Introduction
Lung cancer is a leading cause of cancer-associated
mortality, and has an increased incidence and mortality compared
with other cancers worldwide (1).
Among the various histological subtypes of lung cancer,
adenocarcinoma is the most common subtype in the majority of
countries, accounting for approximately one-half of lung cancers
(2). At present, numerous studies
indicate that there is a particular subpopulation of tumor cells
responsible for the prosperity and metastasis of tumors, although
these cells are rare. These cells are commonly termed cancer stem
cells (CSCs) or cancer initiating cells. The possible presence of
CSCs has now been shown in the majority of cancers (3). Characterized by the abilities of
self-renewal, differentiation and tumorigenicity, CSCs are
considered to be able to form tumors with high efficiency, and also
re-build the tumor with the whole lineage of tumor cells (4). Although methods of identifying CSCs
remain to be elucidated, it is known that CSCs express surface
molecules, in addition to the identified functional characteristics
of stem cells; identification methods include measuring the
activity of aldehyde dehydrogenase (ALDH) by ALDEFLUOR assay. At
present, ALDH has been well accepted as a reliable CSC marker
(5). In addition to the role of
identifying the CSCs, the family of ALDH enzymes, which consists of
19 isoforms that localize to the cytoplasm and mitochondria of the
nucleus, also manifest the capability to oxidize endogenous and
exogenous aldehydes to carboxylic acids (6). Thus, ALDH also has a protective role in
the process of maintaining the homeostasis of cells, which may
benefit the survival of cancer cells. However, ALDHs also perform
other roles that include ester hydrolysis, acting as binding
proteins for cholesterol and acting as antioxidants in the process
of NADPH production (7). In general,
the diverse characteristics of ALDH are associated with the
stemness of CSCs.
Therefore, investigating the regulation of
expression and enzyme activity of ALDH is extremely important. In
the present study, the expression of ALDH1 and Nodal was assessed
in 28 cases of lung mixed adenocarcinoma, and the enzyme activity
of ALDH was also investigated in the lung adenocarcinoma A549 cell
line treated with histamine, rhNodal and the agonists and
antagonists of histamine H1 receptor (H1R) and H2 receptors by flow
cytometric assay.
Materials and methods
Lung mixed adenocarcinoma samples
A total of 28 lung mixed adenocarcinoma tissue
samples were obtained from the Pathology Department of Osaka
University Affiliated Hospital (Suita, Japan). All cases were
graded according to the 2011 World Health Organization
classification (8). The study was
approved by the Osaka University Ethical Review Board (approval no.
15234). The requirement for patient consent was waived by the
Ethical Review Board, due to the anonymized retrospective nature of
the present study.
Cell culture
The A549 cell line was obtained from the Cell
Repository of Osaka University and cultured at 37°C with 5%
CO2, high glucose Dulbecco's modified Eagle's medium
(Wako Pure Chemical Industries, Ltd., Osaka, Japan), 10% fetal calf
serum (HyClone; GE Healthcare Life Sciences, Logan, UT, USA), 100
U/ml penicillin and 100 µg/ml streptomycin (Invitrogen; Thermo
Fisher Scientific, Inc., Waltham, MA, USA).
Immunohistochemistry
Lung mixed adenocarcinoma tissues (28 samples) were
fixed using 10% formalin for 72 h at room temperature, embedded in
paraffin and then cut into 5 µm sections and mounted onto slides.
Subsequent to deparaffinization in xylene and rehydration in graded
ethanol, slides were pretreated for antigen retrieval using citrate
buffer (pH 6) under 120°C, 2,020 kPa for 20 min. Sections were
incubated with Dako Real Peroxidase-Blocking solution, which
contained 3% hydrogen peroxide, 15 mmol/l NaN3 and Tween-20
(Agilent Technologies, Inc., Santa Clara, CA, USA) for 5 min at
room temperature, and then washed three times with PBS containing
0.01% Tween (PBST) for 5 min each time. ALDH1 and Nodal were
detected by incubating the slides with mouse anti-human ALDH1
monoclonal antibodies (cat. no. 611194; BD Biosciences, San Jose,
CA, USA) and mouse anti-human Nodal monoclonal antibodies (cat. no.
ab55676; Abcam, Tokyo, Japan) at a dilution of 1:200 at room
temperature for 1 h. Subsequent to washing, the slides were treated
with horseradish peroxidase-conjugated rabbit anti mouse IgG (cat.
no. 354, Medical & Biological Laboratories Co., Ltd., Nagoya,
Japan) for 30 min at room temperature. Subsequent to washing with
PBST, the slides were incubated with Dako Detection Reagent
Envision kit (Agilent Technologies, Inc.) for 5–15 sec at room
temperature and were counterstained with hematoxylin for 1 min,
followed by dehydration and sealing.
