Introduction
Prostate cancer (PCa) is a common malignancy of the
urinary system in men aged >50 years old. In 2012, PCa was the
second leading cause of cancer-associated mortality in the USA
(1). The incidence of PCa has
continued to increase in the past decades due to the wide use of
serum prostate-specific antigen (PSA) screening throughout the
world (2). Although the survival
rates for patients with PCa are high, 30–40% of patients experience
PSA recurrence within 10 years of surgery or radiation treatment
(3). However, previous studies have
raised concern that PSA may not be an optimal prognostic biomarker.
For example, the overall sensitivity of PSA may be too low to
predict the morbidity and mortality rate accurately (4). By contrast, since the sensitivity of PSA
is too low, false-positive diagnosis of PCa commonly occurs and
results in over-detecting and overtreating, with, the majority of
males with PCa succumbing to unrelated causes of mortality
(5). Therefore, there is an urgent
need to identify novel, robust biomarkers for PCa.
Tumor protein D54 (TPD54), also termed TPD52L2, is
the third identified member of the tumor protein D52 (TPD52) family
(6). The family members are
characterized by an N-terminal coiled-coil motif that is used to
form homo- and heterologous complexes with other tumor protein
D52-1ike proteins (7). The most
important physiological function of this family protein is its
involvement in Ca2+-dependent vesicular transport
processes (8). A previous study
demonstrated that TPD52 is highly expressed in multiple types of
human solid tumor, including lung, colon, ovary and breast cancer
(9), which indicates that TPD52
family members may serve as novel markers of malignant tumors. A
previous study demonstrated that TPD54 is a negative regulator of
extracellular matrix-dependent migration and attachment in oral
squamous cell carcinoma (10). In
addition, lentivirus-mediated knockdown of TPD54 inhibits cell
proliferation of glioma (11), oral
squamous cell carcinoma (OSCC) (12),
gastric cancer (13), breast cancer
(14) and liver cancer (15) cells in vitro. The association
between the expression of TPD54 and clinical outcome in prostate
cancer (16) and breast childhood
leukemia (17) has also been
reported. In spite of these results, however, the associations
between TPD54 and the clinicopathological characteristics of
prostate cancer have not been fully explored.
The present study aimed to evaluate the expression
of TPD54 in PCa tumor tissues and adjacent noncancerous tissues,
and to compare TPD54 expression with clinicopathological
characteristics. Finally, the value of TPD54 as a prognostic
biomarker for patients with PCa was evaluated.
Materials and methods
Patients and tissue samples
The present study was conducted in accordance with
the ethical principles for human experimentation stated in the
Declaration of Helsinki and all applicable amendments that have
been defined by the International Conference of Harmonization Good
Clinical Practice Guidelines (E6 R2), ensuring that the rights,
safety and well-being of the subjects are safeguarded and that the
integrity of the data acquired during the study was preserved. The
study protocol was approved by the Ethics Committee of the Tongde
Hospital of Zhejiang Province (Zhejiang, China; approval number:
2012-025) and written informed consent was obtained for all
patients prior to their participation in the present study. The
present study consisted of samples from 117 patients with PCa who
underwent radical retropubic prostatectomy between January 2011 and
January 2013 at the Department of Urology, Tongde Hospital of
Zhejiang Province. All the samples were obtained with informed
consent from the patients, which was provided prior to surgery. The
clinicopathological characteristics of patients, including age at
diagnosis, preoperative PSA, postoperative pathological state,
Gleason score, pathological T stage, capsule penetration, surgical
margins, perineural invasion and seminal vesicle invasion are
presented in Table I. All the
diagnoses of patients were confirmed by pathologists. All of the
tumor specimens consisted of primary prostate tumor tissues and
adjacent normal prostate tissues, and each specimen was divided
into two parts in a standard process. Briefly, surgically resected
specimens were placed in the correct orientation. The prostate was
cut along the coronal plane into 6–8 parts; each section was ~5 mm
thick. PCa tissues and adjacent normal tissues were carefully
recognized and cut into two parts, and each specimen was divided
into two parts along the coronal plane. One was snap-frozen in
liquid nitrogen, and stored at −180°C for extraction of RNA, while
the other part of the specimen was fixed in 10% buffered formalin
at room temperature overnight for immunohistochemistry (IHC). A
total of 111 patients with PCa were successfully followed-up,
whereas 6 patients who succumbed to diseases other than PCa or from
unexpected events were excluded. The mean follow-up time was 42
(9–61) months. For the analysis of biochemical recurrence
(BCR)-free survival, the date of prostatectomy was used to
represent the beginning of the follow-up period. The endpoint was
the time to biochemical relapse, which was defined as the period
between surgical treatment and two consecutive measurements of
serum PSA ≥0.2 ng/ml.
 | Table I.Clinicopathological characteristics of
117 patients with prostate cancer. |
Table I.
