Introduction
Tumorigenesis results from dysregulation of
oncogenes and tumor suppressors that influence cellular
proliferation, differentiation, senescence or apoptosis (1). Protein degradation is essential for
homeostasis and the survival of cells (2). Ubiquitination targets proteins for
proteasome-mediated degradation by the covalent attachment of one
or more ubiquitin molecules to a lysine residue (2–4). In
addition to regulating protein turnover, this post-translational
modification contributes to other cellular processes, including the
regulation of protein trafficking and subcellular distribution,
signal transduction, cell cycle, apoptosis and DNA repair (5–7).
Protein ubiquitination is performed by a trio of
enzymes, termed ubiquitin-activating enzyme, ubiquitin-conjugating
enzyme and ubiquitin protein ligase (E3). The E3 ligases determine
the substrate specificity of the ubiquitination reactions (3,6,8). E3 ubiquitin ligases are a large family
of proteins that are classified into three major structurally
distinct types: N-end rule E3s; E3s containing the homology to E6AP
C-terminus (HECT) domain; and E3s with a Really Interesting New
Gene finger domain, including its derivatives, the Plant
Homeo-Domain and the U-Box (9). HECT
E3s were first reported in 1995, as the first E3s described
(2). HECT E3s serve important roles
in sporadic and hereditary human diseases, including cancer,
cardiovascular (Liddle's syndrome) and neurological (Angelman
syndrome) disorders, and in disease-relevant processes, including
bone homeostasis, immune responses and retroviral budding (10).
E3 ubiquitin-protein ligase (HUWE1) is a
recently-identified 500 kDa HECT domain-containing E3 ligase, that
serves a critical role in proteasomal degradation of several
proteins, including p53, c-Myc, myeloid cell leukemia sequence 1
(Mcl-1), cell division cycle 6 (Cdc6), histones, N-Myc,
Msx-interacting-zinc finger 1(Miz1), DNA topoisomerase 2-binding
protein 1, DNA polymerase β, mitofusin 2, histone deacetylase 2 and
Ras association domain family member 1 (1,11–21).
However, whether any of these existing or other unidentified
substrates mediate the major function of HUWE1 in cell
proliferation remains unclear. One study demonstrated that
silencing HUWE1 increased apoptosis by ubiquitination and
degradation of Mcl-1, while another study reported that silencing
HUWE1 increased survival by ubiquitination and degradation of p53
in the same cell line (1,6,14).
Experiments performed in a HDM2-null genetic background confirmed
that ARF-induced stabilization of p53 involved HUWE (22). However, other data could not
demonstrate the inhibitory activity of ARF toward HUWE1 (15). Furthermore, HUWE1 was incapable of
regulating p53 abundance in response to DNA-damage stress, while
other substrates, including Mcl-1 and Cdc6, were ubiquitylated and
degraded (16). In addition, the
steady-state protein levels of p53 were not increased by depletion
of HUWE1 in neuroblastoma cells (6,21).
Another study demonstrated that HUWE1 assembled Lysine (Lys)
63-linked polyubiquitin chains on c-Myc and this modification was
revealed to be required for gene activation by c-Myc (15). However, other data revealed that
HUWE1 ubiquitylates N-Myc via Lys48-mediated linkages and targets
it for destruction by the proteasome (21), and post-translational modifications
of c-Myc do not appear to be required for its interaction with p300
(23,24).
Previous studies have indicated that HUWE1 may
either serve an oncogenic or anti-oncogenic function depending on
the type of cancer (14,25,26). In
addition, it has been demonstrated that HUWE1 serves important
roles in proteasomal degradation of several proteins in various
types of cancer (1,11–21).
However, the precise role of HUWE1 expression remains controversial
due to conflicting evidence. Notably, the HUWE1 gene exhibits an
important role in various types of tumor, including lung, breast
and colorectal carcinomas (1,8,14,15).
Compared with normal tissue the expression levels of HUWE1 are
higher in numerous types of primary human tumors, predominantly
solid tumor, including lung, breast, colon and prostate carcinoma,
as well as glioblastoma (14). By
contrast, HUWE1 is undetectable or expressed at very low levels in
normal epithelium, in benign polyps, and pancreatic cancer
(14,27). These results suggest that HUWE1 may
serve an important role depending on the type of cancer.
In order to analyze the expression of HUWE1 in
various types of cancer and to investigate the molecular mechanism
of HUWE1 in the regulation of tumor development, the present study
evaluated HUWE1 expression using the Oncomine database. Gene
alterations during carcinogenesis, copy number alterations and
mutations of HUWE1 were also examined using cBioPortal, which is
the International Cancer Genome Consortium and the Tumorscape
database.
