Introduction
Cervical carcinoma is one of the most common types
of cancer and fourth leading cause of cancer-associated mortality
in women worldwide (1). There are
~529,800 newly diagnosed cervical carcinomas and 275,100 cervical
carcinoma-associated mortalities every year, accounting for ~9% of
all female cancer incidence and mortality (2). It is estimated that >80% of these
cases occur in developing countries (3). Despite recent advances in diagnostic and
treatment strategies in clinical and experimental oncology, the
5-year overall survival rate for patients with advanced disease
remains low (4). Therefore, it is
critical to improve the understanding of the molecular mechanisms
of cervical carcinoma tumorigenesis and progression in order to
facilitate the development of efficient methods for individualized
early diagnosis and treatment of the disease.
MicroRNAs (miRNAs) are an abundant group of small
noncoding RNAs (~22 nucleotides each). They control the expression
of target genes by binding to the 3′-untranslated region (UTR) of
their associated mRNAs and serve an important role in a variety of
biological processes, including cell proliferation, apoptosis,
differentiation, invasion and migration (5–8).
Accumulating studies are demonstrating that miRNAs are dysregulated
in a variety of cancers and serve a critical role in tumorigenesis
(9–14). Recent studies demonstrated that miRNAs
are critical regulators in the development and progression of
cancer, including cervical cancer (15,16).
Therefore, identification of novel miRNAs involved in cervical
cancer progression may contribute to the development of prognostic
biomarkers and therapeutic strategies for cervical cancer.
The miR-30 family contains six distinct mature miRNA
sequences: miR-30a/miR-30c-2, miR-30d/miR-30b, and
miR-30e/miR-30c-1 (17). Accumulating
evidence indicates that the dysregulation of miR-30a contributes to
various malignant tumors, including lung, thyroid, gastric, breast
and colon cancer (18–23). miR-30a promotes tumorigenesis in these
cancers by directly targeting tumor-associated proteins. Zhang
et al (24) demonstrated that
miR-30c inhibited the growth and lung metastasis of colon cancer by
targeting ADAM metallopeptidase domain 19. miR-30a has also been
reported to target insulin receptor substrate 2 in colorectal
tumorigenesis (18). However, the
expression and role of miR-30a in the progression of cervical
cancer remains unclear.
In the present study, the dysregulated expression of
miR-30a in cervical cancer was revealed, and the effect of miR-30a
on cervical cancer cell proliferation and invasion was
investigated. Furthermore, myocyte enhancer factor 2D
(MEF2D), which promotes cervical cancer progression, was
identified as a direct target of miR-30a. In conclusion, miR-30a
acts as a tumor suppressor and may serve as a potential therapeutic
target in cervical cancer.
Materials and methods
Human tissue specimens
Paired cervical cancer and matched normal tissue
specimens were obtained with informed consent from 20 cervical
cancer patients (age range 30–62; mean age 45 years) that had not
undergone preoperative chemotherapy or radiotherapy between January
2014 to December 2015 at the Hunan Provincial Maternal and Child
Health Hospital (Changsha, China). All tissues were obtained during
surgery and immediately stored in liquid nitrogen until use. The
Institute Research Medical Ethics Committee of Hunan Provincial
Maternal and Child Health Hospital granted approval for this
study.
Cell culture and transfection
The cervical cancer cell lines HeLa, SiHa and
Ca-Ski, as well as the normal human cervical epithelial GH329 and
human embryonic kidney 293 cell lines, were obtained from the
American Type Culture Collection (Manassas, VA, USA). All cells
were cultured in Dulbecco's modified Eagle's medium (DMEM;
Invitrogen; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
supplemented with 10% fetal bovine serum (FBS; Invitrogen; Thermo
Fisher Scientific, Inc.). Cultures were maintained at 37°C in a
humidified atmosphere with 5% CO2.
miR-30a mimic, miR-30a inhibitor, mimic negative
control, and inhibitor negative control were synthesized by
Shanghai GenePharma Co., Ltd. (Shanghai, China). For transfection,
HeLa cells were seeded in 12-well plates and transiently
transfected with 100 nM of the following mimics/inhibitors: miR-30a
(hsa-miR-30a) mimic, miR-30a inhibitor, mimic negative control
(designated as mimic control), and inhibitor negative control
(designated as inhibitor control) for 8 h using Lipofectamine 2000
according to the manufacturer's protocol (Invitrogen; Thermo Fisher
Scientific, Inc.). A total of 24 h after transfection, cells were
collected for qPCR, cell viability and invasion. After 48 h
transfection, cells were collected for western blotting.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
The RNA was extracted from human cervical cancer
cells and tissue specimens, as well as normal cervical cells and
tissues, using TRIzol solution (Sigma-Aldrich; Merck Millipore,
Darmstadt, Germany). RT was performed to obtain complementary DNA
using a PrimeScript RT Reagent kit (Takara Bio, Inc., Otsu, Japan)
according to the manufacturer's protocol. gDNA eraser was added to
1 µg RNA at 42°C for 2 min, and RT Primer mix was added at 37°C for
15 min, followed by 85°C for 5 sec. qPCR was performed with SYBR
Premix Ex Taq II (Takara Bio, Inc.) using 300 ng cDNA with the
CFX96™ Real-Time PCR Detection System (Bio-Rad Laboratories, CA).
