Introduction
Human melanoma is one of the most aggressive and
frequently diagnosed cancers in humans (1). For numerous years, the incidence of
melanoma has continually increased, despite advancements in the
understanding of the initiation and progress of melanoma (2). Therefore, the identification of
important molecules in the progression of melanoma is urgently
required, in order to develop new preventive and diagnostic
strategies targeting these markers (3,4).
An increasing number of studies have documented that
microRNAs (miRNAs) play essential roles in multiple cancers, which
lead to messenger (m)RNA degradation by targeting the
3′-untranslated region of target mRNAs (5,6). Among
several miRNAs regulating cell proliferation, miRNA-186 (miR-186)
has been shown to be one of the important determinants of cell
proliferation in various types of cancers (7–10). The
results of the present study revealed that upregulation of miR-186
in melanoma is associated with development of melanoma. Additional
findings that miR-186 directly targets cylindromatosis (CYLD),
thereby promoting cell proliferation of human melanoma cells. In
summary, the present findings suggested that miR-186 is mediated
with progression of melanoma and may serve as a new target for
treatment of melanoma.
Materials and methods
Clinical specimens
Skin tissues were obtained from 8 patients and
histopathologically diagnosed following surgery at Guangzhou First
People's Hospital, Guangzhou Medical University (Guangzhou, China).
The present study was approved by the Ethics Committee of Guangzhou
First People's Hospital, Guangzhou Medical University (Guangzhou,
China). All samples were collected and analyzed with the written
informed consent of the patients.
Cell culture
The human melanoma A375-S2, SKMEL-28, SKMEL-5, MeWO
and RPMI-7951 cell lines were purchased from the National Rodent
Laboratory Animal Resource (Shanghai, China). All melanoma cell
lines were grown in Gibco Dulbecco's modified Eagle's medium
(Thermo Fisher Scientific, Inc., Waltham, MA, USA) supplemented
with 10% fetal bovine serum (FBS; Sigma-Aldrich, St. Louis, MO,
USA) and 100 units/ml Invitrogen penicillin-streptomycin (Thermo
Fisher Scientific, Inc.). Normal human epidermal melanocytes
(NHEMs) from adult skin (PromoCell GmbH, Heidelberg, Germany) were
maintained in serum- and phorbol myristate acetate-free melanocyte
growth medium M2 (PromoCell GmbH). The cell lines were cultured in
a humidified incubator at 37°C in a 5% CO2 and 95% air
atmosphere for 2–3 days.
Plasmids and transfection
To induce the ectopic expression of CYLD, CYLD open
reading frames containing a 3′-UTR was amplified by polymerase
chain reaction (PCR) and then cloned into pGL3 vectors (Promega
Corporation, Madison, WI, USA) downstream of the the Renilla
luciferase complementary DNA. miR-186 mimic, miR-186 inhibitor,
miR-186-mut and negative control (NC) were purchased from
GeneCopoeia, Inc. (Guangzhou, China) and transfected into melanoma
cells using Invitrogen Lipofectamine 2000 reagent (Thermo Fisher
Scientific, Inc.), according to the manufacturer's
instructions.
RNA extraction and reverse
transcription-quantitative PCR (RT-qPCR)
Total RNA was isolated using Invitrogen TRIzol
Reagent (Thermo Fisher Scientific, Inc.) and RT was performed using
the miScript Reverse Transcription kit (Qiagen, Hilden, Germany).
For the quantification of miRNA expression, qPCR was performed
using the miScript Reverse Transcription and miScript SYBR Green
PCR kits, according to the manufacturer's instructions (Qiagen).
The relative miR-186 expression levels following normalization to
the expression of U6 small nuclear RNA were calculated using 2-ΔΔCq
(11). Quantitative PCR was performed
using the QuantiNova SYBR Green PCR Kit (Qiagen China Co., Ltd.,
Shanghai, China) and a 7500 Sequence Detection system (Applied
Biosystems Life Technologies, Foster City, CA, USA). The primers
selected were as follows: Cyclin D1 forward,
5′-TCCTCTCCAAAATGCCAGAG-3′ and reverse, 5′-GGCGGATTGGAAATGAACTT-3′;
p21 forward, 5′-CATGGGTTCTGACGGACAT-3′ and reverse,
5′-AGTCAGTTCCTTGTGGAGCC-3′; and glyceraldehyde-3-phosphate
dehydrogenase (GAPDH) forward, 5′-GACTCATGACCACAGTCCATGC-3′ and
reverse, 3′-AGAGGCAGGGATGATGTTCTG-5′. The PCR reaction conditions
for all assays were as follows: 95°C for 30 sec, followed by 40
cycles of amplification (95°C for 5 sec, 59°C for 30 sec and 72°C
for 30 sec). The expression data was normalized to the geometric
mean of GAPDH to control the variability in expression levels and
calculated using 2-ΔΔCq.
