Introduction
Breast cancer is known as the most frequent
malignancy among women worldwide, with an increasing incidence in
recent years (1). In the past
decades, the expression levels of estrogen receptor, progesterone
receptor and human epidermal growth factor receptor 2 have been
considered as the most important prognostic markers for the
invasiveness of breast tumor (2). The
glycosylated transmembrane protein cluster of differentiation
CD133, which is associated with shorter disease-free and overall
survival times, appears to be a novel promising biomarker for
prognosis (3). CD133 was observed to
present an association between its expression level and aggressive
cellular behavior, including resistance to chemotherapy and
radiotherapy in breast tumors, as well as hepatic and colon cancer
(4,5).
Notably, a wide range of studies has demonstrated that CD133
positively identifies cancer stem cells (6), indicating its association with the
stem-like subpopulation in breast tumors.
Cancer stem-like cells (CSCs) derived from breast
cancer are considered as the key initiator of breast tumor, and are
responsible for maintenance and metastasis. Functionally, CSCs are
defined as the subpopulation derived from breast cells, which
present the capacity to initiate a tumor in immunocompromised mice,
to renew themselves when passaged, and to differentiate into
non-self-renewing cells for formation of solid tumors (7). CSCs are also resistant to the effects of
radiotherapy, chemotherapy and endocrine therapy in breast tumors,
and consequently lead to tumor recurrence (8–10). Thus,
the CSC properties of self-renewal, multipotency and
chemotherapeutic resistance not only drive tumorigenesis, but also
provide resistant advantages for surviving the hostile environment
of the circulation, and proliferate in distal tissues (11).
Subsequent to the removal of breast tumors by
surgery, an increasing number of patients employ breast
reconstruction surgery for improving the quality of lives. However,
one of the major concerns regarding breast reconstruction is local
recurrence (12–15). Several essential factors are involved
in predicting the recurrence, including the subtypes and the
malignancy of breast tumors. Thus, more focus on the study of
stemness and malignancy of breast tumors was devoted by plastic
surgery and breast surgeons.
Metastasis-associated lung adenocarcinoma
transcript-1 (MALAT-1) is generally defined as a long non-coding
RNA (lncRNA) that consists of ~8,000 nt and is located on
chromosome 11q13 (16). It is
associated with malignancy in several different types of cancers,
including bladder, gallbladder, liver and gastric cancer (16). In breast tumor, MALAT-1 was identified
to be significantly downregulated, and thus caused the inhibition
of proliferation, colony formation, migration and invasion
(14). The underlying mechanisms are
possibly involved in inducing cell cycle arrest at G2/M, promoting
apoptosis, suppressing epithelial-mesenchymal transition (EMT) and
reducing the stem-like properties (15). The accumulating evidence indicates
that MALAT-1 acts as an oncogenic lncRNA through regulating
self-renewal capacity, which leads to malignancy in breast
tumors.
In the present study, it was revealed that MALAT-1
is upregulated in the CSC subpopulation in breast tumor cells, and
the epigenetic expression of MALAT-1 significantly reduced the
proportion of CSCs in MCF7. In addition, it was identified that
MALAT-1 regulates the proliferation, colony formation, migration
and invasion of CSCs in vitro. Therefore, the present study
demonstrated that MALAT-1 may serve as a novel biomarker for
predicting the malignancy of breast tumor and a potential
therapeutic target for breast tumor metastasis.
Materials and methods
Human breast cancer cells
MCF7 cells were previously bought from the American
Type Culture Collection (Manassas, VA, USA). Cells were maintained
in Dulbecco's modified Eagle's medium (DMEM; Gibco; Thermo Fisher
Scientific, Inc., Waltham, MA USA) supplemented with 10% fetal
bovine serum (FBS; Gibco; Thermo Fisher Scientific, Inc.) at 37°C
in a 5% CO2 incubator. The CSCs derived from MCF7 were
described previously (17). Single
cells were suspended and plated at 1,000 cells/cm2, and
the following treatments were added to the culture media: 100 ng/ml
recombinant interleukin-8 (R&D Systems, Inc., Minneapolis, MN,
USA), 100 nM SCH563705 (C-X-C chemokine receptor type 1/2
inhibitor; Merck KGaA, Darmstadt, Germany) or 1 µM lapatinib
(GlaxoSmithKline Plc., Brentford, UK). The medium was
half-refreshed every 3 days. After 21 days, spheres formed and were
collected for subsequent culture.
Design and cloning of short hairpin
RNA (shRNA) targeting MALAT-1
The lentivirus-encoded shRNA was employed for
MALAT-1-knockdown, which was synthesized by Shanghai ShengGong Co.,
Ltd. (Shanghai, China). The sequences of shRNA targeting to MALAT-1
was 5′-GGGCTTCTCTTAACATTTA-3′ and the scrambled control sequence
was 5′-TTCTCCGAACGTGTCACGT-3′. shRNAs were cloned into Plko.1
(GV248; Addgene, Inc., Cambridge, MA, USA) lentiviral vectors and
confirmed by sequencing. For each transfection,
Lipofectamine® 2000 (Thermo Fisher Scientific, Inc.) was
employed to introduce 0.8 µg plasmid into target cells.
