Introduction
Gastric cancer (GC) is the fourth most common
malignant tumor globally (1) and the
third leading cause of cancer-associated mortality worldwide
(2). The highest incidence rate of GC
is found in Eastern Asia, and particularly in China (3). GC is a consequence of multi-factors,
including environmental and genetic factors. Infection with
Helicobacter pylori (H. pylori) is a major risk factor,
accounting for approximately 75% risk (4). It is accepted that H. pylori
causes general inflammatory stress leading to malignancy (5), and activates multiple critical pathways
in gastric epithelial cells (6).
Although H. pylori infection is extremely prevalent, only a
small minority of infected individuals are likely to develop
gastric cancer. Thus, only H. pylori infection is not
sufficient to cause cancer. Previous findings showed that genetic
factors synergize in the development of gastric cancer (4). Therefore, it is of vital importance to
elucidate the molecular pathogenesis of GC in order to identify new
biomarkers and targets for effective therapy.
MicroRNAs (miRNA) are a class of endogenous,
non-coding RNAs that are approximately 20–23 nucleotides in length.
At present, over 1500 miRNAs have been identified in humans. miRNAs
negatively regulate gene expression at the post-transcriptional
level mainly by binding to the 3′-untranslated region (3′-UTR) of a
target mRNA (7). miRNAs are involved
in many cancer-related cell processes, including apoptosis, as well
as migration/invasion and differentiation. miRNAs are aberrantly
expressed in various types of cancer, including gastric, lung,
liver and esophageal cancer (8–11). In
addition, miRNAs function as either oncogenes or tumor suppressors
(12). Increasing evidence has shown
that miRNAs are involved in the oncogenesis and progression of GC.
The microRNA-7/NF-κB signaling regulatory feedback circuit
regulates gastric carcinogenesis (8).
Chemotherapy-induced miRNA-29c/catenin-δ signaling suppresses
metastasis in gastric cancer (13).
MicroRNA-29c mediates the initiation of gastric carcinogenesis by
directly targeting ITGB1 (14).
miR-148a is reported to suppress tumor cell invasion and metastasis
by downregulating ROCK1 (15).
However, the mechanism of elucidating mRNA in gastric cancer
remains to be elucidated.
DNA methylation is an epigenetic modification,
mainly regulated by DNA methyltransferase family, which includes
the subtypes DNMT1, DNMT2 and DNMT3. DNMT1 is considered the most
abundant DNMT and is primarily involved in the maintenance of
methylation during DNA replication (16). DNMT1 is overexpressed in various types
of cancer, including gastric cancer (17). It was previously reported that DNMT1
is crucial in the silencing of several tumor suppressor genes
(18).
In the present study, we aimed to determine and
showed that miR-148a was aberrantly downregulated in gastric cancer
tissues and was significantly correlated with aggressive
clinicopathological characteristics. The re-expression of miR-148a
significantly suppressed migration and invasion in vitro. We
identified DNMT1 as a functional and direct target gene of
miR-148a and DNMT1 attenuated the suppression of miR-148a-medicated
inhibition on cell migration and invasion. Therefore, miR-148a was
able to suppress the migration and invasion of GC by directly
targeting DNMT1, thus, providing a new potential therapeutic target
for GC treatment.
Materials and methods
Tissue samples and cell lines
Sixty patients were involved in the present study.
Gastric cancer tissues and their corresponding non-tumorous gastric
tissues were collected from patients between September 2010 and
July 2015 at the Department of Gastrointestinal Surgery, Weihai
Central Hospital, Weihai, Shandong, China. Human tissues were
immediately frozen in liquid nitrogen and stored at −80°C
refrigerator.
Written informed consent for samples was obtained
from the patients. The study was approved by the Clinical Research
Ethics Committee of Weihai Central Hospital.
Two gastric cancer cell lines (MGC-803, HGC-27), one
normal gastric epithelial cell line (GES-1, as control), and
HEK293T cells were used in the present study. The cells were
cultured in RPMI-1640 medium (MGC-803 and HGC-27) or Dulbecco's
modified Eagle's media (HEK293T and GES-1) supplemented with 10%
FBS (HyClone, South Logan, UT, USA) at 37°C in a humidified air
atmosphere containing 5% CO2.
Cell transfection
MGC-803 cells were plated in 6-well plates and
transfection was performed after 24 h. DNMT1 (plasmid no. 36939)
was purchased from Addgene (Cambridge, MA, USA), and was
transfected into cells using X-tremeGENE HP Transfection Reagent
(Roche Applied Science, Indianapolis, IN, USA). miR-148a mimics or
inhibitor and the control were transfected using Lipofectamine 2000
(Invitrogen, Carlsbad, CA, USA) according to the manufacturer's
protocol. Cells were collected at 48 h after transfection for
reverse transcription-quantitative PCR (RT-qPCR) and western blot
analysis. miR-148a mimics or inhibitor and the control were
purchased from RiboBio (Guangzhou, China).
RT-qPCR
miRNA was extracted from tissue samples and cultured
cells using mirVana™ miRNA isolation kit (Ambion, Austin, TX, USA).
