Introduction
Pancreatic cancer (PC) is one of the most aggressive
malignancies in the world with an extremely low 5-year survival
rate (1,2). To date, detection and diagnosis of PC is
difficult on account of the lack of diagnostic markers. When the
patients were diagnosed with PC, most of them developed an
aggressive form of disease which limits the potential for
therapeutic intervention (3).
Although surgery is the primary way of treatment, pharmacological
treatments are also an important adjuvant to surgery. During the
past decades, studies have tried to understand the undelying
molecular and signaling mechanisms that regulate the development of
PC so that better therapy strategies may be developed (4–6).
At this stage, plenty of researches have
demonstrated that, during the the occurrence and development of PC,
several genetic and epigenetic changes have taken place, such as
DNA methylation, microRNA (miRNAs) expression profile and so on
(7–9).
However, minimal improvements have been made in the prevention and
treatment of PC due to the lack of a fundamental target. Therefore,
there is an urgent need to develope a new associated factors or
novel therapeutic targets in PC.
The human ether-a-go-go-related potassium channel
(hERG), also known as KCNH2 or Kv11.1, plays an important role on
the terminal repolarization in human ventricular myocytes (10). Several clinically successful drugs
have a tendency to inhibit hERG, leading to the risk of adverse
drug reactions with QT interval prolongation syndrome or sudden
death during use (11,12). In recent years, increasing evidence
has demonstrated that plasma membrane ether-a-go-go-related
potassium channel 1 (hERG1) potassium (K+) channels are required
for cell proliferation and have essential roles in many crucial
cellular events such as apoptosis, migration and invasion (13–15).
However, research examining the function of hERG1 in human PC are
rare and the underlying mechanisms regulating hERG1 expression in
PC progression remian largely unknown.
miRNAs are highly conserved small noncoding RNAs
that recognize and bind to the 3′UTR of targeted mRNAs resulting in
translational repression. More than half of the known miRNAs have
been shown to participate in human tumorigenesis and/or metastasis
by directly targeting oncogenes or tumor suppressor genes (16,17).
Numerous profiling studies over the past decade have shown that the
miRNAs play pivotal roles in multiply human cancers, including PC
(18–20), which thereby being potential cancer
biomarkers in PC. Based on the previous research about the miRNA
expression profile in PC (21) and
the result that we profiled Targetscan and miRBase databases to
explore miRNAs which has putative binding sites with hERG1 gene
(KCNH2), we selected miR-493 as a potential regulator in PC
progression.
In the present study, we provide evidences that
miR-493 was essential in the regulation of proliferation and
invasion of PC cells, and the underlying molecular mechanism
potentailly lied in the regulation of hERG channel expression,
which shed a new light in understanding of how miRNAs act in
tumorigenesis and provide novel therapeutic strategies in PC
treatment.
Materials and methods
Tissue samples
Tissue samples were collected from patients
diagnosed with PC and underwent surgery at The Third Affiliated
Hospital of Harbin Medical University (Harbin, China). The samples
were frozened and stored at −80°C until total RNA or protein was
extracted. All patients have signed the consent to the research.
The Research Ethics Committee of Harbin Medical University (Harbin,
China) approved the study.
Cell culture and transfection
Human PC cell lines PANC-1 and CFPAC-1 and the human
pancreatic cell line (HPDE) were purchased from the Chinese Cell
Bank of the Chinese Academy of Sciences (Shanghai, China). Cells
were maintained in DMEM (Hyclone; GE Healthcare Life Sciences,
Logan, UT, USA) supplemented with 10% (v/v) fetal bovine serum
(FBS; Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA) at
37°C in a humidified atmosphere of 5% CO2. miR-493 mimic
and antisense oligodeoxyribonucleotide (AMO) were obtained from
GenePharma (Shanghai, China). All the cell transfections were
performed according to the kit instruction (X-tremeGENE; Roche
Diagnostics, Indianapolis, IN, USA). Cells were studied within 8 h
of harvest.
Luciferase assay
As miR-493 was identified as the potent regulator of
HERG1 levels by Targetscan database, next it was examined whether
HERG1 is a direct target of miR-493. Briefly, the miR-493-binding
site in the 3′-UTR region of HERG1 (wild or mutant type) was cloned
into the downstream of the firefly luciferase gene in a
pGL3-promoter vector (Promega, Madison, WI, USA). Next, the plasmid
DNA (wild-type or mutant pGL3-hERG1-3′ UTR; 500 ng/ml) and miR-297
mimics (100 nM) were transfected into cells using Lipofectamine
2000 for 48 h. Firefly luciferase activity was analyzed by
Dual-Luciferase Reporter Assay System according to the
manufacturer's protocol (Promega) and was normalized to the
Renilla luciferase activity.
Cell proliferation assay
The cell proliferation was determined by MTT assay.
Briefly, cells were seeded in 96-well plates at 1×105
cells/well and maintained for 24 h to allow cell adhesion.
Subsequently, the cells were incubated with 30 µl of MTT (5 mg/ml)
for 4 h. Afterwards, the foramazan crystals were dissolved in 100
µl DMSO, and the absorbance was measured at 570 nm by using a
microplate reader (Olympus, USA). The viability of treated samples
was assessed by comparison with negative control. Each experiment
was tested in triplicate.