Clinicopathological characteristics of
117 patients with prostate cancer.
| Features | Patients |
|---|
| Median age (range),
years | 68 (42–80) |
| PSA, ng/ml |
|
|
<10 | 34 (29) |
|
10–20 | 33 (28) |
|
>20 | 50 (43) |
| GS |
|
|
<7 | 41 (35) |
| =7 | 45 (38) |
|
>7 | 31 (26) |
| pT stage |
|
| pT2 | 97 (83) |
| pT3 | 20 (17) |
| pN stage |
|
| N0 | 109 (93) |
| N1 | 8 (7) |
| CP |
|
| + | 7 (6) |
| − | 110 (94) |
| SM |
|
| + | 14 (12) |
| − | 103 (88) |
| PNI |
|
| + | 24 (21) |
| − | 93 (79) |
| SVI |
|
| + | 18 (15) |
| − | 99 (85) |
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA from the primary tumor and adjacent normal
tissues was extracted using TRIzol reagent (Thermo Fisher
Scientific, Inc., Waltham, MA, USA) according to the manufacturer's
protocol. RNA (1 µg) from each sample was used for cDNA synthesis
using M-MLV Reverse Transcriptase (Thermo Fisher Scientific, Inc.)
according to the manufacturer's protocol. The reverse transcription
system was as follows: Oligo (dT)12-18 (1 µl), total RNA
(1 µg), 10 mM dNTP (1 µl) and distilled water up to 12 µl. The
mixture was heated to 65°C for 5 min and chilled on ice. To the
mixture was then added 5X First-Strand Buffer (4 µl), 0.1 M DTT (2
µl), recombinant ribonuclease inhibitor (1 µl), M-MLV RT (1 µl),
which was mixed and incubated for 50 min at 37°C. The mRNA levels
were quantified by qPCR with the Applied Biosystems 7500 PCR system
using SYBR Master Mix (Thermo Fisher Scientific, Inc.). The
sequences of the primers were as follows: TPD54 forward,
5′-CATGACGTGCAGGTCTCTAGC-3′ and reverse,
5′-GCCTGTGAAAGAGTTTCCTGAGT-3′; GAPDH forward,
5′-GCACCGTCAAGGCTGAGAAC-3′ and reverse, 5′-TGGTGAAGACGCCAGTGGA-3′.
The PCR conditions were as follows: 50°C for 2 min and 95°C for 10
min, followed by 40 cycles at 95°C for 15 sec and 60°C for 1 min.
Relative quantification of TPD54 mRNA expression was calculated
using the 2−ΔΔCq method (17). Normalization was performed against
GAPDH mRNA expression.
Western blotting. Total proteins from 8 pairs of
fresh prostate cancer and adjacent normal tissues were extracted
using RIPA buffer (cat. no., 9806; Cell Signaling Technology, Inc.,
Danvers, MA, USA). Briefly, tissues were cut into small pieces, and
20 mg of tissue was homogenized with a glass tissue grinder in
100–200 µl RIPA buffer on ice for 30 min. Lysates were then
centrifuged at 8,000 × g at 4°C for 20 min, and the supernatants
were collected. The protein concentrations were determined using a
bicinchoninic acid assay kit (Beyotime Institute of Biotechnology,
Haimen, China). Next, equal amounts of loading buffer were added
and the protein was boiled. Proteins were resolved using a 12% gel
and SDS-PAGE, transferred to polyvinylidene fluoride membranes and
incubated with the following antibodies: Anti-TPD52L2 (dilution,
1:500; cat. no., 11795-1-AP, ProteinTech Group, Inc., Chicago, IL,
USA) and anti-β-actin (dilution, 1:2,000; cat. no., 60008-1-Ig,
ProteinTech Group, Inc.) at room temperature for 2 h. Following
washing with TBS-Tween-20 (TBST) three times, the membranes were
incubated with corresponding horseradish peroxidase-conjugated goat
anti-mouse IgG (dilution, 1:2,000; cat. no. A0216; Beyotime
Institute of Biotechnology) or goat anti-rabbit IgG (dilution,
1:2,000; cat. no. A0239; Beyotime Institute of Biotechnology)
antibodies were used as secondary antibodies for 1 h at room
temperature, and then washed three times with TBST. The final band
was visualized using enhanced chemiluminescence reagents (Thermo
Fisher Scientific, Inc.) and detected by an Alpha Imager v.FC800
(Alpha Innotech, San Leandro, CA, USA). The statistical analyses
were conducted from three independent experiments.