The expression level of miR-148a was quantified with specific
primers and probes using TaqMan miRNA assays (Applied Biosystems,
Carlsbad, CA, USA) and U6 small nuclear RNA was used as an internal
control as per the manufacturer's protocol. The experiments were
repeated three times. Total RNA from cultured cells and tissues was
extracted using TRIzol Reagent (Invitrogen) as per the
manufacturer's protocol. RT-qPCR assays were carried out to detect
DNMT1 relative expression by using the PrimeScript RT Reagent kit
(Takara, Dalian, China) and SYBR Premix Ex Taq (Takara) with the
Roche LightCycler 480 system (Roche, Basel, Switzerland) according
to the manufacturer's protocol. GAPDH was used as an internal
control and was analyzed using the 2−∆∆ct method. The
experiments were repeated three times.
Western blot analysis
Cell proteins were extracted and separated using 10%
SDS-PAGE and then transferred to nitrocellulose filter membranes
(Millipore, Darmstadt, Germany). After blocking in 5% non-fat milk,
the membranes were incubated overnight at 4°C with rabbit
polyclonal anti-DNMT1 antibody (cat. no. ab87654; dilution,
1:1,000; Abcam, Cambridge, MA, USA), and rabbit monoclonal
anti-GAPDH antibody (cat. no. EPR16891; dilution, 1:1,000;
Epitomics, Burlingame, CA, USA). The membranes were subsequently
washed and incubated with goat polyclonal anti-rabbit IgG H&L
secondary antibody (cat. no. ab150077; dilution, 1:2,000; Abcam).
The proteins were visualized using a LI-COR Odyssey Infrared
Imaging System (LI-COR Biosciences, Lincoln, NE, USA).
Luciferase reporter assay
HEK-293T cells were used for the luciferase reporter
assay. Cells were cultured in a 24-well plate and co-transfected
with the pGL3-3′-UTR (500 ng) of DNMT1 or mutated 3′-UTR and
miR-148a expressing vector or negative control vector. The 3′-UTR
was amplified using the primers: Forward, CCGCTCGAGAAATAAAGGAGGAGGA
AGCTGC and reverse, GGGTTTAAACGGTGGTTTATAG GAGAGAT (19). The 3′-UTR mutation was generated using
the QuikChange II XL Site-Directed Mutagenesis kit (Stratagene,
Santa Clara, CA, USA). At 48 h after transfection, the cells were
collected and analyzed using the Dual Luciferase Assay System