In vitro cell invasion assays
The invasion assay was performed with 24-well plates
coated with 100 µl of Corning Matrigel Basement Membrane Matrix (BD
Biosciences, San Diego, CA, USA). Cells were seeded on to the
Matrigel coated wells (3×104 cells/cm2).
After 24 h, the cells migrated to the bottom surface of the
membrane were fixed in 4% paraformaldehyde in PBS. Once fixed, the
cells were stained with crystal violet for 10 min at room
temperature and then the pictures were captured by using a
fluorescence microscope (Nikon Corp., Tokyo, Japan). Data were
expressed as the average number of cells per insert.
RNA extraction and real-time
quantitative PCR
Total RNA from mouse tissues was extracted using
TRIzol reagent (Invitrogen, the Netherlands). The concentration of
RNA was detected by a NanoDrop Spectrophotometer (NanoDrop
Technologies; Thermo Fisher Scientific, Inc., Wilmington, DE, USA).
Synthesis of hERG1 cDNA was carried out using a universal reverse
transcription kit (Takara, Dalian, China), while a poly(A) tail was
added to the miRNAs for miR-493 cDNA synthesis. Real-time PCR was
performed using the SYBR-Green PCR Master Mix (Roche) and the ABI
7500 Real-Time PCR System (Life Technologies, Grand Island, NY,
USA). PCR amplification was performed in a total volume of 20 µl
with 2 µl cDNA, 6 µl DEPC, 10 µl SYBR-Green Master Mix, 1 µl
forward primer and 1 µl reverse primer. PCR conditions were as
follows: 40 cycles of 95°C for 15 sec, 60°C for 15 sec, and 72°C
for 45 sec. After circle reaction, the threshold cycle (Ct) was
determined, and relative miR-493 and hERG1 were calculated based on
the Ct values and normalized by U6 or GAPDH level respectively in
each sample.
Western blot analysis
For the western blot analysis, the total protein of
the tissues and cells were harvested with radioimmunoprecipitation
assay buffer (RIPA) containing 1% protease inhibitor (Sigma, St.
Louis, MO, USA) and the protein concentration was determined by BCA
Protein Assay kit (Thermo Scientific, USA) Samples were loaded to
8% SDS-PAGE gels and transferred onto PVDF (Pall Life Sciences,
Port Washington, NY, USA) membrane by electrophoresis. After 2 h
blocking with 5% nonfatmilk in PBS, membranes were incubated with
hERG1 antibody (1:1,000 dilution) or monoclonal GAPDH antibody
(1:3,000) overnight at 4°C with gentle shaking. Afterwards,
secondary antibody labelled with fluorochrome dyes (Alexa Fluor
