Introduction
Liver cancer is the sixth-leading cause of cancer
and the second-leading cause of cancer-associated mortality
worldwide (1), leading to 746,000
mortalities in 2012 (1).
Hepatoblastoma, an embryonal malignancy originated in hepatocytes,
is the most common primary liver tumor in children, and is usually
diagnosed in the first 3 years of life (2). The incidence of hepatoblastoma in
children has gradually increased, and the overall 5-year survival
rate in children is ~70% (3,4). The most common manifestation of
hepatoblastoma is abdominal distension or presence of a mass
(2). Hepatoblastoma can generally be
cured via tumor resection if the tumor is single, smaller and
non-metastasic, but a number of children cannot be cured due to a
late diagnosis, multiple subtypes and metastases, which cause a
poor prognosis (5). Therefore, it is
necessary to investigate the molecular mechanisms of hepatoblastoma
and lay the basis for refining the current diagnosis or treatment
interventions. The HepG2 cell line is originated from
hepatoblastoma (6); thus, HepG2 cells
were selected in the present study to investigate the molecular
mechanisms of hepatoblastoma.
MicroRNAs (miRNAs/miRs) are small non-coding RNA
molecules involved in diverse cell activities, including regulation
of cell proliferation, differentiation, apoptosis and
carcinogenesis (5,6). miR-124 is the most abundant miRNA in the
adult brain and serves a tumor-suppressive role in diverse types of
cancer, including those of the breast and prostate (7,8). miR-124
is potentially involved in liver cancer: miR-124 suppresses
aggressive hepatocellular carcinoma by repressing Rho-associated,
coiled-coil containing protein kinase 2 and enhancer of zeste 2
polycomb repressive complex 2 subunit (9). miR-124 overexpression appears to repress
the migration and invasion of intrahepatic cholangiocarcinoma cells
in vitro (10). This evidence
indicates that miR-124 may serve a complicated tumor-suppressive
role in hepatoblastoma.; however, its underlying molecular
mechanism remains elusive.
The activity of miRNAs is negatively associated with
the expression of their target mRNAs (11–13). On
this basis, miR-124 transfection has been used to identify its
downregulated target mRNAs by microarray time-course experiments
(14). These target genes may be
implicated in various biological functions and signaling pathways,
thereby contributing to the progression of hepatoblastoma. To
further investigate the underlying mechanisms of miR-124
overexpression in hepatoblastoma, the present study applied a range
of bioinformatics approaches to a microarray dataset (GSE6207).
Differentially expressed genes (DEGs) between human hepatoblastoma
HepG2 cells transfected with a miR-124 duplex or a negative control
RNA duplex at different time points (4, 8, 16, 24, 32, 72 and 120
h) were screened, followed by Kyoto Encyclopedia of Genes and
Genomes (KEGG) pathway enrichment analysis for the screened DEGs.
Changes in miR-124 activity over the time course were also
analyzed. miR-124-target mRNA pairs were then selected to construct
miR-124-target mRNA networks for each time point. Gene ontology
(GO) function and KEGG pathway enrichment analysis were performed
for the targets genes of miR-124. The present study therefore
provides additional data concerning the molecular mechanisms of
miR-124 in hepatoblastoma.
Materials and methods
Microarray data and preprocessing
The present study used the GSE6207 microarray
dataset, downloaded from the National Center for Biotechnology
Information Gene Expression Omnibus (GEO; https://www.ncbi.nlm.nih.gov/geo/) repository based on
the Affymetrix Human Genome U133 Plus 2.0 Array platform (14). It included RNA samples extracted from
HepG2 cells transfected with a miRNA-124 duplex or a negative
control, with one sample for each time point (4, 8, 16, 24, 32, 72
and 120 h). The gene expression data were sequentially preprocessed
with background correction, quantile normalization and probe
summarization using robust multiarray analysis (RMA) of the affy
package in R (15). The expression
values of several probes that correspond to a certain gene were
averaged, and the averaged expression value was defined as the
expression value of the gene.
DEG screening
DEGs at each time point were screened in the HepG2
cells transfected with the miRNA-124 duplex and those transfected
with the negative control RNA duplex. As there was only one sample
for each experimental condition at each time point,
|log2 fold change| ≥0.58 was set as the strict cutoff
(16). The expression values of DEGs
at different time points were hierarchically clustered using
Cluster software (version 3.0; http://bonsai.hgc.jp/~mdehoon/software/cluster/software.htm),
which was originally developed by Michael Eisen while at Stanford
University (17), and displayed in a
heat map.
KEGG pathway enrichment analysis
To unravel the pathways involving the DEGs at each
time point, KEGG pathway enrichment analysis was performed for the
DEGs using Database for Annotation, Visualization and Integration
Discovery (DAVID), with P<0.05 as the cutoff value (18).
Inferred activity of miR-124
The regulatory activity of miRNAs is negatively
associated with the expression of their target mRNAs, which could
be measured by microarray. When the expression of the target mRNA
is downregulated, the activity of the miRNAs is enhanced, and when
the target mRNA expression is upregulated, the activity of the
corresponding miRNA is inhibited (12). On this basis, the miR-124 activity at
each time point in the study was inferred by combing microarray
expression data with miRNA target predictions, as described in a
previous study [the number of permutations for binding vectors
(k)=1,000] (12).
Construction of miRNA-target mRNA
networks
The target mRNAs of miRNA-124 at each time point
were predicted using five repositories: miRanda (http://www.microrna.org/microrna/home.do), MirTarget2
(http://nar.oxfordjournals.org/cgi/content/abstract/34/5/1646),
PicTar (http://pictar.mdc-berlin.de/), PITA
(https://genie.weizmann.ac.il/pubs/mir07/mir07_data.html)
and TargetScan (http://www.targetscan.org/vert_71/) (19–23). These
repositories provide information on the integration of miRNA target
prediction with expression files. Only the miR-124-target mRNA
pairs that were validated in at least three out of the five
databases were selected and included in the miR-124-target gene
network. Expression values of miRNA-124-target genes at different
time points were hierarchically clustered using Cluster software.
Furthermore, GO function (24) and
KEGG pathway enrichment analysis were performed for functional
annotation of the obtained target genes in the network, with
P<0.05 as the strict threshold.
