Introduction
Breast cancer is the most commonly occurring type of
cancer in females, and excessive proliferation of tumor cells is an
important factor affecting the prognosis of breast cancer (1,2). In recent
years, an increasing number of studies have demonstrated that
microRNAs (miRNA/miR) are involved in the diagnosis, treatment and
prognosis of breast cancer (3–5), including
a large-scale scientific investigation into molecular biological
markers for monitoring the tumor progression, diagnosis or therapy
(6,7).
Previous studies have revealed that miR-194 is highly expressed in
numerous types of human breast cancer and radiation-induced breast
cancer (8,9); however, the underlying mechanisms
require further investigation.
F-box/WD repeat-containing protein 7
(Fbxw-7), is a tumor suppressor that has been demonstrated
to regulate several oncoproteins (10) including, cyclin D, cyclin E (11–13), c-Myc
(14,15), aurora kinase A (16), Notch (17–20) and
the proto-oncogene c-Jun (21,22) c-Jun,
an extensively-studied member of the JUN family, has been
associated with cell proliferation, tumor cell survival and
apoptosis (23). Therefore, it is
important to investigate the Fbxw-7-associated signaling
pathways involved in the proliferation of breast cancer. Previous
studies have focused on the signal factors for the expression of
Fbxw-7 prior to and following its transcription (24,25). It
has previously been revealed that the Fbxw-7 gene may be
suppressed due to methylation of its promoter region (26). In addition, it has been demonstrated
that miR-223 serves a critical role in human gastric cancer by
targeting Fbxw-7 (27). These
studies suggest that Fbxw-7 is important in human types of
cancer for its interactions with alternative genes.
The present study demonstrated that the serum
expression levels of miR-194 were significantly higher in patients
with breast cancer compared with in healthy adults. Additionally,
the overexpression of miR-194 significantly inhibited the
proliferation of breast cancer cells by directly targeting the
3′-untranslated region (3′-UTR) of Fbxw-7 and by regulating
the cell cycle. Overall, the results of the present study
demonstrated that miR-194 expression has a significant impact on
the progression of breast cancer and may be a potential therapeutic
target for breast cancer.
Materials and methods
Patients and samples
Blood samples were obtained by EDTA vacutainer blood
collection tubes (BD Biosciences, Franklin Lakes, NJ, USA) and
immediately stored at −80°C from 43 patients with breast cancer
(aged 18–80 years) at the First Affiliated Hospital of Zhejiang
University (Hangzhou, China) between April 2013 and April 2015. The
samples were immediately stored at −80°C until use. All of the
patients were female and >18 years old. A total of 26 of the
patients were identified to have well differentiated breast cancer
(grade I–II) and 17 had poorly differentiated breast cancer (grade
III). A total of 20 healthy adults were recruited from the First
Affiliated Hospital of Zhejiang University to participate in the
study as the control group. The Ethics Committee of The First
Affiliated Hospital of Zhejiang University approved the present
study. Written informed consent was obtained from all participants
included in the study.
Cell lines
Two breast cancer cell lines, MCF-7 and MDA-MB-231,
were obtained from the American Type Culture Collection (Manassas,
VA, USA). They were cultured in Dulbecco's modified Eagle's medium
(DMEM) with high glucose (Gibco; Thermo Fisher Scientific, Inc.,
Waltham, MA, USA), supplemented with 10% fetal bovine serum (FBS)
and 100 U/ml penicillin and 100 µg/ml streptomycin, at 37°C in 5%
CO2.
Transfection of miRNA or the
inhibitor
Breast cancer cells were seeded in a 24-well plate
with a density of 4×104 cells/well and cultured for 24 h
at 37°C prior to transfection. Subsequently, the cells were starved
for 1 h at 37°C using FBS-free medium without antibiotics.
Transfection with the negative control oligonucleotides (NC mimic),
miR-194, antagomiR-194 or miR-194/antagomiR-194 was performed using
Lipofectamine® 2000 reagent (Invitrogen; Thermo Fisher
Scientific, Inc.), according to the manufacturer's instructions.
Briefly, 0.5 µg miRNA was diluted with 100 µl Opti-MEM (Thermo
Fisher Scientific, Inc.) without FBS, which was mixed with 1.25 µl
Lipofectamine® LTX reagent (Invitrogen; Thermo Fisher
Scientific, Inc.). The mixture (containing the transfection
reagents) was incubated for 25 min at room temperature, and then
added to wells containing cells and incubated at room temperature
for 4 h. Following 48 h transfection at 37°C, the cells from the
various groups were seeded (1×104 cells/ml) onto 96-well
plates in triplicate, and then cultured at 37°C for 24 h.