Immunohistochemical staining
Paraffin-embedded samples were cut into 4-µm thick
sections and stained with hematoxylin and eosin for 10 min at room
temperature for tumor confirmation. The expression level of TPD52L2
was determined using a Strept Avidin-Biotin Complex
immunohistochemical assay kit (catalog no. SA1027, Wuhan Boster
Biological Technology, Ltd., Wuhan, China) according to the
manufacturer-s protocol. Briefly, antigen retrieval was performed
by immersing the slides in a solution of 0.01% sodium citrate pH
6.0 for 5 min in boiling water. Endogenous peroxidase activity was
inhibited by immersing the slides in 3%
H2O2-methanol at room temperature for 10 min
and then incubated in 5% goat antiserum (cat. no. C0265; Beyotime
Institute of Biotechnology) for 15 min at 37°C. The sections were
then sequentially incubated overnight at 4°C with the rabbit
anti-human TPD52L2 polyclonal antibody (dilution, 1:100; cat. no.,
11795-1-AP, ProteinTech Group, Inc.), biotinylated goat anti-rabbit
immunoglobulin G (cat. no. TA130016; dilution, 1:1,000; OriGene
Technologies, Inc., Beijing, China) and avidin biotin-peroxidase
complex. Following staining with diaminobenzidine for 3-10 min at
room temperature, sections were analyzed under a light microscope
(magnification, ×100; Olympus BX-40; Olympus Corporation, Tokyo,
Japan).
Immunohistochemical staining was evaluated
independently by two pathologists in Department of Pathology,
Tongde Hospital of Zhejiang Province (Hangzhou, China). The level
of TPD54 staining was based on the intensity of staining and the
proportion of positively stained cancer cells. Intensity of
staining was scored as follows: 0, no staining; 1, light yellow; 2,
yellow brown; 3, strong brown color. The proportion positive tumor
cells was scored as follows: 0, ≤5% positive tumor cells; 1,
6<25% positive tumor cells; 2, 26–50% positive tumor cells; 3,