800; LI-COR Biosciences, Lincoln, NE, USA) was used. Finally,
immunoreactivity was detected with the Odyssey fluorescent scanning
system (LI-COR Biosciences) and the band densities were quantified
by densitometry, using Scion Image software (Scion, Frederick, MD,
USA). Data were normalized to GAPDH expression levels.
Statistical analysis
Data are expressed as mean ± standard error of mean
(mean ± SEM) in triplicate experiments and analyzed with SPSS 13.0
software. A one-way analysis of variance or Student's t test was
used to determine the significance of differences between control
and test groups. P≤0.05 was considered to indicate a statistically
significant difference (two-tailed).
Results
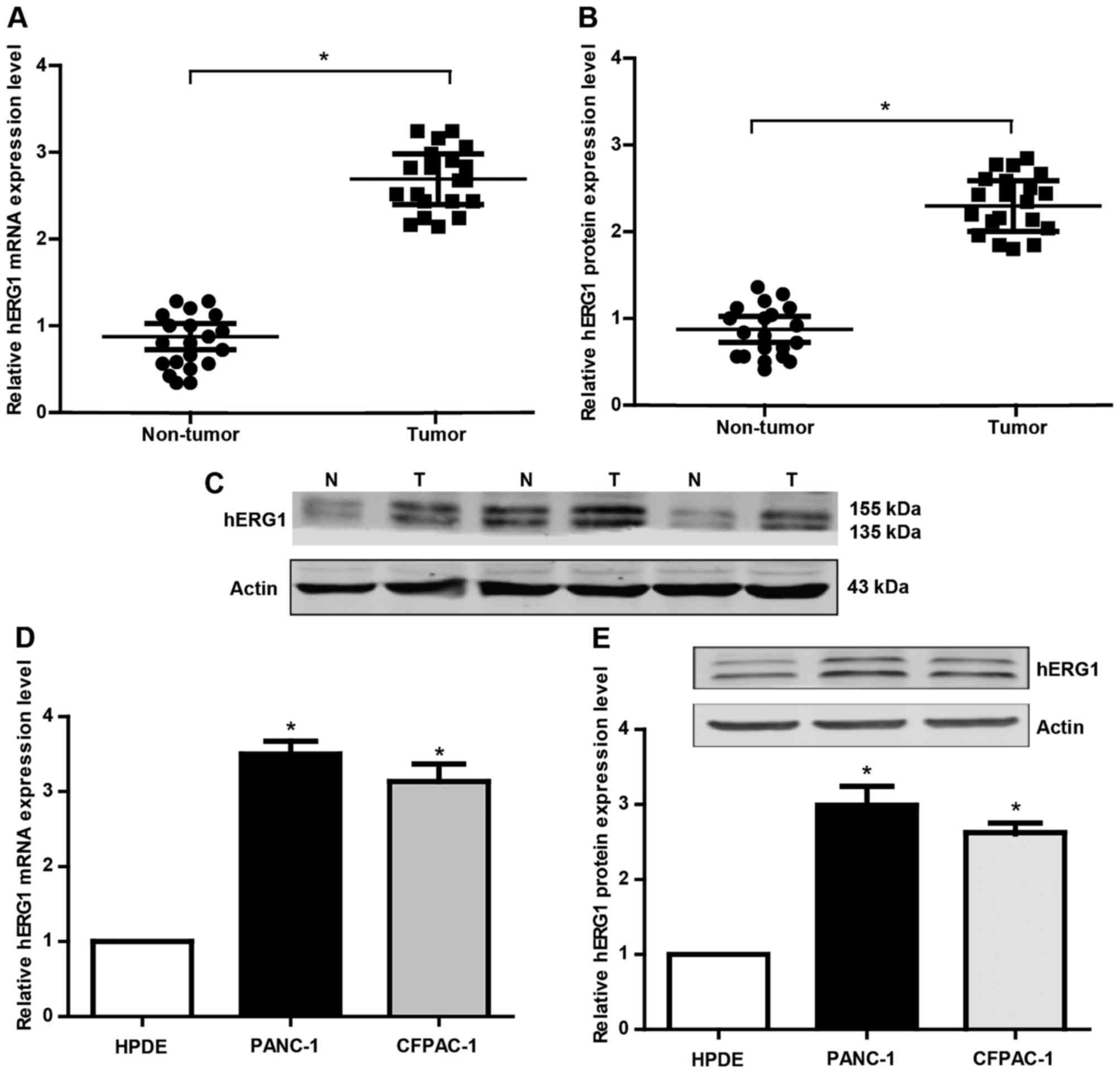
hERG1 is upregulated in human PC
In order to explore the role of hERG1 in pancreatic
carcinogenesis, the mRNA and protein expression of hERG1 was
detected by qRT-PCR and western blot analyses. Twenty pairs of PC
patient tissues and matched noncancerous tissues were collected in
this research. There were 14 males and 6 females with a medium age
at 63 years old. As shown in Fig.
1A-B, the hERG1 was found to be significantly upregulated in PC
tissues compared to the noncancerous tissues. Meanwhile, we also
evaluated hERG1 levels in HPDE, PANC-1, CFPAC-1 cells. The results
(Fig. 1D and E) showed that the
expression level of hERG1 in the PC cell lines (PANC-1, CFPAC-1) is
obviously higher than that in the normal HPDE. These findings
suggested that hERG1level may has a potential correlation with the
pathogenesis of PC.
MiR-493 is downregulated in PC and
directly targets hERG1
miRNA is one of the most important epigenetic
regulators in human cancers. Based on the predictive results of
bioinformatics tools and the expression profile of miRNA from
previous study in PC, we found that miR-493 has potential biding
sites with the 3′UTR of KCNH2 mRNA, which is the coding gene for
hERG1. To validate the actual relationship of miR-493 and hERG1 in
PC, we firstly explored the level of miR-493 in 20 pairs of PC
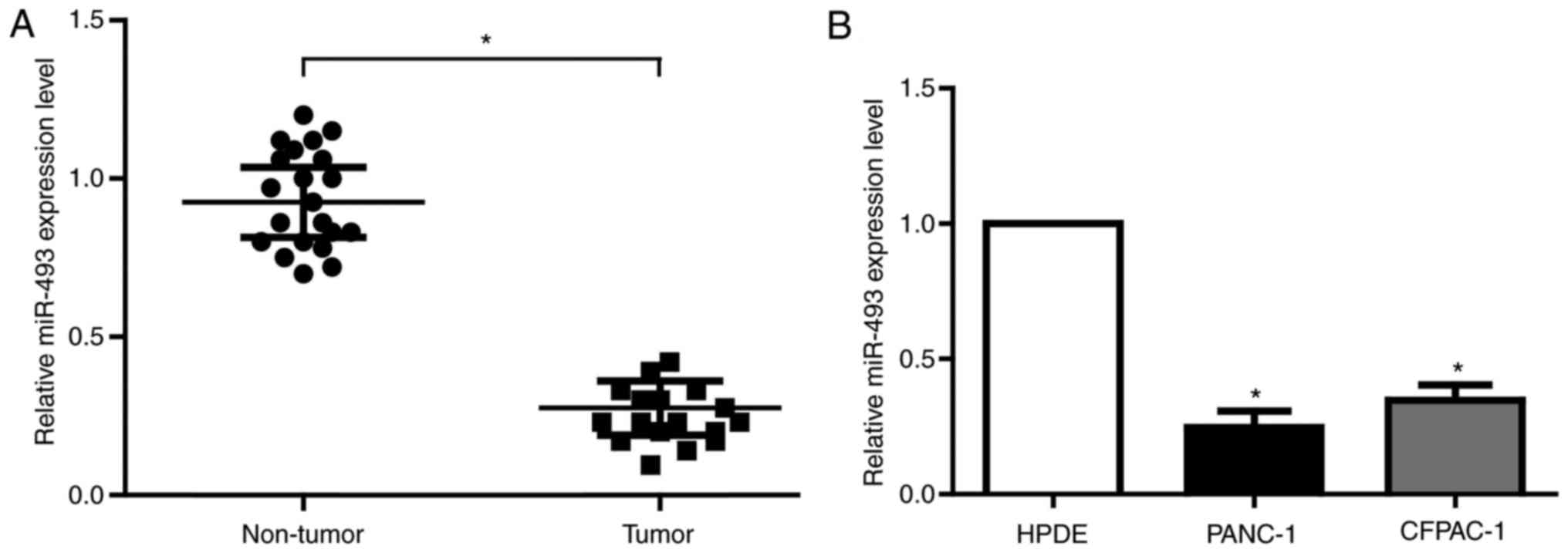
tissues and matched noncancerous tissues. As is expected, miR-493
showed obvious decrease in all the tumor tissues (Fig. 2A). The results from PC cell lines were
in accordance with that in PC tissues (Fig. 2B). Therefore, miR-493 is most likely
to be the important regulator of hERG1 in PC cells.