RNA isolation and reverse
transcription-polymerase chain reaction (RT-PCR)
Total RNA was isolated from breast cancer cell lines
using TRIzol® (Invitrogen; Thermo Fisher Scientific,
Inc.), chloroform, isopropanol and ethanol, following the
manufacturer's instructions. The isolated RNA, including miRNA, was
dissolved in 30 µl diethyl pyrocarbonate-treated (high temperature
sterilization at 121°C for 20 min) water. In order to detect the
serum levels of miR-194, the total RNA was transcribed into cDNA
using the miScript II RT kit (Qiagen GmbH, Hilden, Germany). cDNA
was synthesized with the Prime-Script RT reagent kit (Takara Bio
Inc., Otsu, Japan) from 500 ng of total RNA. RT-PCR was performed
in triplicate using Fast SYBR® Green Master Mix (Thermo
Fisher Scientific, Inc.), according to the manufacturer
instructions of the ABI Prism® 7900HT Sequence Detection
System (Applied Biosystems; Thermo Fisher Scientific, Inc.). The
primer pairs for miR-194 were as follows: Forward,
5′-CTAAGCTTAGTGGGCATGGGACACTCT-3′ and reverse,
5′-CTGAATTCACCTGCCTCTCCTTCTTCGT-3′. U6 was used as an internal
control. The primers for U6 were as follows: Forward,
5′-CTCGCTTCGGCAGCACA-3′ and reverse, 5′-AACGCTTCACGAATTTGCGT-3′.
The PCR reaction conditions were as follows: 95°C for 5 min
heating; 40 cycles of 95°C for 10 sec; 59°C for 40 sec; and 72°C
for 1 min for application. The 2−∆∆Cq method was used
for quantification of the relative expression levels of miR-194 in
the samples of various groups (28).
Western blot analysis
Total protein was extracted from breast cancer cell
lines following a 48-h transfection using the
ProteoPrep® Total Extraction Sample kit (Sigma-Aldrich;
Merck Millipore, Darmstadt, Germany). The protein concentrations
were evaluated using an enhanced Bicinchoninic Acid assay kit
(Beyotime Institute of Biotechnology, Haimen, China), according to
the manufacturer's instructions. Equal concentrations of the
extracted proteins and the 5X SDS-PAGE sample loading buffer were
then heated at 95°C for 10 min. Equal quantities of the protein
samples (25 µg/lane) were loaded onto the gel for 10% SDS-PAGE.
Subsequently, the proteins were transferred to a polyvinylidene
fluoride membrane. Following washing with TBS and Tween-20, the
membranes were blocked with 5% nonfat milk and 1% Tween-20 in TBS
at room temperature for 2 h, and subsequently incubated with the
following primary antibodies: Anti-Fbxw-7 (cat. no. ab74054;
1:1,000), anti-cyclin D (cat. no. ab7958; 1:500), anti-cyclin E
(cat. no. ab7958; 1:500) and β-actin (cat. no. ab8227; 1:5,000).
All primary antibodies were purchased from Abcam (Cambridge, UK).
Following washing with TBST three times, the membranes were
incubated with the corresponding anti-rabbit IgG (cat. no. A0545;
1:20; Sigma-Aldrich; Merck KGaA) conjugated with horseradish
peroxidase (OriGene Technologies, Inc., Rockville, MD, USA) for 1.5
h at room temperature. The membranes were visualized using an
enhanced chemiluminescence kit (Pierce; Thermo Fisher Scientific,
Inc.).
MTT assay
Breast cancer cells (1×104 cells/well) in
the various groups were seeded into a 96-well plate with 100 µl of
medium/well. The cells were plated in quintuplicate each day.
Following culture at 37°C for 24 h, 20 µl MTT solution (5 mg/ml;
Invitrogen; Thermo Fisher Scientific, Inc.) was added to each well
and incubated at 37°C for 4 h. Subsequently, the supernatant in
each well was removed and 150 µl dimethyl sulfoxide was added in
order to dissolve the formazan crystals. The absorbance wavelengths
were read at 570 nm with a microplate spectrophotometer. The cell
growth curve in each group was drawn according to the optical
density values.
Bioinformatic predictions and
luciferase assay
The prediction of the miR-194 target genes was
performed using TargetScan version 3.1 (29). The results revealed that the 3′-UTR of
Fbxw-7 is complementarily base paired with miR-194. In order
to confirm the prediction, a luciferase assay was performed.
Briefly, 293T cells were seeded (1×104 cells/ml) in a
6-well plate and incubated at room temperature for 24 h.
Subsequently, the cells divided into 3 groups and co-transfected
with PGL3-Fbxw-7 3′UTR, miR-194 or an NC mimic using
Lipofectamine® 2000. Following 40 h, the medium in each
well was discarded and 500 µl cell lysis solution was added.
Following lysis, the supernatant was centrifuged at 15,000 × g for
5 min at room temperature. A total of 100 µl luciferase assay
reagent (Promega Corporation, Madison, WI, USA) was added into 100
µl supernatant from each well. The relative intensity of
fluorescence was determined using a multifunctional microplate
reader (Tecan Group, Ltd., Mannedorf, Switzerland) with 2 sec
measurement intervals at an absorbance wavelength of 366 nm and 10
sec measurement duration.
Statistical analysis
All data were analyzed using SPSS 16.0 (SPSS, Inc.,
Chicago, IL, USA). Data are presented as the mean ± standard
deviation. The miR-194 expression levels and MTT data were
statistically analyzed using a two-tailed Student's t-test.
P<0.05 was considered to indicate a statistically significant
difference.