Methyl thiazolyl tetrazolium (MTT) and
colony formation assays
The viability of the SKMEL-28 cells was measured by
MTT assay. The SKMEL-28 cells (3,000 cells/well) were seeded in
96-well plates in medium containing 10% FBS. The cell cultures were
stained on days 1–5. At the end of the stipulated time, 20 µl of 5
mg/ml MTT solution (Sigma-Aldrich) was added to each well and
incubated for 4 h at 37°C. Following the incubation period, the
culture medium was removed and 150 µl dimethyl sulfoxide
(Sigma-Aldrich) was added. The absorbance at 490 nm was measured
using a Multiskan microplate reader (Thermo Fisher Scientific,
Inc.).
For the colony formation assay, the SKMEL-28 cells
were plated in 6-well plates at a density of 1,000 cells per well,
and then incubated for 10 days in medium containing 10% FBS. The
colonies were stained with crystal violet (Beyotime Biotechnology
Institute of Biotechnology, Inc., Shanghai, China) and counted
using an Olympus BX41 microscope fitted with the cellSens software
(Olympus, Center Valley, PA, USA).
Anchorage-independent growth
assay
In total, 1,000 SKMEL-28 cells were resuspended in 2
ml complete medium supplemented with 0.3% agar (Sigma-Aldrich). The
agar-cell mixture was plated on top of a bottom layer consisting of
1% agar in complete medium. The cells were incubated for 14 days at
37°C until colony formation occurred, and the colonies were then
stained with 0.5% crystal violet prior to counting under a
microscope (DM IRBE; Leica, Heidelberg, Germany), and images of the
cell colonies were captured at an original magnification of ×100.
Colonies >0.1 mm in diameter were counted.
Flow cytometric analysis
A total of 48 h subsequent to transfection, the
SKMEL-28 cells were harvested and washed with PBS, and fixed with
70% ethanol. The fixed cells were then treated with 20 µg/ml RNaseA
and 50 µg/ml propidium iodide (Sigma-Aldrich) for 30 min, and
stained cells were immediately analyzed by a FACScan flow cytometer
(BD Biosciences, Franklin Lakes, NJ, USA).
Luciferase assays
The pGL3-luciferase reporter gene plasmid
pGL3-CYLD-3′-UTR (GeneCopoeia, Guangzhou, China) was cotransfected
into the cells with 15 pmol of the miR-186 mimic, miR-186 control
or miR-186-mut and 5 ng pRL-TK TKRenilla plasmid (Promega
Corporation) using Invitrogen Lipofectamine 2000 reagent. The
activity of firefly and andRenilla luciferase was assessed 48 h
subsequent to transfection using the dual luciferase assay kit
(Beyotime Biotechnology Institute of Biotechnology, Inc.),
according to the manufacturer's instructions. The results are
expressed as the ratio of of Renilla luciferase activity to firefly
luciferase activity.
Western blotting
Protein lysates were prepared using
radioimmunoprecipitation assay buffer (Cell Signaling Technology,
Inc., Danvers, MA, USA) and equal quantities of protein were
resolved using sodium dodecyl sulfate-polyacrylamide gel
electrophoresis (Beyotime Biotechnology Institute of Biotechnology,
Inc.). The gels were transferred onto polyvinylidene difluoride
membranes (EMD Millipore, Billerica, MA, USA) for 2 h at 4°C, run
at a current of 125 mA and blocked for 2 h. The membrane was
incubated overnight with the following rabbit antibodies anti-CYLD
(#8462), anti-cyclin D1 (#2978) and anti-p21 antibodies (#2947; all
primary antibodies dilution, 1:1,000; Cell Signaling Technology,
Inc.) and washed with Tris-buffered saline and Tween-20 (TBST;
Beyotime Biotechnology Institute of Biotechnology, Inc.) three
times, for 5 min each time. For control sample loading, the
blotting membranes were stripped and re-probed with an anti-β-actin
antibody (dilution, 1:5,000; A5316; Sigma-Aldrich). Subsequent to
washing with TBST and incubation with anti-rabbit horseradish
peroxidase-conjugated secondary antibody (Sigma-Aldrich) for 2 h at
room temperature, immunocomplexes were visualized using
chemiluminescence (GE Healthcare Bio-Sciences, Pittsburgh, PA,
USA), according to the manufacturer's instructions.