The cycling conditions for the qPCR were 95°C for 2 min followed by
45 cycles of 95°C for 15 sec and 60°C for 30 sec. The primer
sequences for miR-30a and MEF2D were described previously (18,25).
miR-30a expression in each sample was calculated by normalizing
with U6 and the MEF2D expression in each sample was calculated by
normalizing with GAPDH. U6 forward, 5′-CTCGCTTCGGCAGCACA-3′ and
reverse, 5′-AACGCTTCACGAATTTGCGT-3′; and GAPDH forward,
5′-CGGAGTCAACGGATTTGGTCGTAT-3′ and reverse,
5′-AGCCTTCTCCATGGTGGTGAAGAC-3′ The expression level of each miRNA
was measured using the 2ΔΔCq method (26). All samples were run in triplicates in
the same culture plate.
Luciferase reporter gene assays
miRNA target prediction websites www.microRNA.org and TargetScan (www.targetscan.org) were used to predict the target
gene of miR-30a. The 3′-UTR of MEF2D containing the putative
binding site of miR-30a was amplified and subcloned into a pGL3
luciferase promoter vector (Promega Corporation, Madison, WI, USA),
as described previously (27); the
putative binding site was mutated as negative control (MEF2D-Mut).
The vector was co-transfected with miR-30a mimic into 293 cells for
48 h. The cells were harvested and relative luciferase activity was
detected using a Dual-Luciferase Reporter Assay System (Promega
Corporation) according to the manufacturer's protocol. All
experiments were performed at least three times.
Western blot analysis
Whole cell extracts were prepared with
radioimmunoprecipitation assay buffer according to the
manufacturer's protocol (Sigma-Aldrich; Merck Millipore), and the
protein was quantified using a Pierce BCA Protein Assay kit (Thermo
Fisher Scientific, Inc.). The protein samples (30 µg) were
separated by SDS-PAGE (10%), transferred to a methanol-activated
polyvinylidene fluoride membrane and blocked with 5% milk at room
temperature for 1 h, and then detected by western blot using a
polyclonal rabbit anti-MEF2D antibody (1:1,000, HPA004807, Santa
Cruz Biotechnology, Inc., Santa Cruz, CA, USA), incubated at 4°C
overnight. The membranes were subsequently incubated with a goat
anti-rabbit IgG secondary antibody (1:5,000; cat. no. 65-6120;
Pierce; Thermo Fisher Scientific, Inc.) at room temperature for 1 h
conjugated to horseradish peroxidase. Blots were then developed
using an Enhanced Chemiluminescence kit (Pierce; Thermo Fisher
Scientific, Inc.) following manufacturer's instructions. ImageJ
v2.1.4.7 (National Institutes of Health, Bethesda, MD, USA) was
used to quantify band density of western blot experiments.
Cell viability assay
An MTT assay was employed to assess cell viability,
as described previously (28), in
HeLa cells transfected with miR-30a mimic, inhibitor, mimic control
or inhibitor control.
Invasion assay
A total of 24 h after HeLa cells were transiently
transfected with the aforementioned mimics/inhibitors, cell
invasion ability was examined by Transwell invasion assay. Cells
were seeded (2×105 cells/well) into 12-well plates. The
Transwell migration chambers were used to evaluate cell invasion.
Transwell insert chambers were covered with Matrigel (BD
Biosciences, Franklin Lakes, NJ, USA) to detect the ability of
invasion. Matrigel (10 mg/ml) was coated on the upper side of the
filter, and collagen was coated on the lower side of the filter.
The upper chamber was filled with cells in serum-free DMEM, and the
lower chamber was filled with DMEM containing 5% FBS. Cells were
incubated for 48 h at 37°C, and then non-invading cells were
removed by swabbing the top layer of Matrigel with a cotton swab.
The cells were stained with 0.1% Crystal Violet Staining for 15 min
at 37°C. For each well, ten random fields were counted and the
average number of cells was determined under a light microscope at
×100 magnification. The invasion ratio was equal to the cell number
in the experimental group divided by the cell number in the control
group. All the experiments were repeated in triplicate.
Statistical analysis
Each experiment was repeated at least three times.
Data are presented as the mean ± SD and analyzed using SPSS 19.0
(IBM SPSS, Armonk, NY, USA). Comparison of more than two groups was
made using one-way analysis of variance with Tukey's post hoc test.
