Introduction
Ovarian serous carcinoma is a common cause of cancer
deaths of females worldwide. Patients are generally diagnosed at an
advanced stage and consequently have a high mortality rate. The
standard treatment comprises maximum debulking surgery and
platinum-taxane combination therapy. Despite the high response rate
to chemotherapy, the majority of patients will be resistant to
first-line agents, and the prognosis of resistant patients is
particularly poor. Upon recurrence, the probability of a response
to retreatment with platinum-based chemotherapy depends on the
platinum-free interval, defined as the duration from the last
platinum administration to cancer recurrence (1). If the ovarian carcinoma recurs within 6
months from the last platinum administration, it is defined as
‘platinum-resistant.’ If the carcinoma recurs after 6 months from
the last platinum administration, it is defined as
‘platinum-sensitive’ (2).
Platinum-sensitivity or resistance is an independent prognostic
factor for overall and progression-free survival of patients with
ovarian carcinoma (3).
However, it is difficult to determine sensitivity to
platinum-based chemotherapy at the first administration of
chemotherapy and before the first recurrence. Therefore,
‘platinum-resistant’ patients are identified retrospectively after
recurrence or unresponsiveness to initial platinum-based
chemotherapy. Knowledge of the predictors of the response to
platinum-based chemotherapy may allow selection of sensitive
patients who are candidates for chemotherapy, as well as to avoid
administering platinum-based chemotherapy to platinum-resistant
patients. Further, customized treatment can be designed according
to clinical stratification according to drug resistance.
Unfortunately, a reliable method is not available
that determines or predicts platinum resistance. To improve the
prognosis of platinum-resistant patients with ovarian serous
carcinoma, we aimed here to identify new biomarkers with prognostic
and predictive potential and to identify new therapeutic
targets.
T-box 2 (TBX2) is a member of the T-box family of
transcription factors that are involved in embryonic development,
cell cycle regulation, and cancer (4,5). TBX2
allows cells to bypass senescence through its ability to repress
the activities of the cell cycle regulators p21 and
p14ARF, and silencing TBX2 expression induces senescence
(5–8).
Overexpression of TBX2 occurs in breast cancer (7,9), melanoma
(8) pancreatic cancer (10), gastric cancer (11), prostate cancer (12), laryngeal squamous cell carcinoma
(13), and non-small cell lung cancer
(14). The ectopic expression of TBX2
is associated with resistance to the DNA-damaging chemotherapeutic
drugs cisplatin and doxorubicin (15,16).
However, the mechanism of regulation of expression and the role of
TBX2 in ovarian cancer remain to be determined.
Here we assessed the association between TBX2
expression and the sensitivity of ovarian serous carcinoma to
platinum-based chemotherapy. We aimed to identify new biomarkers
with prognostic and predictive potential and searched for new
therapeutic targets to improve the prognosis of patients with
platinum-resistant ovarian serous carcinoma.
Patients and methods
Patients and samples
We reviewed the records of 54 patients with ovarian
serous carcinoma, stages III–IV, treated at our hospital from
January 2005 to December 2013. Patients were allocated to the
groups as follows: i) Platinum-sensitive group (n=27), subjected to
maximum debulking surgery followed by platinum-based chemotherapy
whose tumors did not recur within 6 months from the last platinum
administration; ii) platinum-resistant group (n=27) subjected to
maximum debulking surgery followed by platinum-based chemotherapy
whose tumors recurred within 6 months. Written informed consent was
obtained from all patients prior to treatment and the Institutional
Review Board (IRB) of Osaka City University Hospital approved this
study (IRB no. 3525).
Immunohistochemistry
TBX2 expression was determined by conducting
immunohistochemical analysis of paraffin-embedded sections using a
Dako LSAB2 Peroxidase kit (cat. no. K0675; Agilent Technologies,
Inc., Santa Clara, CA, USA). Sections (4 µm-thick) were
deparaffinized, rehydrated, and immersed in 3% hydrogen peroxide at
room temperature for 10 min to block endogenous peroxidase
activity. Antigen retrieval was performed by immersing sections in
10 mM citrate buffer (pH 6.0) and heating to 110°C for 20 min in an
autoclave. Tissue sections were then washed in phosphate-buffered
saline (PBS) and incubated overnight at 4°C with a 1:32 dilution of
a rabbit polyclonal anti-TBX2 antibody (LS-C402301; LifeSpan
BioSciences, Inc., Seattle, WA, USA). Next, sections were washed in
PBS for 15 min and then incubated for 10 min with biotinylated goat
immunoglobulin G secondary antibodies (Dako; Agilent Technologies,
Inc.). Sections were then incubated with a streptavidin-peroxidase
complex, and 3,3′-diaminobenzidine was used as the chromogen.
Finally, tissue sections were counterstained with hematoxylin, and
the specificity of the immunohistochemical reactions was verified
by omitting the primary antibody.
TBX2 expression scores were calculated using the
weighted score of Sinicrope et al (17). The percentage positivity was scored
as: 0 (<5%), 1 (5–25%), 2 (25–50%), 3 (50–75%), and 4 (>75%).
