|
1
|
Xu YJ, Du Y and Fan Y: Long noncoding RNAs
in lung cancer: What we know in 2015. Clin Transl Oncol.
18:660–665. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kang CG, Lee HJ, Kim SH and Lee EO:
Zerumbone suppresses osteopontin-induced cell invasion through
inhibiting the FAK/AKT/ROCK pathway in human non-small cell lung
cancer A549 cells. J Nat Prod. 79:156–160. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jemal A, Murray T, Ward E, Samuels A,
Tiwari RC, Ghafoor A, Feuer EJ and Thun MJ: Cancer statistics,
2005. CA Cancer J Clin. 55:10–30. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Xu G, Chen J, Pan Q, Huang K, Pan J, Zhang
W, Chen J, Yu F, Zhou T and Wang Y: Long noncoding RNA expression
profiles of lung adenocarcinoma ascertained by microarray analysis.
PLoS One. 9:e1040442014. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Choe C, Shin YS, Kim C, Choi SJ, Lee J,
Kim SY, Cho YB and Kim J: Crosstalk with cancer-associated
fibroblasts induces resistance of non-small cell lung cancer cells
to epidermal growth factor receptor tyrosine kinase inhibition.
Onco Targets Ther. 8:3665–3678. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ferlay J, Shin HR, Bray F, Forman D,
Mathers C and Parkin DM: Estimates of worldwide burden of cancer in
2008: GLOBOCAN 2008. Int J Cancer. 127:2893–2917. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Chen G, Umelo IA, Lv S, Teugels E, Fostier
K, Kronenberger P, Dewaele A, Sadones J, Geers C and De Grève J:
miR-146a inhibits cell growth, cell migration and induces apoptosis
in non-small cell lung cancer cells. PLoS One. 8:e603172013.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Richards EJ, Permuth-Wey J, Li Y, Chen YA,
Coppola D, Reid BM, Lin HY, Teer JK, Berchuck A, Birrer MJ, et al:
A functional variant in HOXA11-AS, a novel long non-coding RNA,
inhibits the oncogenic phenotype of epithelial ovarian cancer.
Oncotarget. 6:34745–34757. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Wang Q, Zhang J, Liu Y, Zhang W, Zhou J,
Duan R, Pu P, Kang C and Han L: A novel cell cycle-associated
lncRNA, HOXA11-AS, is transcribed from the 5-prime end of the HOXA
transcript and is a biomarker of progression in glioma. Cancer
Lett. 373:251–259. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hu B, Zhang H, Wang Z, Zhang F, Wei H and
Li L: LncRNA CCAT1/miR-130a-3p axis increases cisplatin resistance
in non-small-cell lung cancer cell line by targeting SOX4. Cancer
Biol Ther. Oct 11–2017.(Epub ahead of print). View Article : Google Scholar
|
|
12
|
Qu CH, Sun QY, Zhang FM and Jia YM: Long
non-coding RNA ROR is a novel prognosis factor associated with
non-small-cell lung cancer progression. Eur Rev Med Pharmacol Sci.
21:4087–4091. 2017.PubMed/NCBI
|
|
13
|
Wilusz JE: Long noncoding RNAs: Re-writing
dogmas of RNA processing and stability. Biochim Biophys Acta.
1859:128–138. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Yuan X, Wang J, Tang X, Li Y, Xia P and
Gao X: Berberine ameliorates nonalcoholic fatty liver disease by a
global modulation of hepatic mRNA and lncRNA expression profiles. J
Transl Med. 13:242015. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Ponting CP, Oliver PL and Reik W:
Evolution and functions of long noncoding RNAs. Cell. 136:629–641.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Wei Y and Niu B: Role of MALAT1 as a
prognostic factor for survival in various cancers: A systematic
review of the literature with meta-analysis. Dis Markers.
2015:1646352015. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Wang X, Song X, Glass CK and Rosenfeld MG:
The long arm of long noncoding RNAs: Roles as sensors regulating
gene transcriptional programs. Cold Spring Harb Perspect Biol.