(Promega, Fitchburg, WI, USA) according to the manufacturer's
instructions. The relative firefly luciferase activities were
normalized to Renilla luciferase activities. The experiments
were repeated at least three times.
In vitro migration and invasion
assays
For the migration and invasion assay,
5×104 cells were cultured in 24-well plates with 8-µm
pore size chamber inserts (Corning Inc., Corning, NY, USA). For the
migration assay, cells resuspended in 200 µl of serum-free medium
were placed into the upper chamber with the non-coated membrane.
For the invasion assay, the cells were placed into the upper
chamber with the coated membrane. The lower chamber was filled with
10% FBS as a chemoattractant and incubated for 48 h for the
migration assay and 72 h for the invasion assay. The cells on the
upper surface of the membrane were removed and cells on the bottom
surface of the chamber were fixed with 100% methanol for 20 min and
stained with 0.1% crystal violet for 30 min. The assays were
conducted in independent triplicates.
Statistical analysis
Statistical analysis was performed with GraphPad
Prism version 6 for Windows (GraphPad Software Inc., La Jolla, CA,
USA). Data were presented as means ± SD. The difference between two
groups was analyzed using ANOVA and Chi-squared test. Differences
were considered significant for P<0.05.
Results
miR-148a is downregulated in gastric
cancer tissues
To investigate the role of miR-148a in the
progression of gastric cancer, we first analyzed the expression
level of miR-148a in 60 pairs of gastric cancer tissues and their
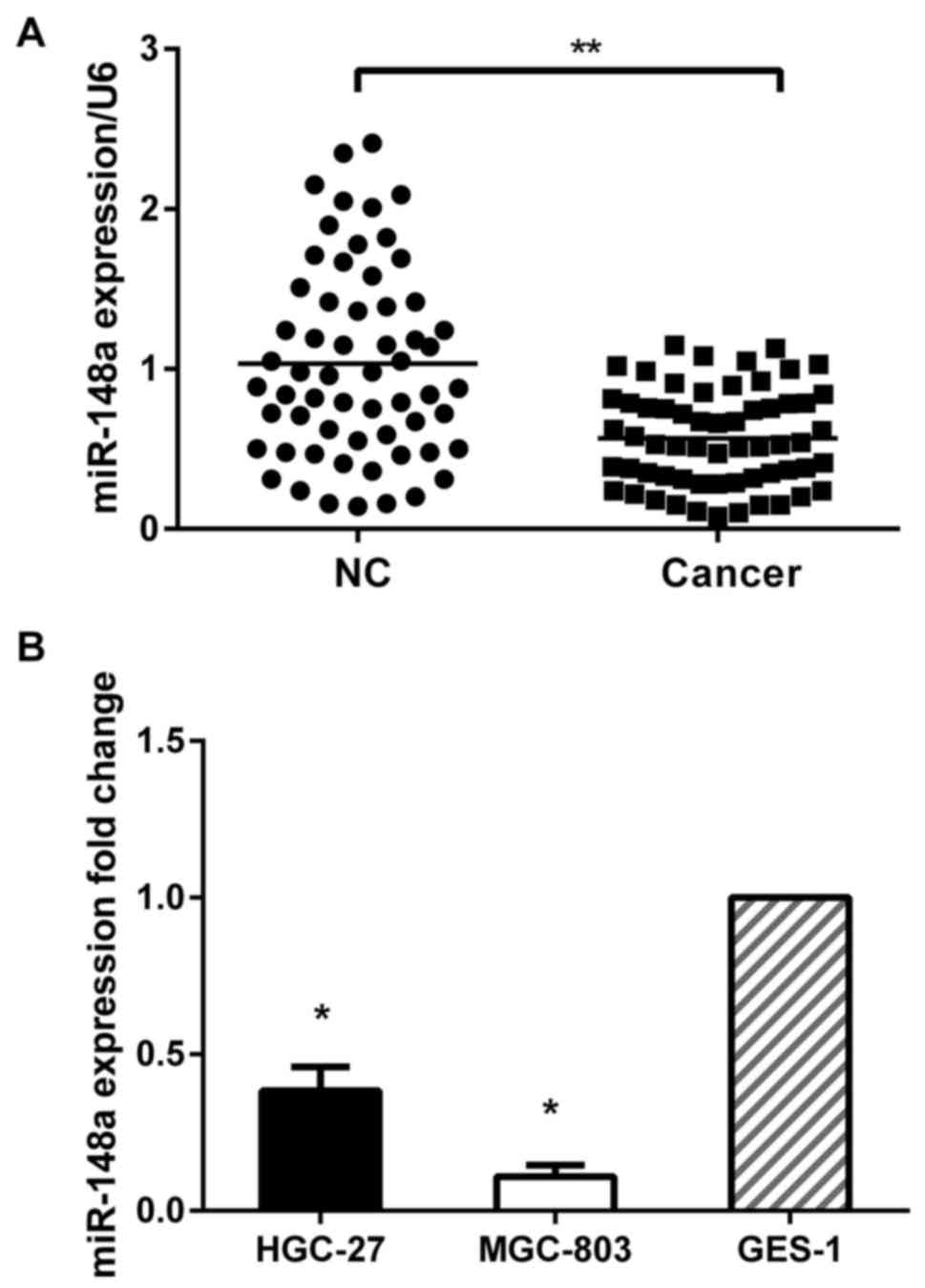
corresponding normal tissues. The RT-qPCR analysis revealed that
the expression of miR-148a was significantly downregulated in
cancer tissues compared with the normal control (P<0.01)
(Fig. 1A). We further analyzed the
relative expression of miR-148a in HGC-27 and MGC-803 cells. The
two cell lines had a low expression of miR-148a. Moreover, MGC-803
had a relatively lower expression of miR-148a than HGC-27 (Fig. 1B).
The clinicopathological analysis revealed that no
significant correlations were observed between the miR-148a