Results
miR-194 expression associated with the
differentiation degree of breast cancer
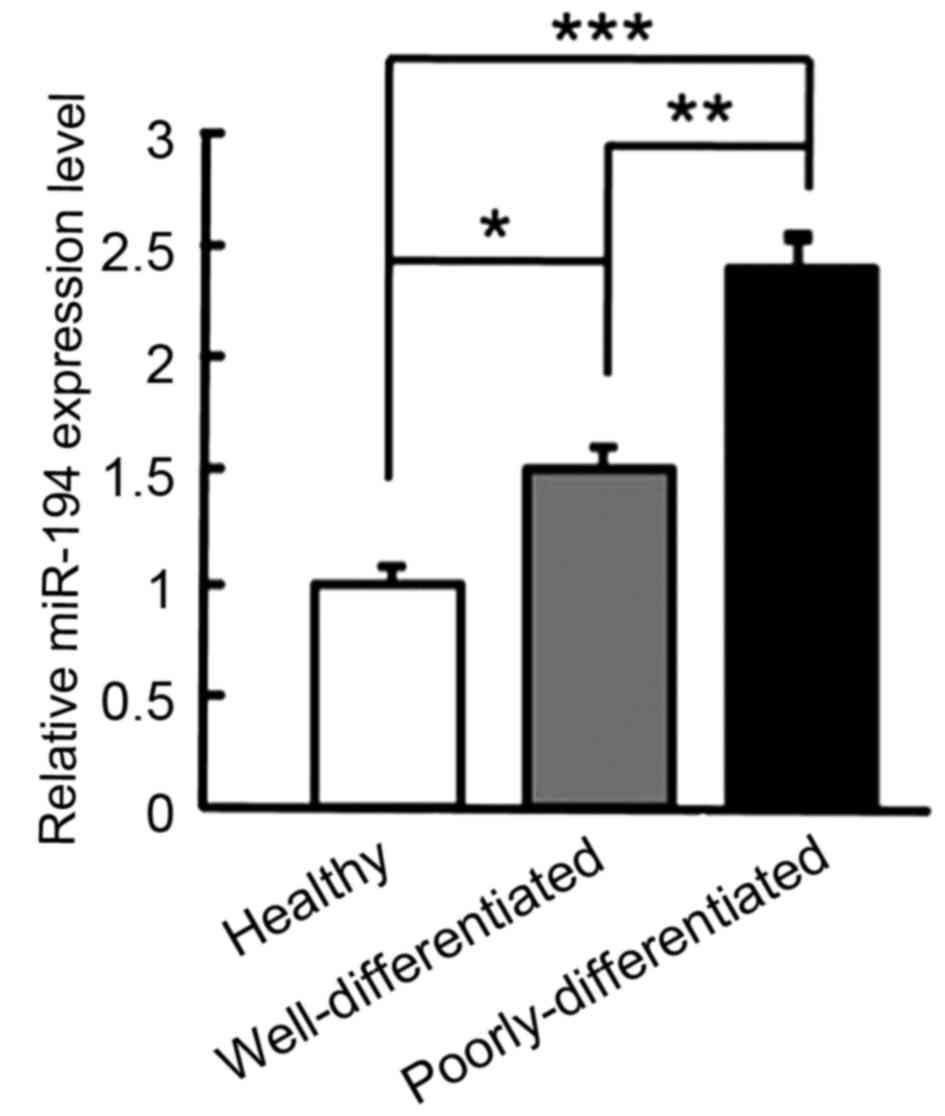
In order to explore the association between miR-194
expression levels and breast cancer, total RNA was extracted from
the sera of 20 healthy adults, 26 patients with well differentiated
breast cancer and 17 patients with poorly differentiated breast
cancer. The results indicated that the serum expression levels of
miR-194 were significantly higher in the well differentiated group
(P<0.05; Fig. 1) and the poorly
differentiated group (P<0.05; Fig.
1), as compared with in the control group. Additionally, the
serum expression level of miR-194 was significantly higher in the
poorly differentiated group, as compared with in the well
differentiated group (P<0.01; Fig.
1). Therefore, the serum expression level of miR-194 was
closely associated with the differentiation degree of breast
cancer, suggesting that miR-194 may be involved in the progression
of breast cancer.
miR-194 expression enhances the
proliferation of breast cancer cells
In order to determine the effect of miR-194
expression on breast cancer cell proliferation, MCF-7 and
MDA-MB-231 breast cancer cell lines were transfected with the
control, miR-194, antagomiR-194 or miR-194/antagomir-194. Following
a 48-h transfection, the cells from the various groups were seeded
into a 96-well plate in triplicate, and then cultured for 24 h.
Subsequently, an MTT assay was performed in order to determine the
proliferation rate of breast cancer cells in the various groups.