3:a0037562011. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Liz J and Esteller M: lncRNAs and
microRNAs with a role in cancer development. Biochim Biophys Acta.
1859:169–176. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Li DS, Ainiwaer JL, Sheyhiding I, Zhang Z
and Zhang LW: Identification of key long non-coding RNAs as
competing endogenous RNAs for miRNA-mRNA in lung adenocarcinoma.
Eur Rev Med Pharmacol Sci. 20:2285–2295. 2016.PubMed/NCBI
|
|
20
|
Xie X, Pan J, Wei L, Wu S, Hou H, Li X and
Chen W: Gene expression profiling of microRNAs associated with UCA1
in bladder cancer cells. Int J Oncol. 48:1617–1627. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Liang WC, Fu WM, Wong CW, Wang Y, Wang WM,
Hu GX, Zhang L, Xiao LJ, Wan DC, Zhang JF and Waye MM: The lncRNA
H19 promotes epithelial to mesenchymal transition by functioning as
miRNA sponges in colorectal cancer. Oncotarget. 6:22513–22525.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Salmena L, Poliseno L, Tay Y, Kats L and
Pandolfi PP: A ceRNA hypothesis: The rosetta stone of a hidden RNA
language? Cell. 146:353–358. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Sun C, Li S, Zhang F, Xi Y, Wang L, Bi Y
and Li D: Long non-coding RNA NEAT1 promotes non-small cell lung
cancer progression through regulation of miR-377-3p-E2F3 pathway.
Oncotarget. 7:51784–51814. 2016.PubMed/NCBI
|
|
24
|
Lu L, Xu H, Luo F, Liu X, Lu X, Yang Q,
Xue J, Chen C, Shi L and Liu Q: Epigenetic silencing of miR-218 by
the lncRNA CCAT1, acting via BMI1, promotes an altered cell cycle
transition in the malignant transformation of HBE cells induced by
cigarette smoke extract. Toxicol Appl Pharmacol. 304:30–41. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Nie W, Ge HJ, Yang XQ, Sun X, Huang H, Tao
X, Chen WS and Li B: LncRNA-UCA1 exerts oncogenic functions in
non-small cell lung cancer by targeting miR-193a-3p. Cancer Lett.
371:99–106. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Keniry A, Oxley D, Monnier P, Kyba M,
Dandolo L, Smits G and Reik W: The H19 lincRNA is a developmental
reservoir of miR-675 that suppresses growth and Igf1r. Nat Cell
Biol. 14:659–665. 2012. View
Article : Google Scholar : PubMed/NCBI
|
|
27
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
He L and Hannon GJ: MicroRNAs: Small RNAs
with a big role in gene regulation. Nat Rev Genet. 5:522–531. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Tay Y, Zhang J, Thomson AM, Lim B and
Rigoutsos I: MicroRNAs to Nanog, Oct4 and Sox2 coding regions
modulate embryonic stem cell differentiation. Nature.
455:1124–1128. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Yang H, Tang Y, Guo W, Du Y, Wang Y, Li P,
Zang W, Yin X, Wang H, Chu H, et al: Up-regulation of microRNA-138
induce radiosensitization in lung cancer cells. Tumour Biol.
35:6557–6565. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Tang R, Liang L, Luo D, Feng Z, Huang Q,
He R, Gan T, Yang L and Chen G: Downregulation of MiR-30a is
associated with poor prognosis in lung cancer. Med Sci Monit.
21:2514–2520. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wu S, Shen W, Pan Y, Zhu M, Xie K, Geng L,
Wang Y, Liang Y, Xu J, Cao S, et al: Genetic variations in key
MicroRNAs are associated with the survival of nonsmall cell lung
cancer. Medicine (Baltimore). 94:e20842015. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Huang T, She K, Peng G, Wang W, Huang J,
Li J, Wang Z and He J: MicroRNA-186 suppresses cell proliferation
and metastasis through targeting MAP3K2 in non-small cell lung
cancer. Int J Oncol. 49:1437–1444. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Lu YJ, Liu RY, Hu K and Wang Y: MiR-541-3p
reverses cancer progression by directly targeting TGIF2 in
non-small cell lung cancer. Tumour Biol. 37:12685–12695. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Xu X, Wang X, Fu B, Meng L and Lang B:
Differentially expressed genes and microRNAs in bladder carcinoma
cell line 5637 and T24 detected by RNA sequencing. Int J Clin Exp
Pathol. 8:12678–12687. 2015.PubMed/NCBI
|
|
36
|
Li Q, Ge X, Xu X, Zhong Y and Qie Z:
Comparison of the gene expression profiles between gallstones and
gallbladder polyps. Int J Clin Exp Pathol. 7:8016–8023.