Statistical analysis
All statistical analyses, with the exception of
microarray data, were performed using SPSS 17.0 (SPSS, Inc.,
Chicago, IL, USA). Student's t-test was used to evaluate the
significance of the differences between two groups of data in all
the relevant experiments. P<0.05 was considered to indicate a
statistically significant difference.
Results
miR-186 expression was upregulated in
melanoma tissues and melanoma cell lines
To investigate the role of miR-186 expression in
melanoma development, the present study evaluated the expression
levels of miR-186 in melanoma tissues and melanoma cells by
RT-qPCR.
The current results demonstrated that the expression
levels of miR-186 in the melanoma tissues were consistently
increased compared with the matched normal tissues adjacent to the
tumor. Additionally, all 5 tested melanoma cell lines, consisting
of the A375-S2, SKMEL-28, SKMEL-5, MeWO and RPMI-7951 cell lines,
demonstrated significantly increased miR-186 levels compared with
the levels in NHEMs. These results revealed that miR-186 played an
important role in melanoma. Overall, these results suggest that
miR-186 expression is significantly increased in melanoma and may
act as a prognostic marker for patients with melanoma.
miR-186 promoted melanoma cell
proliferation
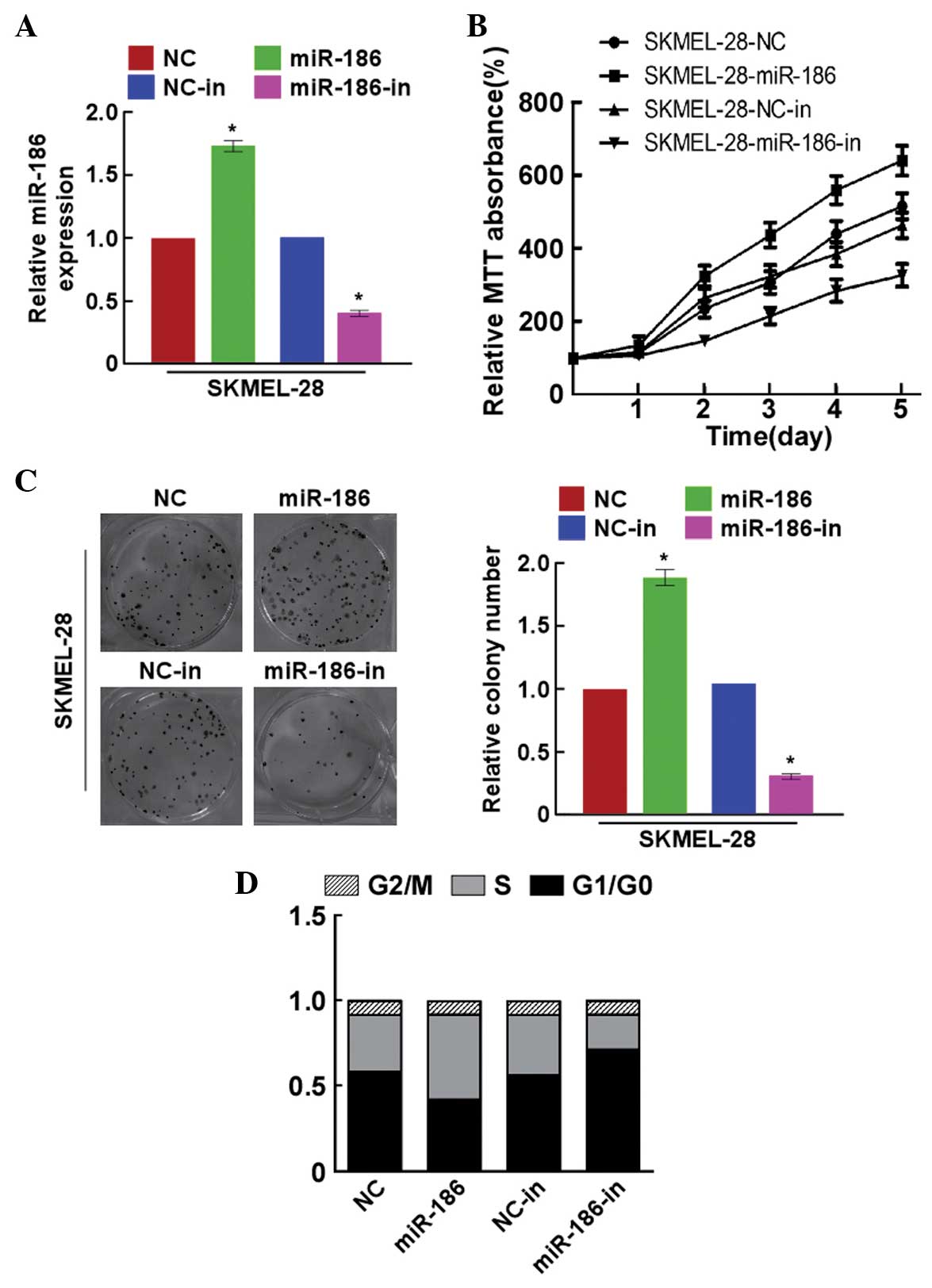
In order to explore the effects of miR-186 on the
growth of melanoma cells, the SKMEL-28 cells were transfected with
miR-186 mimics, miR-186 inhibitor or the respective controls. The
miR-186 precursor was found to upregulate miR-186 expression