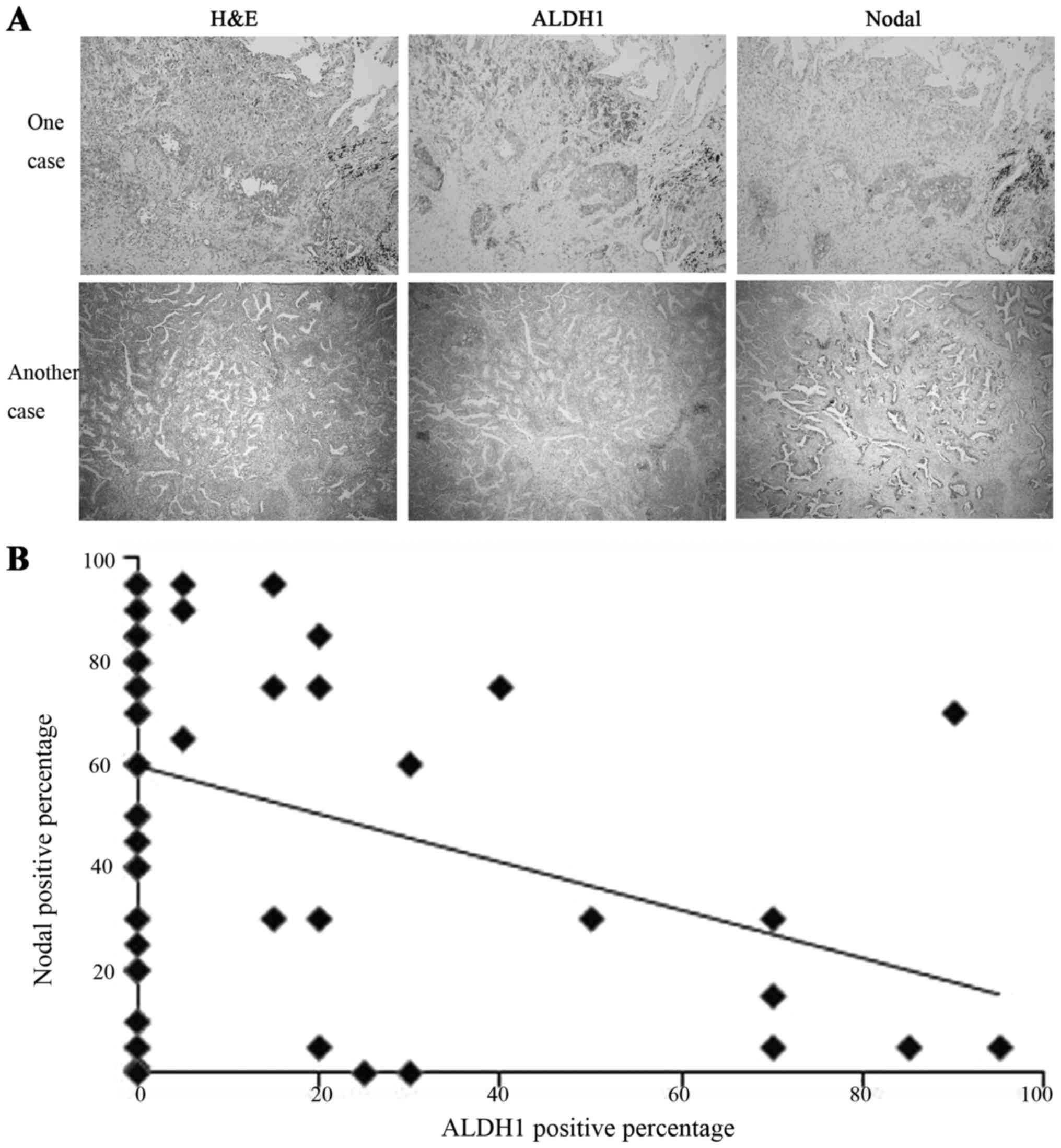
Antibody binding was recognized as brown cytoplasmic
staining by light microscope (magnification, ×100; Fig. 1A). Each sample of lung mixed
adenocarcinoma consisted of multiple subtypes, such as the lepidic,
acinar, papillary, micropapillary and solid with mucin production
subtypes. The percentage of ALDH1-positive tumor cells among each
subtype, including lepidic, acinar, and papillary subtypes, was
calculated. Similarly, the percentage of Nodal-positive tumor cells
was calculated. In each subtype, the percentage of ALDH-positive
and that of Nodal-positive cells were plotted (Fig. 1B). Based on the pots, the association
between ALDH1 and Nodal could be measured using a nonparametric
statistical method. P<0.05 was considered to indicate a
statistically significant difference.
Flow cytometry
In the present study, the living A549 cells were
directly stained without fixation. Reagents used to treat A549
cells at 37°C included: Recombinant human Nodal (rhNodal; 100, 300
or 500 ng/ml; duration, 2 h; cat. no. OLF0812051; R&D Systems,
Inc., Minneapolis, MN, USA); histamine dihydrochloride (10, 100,
500 µg/ml; duration, 2,4,18 h; Sigma-Aldrich; Merck KGaA); and the
agonists and antagonists of human histamine receptors H1R and H2R.
The agonist of H1R [histamine trifluoromethyltoluididedimaleate
(HTM); concentration, 100 µM; duration, 2 h] agonist of H2R
[dimapritdihydrochloride (DIM); concentration, 100 µM; duration, 2
h] and the antagonist of H2R [cimetidine (CIM); concentration, 100
µM; duration, 2 h] were all purchased from Tocris Bioscience
(Bristol, UK). The antagonist of H1R [pyrilamine maleate salt
(PYR); concentration, 100 µM; duration, 2 h] was purchased from
Sigma-Aldrich (Merck KGaA).
A549 cells were plated onto a 6-well plate and were
cultured for 2, 4 or 18 h with the reagents (10, 100 or 500 µg/ml
histamine; 100, 300 or 500 ng/ml rhNodal, 100 µM MHTM, 100 µM DIM,
100 µM PYR and 100 µM CIM). Adherent A549 cells were then harvested
with trypsin containing 0.25% EDTA (Gibco; Thermo Fisher
Scientific, Inc.) at a concentration of 2×105 viable
cells for each sample, and were applied in the detection of the
enzymic activity of ALDH. The percentage of ALDH-positive A549
cells was used to reflect the ALDH activity (9) by using the ALDEFLUOR assay kit, which
contained fluorescent substrate for ALDH, termed
boron-dipyrromethene (BODIPY) aminoacetaldehyde, that consisted of
an aminoacetaldehyde moiety bonded to the BODIPY fluorochrome
(Stemcell Technologies, Inc., Vancouver, BC, Canada). The labeled
cells were analyzed using a BD FACSCantoII (BD Biosciences) and
analyzed using FACSDiva6.1.3 software (BD Biosciences). Each sample
was measured three times.