RNA isolation and reverse
transcription-quantitative PCR (RT-qPCR)
Target cells were suspended using TRIzol®
reagent (Thermo Fisher Scientific, Inc.) according to the
manufacturer's protocol. Total RNA (500 ng) was reverse transcribed
into cDNA using the Transcriptional First Strand cDNA Synthesis kit
(Omega Bio-Tek, Inc., Norcross, GA, USA) according to the
manufacturer's protocol. The relative expression level of MALAT-1
to control GAPDH transcripts was detected by SYBR Green qPCR using
the ABI7500 Fast Real-time PCR system (Thermo Fisher Scientific,
Inc.). The primer sequences were synthesized as follows: MALAT-1
forward, 5′-AAAGCAAGGTCTCCCCACAAG-3′ and reverse,
5′-GGTCTGTGCTAGATCAAAAGGCA-3′; GAPDH forward,
5′-ACCACAGTCCATGCCATCAC-3′ and reverse, 5′-TCCACCCTGTTGCTGTA-3′.
lncRNA-regulator of reprogramming (ROR) and H19 were employed as
internal controls, the primer sequences were as follows: ROR
forward, 5′-TCCAAACACATCGCCACTCT-3′ and reverse,
5′-TCCTAGGCCATGAGGAGTCA-3′; H19 forward, 5′-GGAGACTAGGCCAGGTCTC-3′
and reverse, 5′-GCCCATGGTGTTCAAGAAGGC-3′. The qPCR amplification
was performed in triplicate reactions beginning at 98°C for 5 min,
followed by 35 cycles of 98°C for 30 sec, and 60°C for 50 sec.
Quantitative normalization of MALAT-1 cDNA was performed in each
sample using the expression of the GAPDH as an internal control.
The relative level of MALAT-1 transcripts to control GAPDH was
determined by the 2−ΔΔCq method (18).
Flow cytometric analysis of
CD133+
The singularized cells were incubated with CD133-PE
antibodies (Cell Signaling Technology, Inc., Danvers, MA, USA)
according to the manufacturer's protocol. Briefly,
~1×106 cells were centrifuged at 800 × g for 5 min at
4°C. The supernatant was removed and the pellet was washed three
times with ice-cold PBS. CD133 antibody (1 µl) was added, mixed
well and incubated for 10 min in the dark at 4°C. The stained cells
were then washed three times with ice-cold PBS and analyzed by flow
cytometry.
Self-renewal assay
To assess self-renewal capacity, single target cells
were seeded on 6-well plates at 500 cells/well and maintained for
21 days. The secondary spheres were counted and the self-renewal
was calculated by dividing the number of secondary spheres formed
by the number of primary spheres formed.
Western blot analysis
Total protein was extracted using RIPA buffer
(Guangzhou RiboBio Co., Ltd., Guangzhou, China) according to
manufacturer's protocol and determined using Pierce™ BCA
Protein Assay Kit (cat. no. 23225, Life Technologies, Grand Island,
NY, USA) according to the manufacturer's protocol. The primary
antibodies against GAPDH (cat. no. ab8245), sex-determining region
Y-box 2 (Sox-2; cat. no. ab171380), octamer-binding transcription
factor 4 (Oct4; cat. no. ab19857) and stem cell antigen-1 (Sca-1;
cat. no. ab51317) (Abcam, Cambridge, UK) were used in the present
study. All antibodies were diluted in PBS at 1:2,000 and membranes
were incubated for 2 h at room temperature. GAPDH was employed as
an internal control. Total protein lysate (50 µg) was separated by
8% SDS-PAGE. The bands were then transferred to polyvinylidene
fluoride membranes (EMD Millipore, Billerica, MA, USA) followed by
membrane blocking using 5% bovin serum albumin (Sigma Aldrich;
Merck KGaA) in PBS for 30 min at room temperature. Secondary goat
anti-rat antibody (cat. no. ab7010; 1:5,000; Abcam), goat
anti-mouse antibody (cat. no. ab97040; 1:5,000; Abcam) or goat
anti-rabbit antibody (cat. no. ab205718; 1:5,000; Abcam) was added
for an additional incubation at room temperature for 1 h and
immunoblotting was performed and visualized with a
chemiluminescence kit (Thermo Fisher Scientific, Inc.), according
to manufacturer's protocol.