expression level and age, sex, location or tumor size. The low
expression of miR-148a was significantly correlated with lymph-node
metastasis (P<0.01) and gastric cancer with advanced stages
(III+IV) had a lower level of miR-148a than early stages (I+II)
(Table I). These findings suggested
that the downregulation of miR-148a may be involved in gastric
cancer development.
 | Table I.Clinicopathological characteristics
and miR-148a expression in 60 patients with gastric cancer. |
Table I.
Clinicopathological characteristics
and miR-148a expression in 60 patients with gastric cancer.
|
|
| miR-148
expression |
|
|---|
|
|
|
|
|
|---|
| Characteristics | Cases (n=60) | High (n=13) | Low (n=47) | P-valuea |
|---|
| Age (years) |
|
|
| 0.3468 |
| ≥55 | 28 | 8 | 20 |
|
|
<55 | 32 | 5 | 27 |
|
| Sex |
|
|
| 0.2134 |
| Male | 29 | 4 | 25 |
|
|
Female | 31 | 9 | 22 |
|
| Tumor size (cm) |
|
|
| 0.5354 |
| ≥5 | 35 | 7 | 30 |
|
|
<5 | 25 | 6 | 17 |
|
|
Differentiation |
|
|
| 0.9301 |
|
Well | 21 | 4 | 17 |
|
|
Moderately | 18 | 4 | 14 |
|
|
Poor | 21 | 5 | 16 |
|
| TNM stage |
|
|
| 0.0242 |
| I +
II | 24 | 9 | 15 |
|
| III +
IV | 36 | 4 | 32 |
|
| Lymph-node
metastasis |
|
|
| 0.0022 |
|
Yes | 38 | 3 | 35 |
|
| No | 22 | 10 | 12 |
|
Re-expression of miR-148a suppresses
gastric cancer migration and invasion in vitro
To understand better the biological function of
miR-148a on the development of gastric cancer, we re-expressed
miR-148a in MGC-803 cells. Transwell assays were performed to
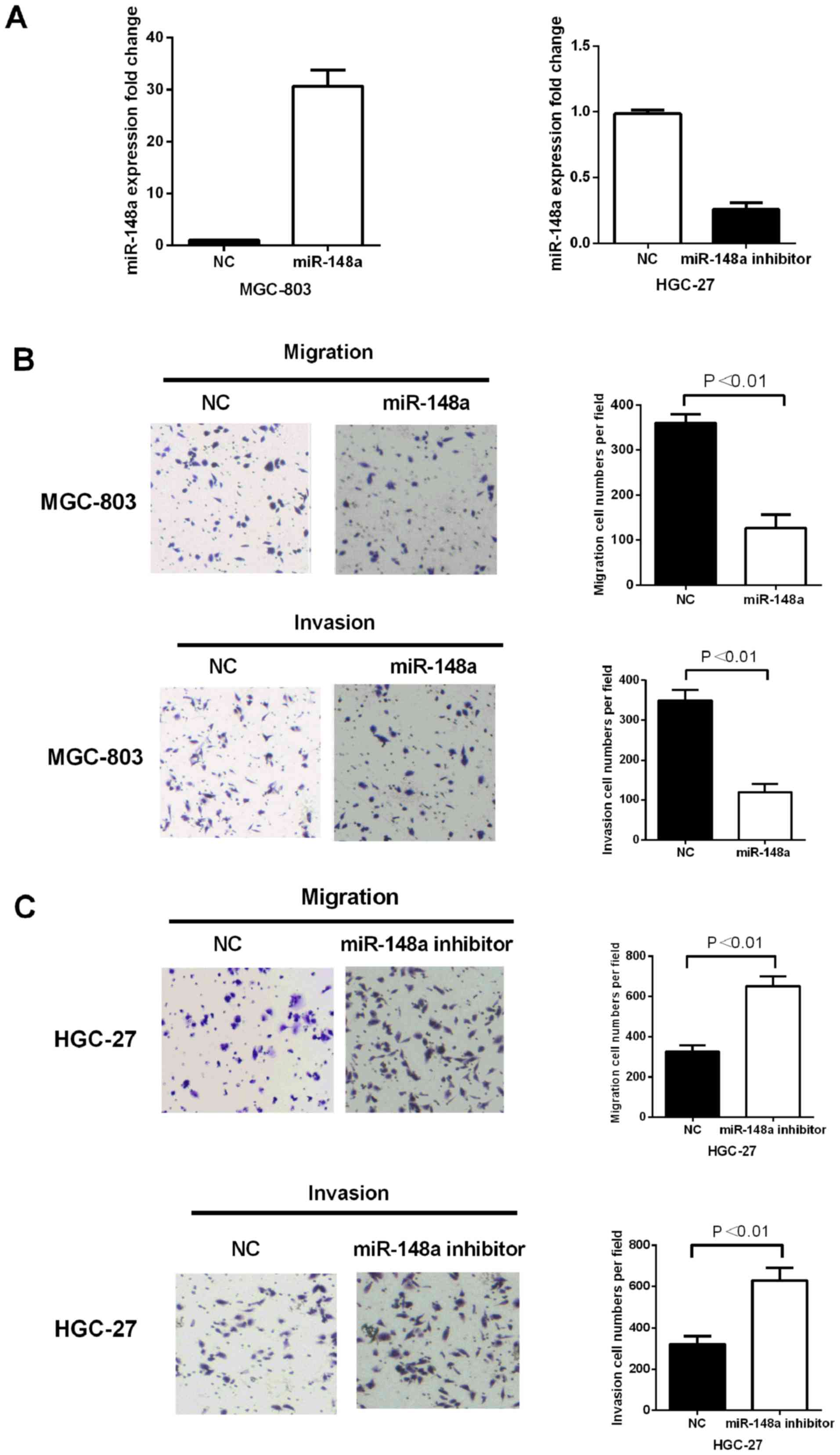
detect migratory and invasive ability induced by miR-148a. The
successful re-expression of mature miR-148a in MGC-803 cells was
confirmed by RT-qPCR (Fig. 2A). For
the migration assay, the re-expression of miR-148a significantly
reduced the migratory ability and cell numbers in MGC-803 cells
(P<0.01) (Fig. 2B). For the
invasion assay, the re-expression of miR-148a significantly
decreased the invasive ability and cell numbers in MGC-803 cells
(P<0.01) (Fig. 2B). Furthermore,
deletion of miR-148a by a specific miR-148a inhibitor increased
cell migratory and invasive ability in GC cell lines (P<0.01)
(Fig. 2C).