51–75% positive tumor cells and 4, ≥76% positive tumor cells. The
final basis (IS; immunoreactivity score) for grouping was the
product of the staining area score and staining intensity as
follows: Negative (−), score 0; weak (+), scores 1–4; moderate
(++), scores 5–8; strong positive (++), scores 9–12.
Statistical analysis
Statistical analyses were performed using SPSS 19.0
software (IBM SPSS, Armonk, NY, USA). Results are expressed as the
mean ± standard deviation. The differences in TPD54 expression
between PCa and adjacent normal prostate tissues were analyzed by
the non-parametric Mann-Whitney U-test. The distribution of TPD54
expression in patients with PCa to clinicopathological features was
analyzed using non-parametric Kruskal-Wallis one-way ANOVA and
Dunn´s post hoc test. Univariate and multivariate Cox regression
analyses were performed to analyze the survival data. Survival
curves were plotted using the Kaplan-Meier method, and differences
were tested using the log-rank test. P<0.05 was considered to
indicate a statistically significant difference.
Results
TPD54 protein is overexpressed in
patients with PCa
The mRNA and protein expression of TPD54 was
initially examined by RT-qPCR and western blotting in 8 randomly
selected pairs of primary prostate cancer and adjacent normal
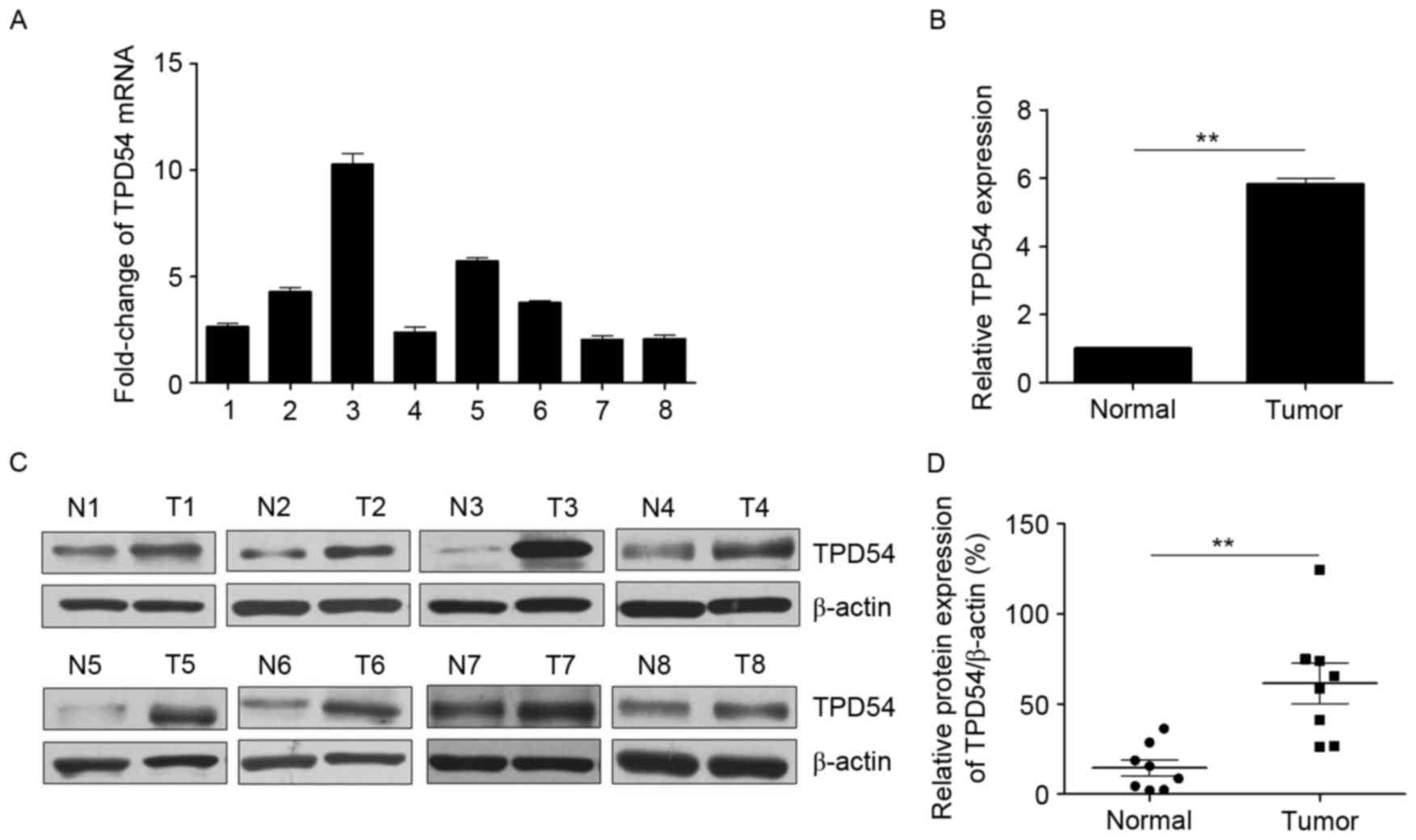
prostate tissues. As presented in Fig.
1, a significant increase in mRNA and protein expression of
TPD54 was observed in prostate cancer tissues compared with
adjacent normal tissues. In addition, TPD54 expression was
evaluated in 117 paired prostate cancer cases by
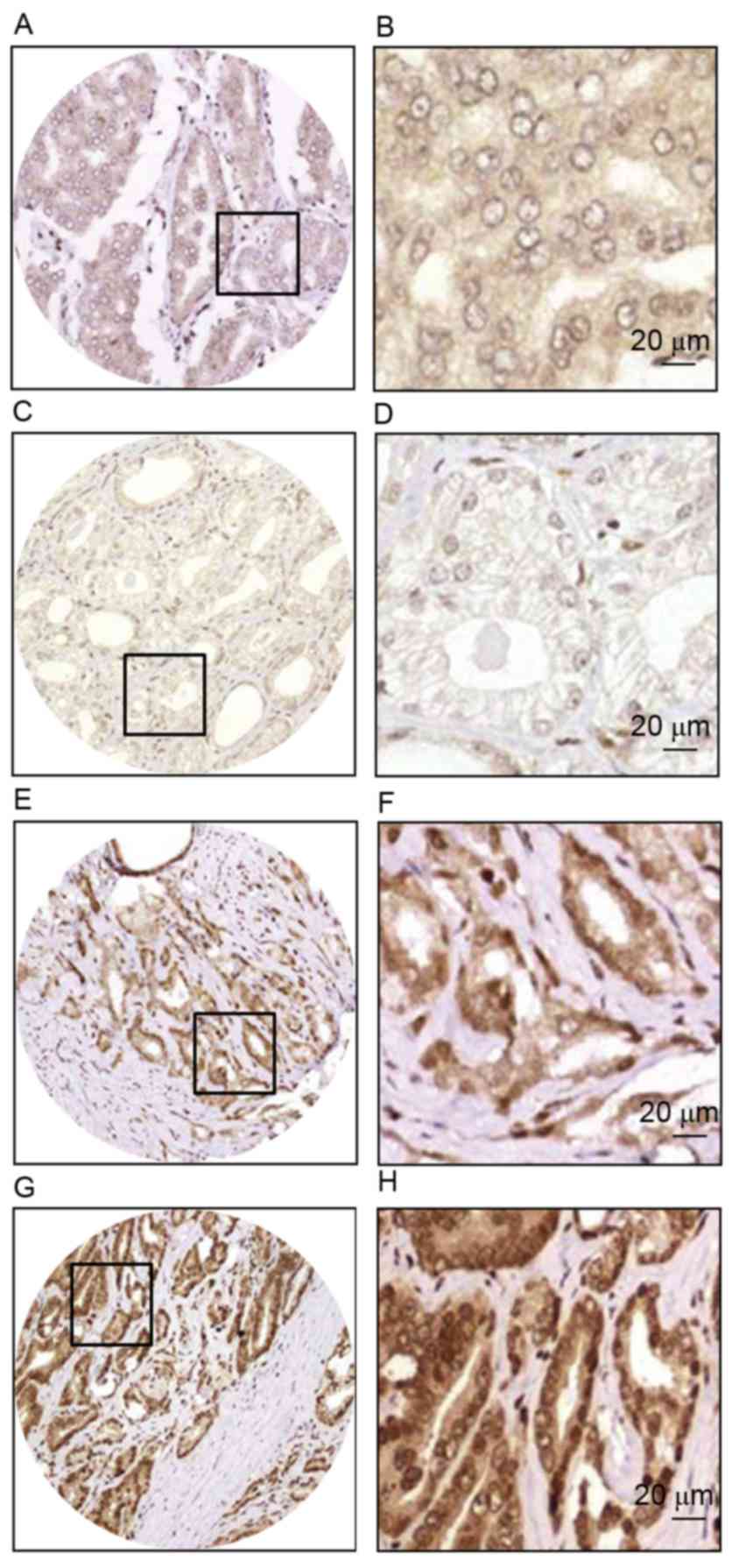
immunohistochemistry (IHC). The IHC data revealed that TPD54 was
mainly localized in the cytoplasm of the prostate cancer cells, and
immunoreactivity ranged between 0 and 100% (Fig. 2). According to the IHC score analysis,
in prostate tumor tissues, 1/117 (0.9%) exhibited negative TPD54