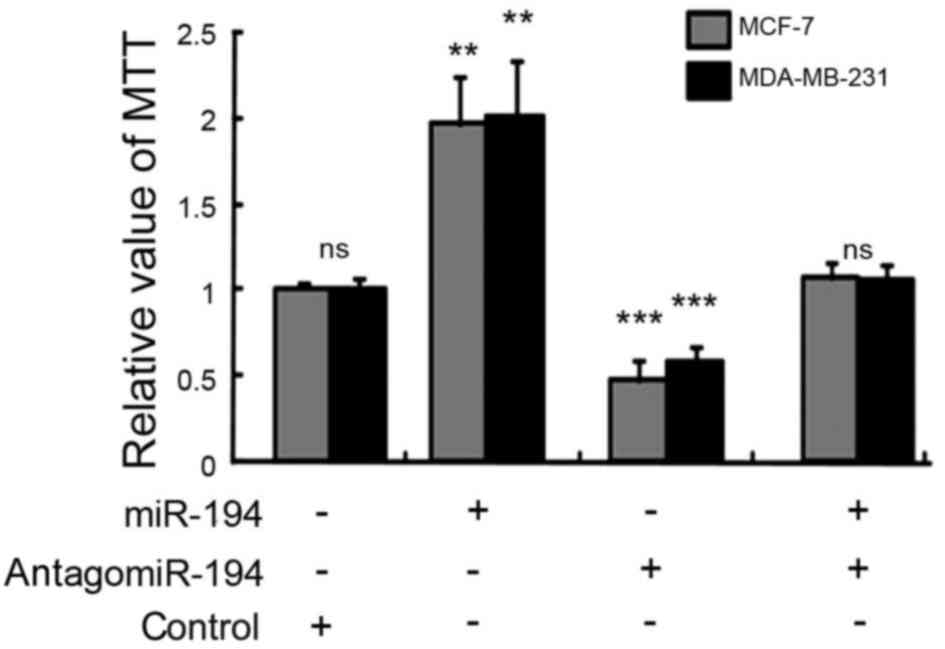
The results demonstrated that the overexpression of miR-194
significantly enhanced the proliferation of MCF-7 (P<0.01;
Fig. 2) and MDA-MB-231 cells
(P<0.01; Fig. 2). Additionally,
the inhibition of miR-194 expression significantly decreased the
proliferation of MCF-7 (P<0.005; Fig.
2) and MDA-MB-231 cells (P<0.005; Fig. 2). No significant change was observed
in the proliferation of cells co-transfected with miR-194 and
antagomiR-194 (Fig. 2).
miR-194 may directly target its
downstream gene, Fbxw-7, in breast cancer cells
According to the bioinformatic database (Targetscan)
analysis and a previous study (30),
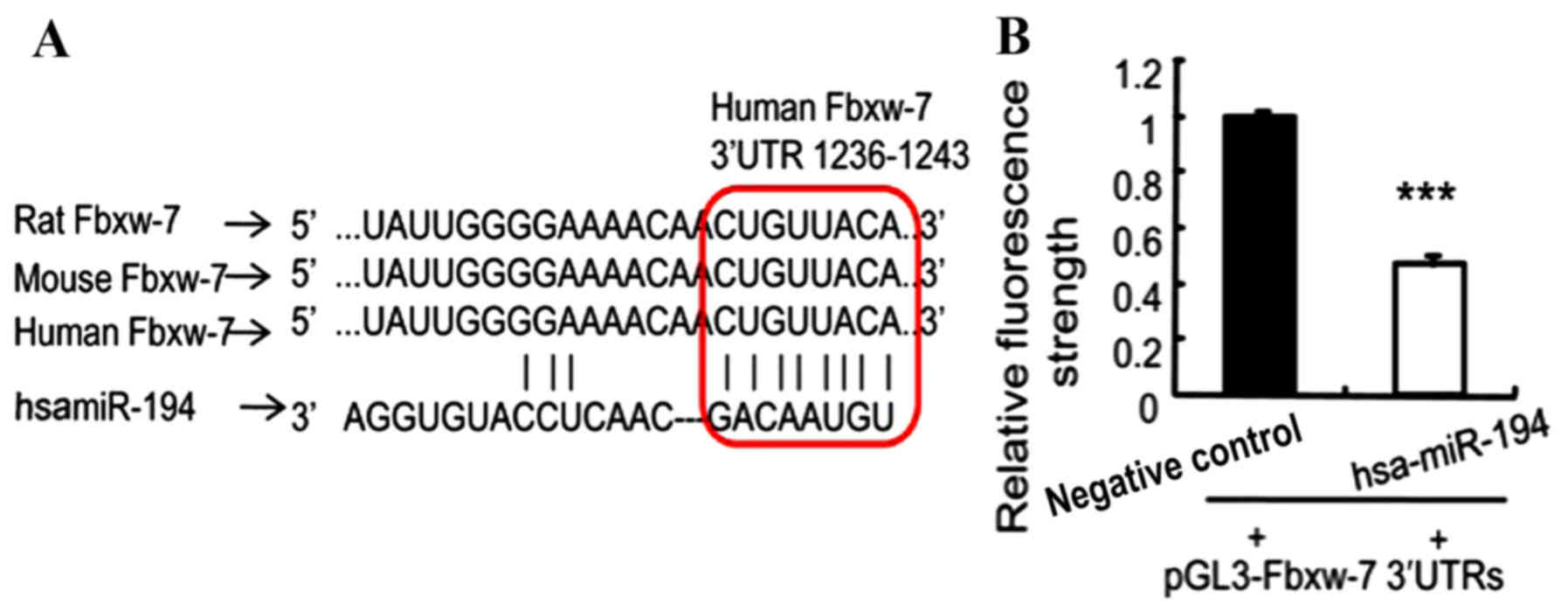
Fbxw-7 may be a potential target of miR194. The binding
sites for miR-194 were detected in the 3′-UTR of Fbxw-7, and
were highly conserved and consistent between rats, mice and humans
(Fig. 3A). Subsequently, the present
study performed a luciferase assay to confirm the prediction made
using the results of the Targetscan analysis. The assay results
revealed that luciferase activity was significantly reduced in
breast cancer cells with overexpressed miR-194, compared with in
the control cells, following transfection of PGL3-Fbxw-7
3′UTRs (P<0.005; Fig. 3B).