The staining intensity was scored as 0 (no staining), 1 (weak
staining), 2 (moderate staining), or 3 (strong staining). The
percentage positivity of cells and staining intensity were
determined in a double-blinded manner. The TBX2 expression score
was calculated by multiplying the percentage positivity score and
the staining intensity score, which ranged from 0 to 12.
Cell culture
The human ovarian serous carcinoma cell line OVSAHO
(cat no. JCRB1046; National Institutes of Biomedical Innovation,
Health and Nutrition, Osaka, Japan) was cultured in RPMI medium
(Gibco; Thermo Fisher Scientific, Inc., Waltham, MA, USA)
containing 10% fetal bovine serum (Gibco; Thermo Fisher Scientific,
Inc.), penicillin (100 units/ml), and streptomycin (100 units/ml)
at 37°C in a humidified incubator containing an atmosphere of 5%
CO2. The medium was changed daily. For real-time PCR
(RT-qPCR) analysis, cells were directly cryopreserved in a
refrigerator at −20°C.
Cell survival assay and siRNA
procedures
OVSAHO cells were seeded into 96-well plates at
10,000 cells per well. We divided cells into a control group that
was not transfected with the TBX2-specific siRNA (siTBX2) and an
siTBX2 group that was transfected with siTBX2 for 24 h. The siTBX2
sequence was sense: 5′rCr CrA rAU rGr ArA rCU rGr CrA rGr ArG rCr
AUT T, antisense: 5′rAU rGr CUr CUr GrC rAr GUU rCr AUU rGr GTT.
Cells were transfected with siTBX2 using Lipofectamine RNAiMax
(Invitrogen, Carlsbad, CA, USA) according to manufacturer's
instructions. Twenty-four h after cell adhesion in the control
group and 24 h after transfection with siTBX2 of the siTBX2 group,
the medium was replaced with the fresh media containing 0, 1.0,
2.5, or 5.0 µM cisplatin or 0, 10, 50, or 100 µM carboplatin, and
the cells were incubated for 48 h. Cells were prepared in six wells
for each treatment and were incubated for 24 h prior to 48 h
treatment with cisplatin or carboplatin. Cell viability was
measured using a Cell Counting kit-8 (CCK-8; Dojindo Molecular
Technologies, Inc., Kumamoto, Japan). Specifically, 10 µl CCK-8 and
100 µl RPMI1640 were added to each well and incubated for 2 h at
37°C. Absorbance was measured at 450 nm using a microplate reader
(Corona Electric Co., Ltd., Ibaraki, Japan). Dose-response curves
were generated to determine the percentage of viable cells compared
with that of control cells.
RT-qPCR
Total RNA was extracted from OVSAHO cells using an
RNeasy mini kit (QIAGEN GmbH, Hilden, Germany) following
manufacturer's instructions. RNA was reverse-transcribed using a
High Capacity cDNA Reverse Transcription kit (Thermo Fisher
Scientific, Inc.). Expression of TBX2 mRNA was performed using the
TaqMan Gene Expression Assay (Applied Biosystems; Thermo Fisher
Scientific, Inc.) with an ABI7500 Fast System. The mRNA levels were
normalized to those of GAPDH mRNA. The RT-qPCR assays
(Thermo Fisher Scientific, Inc.) employed TaqMan TBX2
(Hs00911929_m1) and GAPDH (Hs99999905_m1) assays.
Statistical analysis
The data are expressed as the mean ± standard
deviation. Kaplan-Meier and log-rank analyses were performed to
evaluate prognosis. Weighted scores were compared using the
Mann-Whitney test. The Student's t-test was performed to evaluate
the significance of differences between the mean values of two
groups, and the χ2 test was performed to identify
significant associations between the categorical variables of the
two groups. SPSS software version 21.0 (IBM SPSS, Armonk, NY, USA)
was used for all statistical analyses. P<0.05 indicates a
statistically significant difference.
Results
Patients' characteristics
We investigated the associations of age, FIGO stage,
CA125 levels, and postoperative residual disease. There was no
significant difference between the former three variables. In
contrast, the size of postoperative residual disease was
significantly higher in the platinum-resistant group (P=0.004)
(Table I).
 | Table I.Characteristics of patients in the
platinum-sensitive and platinum-resistant groups. |
Table I.
Characteristics of patients in the
platinum-sensitive and platinum-resistant groups.