2014.PubMed/NCBI
|
|
37
|
Jiang CM, Wang XH, Shu J, Yang WX, Fu P,
Zhuang LL and Zhou GP: Analysis of differentially expressed genes
based on microarray data of glioma. Int J Clin Exp Med.
8:17321–17332. 2015.PubMed/NCBI
|
|
38
|
Chen L, Zhuo D, Chen J and Yuan H:
Screening feature genes of lung carcinoma with DNA microarray
analysis. Int J Clin Exp Med. 8:12161–12171. 2015.PubMed/NCBI
|
|
39
|
Wang X: Improving microRNA target
prediction by modeling with unambiguously identified
microRNA-target pairs from CLIP-ligation studies. Bioinformatics.
32:1316–1322. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Wong N and Wang X: miRDB: An online
resource for microRNA target prediction and functional annotations.
Nucleic Acids Res. 43(Database Issue): D146–D152. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chou CH, Chang NW, Shrestha S, Hsu SD, Lin
YL, Lee WH, Yang CD, Hong HC, Wei TY, Tu SJ, et al: miRTarBase
2016: Updates to the experimentally validated miRNA-target
interactions database. Nucleic Acids Res. 44:D239–D247. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Paraskevopoulou MD, Georgakilas G,
Kostoulas N, Vlachos IS, Vergoulis T, Reczko M, Filippidis C,
Dalamagas T and Hatzigeorgiou AG: DIANA-microT web server v5.0:
Service integration into miRNA functional analysis workflows.
Nucleic Acids Res. 41(Web Server Issue): W169–W173. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Reczko M, Maragkakis M, Alexiou P, Grosse
I and Hatzigeorgiou AG: Functional microRNA targets in protein
coding sequences. Bioinformatics. 28:771–776. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Ashburner M, Ball CA, Blake JA, Botstein
D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT,
et al: Gene ontology: Tool for the unification of biology. The Gene
Ontology Consortium. Nat Genet. 25:25–29. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Huang da W, Sherman BT and Lempicki RA:
Systematic and integrative analysis of large gene lists using DAVID
bioinformatics resources. Nat Protoc. 4:44–57. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Huang da W, Sherman BT and Lempicki RA:
Bioinformatics enrichment tools: Paths toward the comprehensive
functional analysis of large gene lists. Nucleic Acids Res.
