on the ECM1 gene
sequence. Primer sequence for ECM1 were: Forward,
5′-GGCTTTTGCTTACTCCTTCTACCC-3′; Reverse,
5′-AGTAGCTGGCAGGTTGCGTGG-3′. The primer sequence for b-actin was
Forward, 5′-CGTTGCTATCCAGGCTGTGCTAT-3′; Reverse,
5′-CAGCTTCTCCTTAATGTCACGC-3′.
PCR was performed in a 25 µl system containing 1 µl
genomic DNA template, 2.5 µl PCR buffer (10X), 2 µl dNTP mixture (1
mM), 1 µl of each primer (20 µM), 1 µl Taq DNA polymerase, 20 mM
MgCl2 and 16 µl H2O. PCR was conducted under
the following conditions: 95°C denaturing for 5 min, 30 cycles of
95°C for 45 sec, 55°C for 45 sec and finally 72°C for 6 min.
Products were kept at −20°C.
Gel electrophoresis
Agarose gel electrophoresis was performed using
standard procedure; 5 µl PCR products were mixed with 1 µl loading
buffer (6X) and loaded into 0.8% agarose gel. The electrophoresis
was performed in a 120 V electrical field for 10 min as previously
described (19).
PCR products sequencing
Following agarose gel separation, PCR products were
sequenced using directly using Big Dye® Terminator v3.1
cycle sequencing kit (Thermo Fisher Scientific, Waltham, MA, USA).
The sequencing was performed by Beijing Sanbo Yuanzhi Biotechnology
Co., Ltd. The sequencing analyzer was from Applied Biosystems
(3730XLDNA; Thermo Fisher Scientific, Inc.).
Sequence alignment
Sequencing results were aligned with known gene
fragments using a Pubmed database as previously described (20).
Results
Pathological examination of patients
with LP
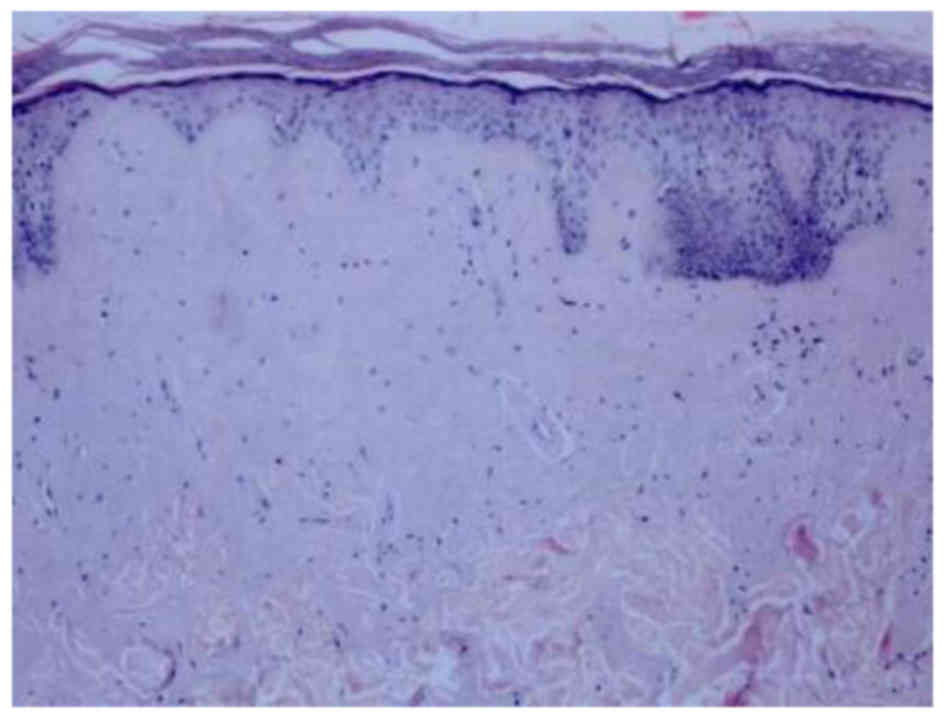
In all 4 patients with LP who took part in the
present study, similar histopathological features were observed;
over-cutinization and a thickening of the spinuous layer with
atypical arrangement. Homologous red precipitation was observed in
sweat gland, vessels, hair follicles and the middle layer. PAS
staining demonstrated a positive reaction, with an onion-shaped
arrangement of PAS-positive substances in epidermal layers of the
skin (Fig. 1).
