Introduction
Gliomas are the most frequent type of malignant
tumor in central nervous system, leading to significant mortality
worldwide annually (1,2). It was classified into low-grade gliomas
(grade II, e.g., infiltrative astrocytomas and oligodendrogliomas)
and high-grade gliomas (grade III and IV, e.g., anaplastic gliomas
and glioblastomas) according to the World Health Organization
grading system (3). The current
therapeutic strategy to glioma consists of surgical resection,
radiotherapy, chemotherapy and antiangiogenesis. However, the
prognosis of the patients with glioma is still poor in spite of
these interventions. Only one third of patients with glioblastoma
(GBM) survival for a year and the 5-year survival rate for grade
III gliomas is 27% (4). The epidermal
growth factor receptor (EGFR) plays a key role in proliferation and
pathogenesis of multiple cancer types. Expression of EGFR
correlates with poor prognosis and radioresistance in glioma
(5).
EGFR amplification and overexpression have been
identified in ~50% of GBM patients. EGFR overexpression is often
accompanied by the expression of a mutant EGFR named as EGFRvIII
(EGFR type III, ΔEGFR), which is expressed in 30% of GBM tumors
(6,7).
EGFRvIII is generated from a genomic deletion involving exons 2–7
of the EGFR gene, which results in deletion of the extracellular
ligand-binding domain. It is unable to bind ligand, yet
constitutively activated in a ligand-independent manner leading to
overproliferation, survival, resistance to treatment and
invasiveness of GBM cells (8,9). Both of the classical subtypes classified
by the TCGA working group and the EM subgroup classified by the
CGGA working group showed that the characteristics of EGFR
activation associated with amplification and mutation displayed the
worst prognosis (10–12).
Centrosomal protein 55 (CEP55) is the latest found
member in the centrosomal relative protein family, which
participates in the regulation of the cell cycle. CEP55 is located
in the centrosome in interphase cells and is recruited into the
midbody during cytokinesis (13). In
addition, CEP55 is required for midbody structure and the
completion of cytokinesis at the terminal stage (14). Overexpressing of CEP55 were found in
several human tumors and various tumor cell lines (15–17).
However, the relationship between EGFRvIII and CEP55 in
proliferation regulation of glioma cells and the effect of CEP55 on
these malignant features of GBM is poorly understood. Therefore, we
detected the expression of CEP55 in human glioma tissues and
investigated the proliferation of glioblastoma U251 cells after
transfected with EGFRvIII and CEP55 siRNA.
Materials and methods
Tissue samples and cells line
Glioma tissues of 40 patients undergoing resection
of lesion were collected from the department of pathology in the
Xinqiao hospital from 2010 to 2016. Tumor samples were diagnosed
according to the WHO classification of central nervous system
tumors. Forty patients with glioma were newly diagnosed and didn't
received chemotherapy or radiotherapy. Twenty gliomas were
classified as low-grade glioma (WHO I and II) and others were
high-grade glioma (WHO III and IV). Ten non-tumor brain tissue were
obtained from patients with epilepsy undergoing surgical treatment
in our hospital. All patients provided written informed consent
according to the research proposals approved by the ethical
committee of the Xinqiao Hospital.
Human glioblastoma U251 cells were cultured in
Dulbecco's modified Eagle's medium (DMEM; Gibco; Thermo Fisher
Scientific, Inc., Waltham, MA, USA) supplemented with 10% fetal
bovine serum (FBS), 50 IU/ml benzyl penicillin G potassium, and 100
µg/ml streptomycin sulfate in a humidified incubator at 37°C and 5%
CO2. U251 glioma cells were purchased from Riken Cell
Bank (Tsukuba, Japan).
TCGA data analysis
Gene expression data from TCGA was analyzed in
Firebrowse (http://firebrowse.org/). Input
‘CEP55’ in View Expression Profile box; click ‘Filter on’ and
choose ‘GBMLGG’; submit. The Kaplan Meier Plot was analyzed in web
(https://xenabrowser.net/heatmap/#).
Firstly, choose ‘Visualization’ at the top of the web. Then, choose
‘TCGA lower grade glioma and glioblastoma (GBMLGG)’ in ‘Cohort’;
choose ‘gene expression RNAseq (polyA+ Illumina HiSeq)’ in ‘samples
in’; click ‘+Date’, ‘gene expression RNAseq’, ‘gene expression
RNAseq (polyA+ Illumina HiSeq)’ in turn; click ‘next’; input CEP55
in Genes box; click ‘Done’. Finally, click ‘Column menu (Inverted
triangle symbol)’ and choose ‘Kaplan Meier plot’.
Western-blot analysis
For western blot analyses, protein was harvested
from cells plated to 70–80% confluence. Cell lysates were prepared
using a RIPA buffer (25 mM Tris-HCl, 150 mM NaCl, 1% NP-40, 1%
sodium deoxycholate, and 0.1% sodium dodecyl sulfate; pH 7.6).
Protease inhibitors were added prior to use. The protein extracts
were loaded, size-fractionated by SDS-polyacrylamide gel
electrophoresis and transferred to PVDF membranes (Bio-Rad
Laboratories, Inc., Hercules, CA, USA). After blocking, the
membranes were incubated with the specific first antibodies (Abcam,
Cambridge, UK) in dilution buffer at 4°C overnight. The blots were
incubated with horseradish peroxidase (HRP)-conjugated secondary
antibodies (1:5,000, anti-rabbit/anti-mouse; Boster Biological
Technology Co., Ltd., Wuhan, China) at room temperature for 2 h and
then chemiluminescence signals were detected using a UVI gel
imaging system camera.