Comparison of two groups was made using Student's t-test for
unpaired data. P<0.05 was considered to indicate a statistically
significant difference.
Results
miR-30a is downregulated in cervical
cancer tissues and cell lines
miR-30a expression was detected in 20 human cervical
cancer and adjacent normal tissues, as well as in cervical cancer
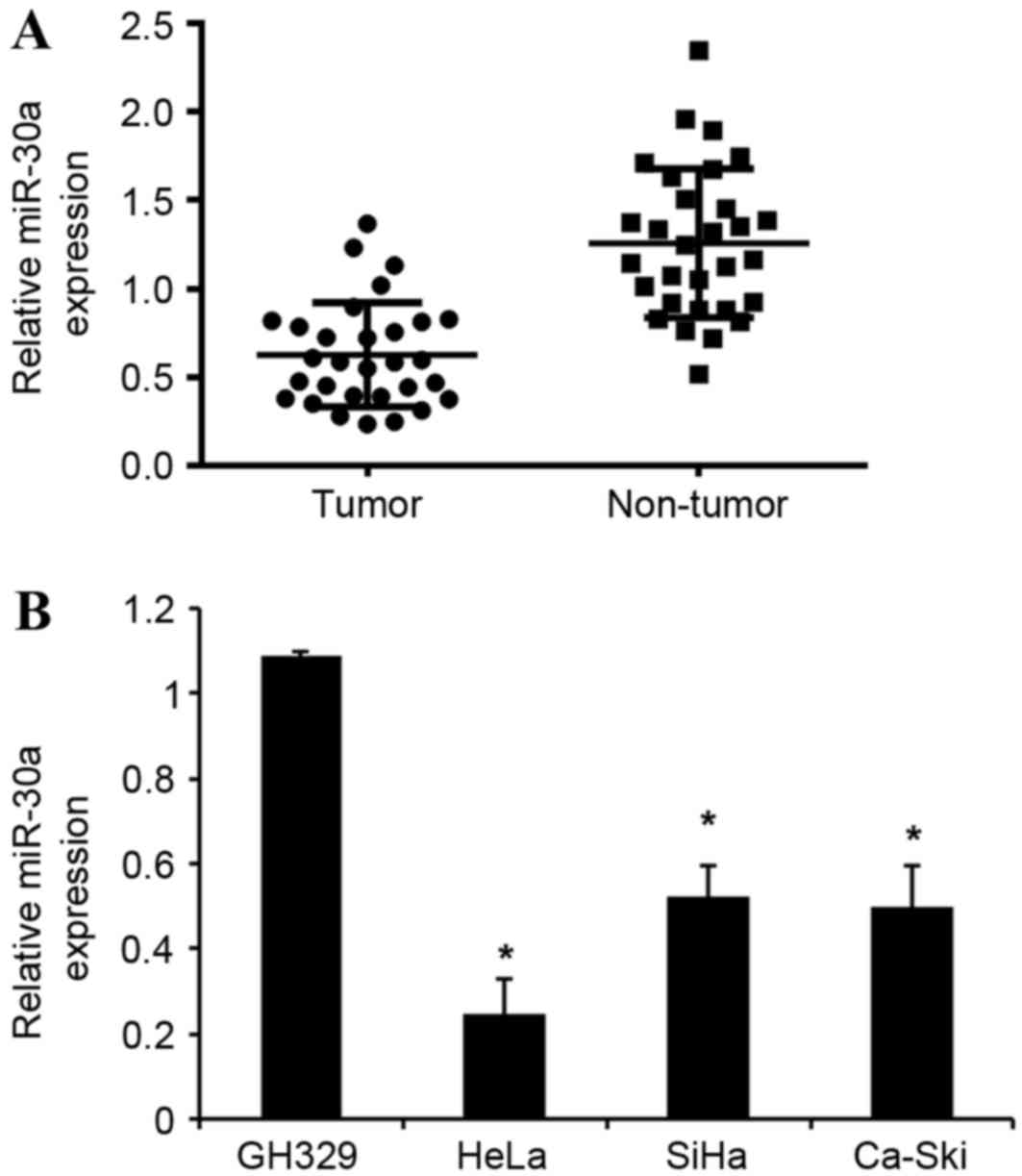
and normal cervical cell lines, using an RT-qPCR assay. As shown in
Fig. 1A, miR-30a expression was
significantly downregulated in cervical cancer tissues compared
with corresponding adjacent normal tissues. Furthermore, the
expression levels of miR-30a in the three cervical cancer cells
(HeLa, SiHa and Ca-Ski) were significantly downregulated compared
with that of human normal cervical cell line GH329 (Fig. 1B). Collectively, these data suggest
that the downregulation of miR-30a may be involved in the
tumorigenesis of cervical cancer.