miR-194 promotes the expression levels
of cyclin D and cyclin E and inhibits the expression level of
Fbxw-7
It has previously been reported that the inhibition
of Fbxw-7 may increase cell proliferation by increasing the
number of ESCC cells in the S phase and decreasing the number of
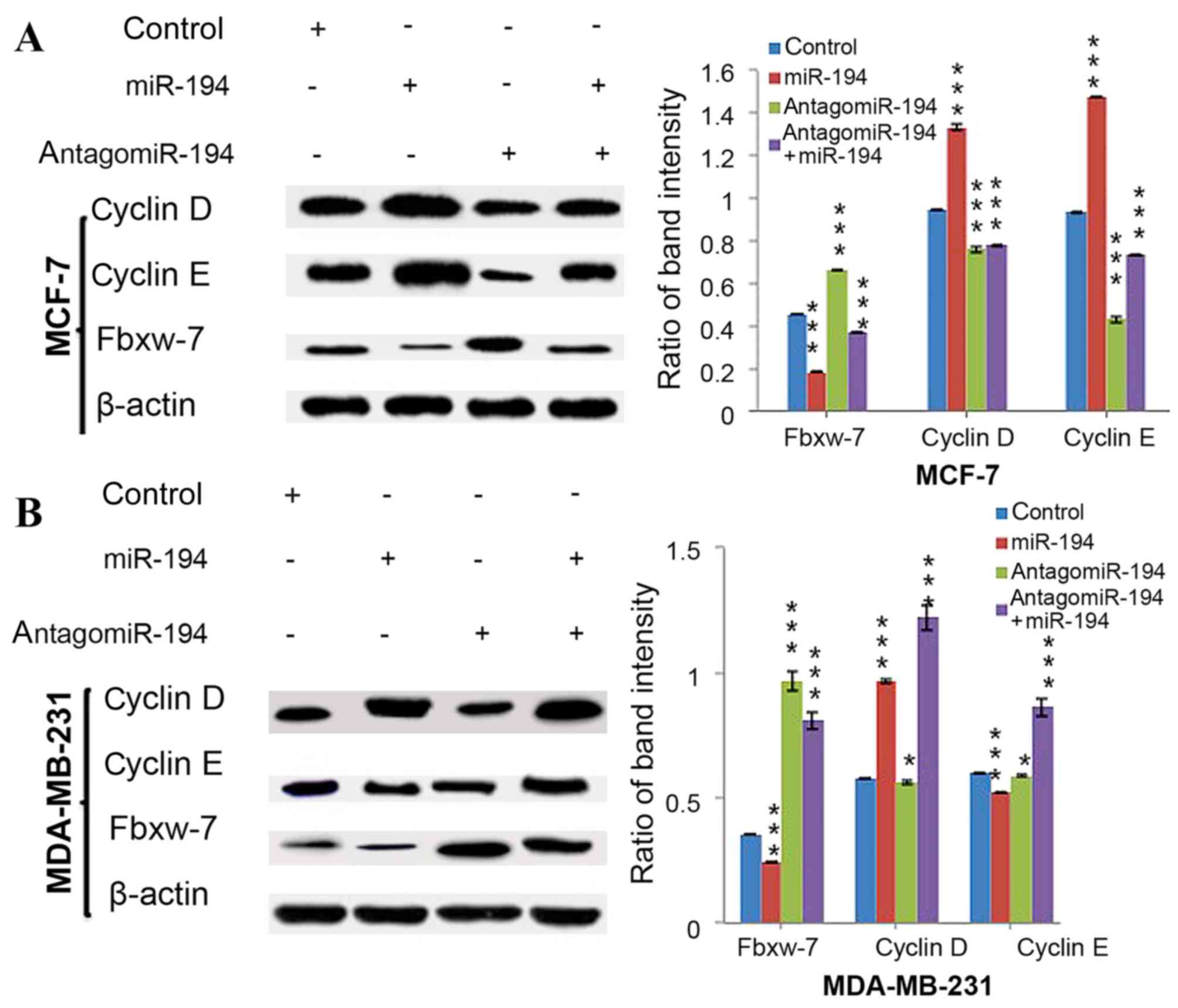
ESCC cells in the G1/G0 phase (31). The present study performed western
blot analysis in order to determine the expression levels of the
cyclin proteins, cyclin D and cyclin E. The results revealed that
miR-194 expression inhibited the expression of Fbxw-7, which
additionally significantly increased the expression levels of
cyclin D and cyclin E in MCF-7 (P<0.05, Fig. 4A) and MDA-MB-231 cells (P<0.05,
Fig. 4B).