Family survey of LP patients
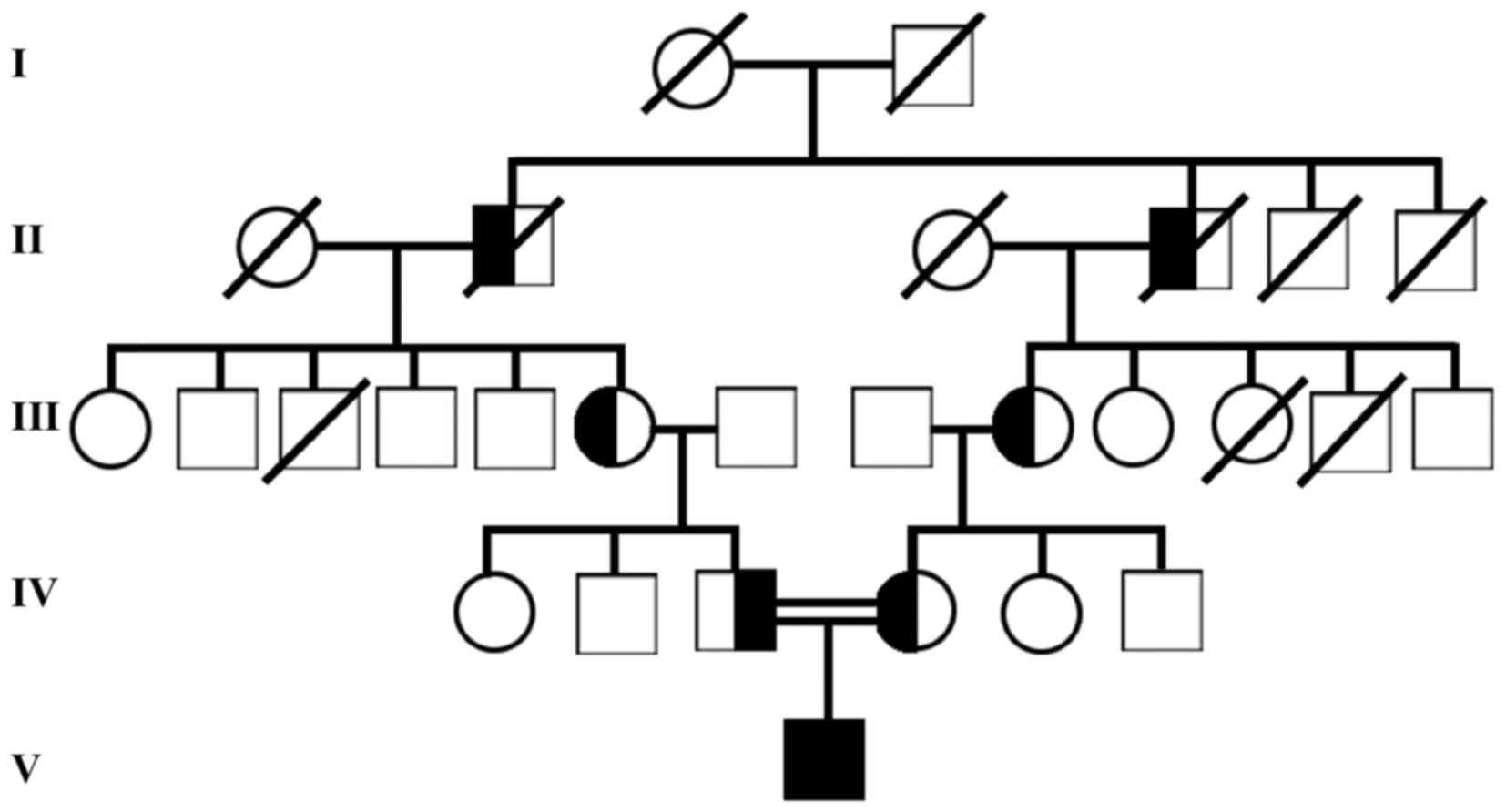
Only 1 out of the 4 patients with LP agreed to have
a family survey performed. As demonstrated in Fig. 2, the incidence of LP fitted the
autosomal recessive inheritance pattern.