| Characteristics | Platinum sensitive
(n) | Platinum resistant
(n) | P-value |
|---|
| No. of patients | 27 | 27 |
|
| Age (years) |
|
|
0.725a |
| Mean ±
SD | 61.0±12.2 | 60.0±10.0 |
|
| FIGO stage |
|
| 0.277b |
| IIIA | 1 | 0 |
|
| IIIB | 3 | 1 |
|
| IIIC | 21 | 19 |
|
| IVA | 1 | 4 |
|
| IVB | 1 | 3 |
|
| Tumor marker |
|
| 0.374a |
| Mean
CA125, U/ml | 3,343.3 | 2,180.1 |
|
| Postoperative |
|
| 0.004b |
| residual disease |
|
|
|
| None | 5 | 0 |
|
| <1
cm | 10 | 4 |
|
| >1
cm | 12 | 23 |
|
Expression of TBX2 in ovarian serous
carcinoma tissue
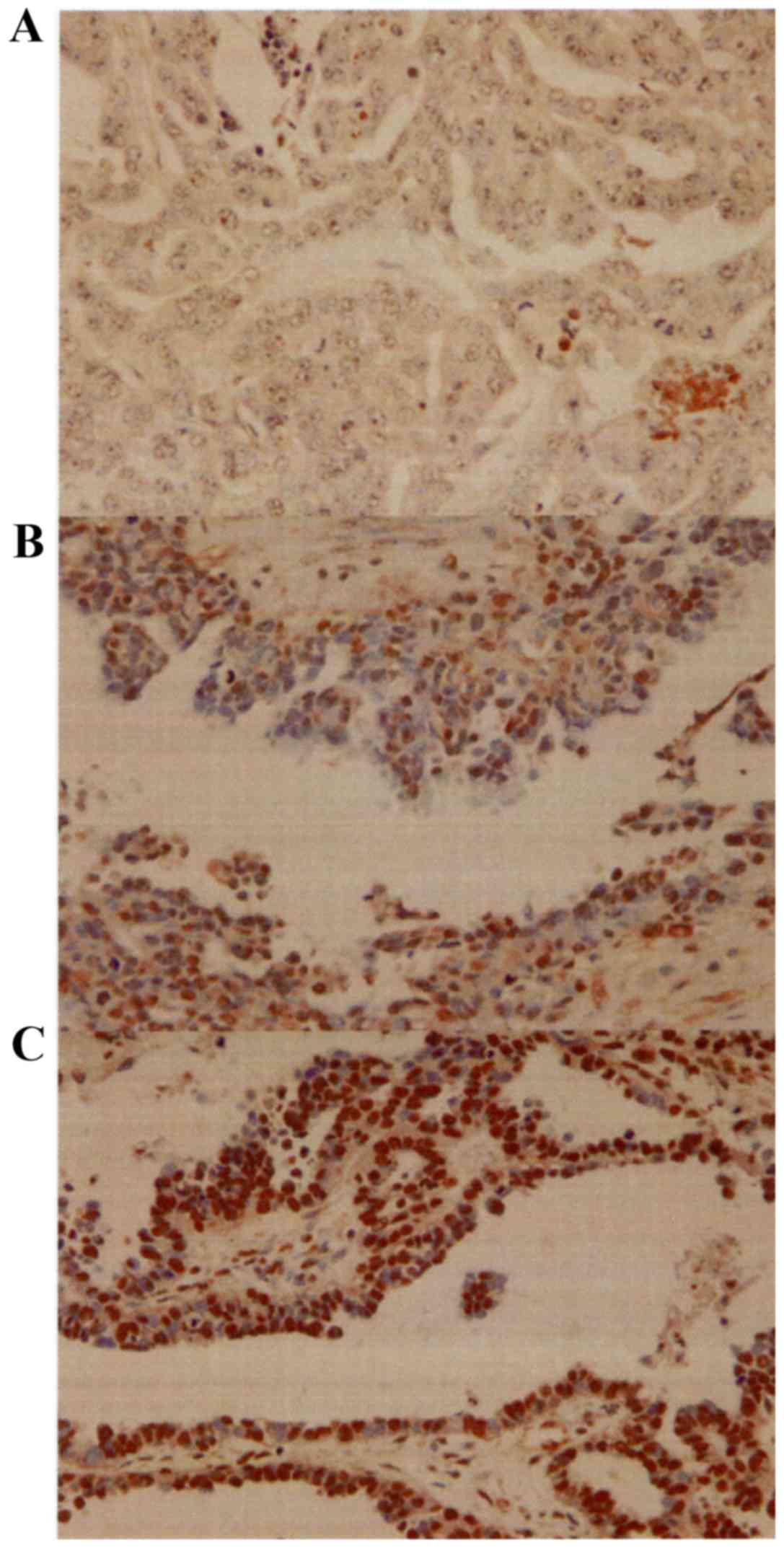
TBX2 was detected predominantly in the nucleus
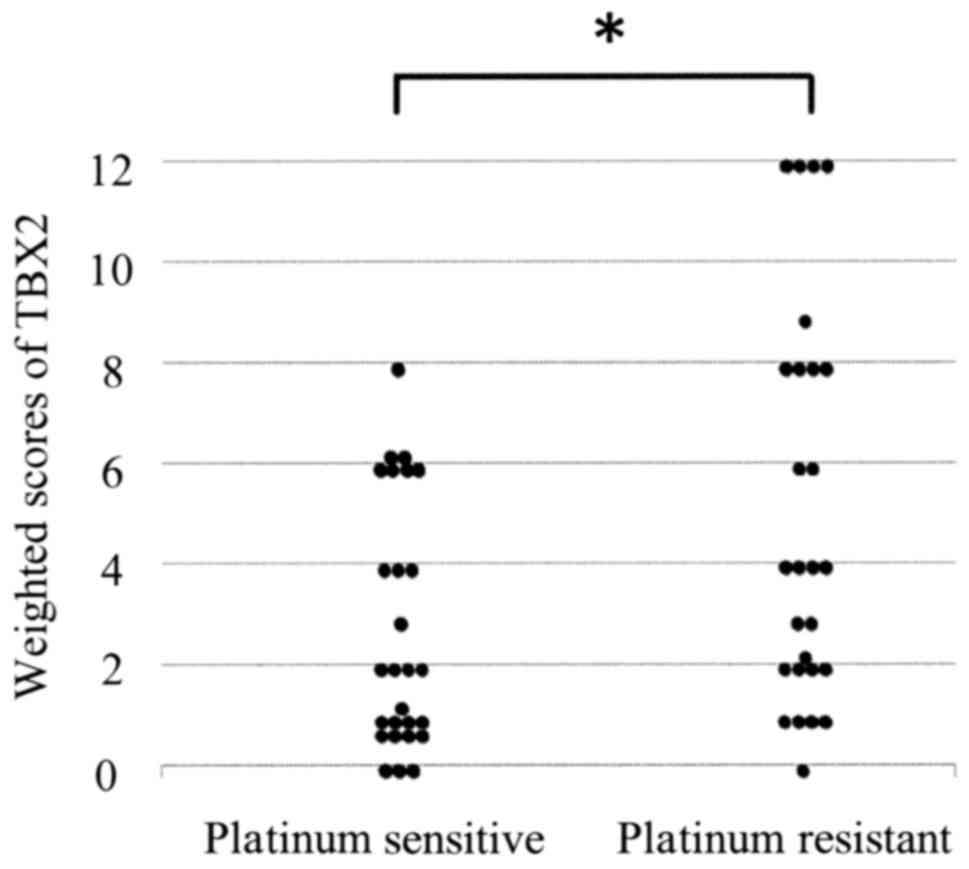
(Fig. 1). The mean weighted scores of
the platinum-sensitive and platinum-resistant groups were 2.7 and
5.4, respectively. The mean weighted score for TBX2 expression was
significantly lower in the platinum sensitive group (P=0.005)
(Table II and Fig. 2). Next, we divided the patients into
two groups according to their TBX2 expression scores as follows:
Low TBX2 expression (weighted score ≤6, n=44) and high TBX2
expression (weighted score ≥8, n=10). Table III lists the characteristics of the
high and low expression groups. There was no significant difference
between them.
 | Table II.Weighted scores of TBX2 expression in
the platinum-sensitive and platinum-resistant groups. |
Table II.
Weighted scores of TBX2 expression in
the platinum-sensitive and platinum-resistant groups.
|
| No. of patients |
|---|
|
|
|
|---|
| Weighted score | Platinum
sensitive | Platinum