Reverse transcription-polymerase chain
reaction
Total RNA was extracted from A549 cells using a
RNeasy Mini kit (Qiagen, Inc., Valencia, CA, USA) and converted
into cDNA and amplified using a Super Script III RNase H-Reverse
Transcriptase kit (cat. no. 1244390; Invitrogen; Thermo Fisher
Scientific, Inc.). The resulting cDNA was amplified in a final
volume of 25 µl containing 2.5 pmol of each primer and 12.5 µl of
PrimeSTAR MAX DNA polymerase (Takara Bio, Inc., Otsu, Japan). The
amplification conditions were denaturation at 98°C for 10 sec (5
min for the first cycle), annealing at 55°C for 5 sec, and
extension at 72°C for 1 min for 40 cycles for the amplification of
H1R, H2R and H4R. The amplification conditions of H3R were
denaturation at 94°C for 30 sec (5 min for the first cycle),
annealing at 59°C for 30 sec, and extension at 72°C for 1min (5 min
for the last cycle) for 40 cycles. The amplification products (each
10 µl) were separated on 2% agarose and stained by ethidium
bromide. The primers (Applied Biosystems; Thermo Fisher Scientific,
Inc.) of H1R, H2R, H3R and H4R were used as below (10): H1R forward,
5′-CATTCTGGGGGCCTGGTTTCTCT-3′ and reverse,
5′-CTTGGGGGTTTGGGATGGTGACT-3′; H2R forward, 5′-CCGGCTCCGCAACCT-3′
and reverse, 5′-CTGATCCCGGGCGACCTTGA-3′; H3R forward,
5′-TCAGCTACGACCGCTTCCTGTCGGTCAC-3′ and reverse,
5′-TTGAGTGAGCGCGGCCTCTCAGTGCCCC-3′; and H4R forward,
5′-GAATTGTCTGGCTGGATTAATTTGCTAATTTG-3′ and reverse,
5′-AAGAATGATGTGATGGCAAGGATGTACC-3′.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
RT-qPCR was performed using 200 ng cDNA derived from
RNA extracted from A549 cells using a RNeasy Mini kit (Qiagen,
Inc.) and reverse transcribed using a Super Script III RNase
H-Reverse Transcriptase kit (cat. no. 1244390; Invitrogen; Thermo
Fisher Scientific, Inc.), and 1.25 µl of specific primer pairs in
TaqMan Gene Expression Master mix (cat. no. 1110130; Applied
Biosystems; Thermo Fisher Scientific, Inc.) in a 25 µl reaction
mixture. The human GAPDH primer was purchased from Applied
Biosystems (cat. no. 4326317E; Thermo Fisher Scientific, Inc.). The
aldehyde dehydrogenase 1A1 (ALDH1A1) primer was also purchased from
Applied Biosystems (cat. no. 1164131; Thermo Fisher Scientific,
Inc.). Thermocycling conditions were as according to the
manufacturer's protocol: 1 cycle at 48°C for 30 min, 1 cycle at
95°C for 10 min and 50 cycles each at 95°C for 15 sec and 60°C for
1 min. Comparative gene expression analysis was performed using
normalization to the level of the internal control gene, GAPDH
using the 2−ΔΔCq method (11).
Statistical analysis
All data are expressed as the mean ± standard
deviation, and statistical analyses were carried out by SPSS 17.0
(SPSS, Inc., Chicago, IL, USA). Statistical analysis was performed
using one-way ANOVA and correlations were analyzed using Spearman's
rank correlation coefficient. P<0.05 was considered to indicate
a statistically significant difference. Experiments data were
conducted in triplicate.
Results
Expression of ALDH and Nodal is
negatively correlated in lung mixed adenocarcinomas, and rhNodal
downregulated the expression of ALDH on A549 cells
Immunostaining of the expression of ALDH1 and Nodal
in lung mixed adenocarcinomas is shown in Fig. 1 and Table
I. ALDH1 and Nodal were each expressed in the cytoplasm of the
lung cancer cells. For a diagnosis of lung mixed adenocarcinoma,
each case must consist of more than one subtype; the percentage of
different histology types, and percentage of the positive
expression of ALDH1 and Nodal in their respective histology types
were evaluated and compared to identify the statistical
significance. The results indicated that the expression of ALDH1
and Nodal was inversely correlated (Fig.
1B). In the ALDH1-positive expression area, the expression of
Nodal was negative or weak, and vice versa.
 | Table I.Expression of ALDH1 and Nodal in 28
cases of lung mixed adenocarcinoma. |
Table I.
Expression of ALDH1 and Nodal in 28
cases of lung mixed adenocarcinoma.