Transwell assay
A Transwell assay without matrix gel was employed
for migration analysis, whereas a Transwell assay with matrix gel
was employed for invasion analysis. Chambers were covered with 80
µl Matrigel (BD Biosciences, Franklin Lakes, NJ, USA), or not,
diluted with 200 µl DMEM and incubated at 37°C for 6 h. A total of
2×104 cells were plated in the upper chambers, and 600
µl DMEM supplemented with 10% FBS was added to the lower chamber.
Following incubation at 37°C for 24 h, the cells were fixed with 4%
paraformaldehyde for 15 min at room temperature, and stained with
0.5% crystal violet for 30 min at room temperature, and imaged with
a fluorescence microscope using a magnification ×100.
Cell proliferation assay
Cell Counting Kit-8 (CCK-8; Sigma-Aldrich) was
employed for a simple and accurate cell viability assay. In the
assay, the WST-8 contained in the reagent was reduced to formazan
dye by a dehydrogenase enzyme of the cell mitochondria through
electron carrier 1-methoxy phenazinium methylsulfate. Singularized
cells at different time points were incubated with 10 µl of CCK-8
solution in separated wells and maintained in 37°C for 3 h. Cell
viability was measured as the absorbance at 450 nm with a
microplate reader (Synergy 2 Multi-Mode microplate reader; BioTek
Instruments, Inc., Winooski, VT, USA). The mean optical density
values from triplicate wells were used as the index of cell
viability.
Statistical analysis
All statistical analyses were performed using SPSS
software (version 16; SPSS, Inc., Chicago, IL, USA). Data are
presented as the mean ± standard deviation from three independent
experiments. Paired two tailed Student's t-tests and one-way
analysis of variance followed by Tukey's post hoc test were applied
for comparisons. P<0.05 was considered to indicate a
statistically significant difference.
Results
MALAT-1 is upregulated in CSCs and
positively regulated with the proportion of CSCs in breast cancer