miR-148a downregulates DNMT1
expression by directly targeting its 3′-UTR
To explore downstream targets of miR-148a, two
online algorithms, TargetScan and miRanda, were used to predict its
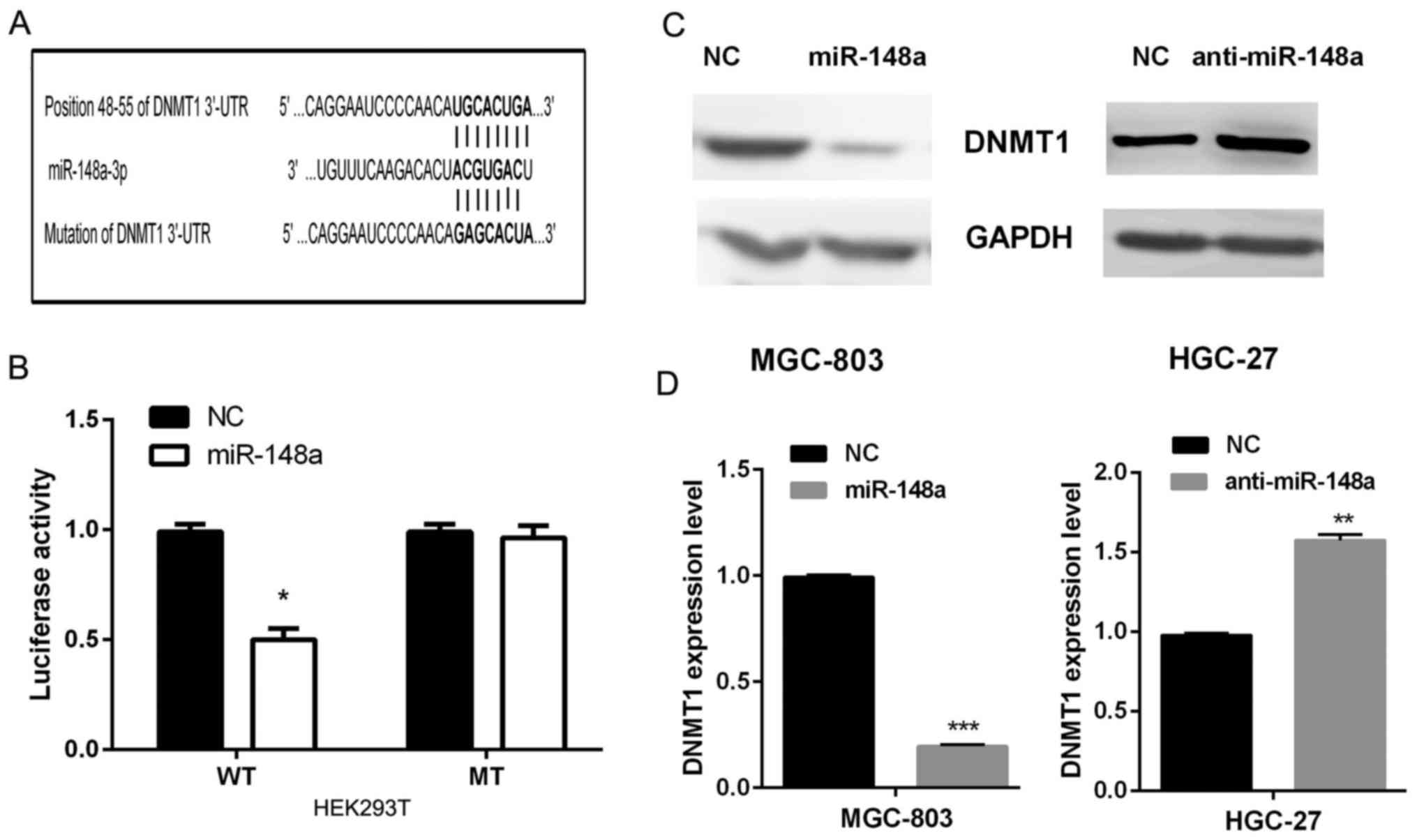
targets. DNMT1 was identified as one candidate target (Fig. 3A). To verify whether miR-148a could
bind to 3′-UTR of DNMT1, luciferase report assays were performed in
HEK293T cells. Our results showed that the luciferase activity of
the wild-type (WT) 3′-UTR of DNMT1 was significantly decreased in
cells co-transfected with miR-148a compared with that in miRNA
control. While the luciferase activity of the mutated type (MT)
3′-UTR of DNMT1 was not obviously altered in cells co-transfected
with miR-148a compared with that in miRNA control (Fig. 3B). Additionally, we detected whether
the expression of DNMT1 is regulated by miR-148a. In MGC-803 cells,
the re-expression of miR-148a reduced the expression of DNMT1 at
the protein and mRNA level, and in HGC-27 cells, the miR-148a
inhibitor increased the level of DNMT1 at protein and mRNA levels
(Fig. 3C and D).
