Introduction
Human papillomaviruses (HPVs) are DNA viruses that
induce epithelial proliferative lesions of skin and mucosa in
humans (1). Based on their oncogenic
potential, HPV genotypes are classified as either high- or
low-risk. High-risk genotypes, including genotypes 16, 18, 31, 33,
35, 39, 45, 51, 52, 56, 58, 59 and 66, can stimulate the
development of cervical cancer and are associated with other types
of mucosal cancer, including anogenital, head and neck cancer.
HPV-16 and HPV-18 are the most prevalent high-risk genotypes that
induce cervical cancer (2).
Infections with low-risk genotypes, including HPV-6 and HPV-11,
cause benign or low-grade cervical tissue changes and genital warts
(3). The molecular mechanisms that
underlie the oncogenic potential of high-risk HPVs have been
extensively studied and it has been determined that the HPV-E6
protein acts as an oncoprotein that interacts with tumor suppressor
p53, leading to the transformation and dysregulated proliferation
of transfected cells (4). However, it
remains unknown why low-risk HPVs only induce benign warts and not
malignant transformation; thus, further studies are required to
elucidate this.
A previous study assessed the cellular localization
of high-risk HPV-16E6 and low-risk HPV-11E6 in 293T and MCF-7 cell
lines. High-risk HPV-16E6 was primarily located in the nucleus,
whereas low-risk HPV-11E6 was primarily located in the cytoplasm
(5). In accordance with this, the
present study aimed to investigate the cellular localization and
expression efficiency of low-risk HPV-6E6 in dendritic cells (DCs).
This approach may identify the mechanism that explains why low-risk
HPV is unable to induce malignant transformation.
Materials and methods
DC isolation and culture
DCs were generated from mouse bone marrow cells as
previously described (6). Bone marrow
cells were obtained from 4–6-week-old female C57BL/6 mice (Vital
River Laboratory Animal Technology Co., Ltd., Beijing, China) which
were bred in house, at Laboratory Animal Facility at the Chinese
Center for Disease Prevention and Control. A total of 158 mice were
used in the present study, with a weight range between 18–22 g.
Mice were kept in a controlled room with a 12/12 light/dark cycle
with access to food and water ad libitum and maintained at a
temperature (22±1°C) and humidity (20–23%). Cells were cultivated
in RPMI-1640 medium (Hyclone; GE Healthcare Life Sciences Logan,
UT, USA) supplemented with 10% fetal bovine serum (FBS; Thermo
Fisher Scientific Inc., Waltham, MA, USA) and
penicillin/streptomycin at 37°C and 5% CO2 for 3 h.
Following the removal of suspended cells, the remaining adherent
cells which had firmly adhered to the slides were washed with PBS,
in order to remove nonadherent cells without dislodging clusters of
developing dendritic cells. Following this, cells were cultured in
RPMI-1640 medium containing growth factors of 10 ng/ml recombinant
granulocyte macrophage colony-stimulating factor (Prospec-Tany
TechnoGene Ltd., East Brunswick, NJ, USA) and 2 ng/ml interleukin-4
(Prospec-Tany TechnoGene Ltd., East Brunswick, NJ, USA), and 10%
FBS, for 6 days with replacement of medium on days 3 and 5.
Immature DCs were harvested on day 6 for further experimentation.
All experiments involving animals were approved by the Chinese
Center for Disease Control and Prevention Laboratory Animal Welfare
& Ethics Committee (Dec 12, 2012; no. 2012-0022).
Construction of the expression vector
and transfection
The full-length HPV-6E6 sequence was amplified using
polymerase chain reaction (PCR) from the HPV type 6 complete
genome. The Q5 High-Fidelity DNA polymerase (New England BioLabs,
Inc., Ipswich, MA, USA) was used in this study. The PCR primer
sequences were used to amplify the full-length HPV-6E6 sequence:
Forward, 5′-GGAGAAAGTGCAAATGCCTCC-3′; and reverse,
5′-GGGTAACATGTCTTCCATGTC-3′. The PCR conditions were as follows: 5
min at 95°C, 30 cycles of 50 sec denaturation at 94°C, 30 sec
annealing at 61°C and extension at 72°C for 1 min. A final
extension step was performed at 72°C for 10 min. The products were
analyzed with 1% agarose gel electrophoresis, stained with 1
µg/µl ethidium bromide (Santa Cruz Biotechnology, Inc.,
Dallas, TX, USA) at room temperature for 30 min, and visualized
with the Molecular Imager FX (Bio-Rad Laboratories Inc., Hercules,
CA, USA). The HPV-6E6 sequence was cloned in frame within the C
terminus of the green fluorescent protein (GFP) at the Bgl
II and EcoR I sites of the polylinker region of the
mammalian expression vector pGFP (Clontech Laboratories Inc.,
Mountainview, CA, USA), producing a GFP-6E6 plasmid. DCs were
transiently transfected overnight with 1 µg/µl
plasmid, either pGFP-6E6 or pGFP using Lipofectamine®
2000 transfection reagent (Invitrogen; Thermo Fisher Scientific
Inc.) according to the manufacturer's protocol. The reagent: DNA
ratio was 2:1.