Quantitative real-time PCR
Total RNA was extracted from frozen tissues and cell
lines using TRIzol reagent (Invitrogen, Carlsbad, CA, USA)
according to the manufacturer's instructions and was reverse
transcribed into cDNA using PrimeScript RT reagent (Takara Bio
Inc., Shiga, Japan). The reaction system (20 µl) contained the
corresponding cDNA. The CEP55 primers used in quantitative
real-time PCR were as follows: forward primer of
5′-TTGGAACAACAGATGCAGGC-3′ and reverse primer of
5′-GAGTGCAGCAGTGGGACTTT-3′. GAPDH, used as an internal control, was
amplified with forward primer 5′-TGGACTCCACGACGTACTCAG-3′ and
reverse primer 5′-CGGGAAGCTTGTCATCAATGGAA-3′. All procedures were
performed in triplicate.
RNAi and transfection
To further identify the role of CEP55 in tumor and
the relationship of CEP55 and EGFRvIII, the CEP55 knockdown
lentiviral vector (siCEP55) was constructed (Shanghai GeneChem Co.,
Ltd., Shanghai, China). A GFP lentiviral vector was used as
negative control (NC). The day before transfection, cells were
seeded in 24-well plates at a density of 50,000 cells/well. The
lentiviruses were transfected according to the manufacturer's
instruction with MOI=10, and stably transfection cells were
selected by puromycin (6 µg/ml). All lentiviral vectors expressed
GFP and Puromycin, which enabled us to select stably transfection
cells. The siRNA sequences are as following:
5′-GTGGGAAAGGAAAGCTGAC-3′. The high expression of EGFRvIII
lentiviral vector was also constructed (Shanghai GeneChem Co.,
Ltd.). The sequences resulted from NCBI. The following are the NCBI
reference sequence: NM_001346941.1. A RFP lentiviral vector was
used as negative control.
Proliferation assay of U251 cells
The Cell Counting Kit-8 (CCK-8)
assay
U251 cells (3,000 cells/well) were seeded into
96-well plates in 100 µl complete medium. The CCK-8 (Dojindo
Laboratories, Kumamoto, Japan) was used to measure cell viability
according to the manufacturer's instructions. The plates were