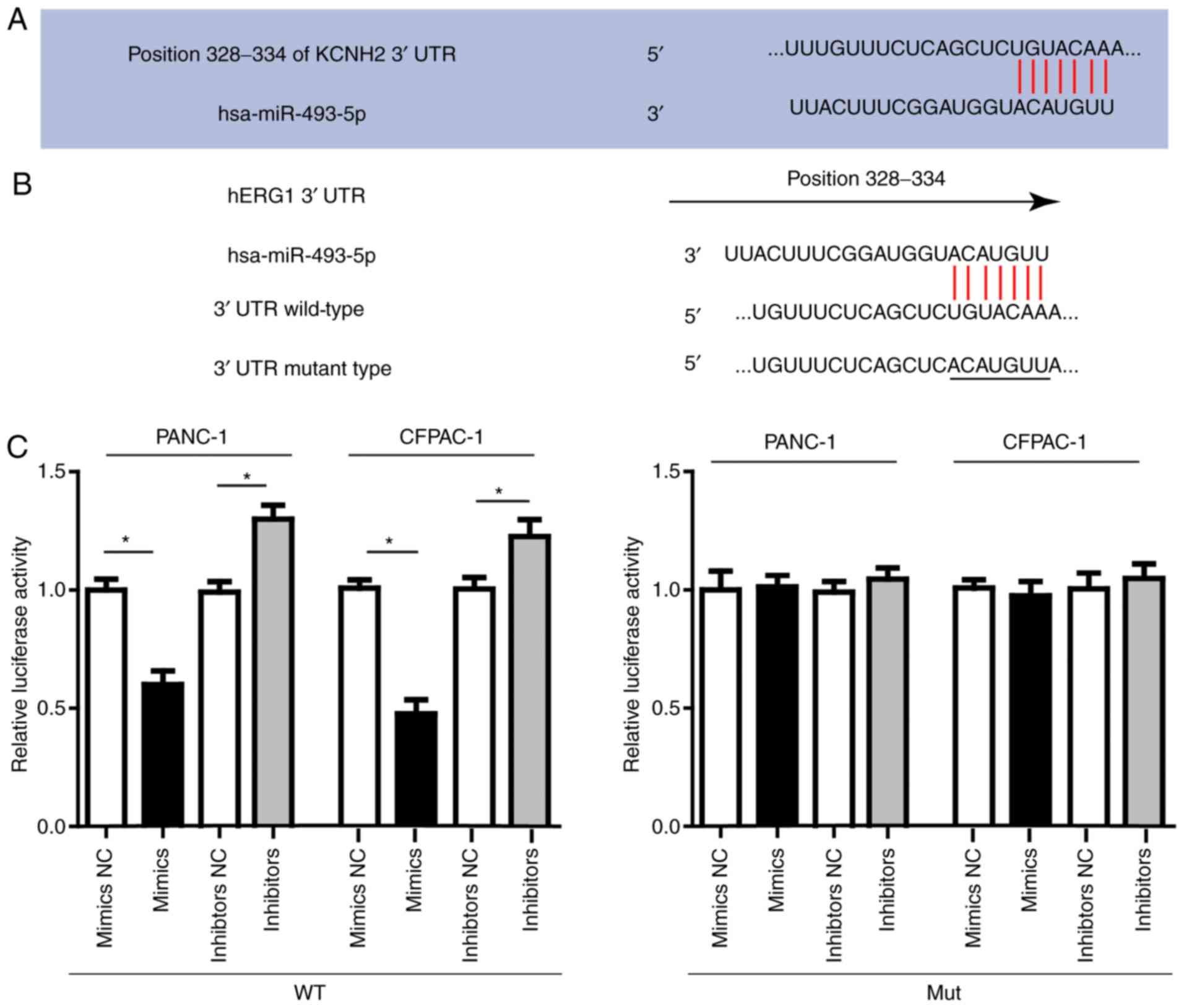
Then we conducted luciferase assay to investigate
the direct interaction between miR-493 and hERG1. In order to
verify this direct targeting relationship, the miR-493 binding
sequence in the 3′-UTR of hERG1 and the mutated 3′-UTR sequence
were inserted into the downstream of the firefly luciferase
reporter gene in a pGL3-promoter vector and then co-transfected
with miR-493 mimics, inhibitors or negative control, respectively
into PANC-1, CFPAC-1 cells. As shown in Fig. 3, the relative luciferase activity in
PANC-1 or CFPAC-1 cells co-transfected with pGL3-hERG1 and miR-493
mimics was significantly decreased by nearly 40% compared with that
in the negative control (NC) group, while the relative luciferase
activity in PANC-1 or CFPAC-1 cells co-transfected with mutated
pGL3-hERG1 and miR-493 mimics or miRNA NC was no different. The
luciferase activity showed relative increase when miR-493
inhibitors were used instead. In conclusion, our results confirmed
that miR-493 could suppress hERG1 expression through direct binding
sequences at the 3′UTR of hERG1 mRNA.