DNMT1 is involved in cell migration
and invasion
As miR-148a is downregulated in gastric cancer and
targets DNMT1 by binding to its 3′-UTR, we then examined whether
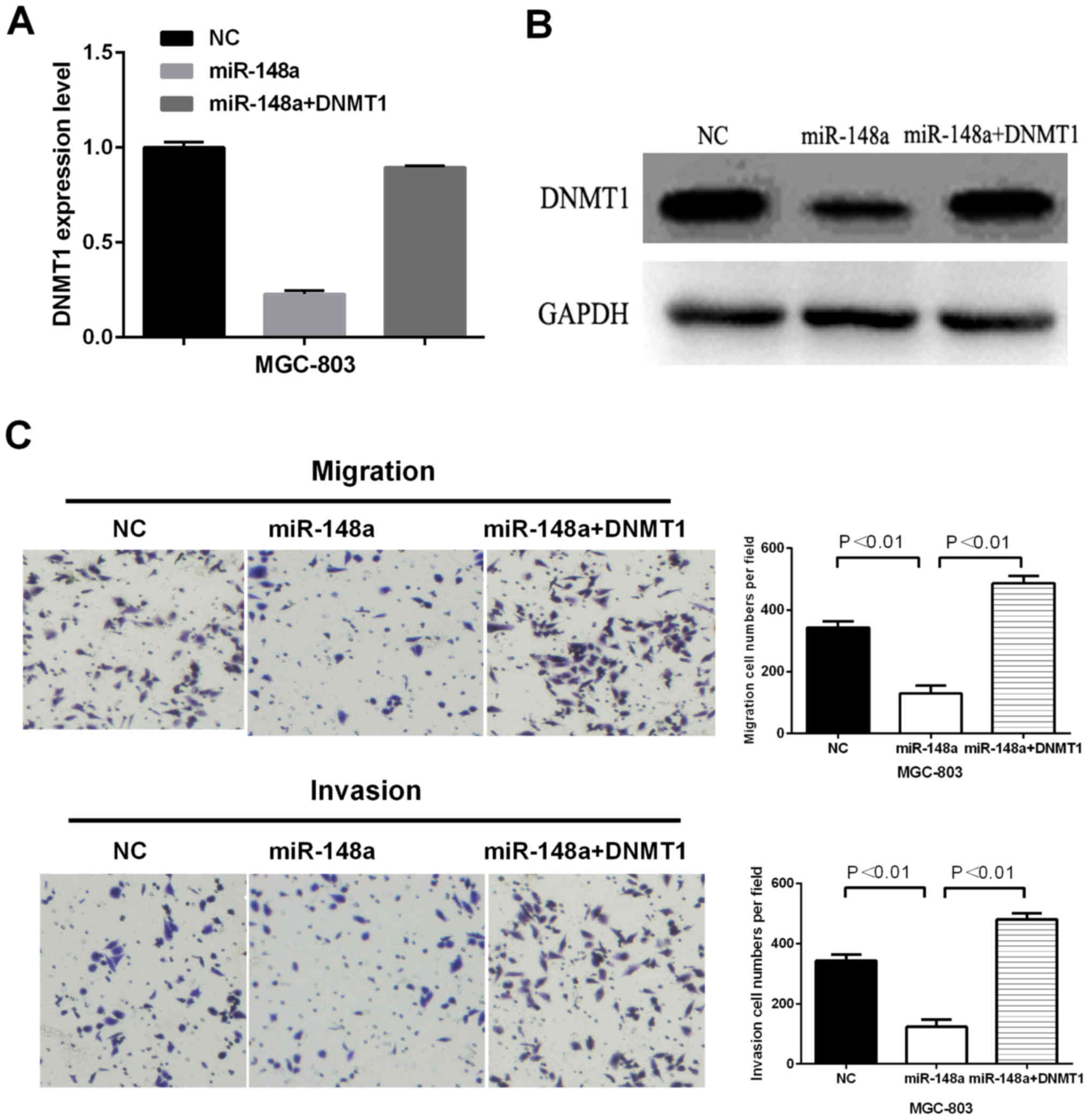
DNMT1 is involved in cell migration and invasion. MGC-803 cells
were co-transfected with DNMT1 overexpression plasmid and miR-148a
and then detected the mRNA and protein levels using RT-qPCR and
western blot analysis. As shown in Fig.
4A and B, the downregulation of DNMT1 mediated by miR-148a was
partially reversed by the ectopic expression of DNMT1. The
transwell assay showed that the ectopic expression of DNMT1
partially attenuated miR-148-mediated inhibition on cell migratory
and invasive abilities (Fig. 4C).
Moreover, the relative cell numbers were presented beside the
images. Thus, DNMT1 may play an important role in gastric cancer
cell migration and invasion.
Discussion
In the present study, we showed that miR-148a was
aberrantly downregulated in GC tissues and was significantly
correlated with lymph node metastasis and gastric cancer with
advanced stages (III+IV) had a lower level of miR-148a than early
stages (I+II). Re-expression of miR-148a was observed to suppress
gastric cancer migration and invasion in vitro. In addition,
DNMT1 was identified as a functional and direct target of miR-148a.
Our results suggest that miR-148a plays important roles in
suppressing tumor progression in GC by directly targeting
DNMT1.
miRNAs perform as tumor suppressors or oncomiR. It
is reported that miR-148a is downregulated in various types of
cancer, including gastric cancer (15), and non-small cell lung cancer
(20), suggesting that miR-148a may
play a key role in tumorigenesis and tumor progression. In early
pancreatic carcinogenesis, it is demonstrated that the silencing of
miR-148a was induced by DNA hypermethylation, which was regulated
by DNA methyltransferase-1 (21). In
gastric cancer, it was revealed that the downregulation of miR-148a
was significantly correlated with an advanced clinical stage, lymph
node metastasis, and poor clinical outcome (22). In addition, the ectopic expression of
miR-148a was validated by inhibiting tumor cell proliferation and
migration in vitro, and inhibiting tumor formation in
vivo (23).
miRNAs perform their function by downregulating the
expression of their target mRNAs at the post-transcriptional level
(24). Many target genes are involved
in the promotion of cancer progression. Song et al reported
that miR-148b can suppress cell growth by targeting cholecystokinin
B receptor (CCK-BR) in colorectal cancer (25). Yu et al revealed that miR-148a
functions as a tumor suppressor by targeting CCK-BR through the
inactivation of STAT3 and Akt in human gastric cancer (23). Sakamoto et al demonstrated that
microRNA-148a is downregulated in gastric cancer and targets MMP7
(22). By contrast, in this study,
DNMT1 was identified as a functional target of miR-148a. The
re-expression of miR-148a reduced DNMT1 expression in gastric
cancer. In addition, the downregulation of miR-148a increased DNMT1
expression. The luciferase report assay showed that luciferase
activity was recovered from mutated-type 3′-UTR construct.