cells
In order to investigate the roles of MALAT-1 in
regulating the stem cell-like phenotypes in breast cancer cells,
the expression levels of MALAT-1 were stably knocked down in
cultured MCF7 cells (MCF7-MALAT-1KD), as well as that of
control cells (MCF7-CtlKD). The knockdown efficacy in
the target cells was firstly examined by RT-qPCR, and the results
confirmed that MALAT-1 was significantly silenced compared with the
negative control MCF7-CtlKD (Fig. 1A). The alteration of CD133 and ALDH
expression, a putative CSC marker in cancer cells, was then
assessed, following MALAT-1-knockdown by flow cytometry. The
results revealed that the percentage of
CD133+/ALDH+ subpopulation was markedly
reduced compared with control cells (Fig.
1B). Collection of CSCs subpopulation from MCF7 was then
performed and involved into RT-qPCR for detecting the expression
level of MALAT-1 RNA (Fig. 1C). It
was revealed that, the subpopulation of CSCs exhibited
significantly higher MALAT-1 expression levels compared with the
overall MCF7 population (Fig. 1D).
For confirmation of the specific transcriptional alteration of
MALAT-1, lncRNA-regulator of reprogramming (ROR) and H19 were
employed as controls. As expected, no detectable change was
observed using these two lncRNAs in CSCs (Fig. 1D).
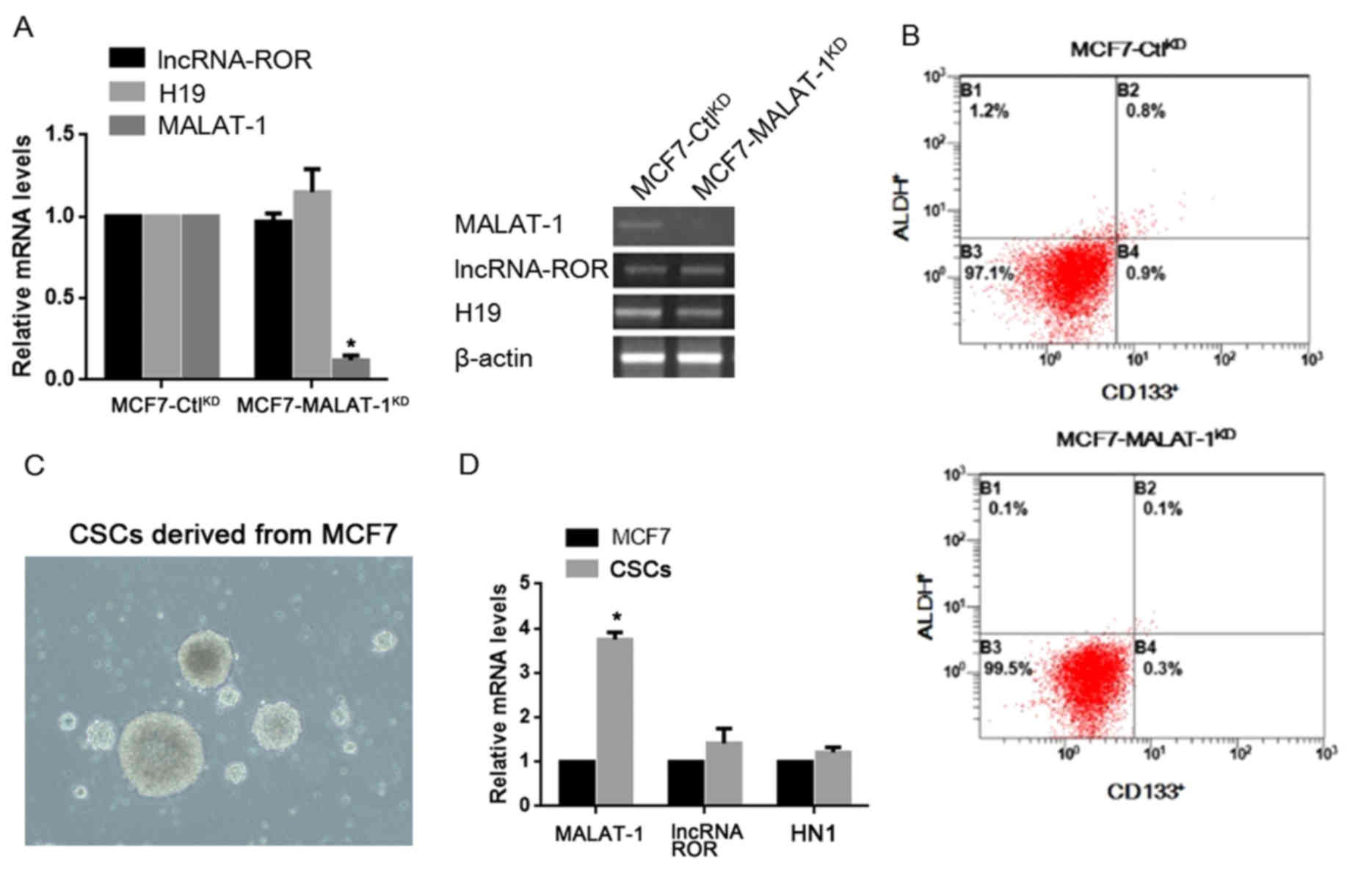 | Figure 1.Expression level of MALAT-1 in CSCs
and its effects on the proportion of CD133/ALDH positive
subpopulation. (A) RT-qPCR and semi-quantitative PCR were performed
for detecting lncRNA-ROR, H19 and MALAT-1. (B) CD133 and ALDH
staining was performed in MCF7-CtlKD and
MCF7-MALAT-1KD cells by flow cytometry. (C) Culture of
CSCs derived from MCF7 (magnification, ×40). (D) Detection of
MALAT-1, lncRNA-ROR and HN-1 in MCF7 and CSCs by RT-qPCR.
*P<0.05. MALAT-1, metastasis-associated lung adenocarcinoma
transcript-1; CSC, cancer stem cells; ALDH, aldehyde dehydrogenase;