Materials and methods
Oncomine database analysis
Expression levels of HUWE1 in cancer vs. normal
tissues were analyzed using the ‘cancer vs. normal’ filter in the
Oncomine database (https://www.oncomine.org/resource/login.html)
(28,29). All data conforming to the criteria of
P<0.01, fold-change >2 and a gene rank percentile <10%,
were included in the present study (20–33). The
advanced analysis criteria were adjusted as follows: P<0.0001,
fold-change >2 and gene ranking in the top 10%. A heat map was
used to present the expression profile of HUWE1 in various cancer
types.
Kaplan-Meier analysis
The Kaplan-Meier plotter database (http://kmplot.com/analysis/) (34) was used to assess the effect of 54,675
genes on survival using 10,461 cancer samples. This included 5,143
breast, 1,816 ovarian, 2,437 lung and 1,065 gastric cancer cases
with a mean follow-up of 69, 40, 49 and 33 months, respectively.
The primary purpose of the tool is for meta-analysis-based
biomarker assessment. The association between the expression levels
of HUWE1 and survival rates in breast, gastric, ovarian and lung
cancer were analyzed using the Kaplan-Meier plotter. The hazard
ratio with a 95% confidence interval and log rank P-value were
calculated.
Prognoscan database analysis
The PrognoScan database (http://dna00.bio.kyutech.ac.jp/PrognoScan/) (35) was searched to determine the
association between the expression levels of HUWE1 and survival
rates in various cancer types. The threshold was adjusted to Cox
P<0.05.
Identifying the proteins that interact
with HUWE1
The search tool for the retrieval of interacting
genes/proteins (STRING v.10) analysis tool (https://string-db.org/) was used to identify
interacting proteins, with ‘HUWE1 (Homo sapiens)’ used as