miR-493 regulates hERG1 expression in
PC cells
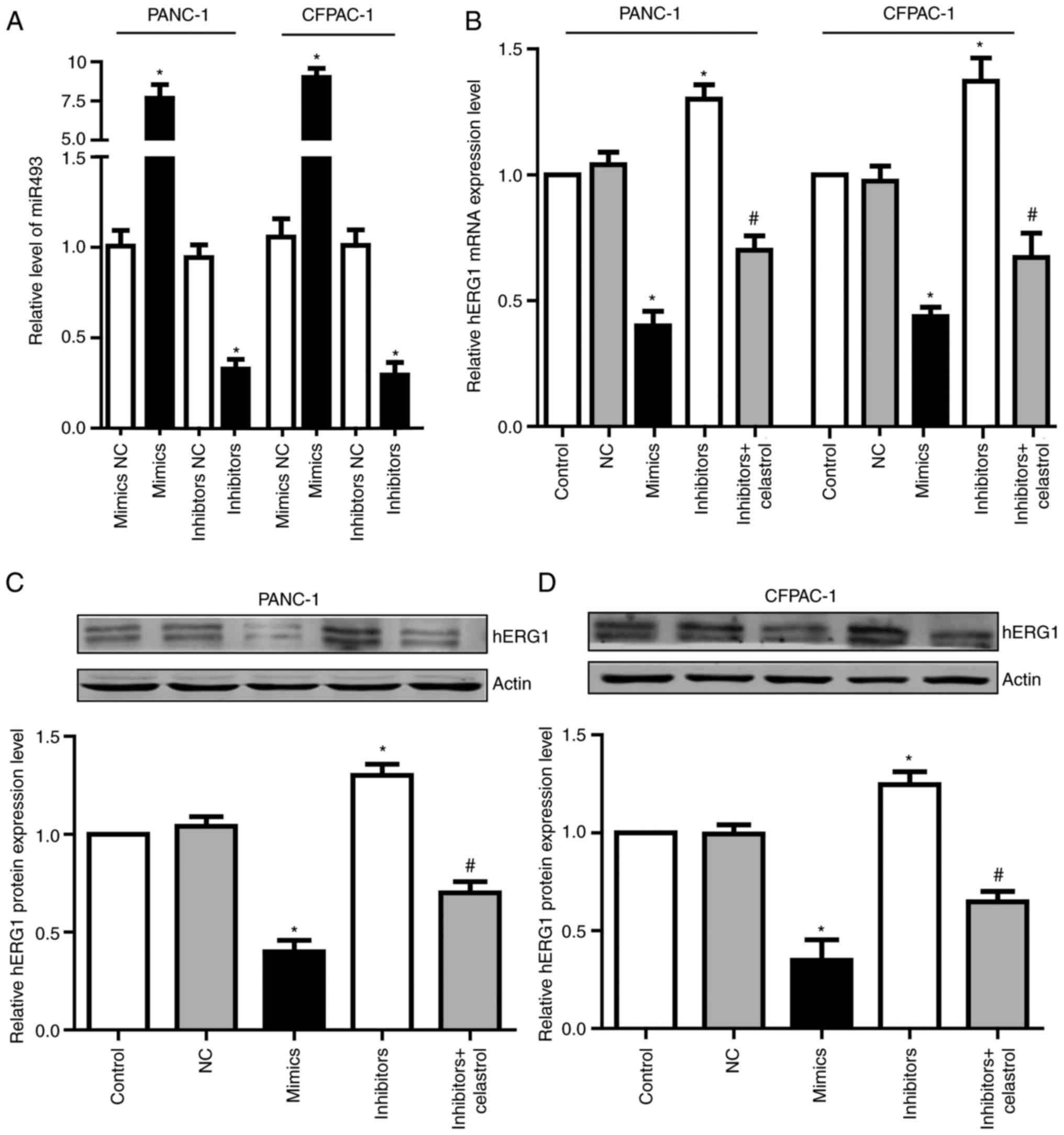
To determine the function of miR-493, we transfected
miR-493 mimics or inhibitors into PANC-1 and CFPAC-1 cells. After
24 h, qRT-PCR assay was performed to detect miR-493 levels. Results
showed that miR-493 mimics could significantly increase miR-493
level in both PANC-1 and CFPAC-1 cells, while miR-493 inhibitors
dramatically decreased miR-493 level (Fig. 4A). To further investigate if miR-493
affected hERG1expression in PC cells, the mRNA and protein level of
hERG1 were measured after the transfection of miR-493 mimics or
inhibitors. As shown in Fig. 4B-D,
overexpression of miR-493 led to the obvious suppression of hERG1,
while knockdown of miR-493 enhances the expression of hERG1 in both
mRNA and protein levels. Additionally, the elevated hERG1 level by
miR-493 inhibitors could be partially reversed by celastrol, which
is a well-known hERG channel inhibitor. These data demonstrated
that miR-493 is an important post-transcriptional regulator of
hERG1 in PC cells.