Results
Screening of DEGs at different time
points
DEGs in HepG2 cells transfected with the miRNA-124
duplex and negative control were screened at 4, 8, 16, 24, 32, 72
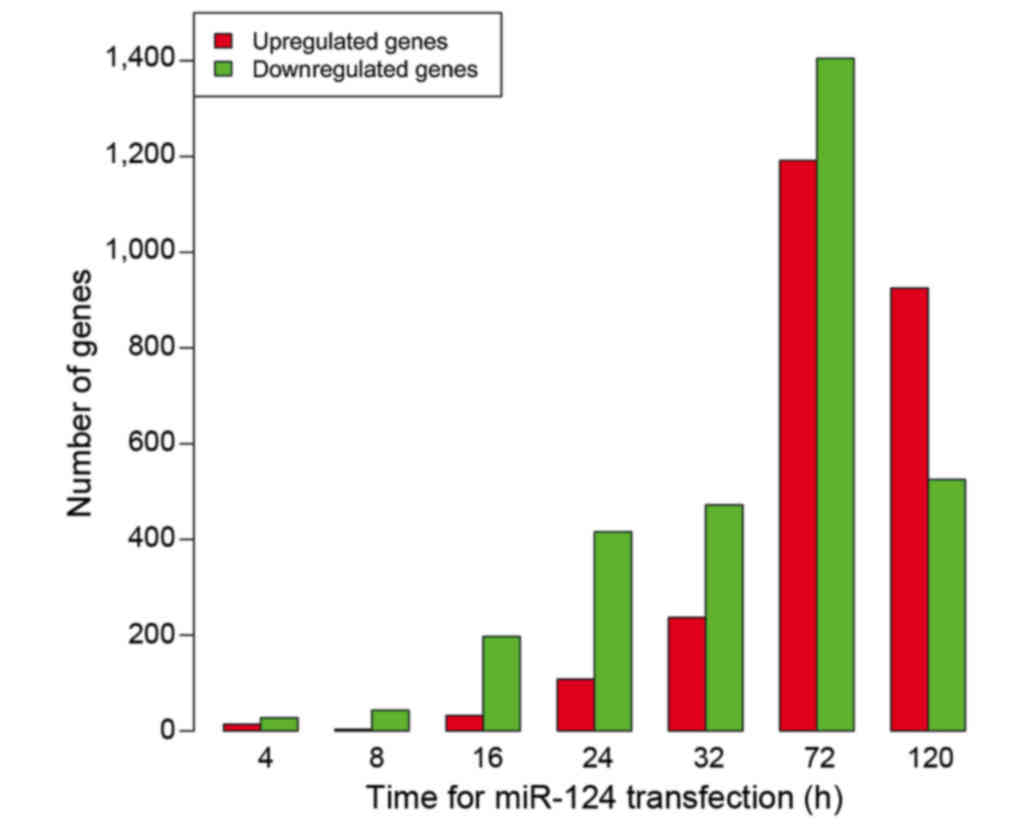
and 120 h. As shown in Fig. 1, the
total count of DEGs increased following transfection, reaching its
peak at 72 h and then decreasing. Of the DEGs, there were more
downregulated genes than upregulated genes at 4, 8, 16, 24, 32 and
72 h after transfection. The exception was 120 h, when there were
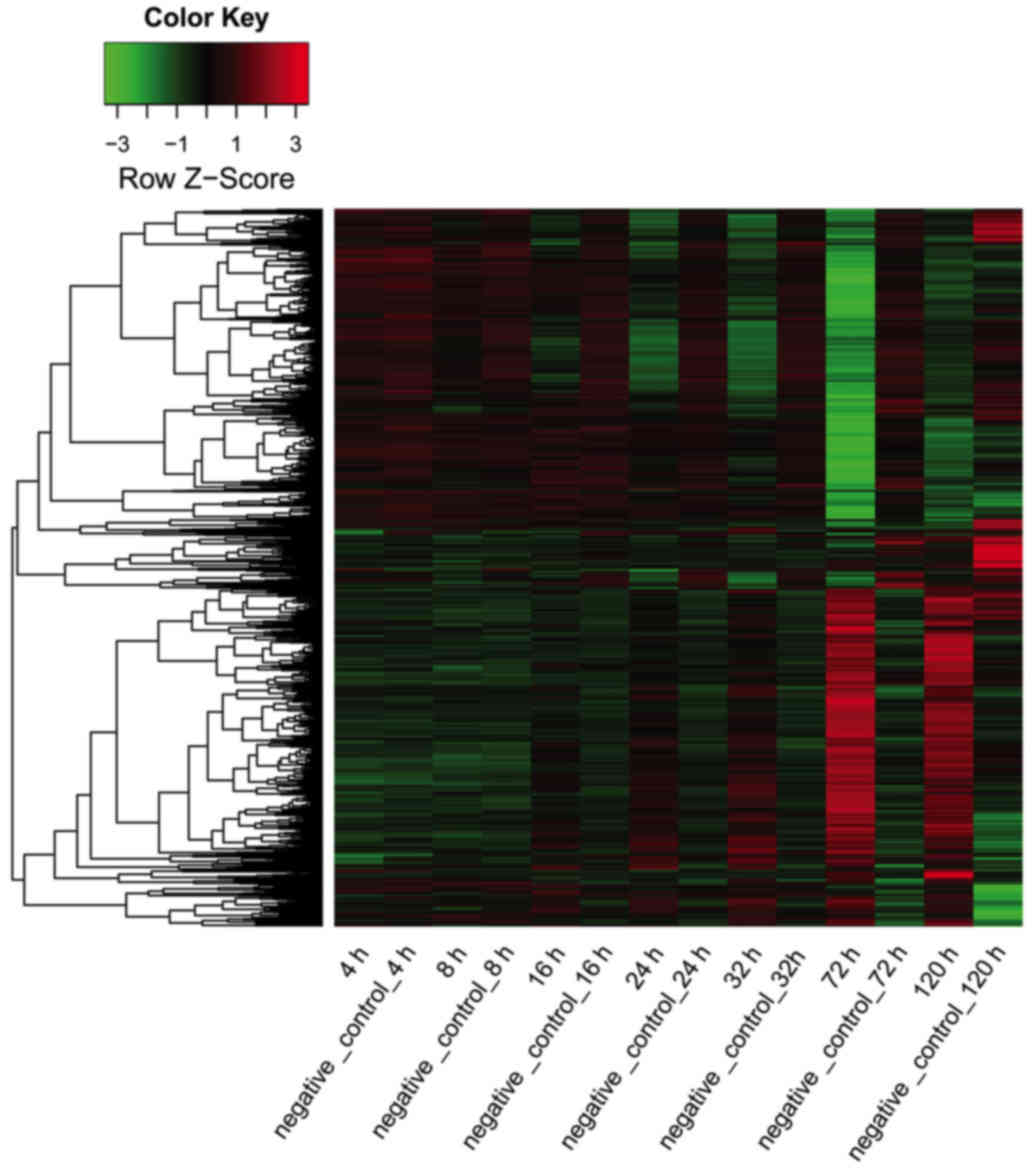
more upregulated genes than downregulated genes. The expression
data of all DEGs at seven time points is displayed in Fig. 2.
KEGG pathway enrichment analysis for
DEGs
Pathways significantly enriched for upregulated DEGs
at different time points are shown in Table I. No significant pathways were
detected at 4, 8 or 16 h; genes involved in the tumor protein p53
signaling and retinol metabolism pathways were significantly
associated with upregulated genes at 24 h after transfection.
Expression of genes involved in the p53, systemic lupus
erythematosus and mitogen-activated protein kinase (MAPK) signaling
pathways were significantly upregulated at 32 h; those involved in
axon guidance and adherens junction pathways were significantly
upregulated at 72 h; those involved in glycerolipid metabolism and
p53 signaling pathways were significantly upregulated at 120 h.
With respect to downregulated DEGs (Table II), no significant pathway was
detected at 4 or 16 h; those involved in the ribosome pathway
(hsa03010: ribosome) were significantly downregulated at 8 h; those
involved in small cell lung cancer, sphingolipid metabolism and the
cell cycle were significantly downregulated at 24 h; those involved
in small cell lung cancer, the cell cycle and pathways in cancer
were significantly downregulated at 32 h; those involved in the
cell cycle and DNA replication pathways were significantly
associated with downregulated genes at 72 h; and those involved in
the complement and coagulation cascades, cell cycle, and valine,
leucine and isoleucine degradation pathways were significantly
downregulated at 120 h.
 | Table I.KEGG pathways enriched with
upregulated differentially expressed genes at different time
points. |
Table I.