the query. Several previously identified partners have been
genetically verified and therefore served as the foundation for
revealing other protein partners in the network. If any identified
proteins were not specific to the HUWE1 network, they were excluded
from the gene signature (36).
cBioPortal database analysis
The cBioPortal (http://cbioportal.org) was used to investigate
mutations and copy number alterations (CNAs) of the HUWE1 gene and
the predicted protein partners in various cancer types. The
cBioPortal is a website used for investigating, visualizing and
analyzing multidimensional cancer genomics data (37,38). The
threshold criteria of studies were dataset ≥100 samples and samples
with >20% alteration frequency (37).
Statistical analysis
The Prognoscan and Kaplan-Meier plots were used to
generate survival curves. All results were reviewed with a P-value
from a log-rank test, and with Oncomine and heat maps. Oncomine
reported the statistical significance with a P-value.
Results
HUWE1 transcript expression by cancer
type
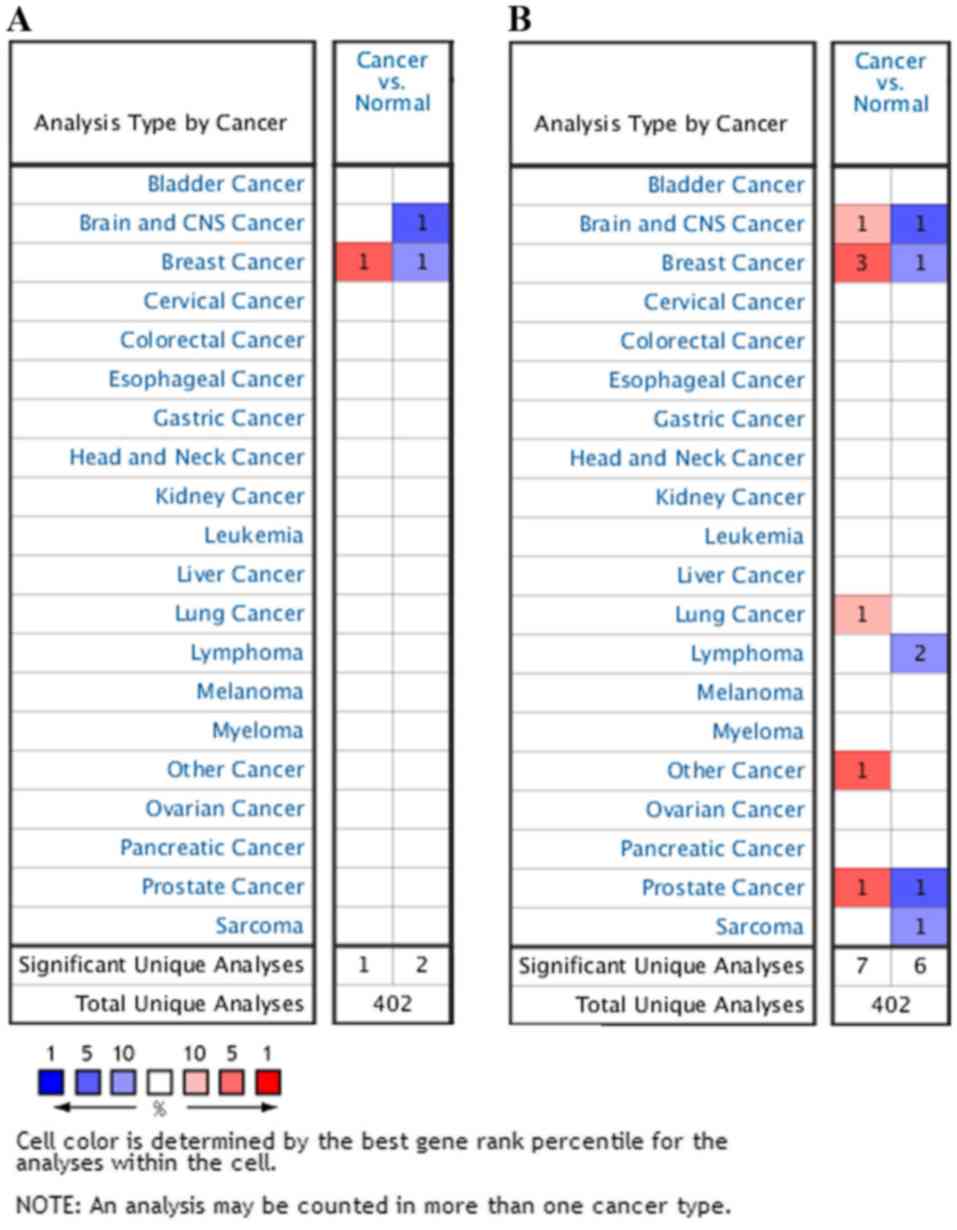
To investigate the function of HUWE1 in different
cancer types, the Oncomine database was used to compare HUWE1
expression levels between tumor and normal tissues. The HUWE1 mRNA
expression levels in the tissue of origin were selected and
compared using the ‘cancer vs. normal’ filter. For inclusion and
further evaluation, data matching the following criteria were
selected: P<0.01 and fold-change >2, or P<0.0001 and
fold-change >2. Statistical analyses, including P-values,
two-tailed Student's t-test and multiple testing corrections, were
performed using the Oncomine default algorithms. Compared with
normal tissue, HUWE1 was overexpressed in various tumor types,