Live cell imaging using a confocal
microscope
A total of 2×105 cells/well were seeded
on glass coverslips and then transfected with either the pGFP-6E6
or pGFP plasmid following the aforementioned procedure. At 24 h
post-transfection, coverslips were mounted on modified glass slides
with 10% fetal calf serum (Gibco; Thermo Fisher Scientific, Inc.)
containing RPMI-1640 medium and immediately subjected to imaging.
Live cell images were obtained using Leica confocal microscopy at a
×400 magnification. The spectra were excited at 488 nm and emission
was detected at 507 nm.
Immunocytochemistry
DCs transfected with pGFP-6E6 and pGFP were seeded
on glass coverslips at a density of 2×105 cells/well.
Cells were then washed with PBS and fixed with 4% paraformaldehyde
for 10 min at room temperature. Then cells were permeabilized with
0.3% Triton X-100 (Sigma-Aldrich; Merck KGaA, Darmstadt, Germany)
for 30 min at room temperature. Cells were then blocked by
incubating with 1% bovine serum albumin (Sigma-Aldrich; Merck KGaA)
in PBS at room temperature for 30 min. Cells were then incubated
with a primary antibody against p53 (cat. no. 2524, 1:500, Cell
signaling Technology Inc., Danvers, MA, USA) overnight at 4°C. The
signal detection was performed using Cy3-conjugated goat anti-mouse
IgG secondary antibody (cat. no. C2181, 1:200 dilution,
Sigma-Aldrich, Wisconsin, USA), in a blocking solution (5% fat-free
milk and 0.1% Tween-20 in PBST) for 30 min at room temperature in
the dark. Cell images were examined using Leica TCS SMD FCS
confocal microscopy (Leica Microsystems GmbH, Wetzler, Germany) at
a magnification of ×400. The confocal microscopy measurements were
all performed with Leica SP8 software (Leica Microsystems GmbH,
Wetzler, Germany).
Analysis of apoptosis using DAPI
staining
To detect cell apoptosis, nuclear staining was
performed using 1 mg/ml DAPI at 37°C in the dark for 10 min. Cells
were washed twice with PBS and mounted on slides prior to analysis
with a CX41-32RFL fluorescence microscope (×200 magnification,
Olympus Corporation, Tokyo, Japan). Apoptotic cells were identified
by their morphology of large amounts of cell debris and by
fragmentation of their nuclei.
Analysis of apoptosis by flow
cytometry using Annexin V/propidium Iodide (PI) double
staining
Transfected DCs were harvested 24 h
post-transfection using trypsinization and apoptosis was analyzed
using an Annexin V-APC Apoptosis detection kit (Thermo Fisher
Scientific Inc., Waltham, MA, USA). A total of 1×106
cells/well in 100 µl binding buffer were stained with a mixture of
5 µl Annexin V-APC and 10 µl PI (final concentration, 1 µg/ml) and
incubated on ice for 10 min in the dark. Cells were analyzed using
a flow cytometer (FACSCalibur; BD Biosciences, Franklin Lakes, NJ,
USA) and data were processed using Cell Quest software v. 5.1 (BD
Biosciences). A minimum of 20,000 cells per sample was collected.
The results presented are expressed as percentage of apoptotic cell
number and data from three independent experiments are
presented.
Western blotting
For each sample, 1×106 cells were
collected by centrifugation (1,000 × g for 5 min, at 4°C), washed
once with ice cold PBS and lysed in 100 µl radioimmunoprecipitation
assay buffer (Complete Mini; Roche Diagnostics GmbH, Mannheim,
Germany). Protein concentration was determined using a
bicinchoninic acid assay (Pierce; Thermo Fisher Scientific Inc.).
Total proteins (30 µg) were separated on 12% SDS polyacrylamide
gels and transferred to polyvinylidene difluoride membranes
(Invitrogen; Thermo Fisher Scientific, Inc.,). Membranes were
blocked with 5% fat-free milk and 0.1% Tween20 in PBST for 1 h at
room temperature. Membranes were incubated with primary antibodies
anti-p53 (cat. no. 2524, 1:500, Cell Signaling Technology Inc.),
anti-BCL2 associated X apoptosis regulator (cat. no. ab32503,
1:1,000, Abcam, Cambridge, UK), anti-BCL2 homologous
antagonist/killer (cat. no. ab2371, 1:1,000, Abcam),
anti-cytochrome c (cat. no. ab13575, 1:50, Abcam) and anti-β-actin
(cat. no. sc130300, 1:1,000; Santa Cruz Biotechnology Inc.), at 4°C
overnight. Blots were counterstained with secondary antibodies,
horseradish peroxidase conjugated goat anti-mouse (1:1,000,
PA1-28887) or goat anti-rabbit IgG (1:1,000. YX-1598P; Pierce;
Thermo Fisher Scientific, Inc.), at room temperature for 30 min.
Proteins were visualized using an enhanced chemiluminescence kit
(GE Healthcare, Chicago, IL, USA).