(Fig. 2A). The results of the MTT
assays showed that miR-186 significantly promoted the proliferation
of SKMEL-28 cells (Fig. 2B), and this
was additionally confirmed by a colony formation assay (Fig. 2C). By contrast, the cell growth rates
and colony numbers of the SKMEL-28 cells transfected with
miR-186-in were significantly inhibited compared with the growth
rates and colony numbers of cells transfected with the NC (Fig. 2B and C). In addition,
miR-186-overexpressing SKMEL-28 cells had a significantly lower
percentage of cells in the G1/G0 phase and
increased percentage of cells in the S phase compared with the
NC-transfected cells (Fig. 2D).
miR-186-in demonstrated the opposite effects to miR-186. These
results revealed that miR-186 promoted melanoma tumorigenicity
in vitro.
miR-186 directly targets CYLD by
binding to its 3′-UTR and alters the levels of proteins associated
with cell proliferation and cell cycle in melanoma cells
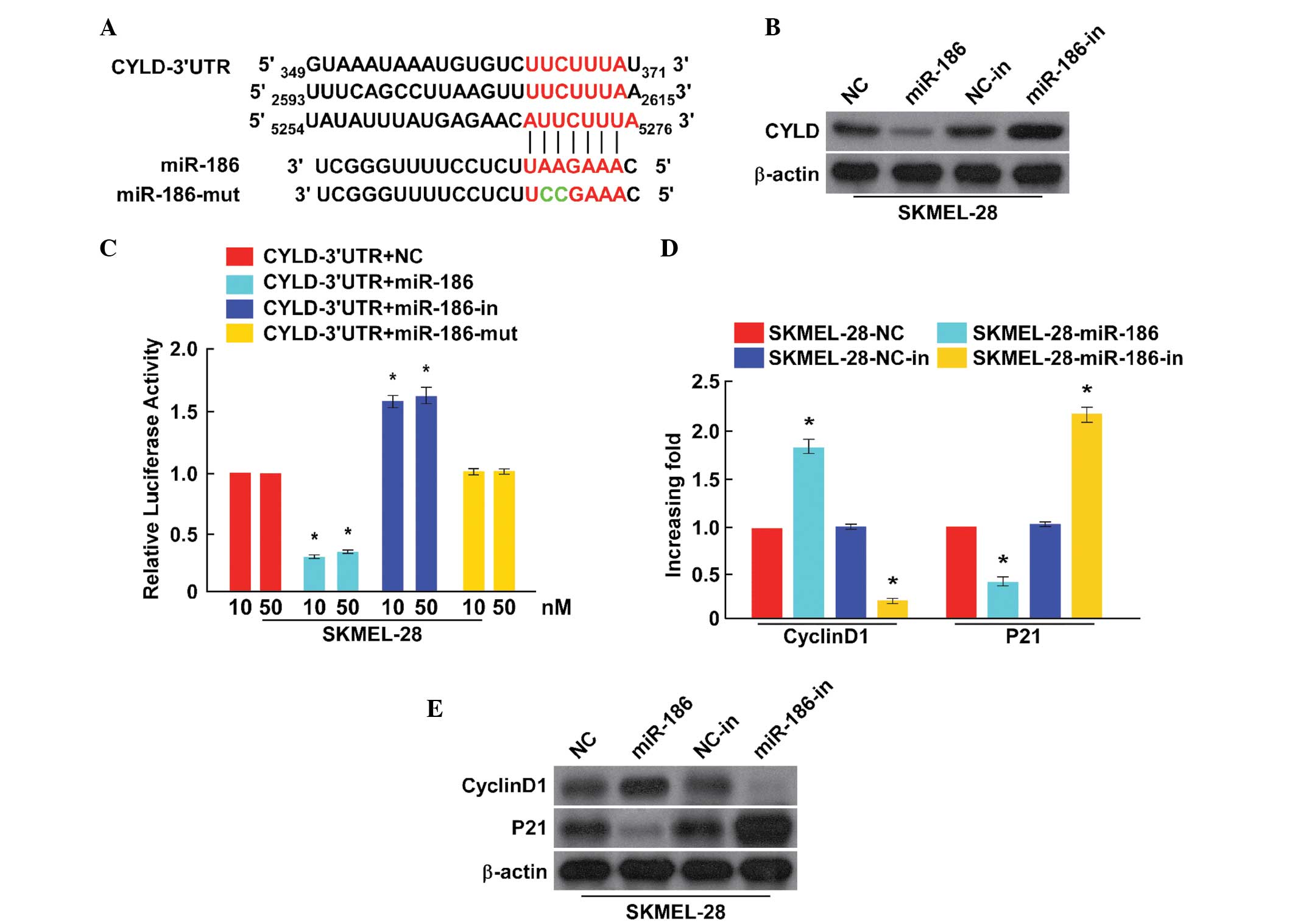
To explore the mechanism that facilitates the
effects on cell proliferation and cell cycle induced by miR-186,
the putative targets of miR-186 were analyzed by a bioinformatics
screen that was developed using TargetScan. The present study
focused on CYLD, from which originated a 3′-UTR containing a
binding site of miR-186 (Fig.
3A).
To verify the oncogenic role of miR-186, the present
study investigated the expression level of CYLD protein through
transfection with miR-186 mimics/inhibitors in SKMEL-28 cells. The
over-expression of miR-186 was found to result in a significant