while it exhibited lower expression levels in other tumor types.
These results indicated that HUWE1 may serve either an oncogenic or
tumor suppressor function depending on the cancer type (Fig. 1). Accordingly, further detailed
analyses of HUWE1 were performed.
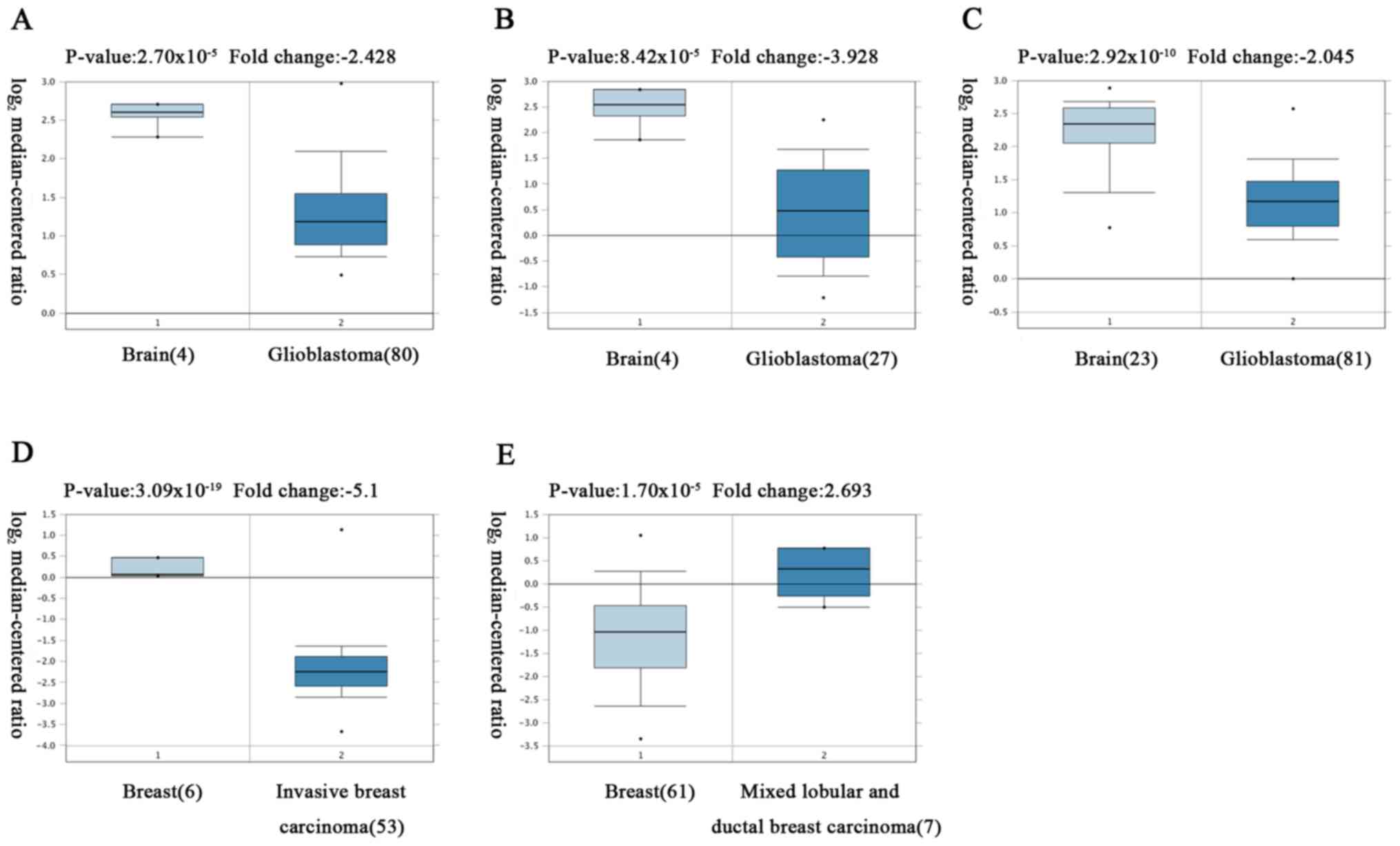
The analysis revealed that HUWE1 was
overexpressed in leukemia and lung cancer, but was under expressed
in glioblastoma, lymphoma, sarcoma and testicular seminoma tissues
compared with normal tissues
However, the expression of HUWE1 in breast and
prostate cancers remains controversial (Fig. 2; Table
I).
 | Table I.Association of HUWE1 expression with
survival of patients with cancer. |
Table I.
Association of HUWE1 expression with
survival of patients with cancer.
| Cancer type | n | P-value | HR (95% CI) | Endpoint | Dataset | Probe ID | ln (HR) |
|---|
| Bladder cancer | 30 |
3.05×10−2 | 3.42
(1.12–10.42) | Overall
survival | GSE5287 | 208598_s_at | 1.2296 |
| Brain cancer | 74 |
3.16×10−2 | 0.23
(0.06–0.88) | Overall
survival | GSE4412-GPL96 | 208598_s_at | −1.4783 |
| Breast cancer | 158 |
2.07×10−2 | 3.69
(1.22–11.15) | Overall
survival | GSE3143 | 34372_at | 1.3054 |
| Breast cancer | 204 |
2.68×10−2 | 0.47
(0.24–0.92) | Relapse free
survival | GSE12276 | 207783_x_at | −0.7499 |
| Breast cancer | 77 |
2.47×10−2 | 0.01
(0.00–0.57) | Relapse free
survival | GSE9195 | 207783_x_at | −4.4615 |
| Breast cancer | 77 |
2.07×10−2 | 0.01
(0.00–0.45) | Distant metastasis
free survival | GSE9195 | 207783_x_at | −5.2755 |
| Breast cancer | 136 |
3.91×10−3 | 0.26
(0.10–0.65) | Distant metastasis
free survival | GSE12093 | 208598_s_at | −1.3632 |
| Breast cancer | 200 |
1.84×10−2 | 1.54
(1.08–2.21) | Distant Metastasis
Free survival | GSE11121 | 208599_at | 0.4321 |