KEGG pathways enriched with
upregulated differentially expressed genes at different time
points.
| Time after
transfection, h | KEGG pathway
identity | Enriched genes,
n | P-value |
|---|
| 24 | hsa04115:p53
signaling pathway | 5 |
1.00×10−3 |
|
| hsa00830:Retinol
metabolism | 3 |
5.00×10−2 |
| 32 | hsa04115:p53
signaling pathway | 5 |
1.02×10−2 |
|
| hsa05322:Systemic
lupus erythematosus | 5 |
3.53×10−2 |
|
| hsa04010:MAPK
signaling pathway | 8 |
4.90×10−2 |
| 72 | hsa04360:Axon
guidance | 17 |
4.41×10−3 |
|
| hsa04520:Adherens
junction | 12 |
6.03×10−3 |
|
|
hsa04144:Endocytosis | 20 |
1.44×10−2 |
|
| hsa00051:Fructose
and mannose metabolism | 7 |
1.48×10−2 |
|
| hsa05130:Pathogenic
Escherichia coli infection | 9 |
2.00×10−2 |
|
| hsa04115:p53
signaling pathway | 10 |
2.00×10−2 |
|
|
hsa05410:Hypertrophic cardiomyopathy
(HCM) | 11 |
3.09×10−2 |
|
|
hsa05412:Arrhythmogenic right ventricular
cardiomyopathy | 10 |
3.81×10−2 |
|
| hsa00983:Drug
metabolism | 7 |
4.26×10−2 |
|
| hsa05414:Dilated
cardiomyopathy | 11 |
4.93×10−2 |
| 120 |
hsa00561:Glycerolipid metabolism | 8 |
4.35×10−3 |
|
| hsa04115:p53
signaling pathway | 9 |
1.30×10−2 |
|
| hsa00140:Steroid
hormone biosynthesis | 7 |
1.89×10−2 |
|
|
hsa04070:Phosphatidylinositol signaling
system | 9 |
2.09×10−2 |
|
| hsa05120:Epithelial
cell signaling in Helicobacter pylori infection | 8 |
3.77×10−2 |
|
|
hsa00564:Glycerophospholipid
metabolism | 8 |
3.77×10−2 |
|
| hsa05211:Renal cell
carcinoma | 8 |
4.32×10−2 |
|
| hsa04650:Natural
killer cell-mediated cytotoxicity | 12 |
4.49×10−2 |
|
| hsa00983:Drug
metabolism | 6 |
4.80×10−2 |
 | Table II.KEGG pathways enriched with
downregulated differentially expressed genes at different time
points. |
Table II.
KEGG pathways enriched with
downregulated differentially expressed genes at different time
points.
| Time after
transfection, h | Term | Enriched genes,
n | P-value |
|---|
| 8 |
hsa03010:Ribosome | 3 |
2.30×10−2 |
| 24 | hsa05222:Small cell
lung cancer | 9 |
7.02×10−4 |
|
|
hsa00600:Sphingolipid metabolism | 5 |
1.25×10−2 |
|
| hsa04110:Cell
cycle | 8 |
2.65×10−2 |
|
| hsa00565:Ether
lipid metabolism | 4 |
4.75×10−2 |
| 32 | hsa05222:Small cell
lung cancer | 11 |
6.88×10−5 |
|
| hsa04110:Cell
cycle | 10 |
5.95×10−3 |
|
| hsa05212:Pancreatic
cancer | 7 |
1.21×10−2 |
|
| hsa05200:Pathways
in cancer | 16 |
2.79×10−2 |
|
|
hsa00480:Glutathione metabolism | 5 |
4.37×10−2 |
| 72 | hsa04110:Cell
cycle | 46 |
7.63×10−20 |
|
| hsa03030:DNA
replication | 20 |
1.43×10−12 |
|
|
hsa04914:Progesterone-mediated oocyte
maturation | 19 |
8.17×10−5 |
|
|
hsa03040:Spliceosome | 24 |
8.48×10−5 |
|
| hsa00240:Pyrimidine
metabolism | 20 |
9.97×10−5 |
|
| hsa03430:Mismatch
repair | 9 |
2.18×10−4 |
| 120 | hsa04610:Complement
and coagulation cascades | 16 |
1.39×10−9 |
|
| hsa04110:Cell
cycle | 11 |
4.75×10−3 |
|
| hsa00280:Valine,
leucine and isoleucine degradation | 6 |
1.05×10−2 |
|
| hsa00260:Glycine,
serine and threonine metabolism | 5 |
1.41×10−2 |
|
| hsa05020:Prion
diseases | 5 |
2.13×10−2 |
|
| hsa00910:Nitrogen
metabolism | 4 |
3.20×10−2 |
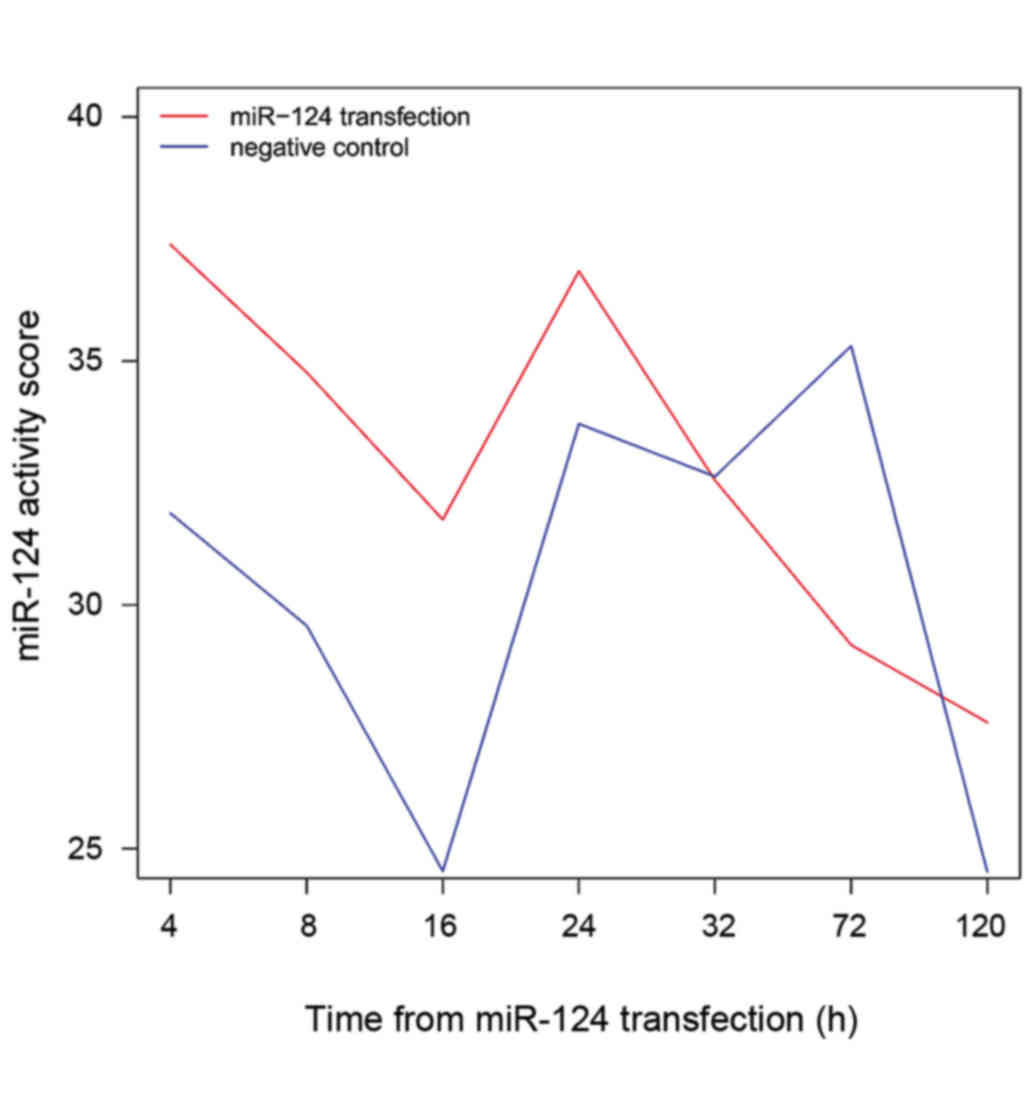
Activity changes of miRNA-124
The changes in activity of miR-124 following
transfection are shown in Fig. 3. The
activity of miR-124 varied with time after transfection: It was
highest at 4 h after transfection. The activity of miR-124 was
higher in the experimental group than that in the control group at
4, 8, 16 and 24 h after transfection. By contrast, the experimental
group exhibited decreased miR-124 activity compared with that of
the control group at 72 h after transfection. Notably, at 32 h, the
activity of miR-124 in the experimental group was similar to that
in the control group.