PCR results
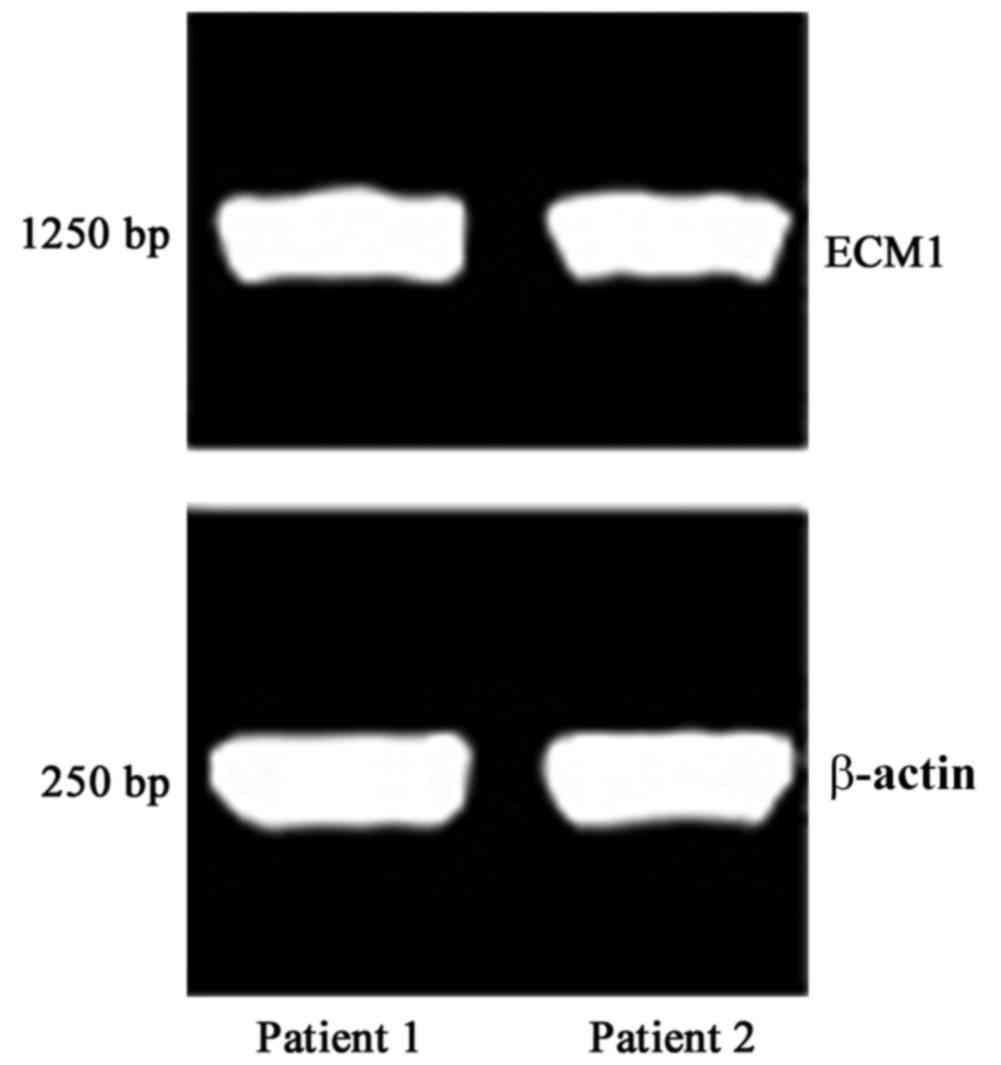
PCR and agarose gel electrophoresis were performed
on the genomic DNA of patients with LP. As demonstrated in Fig. 3, the PCR products had a length of
~1,250 bp, consistent with the prediction, suggesting successful
amplification.