incubated for 7 days. The number of viable cells was assessed by
measurement of the spectrophotometric absorbance at 450 nm with a
microplate reader iMARK (Bio-Rad Laboratories, Inc.).
EdU assay
Cells that had undergone different interventions
were seeded in 96-well plates at a density of 3×103 cells/well. EdU
assay was done following manufacturer's instructions after
culturing for 48 h Cell. The proliferation was assessed by
calculating the percentage of EdU-positive cells in all cells.
Statistical analysis
Statistical analysis for TCGA was described above.
SPSS software program (version 18.0; SPSS Inc., Chicago, IL, USA)
and Graphpad Prism 5 software program (Graphpad Software, Inc., San
Diego, CA, USA) were used for statistical analyses. All results
were presented as mean ± standard deviation. The horizontal bars in
histograms represented mean values. The one-way ANOVA was used to
analyze the differences between groups for in vivo or in
vitro studies. If the variance was homogeneous, Tukey's test
was used for the post-hoc test; If not, Dunnett's test was used.
P<0.05 was considered to indicate a statistically significant
difference.
Results
Expression of CEP55 increased in human
glioma cells and related to survival of patients with glioma
CEP55 played an important role in proliferation of
cells and overexpressed in several human tumors. The mRNA and
protein of CEP55 were detected by qPCR and western blot analysis in
40 cases of glioma tissues and 10 cases of normal brain tissues.
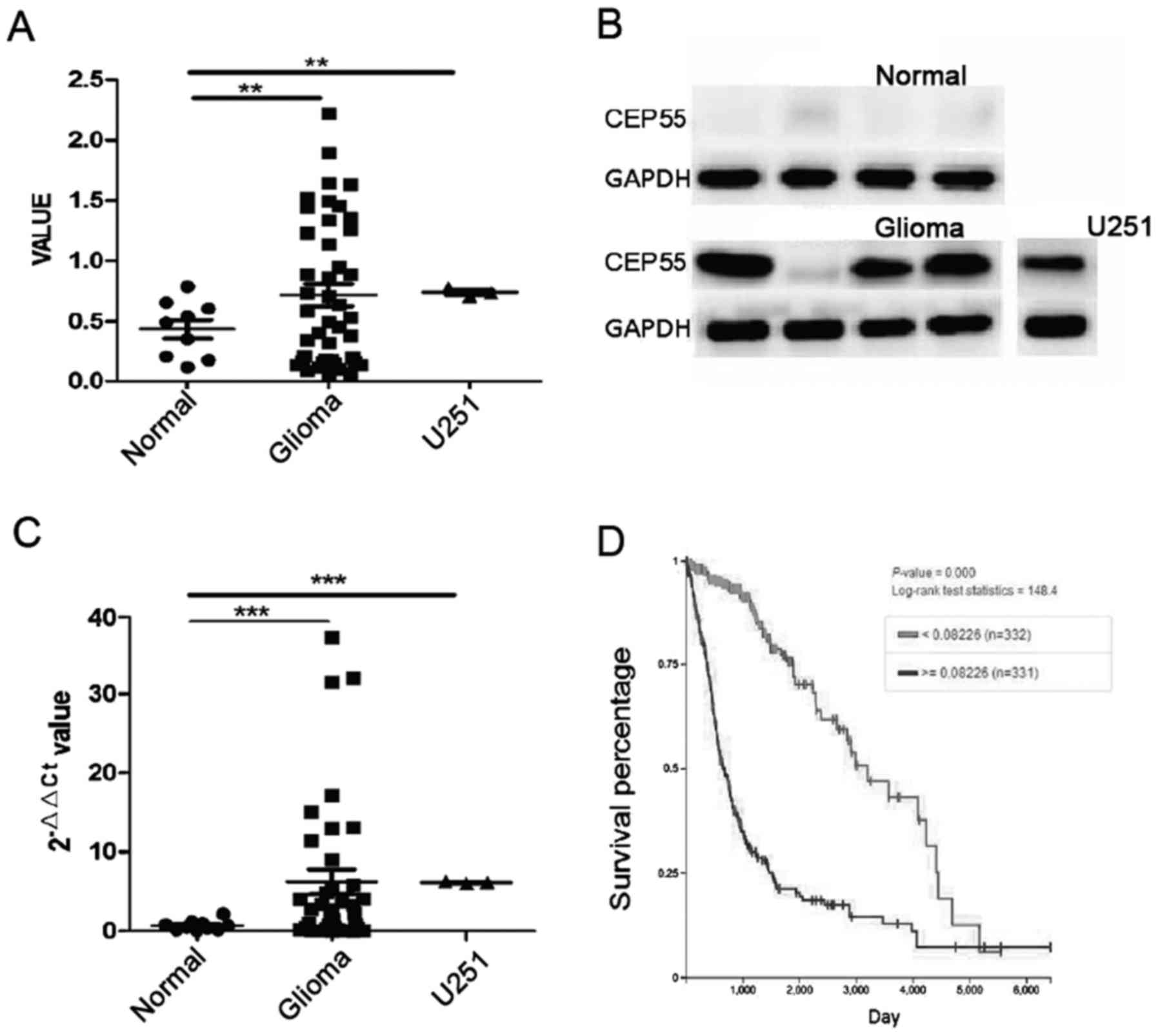
The mRNA of CEP55 in glioma tissues increased significantly than it
in normal brain tissues (P<0.01) (Fig.
1A). The result of western blot analysis showed that CEP55
proteins increased in 40 cases of glioma tissues accompany to
improving of CEP55 mRNA. The level of CEP55 proteins in glioma was
7 times higher than it in normal brain tissues (P<0.001)
(Fig. 1B and C). There was