miR-30a inhibits cervical cancer cell
viability and invasion
To evaluate the role of miR-30a in cervical cancer
progression, HeLa cells were transfected with miR-30a mimic,
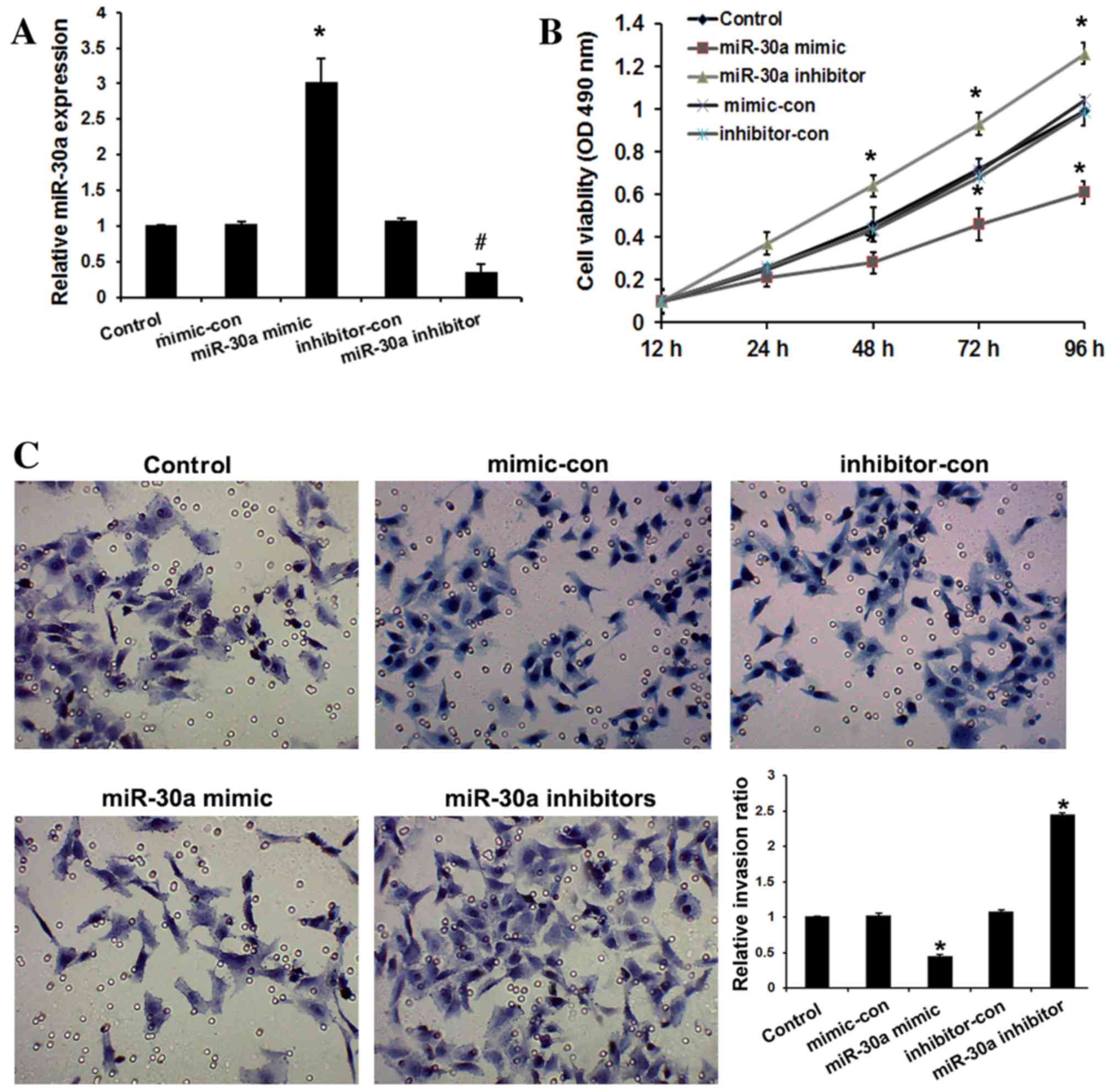
inhibitors, and their respective negative controls. As shown in
Fig. 2A, miR-30a mimic significantly
upregulated miR-30a expression, while miR-30a inhibitors
significantly downregulated miR-30a expression in HeLa cells
compared with the controls (P<0.05). The effect of miR-30a on
the viability of HeLa cells was examined by MTT assay. It was
observed that overexpression of miR-30a resulted in significantly
decreased viability of cervical cancer cells relative to the
controls after 48 h (P<0.05), whereas miR-30a inhibitors
increased the viability of HeLa cells relative to the controls
(P<0.05; Fig. 2B). To determine
the function of miR-30a in cervical cancer progression, the
invasive abilities of transfected cell were determined. Compared
with the controls, overexpression of miR-30a in cervical cancer
cells significantly suppressed cell invasion (P<0.01), whereas
loss of its expression promoted invasion (P<0.01; Fig. 2C). These observations suggest that
miR-30a serves an important role in inhibiting the invasiveness of
cervical cancer cells.