RT-qPCR, reverse transcription-quantitative polymerase chain
reaction; lncRNA-ROR, long non-coding RNA-regulator of
reprogramming; CD133, cluster of differentiation 133; KD,
knockdown. |
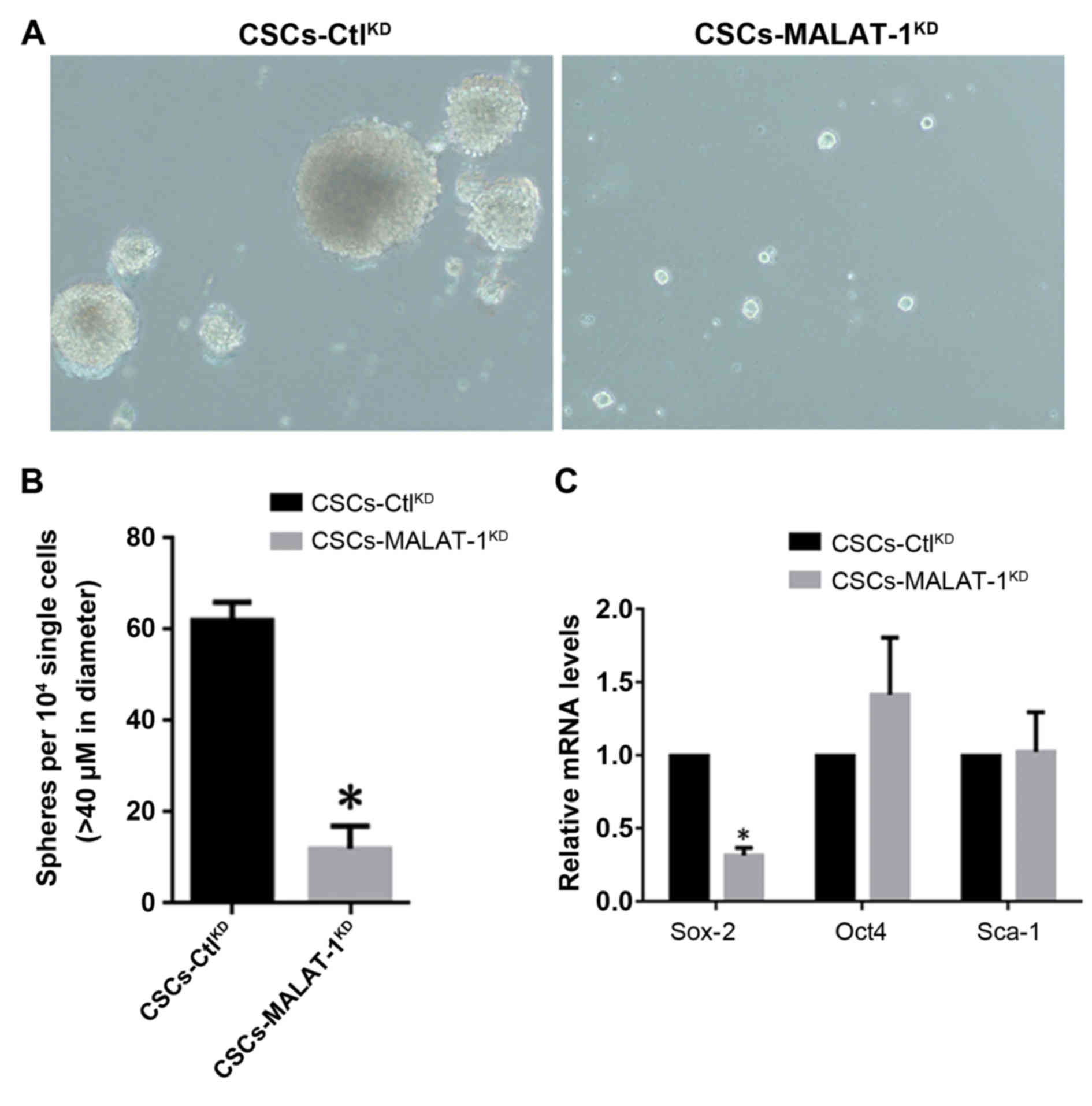
MALAT-1 promotes self-renewal capacity
in CSCs derived from MCF7
By considering the association of MALAT-1 RNA
expression level with the existence of CSCs derived from MCF7, an
in vitro sphere formation assay was performed to investigate
whether MALAT-1 functions in the self-renewal process of CSCs. The
CSCs derived from MCF7, which were MALAT-1-knockdown
(CSC-MALAT-1KD) or negative control
(CSC-CtlKD), were established by lentivirus infection.
It was shown that in CSC-MALAT-1KD, MALAT-1-knockdown
markedly reduced the formation of spheres and the diameter of
spheres was significantly smaller compared with the
CSC-CtlKD (Fig. 2A and B).
The expression levels of CSCs, Sox-2, Oct4 and Sca-1, which are
considered as the biomarkers of stem-like cells, were then
detected. The results demonstrated that Sox-2 exhibited a
significant decrease following knockdown of MALAT-1. However,
inconsistently, Oct4 and Sca-1 showed no significant change,
indicating that the alteration of Sox-2 was achieved specifically
(Fig. 2C).
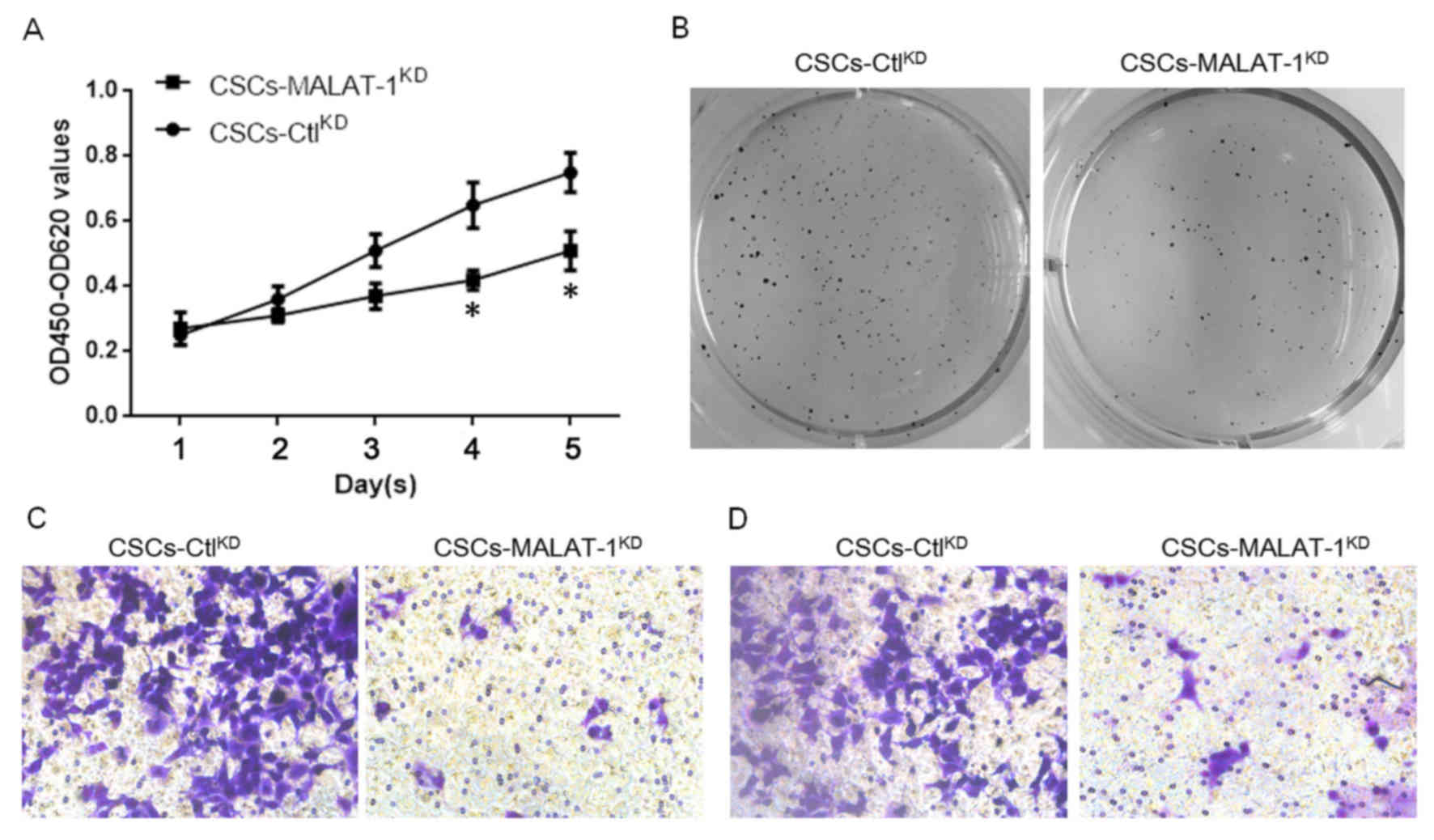
MALAT-1 regulates proliferation,
colony formation, migration and invasion in CSCs derived from
MCF7
The effects of MALAT-1 on the physiological
processes of CSCs derived from MCF7 were then examined. In order to
evaluate its effect on cell proliferation, the CCK-8 assay was
employed. The results demonstrated that knockdown of MALAT-1
significantly decreased the proliferation of CSCs following 4 days
(Fig. 3A). Consistent with the
proliferation, the results of colony formation assay on soft agar
of CSCs also indicated that MALAT-1 expression promoted the colony
formation of CSCs derived from MCF7 (Fig.
3B). Due to a previous study demonstrating that CSCs interact