| Breast cancer | 117 |
3.52×10−2 | 7.17
(1.15–44.82) | Distant metastasis
free survival | E-TABM-158 | 208599_at | 1.9697 |
| Breast cancer | 236 |
1.38×10−2 | 1.57
(1.10–2.24) | Disease specific
survival | GSE3494-GPL96 | 214673_s_at | 0.44818 |
| Breast cancer | 236 |
1.25×10−2 | 5.04
(1.42–17.91) | Disease specific
survival | GSE3494-GPL96 | 208598_s_at | 1.6168 |
| Breast cancer | 249 |
4.97×10−3 | 4.08
(1.53–10.89) | Disease Free
Survival | GSE4922-GPL96 | 208598_s_at | 1.4066 |
| Breast cancer | 198 |
7.58×10−3 | 0.58
(0.39–0.87) | Distant metastasis
free survival | GSE7390 | 207783_x_at | −0.5376 |
| Breast cancer | 198 |
5.87×10−3 | 0.57
(0.39–0.85) | Overall
survival | GSE7390 | 207783_x_at | −0.5539 |
| Breast cancer | 198 |
2.54×10−2 | 0.63
(0.43–0.95) | Relapse free
survival | GSE7390 | 207783_x_at | −0.4546 |
| Colorectal
cancer | 177 |
3.56×10−2 | 0.22
(0.05–0.90) | Overall
survival | GSE17536 | 214673_s_at | −1.5180 |
| Colorectal
cancer | 55 |
3.20×10−4 | 0.01
(0.00–0.11) | Overall
survival | GSE17537 | 214673_s_at | −4.8093 |
| Colorectal
cancer | 55 |
2.01×10−4 | 0.00
(0.00–0.06) | Disease free
survival | GSE17537 | 214673_s_at | −6.1017 |
| Colorectal
cancer | 55 |
2.31×10−2 | 0.02
(0.00–0.58) | Overall
survival | GSE17537 | 208599_at | −3.9563 |
| Colorectal
cancer | 49 |
7.78×10−3 | 0.01
(0.00–0.32) | Disease specific
survival | GSE17537 | 214673_s_at | −4.2781 |
| Esophagus
cancer | 34 |
4.49×10−2 | 14.93
(1.06–209.58) | Overall
survival | GSE11595 | 66423 | 2.7033 |
| Lung cancer | 104 |
1.09×10−2 | 0.01
(0.00–0.32) | Overall
survival |
jacob-00182-MSK | 207783_x_at | −4.9303 |
| Lung cancer | 204 |
4.40×10−3 | 18.26
(2.47–134.83) | Overall
survival | GSE31210 | 207783_x_at | 2.9050 |
| Lung cancer | 204 |