miR-30a inhibits MEF2D expression by
binding to its 3′-UTR
In the present study, miRNA target prediction
websites www.microRNA.org and TargetScan
(www.targetscan.org) predicted that
MEF2D is one of the targets of miR-30a. A conserved
miR-30a-binding site in the 3′-UTR of MEF2D mRNA was
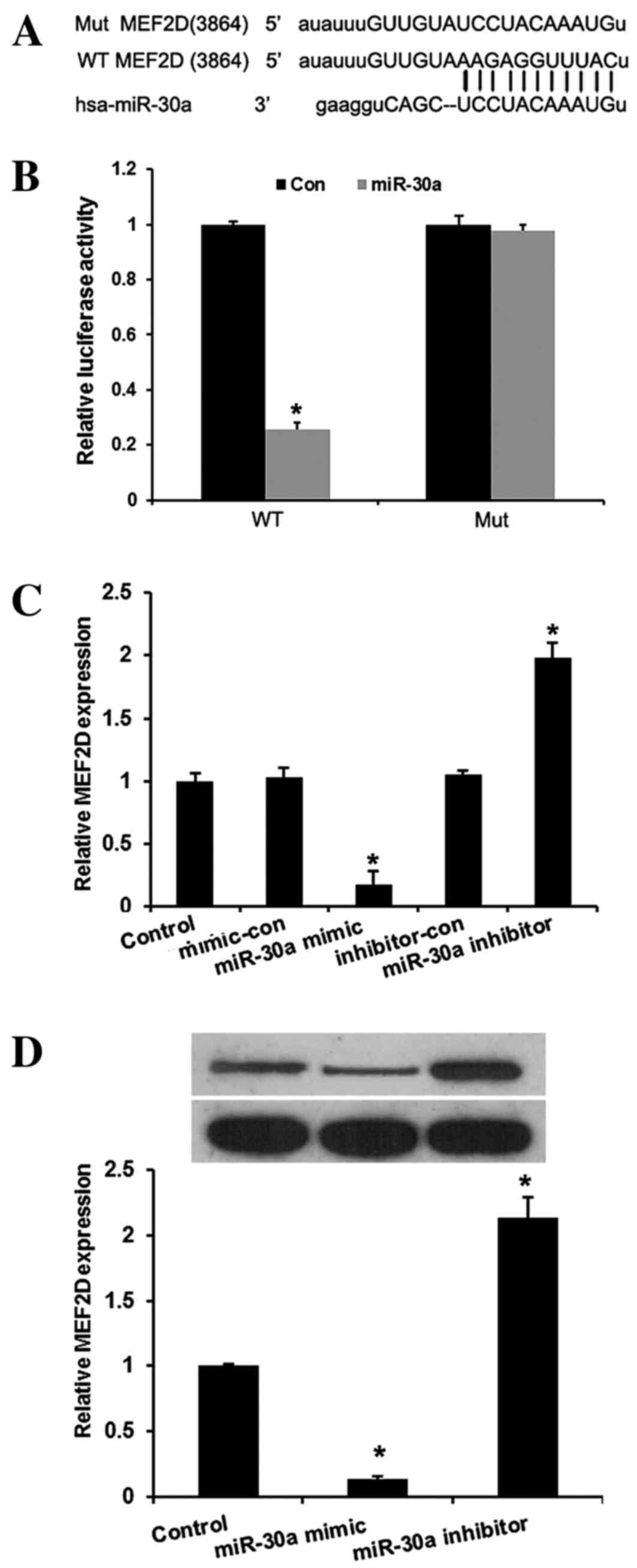
identified. To verify this, a wild-type or mutant target region
sequence of the MEF2D 3′-UTR was cloned and inserted into a
luciferase reporter vector (Fig. 3A).
These reporter vectors were co-transfected along with the miR-30a
mimic and mimic control into the HEK293 cell line. As shown in
Fig. 3B, the Dual-Luciferase reporter
assay revealed that miR-30a mimic suppressed the luciferase
activity of the reporter with wild-type MEF2D 3′-UTR
sequence and failed to inhibit that of mutated MEF2D,
indicating that MEF2D is one of the direct targets of
miR-30a in cervical cancer cells. To further confirm MEF2D
as a direct target of miR-30a, RT-qPCR and western blot assays were
used to detect the expression of MEF2D in HeLa cells. As
shown in Fig. 3C and D, the mRNA and
protein levels of MEF2D were significantly downregulated by
miR-30a mimic and upregulated by miR-30a inhibitors in HeLa cells
compared with the controls (P<0.01).