Discussion
Previous studies demonstrated that various
expression levels of miRNA have been identified in cancer tissues
(6,32). miRNA may have clinical applications
for use as a targeted therapy or for molecular diagnosis and
prognosis (5,7). The regulatory mechanisms underlying
miRNA function in breast cancer have previously been studied
(4,33–35).
miR-194 was first identified as a suppressor gene in
the hepatic epithelial cells of mice by inhibiting the metastasis
of liver cancer cells (36). In
addition, in cancer of the gut (37,38) and
kidney (39), miR-194 served a
similar inhibitory role. By contrast, it was suggested that miR-194
served a role in the promotion and progression of breast cancer
(9) and pancreatic ductal
adenocarcinoma (40). These studies
indicated that miR-194 had various target genes (31–40).
Previous studies have demonstrated that miR-194 inhibited cell
migration and invasion, which may have contributed to the
anti-tumor activity of trastuzumab on HER2-overexpressing breast
cancer cells (41,42). Iizuka et al (9) demonstrated that the expression level of
miR-194 was increased in human breast cancer cell lines and a rat
mammary cancer model induced by radiation. However, the downstream
pathways of miR-194 in breast cancer are not currently known.
Therefore, it is important to investigate its specific target genes
in breast cancer in order to understand the regulatory mechanisms
underlying miR-194.
In the present study, it was demonstrated that the
E3 ubiquitin ligase Fbxw-7 was the target gene for miR-194
in breast cancer. The results of the bioinformatic prediction
analysis revealed that miR-194 had 367 potential target genes,
which may be associated with the numerous functions of miR-194 in a
variety of cancer subtypes. According to the ranking results of the
bioinformatic analysis and previous studies investigating breast
cancer cell proliferation (41,42), the
present study focused on the target gene of miR-194, Fbxw-7,
which was identified by performing a luciferase assay. Furthermore,
the present study demonstrated that miR-194 promoted the
proliferation of breast cancer cells by targeting Fbxw-7 and
upregulating the expression levels of cyclins D and E. Previous
studies revealed that Fbxw-7 acted as a tumor suppressor
gene in a number of cancer subtypes (16,43). The
mutation or deletion of Fbxw-7 may induce the accumulation
of cancer proliferation genes, including cyclin E, c-Myc and aurora
kinase A (10). A previous study
revealed that the expression level of Fbxw-7 was
significantly reduced in colorectal cancer tissues compared with in
normal tissues, and the inhibition of Fbxw-7 increased the
proliferation of colorectal cancer cells (37). These results are similar to those of
the present study. However, the expression level of miR-194 in the
sera of patients with breast cancer of various differentiation
grades was determined in the present study, and it was revealed
that miR-194 expression is closely associated with the grade of
breast cancer. The serum expression level of miR-194 was highest in
patients with poorly differentiated breast cancer, which suggested
that miR-194 may be a promising prognostic factor for breast
cancer. In the present study, the inhibition of miR-194 expression
by antagomiR-194 decreased the proliferation of breast cancer
cells, which suggested that antagomiR-194 may be a potential
therapeutic drug.
In conclusion, the present study indicated that
miR-194 served an important role in the proliferation of breast
cancer cells by directly inhibiting the expression of Fbxw-7
and regulating the cell cycle. As a result, miR-194 may be a
potential therapeutic target for inhibiting the progression of
breast cancer. Therefore, it is necessary for future studies to
verify the clinical values of miR-194 and determine the effects of
antagomiR-194 as a treatment for breast cancer.
References
|
1
|
Van Diest PJ, van der Wall E and Baak JP:
Prognostic value of proliferation in invasive breast cancer: A
review. J Clin Pathol. 57:675–681. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Yu H, Yang J, Jiao S, Li Y, Li L and Wang
J: A proliferation-inducing ligand expression in breast cancer and
its relationship with prognosis. Nan Fang Yi Ke Da Xue Xue Bao.
35:185–190. 2015.(In Chinese). PubMed/NCBI
|
|
3
|
Wu X, Zeng R, Wu S, Zhong J, Yang L and Xu
J: Comprehensive expression analysis of miRNA in breast cancer at
the miRNA and isomiR levels. Gene. 557:195–200. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Xue J, Chi Y, Chen Y, Huang S, Ye X, Niu
J, Wang W, Pfeffer LM, Shao ZM, Wu ZH and Wu J: MiRNA-621
sensitizes breast cancer to chemotherapy by suppressing FBXO11 and
enhancing p53 activity. Oncogene. 35:448–458. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Zaleska K: miRNA - Therapeutic tool in
breast cancer? Where are we now? Rep Pract Oncol Radiother.