reduction of CYLD protein expression in SKMEL-28 cells, while
miR-186-in clearly promoted CYLD protein expression (Fig. 3B). A luciferase reporter assay was
performed to determine whether miR-186 directly regulates the
expression of CYLD in SKMEL-28 cells. As shown in Fig. 3C, the results indicated that a
consistent and dose-dependent reduction of luciferase activity was
observed in SKMEL-28 cells transfected with the miR-186 mimic,
whereas the repressive effect of miR-186-in increased wild-type
CYLD luciferase activity. However, overexpressing miR-186-mut had
no effect on the luciferase activity of CYLD 3′-UTR. Overall, these
results demonstrated that CYLD is a target of miR-186.
It was demonstrated that miR-186 promoted the growth
and proliferation of melanoma and CYLD was a direct target of
miR-186 (12). Previously, it has
been reported that CYLD expression is associated with cell
proliferation and the cell cycle. In addition, the present study
observed alteration in the mRNA expression of the CYLD downstream
genes cyclin D1 and p21, which are critical cell-proliferation and
cell-cycle regulators. As shown in Fig. 4D, the cyclin D1 levels in
SKMEL-28 cells were all significantly upregulated by miR-186, and
downregulated by miR-186-in, while the expression of p21 was
downregulated in cells transfected with miR-186 and upregulated in
cells transfected with miR-186-in. Altogether, the present results
indicated that miR-186 functionally modulates cellular
proliferation and the cycle regulators cyclin D1 and p21, and are
therefore relevant to cell proliferation and the cell cycle.