| Case number | Histology
subtypes | Histology
percentage | ALDH1, % | Nodal, % |
|---|
| 1 | Lepidic | 40 | 70 | 30 |
|
| Acinar | 60 | 85 | 5 |
| 2 | Lepidic | 70 | 0 | 40 |
|
| Papillary | 30 | 50 | 30 |
| 3 | Papillary | 80 | 15 | 30 |
|
| Acinar | 20 | 25 | 0 |
| 4 | Papillary | 10 | 0 | 95 |
|
| Micropapillary | 2 | 0 | 95 |
|
| Acinar | 8 | 0 | 95 |
|
| Solid with mucin | 80 | 0 | 75 |
| 5 | Papillary | 90 | 90 | 65 |
|
| Micropapillary | 5 | 5 | 20 |
|
| Solid with mucin | 5 | 5 | 60 |
| 6 | Lepidic | 70 | 0 | 10 |
|
| Papillary | 30 | 0 | 50 |
| 7 | Lepidic | 20 | 0 | 20 |
|
| Papillary | 80 | 0 | 40 |
| 8 | Lepidic | 90 | 0 | 1 |
|
| Acinar | 10 | 0 | 0 |
| 9 | Lepidic | 70 | 0 | 45 |
|
| Papillary | 10 | 30 | 0 |
|
| Acinar | 20 | 95 | 5 |
| 10 | Papillary | 10 | 0 | 5 |
|
| Micropapillary | 10 | 0 | 30 |
|
| Solid with
mucin | 80 | 0 | 20 |
| 11 | Papillary | 70 | 0 | 85 |
|
| Acinar | 30 | 0 | 60 |
| 12 | Papillary | 70 | 70 | 5 |
|
| Micropapillary | 10 | 5 | 95 |
|
| Solid with
mucin | 20 | 0 | 85 |
| 13 | Papillary | 40 | 15 | 95 |
|
| Micropapillary | 30 | 0 | 95 |
|
| Solid with
mucin | 30 | 0 | 80 |
| 14 | Lepidic | 10 | 0 | 30 |
|
| Papillary | 45 | 0 | 75 |
|
| Acinar | 45 | 0 | 40 |
| 15 | Lepidic | 10 | 0 | 85 |
|
| Papillary | 40 | 0 | 90 |
|
| Acinar | 35 | 0 | 95 |
|
| Others | 15 | 0 | 0 |
| 16 | Papillary | 95 | 0 | 70 |
|
| Solid with
mucin | 5 | 0 | 80 |
| 17 | Lepidic | 40 | 0 | 90 |
|
| Papillary | 60 | 0 | 85 |
| 18 | Lepidic | 30 | 30 | 60 |
|
| Papillary | 70 | 20 | 75 |
| 19 | Lepidic | 15 | 0 | 75 |
|
| Papillary | 85 | 0 | 90 |
| 20 | Lepidic | 50 | 0 | 70 |
|
| Papillary | 20 | 0 | 5 |
|
| Micropapillary | 30 | 0 | 25 |
| 21 | Lepidic | 60 | 20 | 85 |
|
| Papillary | 20 | 90 | 70 |