Statistics
All data were expressed as the mean ± standard
deviation and analyzed using one-way analysis of variance for
multiple comparisons, and Student-Newman-Keuls post-hoc test was
used in this analysis. All analyses were performed using SPSS 19.0
software (IBM Corp., Armonk, NY, USA) and P<0.05 was considered
to indicate a statistically significant difference.
Results
Constructed GFP-6E6 expressing plasmid
and transfection of bone marrow-derived DCs
The coding region of HPV-6E6 was inserted within the
C terminus of the pEGFP-C1 vector, allowing E6 proteins to be
expressed as GFP-6E6 fusion proteins. The different locations of
protein in cells may be indicative of variations in function.
Therefore, it was important to start our research with the location
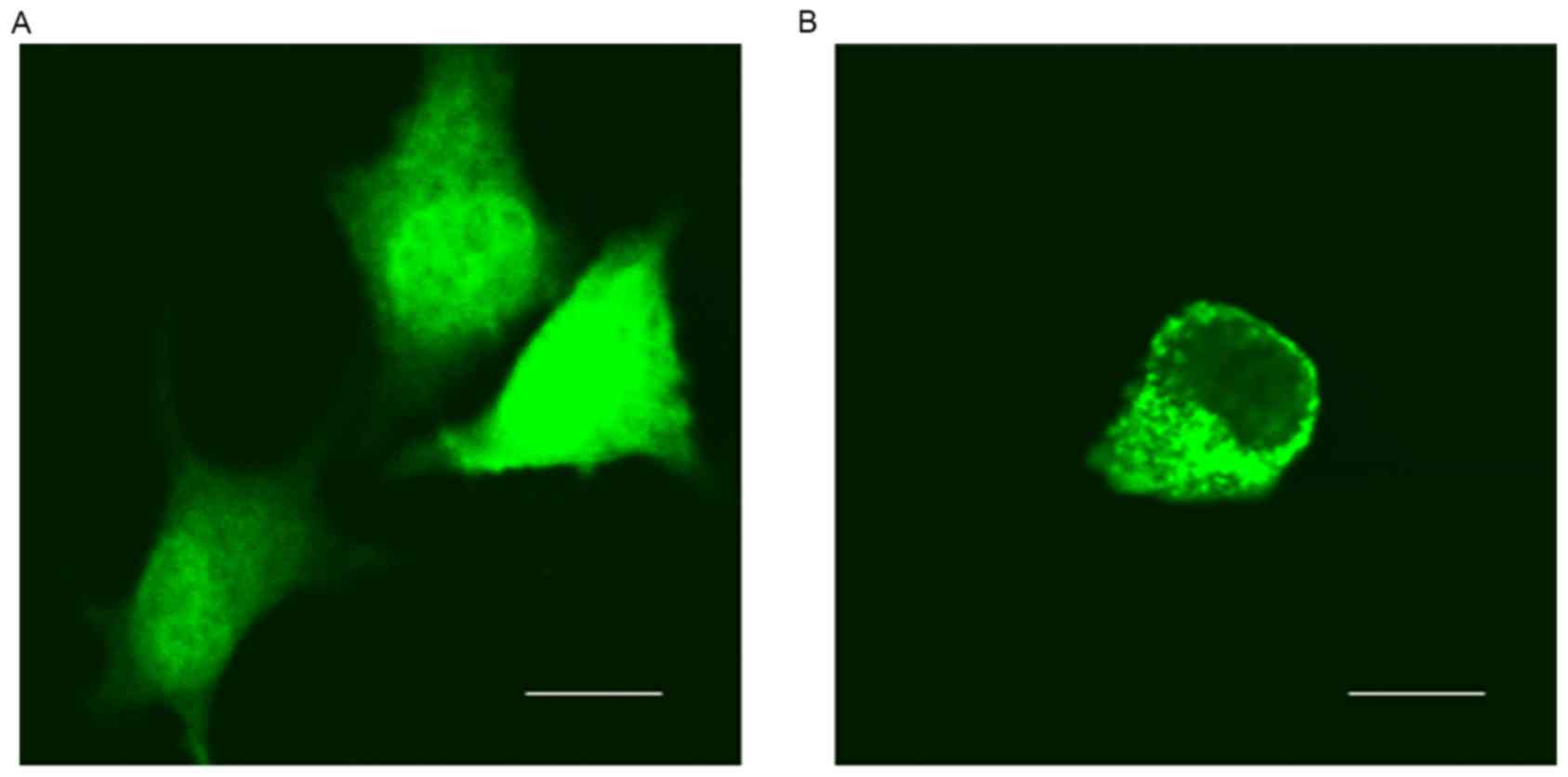
of GFP-6E6 in DCs. Using fluorescent microcopy, the subcellular
localization of GFP-6E6 and GFP proteins in DCs were determined.
GFP-6E6 was observed to be primarily located in the cytoplasm of
DCs, whereas the expression of GFP alone was present in the nuclei
and cytoplasm in DCs (Fig. 1).