staining, 9/117 (7.7%) weak staining, 75/117 (64.1%) moderate
staining and 32/117 (27.3%) strong staining; whereas in adjacent
normal prostate tissues, 15/117 (12.8%) demonstrated negative TPD54
staining, 64/117 (51.7%) weak staining, 37/117 (31.6%) moderate
staining and 1/117 (0.9%) strong staining. The Mann-Whitney U test
revealed that the expression level of TPD54 in prostate cancer
tissues was significantly higher than that in adjacent normal
prostate tissues (P=0.0001). Taken together, these results
demonstrated that TPD54 expression was upregulated in PCa tissues,
and may be involved in PCa progression.
TPD54 expression is associated with
clinical data in patients with PCa
Whether TPD54 expression was associated with
patient-associated clinical factors was subsequently explored. The
associations between TPD54 expression and clinicopathological
features are summarized in Table II.
The results indicated that TPD54 expression was significantly
associated with Gleason score (P=0.0001). However, no significant
associations were identified with age, preoperative PSA,
pathological T stage, postoperative pathological state, capsule
penetration, surgical margins, perineural invasion and seminal
vesicle invasion. These data demonstrated that increased TPD54
expression may promote tumor growth of PCa.
 | Table II.Association of intra-tumoral tumor
protein D54 expression and clinicopathological variables of 117
patients with prostate cancer. |
Table II.
Association of intra-tumoral tumor
protein D54 expression and clinicopathological variables of 117
patients with prostate cancer.
|
| TPD54 expression,
n |
|
|---|
|
|
|
|
|---|
| Variables | Negative | Weak | Moderate | Strong | P-value |
|---|
| Age, years |
|
|
|
|
|
|
<65 | 0 | 3 | 23 | 12 | 0.738 |
|
65–75 | 1 | 4 | 36 | 13 |
|
|
>75 | 0 | 2 | 13 | 7 |
|
| PSA, ng/ml |
|
|
|
|
|
|
<10 | 0 | 5 | 21 | 8 | 0.126 |
|
10–20 | 0 | 4 | 22 | 7 |