MEF2D expression is increased in
cervical cancer and inversely correlated with miR-30a levels
The association between MEF2D and miR-30a was
further analyzed by detecting the mRNA expression levels of
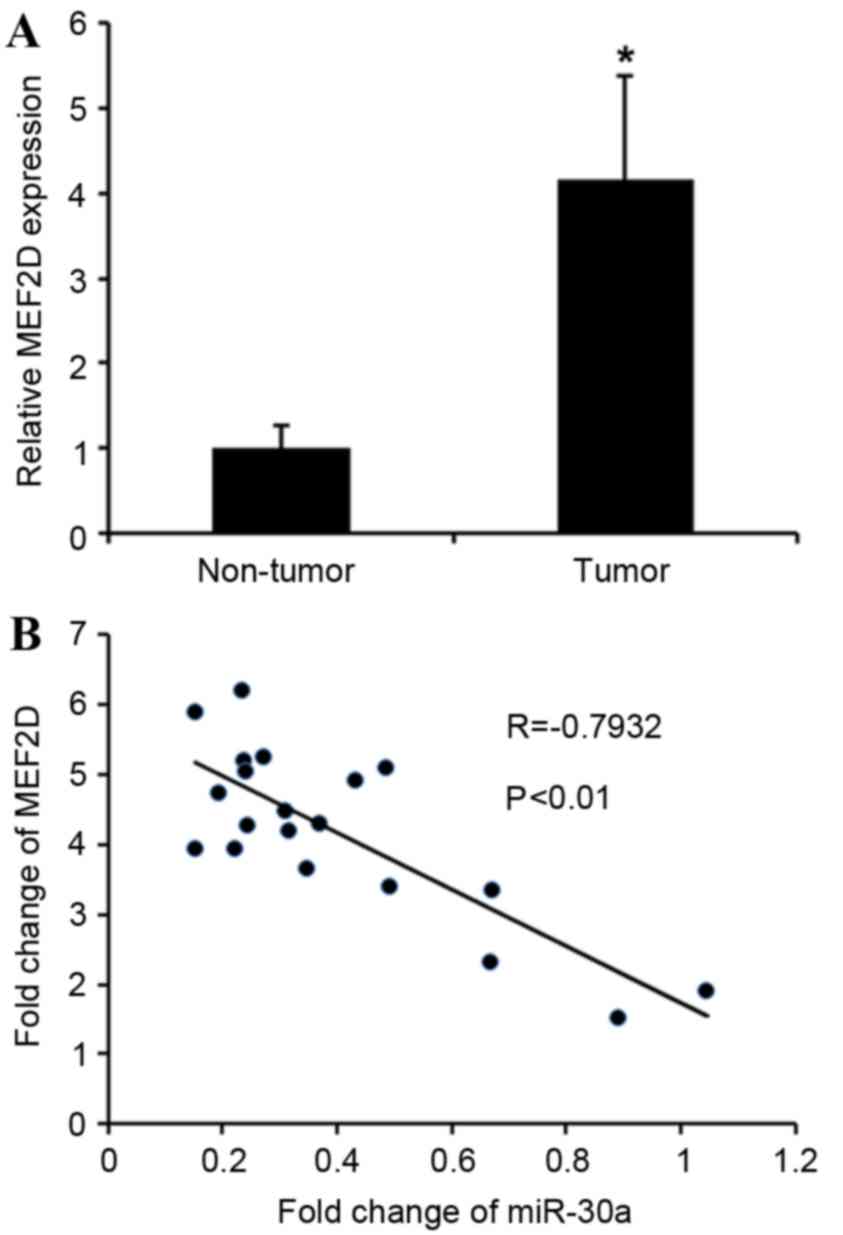
MEF2D in cervical cancer tissues. MEF2D levels in
cervical cancer tissues were greatly upregulated compared with
adjacent non-tumor tissues (Fig. 4A).
MEF2D levels were compared with miR-30a expression in the
same cervical cancer specimens. As shown in Fig. 4B, Spearman's correlation analysis
demonstrated a significant inverse correlation between the levels
of miR-30a and MEF2D mRNA (r=−0.7932; P<0.01).
Taken together, these data strongly support the hypothesis that
MEF2D is a direct target of miR-30a.
Discussion
Increasing evidence has indicated that miRNAs are
involved in tumorigenic processes by targeting a variety of
tumor-associated genes (29).
Previous studies have demonstrated that miR-30a is involved in the
progression of various malignant tumors (20,21,23).
However, the mechanism of miR-30a in the progression of cervical
cancer remains unclear. In the present study, it was revealed that
miR-30a expression was significantly downregulated in cervical
cancer tissue and cell lines. Further investigation indicated that
miR-30a was able to regulate the viability and invasion of cervical
cancer cells by targeting MEF2D. Therefore, for the first
time, miR-30a has been revealed as a tumor suppressor in the
progression of cervical cancer.