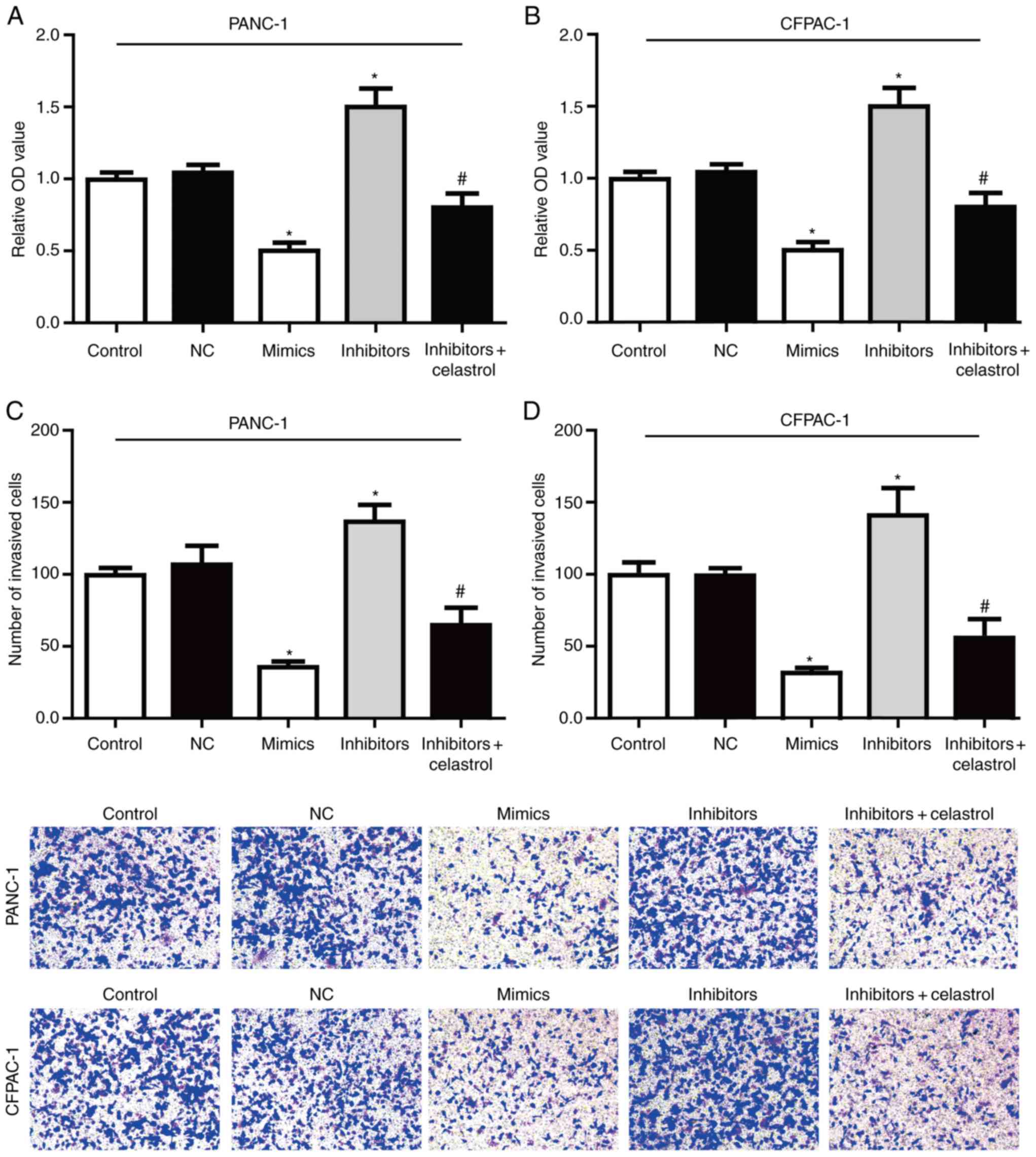
miR-493/hERG1 axis regulates PC cell
proliferation and invasion
The effect of miR-493 on the proliferation of PC
cells was detected in vitro by the MTT assay (Fig. 5A-B). The results showed that
overexpression of miR-493 markedly reduced the cell proliferation
activity in both PANC-1 and CFPAC-1 cells, while knockdown of
miR-493 showed a sharp increased proliferation activity. All these
data suggested that miR-493 could enhance the proliferation of
PANC-1 and CFPAC-1 cells.