resistant |
|---|
| 0 | 3 | 1 |
| 1 | 9 | 4 |
| 2 | 4 | 5 |
| 3 | 1 | 2 |
| 4 | 3 | 4 |
| 6 | 6 | 2 |
| 8 | 1 | 4 |
| 9 | 0 | 1 |
| 12 | 0 | 4 |
| Total | 27 | 27 |
| Mean | 2.7 | 5.4 |
 | Table III.Characteristics of patients in the
low and high TBX2 expression groups. |
Table III.
Characteristics of patients in the
low and high TBX2 expression groups.
|
| No. of
patients |
|
|---|
|
|
|
|
|---|
|
Characteristics | Low TBX2 expression
(score ≤6) | High TBX2
expression (score ≤8) | P-value |
|---|
| No. of
patients | 44 | 10 |
|
| Age (years) |
|
| 0.685a |
| Mean ±
SD | 60.2±11.4 | 61.8±10.0 |
|
| FIGO stage |
|
| 0.649b |
|
IIIA | 1 | 0 |
|
|
IIIB | 4 | 0 |
|
|
IIIC | 31 | 9 |
|
|
IVA | 4 | 1 |
|
|
IVB | 4 | 0 |
|
| Tumor marker |
|
| 0.593a |
| Mean
CA125, U/ml | 2,931.0 | 2,017.1 |
|
| Postoperative
residual disease |
|
| 0.419b |
|
None | 5 | 0 |
|
| <1
cm | 12 | 2 |
|
| >1
cm | 27 | 8 |
|
Association of platinum sensitivity
with TBX2 expression
In the low TBX2 expression group, there were 26
(59.1%) and 18 (40.9%) patients in the platinum-sensitive and
platinum-resistant groups, respectively. In the high TBX2
expression group, one patient (10.0%) and nine (90%) patients were
in the platinum-sensitive and platinum-resistant groups,
respectively. The low TBX2 expression group was more sensitive to
platinum-based chemotherapy compared with the high TBX2 expression
group (P=0.004) (Table IV).
 | Table IV.Number of patients with low and high
TBX2 expression in the platinum-sensitive and platinum-resistant
groups. |
Table IV.