The MEF2 transcription factors serve roles in
muscular, cardiac, skeletal, vascular, neural, blood and immune
cell development through their effects on cell differentiation,
proliferation, apoptosis, migration, morphology and metabolism
(30). Altered MEF2 activity serves a
role in human diseases and it has recently been implicated in the
initiation and progression of various types of cancer in humans
(31). Recently, it was reported that
MEF2D, one member of MEF2 family, is involved in the progression of
several cancer types, including lung carcinoma (27), rhabdomyosarcoma (32), hepatocellular carcinoma (31) and osteosarcoma (25).
In the present study, a luciferase reporter assay
revealed that MEF2D is a direct target of miR-30a in
cervical cancer cells. Overexpression of miR-30a reduced
MEF2D mRNA and protein levels in cervical cancer cells. A
significant negative correlation was observed between the levels of
miR-30a and MEF2D mRNA in the same cervical cancer
specimens. These results indicated that miR-30a functions as a
tumor suppressor in cervical cancer by targeting MEF2D.
Taken together, the results of the present study
have revealed for the first time that miR-30a is a tumor suppressor
in cervical cancer; that the expression levels of miR-30a were
significantly decreased in tumor tissues and cell lines, and that
its ectopic expression inhibited cell proliferation and invasion.
Furthermore, a Dual-Luciferase reporter assay revealed that
MEF2D is a direct target of miR-30a, and MEF2D mRNA
expression was shown to be negatively correlated with miR-30a
expression in cervical cancer. These results indicate that miR-30a
dysregulation may serve important roles in cervical cancer
progression and that the interaction between miR-30a and
MEF2D may be a therapeutic target in the treatment of
cervical cancer.
References
|
1
|
Kanavos P: The rising burden of cancer in
the developing world. Ann Oncol. 17 Suppl 8:viii15–viii23. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Reeler A, Qiao Y, Dare L, Li J, Zhang AL
and Saba J: Women's cancers in developing countries: From research
to an integrated health systems approach. Asian Pac J Cancer Prev.
10:519–526. 2009.PubMed/NCBI
|
|
3
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Sharma V, Kerr SH, Kawar Z and Kerr DJ:
Challenges of cancer control in developing countries: Current
status and future perspective. Future Oncol. 7:1213–1222. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li Z, Lei H, Luo M, Wang Y, Dong L, Ma Y,
Liu C, Song W, Wang F, Zhang J, et al: DNA methylation
downregulated mir-10b acts as a tumor suppressor in gastric cancer.
Gastric Cancer. 18:43–54. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Xiao X, Tang C, Xiao S, Fu C and Yu P:
Enhancement of proliferation and invasion by MicroRNA-590-5p via
targeting PBRM1 in clear cell renal carcinoma cells. Oncol Res.
20:537–544. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Yin WZ, Li F, Zhang L, Ren XP, Zhang N and
Wen JF: Down-regulation of microRNA-205 promotes gastric cancer
cell proliferation. Eur Rev Med Pharmacol Sci. 18:1027–1032.
2014.PubMed/NCBI
|
|
8
|
Yang X, Ni W and Lei K: miR-200b
suppresses cell growth, migration and invasion by targeting Notch1
in nasopharyngeal carcinoma. Cell Physiol Biochem. 32:1288–1298.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Liu Z, Mai C, Yang H, Zhen Y, Yu X, Hua S,
Wu Q, Jiang Q, Zhang Y, Song X and Fang W: Candidate tumour
suppressor CCDC19 regulates miR-184 direct targeting of C-Myc
thereby suppressing cell growth in non-small cell lung cancers. J
Cell Mol Med. 18:1667–1679. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yang Q, Wang Y, Lu X, Zhao Z, Zhu L, Chen
S, Wu Q, Chen C and Wang Z: MiR-125b regulates
epithelial-mesenchymal transition via targeting Sema4C in
paclitaxel-resistant breast cancer cells. Oncotarget. 6:3268–3279.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Gong B, Hu H, Chen J, Cao S, Yu J, Xue J,
Chen F, Cai Y, He H and Zhang L: Caprin-1 is a novel microRNA-223
target for regulating the proliferation and invasion of human
breast cancer cells. Biomed Pharmacother. 67:629–636. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang J, Raimondo M, Guha S, Chen J, Diao
L, Dong X, Wallace MB, Killary AM, Frazier ML, Woodward TA, et al:
Circulating microRNAs in pancreatic juice as candidate biomarkers
of pancreatic cancer. J Cancer. 5:696–705. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Duan HF, Li XQ, Hu HY, Li YC, Cai Z, Mei
XS, Yu P, Nie LP, Zhang W, Yu ZD and Nie GH: Functional elucidation
of miR-494 in the tumorigenesis of nasopharyngeal carcinoma. Tumour
Biol. 36:6679–6689. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Lu J, He ML, Wang L, Chen Y, Liu X, Dong
Q, Chen YC, Peng Y, Yao KT, Kung HF and Li XP: MiR-26a inhibits
cell growth and tumorigenesis of nasopharyngeal carcinoma through
repression of EZH2. Cancer Res. 71:225–233. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zheng W, Liu Z, Zhang W and Hu X: miR-31
functions as an oncogene in cervical cancer. Arch Gynecol Obstet.