PCR product sequencing and
alignment
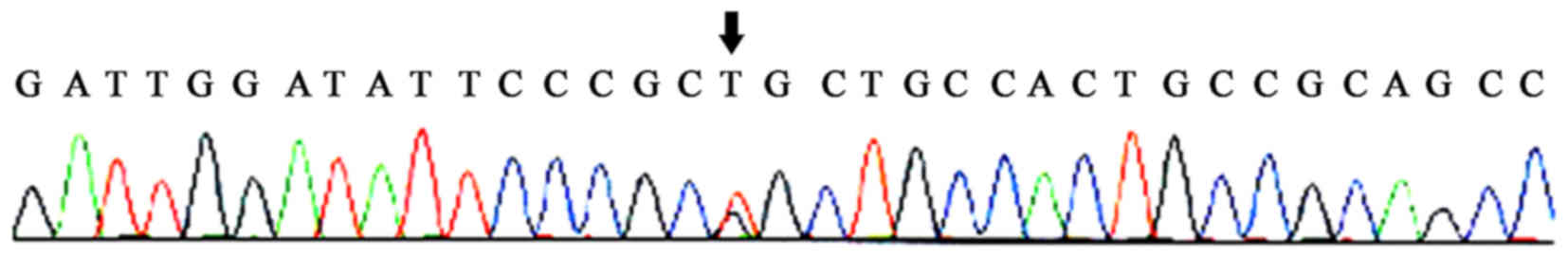
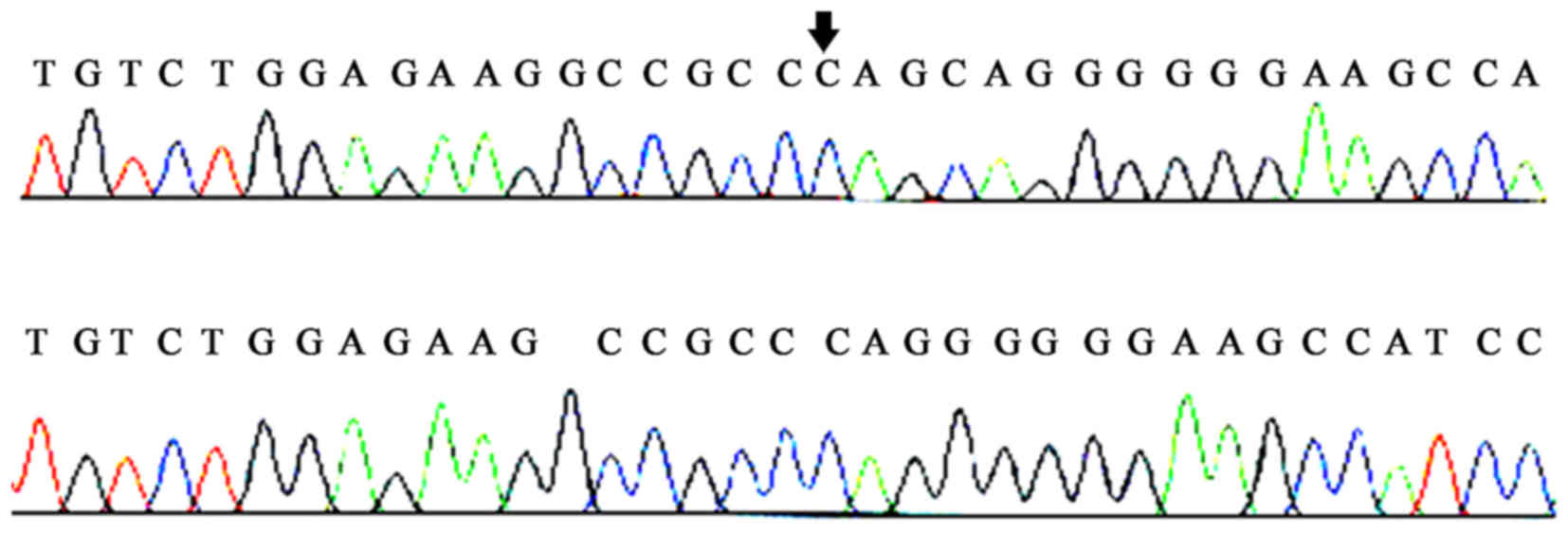
Following the DNA sequencing of PCR products, a
novel mutation locus was identified at the 6th exon of the ECM1
gene. Specifically, a CAG-insertion at position 508 bp of the ECM1
gene: 5′-AAGGCCGCCCAG-3′ 3′-TTCCGGCGGGTC-5′ 5′-CTGGGCGGCCTT-3′.
From these sequencing results a CTG-insertion mutation at position
508 bp of the ECM1 gene may be identified. This frameshift mutation
was designated c.508insCTG for short. Such a mutation caused the
transition of the 169th amino acid of the ECM1 protein to proline
instead of leucine.