In order to determine whether miR-493 affected cell
invasion, Transwell assay were performed after transfection with
miR-493 mimics, inhibitors or NC in PC cell lines. As shown in
Fig. 5C-D, PANC-1 and CFPAC-1 cells
transfected with miR-493 mimics showed a higher ratio in invasion.
On the contrary, knockdown of miR-493 strongly inhibited the cell
invasive ability. Furthermore, the enhanced invasive ability by
miR-493 inhibitors could be largely reversed by co-incubation with
celastrol. Therefore, our data suggested that miR-493 acts as a
tumor suppressor in PC that could inhibit proliferation and
invasion potentially through targeting hERG1, and plays a key role
in the tumorigenesis in PC cells.
Discussion
Despite great efforts made in the treatment of PC,
the prognosis of PC is still very poor with an extremely low 5-year
survival rate. Clinically, PC still lacks tumor markers with enough
specificity and sensitivity. Thus, it's urgently needed to explore
new molecular markers to improve the early diagnosis and accurate
therapy for PC.
The potassium channels have long been suggested to
be related to the regulation of a variety of biological activities
such as the control of cell excitability and the regulation of cell
proliferation and invasion ability (22,23).
Indeed, accumulating evidences have demonstrated that hERG1
channel, a human rapid delayed rectifier in the voltage-gated
potassium channel family, is often aberrantly expressed in numerous
carcinomas including PC. Previous study by Feng J has demonstrated
that hERG1 functions as an oncogene in PC and is downregulated by
miR-96 (23). Since the miRNAs
expression profile is complicated in the progression of diseases,
multiply miRNAs except miR-96, may involved in the process.
Therefore, we tried to explore whether there is another miRNA
involved in the development and progression of PC. In the present
study, we provide evidence that miR-493 inhibits proliferation and
invasion in PC cells and inversely regulated hERG1 expression.
These results implied that multi-target therapy may offer the best
hope for developing an effective PC treatment strategy.
It is becoming increasingly evident that miRNAs are
important epigenetic regulators in human cancers acting as
oncogenes or tumor suppressors to regulate cell proliferation,
apoptosis, migration and invasion (24,25). For
these reasons, we explored the potential miRNAs involved in hERG1
regulation. Based on the complementary pairing prediction by the
bioinformatics tools and the expression profile of miRNA from
previous study in PC by miRNA expression arrays (21), we chose miR-493 for further analysis,
which has putative biding sites in the hERG1 3′-UTR sequence.
Indeed, our in vitro luciferase assay study also confirmed
that miR-493 was a potent regulator of the hERG1 gene. More so,
upregulation of miR-493 in low-expressing PC cells decreases their
malignant potential. These results indicate that miR-493 is an
important tumor suppressor in PC.
Overall, our study shows that miR-493 is an
important oncogene in PC development and progression. Additionally,
we demonstrated that human hERG1 gene is the direct downstream
target of miR-493, with its expression inversely regulated by
miR-493, thus presented a novel epigenetic regulator of miR-493 in
PC. Furthermore, we have shown that overexpression of miR-493 can
inhibit the malignant capacity of PC cells via the hERG1
regulation, while knockdown of miR-493 promotes proliferation and
invasion ability. Based on these results, we proposed the
hypothesis that restoration of miR-493 expression or inhibition of
hERG1 in PC might provide a novel therapeutic strategy for PC.
Acknowledgements
The present study was supported by the Project
funded by China Postdoctoral Science Foundation (grant no.
2015M581491); the Scientific Fund of Heilongjiang Province for
Youth (grant no. QC2015099); the Haiyan Foundation from Harbin
Medical University Cancer Hospital (grant no. JJQN2017-12); and the
Harbin Medical University scientific research innovation fund
(grant no. 2016JCZX59).
References
|
1
|
Moffitt RA, Marayati R, Flate EL, Volmar
KE, Loeza SG, Hoadley KA, Rashid NU, Williams LA, Eaton SC, Chung
AH, et al: Virtual microdissection identifies distinct tumor- and
stroma-specific subtypes of pancreatic ductal adenocarcinoma. Nat
Genet. 47:1168–1178. 2015. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Zhou L, Yao LT, Liang ZY, Zhou WX, You L,
Shao QQ, Huang S, Guo JC and Zhao YP: Nuclear translocation of
fibroblast growth factor receptor 3 and its significance in
pancreatic cancer. Int J Clin Exp Pathol. 8:14640–14648.
2015.PubMed/NCBI
|
|
3
|
Onyeaghala G, Nelson HH, Thyagarajan B,
Linabery AM, Panoskaltsis-Mortari A, Gross M, Anderson KE and
Prizment AE: Soluble MICA is elevated in pancreatic cancer: Results
from a population based case-control study. Mol Carcinog.