Number of patients with low and high
TBX2 expression in the platinum-sensitive and platinum-resistant
groups.
| TBX2
expression | Platinum sensitive,
number (%) | Platinum resistant,
number (%) | P-value |
|---|
| Low expression
(score ≤6) | 26 (59.1) | 18 (40.9) | 0.004a |
| High expression
(score ≥8) | 1 (10.0) | 9 (90.0) |
|
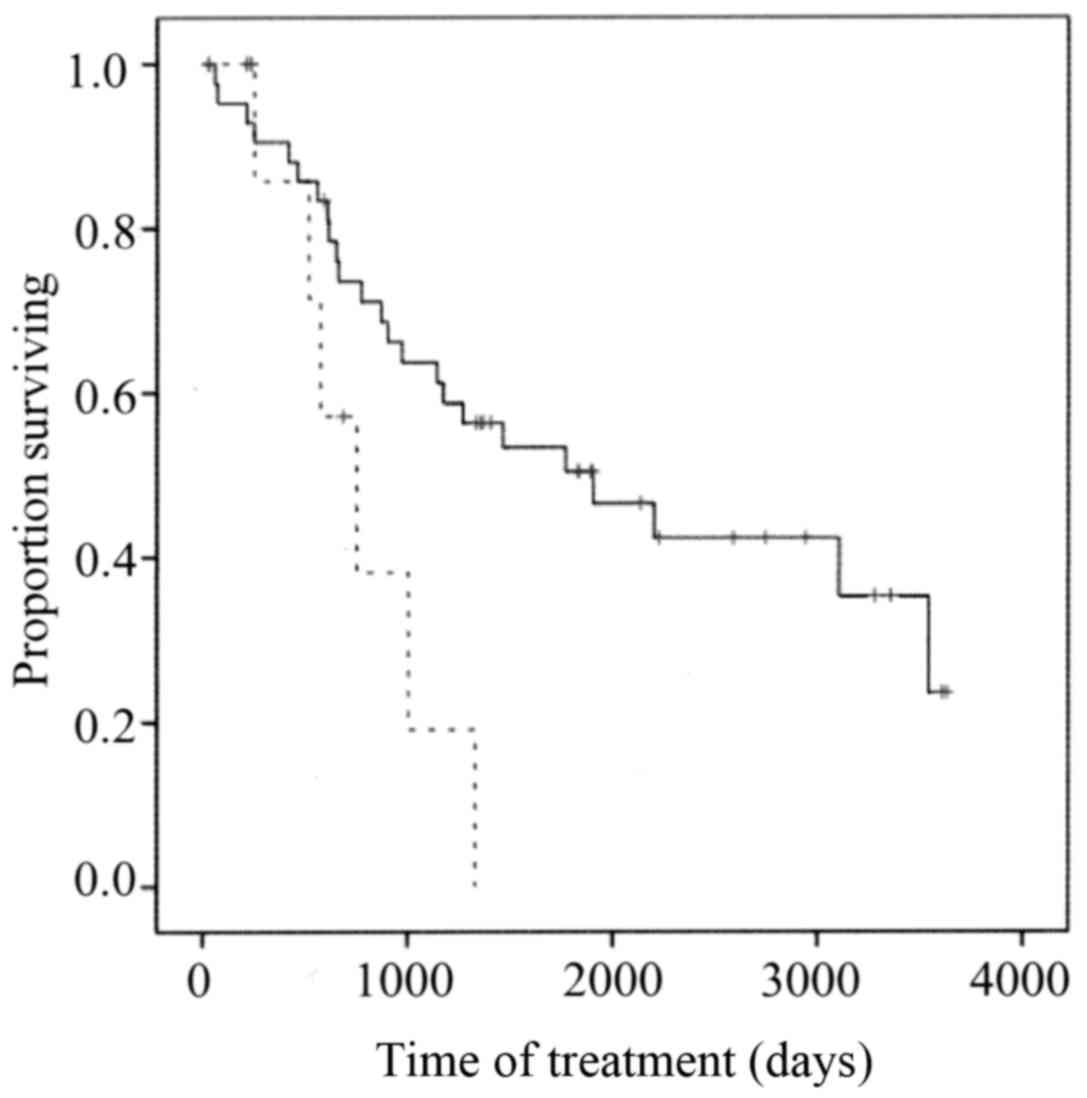
Survival
The overall survival (OS) of members of the low TBX2
expression group was significantly longer compared with that of the
high TBX2 expression group (P=0.023) (Fig. 3).