Expression of low-risk HPV-6E6 in
DCs
GFP-6E6 fusion proteins may have low or high levels
of expression at different times, which may affect the distribution
of E6 (7). Therefore, the
localization and expression of GFP-6E6 between 6 and 72 h
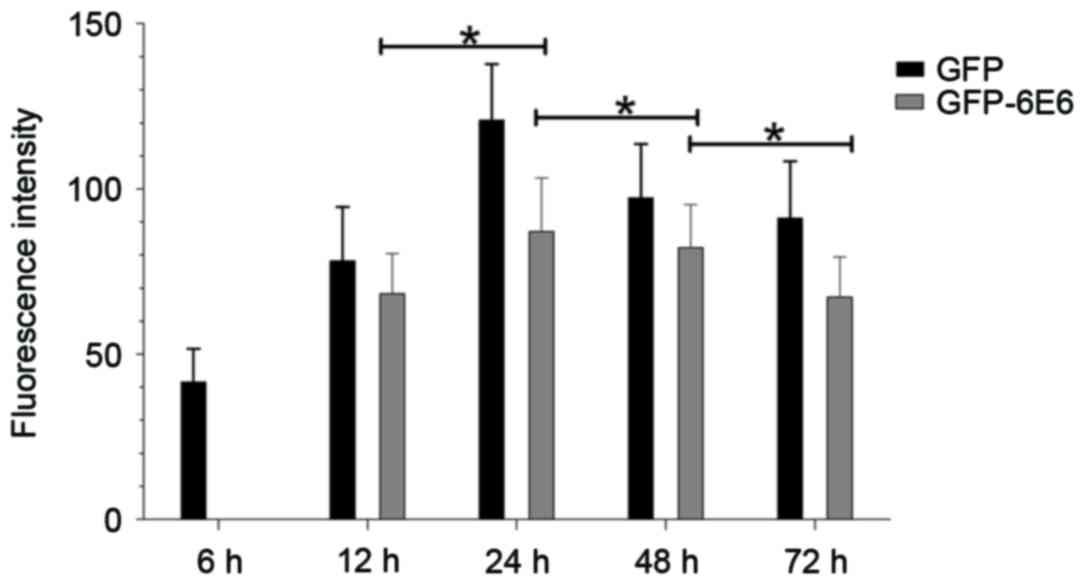
post-transfection was observed dynamically. The results indicated
that the GFP-6E6 protein is primarily expressed in the cytoplasm 12
h post-transfection. Its expression increased gradually and reached
its peak 24 h post-transfection (Fig.
2). GFP-6E6 expression then decreased gradually and was not
detected 1 week after transfection (data not shown). Notably,
during the experimental period, GPF-6E6 was persistently localized
in the cytoplasm (Fig. 2). The
expression of GFP alone was also assessed as a control. It
exhibited a diffused signal, as it was detected in both the nucleus