292:1083–1089. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Song X, Shi B, Huang K and Zhang W:
miR-133a inhibits cervical cancer growth by targeting EGFR. Oncol
Rep. 34:1573–1580. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chang TC, Yu D, Lee YS, Wentzel EA, Arking
DE, West KM, Dang CV, Thomas-Tikhonenko A and Mendell JT:
Widespread microRNA repression by Myc contributes to tumorigenesis.
Nat Genet. 40:43–50. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Zhang Q, Tang Q, Qin D, Yu L, Huang R, Lv
G, Zou Z, Jiang XC, Zou C, Liu W, et al: Role of microRNA 30a
targeting insulin receptor substrate 2 in colorectal tumorigenesis.
Mol Cell Biol. 35:988–1000. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Visone R, Pallante P, Vecchione A,
Cirombella R, Ferracin M, Ferraro A, Volinia S, Coluzzi S, Leone V,
Borbone E, et al: Specific microRNAs are downregulated in human
thyroid anaplastic carcinomas. Oncogene. 26:7590–7595. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Li X, Zhang Y, Zhang Y, Ding J, Wu K and
Fan D: Survival prediction of gastric cancer by a seven-microRNA
signature. Gut. 59:579–585. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Cheng CW, Wang HW, Chang CW, Chu HW, Chen
CY, Yu JC, Chao JI, Liu HF, Ding SL and Shen CY: MicroRNA-30a
inhibits cell migration and invasion by downregulating vimentin
expression and is a potential prognostic marker in breast cancer.
Breast Cancer Res Treat. 134:1081–1093. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Yang C and Pan Y: Fluorouracil induces
autophagy-related gastric carcinoma cell death through Beclin-1
upregulation by miR-30 suppression. Tumour Biol. Jul 25–2015.(Epub
ahead of print).
|
|
23
|
Kumarswamy R, Mudduluru G, Ceppi P,
Muppala S, Kozlowski M, Niklinski J, Papotti M and Allgayer H:
MicroRNA-30a inhibits epithelial-to-mesenchymal transition by
targeting Snai1 and is downregulated in non-small cell lung cancer.
Int J Cancer. 130:2044–2053. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Zhang Q, Yu L, Qin D, Huang R, Jiang X,
Zou C, Tang Q, Chen Y, Wang G, Wang X and Gao X: Role of
microRNA-30c targeting ADAM19 in colorectal cancer. PLoS One.
10:e01206982015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Yu H, Sun H, Bai Y, Han J, Liu G, Liu Y
and Zhang N: MEF2D overexpression contributes to the progression of
osteosarcoma. Gene. 563:130–135. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Song L, Li D, Zhao Y, Gu Y, Zhao D, Li X,
Bai X, Sun Y, Zhang X, Sun H, et al: miR-218 suppressed the growth
of lung carcinoma by reducing MEF2D expression. Tumour Biol.
37:2891–2900. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Han K, Zhao T, Chen X, Bian N, Yang T, Ma
Q, Cai C, Fan Q, Zhou Y and Ma B: microRNA-194 suppresses
osteosarcoma cell proliferation and metastasis in vitro and in vivo
by targeting CDH2 and IGF1R. Int J Oncol. 45:1437–1449. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Xu JQ, Zhang WB, Wan R and Yang YQ:
MicroRNA-32 inhibits osteosarcoma cell proliferation and invasion
by targeting Sox9. Tumour Biol. 35:9847–9853. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Pon JR and Marra MA: MEF2 transcription
factors: Developmental regulators and emerging cancer genes.
Oncotarget. 7:2297–2312. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Ma L, Liu J, Liu L, Duan G, Wang Q, Xu Y,
Xia F, Shan J, Shen J, Yang Z, et al: Overexpression of the
transcription factor MEF2D in hepatocellular carcinoma sustains
malignant character by suppressing G2-M transition genes. Cancer
Res. 74:1452–1462. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhang M, Truscott J and Davie J: Loss of
MEF2D expression inhibits differentiation and contributes to
oncogenesis in rhabdomyosarcoma cells. Mol Cancer. 12:1502013.
View Article : Google Scholar : PubMed/NCBI
|