with angiogenesis (19), the ability
of MALAT-1 to regulate the migration and invasion of CSCs was
examined. A Transwell assay without matrix gel was employed for
migration analysis, whereas a Transwell assay with matrix gel was
employed for invasion analysis. The results demonstrated that, as
expected, knockdown of MALAT-1 markedly inhibited the migration and
invasion ability (Fig. 3C and D).
Discussion
In the past few years, lncRNAs were revealed as
important components of the gene regulatory network that may
perform critical roles in regulating several physiological
processes, including the survival and self-renewal of CSCs
(1,20). Several mechanisms of function of
lncRNAs in stem cell biology have been uncovered. lncRNA-ROR
suppresses the stimulation of p53 under stress and thus inhibits
the expression by activated p53 (2,21).
Notably, lncRNA-ROR includes several miRNA binding sites, including
miR-145, which specifically targets to Oct4, Nanog and Sox-2, and
thus positively regulates their mRNA and protein levels (3,22). Further
studies have confirmed that neutralization of miR-145 by lncRNA-ROR
promotes embryonic stem cell self-renewal (3,22). The
same mechanism was also identified, in that lncRNA H19 interacts
with p53, disturbs its signaling and also acts as a miRNA sponge to
let-7 (4,23). MALAT-1, which serves as an oncogenic
lncRNA, has been shown to be responsible for the malignant
behaviors in several types of human tumors (5,24),
although the mechanisms of its regulatory roles in CSC-like
properties were largely unknown. Among these genes that are
aberrantly expressed in CSCs derived from different types of human
tumors, Oct4, Nanog, BMI1 proto-oncogene polycomb ring finger,
proto-oncogene c-Myc, β-catenin and Sox-2 were significantly
decreased in response to knockdown of MALAT-1 (6,25). Taken
together, lncRNAs may perform critical roles in regulating the
maintenance of CSC-like capacity.