and cytoplasm from 6 h to 1 week post-transfection (data not
shown). The results revealed that GFP-6E6 is primarily located in
the cytoplasm of DCs, whereas the expression of GFP alone was
present in both the nuclei and cytoplasm in DCs. This indicated the
low risk HPV-6E6 trapped the expression of GFP in the cytoplasm of
DCs.
Low-risk HPV-6E6 induces apoptosis in
DCs
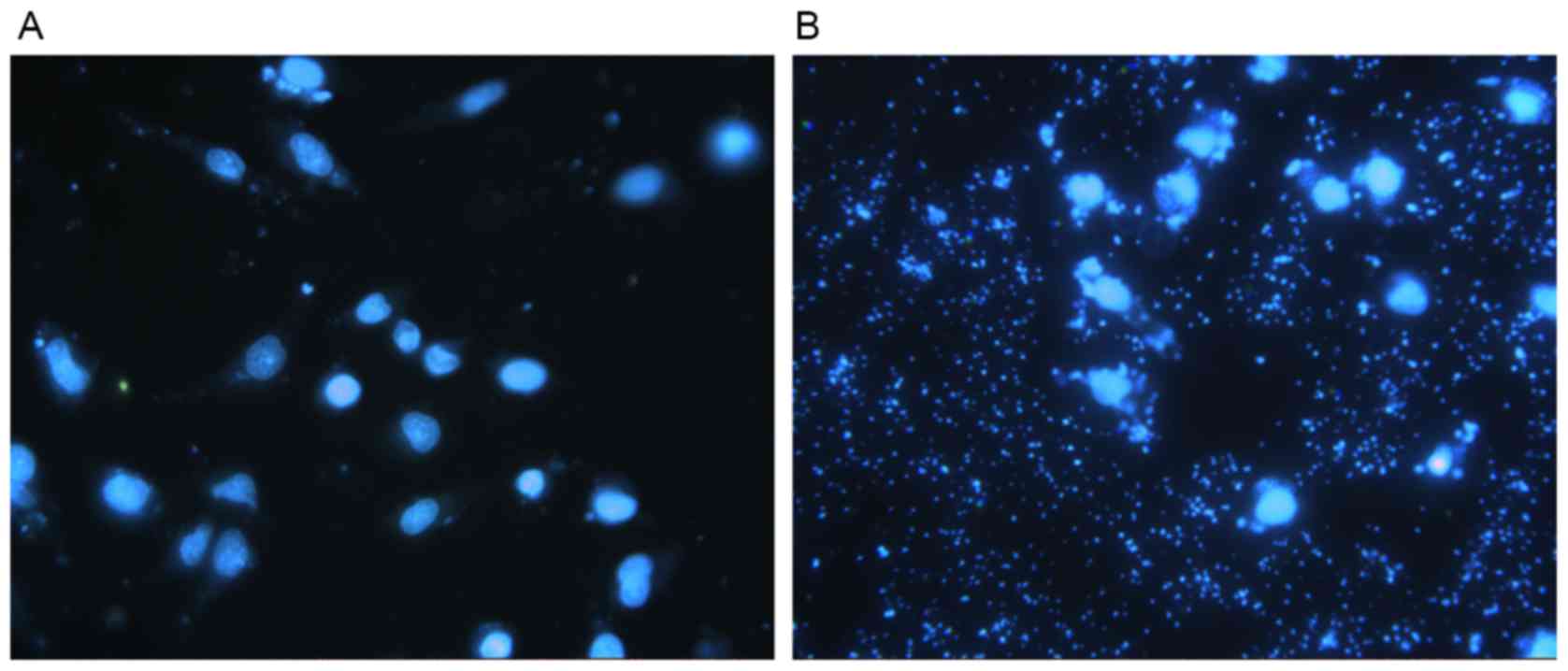
DAPI staining allowed for the observation of nuclear
morphology to detect apoptosis using a fluorescent microscope. DCs
transfected with pGFP-6E6 exhibited large amounts of cell debris
and condensed nuclei, indicating cell apoptosis, whereas the nuclei
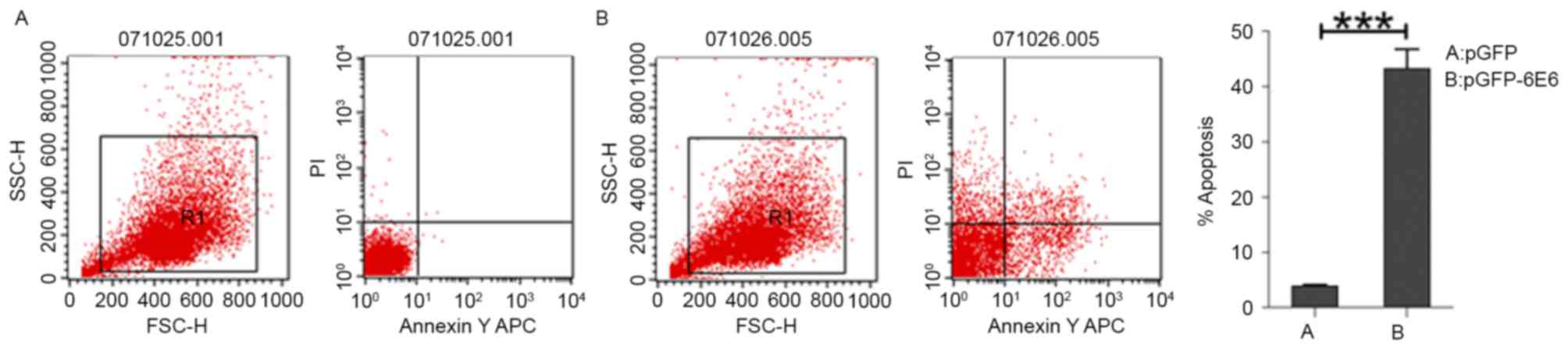
of DCs transfected with control pGFP remained intact (Fig. 3A and B). In addition, flow cytometry
detected marked apoptosis in DCs transfected with pGFP-6E6, but not
in DCs transfected with control pGFP (Fig. 4A and B).