|
| Micropapillary | 20 | 0 | 80 |
| 22 | Papillary | 90 | 40 | 75 |
|
| Micropapillary | 10 | 20 | 30 |
| 23 | Papillary | 70 | 0 | 45 |
|
| Micropapillary | 10 | 0 | 10 |
|
| Acinar | 20 | 0 | 70 |
| 24 | Lepidic | 45 | 70 | 15 |
|
| Acinar | 55 | 0 | 50 |
| 25 | Lepidic | 40 | 15 | 75 |
|
| Papillary | 20 | 0 | 80 |
|
| Acinar | 40 | 5 | 90 |
| 26 | Papillary | 10 | 0 | 80 |
|
| Micropapillary | 5 | 0 | 85 |
|
| Acinar | 10 | 0 | 50 |
|
| Solid with
mucin | 75 | 0 | 75 |
| 27 | Acinar | 75 | 70 | 5 |
|
| Solid with
mucin | 25 | 0 | 85 |
| 28 | Lepidic | 70 | 0 | 70 |
|
| Papillary | 30 | 0 | 80 |
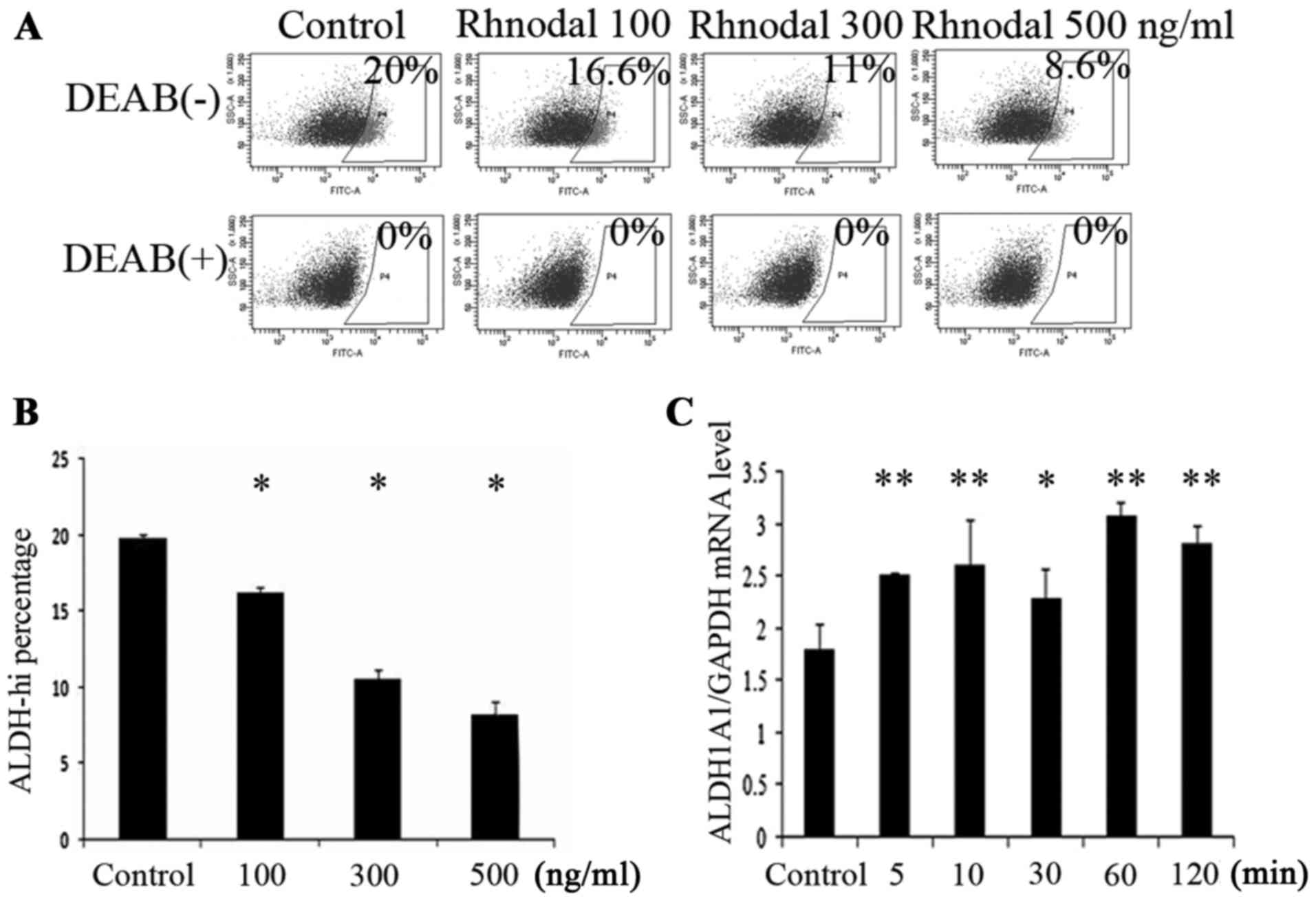
To detect whether the Nodal regulated the enzymatic
activity of ALDH, A549 cells were cultured at different
concentrations (100, 300 and 500 ng/ml) of rhNodal for 2 h, then
the activity of ALDH of the A549 cells was analyzed by flow
cytometry (Fig. 2A and B). The result
indicated that the activity of ALDH was downregulated by
rhNodal.
Activity of ALDH was upregulated by
histamine in A549 cells
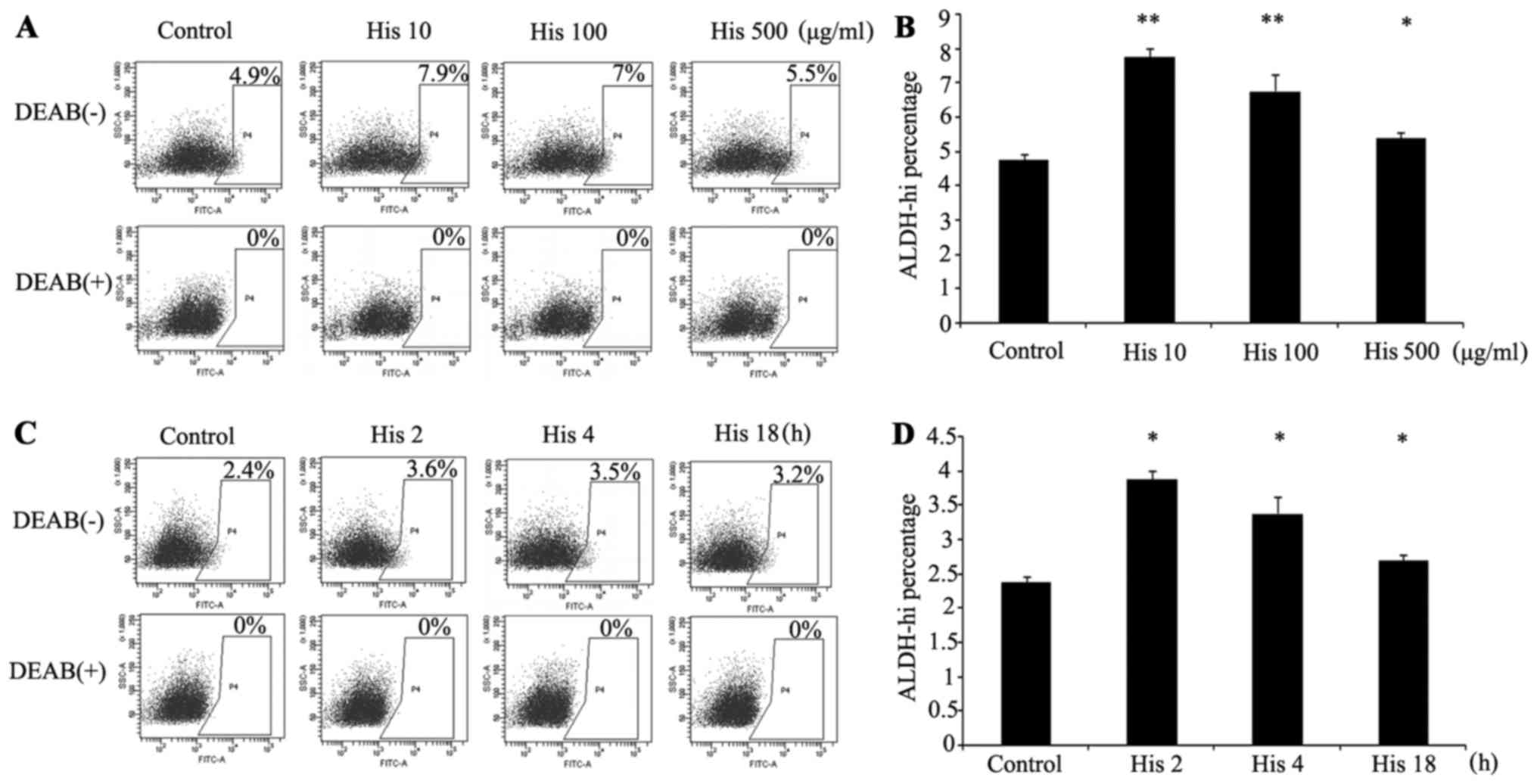
To determine whether the expression of ALDH is
regulated by histamine, subsequent to treatment with histamine at
different concentrations and for different durations, the ALDH
activity of the A549 cells was detected by flow cytometry (Fig. 3). The results indicated that the
activity of ALDH in the A549 cell line could be upregulated by
histamine at a concentration of 10 µg/ml and the suitable culturing
time span was 2 h. The mRNA level of ALDH1A1 was also detected
after treatment with histamine for different durations. The results
showed that the mRNA levels of ALDH1A1 of A549 cells increased with
the duration of histamine treatment (Fig.