overexpression of CEP55 in GBM U251 cells too, compared with normal
brain tissues (Fig. 1A-C). Our
results of qPCR and western blot analysis confirmed that expression
of CEP55 in glioma cells rocket up markedly than it in brain
tissues.
To discover the value of CEP55 overexpression in
glioma cells, we investigated the relationship of CEP55 expression
and survival time in 663 cases of patients with glioma in The
Cancer Genome Atlas (TCGA) database. The patients with glioma were
classified to overexpression of CEP55 group (n=332) and low
expression of CEP55 group (n=331) according to the threshold value
(>=0.00226 and <0.00226) of CEP55 mRNA. The median overall
survival (OS) of patients were 21.5 months in overexpression group
and 107.1 months in low expression group. The result showed that
expression of CEP55 was significantly associated to survival of
patient with glioma and overexpression of CEP55 mean shorter
survival time to patient with glioma (P<0.001) (Fig. 1D).
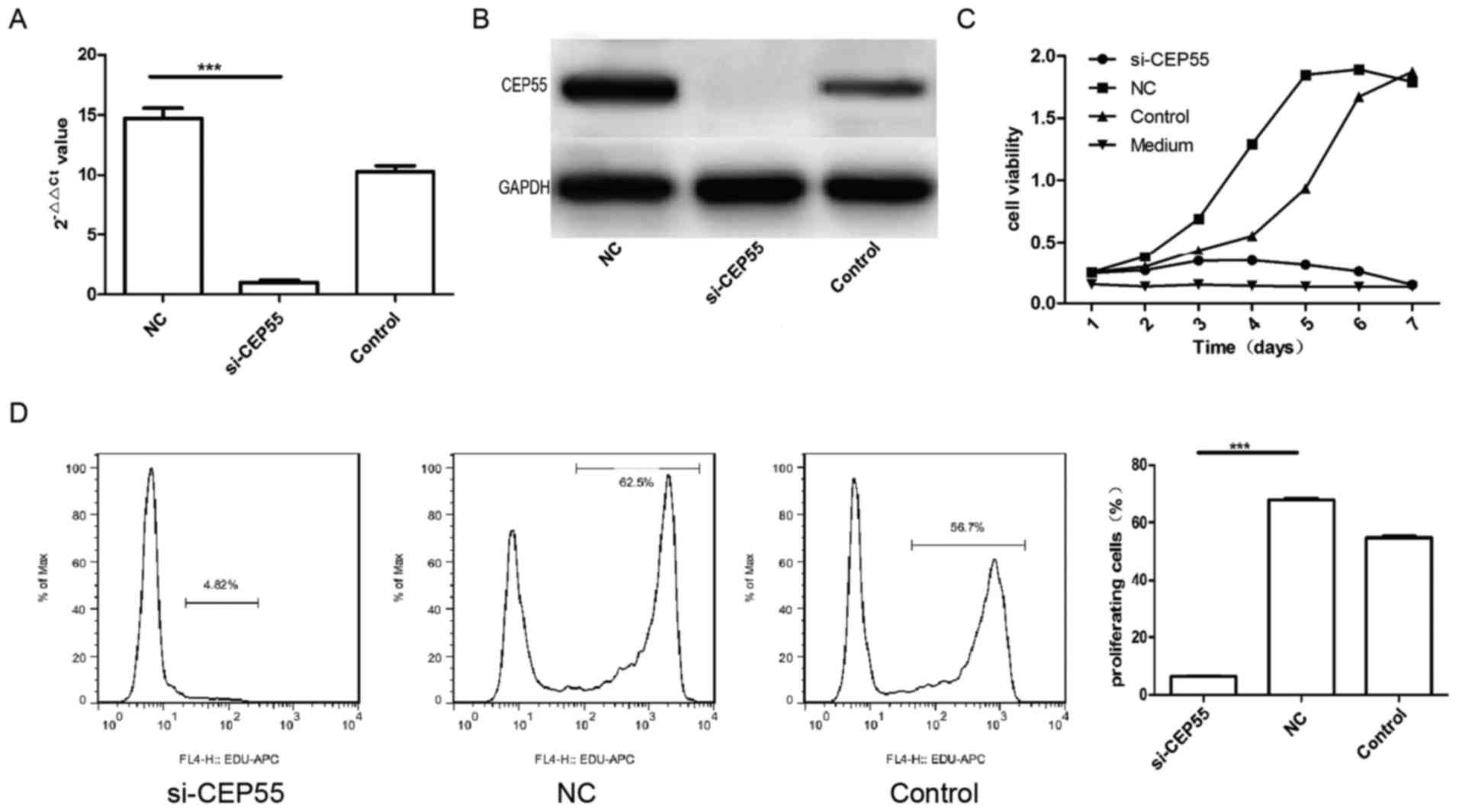
Proliferation of U251 cells was
suppressed by CEP55 RNAi
CEP55 acted as a member of the centrosomal
participated in mitosis of cell. Then, we supposed that expression
of CEP55 was related to proliferation of glioma cells. To survey
the effect of CEP55 on proliferation in glioma cells, the
expression of CEP55 was knockdown with RNAi mediated by lentiviral
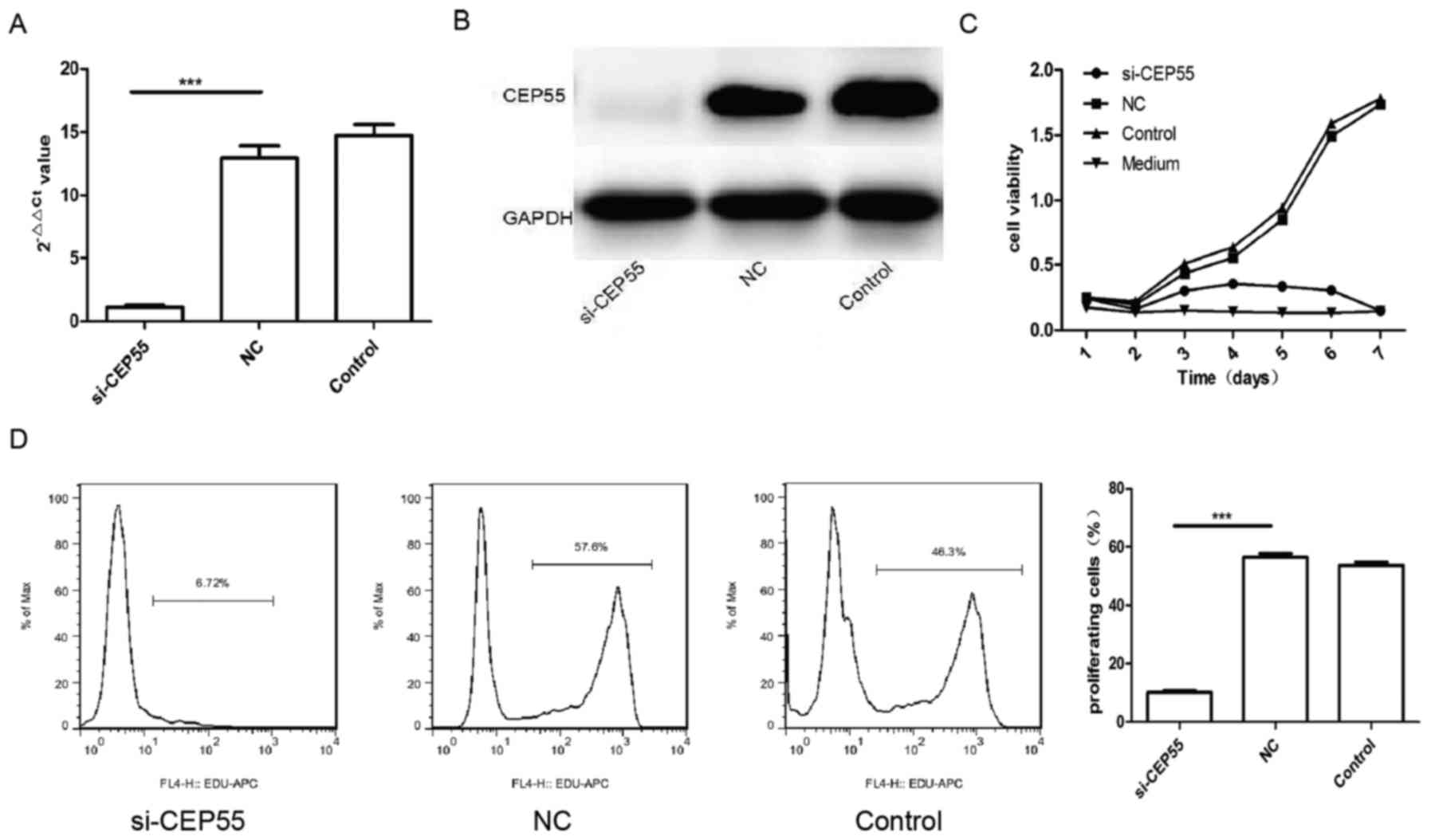
vector in GBM U251 cells. The mRNA and protein of CEP55 in U251
cells decreased remarkably assayed by qPCR and western blot
analysis after the cells infected by CEP55 siRNA lentiviral
(Fig. 2A and B). The proliferation of
three groups of U251 cells (control, NC and si-CEP55) were analyzed
with CCK-8 kit and EdU. The result of CCK-8 showed that the
proliferation of U251 cells with CEP55 knockdown dropped
significantly than the cells without CEP55 intervened. The
proliferation curve of U251 cells with CEP55 RNAi became flat
similar to the medium (Fig. 2C).