Co-localization of GFP-6E6 and
p53
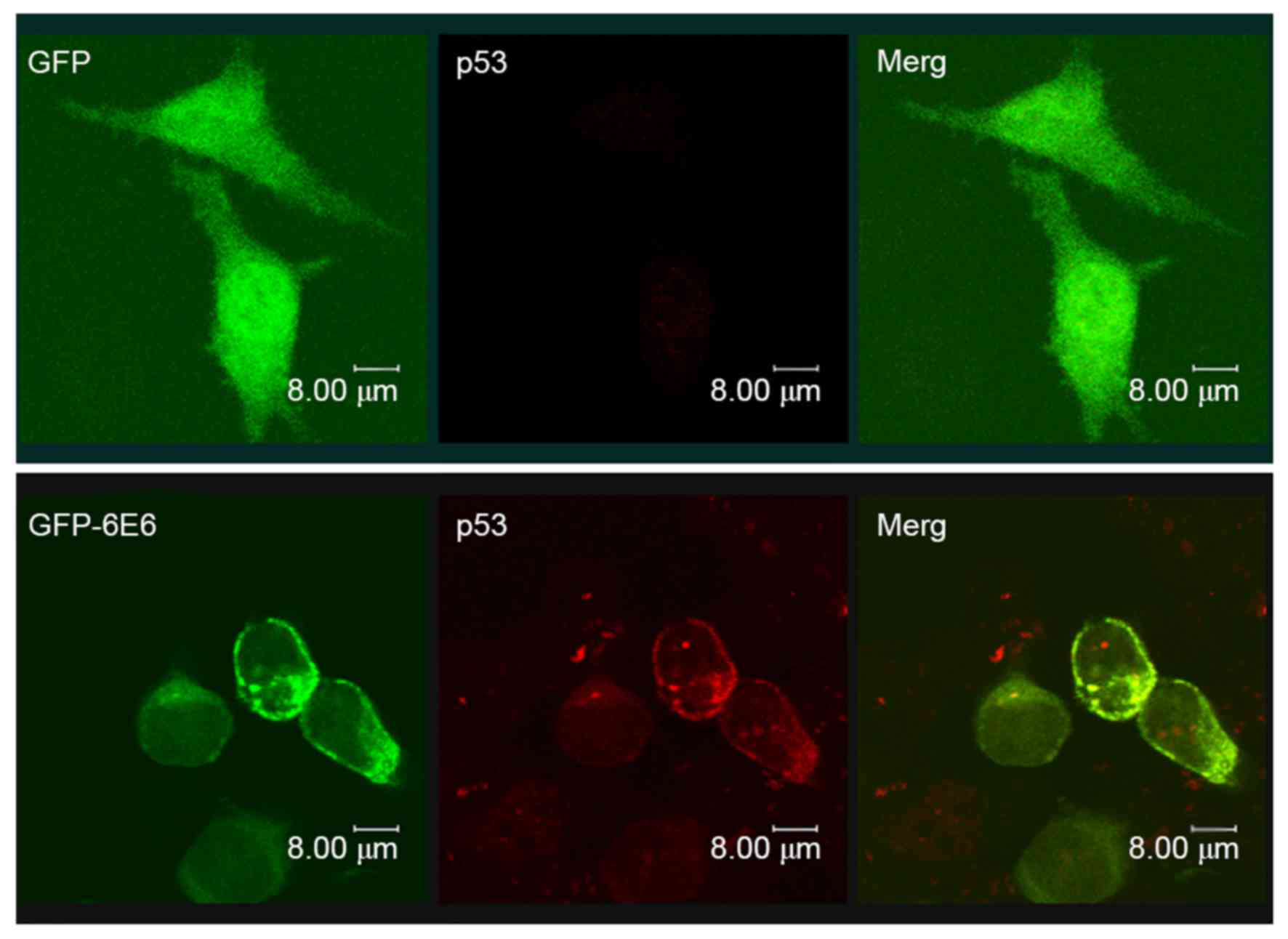
The expression of p53 in DCs transfected with either
pGFP-6E6 or pGFP plasmid were assessed using immunocytochemical
staining (Fig. 5). The results
revealed that there was a higher expression of p53 in DCs 24 h
post-transfection with pGFP-6E6, compared with cells transfected
with pGFP. More importantly, the p53 protein was co-localized with
GFP-6E6 in the cytoplasm, indicating the possibility of protein
interaction.
Expression of apoptosis-associated
proteins in DCs transfected with HPV-6E6
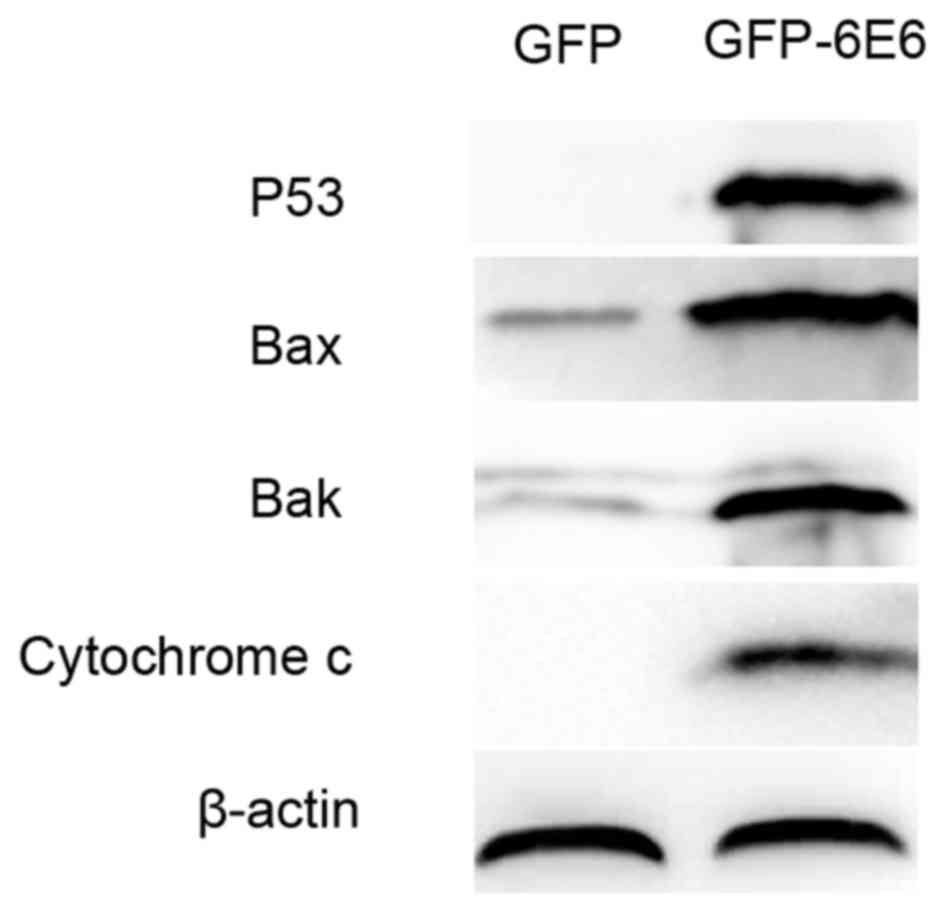
To further assess the apoptotic potential of
HPV-6E6, the expression of apoptosis-associated proteins in DCs
transfected with either pGFP-6E6 or control pGFP plasmids were
assessed using western blotting. The results revealed that there
was a higher expression of apoptosis-associated proteins, including
p53, Bax, Bak and cytochrome c, 24 h post-transfection in DCs
transfected with pGFP-6E6, compared with cells transfected with the
control plasmid (Fig. 6).