RT-qPCR was performed in order to identify the
expression profile of lncRNA-ROR, H19 and MALAT-1 in the breast
cancer MCF7 cell line, and its CSC subpopulation identified as
CD133-positive. It was revealed that MALAT-1, but not of lncRNA-ROR
or H19 was significantly upregulated in CSCs compared with MCF7
cells. Following MALAT-1-knockdown, the subpopulation of
CD133-positive significantly decreased. These results indicated
that MALAT-1 expression level may positively regulate the
expression of CD133 or the portion of CSC subpopulation in MCF7
cells in an uncertain mechanism. Next, the present study provided
evidence that MALAT-1 expression is able to promote the
self-renewal capacity of cells in vitro. Accumulating
evidence has demonstrated that CSCs interact with metastasis and
recurrence (7,25). It is reported that CD44 expression in
breast and pancreatic tumors present enhanced metastatic capacities
(8,9,26,27). Thus, the effects of MALAT-1 on the
migration and invasion of CSCs were demonstrated by Transwell
assay. The results demonstrated that MALAT-1 positively regulates
the migration and invasion of CSCs.
The expression level of MALAT-1 appears to be
critical in regulating the physiological processes of CSCs, and
also of a major population of tumor cells. Compared with adjacent
tissues, in lung (10,28), liver (11,29) and
prostate (12,30) cancer, MALAT-1 has been revealed to be
upregulated, and exhibit a tumor-promoting function. The data in
the current study demonstrated that, in the CSC subpopulation,
MALAT-1 expression was significantly higher compared with that in
the MCF7 cells. However, further studies are required to elucidate
whether the expression level of MALAT-1 is the key factor for
inducing tumorigenesis via the promotion of CSC formation. Taken
together, the results of the present study demonstrated the
potential function of long non-coding RNA MALAT-1 in maintaining
the stem phenotype of CSC from breast cancer, which indicates it as
a potential therapeutic target for treating breast cancer by
targeting to CSCs.
References
|
1
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
von Minckwitz G, Untch M, Nüesch E, Loibl
S, Kaufmann M, Kümmel S, Fasching PA, Eiermann W, Blohmer JU, Costa
SD, et al: Impact of treatment characteristics on response of
different breast cancer phenotypes: Pooled analysis of the German
neo-adjuvant chemotherapy trials. Breast Cancer Res Treat.
125:145–156. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Brugnoli F, Grassilli S, Piazzi M, Palomba
M, Nika E, Bavelloni A, Capitani S and Bertagnolo V: In triple
negative breast tumor cells, PLC-β2 promotes the conversion of
CD133 high to CD133 low phenotype and reduces the CD133-related
invasiveness. Mol Cancer. 12:1652013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Smith LM, Nesterova A, Ryan MC, Duniho S,
Jonas M, Anderson M, Zabinski RF, Sutherland MK, Gerber HP, Van
Orden KL, et al: CD133/prominin-1 is a potential therapeutic target
for antibody-drug conjugates in hepatocellular and gastric cancers.
Br J Cancer. 99:100–109. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Pohl A, El-Khoueiry A, Yang D, Zhang W,
Lurje G, Ning Y, Winder T, Hu-Lieskoven S, Iqbal S, Danenberg KD,
et al: Pharmacogenetic profiling of CD133 is associated with
response rate (RR) and progression-free survival (PFS) in patients
with metastatic colorectal cancer (mCRC), treated with
bevacizumab-based chemotherapy. Pharmacogenomics J. 13:173–180.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Mizrak D, Brittan M and Alison M: CD133:
Molecule of the moment. J Pathol. 214:3–9. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
McDermott SP and Wicha MS: Targeting
breast cancer stem cells. Mol Oncol. 4:404–419. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Phillips TM, McBride WH and Pajonk F: The
response of CD24(−/low)/CD44+ breast cancer-initiating cells to
radiation. J Natl Cancer Inst. 98:1777–1785. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li X, Lewis MT, Huang J, Gutierrez C,
Osborne CK, Wu MF, Hilsenbeck SG, Pavlick A, Zhang X, Chamness GC,
et al: Intrinsic resistance of tumorigenic breast cancer cells to
chemotherapy. J Natl Cancer Inst. 100:672–679. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Creighton CJ, Massarweh S, Huang S,
Tsimelzon A, Hilsenbeck SG, Osborne CK, Shou J, Malorni L and
Schiff R: Development of resistance to targeted therapies
transforms the clinically associated molecular profile subtype of
breast tumor xenografts. Cancer Res. 68:7493–7501. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Dontu G, Abdallah WM, Foley JM, Jackson
KW, Clarke MF, Kawamura MJ and Wicha MS: In vitro propagation and
transcriptional profiling of human mammary stem/progenitor cells.