There were 57.6 and 46.3% cells being reduplication phase
separately in NC group and control group of U251 cells labeled with
EdU. But only 6.72% cells were being proliferation phase in
si-CEP55 group of U251 cells detected with flow cytometry. The rate
of U251 cells being proliferation in si-CEP55 group was lower
markedly than it in NC group and control group (Fig. 2D). The result demonstrated that CEP55
RNAi was able to induce the inhibition of proliferation in U251
cells.
EGFRvIII promoted the proliferation
and expression of CEP55 in U251 cells
CEP55 is a conserved gene to preventing genetic
damage and its protein acts as a regulator of cell cycle
progression ensuring proper abscission, the final stage of
cytokinesis (18). Therefore, the
expression of CEP55 might be regulated by other driver gene. The
EGFRvIII is one frequent driver gene of GBM cells and plays an
important role of proliferation in GBM cells. Then, EGFRvIII gene
were transfected into U251 cells by lentiviral vector and the
expression of EGFRvIII in U251 cells were detected with qPCR and
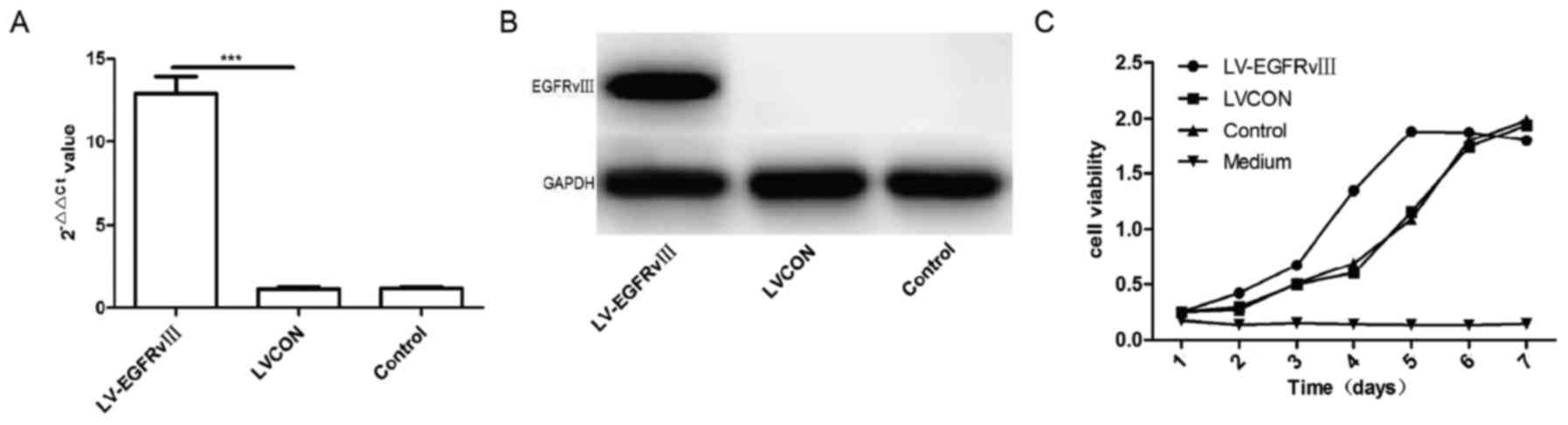
Western-blot. The results of qPCR and western blot analysis
verified that the mRNA and proteins of EGFRvIII increased
significantly in U251 cells infected by EGFRvIII gene lentiviral
vector compared with the cells infected by control lentiviral
vector group (LVCON) and control group (Fig. 3A and B). CCK-8 kit was used to analyze
the proliferation of U251 cells after transduced with EGFRvIII
gene. The results of curve showed that proliferation of U251
transduced with EGFRvIII gene increased considerably compared to
the cells of LVCON group and control group (P<0.001) (Fig. 3C). Expression of EGFRvIII was able to
promote the proliferation of U251 cells.