Discussion
Numerous studies have shown that miRNAs are a class
of diverse, small, noncoding RNAs that function as critical gene
regulators, and then potentially play essential roles in multiple
biological processes, including cell differentiation,
proliferation, angiogenesis, invasion and migration (4,13–15). It is widely recognized that miRNAs
perform important roles in tumorigenesis by targeting various
mRNAs. At present, the abnormal expression of several miRNAs, such
as miRNA-573 (16), miRNA-106b
(17), miRNA-524-5p (18) and miRNA-451a (19), has been identified in melanoma and may
contribute to the development and progression of melanoma. In the
present study, miR-186 was identified as a tumor-promoter miRNA in
human melanoma that acts by repressing CYLD, which has been
previously identified as a tumor suppressive protein, provides
additional evidence of a pivotal role for miRNAs in the
tumorigenesis and progression of melanoma.
Previously, miRNAs have emerged as an important
regulator of various physiological and pathological processes in
cancer cells (10,20,21).
However, the functional role of miR-186 in the tumorigenesis of
melanoma remains largely unknown. In the present study, it was
found that miR-186 expression was markedly upregulated in melanoma
tissues and cells. Additional experiments revealed that miR-186
overexpression dramatically promoted the proliferation of SKMEL-28
cells, accompanied by a decrease in the proportion of cells in the
G1/G0 phase and an increase in the proportion
of cells in the S phase, while miR-186-in showed the opposite
function. These findings suggested that miR-186 was involved in the
processes of the development of melanoma.
CYLD, a deubiquitination enzyme, functions as a
tumor suppressor in various types of cancer (22,23). To
improve the understanding of the underlying mechanisms of
miR-186-induced melanoma cell growth and cell cycle progression,
CYLD was identified as a potential target gene of miR-186 using a
bioinformatics algorithm. The present results confirmed a vital
molecular association between miR-186 and CYLD. The present
experiment revealed that upregulation of miR-186 expression in
SKMEL-28 cells effectively suppressed CYLD expression, while
miR-186 increased the expression of the CYLD protein. Studies have
indicated that CYLD interferes with cell proliferation and cell
survival of cancer (23–25). The expression levels of a number of
critical cell-proliferation regulators were also detected. It has
also been reported that CDK proteins play critical roles in cell
proliferation and cell cycle transition. Among the cell
proliferation and cell cycle-associated genes, the CDK inhibitor
p21 and the CDK regulator cyclin D1 were focused on in the current
study. The present results showed that ectopic miR-186 expression
suppressed the expression level of the cell cycle inhibitor p21,
and increased the expression of the cell cycle regulator cyclin D1,
consequently leading to a decrease in the proportion of SKMEL-28
cells in the G0/G1 phase and an increase the
proportion of SKMEL-28 cells in the S phase.
In conclusion, the current study revealed that
miR-186 expression may potentially act as a method to determine the
progression of melanoma. The present results revealed that miR-186
regulated melanoma cell growth and cell cycle progression by
targeting the suppression of CYLD expression. Therefore, all the
results indicated that miR-186 was an oncomiR and may act as a
potential therapeutic target for the treatment of melanoma.
Acknowledgements
This study was supported by the Department of
Ophthalmology, Nansha Hospital, Guangzhou First People's Hospital,
Guangzhou Medical University.
References
|
1
|
Siegel R, Naishadham D and Jemal A: Cancer
statistics, 2013. CA Cancer J Clin. 63:11–30. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Baroudjian B, Pagès C and Lebbé C:
Melanoma, from diagnosis to treatment. Rev Infirm. 65:16–18. 2016.
View Article : Google Scholar
|
|
3
|
Luo C, Weber CE, Osen W, Bosserhoff AK and
Eichmuller SB: The role of microRNAs in melanoma. Eur J Cell Biol.