Gene sequencing and alignment results in all 4
patients with LP in the present study are presented in Figs. 4–7. These results suggested point mutations
of the ECM1 gene, including one homozygous point mutation (C220G)
as previously reported, one novel homozygous mutation (c.508insCTG)
and two heterozygous mutations (C220G/P.R481X and
c.507delT/c.l473delT).
Discussion
LP is a rare skin disease with autosomal recessive
inheritance. Various studies have indicated that an ECM1 gene
mutation underlies the pathology of LP (1,2). In
the Chinese population, however, the association between ECM1
mutation and occurrence of LP requires further investigation. The
present study discussed the mutation types of the ECM1 gene in
Chinese patients with LP.
The present study obtained three main results: i)
Histopathological examination revealed the consistent features of
LP in the patients; ii) PCR amplification of the ECM1 gene in
patients with LP obtained relevant bands; and iii) sequencing of
PCR products identified point mutations of ECM1 gene, including one
homozygous point mutation (C220G) as previously reported, one novel
homozygous mutation (c.508insCTG) and two heterozygous mutations
(C220G/P.R481X and c.507delT/c.l473delT).
A previous study demonstrated the existence of
LP-associated ECM1 gene mutations mainly in exons 6 and 7 (21). The present study agreed that exons
6 and 7 possessed gene mutations. The difference was that previous
reports identified mainly homozygous mutations (8–10),
while heterozygous mutations occurred in the present study. The
mutation of the ECM1 gene can lead to LP, even by heterozygous
mutation.
A previous study suggested that consanguineous
marriage is a major reason for LP (22,23).
The majority of studies also agree that homozygous mutation of the
ECM1 gene is typically observed. The present study identified the
existence of heterozygous mutations in patients with LP. These data
suggested higher susceptibility even without consanguineous
marriage, stressing the importance of premarital examination and
prepotency.
Certain weaknesses and limitations also existed in
the present study: i) Relatively few cases were included (n=4) and
may bias the final results; ii) only the association between the
ECM1 gene and LP was analyzed, without considering other possible
factors; and iii) the present study did not further investigate the
molecular mechanism of the ECM1 mutation for LP, or the normal
function of ECM1 protein for inducing LP.
In summary, the present study demonstrated the
association between ECM1 gene mutation and patients with LP.
Patients with LP exhibited one homozygous point mutation (C220G) as
previously reported, one novel homozygous mutation (c.508insCTG)
and two heterozygous mutations (C220G/P.R481X and
c.507delT/c.l473delT).
Acknowledgements
Not applicable.
Funding
This work was supported by Yantai University (grant
no. 2012084).
Availability of data and materials
All data generated or analysed during this study are
included in this published article.
Authors' contributions
DG designed the study and wrote the manuscript. DG,
XM, PL, SZ and JC performed the experiments and analysed the
data.
Ethics approval and consent to
participate
All experimental procedures involving animals were
approved by the Ethnic Committee of Yantai Yuhuangding Hospital
(Yantai, China). Not applicable for consent to participate for
human individuals.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Urbach E and Weithe C: Lipoidosis cutis et
mucosae. Virchows Arch Path Anat. 273:285–319. 1929. View Article : Google Scholar
|
|
2
|
Çalıskan E, Açıkgöz G, Tunca M, Koç E,
Arca E and Akar A: Treatment of lipoid proteinosis with ablative
Er:YAG laser resurfacing. Dermatol Ther. 28:291–295. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Antoon JW, Hernandez ML, Roehrs PA, Noah
TL, Leigh MW and Byerley JS: Endogenous lipoid pneumonia preceding
diagnosis of pulmonary alveolar proteinosis. Clin Respir J.
10:246–249. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bakry OA, Samaka RM, Houla NS and Basha
MA: Two Egyptian cases of lipoid proteinosis successfully treated
with acitretin. J Dermatol Case Rep. 8:29–34. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Tewari A, Fassihi H, McGibbon D, Robson A
and Sarkany R: A case of extensive hyaline deposition in facial
skin caused by erythropoietic protoporphyria. Br J Dermatol.