Genes Dev. 17:1253–1270. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Lim W, Ko BS, Kim HJ, Lee JW, Eom JS, Son
BH, Lee TJ and Ahn SH: Oncological safety of skin sparing
mastectomy followed by immediate reconstruction for locally
advanced breast cancer. J Surg Oncol. 102:39–42. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Medina-Franco H, Vasconez LO, Fix RJ,
Heslin MJ, Beenken SW, Bland KI and Urist MM: Factors associated
with local recurrence after skin-sparing mastectomy and immediate
breast reconstruction for invasive breast cancer. Ann Surg.
235:814–819. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Ren D, Li H, Li R, Sun J, Guo P, Han H,
Yang Y and Li J: Novel insight into MALAT-1 in cancer: Therapeutic
targets and clinical applications. Oncol Lett. 11:1621–1630. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zhao Z, Chen C, Liu Y and Wu C:
17β-estradiol treatment inhibits breast cell proliferation,
migration and invasion by decreasing MALAT-1 RNA level. Biochem
Biophys Res Commun. 445:388–393. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Bonasio R and Shiekhattar R: Regulation of
transcription by long noncoding RNAs. Annu Rev Genet. 48:433–455.
2014. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Harrison H, Farnie G, Howell SJ, Rock RE,
Stylianou S, Brennan KR, Bundred NJ and Clarke RB: Regulation of
breast cancer stem cell activity by signaling through the Notch4
receptor. Cancer Res. 70:709–718. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Zhao Y, Bao Q, Renner A, Camaj P, Eichhorn
M, Ischenko I, Angele M, Kleespies A, Jauch KW and Bruns C: Cancer
stem cells and angiogenesis. Int J Dev Biol. 55:477–482. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ng JH and Ng HH: LincRNAs join the
pluripotency alliance. Nat Genet. 42:1035–1036. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhang A, Zhou N, Huang J, Liu Q, Fukuda K,
Ma D, Lu Z, Bai C, Watabe K and Mo YY: The human long non-coding
RNA-RoR is a p53 repressor in response to DNA damage. Cell Res.
23:340–350. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wang Y, Xu Z, Jiang J, Xu C, Kang J, Xiao
L, Wu M, Xiong J, Guo X and Liu H: Endogenous miRNA sponge
lincRNA-RoR regulates Oct4, Nanog and Sox2 in human embryonic stem
cell self-renewal. Dev Cell. 25:69–80. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kallen AN, Zhou XB, Xu J, Qiao C, Ma J,
Yan L, Lu L, Liu C, Yi JS, Zhang H, et al: The imprinted H19 lncRNA
antagonizes let-7 microRNAs. Mol Cell. 52:101–112. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Gutschner T, Hämmerle M and Diederichs S:
MALAT1-a paradigm for long noncoding RNA function in cancer. J Mol
Med (Berl). 91:791–801. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Jiao F, Hu H, Han T, Yuan C and Wang L,
Jin Z, Guo Z and Wang L: Long noncoding RNA MALAT-1 enhances stem
cell-like phenotypes in pancreatic cancer cells. Int J Mol Sci.
16:6677–6693. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Al-Hajj M, Wicha MS, Benito-Hernandez A,
Morrison SJ and Clarke MF: Prospective identification of
tumorigenic breast cancer cells. Proc Natl Acad Sci USA. 100:pp.
3983–3988. 2003; View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Hermann PC, Huber SL, Herrler T, Aicher A,
Ellwart JW, Guba M, Bruns CJ and Heeschen C: Distinct populations
of cancer stem cells determine tumor growth and metastatic activity
in human pancreatic cancer. Cell Stem Cell. 1:313–323. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Schmidt LH, Spieker T, Koschmieder S,
Schäffers S, Humberg J, Jungen D, Bulk E, Hascher A, Wittmer D,
Marra A, et al: The long noncoding MALAT-1 RNA indicates a poor
prognosis in non-small cell lung cancer and induces migration and
tumor growth. J Thorac Oncol. 6:1984–1992. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lai MC, Yang Z, Zhou L, Zhu QQ, Xie HY,
Zhang F, Wu LM, Chen LM and Zheng SS: Long non-coding RNA MALAT-1
overexpression predicts tumor recurrence of hepatocellular
carcinoma after liver transplantation. Med Oncol. 29:1810–1816.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Ren S, Wang F, Shen J, Sun Y, Xu W, Lu J,
Wei M, Xu C, Wu C, Zhang Z, et al: Long non-coding RNA metastasis
associated in lung adenocarcinoma transcript 1 derived miniRNA as a
novel plasma-based biomarker for diagnosing prostate cancer. Eur J
Cancer. 49:2949–2959. 2013. View Article : Google Scholar : PubMed/NCBI
|