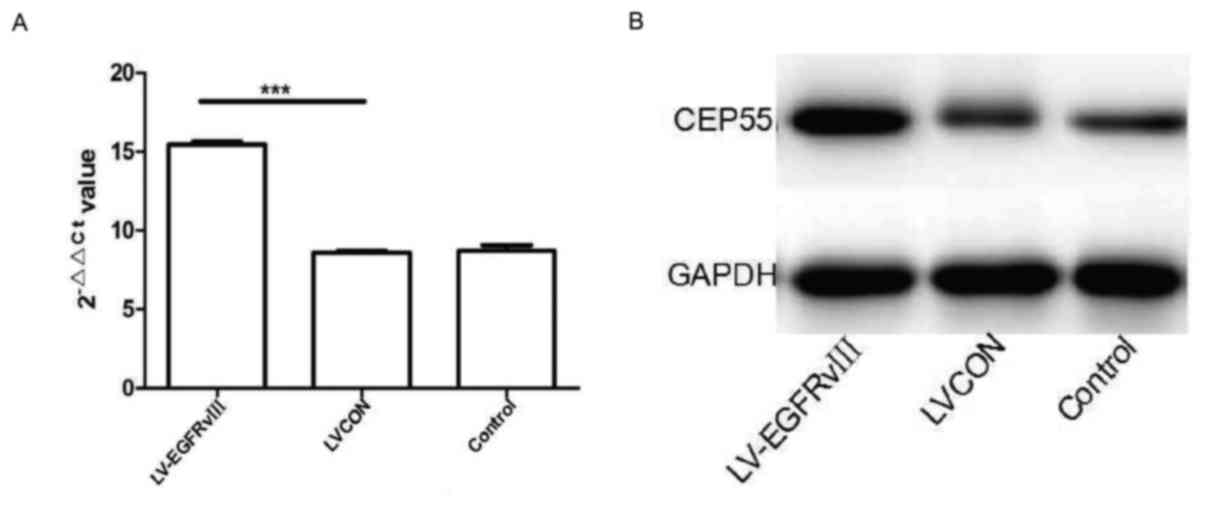
Meanwhile, the qPCR and western blot analysis were
performed to measuring the mRNA and protein of CEP55 in U251 cells
transduced with EGFRvIII gene. The level of CEP55 mRNA was higher
in U251 cells of EGFRvIII group than in cells of LVCON group and
control group. After EGFRvIII gene transduced and expressed in U251
cells, the protein of CEP55 increased drastically compared to the
cells of control group (Fig. 4A and
B). The results of qPCR and western blot analysis revealed that
expression of EGFRvIII promoted the proliferation and expression of
CEP55 in GBM U251 cells.
CEP55 RNAi inhibited the proliferation
induced by EGFRvIII in U251 cells
To detect the relationship of the proliferation
promoted by EGFRvIII and expression of CEP55 in U251 cells, the
expression of CEP55 was suppressed by RNAi in U251 cells transduced
with EGFRvIII. The mRNA and protein of CEP55 assayed by qPCR and
western blot analysis were reduced significantly in U251 cells with
EGFRvIII overexpressing after it transduced with CEP55 siRNA
(Fig. 5A and B). The curve of growth
was analyzed with CCK-8 in U251 cells with EGFRvIII overexpressing
and CEP55 siRNA transduced. The results indicated that the
proliferation of U251 cells declined markedly after the expression
of CEP55 was inhibited by RNAi (Fig.
5C). The proliferation of U251 cells with EGFRvIII
overexpressing and CEP55 siRNA transduced was also measured by flow
cytometry stained with EDU. The rate of proliferation in U251 cells
expressing EGFRvIII declined to 4.82% after the expression of CEP55
was suppressed with CEP55 siRNA, and the rates of proliferation in
U251 cells of NC group and control group were 62.5 and 56.7%
respective (Fig. 5D). The result
implied that the proliferation of U251 cells induced by EGFRvIII
was inhibited by CEP55 RNAi and EGFRvIII promoted proliferation in
U251 cells through regulating CEP55 overexpression.
Discussion
CEP55 was known as a centrosome and midbody
associated protein which might cooperate with members of endosomal
sorting complex required for transport machinery to allow
abscission (19,20). Soon after, overexpression of CEP55 was
found in lung adenocarcinoma, gastric carcinoma and hepatocellular
carcinoma, where it promote cell proliferation and invasion
(17,21,22). It
was among the top 70 most highly overexpressed genes in an analysis
across 12 cancer types including lung, lymphoma, medulloblastoma
and breast, and was identified in prognostic signatures for these
cancers (18,23). In this study, we found that CEP55
expression increased significantly in glioma and GBM U251 cells
than in normal brain tissue. The level of CEP55 expression was
significantly associated to survival of patient with glioma through
analyzing CEP55 mRNA and survival in 663 patients from TCGA
database. These results implied that CEP55 may play an important
role on proliferation or invasion of glioma cells. To observe the
effect of CEP55 overexpression on glioma cells, expression of CEP55
were inhibited by RNAi in GBM U251 cells in this study. The
proliferation of U251 cells was markedly suppressed after CEP55
knockdown. This result was confirmed in Wang et al study,
which provided evidence of inhibiting cell proliferation and
inducing cell apoptosis by knockdown of CEP55 in glioma cells
(24).