37:1–13. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
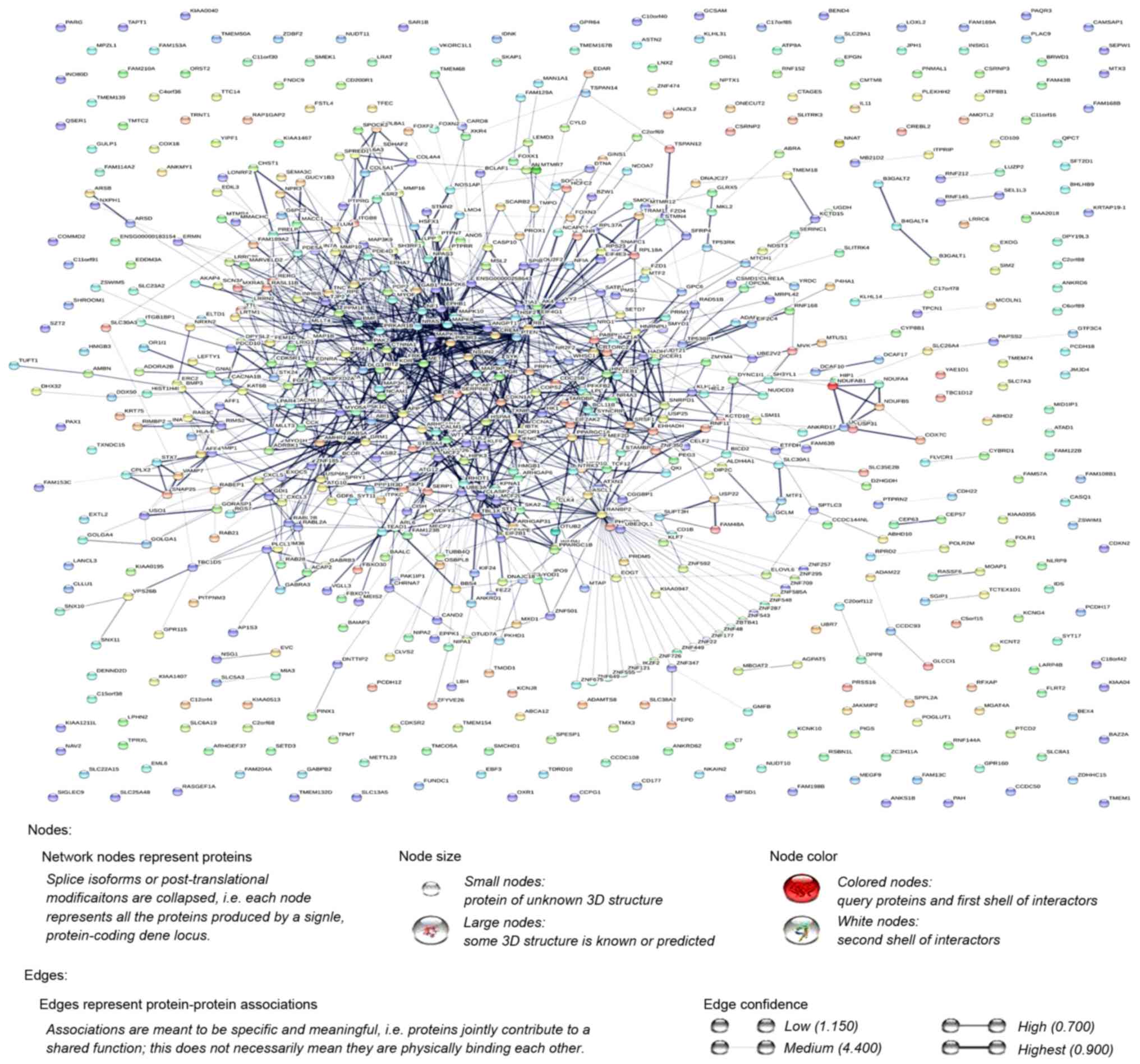
Szklarczyk D, Morris JH, Cook H, Kuhn M,
Wyder S, Simonovic M, Santos A, Doncheva NT, Roth A, Bork P, et al:
The STRING database in 2017: Quality-controlled protein-protein
association networks, made broadly accessible. Nucleic Acids Res.
45:D362–D368. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Shannon P, Markiel A, Ozier O, Baliga NS,
Wang JT, Ramage D, Amin N, Schwikowski B and Ideker T: Cytoscape: A
software environment for integrated models of biomolecular
interaction networks. Genome Res. 13:2498–2504. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9.1: Protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res.
41(Database Issue): D808–D815. 2013.PubMed/NCBI
|
|
51
|
Bornstein S, Schmidt M, Choonoo G, Levin
T, Gray J, Thomas CR Jr, Wong M and McWeeney S: IL-10 and integrin
signaling pathways are associated with head and neck cancer
progression. BMC Genomics. 17:382016. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Li B, Ruotti V, Stewart RM, Thomson JA and
Dewey CN: RNA-Seq gene expression estimation with read mapping
uncertainty. Bioinformatics. 26:493–500. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Wang K, Singh D, Zeng Z, Coleman SJ, Huang
Y, Savich GL, He X, Mieczkowski P, Grimm SA, Perou CM, et al:
MapSplice: Accurate mapping of RNA-seq reads for splice junction
discovery. Nucleic Acids Res. 38:e1782010. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Wu Y, Lyu H, Liu H, Shi X, Song Y and Liu
B: Downregulation of the long noncoding RNA GAS5-AS1 contributes to
tumor metastasis in non-small cell lung cancer. Sci Rep.