1.27×10−5 | 28.08
(6.28–25.48) | Relapse free
survival | GSE31210 | 207783_x_at | 3.3349 |
| Lung cancer | 204 |
1.44×10−2 | 0.38
(0.17–0.82) | Overall
survival | GSE31210 | 236294_at | −0.9750 |
| Lung cancer | 204 |
3.28×10−6 | 0.26
(0.14–0.45) | Relapse free
survival | GSE31210 | 236294_at | −1.3652 |
| Lung cancer | 138 |
6.40×10−3 | 0.00
(0.00–0.22) | Relapse free
survival | GSE8894 | 236294_at | −5.4148 |
| Lung cancer | 138 |
2.50×10−2 | 0.15
(0.03–0.79) | Relapse free
survival | GSE8894 | 208599_at | −1.8913 |
| Ovarian cancer | 133 |
4.32×10−2 | 1.95
(1.02–3.72) | Overall
survival | DUKE-OC | 208599_at | 0.6675 |
| Prostate
cancer | 281 |
5.88×10−3 | 1.80
(1.19–2.75) | Overall
survival | GSE16560 | DAP4_0152 | 0.5900 |
| Skin cancer | 38 |
2.33×10−3 | 11.36
(2.38–54.33) | Overall
survival | GSE19234 | 208599_at | 2.4304 |
| Soft tissue
cancer | 140 |
6.89×10−4 | 3.20
(1.64–6.27) | Distant Recurrence
Free survival | GSE30929 | 208598_s_at | 1.1640 |
Genetic expression levels of HUWE1 and
patient survival
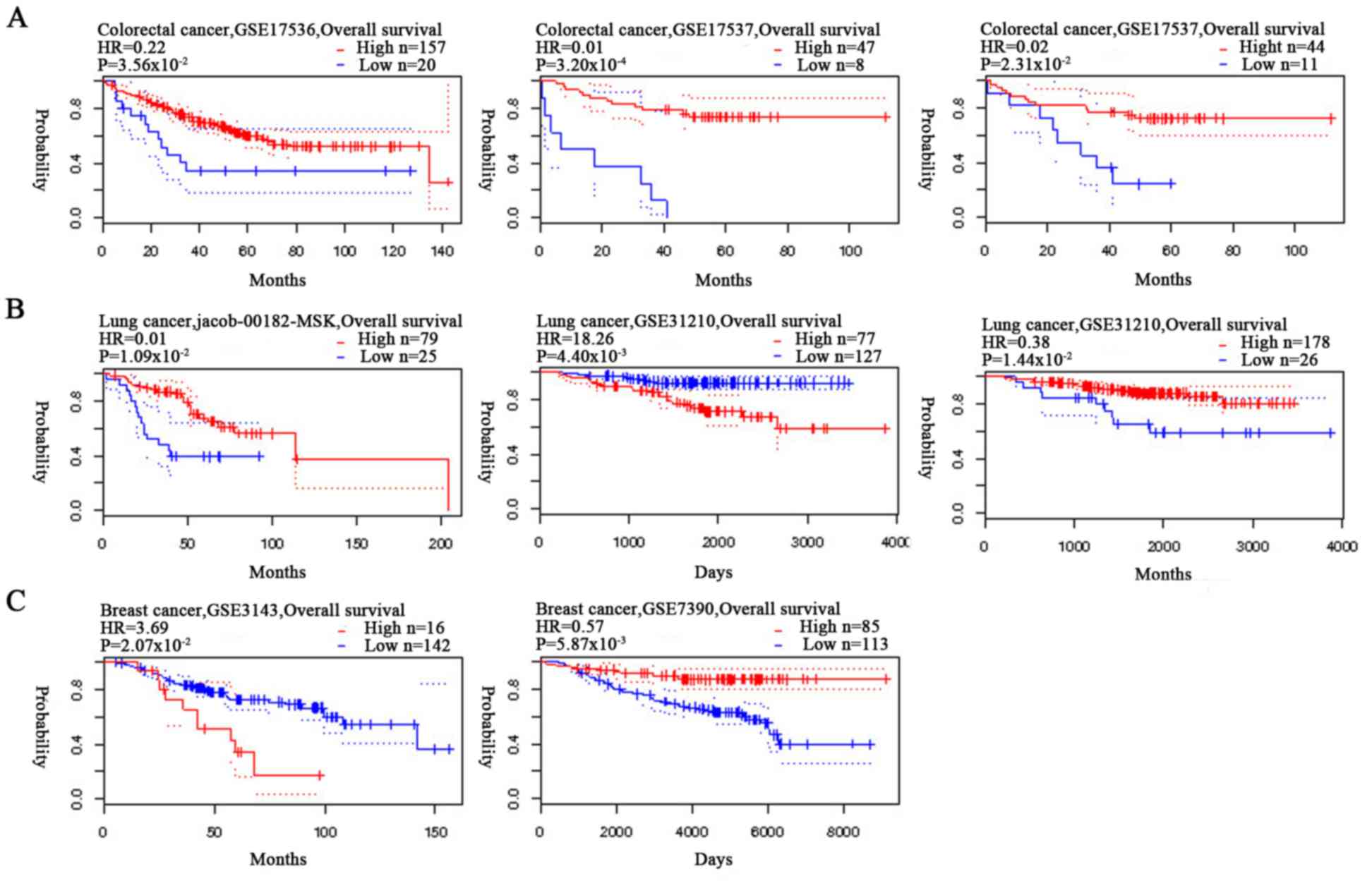
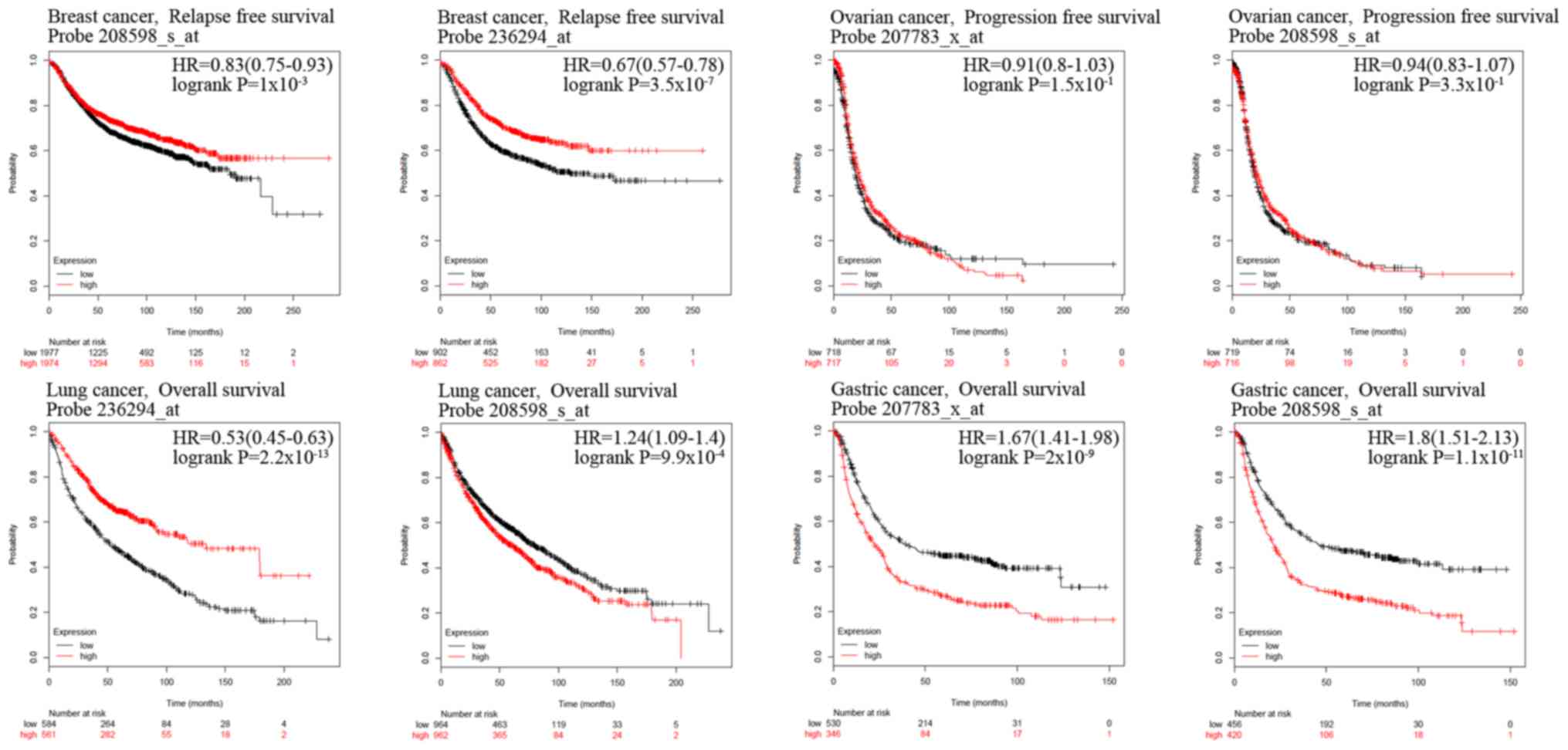
The association between the expression levels of
HUWE1 and OS rate of patients with colorectal cancer was analyzed
using Kaplan-Meier analysis. Patients with colorectal cancer with a
high expression level of HUWE1 exhibited a significantly higher
survival rate (Fig. 3). Analysis of
gastric cancer revealed an association between a high expression
level of HUWE1 and low OS rate. HUWE1 expression and PFS rate of
patients with ovarian cancer were not statistically significant
(P=1.50×10−1 and P=3.30×10−1; Fig. 4). The significance of the association
of HUWE1 expression and survival rates in lung and breast cancer
remains inconclusive (Figs. 3B and
4).