|
|
>20 | 1 | 0 | 32 | 17 |
|
| GS |
|
|
|
|
|
|
<7 | 1 | 7 | 25 | 8 | 0.005 |
| =7 | 0 | 2 | 33 | 10 |
|
|
>7 | 0 | 0 | 17 | 14 |
|
| pT stage |
|
|
|
|
|
| pT2 | 1 | 5 | 63 | 28 | 0.136 |
| pT3 | 0 | 4 | 12 | 4 |
|
| pN stage |
|
|
|
|
|
| N0 | 1 | 9 | 69 | 30 | 0.148 |
| N1 | 0 | 0 | 6 | 2 |
|
| CP |
|
|
|
|
|
| + | 0 | 1 | 5 | 1 | 0.381 |
| − | 1 | 8 | 70 | 31 |
|
| SM |
|
|
|
|
|
| + | 0 | 2 | 7 | 5 | 0.770 |
| − | 1 | 7 | 68 | 27 |
|
| PNI |
|
|
|
|
|
| + | 0 | 0 | 16 | 8 | 0.191 |
| − |
|
|
|
|
|
| 1 |
| 9 | 59 | 24 |
|
| SVI |
|
|
|
|
|
| + | 0 | 1 | 13 | 4 | 0.816 |
| − | 1 | 8 | 62 | 28 |
|
Associations between TPD54 expression
and prognosis in patients with PCa
To determine whether TPD54 expression is a
prognostic factor for patients with PCa, Kaplan-Meier analysis and
log-rank tests were performed to analyze TPD54 expression and
clinical follow-up data in 111 patients. Of the 111 patients with
PCa, biochemical recurrence occurred in 27 cases (24.3%). The
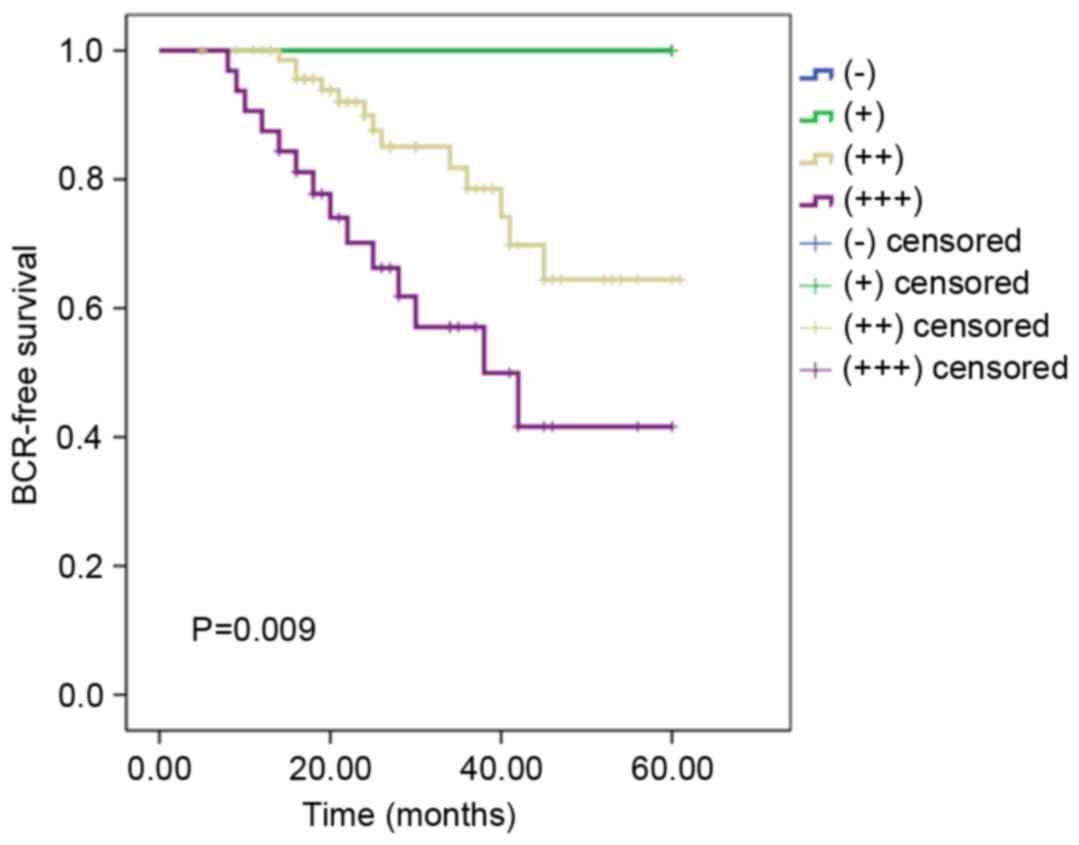
results demonstrated that the BCR-free survival rate for patients
with strong TPD54 expression was 56.3%, which was significantly
lower than that in patients with moderate TPD54 expression (82.7%;
P=0.009) and patients with weak/negative TPD54 expression (100.0%).
These results revealed that a high level of TPD54 expression is a
negative indicator of prognosis for patients with PCa (Fig. 3). In addition, univariate and
multivariate analyses revealed that TPD54 expression, PSA, Gleason
score, pT stage, pN stage, perineural invasion were all independent
prognostic factors in patients with PCa (Table III). Therefore, the present results
demonstrated that TPD54 may be a significant prognostic marker for
patients with PCa.
 | Table III.Univariate and multivariate analyses
of factors associated with biochemical recurrence-free survival of
patients with prostate cancer. |
Table III.