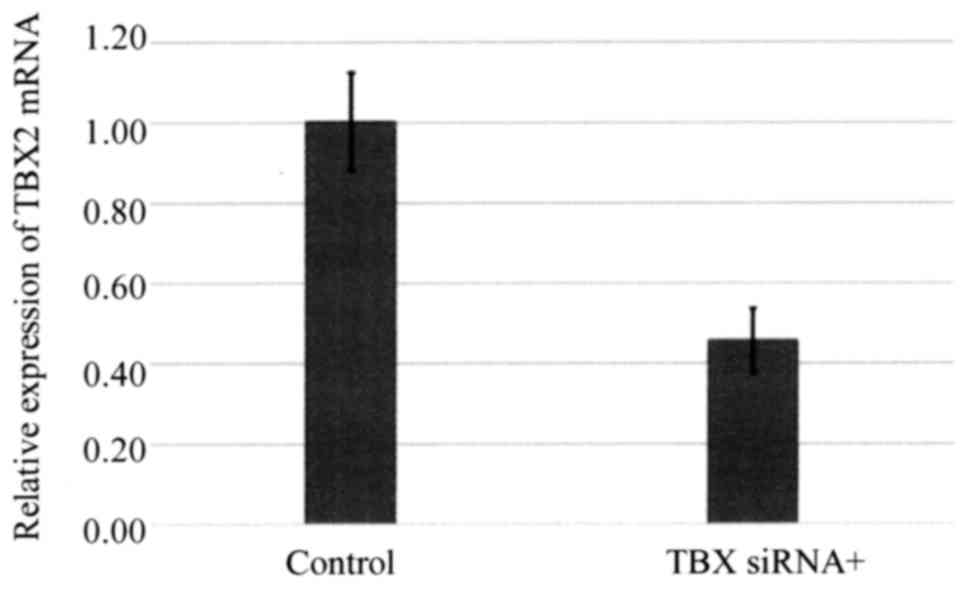
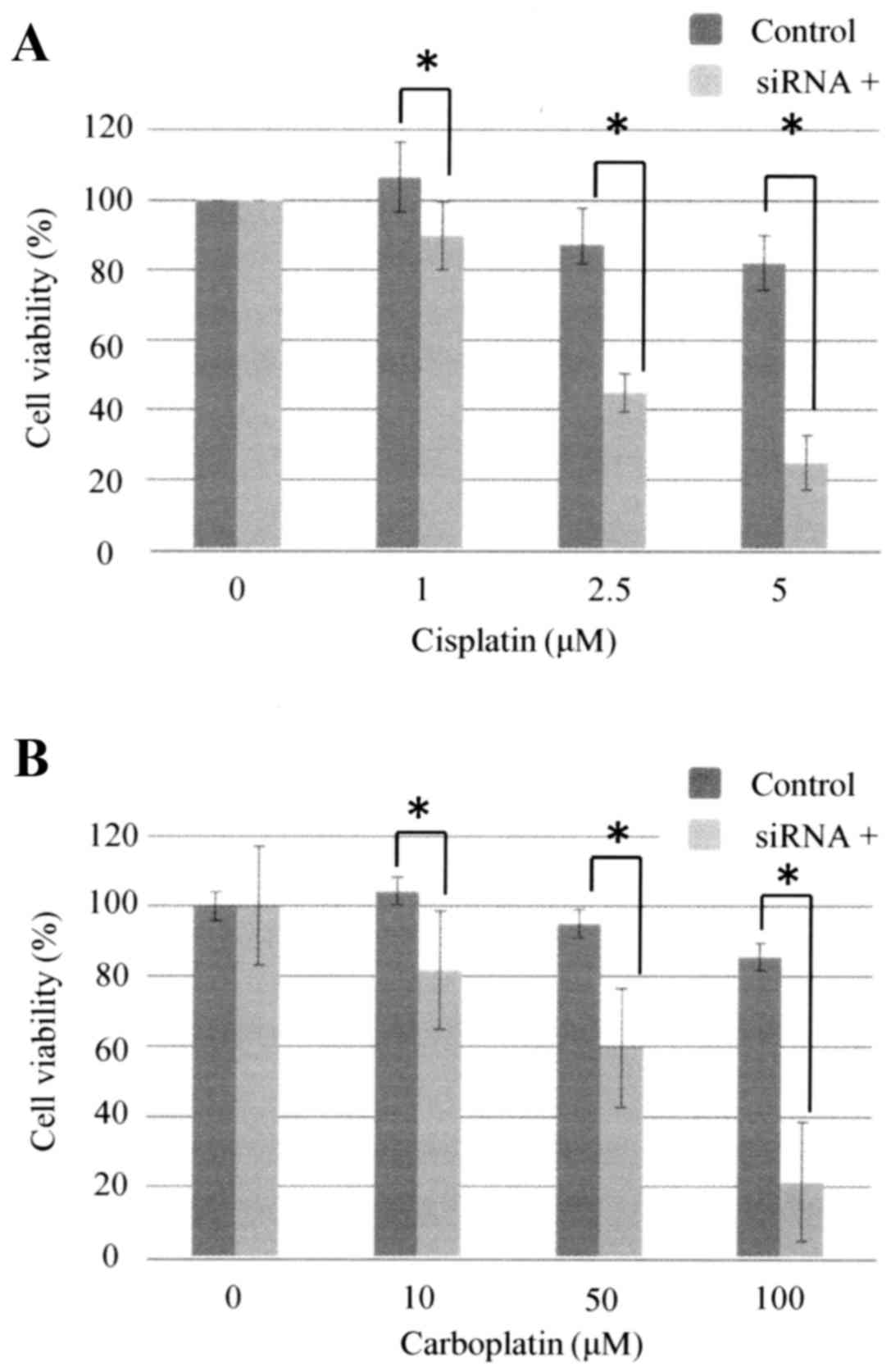
siRNA-mediated silencing of TBX2
expression enhances the sensitivity of ovarian carcinoma cells to
carboplatin
TBX2 mRNA expression by OVSAHO cells was suppressed
48 h after the cells were transfected with siTBX2 (Fig. 4), and cells transfected with siTBX2
were significantly more sensitive to cisplatin and carboplatin
after 48 h (Fig. 5).