20:79–86. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Fiannaca A, La Rosa M, La Paglia L, Rizzo
R and Urso A: Analysis of miRNA expression profiles in breast
cancer using biclustering. BMC Bioinformatics. 16 Suppl 4:S72015.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
McGuire A, Brown JA and Kerin MJ:
Metastatic breast cancer: The potential of miRNA for diagnosis and
treatment monitoring. Cancer Metastasis Rev. 34:145–155. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Le XF, Almeida MI, Mao W, Spizzo R, Rossi
S, Nicoloso MS, Zhang S, Wu Y, Calin GA and Bast RC Jr: Modulation
of MicroRNA-194 and cell migration by HER2-targeting trastuzumab in
breast cancer. PLoS One. 7:e411702012. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Iizuka D, Imaoka T, Nishimura M, Kawai H,
Suzuki F and Shimada Y: Aberrant microRNA expression in
radiation-induced rat mammary cancer: The potential role of miR-194
overexpression in cancer cell proliferation. Radiat Res.
179:151–159. 2013. View
Article : Google Scholar : PubMed/NCBI
|
|
10
|
Fujii Y, Yada M, Nishiyama M, Kamura T,
Takahashi H, Tsunematsu R, Susaki E, Nakagawa T, Matsumoto A and
Nakayama KI: Fbxw7 contributes to tumor suppression by targeting
multiple proteins for ubiquitin-dependent degradation. Cancer Sci.
97:729–736. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Koepp DM, Schaefer LK, Ye X, Keyomarsi K,
Chu C, Harper JW and Elledge SJ: Phosphorylation-dependent
ubiquitination of cyclin E by the SCFFbw7 ubiquitin ligase.
Science. 294:173–177. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Moberg KH, Bell DW, Wahrer DC, Haber DA
and Hariharan IK: Archipelago regulates Cyclin E levels in
Drosophila and is mutated in human cancer cell lines. Nature.
413:311–316. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
13
|
Strohmaier H, Spruck CH, Kaiser P, Won KA,
Sangfelt O and Reed SI: Human F-box protein hCdc4 targets cyclin E
for proteolysis and is mutated in a breast cancer cell line.
Nature. 413:316–322. 2001. View
Article : Google Scholar : PubMed/NCBI
|
|
14
|
Welcker M, Orian A, Jin J, Grim JE, Harper
JW, Eisenman RN and Clurman BE: The Fbw7 tumor suppressor regulates
glycogen synthase kinase 3 phosphorylation-dependent c-Myc protein
degradation. Proc Natl Acad Sci USA. 101:pp. 9085–9090. 2004;
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Yada M, Hatakeyama S, Kamura T, Nishiyama
M, Tsunematsu R, Imaki H, Ishida N, Okumura F, Nakayama K and
Nakayama KI: Phosphorylation-dependent degradation of c-Myc is
mediated by the F-box protein Fbw7. EMBO J. 23:2116–2125. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Mao JH, Perez-Losada J, Wu D, Delrosario
R, Tsunematsu R, Nakayama KI, Brown K, Bryson S and Balmain A:
Fbxw7/Cdc4 is a p53-dependent, haploinsufficient tumour suppressor
gene. Nature. 432:775–779. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hubbard EJ, Wu G, Kitajewski J and
Greenwald I: sel-10, a negative regulator of lin-12 activity in
Caenorhabditis elegans, encodes a member of the CDC4 family of
proteins. Genes Dev. 11:3182–3193. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gupta-Rossi N, Le Bail O, Gonen H, Brou C,
Logeat F, Six E, Ciechanover A and Israël A: Functional interaction
between SEL-10, an F-box protein, and the nuclear form of activated
Notch1 receptor. J Biol Chem. 276:34371–34378. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Oberg C, Li J, Pauley A, Wolf E, Gurney M
and Lendahl U: The Notch intracellular domain is ubiquitinated and
negatively regulated by the mammalian Sel-10 homolog. J Biol Chem.
276:35847–35853. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wu G, Lyapina S, Das I, Li J, Gurney M,
Pauley A, Chui I, Deshaies RJ and Kitajewski J: SEL-10 is an
inhibitor of notch signaling that targets notch for
ubiquitin-mediated protein degradation. Mol Cell Biol.
21:7403–7415. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Nateri AS, Riera-Sans L, Da Costa C and
Behrens A: The ubiquitin ligase SCFFbw7 antagonizes apoptotic JNK
signaling. Science. 303:1374–1378. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Balamurugan K, Wang JM, Tsai HH, Sharan S,
Anver M, Leighty R and Sterneck E: The tumour suppressor C/EBPδ
inhibits FBXW7 expression and promotes mammary tumour metastasis.
EMBO J. 29:4106–4117. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Fu L, Balasubramanian M, Shan J,
Dudenhausen EE and Kilberg MS: Auto-activation of c-JUN gene by
amino acid deprivation of hepatocellular carcinoma cells reveals a
novel c-JUN-mediated signaling pathway. J Biol Chem.
286:36724–36738. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Snijders AM, Liu Y, Su L, Huang Y and Mao
JH: Expression profiling reveals transcriptional regulation by
Fbxw7/mTOR pathway in radiation-induced mouse thymic lymphomas.