CEP55 expresses in various cancers and is barely
detectable in normal tissues except for testis and thymus (24). Its overexpression and activation was
regulated via aberrantly upregulated PI3K/AKT pathways and Forkhead
box protein M1 in lung cancer, hepatocellular carcinoma and
head-neck squamous cell carcinoma (17,21,25). In
glioma, the EGFR and its mutations contributed to tumorigenesis and
biology via various ways. EGFR amplification and mutation were
highly frequent genetic change in gliomas, which resulted in
expression of EGFRvIII and overexpression of EGFR. Several studies
demonstrated that they were associated with proliferation,
progression, and chemo-radiotherapy resistance of gliomas (5–9). The
PI3K/AKT pathway activated by EGFR and EGFRvIII was an important
molecular mechanism which gave rise to promoting proliferation and
resistance of drug and radiation (9,26). Both
EGFR/EGFRvIII and CEP55 were associated with proliferation of
glioma cells and activation of PI3K/AKT pathway. What is the
relationship between CEP55 and EGFR, especially EGFRvIII which is a
ligand-independent constitutively active variant of EGFR, in
regulation of glioma cells proliferation? To investigate the
regulating mechanism of proliferation by EGFRvIII and CEP55 in
glioma cells, U251 cells were firstly transfected with EGFRvIII
gene in this study. Expression of EGFRvIII promoted significantly
the proliferation of U251 cells transfected with EGFRvIII gene.
Meanwhile, the CEP55 expression increased remarkably in U251 cells
expressing EGFRvIII compared with cells of control group. This
result suggested that CEP55 participated the regulating mechanism
of proliferation induced by EGFRvIII in U251 cells.
EGFRvIII, a mutant of EGFR, was able to
over-activate various downstream signaling pathways, including the
PI3K/AKT and RAS-RAF-ERK and regulated many biological outputs that
are beneficial to glioma cell proliferation including their
initiation and progression of cell cycle (27). CEP55 participated directly abscission
in mitosis of cell. To verify CEP55 mediating the proliferation of
U251 cells promoted by EGFRvIII, the expression of CEP55 was
inhibited by CEP55 siRNA in U251 cell with EGFRvIII transfecting.
The proliferation of U251 cells promoted by EGFRvIII declined
significantly after CEP55 was knockdown. These results demonstrated
that CEP55 participated the proliferation regulated by EGFRvIII and
knockdown CEP55 was able to eliminate the effect of promoting
proliferation induced by EGFRvIII in U251 cells. Certainly, CEP55
is a mitotic phosphoprotein, acting as an effector of proliferation
pathways, which required tightly precise regulation in cytokinesis.
The effect of deregulated CEP55 expression in the glioma cells on
upstream factor of regulating proliferation such as EGFR remain
unresolved. The molecular mechanism and pathways of EGFRvIII
upregulating CEP55 expression in detail need further research in
our future study.
In conclusion, in this study, we found that CEP55
overexpression played a key role in proliferation of glioma cells
and mediated EGFRvIII stimulating proliferation in glioma cells.
Knockdown of CEP55 inhibited proliferation of U251 cells induced by
EGFRvIII and CEP55 may be a novel molecular therapeutic target to
patients with gliomas expressing EGFRvIII. The mechanism underlying
proliferation regulated by EGFRvIII-CEP55 in glioma cells need
further investigation.
Acknowledgements
This study was supported by the National Natural
Science Foundation of China (no. 81372408).
References
|
1
|
Frattini V, Trifonov V, Chan JM, Castano
A, Lia M, Abate F, Keir ST, Ji AX, Zoppoli P, Niola F, et al: The
integrated landscape of driver genomic alterations in glioblastoma.
Nat Genet. 45:1141–1149. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
2
|
Cancer Genome Atlas Research Network, ;
Brat DJ, Verhaak RG, Aldape KD, Yung WK, Salama SR, Cooper LA,
Rheinbay E, Miller CR, Vitucci M, et al: Comprehensive, integrative
genomic analysis of diffuse lower-grade gliomas. N Engl J Med.
372:2481–2498. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Louis DN, Ohgaki H, Wiestler OD, Cavenee
WK, Burger PC, Jouvet A, Scheithauer BW and Kleihues P: The 2007
WHO classification of tumours of the central nervous system. Acta
Neuropathol. 114:97–109. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Dolecek TA, Propp JM, Stroup NE and
Kruchko C: CBTRUS statistical report: Primary brain and central
nervous system tumors diagnosed in the United States in 2005–2009.
Neuro Oncol. 14 Suppl 5:v1–v49. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Network TC: Corrigendum: Comprehensive
genomic characterization defines human glioblastoma genes and core
pathways. Nature. 494:5062013. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Viana-Pereira M, Lopes JM, Little S,
Milanezi F, Basto D, Pardal F, Jones C and Reis RM: Analysis of
EGFR overexpression, EGFR gene amplification and the EGFRvIII
mutation in Portuguese high-grade gliomas. Anticancer Res.
28:913–920. 2008.PubMed/NCBI
|
|
7
|
Johnson H, Del Rosario AM, Bryson BD,
Schroeder MA, Sarkaria JN and White FM: Molecular characterization
of EGFR and EGFRvIII signaling networks in human glioblastoma tumor
xenografts. Mol Cell Proteomics. 11:1724–1740. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Ramnarain DB, Park S, Lee DY, Hatanpaa KJ,
Scoggin SO, Otu H, Libermann TA, Raisanen JM, Ashfaq R, Wong ET, et
al: Differential gene expression analysis reveals generation of an
autocrine loop by a mutant epidermal growth factor receptor in
glioma cells. Cancer Res. 66:867–874. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Mukherjee B, McEllin B, Camacho CV,
Tomimatsu N, Sirasanagandala S, Nannepaga S, Hatanpaa KJ, Mickey B,
Madden C, Maher E, et al: EGFRvIII and DNA double-strand break
repair: A molecular mechanism for radioresistance in glioblastoma.