Discussion
The effective treatment of ovarian serous carcinoma
remains a major challenge because of the recurrence of
platinum-resistant tumors. The mechanism of platinum-resistance may
involve decreased cellular uptake caused by abnormalities of
transporters, intracellular cisplatin inactivation (e.g., caused by
glutathione), and increased DNA repair (18). However, no available therapy prevents
platinum-resistance.
TBX2 is overexpressed by numerous human cancers
(7–14). TBX2 may serve as a prognostic factor
of breast cancer (7,9), melanoma (8), gastric cancer (10), prostate cancer (11), laryngeal squamous cell carcinoma
(12), and non-small cell lung cancer
(14). TBX2 is associated with
resistance to therapeutic drugs such as cisplatin and doxorubicin
(15,16), and TBX2 therefore may serve as a
therapeutic target.
One report shows that chromosome 17q12-q24 harbors
strong candidates for ovarian tumorigenesis, such as LASP1 (17q12),
TGF11 (17q21.32), MUL (17q23.2), TBX2 (17q23.2), AXIN2 (17q24.3),
and GRB2 (17q25.1) (19). Further,
TBX2 is upregulated in a subset of breast cancer cell lines, and
breast tumors with mutations in BRCA1 and BRCA2 (7,20–22), which are strongly associated with
ovarian serous carcinoma.
In the present study, the OS of the low TBX2
expression group was significantly longer compared with that of the
high TBX2 expression group. Transfection with siTBX2 increased the
sensitivity of OVSAHO cells to cisplatin and carboplatin. Wansleben
et al found that breast cancer and melanoma cell lines are
sensitive to cisplatin and undergo mitotic catastrophe in a
cisplatin-resistant breast cancer cell line when TBX2 expression is
knocked down (23), which is
consistent with our present results.
Demay et al found that TBX2 has the potential
to recognize mitotic chromatin and can interact with the histone H3
N-terminal tail (24). Further,
Warfel et al indicated that p21 is frequently downregulated
in human cancers, and its expression can either inhibit or promote
carcinogenesis, depending on the cellular context (25). Huang et al reported that TBX2
is overexpressed, while p21 is expressed at relatively lower levels
in laryngeal squamous cell cancer (13). Moreover, TBX2 binds and represses the
p21 promoter in melanoma cells by recruiting histone deacetylase 1
(HDAC1) (8). In contrast, TBX2 and
p21 protein levels increase simultaneously (24).
Another mechanism that accounts for poor prognosis
associated with TBX2 involves the repression of E-cadherin by TBX2,
leading to the epithelial-mesenchymal transition and subsequent
invasion of adjacent tissues by tumor cells (26). Moreover, TBX2 inhibits the tumor
suppressor PTEN by recruiting HDAC1 (27). Despite several studies showing the
association of TBX2 with poor prognosis, the mechanism by which
TBX2 induces resistance of chemotherapy is unknown. This study
included only 54 patients. That small number is one of the
limitations of this study. Further investigations with a larger
number of cases are required to fill the critically important gap
in our knowledge.
In conclusion, TBX2 expression may serve as a marker
that predicts the efficacy of platinum-based chemotherapy
administered to patients with ovarian serous carcinoma. To our
knowledge, the present study is the first to report an association
of TBX2 expression with platinum sensitivity. This knowledge will
be helpful for efforts to improve the prognosis of patients with
ovarian serous carcinoma.
References
|
1
|
Friedlander M, Trimble E, Tinker A,
Alberts D, Avall-Lundqvist E, Brady M, Harter P, Pignata S,
Pujade-Lauraine E, Sehouli J, et al: Clinical trials in recurrent
ovarian cancer. Int J Gynecol Cancer. 21:771–775. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Markman M, Rothman R, Hakes T, Reichman B,
Hoskins W, Rubin S, Jones W, Almadrones L and Lewis JL Jr:
Second-line platinum therapy in patients with ovarian cancer
previously treated with cisplatin. J Clin Oncol. 9:389–393. 1991.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Kyrgiou M, Salanti G, Pavlidis N,
Paraskevaidis E and Ioannidis JP: Survival benefits with diverse
chemotherapy regimens for ovarian cancer: Meta-analysis of multiple
treatments. J Natl Cancer Inst. 98:1655–1663. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Rowley M, Grothey E and Couch FJ: The role
of Tbx2 and Tbx3 in mammary development and tumorigenesis. J
Mammary Gland Biol Neoplasia. 9:109–118. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Abrahams A, Parker MI and Prince S: The
T-box transcription factor Tbx2: Its role in development and
possible implication in cancer. IUBMB Life. 62:92–102.