The expression of HUWE1 was evaluated using the
cBioPortal database. The data demonstrated lower expression levels
of HUWE1 in brain, lymphoma and round cell liposarcoma; however,
its roles in lung and breast cancer were uncertain (Table II). Despite this controversy,
certain data agreed with previously published reports. Previous
studies have demonstrated that HUWE1 is highly expressed in a
significant proportion of lung and breast carcinomas (14,15), and
lowly expressed in colorectal carcinomas; HUWE1 expression was
positively correlated with tumor stage and negatively correlated
with p53 protein expression (26).
 | Table II.HUWE1 expression in cancer types. |
Table II.
HUWE1 expression in cancer types.
| Cancer type | P-value | Fold-change | Sample size, n | (Refs.) |
|---|
| Brain |
|
|
|
|
|
Glioblastoma |
2.70×10−5 | −2.428 | 84 | (30) |
|
Glioblastoma |
8.42×10−5 | −3.928 | 54 | (31) |
|
Anaplastic
oligoastrocytoma |
3.00×10−3 | −2.916 | 54 | (31) |
|
Anaplastic
oligodendroglioma |
2.80×10−2 | −2.667 | 54 | (31) |
|
Glioblastoma |
3.20×10−2 | −3.046 | 38 | (46) |
|
Oligoastrocytoma |
4.00×10−2 | −2.045 | 38 | (46) |
|
Glioblastoma |
2.92×10−10 | −2.045 | 180 | (32) |
| Breast |
|
|
|
|
| Mixed
lobular and ductal breast carcinoma |
1.70×10−5 | 2.693 | 68 | (60) |
|
Invasive breast carcinoma |
3.09×10−19 | −5.100 | 59 | (33) |
| Leukemia |
|
|
|
|
| Chronic
lymphocytic leukemia |
2.00×10−3 | 2.562 | 111 | (47) |
| Lung |
|
|
|
|
|
Squamous cell lung
carcinoma |
9.00×10−3 | 3.019 | 203 | (48) |
| Small
cell lung carcinoma |
2.80×10−2 | 2.137 | 73 | (59) |
| Lymphoma |
|
|
|
|
| Diffuse
large B-cell lymphoma |
1.00×10−2 | −3.503 | 27 | (50) |
|
Marginal zone B-cell
lymphoma |
8.00×10−3 | −3.615 | 27 | (50) |
| Other |
|
|
|
|
|
Testicular seminoma |
2.00×10−3 | 11.159 | 30 | (51) |
| Prostate |
|
|
|
|
|
Prostate carcinoma |
5.00×10−3 | 6.016 | 30 | (52) |
|
Prostate carcinoma |
3.40×10−2 | −2.207 | 19 | (53) |
| Sarcoma |
|
|
|
|
| Round cell
liposarcoma |
6.00×10−3 | −3.047 | 54 | (54) |
Proteins associated with HUWE1
Functional proteins understood to be associated with
HUWE1 were selected by STRING analysis tool for the retrieval of
interacting genes/proteins. The ten predicted protein partners of
HUWE1 were as follows (Fig. 5):
non-POU domain-containing octamer-binding protein, transmembrane
protein 164, protein Jade-3 (PHF16), TGF-β activated kinase 1
(TAB3), TPP synthase 2 (CTPS2), solute carrier family 9 member A7
(SLC9A7), thyroid hormone receptor interactor 12, zinc finger
protein 280C, ubiquitin-like modifier-activating enzyme 1 (UBA1)
and RNA binding motif protein 10 (RBM10). The above ten predicted
proteins were analyzed by the cBioPortal database, each protein
according to the threshold criteria of studies, and datasets ≥100
samples and samples with >20% alteration frequency were selected
for further analysis: RBM10, UBA1, PHF16, TAB3, CTPS2 and
SLC9A7.