miR-124-target gene network
analysis
The target mRNAs of miRNA-124 at each time point
were predicted based on the aforementioned five repositories
(miRanda, MirTarget2, PicTar, PITA and TargetScan). To investigate
the association between miR-124 and its target genes, the
miR-124-target gene pairs validated in at least three out of these
five databases were used to construct miR-124-target gene networks
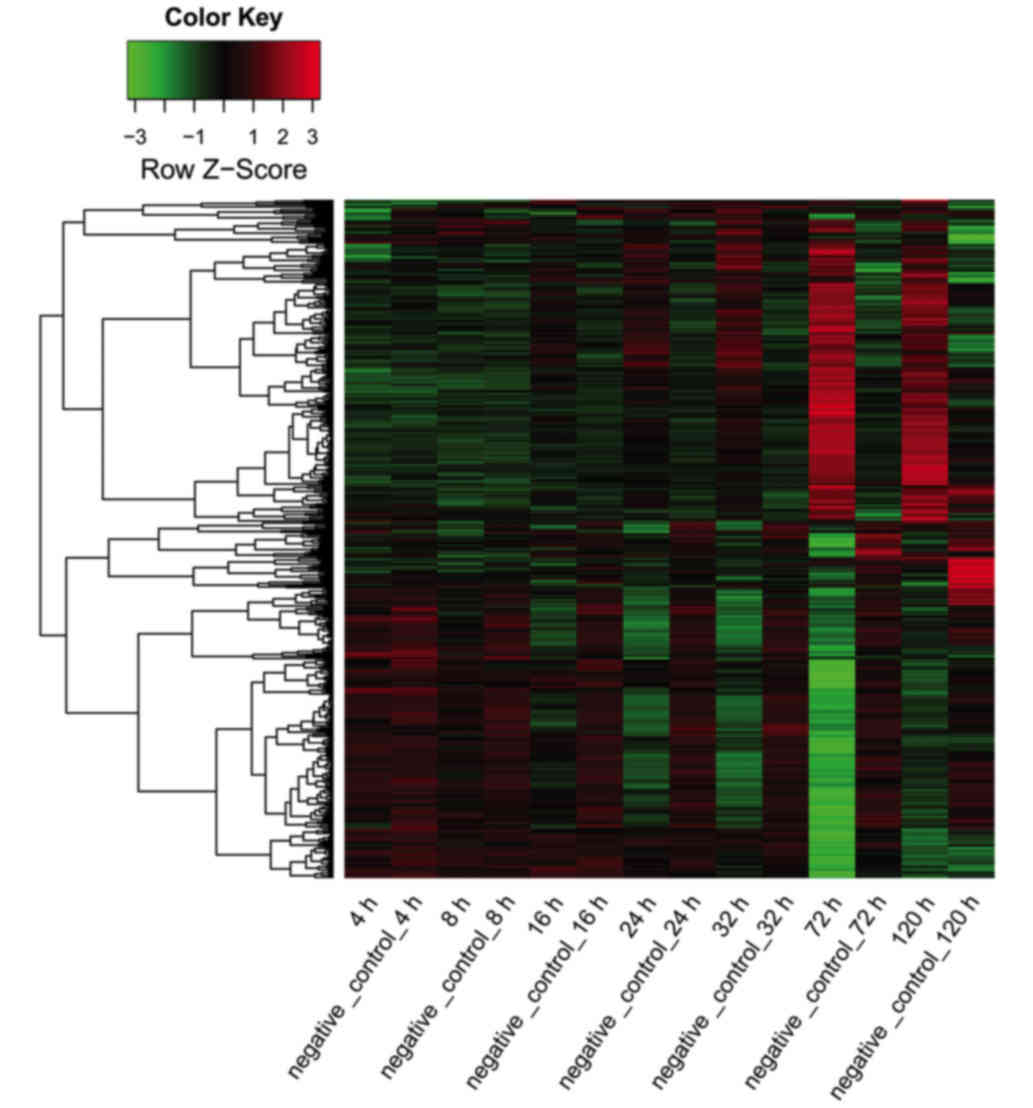
for each time point. The expression values of all the predicted
target genes at different time points are shown in a heat map
(Fig. 4).
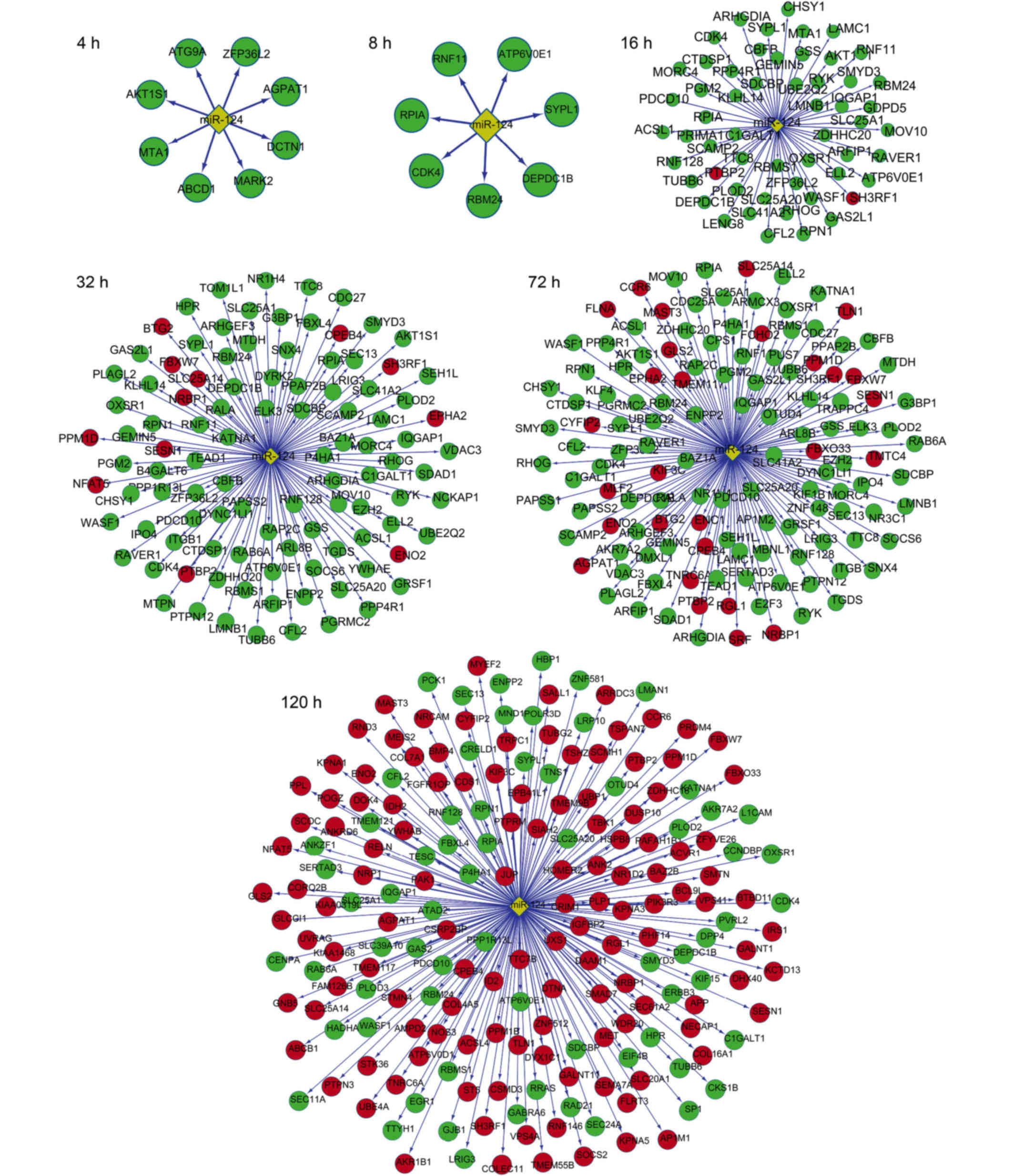
The miR-124-target gene networks are shown in
Fig. 5. At 4 and 8 h after
transfection, all miR-124-target genes were downregulated,
indicating high miR-124 activity. At 16, 24 and 32 h, a small
proportion of the target genes were upregulated, whereas the
majority of the target genes were downregulated. By contrast, more
upregulated genes were observed at 72 and 120 h when compared to
that at 16, 24 and 32 h, indicating a reduction in miR-124
activity.