Furthermore, we revealed that DNMT1 upregulated cell migration and
invasion, and the upregulation of DNMT1 attenuates
miR-148a-mediated inhibition on cell migration and invasion. DNMT1
as a number of DNA methyltransferase, functions as an inhibitor of
several suppressor genes in various types of cancer. Hino et
al demonstrated that the activation of DNA methyltransferase 1
by EBV latent membrane protein 2A leads to promoter
hypermethylation of PTEN gene in gastric carcinoma (26). Zhang et al suggested that the
overexpression of DNMT1 induces tumor suppressor gene
hypermethylation via the Akt-NFκB pathway in gastric cancer
development (27). A high DNMT
protein expression was identified in 83% of primary gastric cancer
patients as detected by Mutze et al (28). DNMT1 expression was also significantly
associated with tumor differentiation, and the overexpression of
DNMT1 was associated with poor outcome after treatment (29). DNMT1 inhibitors have been demonstrated
to be useful as sensitizers for various chemotherapeutic agents or
radiation in different cell lines including gastric cancer cell
lines (29). Collectively DNMT1 was
identified as an important target for gastric cancer development
and treatment.
In conclusion, the present findings show that
miR-148a suppresses cell invasion and migration in gastric cancer
by regulating DNMT1 expression. The miR-148a/DNMT1 axis can be
considered a prospective novel target for gastric cancer
therapy.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
HS conceived and designed the study and wrote the
manuscript; XC contributed significantly to the analysis and
manuscript preparation; HJ performed the data analyses; XW helped
perform the analysis with constructive discussions; HY performed
the histological examination of the stomach; PS sorted out
experimental data; XS contributed to the conception of the study.
All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Written informed consent for samples was obtained
from the patients. The study was approved by the Clinical Research
Ethics Committee of Weihai Central Hospital.
Consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ferro A, Peleteiro B, Malvezzi M, Bosetti
C, Bertuccio P, Levi F, Negri E, La Vecchia C and Lunet N:
Worldwide trends in gastric cancer mortality (1980–2011), with
predictions to 2015, and incidence by subtype. Eur J Cancer.
50:1330–1344. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Siegel R, Ma J, Zou Z and Jemal A: Cancer
statistics, 2014. CA Cancer J Clin. 64:9–29. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bertuccio P, Chatenoud L, Levi F, Praud D,
Ferlay J, Negri E, Malvezzi M and La Vecchia C: Recent patterns in
gastric cancer: A global overview. Int J Cancer. 125:666–673. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Polk DB and Peek RM Jr: Helicobacter
pylori: Gastric cancer and beyond. Nat Rev Cancer. 10:403–414.
2010. View
Article : Google Scholar : PubMed/NCBI
|
|
5
|
Fox JG, Rogers AB, Whary MT, Ge Z, Ohtani
M, Jones EK and Wang TC: Accelerated progression of gastritis to
dysplasia in the pyloric antrum of TFF2 −/− C57BL6 × Sv129
Helicobacter pylori-infected mice. Am J Pathol. 171:1520–1528.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ding SZ, Goldberg JB and Hatakeyama M:
Helicobacter pylori infection, oncogenic pathways and epigenetic
mechanisms in gastric carcinogenesis. Future Oncol. 6:851–862.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Bartel DP: MicroRNAs: Target recognition
and regulatory functions. Cell. 136:215–233. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Zhao XD, Lu YY, Guo H, Xie HH, He LJ, Shen
GF, Zhou JF, Li T, Hu SJ, Zhou L, et al: MicroRNA-7/NF-κB signaling
regulatory feedback circuit regulates gastric carcinogenesis. J
Cell Biol. 210:613–627. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Cui R, Meng W, Sun HL, Kim T, Ye Z, Fassan
M, Jeon YJ, Li B, Vicentini C, Peng Y, et al: MicroRNA-224 promotes
tumor progression in nonsmall cell lung cancer. Proc Natl Acad Sci
USA. 112:pp. E4288–E4297. 2015; View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ohta K, Hoshino H, Wang J, Ono S, Iida Y,
Hata K, Huang SK, Colquhoun S and Hoon DS: MicroRNA-93 activates
c-Met/PI3K/Akt pathway activity in hepatocellular carcinoma by
directly inhibiting PTEN and CDKN1A. Oncotarget. 6:3211–3224. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Liu R, Gu J, Jiang P, Zheng Y, Liu X,
Jiang X, Huang E, Xiong S, Xu F, Liu G, et al: DNMT1-microRNA126
epigenetic circuit contributes to esophageal squamous cell
carcinoma growth via ADAM9-EGFR-AKT signaling. Clin Cancer Res.