56:2158–2164. 2017. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yu Y, Liu L, Ma R, Gong H, Xu P and Wang
C: MicroRNA-127 is aberrantly downregulated and acted as a
functional tumor suppressor in human pancreatic cancer. Tumour
Biol. 37:14249–14257. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Yang XP, Liu SL, Xu JF, Cao SG, Li Y and
Zhou YB: Pancreatic stellate cells increase pancreatic cancer cells
invasion through the hepatocyte growth factor/c-Met/survivin
regulated by P53/P21. Exp Cell Res. 357:79–87. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Jixiang C, Shengchun D, Jianguo Q, Zhengfa
M, Xin F, Xuqing W, Jianxin Z and Lei C: YEATS4 promotes the
tumorigenesis of pancreatic cancer by activating beta-catenin/TCF
signaling. Oncotarget. 8:25200–25210. 2017.PubMed/NCBI
|
|
7
|
Silverman BR and Shi J: Alterations of
epigenetic regulators in pancreatic cancer and their clinical
implications. Int J Mol Sci. 17:pii: E2138. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Mao Y, Shen J, Lu Y, Lin K, Wang H, Li Y,
Chang P, Walker MG and Li D: RNA sequencing analyses reveal novel
differentially expressed genes and pathways in pancreatic cancer.
Oncotarget. 8:42537–42547. 2017.PubMed/NCBI
|
|
9
|
Wang J, Guo XJ, Ding YM and Jiang JX:
miR-1181 inhibits invasion and proliferation via STAT3 in
pancreatic cancer. World J Gastroenterol. 23:1594–1601. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Vandenberg JI, Perry MD, Perrin MJ, Mann
SA, Ke Y and Hill AP: hERG K(+) channels: Structure, function, and
clinical significance. Physiol Rev. 92:1393–1478. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Villoutreix BO and Taboureau O:
Computational investigations of hERG channel blockers: New insights
and current predictive models. Adv Drug Deliv Rev. 86:72–82. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Cubeddu LX: Drug-induced inhibition and
trafficking disruption of ion channels: Pathogenesis of QT
abnormalities and drug-induced fatal arrhythmias. Curr Cardiol Rev.
12:141–154. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Perez-Neut M, Haar L, Rao V, Santha S,
Lansu K, Rana B, Jones WK and Gentile S: Activation of hERG3
channel stimulates autophagy and promotes cellular senescence in
melanoma. Oncotarget. 7:21991–22004. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Becchetti A, Crescioli S, Zanieri F,
Petroni G, Mercatelli R, Coppola S, Gasparoli L, D'Amico M,
Pillozzi S, Crociani O, et al: The conformational state of hERG1
channels determines integrin association, downstream signaling, and
cancer progression. Sci Signal. 10:pii: eaaf3236. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Arcangeli A and Becchetti A: hERG
channels: From antitargets to novel targets for cancer therapy.
Clin Cancer Res. 23:3–5. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Garzon R, Calin GA and Croce CM: MicroRNAs
in cancer. Annu Rev Med. 60:167–179. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Rachagani S, Macha MA, Heimann N,
Seshacharyulu P, Haridas D, Chugh S and Batra SK: Clinical
implications of miRNAs in the pathogenesis, diagnosis and therapy
of pancreatic cancer. Adv Drug Deliv Rev. 81:16–33. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Sun B, Liu X, Gao Y, Li L and Dong Z:
Downregulation of miR-124 predicts poor prognosis in pancreatic
ductal adenocarcinoma patients. Br J Biomed Sci. 73:152–157. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Cai H, An Y, Chen X, Sun D, Chen T, Peng
Y, Zhu F, Jiang Y and He X: Epigenetic inhibition of miR-663b by
long non-coding RNA HOTAIR promotes pancreatic cancer cell
proliferation via up-regulation of insulin-like growth factor 2.
Oncotarget. 7:86857–86870. 2016.PubMed/NCBI
|
|
20
|
Schultz NA, Dehlendorff C, Jensen BV,
Bjerregaard JK, Nielsen KR, Bojesen SE, Calatayud D, Nielsen SE,
Yilmaz M, Holländer NH, et al: MicroRNA biomarkers in whole blood
for detection of pancreatic cancer. JAMA. 311:392–404. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Kato K, Iwama H, Yamashita T, Kobayashi K,
Fujihara S, Fujimori T, Kamada H, Kobara H and Masaki T: The
anti-diabetic drug metformin inhibits pancreatic cancer cell
proliferation in vitro and in vivo: Study of the
microRNAs associated with the antitumor effect of metformin. Oncol
Rep. 35:1582–1592. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Debska G, Kicinska A, Skalska J and
Szewczyk A: Intracellular potassium and chloride channels: An
update. Acta Biochim Pol. 48:137–144. 2001.PubMed/NCBI
|
|
23
|
Feng J, Yu J, Pan X, Li Z, Chen Z, Zhang
W, Wang B, Yang L, Xu H, Zhang G and Xu Z: HERG1 functions as an
oncogene in pancreatic cancer and is downregulated by miR-96.
Oncotarget. 5:5832–5844. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Pengcheng S, Ziqi W, Luyao Y, Xiangwei Z,
Liang L, Yuwei L, Lechen L and Wanhai X: MicroRNA-497 suppress
renal cell carcinoma by targeting VEGFR-2 in ACHN cells. Biosci
Rep. 37:pii: BSR20170270. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Zhang K, Zhang C, Liu L and Zhou J: A key
role of microRNA-29b in suppression of osteosarcoma cell
proliferation and migration via modulation of VEGF. Int J Clin Exp
Pathol. 7:5701–5708. 2014.PubMed/NCBI
|