Functional annotation for target genes
of miR-124
GO functional enrichment analysis was performed for
miR-124-target genes at different time points. No significant GO
term was enriched at 4 or 8 h. As shown in Table III, Ras protein signal transduction
was a significant GO term at 16 h and was enriched with syndecan
binding protein (SDCBP), Ras homolog family member G
(RHOG) and Rho GDP dissociation inhibitor-α
(ARHGDIA). At 24 h, small GTPase-mediated signal
transduction and Ras protein signal transduction were primary GO
terms, and the two were significantly enriched with SDCBP,
RHOG and ARHGDIA, Rho-guanine nucleotide exchange factor
3 (ARHGEF3), GTPase activating protein (SH3 domain) binding
protein 1 (G3BP1) and V-Ral simian leukemia viral oncogene
homolog A (RALA). Similarly, at 32 h, the two GO terms were
also enriched with the above six genes. Cell cycle and
intracellular transport were significant GO terms at 72 h. A total
of 44 genes were significantly associated with the cell cycle, such
as kinesin family member 23 (KIF23), KIF15, cell
division cycle 27 (CDC27) and CDC25A. A total of 38
genes were significantly involved in intracellular transport, such
as adaptor-related protein complex 1, µ-1 subunit (AP1M1),
adaptor-related protein complex 1, µ-2 subunit (AP1M2), KDEL
endoplasmic reticulum protein retention receptor 1 (KDELR1)
and KDELR2. At 120 h, cell motion was significantly
associated with 17 genes, such as talin 1 (TLN1) and protein
tyrosine phosphatase, receptor type, M (PTPRM), while
intracellular transport was significantly associated with 18 genes,
such as karyopherin-α (KPNA)5, KPNA3 and KPNA1.
 | Table III.GO terms enriched with target genes
of microRNA-124 at different time points. |
Table III.
GO terms enriched with target genes
of microRNA-124 at different time points.
| Time after
transfection, h | Term | Enriched genes,
n | P-value | Genes |
|---|
| 16 | GO:0007265~Ras
protein signal transduction | 3 |
4.38×10−2 | SDCBP, RHOG,
ARHGDIA |
| 24 | GO:0007264~small
GTPase mediated-signal transduction | 10 |
1.47×10−4 | ARHGEF3, RAP2C,
G3BP1, RALA, SDCBP, RAB6A, ARL8B, IQGAP1, RHOG, ARHGDIA |
|
| GO:0007265~Ras
protein signal transduction | 6 |
5.71×10−4 | ARHGEF3, G3BP1,
RALA, SDCBP, RHOG, ARHGDIA |
|
|
GO:0046907~intracellular transport | 11 |
9.37×10−3 | SLC25A20,
SLC25A14, NRBP1, SCAMP2, TOM1L1, IPO4, SDCBP, SEC13, SLC25A1,
ARFIP1, YWHAE |
|
| GO:0006928~cell
motion | 9 |
1.13×10−2 | ENPP2, WASF1,
KATNA1, SDCBP, LAMC1, PPAP2B, ITGB1, YWHAE, ARHGDIA |
|
| GO:0015031~protein
transport | 11 |
2.41×10−2 | SDAD1, SCAMP2,
TOM1L1, SEH1L, IPO4, SDCBP, SEC13, SNX4, RAB6A, ARFIP1,
YWHAE |
| 32 | GO:0007264~small
GTPase mediated-signal transduction | 11 |
1.61×10−4 | ARHGEF3, RAP2C,
G3BP1, RALA, SDCBP, RAB6A, ARL8B, IQGAP1, RHOG, ARHGDIA,
RGL1 |
|
| GO:0007265~Ras
protein signal transduction | 6 |
1.52×10−3 | ARHGEF3, G3BP1,
RALA, SDCBP, RHOG, ARHGDIA |
|
| GO:0006928~cell
motion | 11 |
4.64×10−3 | TLN1, CCR6,
ENPP2, WASF1, KATNA1, SDCBP, LAMC1, SRF, PPAP2B, ITGB1,
ARHGDIA |
|
|
GO:0046907~intracellular transport | 13 |
6.11×10−3 | SCAMP2, NRBP1,
AP1M2, ARFIP1, FLNA, SLC25A20, SLC25A14, KIF1B, IPO4, SDCBP, SEC13,
TRAPPC4, SLC25A1 |
|
|
GO:0051329~interphase of mitotic cell
cycle | 5 |
9.28×10−3 | PPM1D, KATNA1,
CDK4, ITGB1, CDC25A |
| 72 | GO:0007049~cell
cycle | 44 |
3.69×10−7 | KIF23, CKS1B,
E2F3, TUBB2B, CDC16, ITGB1, SESN1, CTNNB1, CCNE2, APP, NDE1, RAD21,
DUSP13, SEH1L, CENPA, KATNA1, RB1CC1, TFDP2, VPS4A, PPP3CA, HELLS,
CSRP2BP, NASP, KIF15, SUV39H1, SMAD3, MND1, ILF3, GAS2, MCM2, CDK4,
CDC27, CDC25A, SMC4, MCM6, PPM1G, PA2G4, PPM1D, FANCD2, PLK1,
GAS2L1, SETD8, SIAH2, ACVR1 |
|
|
GO:0046907~intracellular transport | 38 |
1.79×10−6 | NUP98, AP1M1,
AP1M2, SEC24A, NRBP1, EIF5A, BCL2L1, KLC2, LMAN1, SLC25A20, AP1S1,
APP, AP2B1, NDE1, CSE1L, ZFYVE9, VPS4A, SLC25A1, PPP3CA, SAR1B,
KDELR1, XPOT, GABARAPL2, KDELR2, SCAMP2, VPS41, ARFIP1, SLC25A12,
SLC25A14, TOM1L1, IPO4, TRAPPC4, SEC13, SDCBP, KPNA5, KPNA3,
TRAPPC1, SSR2 |
|
| GO:0022402~cell
cycle process | 33 |
8.30×10−6 | KIF23, TUBB2B,
CDC16, ITGB1, SESN1, CTNNB1, APP, NDE1, RAD21, DUSP13, SEH1L,
CENPA, KATNA1, PPP3CA, HELLS, CSRP2BP, KIF15, SMAD3, MND1, ILF3,
GAS2, CDK4, CDC27, CDC25A, SMC4, PPM1G, PA2G4, PPM1D, FANCD2, PLK1,
GAS2L1, SETD8, ACVR1 |
|
| GO:0048193~Golgi
vesicle transport | 13 |
8.58×10−5 | GABARAPL2,
AP1M1, SCAMP2, NRBP1, SEC24A, AP1M2, VPS41, LMAN1, AP1S1, SEC13,
TRAPPC4, TRAPPC1, SAR1B |
|
| GO:0070727~cellular
macromolecule localization | 24 |
2.12×10−4 | KDELR2, AP1M1,
NUP98, SEC24A, AP1M2, EIF5A, VPS41, ARFIP1, CASC3, CTNNB1, AP1S1,