6:310932016. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Fu X, Li H, Liu C, Hu B, Li T and Wang Y:
Long noncoding RNA AK126698 inhibits proliferation and migration of
non-small cell lung cancer cells by targeting Frizzled-8 and
suppressing Wnt/β-catenin signaling pathway. Onco Targets Ther.
9:3815–3827. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Wang Z, Jin Y, Ren H, Ma X, Wang B and
Wang Y: Downregulation of the long non-coding RNA TUSC7 promotes
NSCLC cell proliferation and correlates with poor prognosis. Am J
Transl Res. 8:680–687. 2016.PubMed/NCBI
|
|
59
|
You J, Zhang Y, Liu B, Li Y, Fang N, Zu L,
Li X and Zhou Q: MicroRNA-449a inhibits cell growth in lung cancer
and regulates long noncoding RNA nuclear enriched abundant
transcript 1. Indian J Cancer. 51 Suppl 3:e77–e81. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Hamam R, Ali AM, Alsaleh KA, Kassem M,
Alfayez M, Aldahmash A and Alajez NM: microRNA expression profiling
on individual breast cancer patients identifies novel panel of
circulating microRNA for early detection. Sci Rep. 6:259972016.
View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Deng W, Yan M, Yu T, Ge H, Lin H, Li J,
Liu Y, Geng Q, Zhu M, Liu L, et al: Quantitative proteomic analysis
of the metastasis-inhibitory mechanism of miR-193a-3p in non-small
cell lung cancer. Cell Physiol Biochem. 35:1677–1688. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Zhang C, Ge S, Hu C, Yang N and Zhang J:
MiRNA-218, a new regulator of HMGB1, suppresses cell migration and
invasion in non-small cell lung cancer. Acta Biochim Biophys Sin
(Shanghai). 45:1055–1061. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Zhang R, Li Y, Wang Z, Chen L, Dong X and
Nie X: Interference with HMGB1 increases the sensitivity to
chemotherapy drugs by inhibiting HMGB1-mediated cell autophagy and
inducing cell apoptosis. Tumour Biol. 36:8585–8592. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Feng A, Tu Z and Yin B: The effect of
HMGB1 on the clinicopathological and prognostic features of
non-small cell lung cancer. Oncotarget. 7:20507–20519. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
65
|
Karachaliou N, Codony-Servat J, Teixidó C,
Pilotto S, Drozdowskyj A, Codony-Servat C, Giménez-Capitán A,
Molina-Vila MA, Bertrán-Alamillo J, Gervais R, et al: BIM and mTOR
expression levels predict outcome to erlotinib in EGFR-mutant
non-small-cell lung cancer. Sci Rep. 5:174992015. View Article : Google Scholar : PubMed/NCBI
|
|
66
|
Xu T, Wu S, Yuan Y, Yan G and Xiao D:
Knockdown of phosphodiesterase 4D inhibits nasopharyngeal carcinoma
proliferation via the epidermal growth factor receptor signaling
pathway. Oncol Lett. 8:2110–2116. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
67
|
Powers GL, Hammer KD, Domenech M,
Frantskevich K, Malinowski RL, Bushman W, Beebe DJ and Marker PC:
Phosphodiesterase 4D inhibitors limit prostate cancer growth
potential. Mol Cancer Res. 13:149–160. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
68
|
Ge X, Milenkovic L, Suyama K, Hartl T,
Purzner T, Winans A, Meyer T and Scott MP: Phosphodiesterase 4D
acts downstream of Neuropilin to control Hedgehog signal
transduction and the growth of medulloblastoma. Elife. 4:2015.
View Article : Google Scholar
|