Univariate and multivariate analyses
of factors associated with biochemical recurrence-free survival of
patients with prostate cancer.
|
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|
|---|
| Factors | Hazard ratio | 95% CI | P-value | Hazard ratio | 95% CI | P-value |
|---|
| Age, years (≤70,
>70) | 1.941 | 0.908–4.149 | 0.087 |
|
|
|
| PSA, ng/ml (<10,
10–20, >20) | 2.166 | 1.268–3.698 | 0.005 | 2.047 | 1.547–4.269 | 0.037 |
| GS (<7, 7,
>7) | 4.555 | 2.423–8.566 | <0.001 | 3.832 | 1.966–7.469 | <0.001 |
| pT Stage (pT2,
pT3) | 5.649 | 2.631–12.129 | <0.001 | 8.097 | 3.606–18.180 | <0.001 |
| pN Stage (pN0,
pN1) | 7.858 | 3.090–19.983 | <0.001 | 7.694 | 3.897–10.670 | <0.001 |
| CP (+, -) | 1.328 | 0.314–5.619 | 0.700 |
|
|
|
| SM (+, -) | 1.885 | 0.713–4.985 | 0.201 |
|
|
|
| PNI (+, -) | 2.842 | 1.291–6.258 | 0.009 | 2.364 | 1.841–5.632 | 0.013 |
| SVI (+, -) | 1.239 | 0.469–3.276 | 0.666 |
|
|
|
| TPD54 expression
(−, +, ++, ++) | 2.790 | 1.407–5.534 | 0.003 | 2.259 | 1.090–4.679 | 0.028 |
Discussion
At present, radical prostatectomy is one of the most
effective methods for the treatment of localized PCa (2). However, a significant percentage of
prostate tumors progress rapidly and this is responsible for the
poor survival of patients, irrespective of active treatment
(3). Therefore, identification of
PCa-specific biomarkers may be important for diagnosis, therapy and
prognostic prediction. To the best of our knowledge, the present
study has demonstrated, for the first time, that the expression of
TPD54 in 117 patients with PCa is associated with patient survival
and clinicopathological characteristics. Positive TPD54 expression
was observed in 91.4% of PCa tissues in the present study.
Furthermore, a significant association was observed between
positive TPD54 expression and poor prognosis, independent of other
patient characteristics. These results indicated that TPD54
expression may be a novel prognostic marker for PCa.
TPD54, which was upregulated in several types of
solid malignancies, is an oncogene responsible for the high
proliferation rates observed in cancer cells (9). Several studies have reported that the
suppression of TPD54 gene inhibits the growth of tumor cells. TPD54
was overexpressed in OSCC-derived cell lines. Knockdown of TPD54 in
OSCC cell lines induces cell growth inhibition, promotes cell
apoptosis, and inhibits extracellular matrix-dependent cell
migration and attachment in OSCC (10,12).
Furthermore, Yang et al (14)
reported that Lentivirus-mediated TPD54 inhibited proliferation and
colony formation in breast cancer cell line, ZR-75-30. Furthermore,
knockdown of TPD54 promoted glycogen synthase kinase-3β
phosphorylation in ZR-75-30 cells. Similar results were observed in
glioma (11), gastric cancer
(13) and liver cancer (15). Previous studies have also reported the
association between the expression of TPD54 and clinical outcome in
PCa (16) and childhood leukemia
(17). Accordingly, in the present
study, TPD54 mRNA expression in PCa samples was investigated using
qPCR analysis and TPD54 protein expression was investigated using
western blot analysis. The results demonstrated that TPD54 mRNA and
protein levels were significantly increased in tumor tissues
compared with those in the adjacent normal prostate tissues. These
results indicated that TPD54 may serve a critical function in PCa
progression. However, to learn more about the biological function
of TPD54 in PCa development, knockdown or overexpression assays in
PCa cell lines are urgently required.
In the present study, TPD54 protein expression was
analyzed in tumor tissues by IHC in order to assess its prognostic
significance for PCa. IHC analysis revealed moderate/strong TPD54
expression in 91.4% (107/117) of patients with PCa, markedly higher
than that in adjacent normal prostate tissues (32.5%, 38/117). High
expression of TPD54 was revealed to be significantly associated
with the Gleason score of patients with PCa, indicating that TPD54
expression may promote tumor growth of PCa. These results suggested
that TPD54 may be involved in the tumorigenesis and progression of
PCa. Furthermore, the results demonstrated that patients with high
TPD54 expression presented with a shorter BCR-free survival time
compared with those with low TPD54 expression. Univariate analyses
revealed that increased TPD54 expression in PCa tissues was
significantly associated with BCR-free survival. In addition,
multivariate analysis revealed that TPD54 expression was an
independent prognostic factor for patients with PCa. These results
suggested that TPD54 may be an important prognostic marker for
patients with PCa. Notably, using the Gene Expression Omnibus
database and Top-scoring pairs algorithm, Zhang et al
(16) revealed that TPD52L2/SQLE gene
pair was a top-scoring pair for predicting prostate tumor
progression.