AP2B1, CSE1L, TOM1L1, IPO4, ZFYVE9, SEC13, SDCBP, KPNA5, PPP3CA,
SAR1B, KPNA3, KDELR1, SSR2 |
| 120 | GO:0006928~cell
motion | 17 |
1.86×10−4 | TLN1, NRP1,
PTPRM, ENPP2, WASF1, MET, L1CAM, NRCAM, APP, CCR6, TNS1, KATNA1,
SDCBP, NOS3, PAFAH1B1, RELN, ACVR1 |
|
|
GO:0046907~intracellular transport | 18 |
2.27×10−3 | AP1M1, SEC24A,
NRBP1, YWHAB, VPS41, LMAN1, SLC25A20, APP, SLC25A14, VPS4A, SDCBP,
SEC13, SLC25A1, PAFAH1B1, KPNA5, KPNA3, KPNA1, SEC61A2 |
|
| GO:0032355~response
to estradiol stimulus | 5 |
3.94×10−3 | BMP4, SOCS2,
ENO2, NOS3, IGFBP2 |
|
| GO:0016477~cell
migration | 10 |
6.03×10−3 | NRCAM, TNS1,
NRP1, KATNA1, MET, SDCBP, NOS3, PAFAH1B1, RELN, ACVR1 |
|
|
GO:0006796~phosphate metabolic
process | 22 |
6.12×10−3 | ATP6V0E1, NRBP1,
PTPRM, PTPN3, SLC20A1, ENPP2, ERBB3, SMAD7, TBK1, STK36, MET,
DUSP10, OXSR1, PPM1B, CDK4, MAST3, PPM1D, APP, RELN, PAK1,
ATP6V0D1, ACVR1 |
The potential signaling pathways and target genes
that may be involved in these signaling pathways were also analyzed
(Table IV). No significant pathways
were enriched with target genes at 4, 8 or 16 h. Regulation of the
actin cytoskeleton pathway was a significant pathway at 24 h, and
was enriched with several target genes, including cofilin 2
(CFL2) and WAS protein family, member 1 (WASF1). The
target genes were predominantly associated with small cell lung
cancer and ether lipid metabolism pathways at 32 h. At 72 h, the
D-glutamine and D-glutamate metabolism pathway was significantly
enriched with glutamate dehydrogenase 1 (GLUD1) and
GLUD2. At 120 h, axon guidance, adherens junction and the
transforming growth factor β signaling pathway were enriched. The
axon guidance pathway was significant at 72 and 120 h, and was
enriched with a number of target genes, such as neuropilin 1
(NRP1), MET proto-oncogene, receptor tyrosine kinase
(MET) and semaphorin 7A, GPI membrane anchor
(SEMA7A).
 | Table IV.Kyoto Encyclopedia of Genes and
Genomes pathways enriched with target genes of microRNA-124 at
different time points. |
Table IV.
Kyoto Encyclopedia of Genes and
Genomes pathways enriched with target genes of microRNA-124 at
different time points.
| Time after
transfection, h | Term | Enriched genes,
n | P-value | Genes |
|---|
| 24 | hsa04810:Regulation
of actin cytoskeleton | 5 |
4.87×10−2 | CFL2, WASF1,
ITGB1, IQGAP1, NCKAP1 |
| 32 | hsa05222:Small cell
lung cancer | 4 |
3.14×10−2 | E2F3, LAMC1,
CDK4, ITGB1 |
|
| hsa00565:Ether
lipid metabolism | 3 |
3.34×10−2 | ENPP2, PPAP2B,
AGPAT1 |
| 72 | hsa04360:Axon
guidance | 15 |
6.34×10−6 | NRP1, MET,
DPYSL2, ITGB1, EPHA2, SEMA6A, ROBO1, SEMA7A, SEMA3F, CFL2, NFAT5,
PAK1, PPP3CA, NFATC3, RASA1 |
|
| hsa04110:Cell
cycle | 14 |
2.20×10−5 | E2F3, E2F5,
SMAD3, MCM2, CDC16, CDK4, CDC27, CDC25A, MCM5, MCM6, CCNE2, RAD21,
PLK1, TFDP2 |
|
| hsa05212:Pancreatic
cancer | 9 |
5.74×10−4 | E2F3, RALBP1,
RELA, ARAF, PIK3CD, SMAD3, RALA, BCL2L1, CDK4 |
|
| hsa05222:Small cell
lung cancer | 9 |
1.60×10−3 | CCNE2, CKS1B,
E2F3, RELA, PIK3CD, LAMC1, BCL2L1, CDK4, ITGB1 |
|
|
hsa00471:D-Glutamine and D-glutamate
metabolism | 3 |
4.05×10−3 | GLS2, GLUD2,
GLUD1 |
| 120 | hsa04360:Axon
guidance | 7 |
1.29×10−2 | NRP1, CFL2,
SEMA7A, MET, NFAT5, L1CAM, PAK1 |
|
| hsa04520:Adherens
junction | 5 |
2.86×10−2 | PTPRM, WASF1,
MET, PVRL2, IQGAP1 |
|
| hsa04350:TGF-β
signaling pathway | 5 |
4.21×10−2 | BMP4, ID2, SP1,
SMAD7, ACVR1 |
Discussion
miRNAs participate in a variety of cell activities
via suppression of the expression of their target genes (25). The molecular mechanism by which
miR-124 results in suppression of hepatoblastoma is not entirely
clear. The present study revealed that the total number of DEGs in
hepatoblastoma cells increased with time following miR-124
transfection, reached a peak at 72 h and then decreased, compared
with the findings in cells treated with a negative control. miR-124
overexpression led to more downregulated genes than upregulated
genes at 4, 8, 16, 24, 32 and 72 h, which was partly consistent
with the fact that miR-124 inhibits the expression of its target
genes (26). However, a proportion of
DEGs were upregulated following miR-124 transfection. An
explanation for this observation is that the upregulated DEGs may
be the result of downregulation of DEGs by miR-124 overexpression.
miRNAs may regulate the expression of other miRNAs (12,27), which
could therefore strengthen or inhibit the activity of other
endogenous miRNAs, thus contributing to differential expression of
their target genes. These inhibited miRNAs may cause upregulation
of their target genes.