21:854–863. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Croce CM: Causes and consequences of
microRNA dysregulation in cancer. Nat Rev Genet. 10:704–714. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Wang Y, Liu C, Luo M, Zhang Z, Gong J, Li
J, You L, Dong L, Su R, Lin H, et al: Chemotherapy-Induced
miRNA-29c/Catenin-δ Signaling Suppresses Metastasis in Gastric
Cancer. Cancer Res. 75:1332–1344. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Han TS, Hur K, Xu G, Choi B, Okugawa Y,
Toiyama Y, Oshima H, Oshima M, Lee HJ, Kim VN, et al: MicroRNA-29c
mediates initiation of gastric carcinogenesis by directly targeting
ITGB1. Gut. 64:203–214. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Zheng B, Liang L, Wang C, Huang S, Cao X,
Zha R, Liu L, Jia D, Tian Q, Wu J, et al: MicroRNA-148a suppresses
tumor cell invasion and metastasis by downregulating ROCK1 in
gastric cancer. Clin Cancer Res. 17:7574–7583. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Robertson KD, Keyomarsi K, Gonzales FA,
Velicescu M and Jones PA: Differential mRNA expression of the human
DNA methyltransferases (DNMTs) 1, 3a and 3b during the G(0)/G(1) to
S phase transition in normal and tumor cells. Nucleic Acids Res.
28:2108–2113. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
He M, Fan J, Jiang R, Tang WX and Wang ZW:
Expression of DNMTs and genomic DNA methylation in gastric signet
ring cell carcinoma. Mol Med Rep. 8:942–948. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Hermann A, Gowher H and Jeltsch A:
Biochemistry and biology of mammalian DNA methyltransferases. Cell
Mol Life Sci. 61:2571–2587. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Jiang C and Fang G: MiR-148a promotes
myocardial differentiation of human bone mesenchymal stromal cells
via DNA methyltransferase 1 (DNMT1). Cell Biol Int. Jun
28–2017.(Epub ahead of print). View Article : Google Scholar
|
|
20
|
Joshi P, Jeon YJ, Laganà A, Middleton J,
Secchiero P, Garofalo M and Croce CM: MicroRNA-148a reduces
tumorigenesis and increases TRAIL-induced apoptosis in NSCLC. Proc
Natl Acad Sci USA. 112:pp. 8650–8655. 2015; View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Hanoun N, Delpu Y, Suriawinata AA, Bournet
B, Bureau C, Selves J, Tsongalis GJ, Dufresne M, Buscail L,
Cordelier P, et al: The silencing of microRNA 148a production by
DNA hypermethylation is an early event in pancreatic
carcinogenesis. Clin Chem. 56:1107–1118. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Sakamoto N, Naito Y, Oue N, Sentani K,
Uraoka N, Zarni Oo H, Yanagihara K, Aoyagi K, Sasaki H and Yasui W:
MicroRNA-148a is downregulated in gastric cancer, targets MMP7, and
indicates tumor invasiveness and poor prognosis. Cancer Sci.
105:236–243. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Yu B, Lv X, Su L, Li J, Yu Y, Gu Q, Yan M,
Zhu Z and Liu B: MiR-148a functions as a tumor suppressor by
targeting CCK-BR via inactivating STAT3 and Akt in human gastric
cancer. PLoS One. 11:e01589612016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Fiedler J, Batkai S and Thum T:
MicroRNA-based therapy in cardiology. Herz. 39:194–200. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Song Y, Xu Y, Wang Z, Chen Y, Yue Z, Gao
P, Xing C and Xu H: MicroRNA-148b suppresses cell growth by
targeting cholecystokinin-2 receptor in colorectal cancer. Int J
Cancer. 131:1042–1051. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Hino R, Uozaki H, Murakami N, Ushiku T,
Shinozaki A, Ishikawa S, Morikawa T, Nakaya T, Sakatani T, Takada
K, et al: Activation of DNA methyltransferase 1 by EBV latent
membrane protein 2A leads to promoter hypermethylation of PTEN gene
in gastric carcinoma. Cancer Res. 69:2766–2774. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zhang BG, Hu L, Zang MD, Wang HX, Zhao W,
Li JF, Su LP, Shao Z, Zhao X, Zhu ZG, et al: Helicobacter pylori
CagA induces tumor suppressor gene hypermethylation by upregulating
DNMT1 via AKT-NFκB pathway in gastric cancer development.
Oncotarget. 7:9788–9800. 2016.PubMed/NCBI
|
|
28
|
Mutze K, Langer R, Schumacher F, Becker K,
Ott K, Novotny A, Hapfelmeier A, Höfler H and Keller G: DNA
methyltransferase 1 as a predictive biomarker and potential
therapeutic target for chemotherapy in gastric cancer. Eur J
Cancer. 47:1817–1825. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Qiu H, Yashiro M, Shinto O, Matsuzaki T
and Hirakawa K: DNA methyltransferase inhibitor 5-aza-CdR enhances
the radiosensitivity of gastric cancer cells. Cancer Sci.
100:181–188. 2009. View Article : Google Scholar : PubMed/NCBI
|