Oncotarget. 6:44794–44805. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Hernandez MA, Patel B, Hey F, Giblett S,
Davis H and Pritchard C: Regulation of BRAF protein stability by a
negative feedback loop involving the MEK-ERK pathway but not the
FBXW7 tumour suppressor. Cell Signal. 28:561–571. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Akhoondi S, Lindström L, Widschwendter M,
Corcoran M, Bergh J, Spruck C, Grandér D and Sangfelt O:
Inactivation of FBXW7/hCDC4-β expression by promoter
hypermethylation is associated with favorable prognosis in primary
breast cancer. Breast Cancer Res. 12:R1052010. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Li J, Guo Y, Liang X, Sun M, Wang G, De W
and Wu W: MicroRNA-223 functions as an oncogene in human gastric
cancer by targeting FBXW7/hCdc4. J Cancer Res Clin Oncol.
138:763–774. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Lewis BP, Shih I-H, Jones-Rhoades MW,
Bartel DP and Burge CB: Prediction of mammalian microRNA targets.
Cell. 115:787–798. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Chen P and Yao GD: The role of cullin
proteins in gastric cancer. Tumor Biol. 37:29–37. 2016. View Article : Google Scholar
|
|
31
|
Wu XZ, Wang KP, Song HJ, Xia JH, Jiang Y
and Wang YL: MiR-27a-3p promotes esophageal cancer cell
proliferation via F-box and WD repeat domain-containing 7 (FBXW7)
suppression. Int J Clin Exp Med. 8:15556–15562. 2015.PubMed/NCBI
|
|
32
|
Liu SG, Qin XG, Zhao BS, Qi B, Yao WJ,
Wang TY, Li HC and Wu XN: Differential expression of miRNAs in
esophageal cancer tissue. Oncol Lett. 5:1639–1642. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Tanić M, Yanowski K, Andrés E, Gómez-López
G, Socorro MR, Pisano DG, Martinez-Delgado B and Benítez J: miRNA
expression profiling of formalin-fixed paraffin-embedded (FFPE)
hereditary breast tumors. Genom Data. 3:75–79. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Hu J, Xu J, Wu Y, Chen Q, Zheng W, Lu X,
Zhou C and Jiao D: Identification of microRNA-93 as a functional
dysregulated miRNA in triple-negative breast cancer. Tumour Biol.
36:251–258. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Wu Z, Wang P, Song C, Wang K, Yan R, Li J
and Dai L: Evaluation of miRNA-binding-site SNPs of MRE11A, NBS1,
RAD51 and RAD52 involved in HRR pathway genes and risk of breast
cancer in China. Mol Genet Genomics. 290:1141–1153. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhang W, Qian P, Zhang X, Zhang M, Wang H,
Wu M, Kong X, Tan S, Ding K, Perry JK, et al: Autocrine/paracrine
human growth hormone-stimulated MicroRNA 96-182-183 cluster
promotes epithelial-mesenchymal transition and invasion in breast
cancer. J Biol Chem. 290:13812–13829. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Meng Z, Fu X, Chen X, Zeng S, Tian Y, Jove
R, Xu R and Huang W: miR-194 is a marker of hepatic epithelial
cells and suppresses metastasis of liver cancer cells in mice.
Hepatology. 52:2148–2157. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Song Y, Zhao F, Wang Z, Liu Z, Chiang Y,
Xu Y, Gao P and Xu H: Inverse association between miR-194
expression and tumor invasion in gastric cancer. Ann Surg Oncol. 19
Suppl 3:S509–S517. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Chen X, Wang Y, Zang W, Du Y, Li M and
Zhao G: miR-194 targets RBX1 gene to modulate proliferation and
migration of gastric cancer cells. Tumour Biol. 36:2393–2401. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Khella HW, Bakhet M, Allo G, Jewett MA,
Girgis AH, Latif A, Girgis H, Von Both I, Bjarnason GA and Yousef
GM: miR-192, miR-194 and miR-215: A convergent microRNA network
suppressing tumor progression in renal cell carcinoma.
Carcinogenesis. 34:2231–2239. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Zhang J, Zhao CY, Zhang SH, Yu DH, Chen Y,
Liu QH, Shi M, Ni CR and Zhu MH: Upregulation of miR-194
contributes to tumor growth and progression in pancreatic ductal
adenocarcinoma. Oncol Rep. 31:1157–1164. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Yokobori T, Mimori K, Iwatsuki M, Ishii H,
Onoyama I, Fukagawa T, Kuwano H, Nakayama KI and Mori M:
p53-Altered FBXW7 expression determines poor prognosis in gastric
cancer cases. Cancer Res. 69:3788–3794. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Iwatsuki M, Mimori K, Ishii H, Yokobori T,
Takatsuno Y, Sato T, Toh H, Onoyama I, Nakayama KI, Baba H and Mori
M: Loss of FBXW7, a cell cycle regulating gene, in colorectal
cancer: Clinical significance. Int J Cancer. 126:1828–1837.
2010.PubMed/NCBI
|