In conclusion, the present study demonstrated that
TPD54 was overexpressed in patients with PCa, and its expression
was associated with a worse clinical outcome. TPD54 may serve as a
potential prognostic indicator for PCa, but may not serve as an
optimal biomarker for clinical use currently. Additional studies
are required using a larger cohort of tumor samples to confirm the
prognostic function of TPD54 in PCa, and translational and
prospective studies of TPD54 as a therapeutic target in PCa are
also required.
References
|
1
|
Torre LA, Bray F, Siegel RL, Ferlay J,
Lortet-Tieulent J and Jemal A: Global cancer statistics, 2012. CA
Cancer J Clin. 65:87–108. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Parnes HL, House MG and Tangrea JA:
Prostate cancer prevention: Strategies for agent development. Curr
Opin Oncol. 25:242–251. 2013.PubMed/NCBI
|
|
3
|
Walz J, Joniau S, Chun FK, Isbarn H,
Jeldres C, Yossepowitch O, Chao-Yu H, Klein EA, Scardino PT,
Reuther A, et al: Pathological results and rates of treatment
failure in high-risk prostate cancer patients after radical
prostatectomy. BJU Int. 107:765–770. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Andriole GL, Crawford ED, Grubb RL III,
Buys SS, Chia D, Church TR, Fouad MN, Gelmann EP, Kvale PA, Reding
DJ, et al: Mortality results from a randomized prostate-cancer
screening trial. N Engl J Med. 360:1310–1319. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Shariat SF, Semjonow A, Lilja H, Savage C,
Vickers AJ and Bjartell A: Tumor markers in prostate cancer I:
Blood-based markers. Acta Oncol. 50 Suppl 1:S61–S75. 2011.
View Article : Google Scholar
|
|
6
|
Byrne JA, Nourse CR, Basset P and Gunning
P: Identification of homo- and heteromeric interactions between
members of the breast carcinoma-associated D52 protein family using
the yeast two-hybrid system. Oncogene. 16:873–881. 1998. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Boutros R, Fanayan S, Shehata M and Byrne
JA: The tumor protein D52 family: Many pieces, many puzzles.
Biochem Biophys Res Commun. 325:1115–1121. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Thomas DD, Frey CL, Messenger SW, August
BK and Groblewski GE: A role for tumor protein TPD52
phosphorylation in endo-membrane trafficking during cytokinesis.
Biochem Biophys Res Commun. 402:583–587. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Tennstedt P, Bölch C, Strobel G, Minner S,
Burkhardt L, Grob T, Masser S, Sauter G, Schlomm T and Simon R:
Patterns of TPD52 overexpression in multiple human solid tumor
types analyzed by quantitative PCR. Int J Oncol. 44:609–615. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mukudai Y, Kondo S, Fujita A, Yoshihama Y,
Shirota T and Shintani S: Tumor protein D54 is a negative regulator
of extracellular matrix-dependent migration and attachment in oral
squamous cell carcinoma-derived cell lines. Cell Oncol (Dordr).
36:233–245. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang Z, Sun J, Zhao Y, Guo W, Lv K and
Zhang Q: Lentivirus-mediated knockdown of tumor protein D52-like 2
inhibits glioma cell proliferation. Cell Mol Biol (Noisy-le-grand).
60:39–44. 2014.PubMed/NCBI
|
|
12
|
He Y, Chen F, Cai Y and Chen S: Knockdown
of tumor protein D52-like 2 induces cell growth inhibition and
apoptosis in oral squamous cell carcinoma. Cell Biol Int.
39:264–271. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Xu J, Wang W, Zhu Z, Wei Z, Yang D and Cai
Q: Tumor protein D52-like 2 accelerates gastric cancer cell
proliferation in vitro. Cancer Biother Radiopharm. 30:111–116.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yang M, Wang X, Jia J, Gao H, Chen P, Sha
X and Wu S: Tumor protein D52-like 2 contributes to proliferation
of breast cancer cells. Cancer Biother Radiopharm. 30:1–7. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Pan ZY, Yang Y, Pan H, Zhang J, Liu H,
Yang Y, Huang G, Yin L, Huang J and Zhou WP: Lentivirus-mediated
TPD52L2 depletion inhibits the proliferation of liver cancer cells
in vitro. Int J Clin Exp Med. 8:2334–2341. 2015.PubMed/NCBI
|
|
16
|
Zhao H, Logothetis CJ and Gorlov IP:
Usefulness of the top-scoring pairs of genes for prediction of
prostate cancer progression. Prostate Cancer Prostatic Dis.
13:252–259. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Barbaric D, Byth K, Dalla-Pozza L and
Byrne JA: Expression of tumor protein D52-like genes in childhood
leukemia at diagnosis: Clinical and sample considerations. Leuk
Res. 30:1355–1363. 2006. View Article : Google Scholar : PubMed/NCBI
|