Cancer Res. 69:4252–4259. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Shinojima N, Tada K, Shiraishi S, Kamiryo
T, Kochi M, Nakamura H, Makino K, Saya H, Hirano H, Kuratsu J, et
al: Prognostic value of epidermal growth factor receptor in
patients with glioblastoma multiforme. Cancer Res. 63:6962–6970.
2003.PubMed/NCBI
|
|
11
|
Verhaak RG, Hoadley KA, Purdom E, Wang V,
Qi Y, Wilkerson MD, Miller CR, Ding L, Golub T, Mesirov JP, et al:
Integrated genomic analysis identifies clinically relevant subtypes
of glioblastoma characterized by abnormalities in PDGFRA, IDH1,
EGFR, and NF1. Cancer Cell. 17:98–110. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Sun Y, Zhang W, Chen D, Lv Y, Zheng J,
Lilljebjörn H, Ran L, Bao Z, Soneson C, Sjögren HO, et al: A glioma
classification scheme based on coexpression modules of EGFR and
PDGFRA. Proc Natl Acad Sci USA. 111:pp. 3538–3543. 2014; View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Fabbro M, Zhou BB, Takahashi M, Sarcevic
B, Lal P, Graham ME, Gabrielli BG, Robinson PJ, Nigg EA, Ono Y and
Khanna KK: Cdk1/Erk2- and Plk1-dependent phosphorylation of a
centrosome protein, Cep55, is required for its recruitment to
midbody and cytokinesis. Dev Cell. 9:477–488. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Zhao WM, Seki A and Fang G: Cep55, a
microtubule-bundling protein, associates with centralspindlin to
control the midbody integrity and cell abscission during
cytokinesis. Mol Biol Cell. 17:3881–3896. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Doxsey S: Duplicating dangerously: Linking
centrosome duplication and aneuploidy. Mol Cell. 10:439–440. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Martinez-Garay I, Rustom A, Gerdes HH and
Kutsche K: The novel centrosomal associated protein CEP55 is
present in the spindle midzone and the midbody. Genomics.
87:243–253. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Chen CH, Lu PJ, Chen YC, Fu SL, Wu KJ,
Tsou AP, Lee YC, Lin TC, Hsu SL, Lin WJ, et al: FLJ10540-elicited
cell transformation is through the activation of PI3-kinase/AKT
pathway. Oncogene. 26:4272–4283. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jeffery J, Sinha D, Srihari S, Kalimutho M
and Khanna KK: Beyond cytokinesis: The emerging roles of CEP55 in
tumorigenesis. Oncogene. 35:683–690. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Carlton JG and Martin-Serrano J: Parallels
between cytokinesis and retroviral budding: A role for the ESCRT
machinery. Science. 316:1908–1912. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lee HH, Elia N, Ghirlando R,
Lippincott-Schwartz J and Hurley JH: Midbody targeting of the ESCRT
machinery by a noncanonical coiled coil in CEP55. Science.
322:576–580. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Chen CH, Lai JM, Chou TY, Chen CY, Su LJ,
Lee YC, Cheng TS, Hong YR, Chou CK, Whang-Peng J, et al: VEGFA
upregulates FLJ10540 and modulates migration and invasion of lung
cancer via PI3K/AKT pathway. PLoS One. 4:e50522009. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Tao J, Zhi X, Tian Y, Li Z, Zhu Y, Wang W,
Xie K, Tang J, Zhang X, Wang L and Xu Z: CEP55 contributes to human
gastric carcinoma by regulating cell proliferation. Tumor Biol.
35:4389–4399. 2014. View Article : Google Scholar
|
|
23
|
Carter SL, Eklund AC, Kohane IS, Harris LN
and Szallasi Z: A signature of chromosomal instability inferred
from gene expression profiles predicts clinical outcome in multiple
human cancers. Nat Genet. 38:1043–1048. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Wang G, Liu M, Wang H, Yu S, Jiang Z, Sun
J, Han K, Shen J, Zhu M, Lin Z, et al: Centrosomal protein of 55
regulates glucose metabolism, proliferation and apoptosis of glioma
cells via the Akt/mTOR signaling pathway. J Cancer. 7:1431–1440.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Gemenetzidis E, Bose A, Riaz AM, Chaplin
T, Young BD, Ali M, Sugden D, Thurlow JK, Cheong SC, Teo SH, et al:
FOXM1 upregulation is an early event in human squamous cell
carcinoma and it is enhanced by nicotine during malignant
transformation. PLoS One. 4:e48492009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xu W, Bi Y, Kong J, Zhang J, Wang B, Li K,
Tian M, Pan X, Shi B, Gu J, et al: Combination of an anti-EGFRvIII
antibody CH12 with Rapamycin synergistically inhibits the growth of
EGFRvIII+PTEN-glioblastoma in vivo. Oncotarget. 7:24752–24765.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Wee P and Wang Z: Epidermal growth factor
receptor cell proliferation signaling pathways. Cancers (Basel).
9:E522017. View Article : Google Scholar : PubMed/NCBI
|