171:412–414. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Hamada T, McLean WH, Ramsay M, Ashton GH,
Nanda A, Jenkins T, Edelstein I, South AP, Bleck O, Wessagowit V,
et al: Lipoid proteinosis maps to 1q21 and is caused by mutations
in the extracellular matrix protein 1 gene (ECM1). Hum Mol Genet.
11:833–840. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hamada T, Wessagowit V, South AP, Ashton
GH, Chan I, Oyama N, Siriwattana A, Jewhasuchin P,
Charuwichitratana S, Thappa DM, et al: Extracellular matrix protein
1 gene (ECM1) mutations in lipoid proteinosis and
genotype-phenotype correlation. J Invest Dermatol. 120:345–350.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Izadi F, Mahjoubi F, Farhadi M, Kalayinia
S, Bidmeshkipour A, Tavakoli MM and Samanian S: Extracellular
matrix protein 1 gene (ECM1) mutations in nine Iranian families
with lipoid proteinosis. Indian J Med Res. 143:303–307. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Freedman BR, Bade ND, Riggin CN, Zhang S,
Haines PG, Ong KL and Janmey PA: The (dys)functional extracellular
matrix. Biochim Biophys Acta. 1853:3153–3164. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kim SH, Turnbull J and Guimond S:
Extracellular matrix and cell signalling: The dynamic cooperation
of integrin, proteoglycan and growth factor receptor. J Endocrinol.
209:139–151. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Silver FH, DeVore D and Siperko LM:
Invited review: Role of mechanophysiology in aging of ECM: Effects
of changes in mechanochemical transduction. J Appl Physiol (1985).
95:2134–2141. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Watt FM and Fujiwara H: Cell-extracellular
matrix interactions in normal and diseased skin. Cold Spring Harb
Perspect Biol. 3:a0051242011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Blanpain C and Fuchs E: Epidermal stem
cells of the skin. Annu Rev Cell Dev Biol. 22:339–373. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Mahdieh N and Rabbani B: An overview of
mutation detection methods in genetic disorders. Iran J Pediatr.
23:375–388. 2013.PubMed/NCBI
|
|
15
|
Kaindl T, Rieger H, Kaschel LM, Engel U,
Schmaus A, Sleeman J and Tanaka M: Spatio-temporal patterns of
pancreatic cancer cells expressing CD44 isoforms on supported
membranes displaying hyaluronic acid oligomers arrays. PloS One.
7:e429912012. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bai X, Liu JW and Ma DL: Novel mutations
in extracellular matrix protein 1 gene in a chinese patient with
lipoid proteinosis. Chin Med J (Engl). 129:2765–2766. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Gutte R, Sanghvi S, Tamhankar P and
Khopkar U: Lipoid proteinosis: Histopathological characterization
of early papulovesicular lesions. Indian Dermatol Online J.
3:148–149. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Lajunen TK, Purhonen AK, Haapea M,
Ruokonen A, Puukka K, Hartikainen AL, Savolainen MJ, Morin-Papunen
L, Tapanainen JS, Franks S, et al: Full-length visfatin levels are
associated with inflammation in women with polycystic ovary
syndrome. Eur J Clin Invest. 42:321–328. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Samdani AJ, Azhar A, Shahid SM, Nawab SN,
Shaikh R, Qader SA, Mansoor Q, Khoso BK and Ismail M: Homozygous
frame shift mutation in ECM1 gene in two siblings with lipoid
proteinosis. J Dermatol Case Rep. 4:66–70. 2010.PubMed/NCBI
|
|
20
|
Schuler GD: Sequence alignment and
database searching. Methods Biochem Anal. 39:145–171.
1998.PubMed/NCBI
|
|
21
|
Chelvan HT, Narasimhan M, Subramanian
Shankaran A and Subramaniam S: Lipoid proteinosis presenting with
an unusual nonsense Q32X mutation in exon 2 of the extracellular
matrix protein 1 gene. Australas J Dermatol. 53:e79–e82. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Xu W, Wang L and Zhang L, Han DM and Zhang
L: Otolaryngological manifestations and genetic characteristics of
lipoid proteinosis. Ann Oto Rhinol Laryngol. 119:767–771. 2010.
|
|
23
|
Rizzo R, Ruggieri M, Micali G, Tinè A,
Sanfilippo S and Pavone L: Lipoid proteinosis: A case report.
Pediatr Dermatol. 14:22–25. 1997. View Article : Google Scholar : PubMed/NCBI
|