Small GTPases are intracellular G proteins and
hydrolase enzymes that hydrolyze guanosine triphosphate. They serve
a key role in diverse cell activities, including cell growth,
differentiation, movement and lipid vesicle transport (28,29). Ras
protein family members belong to the small GTPase family. Ras is an
intracellular anti-apoptotic protein that is closely associated
with the phosphoinositide 3-kinase/AKT and MAPK signaling pathways,
regulating various activities, including cell apoptosis and growth
(30,31). Several lines of evidence have
suggested that small GTPases have a regulatory role in
hepatoblastoma (32,33). Consistent with this, the present study
revealed that small GTPase-mediated signal transduction and Ras
protein signal transduction were significant GO categories,
enriched with SDCBP, RHOG and ARHGDIA in
hepatoblastoma at 24 and 32 h. SDCBP encodes syntenin-1,
which is a tandem PDZ domain-containing protein involved in
trafficking, signaling and cancer metastasis (34). Syntenin-1 is implicated in regulating
colon cancer cell migration (35).
However, the role of SDCBP in hepatoblastoma has not been
previously reported. RHOG encodes the Ras homology
growth-related (RhoG) protein, which is a small GTP-binding protein
and a member of the Rac subgroup of the Rho family (36). The Rac signaling pathways serve an
essential role in the motility of hepatocellular carcinoma cells
(37). ARHGDIA encodes Rho
GDP-dissociation inhibitor 1 protein, which participates in
regulating Rho activity. The absence of ARHGDIA has been
reported to enhance the invasion and metastasis of hepatocellular
carcinoma cells (37). However, the
role of these genes in hepatoblastoma has not been reported thus
far. Thus, the present study speculated that miR-124 overexpression
may lead to the inhibition of small GTPase-mediated signal
transduction and Ras protein signal transduction by regulating
SDCBP, RHOG and ARHGDIA, thereby repressing
hepatoblastoma development.
The present study unraveled that regulation of the
actin cytoskeleton pathway was significantly enriched with several
target genes such as CFL2 at 24 h after transfection.
Regulation of the actin cytoskeleton pathway is involved in cancer
cell migration, tumor invasion and metastasis (38). CFL2 is an intracellular
actin-modulating protein that regulates actin-filament dynamics
(39); it indicates that miR-124
overexpression may inhibit cancer cell migration, tumor invasion
and metastasis by regulating actin cytoskeleton pathway genes such
as CFL2. Furthermore, the present study revealed that the
D-glutamine and D-glutamate metabolism pathways were enriched
(GLUD1 and GLUD2) at 72 h. The D-glutamine and
D-glutamate metabolic pathways are critical components of protein
synthesis and cellular energy production (40,41).
GLUD1 and GLUD2 are two mitochondrial matrix enzymes
that serve a key role in glutamate metabolism and energy
homeostasis (42). The findings of
the present study revealed that miR-124 transfection may affect
glutamate metabolism and energy homeostasis.
Notably, the expression of NRP1, SEMA7A and
MET, which are involved in the axon guidance signaling
pathway, was significantly increased at 72 and 120 h. An aberrant
axon guidance pathway is involved in pancreatic carcinogenesis
(43). Nevertheless, the role of the
axon guidance pathway in hepatoblastoma has not been clearly
characterized. Neuropilin-1, encoded by NRP1, is a
co-receptor member for members of the semaphorin family, and is
involved in several cellular processes, including angiogenesis and
cell survival (44). A previous study
revealed that the expression of NRP1 is higher in HepG2 cell
lines than that in a human normal hepatic cell line, indicating the
effect of NRP1 on hepatoblastoma (45). SEMA7A, which is encoded by
SEMA7A, is a membrane-bound semaphorin and promotes axon
outgrowth (46). MET encodes
hepatocyte growth factor receptor (HGFR), which has tyrosine kinase
activity and binds to HGF. It has been reported that invasion of
hepatocellular carcinoma cells is dependent on HGF (47). Furthermore MET/HGF inhibitors have
been increasingly recognized as anticancer therapies for various
types of cancer, including hepatoma (48). These observations indicate that axon
guidance pathway genes may be abnormally regulated due to miR-124
overexpression, thereby suppressing hepatoblastoma cell growth and
invasion. The present study indicates the presence of an
as-yet-undescribed role of the axon guidance pathway in
hepatoblastoma.
The current study demonstrated that miR-124
overexpression may perturb small GTPase-mediated signal
transduction and Ras protein signal transduction, and abnormally
affect the regulation of the actin cytoskeleton, D-glutamine and
D-glutamate metabolic and axon guidance pathways in hepatoblastoma.
SDCBP, RHOG, ARHGDIA, NRP1, SEMA7A and MET may be
promising molecular targets for therapies against hepatoblastoma.
These pathways are not specific to hepatoblastoma, but may also be
involved in other types of cancer. These findings provide useful
information concerning the molecular mechanisms that underlie the
tumor-suppressive role of miR-124 in hepatoblastoma. Further
experimental studies are warranted to validate the findings of the
present study.
References
|
1
|
Stewart BW and Wild CP: World cancer
report 2014. World Health Organization. 2014.
|
|
2
|
Ringe B, Pichlmayr R, Wittekind C and
Tusch G: Surgical treatment of hepatocellular carcinoma: Experience
with liver resection and transplantation in 198 patients. World J
Surg. 15:270–285. 1991. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Maluccio M and Covey A: Recent progress in
understanding, diagnosing, and treating hepatocellular carcinoma.
CA Cancer J Clin. 62:394–399. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Bruix J and Sherman M: American
Association for the Study of Liver Diseases: Management of
hepatocellular carcinoma: An update. Hepatology. 53:1020–1022.
2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Meyers RL, Tiao G, de Ville de Goyet J,
Superina R and Aronson DC: Hepatoblastoma state of the art:
Pre-treatment extent of disease, surgical resection guidelines and
the role of liver transplantation. Curr Opin Pediatr. 26:29–36.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Osada H and Takahashi T: MicroRNAs in
biological processes and carcinogenesis. Carcinogenesis. 28:2–12.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Shi XB, Xue L, Ma AH, Tepper CG,
Gandour-Edwards R, Kung HJ and deVere White RW: Tumor suppressive
miR-124 targets androgen receptor and inhibits proliferation of
prostate cancer cells. Oncogene. 32:4130–4138. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zheng H, Song F, Zhang L, Yang D, Ji P,
Wang Y, Almeida M, Calin GA, Hao X, Wei Q, et al: Genetic variants
at the miR-124 binding site on the cytoskeleton-organizing IQGAP1
gene confer differential predisposition to breast cancer. Int J
Oncol. 38:1153–1161. 2011.PubMed/NCBI
|
|
9
|
Zheng F, Liao YJ, Cai MY, Liu YH, Liu TH,
Chen SP, Bian XW, Guan XY, Lin MC, Zeng YX, et al: The putative
tumour suppressor microRNA-124 modulates hepatocellular carcinoma
cell aggressiveness by repressing ROCK2 and EZH2. Gut. 61:278–289.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zeng B, Li Z, Chen R, Guo N, Zhou J, Zhou
Q, Lin Q, Cheng D, Liao Q, Zheng L and Gong Y: Epigenetic
regulation of miR-124 by hepatitis C virus core protein promotes
migration and invasion of intrahepatic cholangiocarcinoma cells by
targeting SMYD3. FEBS Lett. 586:3271–3278. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Krützfeldt J, Rajewsky N, Braich R, Rajeev
KG, Tuschl T, Manoharan M and Stoffel M: Silencing of microRNAs in
vivo with ‘antagomirs’. Nature. 438:685–689. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cheng C and Li LM: Inferring microRNA
activities by combining gene expression with microRNA target
prediction. PLoS One. 3:e19892008. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Lim LP, Lau NC, Garrett-Engele P, Grimson
A, Schelter JM, Castle J, Bartel DP, Linsley PS and Johnson JM:
Microarray analysis shows that some microRNAs downregulate large
numbers of target mRNAs. Nature. 433:769–773. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang X and Wang X: Systematic
identification of microRNA functions by combining target prediction
and expression profiling. Nucleic Acids Res. 34:1646–1652. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Irizarry RA, Hobbs B, Collin F,
Beazer-Barclay YD, Antonellis KJ, Scherf U and Speed TP:
Exploration, normalization and summaries of high density
oligonucleotide array probe level data. Biostatistics. 4:249–264.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bernhardt V, Hotchkiss MT, Garcia-Reyero
N, Escalon BL, Denslow N and Davenport PW: Tracheal occlusion
conditioning in conscious rats modulates gene expression profile of
medial thalamus. Front Physiol. 2:242011. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Shannon W, Culverhouse R and Duncan J:
Analyzing microarray data using cluster analysis. Pharmacogenomics.