2C).
H1R and H2R are expressed in A549
cells
The aforementioned observations prompted the
investigation of the histamine receptors present in the A549 cell
line. RT-qPCR was used, and the presence and levels of mRNA of
histamine receptors were detected (Fig.
4). The results indicated that the subtypes of histamine
receptor present in A549 cells are H1R and H2R, but not H3R and
H4R.
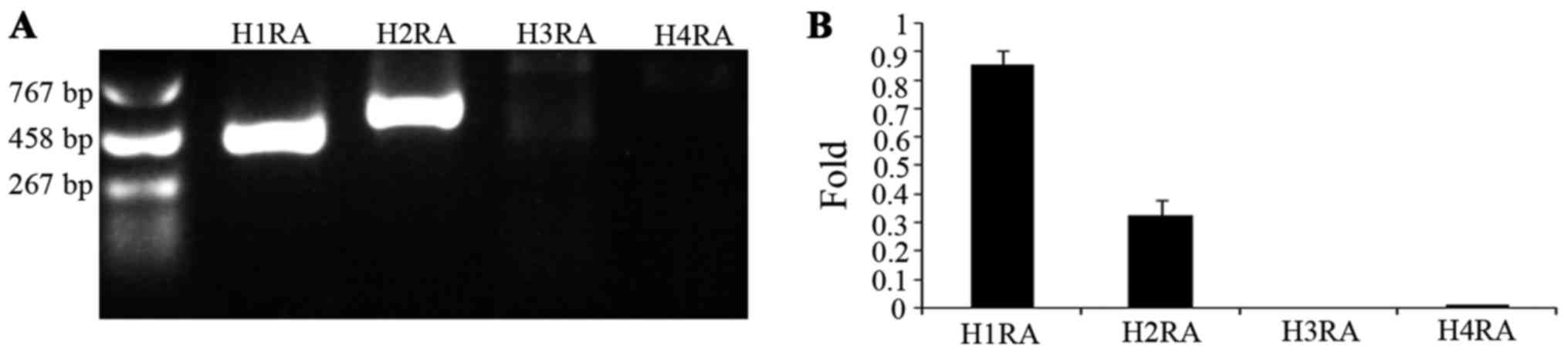 | Figure 4.Expression of human histamine
receptors in A549 cells. (A) Reverse transcription-polymerase chain
reaction analysis of histamine receptor mRNA expression. Lane H1RA,
H1R in A549 cells (402 bp); lane H2RA, H2R in A549 cells (495 bp);
lane H3RA, H3R in A549 cells; lane H4R, H4R in A549 cells. (B)
Ratio of mRNA expression of H1R, H2R, H3R and H4R to the expression
of GAPDH. H1R, H1 receptor; H2R, H2 receptor; H3R, H3 receptor;
H4R, H4 receptor. |
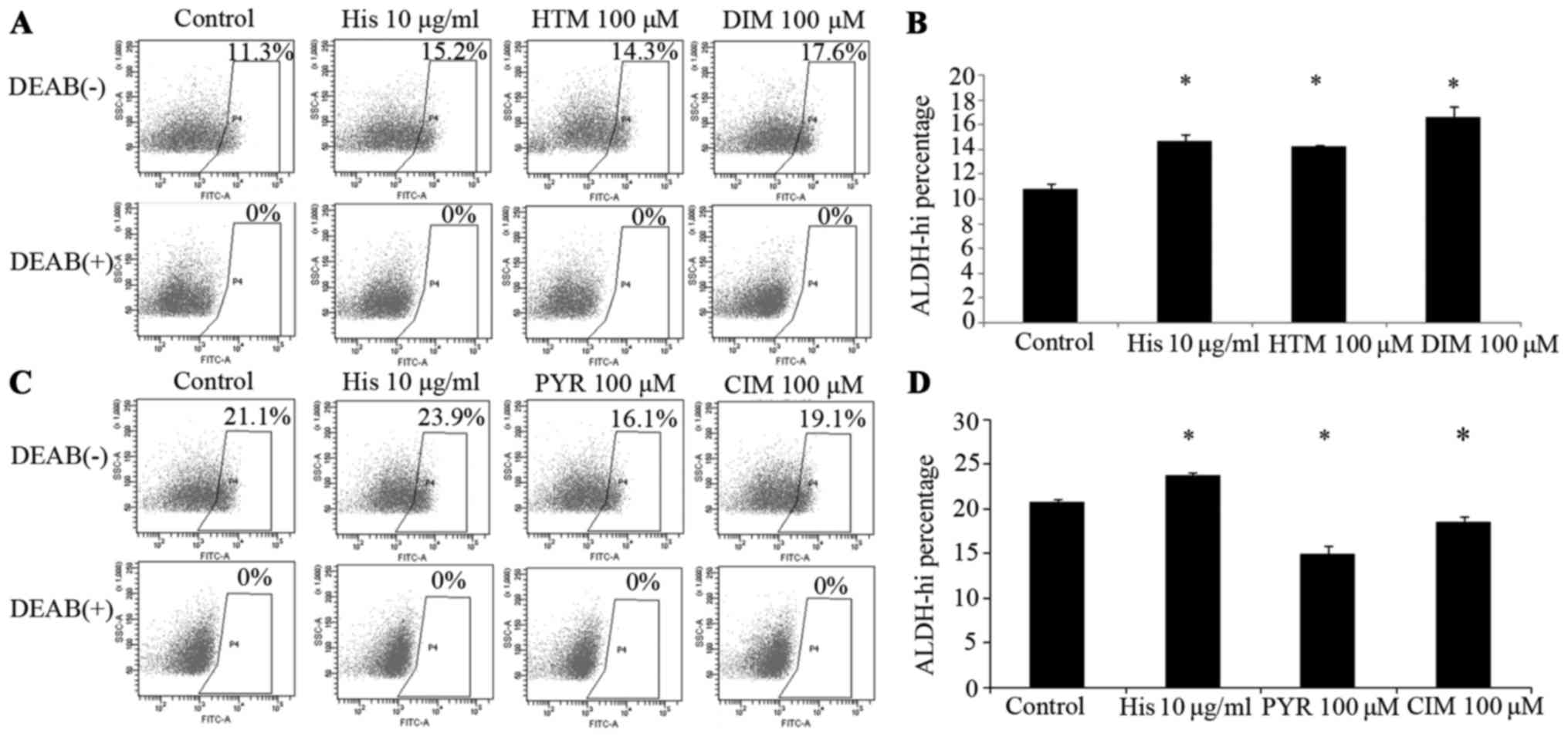
ALDH activity was regulated by H1R and
H2R agonists and antagonists in A549 cells
Histamine has multiple biological roles and exerts
its effects via histamine receptors. Since histamine could
upregulate the expression of ALDH in A549 cells, whether the
agonists and antagonists of H1R and H2R affect the expression of
ALDH was investigated. A549 cells were cultured with agonists and
antagonists of H1R, H2R and histamine at different concentrations
for 2 h, and the activity of ALDH was detected by flow cytometry.
The data showed that ALDH activity was upregulated by H1R and H2R
agonists, and ALDH activity was downregulated by H1R and H2R
antagonists (Fig. 5).
Discussion
In the present study, it was confirmed that the
expression of ALDH1 is inversely associated with Nodal expression
in lung mixed adenocarcinoma, and A549 cells express H1R and H2R,
but not the other histamine receptors. The enzymatic activity of
ALDH could be upregulated by histamine and agonists of H1R and H2R,
as well as downregulated by rhNodal and antagonists of H1R and H2R
in the A549 cell line.
Nodal is a member of the transforming growth
factor-β (TGF-β) family, and it is an important morphogen,
performing regulatory function of cell fate in embryological and
adult systems (12). It has been
reported that TGF-β/activin/Nodal signaling may have an effect in
maintaining pluripotency in human embryonic stem (ES) cells
(13). In addition, Nodal appears to
maintain the stemness of stem cells by inhibiting differentiation
of neuroectodermal differentiation in the human ES cells (14). Mechanically, the activation of the
Nodal receptor may lead to the phosphorylation of Smad2/3.
Subsequently, Smad4 and other transcription factors are activated.
The activated Smad complexes accumulate in the nucleus, then
recognize and bind the Nodal-responsive cis-elements to
regulate the expression of their target genes (15). Smad proteins can also form complexes