93:11–22. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Mimeault M and Batra SK: Novel biomarkers
and therapeutic targets for optimizing the therapeutic management
of melanomas. World J Clin Oncol. 3:32–42. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ambros V: The functions of animal
microRNAs. Nature. 431:350–355. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Cui G, Cui M, Li Y, Liang Y, Li W, Guo H
and Zhao S: MiR-186 targets ROCK1 to suppress the growth and
metastasis of NSCLC cells. Tumour Biol. 35:8933–8937. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Kim SY, Lee YH and Bae YS: MiR-186,
miR-216b, miR-337-3p and miR-760 cooperatively induce cellular
senescence by targeting α subunit of protein kinase CKII in human
colorectal cancer cells. Biochem Biophys Res Commun. 429:173–179.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cai J, Wu J, Zhang H, Fang L, Huang Y,
Yang Y, Zhu X, Li R and Li M: miR-186 downregulation correlates
with poor survival in lung adenocarcinoma, where it interferes with
cell-cycle regulation. Cancer Res. 73:756–766. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ries J, Vairaktaris E, Agaimy A, Kintopp
R, Baran C, Neukam FW and Nkenke E: miR-186, miR-3651 and miR-494:
Potential biomarkers for oral squamous cell carcinoma extracted
from whole blood. Oncol Rep. 31:1429–1436. 2014.PubMed/NCBI
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2-ΔΔCt method. Methods. 25:402–408. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Mathis BJ, Lai Y, Qu C, Janicki JS and Cui
T: CYLD-mediated signaling and diseases. Curr Drug Targets.
16:284–294. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Esquela-Kerscher A and Slack FJ:
Oncomirs-microRNAs with a role in cancer. Nat Rev Cancer.
6:259–269. 2006. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhong Q, Wang T, Lu P, Zhang R, Zou J and
Yuan S: miR-193b promotes cell proliferation by targeting Smad3 in
human glioma. J Neurosci Res. 92:619–626. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Shi L, Wang Z, Sun G, Wan Y, Guo J and Fu
X: miR-145 inhibits migration and invasion of glioma stem cells by
targeting ABCG2. Neuromolecular Med. 16:517–528. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wang HF, Chen H, Ma MW, Wang JA, Tang TT,
Ni LS, Yu JL, Li YZ and Bai BX: miR-573 regulates melanoma
progression by targeting the melanoma cell adhesion molecule. Oncol
Rep. 30:520–526. 2013.PubMed/NCBI
|
|
17
|
Prasad R and Katiyar SK: Down-regulation
of miRNA-106b inhibits growth of melanoma cells by promoting
G1-phase cell cycle arrest and reactivation of p21/WAF1/Cip1
protein. Oncotarget. 5:10636–10649. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liu SM, Lu J, Lee HC, Chung FH and Ma N:
miR-524-5p suppresses the growth of oncogenic BRAF melanoma by
targeting BRAF and ERK2. Oncotarget. 5:9444–9459. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Babapoor S, Fleming E, Wu R and Dadras SS:
A novel miR-451a isomiR, associated with amelanotypic phenotype,
acts as a tumor suppressor in melanoma by retarding cell migration
and invasion. PLoS One. 9:e1075022014. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Rico-Rosillo MG, Vega-Robledo GB and
Oliva-Rico D: The role and importance of the microRNAs in the
diagnosis and development of diseases. Rev Med Inst Mex Seguro Soc.
52:302–307. 2014.(In Spanish). PubMed/NCBI
|
|
21
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Deng LL, Shao YX, Lv HF, Deng HB and Lv
FZ: Over-expressing CYLD augments antitumor activity of TRAIL by
inhibiting the NF-κB survival signaling in lung cancer cells.
Neoplasma. 59:18–29. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Massoumi R: CYLD: A deubiquitination
enzyme with multiple roles in cancer. Future Oncol. 7:285–297.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Lim JH, Jono H, Komatsu K, Woo CH, Lee J,
Miyata M, Matsuno T, Xu X, Huang Y, Zhang W, et al: CYLD negatively
regulates transforming growth factor-β-signalling via
deubiquitinating Akt. Nat Commun. 3:7712012. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Blake PW and Toro JR: Update of
cylindromatosis gene (CYLD) mutations in Brooke-Spiegler syndrome:
Novel insights into the role of deubiquitination in cell signaling.
Hum Mutat. 30:1025–1036. 2009. View Article : Google Scholar : PubMed/NCBI
|