2010.PubMed/NCBI
|
|
6
|
Peres J, Davis E, Mowla S, Bennett DC, Li
JA, Wansleben S and Prince S: The highly homologous T-Box
transcription factors, TBX2 and TBX3, have distinct roles in the
oncogenic process. Genes Cancer. 1:272–282. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jacobs JJ, Keblusek P, Robanus-Maandag E,
Kristel P, Lingbeek M, Nederlof PM, van Welsem T, van de Vijver MJ,
Koh EY, Daley GQ and van Lohuizen M: Senescence bypass screen
identifies TBX2, which represses Cdkn2a (p19(ARF)) and is amplified
in a subset of human breast cancers. Nat Genet. 26:291–299. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Vance KW, Carreira S, Brosch G and Goding
CR: Tbx2 is overexpressed and plays an important role in
maintaining proliferation and suppression of senescence in
melanomas. Cancer Res. 65:2260–2268. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Taneja P, Maglic D, Kai F, Zhu S, Kendig
RD, Fry EA and Inoue K: Classical and novel prognostic markers for
breast cancer and their clinical significance. Clin Med Insights
Oncol. 4:15–34. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Mahlamäki EH, Bärlund M, Tanner M,
Gorunova L, Höglund M, Karhu R and Kallioniemi A: Frequent
amplification of 8q24, 11q, 17q and 20q-specific genes in
pancreatic cancer. Genes Chromosomes Cancer. 35:353–358. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Yu H, Liu BO, Liu A, Li K and Zhao H:
T-box 2 expression predicts poor prognosis in gastric cancer. Oncol
Lett. 10:1689–1693. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Nandana S, Tripathi M, Duan P, Chu CY,
Mishra R, Liu C, Jin R, Yamashita H, Zayzafoon M, Bhowmick NA, et
al: Bone metastasis of prostate cancer can be therapeutically
targeted at the TBX2-WNT signaling axis. Cancer Res. 77:1331–1344.
2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Huang Y, Li Z, Zhong Q, Li G, Zhang Y and
Huang Z: Association of TBX2 and P21 expression with
clinicopathological features and survival of laryngeal squamous
cell carcinoma. Int J Clin Exp Med. 7:5394–5402. 2014.PubMed/NCBI
|
|
14
|
Zhang Z and Guo Y: High TBX2 expression
predicts poor prognosis in non-small cell lung cancer. Neoplasma.
61:476–480. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Davis E, Teng H, Bilican B, Parker MI, Liu
B, Carriera S, Goding CR and Prince S: Ectopic Tbx2 expression
results in polyploidy and cisplatin resistance. Oncogene.
27:976–984. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ismail A and Bateman A: Expression of TBX2
promotes anchorage-independent growth and survival in the
p53-negative SW13 adrenocortical carcinoma. Cancer Lett.
278:230–240. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Sinicrope FA, Ruan SB, Cleary KR, Stephens
LC, Lee JJ and Levin B: bcl-2 and p53 oncoprotein expression during
colorectal tumorigenesis. Cancer Res. 55:237–241. 1995.PubMed/NCBI
|
|
18
|
Amable L: Cisplatin resistance and
opportunities for precision medicine. Pharmacol Res. 106:27–36.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dimova I, Orsetti B, Negre V, Rouge C,
Ursule L, Lasorsa L, Dimitrov R, Doganov N, Toncheva D and Theillet
C: Genomic markers for ovarian cancer at chromosomes 1, 8 and 17
revealed by array CGH analysis. Tumori. 95:357–366. 2009.PubMed/NCBI
|
|
20
|
Redmond KL, Crawford NT, Farmer H, D'Costa
ZC, O'Brien GJ, Buckley NE, Kennedy RD, Johnston PG, Harkin DP and
Mullan PB: T-box 2 represses NDRG1 through an EGR1-dependent
mechanism to drive the proliferation of breast cancer cells.
Oncogene. 29:3252–3262. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Sinclair CS, Adem C, Naderi A, Soderberg
CL, Johnson M, Wu K, Wadum L, Couch VL, Sellers TA, Schaid D, et
al: TBX2 is preferentially amplified in BRCA1- and BRCA2-related
breast tumors. Cancer Res. 62:3587–3591. 2002.PubMed/NCBI
|
|
22
|
Adem C, Soderberg CL, Hafner K, Reynolds
C, Slezak JM, Sinclair CS, Sellers TA, Schaid DJ, Couch F, Hartmann
LC and Jenkins RB: ERBB2, TBX2, RPS6KB1 and MYC alterations in
breast tissues of BRCA1 and BRCA2 mutation carriers. Genes
Chromosomes Cancer. 41:1–11. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Wansleben S, Davis E, Peres J and Prince
S: A novel role for the anti-senescence factor TBX2 in DNA repair
and cisplatin resistance. Cell Death Dis. 10:4:e8462013.
|
|
24
|
Demay F, Bilican B, Rodriguez M, Carreira
S, Pontecorvi M, Ling Y and Goding CR: T-box factors: Targeting to
chromatin and interaction with the histone H3 N-terminal tail.
Pigment Cell Res. 20:279–287. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Warfel NA and El-Deiry WS: P21WAF1 and
tumourigenesis: 20 years after. Curr Opin Oncol. 25:52–58. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Wang B, Lindley LE, Fernandez-Vega V,
Rieger ME, Sims AH and Briegel KJ: The T box transcription factor
TBX2 promotes epithelial-mesenchymal transition and invasion of
normal and malignant breast epithelial cells. PLoS One.
7:e413552012. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhu B, Zhang M, Williams EM, Keller C,
Mansoor A and Davie JK: TBX2 represses PTEN in rhabdomyosarcoma and
skeletal muscle. Oncogene. 35:4212–4224. 2016. View Article : Google Scholar : PubMed/NCBI
|