Discussion
DCs serve a crucial role in the initiation and
regulation of the anti-tumor immune response. DC vaccines loaded
with high-risk HPV-E6 have been assessed in animal tumor models and
clinical trials (8,9). However, the effects of the interaction
between low-risk HPV-6E6 and DCs has not yet been examined. The
present study aimed to determine how DCs respond to low-risk
HPV-6E6 and which biochemical mechanisms are responsible for
inducing apoptosis. The function of the E6 protein is partly
dependent on its location within the cell (10). However, previous studies investigating
the localization of HPV-E6 proteins have revealed contradictory
results, which may be due to low levels of endogenous E6 and the
poor reactivity of anti-E6 antibodies (11,12).
Therefore, the present study used the GFP fusion protein to
dynamically trace the traffic and location of low-risk HPV-6E6 in
bone marrow-derived DCs. The results revealed that HPV-6E6 is
primarily located in the cytoplasm of DCs. This was consistent with
the results of a study by Tao et al (11), which demonstrated that full-length
low-risk HPV-E6 proteins were distributed predominantly in the
cytoplasm in COS-1 cells.
The present study assessed the effect of low-risk
HPV-6E6 transfected into DCs. The results of DAPI nuclear staining
and apoptosis-based flow cytometry assays indicated that DCs
expressed low-risk HPV-6E6 and underwent apoptosis. It has been
demonstrated that the elimination of DCs may improve the chance of
effective viral colonization of the host body (13). However, DC apoptosis does not indicate
a weakness of the host immune system in the fight against
pathogens; it is one of the body's defense strategies (14). The quick death of DCs following the
induction of programmed cell death effectively protects against
viral propagation and spread to other cells. The present study
therefore assessed the molecular mechanism that may underlie this
observation. It was demonstrated that tumor suppressor p53 was
significantly upregulated in DCs that express HPV-6E6 and that
these proteins were co-localized in the cytoplasm. Therefore, it
was determined that the HPV-E6 protein interacts with the tumor
suppressor p53. High-risk HPV-E6 binds to the core of p53, a
process mediated by the E6 associated protein, which can stimulate
the degradation of p53 (15).
However, it has also been identified that low-risk HPV-E6 binds to
the C-terminal region of p53, which does not initiate p53
degradation (16). The results of the
present study support the latter observation and suggest that
low-risk HPV-E6 cannot degrade p53, which may be the primary reason
that infection with low-risk HPV does not lead to the development
of malignant cancer (17,18). Furthermore, DC exposure to low-risk
HPV-6E6 induces apoptosis via the upregulation of
apoptosis-associated proteins, including p53, Bax, Bak and
cytochrome c. This is supported by the results of a study by
Manjarrez et al (19) who
demonstrated that p53 levels were higher in papillomatosis than in
normal larynxes.
In conclusion, the present study revealed that low
risk HPV-6E6 is located in cytoplasm of bone marrow-derived DCs.
Low risk HPV-6E6 induces DC apoptosis and thus upregulates the
expression of apoptosis-associated proteins, particularly p53. The
results of the present study provide novel insights into the
pathogenesis of low risk HPV-6: E6 is unable to downregulate p53
and instead induces cell apoptosis rather than proliferation. This
may be one of the reasons why low-risk HPV does not induce
malignancy.
Acknowledgements
The present study was supported by the National
Natural Foundation of China (grant no. 81201273), the National High
Research and Development Program of China (863 Program; grant no.
2014AA021404) and the National Key Research and Development program
(grant no. 2016YFC1200701).