4:41–52. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Lewis BP, Shih IH, Jones-Rhoades MW,
Bartel DP and Burge CB: Prediction of mammalian microRNA targets.
Cell. 115:787–798. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kertesz M, Iovino N, Unnerstall U, Gaul U
and Segal E: The role of site accessibility in microRNA target
recognition. Nat Genet. 39:1278–1284. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lall S, Grün D, Krek A, Chen K, Wang YL,
Dewey CN, Sood P, Colombo T, Bray N, Macmenamin P, et al: A
genome-wide map of conserved microRNA targets in C. elegans. Curr
Biol. 16:460–471. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang X: miRDB: A microRNA target
prediction and functional annotation database with a wiki
interface. RNA. 14:1012–1017. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Betel D, Wilson M, Gabow A, Marks DS and
Sander C: The microRNA.org resource: targets and expression.
Nucleic Acids Res. 36:D149–D153. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Harris MA, Clark J, Ireland A, Lomax J,
Ashburner M, Foulger R, Eilbeck K, Lewis S, Marshall B, Mungall C,
et al: The gene ontology (GO) database and informatics resource.
Nucleic Acids Res. 32:(Database Issue). D258–D261. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Weis SM and Cheresh DA: Tumor
angiogenesis: Molecular pathways and therapeutic targets. Nat Med.
17:1359–1370. 2011. View
Article : Google Scholar : PubMed/NCBI
|
|
26
|
Pillai RS: MicroRNA function: Multiple
mechanisms for a tiny RNA? RNA. 11:1753–1761. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hobert O: Common logic of transcription
factor and microRNA action. Trends Biochem Sci. 29:462–468. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Etienne-Manneville S and Hall A: Rho
GTPases in cell biology. Nature. 420:629–635. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Jaffe AB and Hall A: Rho GTPases:
Biochemistry and biology. Annu Rev Cell Dev Biol. 21:247–269. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yoshimura T, Arimura N, Kawano Y, Kawabata
S, Wang S and Kaibuchi K: Ras regulates neuronal polarity via the
PI3-kinase/Akt/GSK-3beta/CRMP-2 pathway. Biochem Biophys Res
Commun. 340:62–68. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Chang L and Karin M: Mammalian MAP kinase
signalling cascades. Nature. 410:37–40. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wong CM, Yam JW, Ching YP, Yau TO, Leung
TH, Jin DY and Ng IO: Rho GTPase-activating protein deleted in
liver cancer suppresses cell proliferation and invasion in
hepatocellular carcinoma. Cancer Res. 65:8861–8868. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Calvisi DF, Ladu S, Conner EA, Seo D,
Hsieh JT, Factor VM and Thorgeirsson SS: Inactivation of Ras
GTPase-activating proteins promotes unrestrained activity of
wild-type Ras in human liver cancer. J Hepatol. 54:311–319. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Beekman JM and Coffer PJ: The ins and outs
of syntenin, a multifunctional intracellular adaptor protein. J
Cell Sci. 121:1349–1355. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Lee H, Kim Y, Choi Y, Choi S, Hong E and
Oh ES: Syndecan-2 cytoplasmic domain regulates colon cancer cell
migration via interaction with syntenin-1. Biochem Biophys Res
Commun. 409:148–153. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Bustelo XR, Sauzeau V and Berenjeno IM:
GTP-binding proteins of the Rho/Rac family: Regulation, effectors
and functions in vivo. Bioessays. 29:356–370. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Liang L, Li Q, Huang LY, Li DW, Wang YW,
Li XX and Cai SJ: Loss of ARHGDIA expression is associated with
poor prognosis in HCC and promotes invasion and metastasis of HCC
cells. Int J Oncol. 45:659–666. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yamaguchi H and Condeelis J: Regulation of
the actin cytoskeleton in cancer cell migration and invasion.
Biochim Biophys Acta. 1773:642–652. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Maciver SK and Hussey PJ: The ADF/cofilin
family: Actin-remodeling proteins. Genome Biol. 3:reviews30072002.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Aledo JC: Glutamine breakdown in rapidly
dividing cells: Waste or investment? Bioessays. 26:778–785. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Le Bacquer O, Laboisse C and Darmaun D:
Glutamine preserves protein synthesis and paracellular permeability
in Caco-2 cells submitted to ‘luminal fasting’. Am J Physiol
Gastrointest Liver Physiol. 285:G128–G136. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Spanaki C, Kotzamani D and Plaitakis A:
Widening spectrum of cellular and subcellular expression of human
Glud1 and Glud2 glutamate dehydrogenases suggests novel functions.
Neurochem Res. 42:92–107. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Biankin AV, Waddell N, Kassahn KS, Gingras
MC, Muthuswamy LB, Johns AL, Miller DK, Wilson PJ, Patch AM, Wu J,
et al: Pancreatic cancer genomes reveal aberrations in axon
guidance pathway genes. Nature. 491:399–405. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Gu C, Rodriguez ER, Reimert DV, Shu T,
Fritzsch B, Richards LJ, Kolodkin AL and Ginty DD: Neuropilin-1
conveys semaphorin and VEGF signaling during neural and
cardiovascular development. Dev Cell. 5:45–57. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Xu J and Xia J: NRP-1 silencing suppresses
hepatocellular carcinoma cell growth in vitro and in vivo. Exp Ther
Med. 5:150–154. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Pasterkamp RJ, Peschon JJ, Spriggs MK and
Kolodkin AL: Semaphorin 7A promotes axon outgrowth through
integrins and MAPKs. Nature. 424:398–405. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Monvoisin A, Neaud V, Lédinghen V,
Dubuisson L, Balabaud C, Bioulac-Sage P, Desmoulière A and
Rosenbaum J: Direct evidence that hepatocyte growth factor-induced
invasion of hepatocellular carcinoma cells is mediated by
urokinase. J Hepatol. 30:511–518. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Scagliotti GV, Novello S and von Pawel J:
The emerging role of MET/HGF inhibitors in oncology. Cancer Treat
Rev. 39:793–801. 2013. View Article : Google Scholar : PubMed/NCBI
|