with the other proteins, such as p53, and contribute to the
specific recognition and regulation of subsets of Nodal target
genes (16). In the present study,
Nodal inhibited the expression of ALDH, possibly through a
signaling pathway (e.g., TGF-β), which is consistent with a
previous study in which ALDH1 was revealed to occur in diffuse-type
gastric carcinoma-initiating cells and the expression of ALDH1 and
the size of the ALDH1-positive cell population were reduced by
TGF-β (17). In agreement with this,
rhNodal was found to reduce ALDH activity in the present study.
Histamine has diverse biological roles and exerts
its effects via distinct receptor subtypes, consisting of H1, H2,
H3 and H4 receptors. In the majority of human cells, multiple types
of histamine receptors coexist. However, H3R is exclusively
expressed in neurons (18).
Furthermore, it has been reported that no human cell culture model
expresses sufficient endogenous levels of H3R to be detected
(19). In this previous study
(19), the human lung adenocarcinoma
A549 cell line was demonstrated to express H1R and H2R mRNA.
Histamine is known to have a notable role in acute
and chronic allergic inflammation (20) and also contributes to the
proliferation and differentiation of certain cells (21). The binding of histamine receptors and
their ligands could trigger serious of reactions, including
Ca2+ inflow (22), the
activation of phosphatidylinositol (3–5)-trisphosphate signaling pathway (23) and the activation of nuclear factor-κB
(NF-κB) signaling pathway (24), and
they consequently fulfill their various biological functions. It
has been reported by Muzio et al (25) that the treatment of A549 cells with
arachidonic acid led to a decrease in the enzymatic activity,
protein and mRNA levels of ALDH3A1, whereas the expression of
proliferator-activated receptor-γ increased, and the NF-κB binding
activity was inhibited. Therefore, the expression of ALDH1 may be
indirectly associated with the histamine in A549 cells through a
NF-κB-associated signaling pathway. In the present study, the
activity of ALDH was observed to be slightly enhanced by treatment
with histamine, and this process was not only promoted by agonists
of H1R and H2R, also suppressed by the antagonists of H1R and
H2R.
To conclude, ALDH serves an important function in
labeling and protecting CSCs. The exact regulatory mechanism of
ALDH expression requires delineation, and therefore further
investigation in detail.
References
|
1
|
Parkin DM, Bray F, Ferlay J and Pisani P:
Global cancer statistics, 2002. CA Cancer J Clin. 55:74–108. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Egevad L, Heanue M, Berney D, Fleming K
and Ferlay J: Histological groupsCancer Incidence in Five
Continents. IX. Curado MP, Edwards B, Shin HR, Storm H, Ferlay J,
Heanue M and Boyle P: IARC Scientific Publications; Lyon: pp.