References
|
1
|
Zur Hausen H: Papillomaviruses in human
cancer. Cancer. 15:1692–1996. 1987.
|
|
2
|
Kumvongpin R, Jearanaikool P, Wilailuckana
C, Sae-Ung N, Prasongdee P, Daduang S, Wongsena M, Boonsiri P,
Kiatpathomchai W, Swangvaree SS, et al: High sensitivity,
loop-mediated isothermal amplification combined with colorimetric
gold-nanoparticle probes for visual detection of high risk human
papillomavirus genotypes 16 and 18. J Virol Methods. 234:90–95.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Shin TH, Pankhong P, Yan J, Sardesai NY
and Weiner DB: Induction of robust cellular immunity against HPV6
and HPV11 in mice by DNA vaccine encoding for E6/E7 antigen. Hum
Vaccin Immunother. 8:470–478. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
DasGupta T, Nweze EI, Yue H, Wang L, Jin
J, Ghosh SK, Kawsar HI, Zender C, Androphy EJ, Weinberg A, et al:
Human papillomavirus oncogenic E6 protein regulates human
β-defensin 3 (hBD3) expression via the tumor suppressor protein
p53. Oncotarget. 7:27430–27444. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Sun L, Zhang G, Lei T, Huang C, Song T and
Si L: Two different HPV-11E6 fusion proteins trap p53 in the
cytoplasm and induce apoptosis. Cancer Biol Ther. 7:1909–1915.
2008. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Liu Y, Chiriva-Internati M, Grizzi F,
Salati E, Roman JJ, Lim S and Hermonat PL: Rapid induction of
cytotoxic T-cell response against cervical cancer cells by human
papillomavirus type 16 E6 antigen gene delivery into human
dendritic cells by an adeno-associated virus vector. Cancer Gene
Ther. 8:948–957. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Filippova M, Evans W, Aragon R, Filippov
V, Williams VM, Hong L, Reeves ME and Duerksen-Hughes P: The small
splice variant of HPV16 E6, E6, reduces tumor formation in cervical
carcinoma xenografts. Virology 450–451. 1–164. 2014.
|
|
8
|
Shi GN, Zhang CN, Xu R, Niu JF, Song HJ,
Zhang XY, Wang WW, Wang YM, Li C, Wei XQ and Kong DL: Enhanced
antitumor immunity by targeting dendritic cells with tumor cell
lysate-loaded chitosan nanoparticles vaccine. Biomaterials.
113:191–202. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Baert T, Garg AD, Vindevogel E, VAN
Hoylandt A, Verbist G, Agostinis P, Vergote I and Coosemans AN: In
vitro generation of murine dendritic cells for cancer
immunotherapy: An optimized protocol. Anticancer Res. 36:5793–5801.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hiraiwa A, Kiyono T, Segwa K, Utsumi KR,
Ohashi M and Ishibashi M: Comparative study on E6 and E7 genes of
some cutaneous and genital papillomaviruses of human origin for
their ability to transform 3Y1 cells. Virology. 192:102–111. 1993.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Tao M, Kruhlak M, Xia S, Androphy E and
Zheng ZM: Signals that dictate nuclear localization of human
papillomavirus type 16 oncoprotein E6 in living cells. J Virol.
77:13232–13247. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Liang XH, Volkmann M, Klein R, Herman B
and Lockett SJ: Co-localization of the tumor-suppressor protein p53
and human papillomavirus E6 protein in human cervical carcinoma
cell lines. Oncogene. 8:2645–2652. 1993.PubMed/NCBI
|
|
13
|
Kubicka-Sierszen A and Grzegorczyk JŁ: The
influence of infectious factors on dendritic cell apoptosis. Arch
Med Sci. 11:1044–1051. 2015.PubMed/NCBI
|
|
14
|
Wang JJ, Li YF, Jin YY, Wang X and Chen
TX: Effects of Epstein-Barr virus on the development of dendritic
cells derived from cord blood monocytes: An essential role for
apoptosis. Braz J Infect Dis. 16:19–26. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Kehmeier E, Rühl H, Voland B, Stöppler MC,
Androphy E and Stöppler H: Cellular steady-state levels of ‘high
risk’ but not ‘low risk’ human papillomavirus (HPV) E6 proteins are
increased by inhibition of proteasome-dependent degradation
independent of their p53 and E6AP binding capabilities. Virology.
299:72–87. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Li X and Coffino P: High-risk human
papillomavirus E6 protein has two distinct binding sites within
p53, of which only one determines degradation. J Virol.
70:4509–4516. 1996.PubMed/NCBI
|
|
17
|
Martinez-Zapien D, Ruiz FX, Poirson J,
Mitschler A, Ramirez J, Forster A, Cousido-Siah A, Masson M, Vande
Pol S, Podjarny A, et al: Structure of the E6/E6AP/p53 complex
required for HPV-mediated degradation of p53. Nature. 28:541–545.
2016. View Article : Google Scholar
|
|
18
|
Huibregtse JM, Scheffner M and Howley PM:
Cloning and expression of the cDNA for E6-AP, a protein that
mediates the interaction of the human papillomavirus E6 oncoprotein
with p53. Mol Cell Biol. 13:775–784. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Manjarrez ME, Ocadiz R, Valle L, Pacheco
C, Marroquin A, De la Torre C, Selman M and Gariglio P: Detection
of human papillomavirus and relevant tumor suppressors and
oncoproteins in laryngeal tumors. Clin Cancer Res. 12:6946–6951.
2006. View Article : Google Scholar : PubMed/NCBI
|