62–64. 2007
|
|
3
|
Marcato P, Dean CA, Giacomantonio CA and
Lee PW: Aldehyde dehydrogenase: Its role as a cancer stem cell
marker comes down to the specific isoform. Cell Cycle.
10:1378–1384. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Al-Hajj M, Wicha MS, Benito-Hernandez A,
Morrison SJ and Clarke MF: Prospective identification of
tumorigenic breast cancer cells. Pro Natl Acad Sci USA.
100:3983–3988. 2003. View Article : Google Scholar
|
|
5
|
Ginestier C, Hur MH, Charafe-Jauffret E,
Monville F, Dutcher J, Brown M, Jacquemier J, Viens P, Kleer CG,
Liu S, et al: ALDH1 is a marker of normal and malignant human
mammary stem cells and a predictor of poor clinical outcome. Cell
Stem Cell. 1:555–567. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Marchitti SA, Brocker C, Stagos D and
Vasiliou V: Non-P450 aldehyde oxidizing enzymes: The aldehyde
dehydrogenase superfamily. Expert Opin Drug Metab Toxicol.
4:697–720. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Black W and Vasiliou V: The aldehyde
dehydrogenase gene superfamily resource center. Hum Genomics.
4:136–142. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Travis WD, Brambilla E, Noguchi M,
Nicholson AG, Geisinger KR, Yatabe Y, Beer DG, Powell CA, Riely GJ,
Van Schil PE, et al: International association for the study of
lung cancer/American thoracic society/European respiratory society
international multidisciplinary classification of lung
adenocarcinoma. J Thorac Oncol. 6:244–285. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Huang EH, Hynes MJ, Zhang T, Ginestier C,
Dontu G, Appelman H, Fields JZ, Wicha MS and Boman BM: Aldehyde
dehydrogenase 1 is a marker for normal and malignant human colonic
stem cell (SC) and tracks SC overpopulation during colon
tumorgenesis. Cancer Res. 69:3382–3389. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fujimoto S, Komine M, Karakawa M, Uratsuji
H, Kagami S, Tada Y, Saeki H, Ohtsuki M and Tamaki K: Histamine
differentially regulates the production of Th1 and Th2 chemokinesby
keratinocytes through histamine H1 receptor. Cytokine. 54:191–199.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Strizzi L, Postovit LM, Margaryan NV,
Lipavsky A, Gadiot J, Blank C, Seftor RE, Seftor EA and Hendrix MJ:
Nodal as a biomarker for melanoma progression and a new therapeutic
target for clinical intervention. Expert Rev Dermatol. 4:67–78.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
James D, Levine AJ, Besser D and
Hemmati-Brivanlou A: TGFbeta/activin/nodal signaling is necessary
for the maintenance of pluripotency in human embryonic stem cells.
Development. 132:1273–1282. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Vallier L, Reynolds D and Pedersen RA:
Nodal inhibits differentiation of human embryonic stem cell salong
the neuroectodermal default pathway. Dev Biol. 275:403–421. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Massagué J, Seoane J and Wotton D: Smad
transcription factors. Genes Dev. 19:2783–2810. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Schier AF: Nodal morphogens. Cold Spring
Harb Perspect Biol. 1:a0034592009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Katsuno Y, Ehata S, Yashiro M, Yanagihara
K, Hirakawa K and Miyazono K: Coordinated expression of REG4 and
aldehyde dehydrogenase 1 regulating tumourigenic capacity of
diffuse-type gastric carcinoma-initiating cells is inhibited by
TGF-β. J Pathol. 228:391–404. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Leurs R, Bakker RA, Timmerman H and de
Esch IJ: The histamine H3 receptor: From gene cloning to H3
receptor drugs. Nat Rev Drug Discov. 4:107–120. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Seifert R, Strasser A, Schneider EH,
Neumann D, Dove S and Buschauer A: Molecular and cellular analysis
of human histamine receptor subtypes. Trends Pharmacol Sci.
34:33–58. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Akdis CA and Blaser K: Histamine in the
immune regulation of allergic inflammation. J Allergy Clin Immunol.
112:15–22. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Falus A, Pós Z and Darvas Z: Histamine in
normal and malignant cell proliferation. Adv Exp Med Biol.
709:109–123. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Hill SJ, Ganellin CR, Timmerman H,
Schwartz JC, Shankley NP, Young JM, Schunack W, Levi R and Haas HL:
International Union of Pharmacology. XIII. Classification of
histamine receptors. Pharmacol Rev. 49:253–278. 1997.PubMed/NCBI
|
|
23
|
Robinson AJ and Dickenson JM: Activation
of the p38 and p42/p44 mitogen-activated protein kinase families by
the histamine H(1) receptor in DDT(1)MF-2 cells. Br J Pharmacol.
133:1378–1386. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Bakker RA, Schoonus SB, Smit MJ, Timmerman
H and Leurs R: Histamine H(1)-receptor activation of nuclear
factor-kappa B: Roles for G beta gamma- and G alpha(q/11)-subunits
in constitutive and agonist-mediated signaling. Mol Pharmacol.
60:1133–1142. 2001.PubMed/NCBI
|
|
25
|
Muzio G, Trombetta A, Maggiora M,
Martinasso G, Vasiliou V, Lassen N and Canuto RA: Arachidonic acid
suppresses growth of human lung tumor A549 cellsthrough
down-regulation of ALDH3A1 expression. Free Radic Biol Med.
40:1929–1938. 2006. View Article : Google Scholar : PubMed/NCBI
|