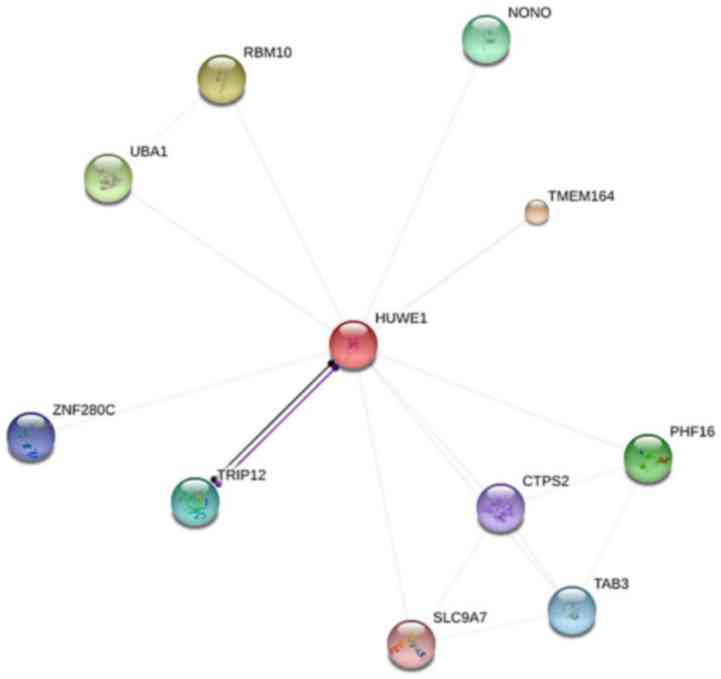 | Figure 5.Identification of proteins associated
with HUWE1 using STRING. Interacting nodes are presented in colored
circles. HUWE1, E3 ubiquitin-protein ligase; RBM10, RNA binding
motif protein 10; UBA1, ubiquitin-like modifier-activating enzyme
1; ZNF280C, zinc finger protein 280C; TRIP12, thyroid hormone
receptor interactor 12; SLC9A7, solute carrier family 9 member A7;
NONO, non-POU domain-containing octamer-binding protein; TMEM164,
transmembrane protein 164; PHF16, protein Jade-3; CTPS2, TPP
synthase 2; TAB3, TGF-β activated kinase 1. |
Alterations of HUWE1 in different
cancer types
To investigate mutations and CNAs of the HUWE1 gene
and the six predicted protein partners in various cancer types, the
cBioPortal tool was used to analyze 91 cancer studies. In total, 11
of these studies included ≥100 samples in the dataset and samples
with >20% alteration frequency. The alteration frequency ranged
between 20.00 and 42.10% (Table
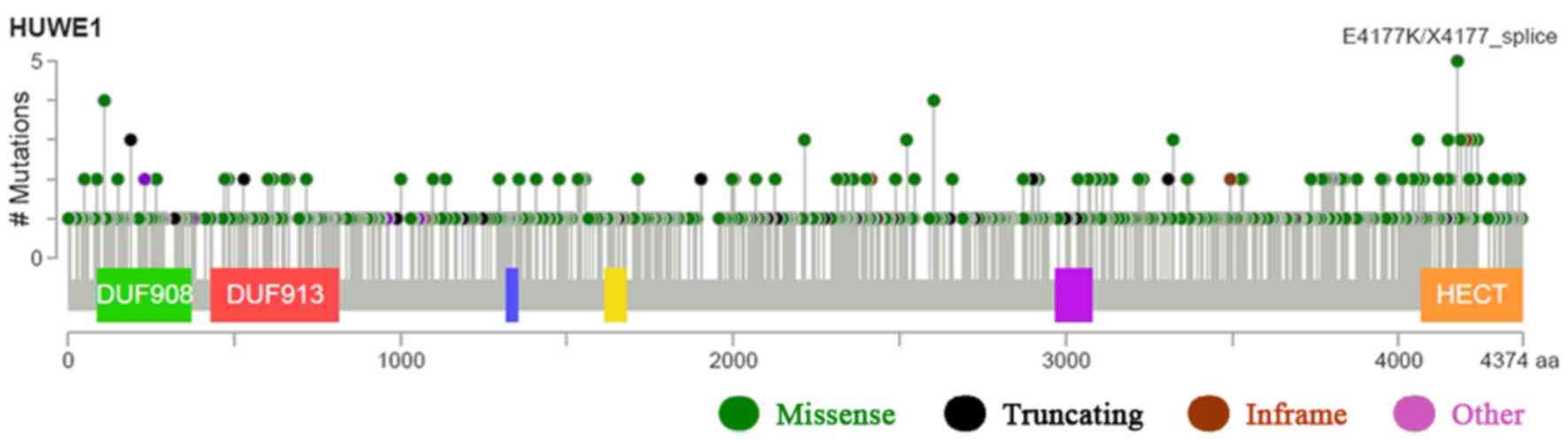
III)(39–45). HUWE1 mutations occurred in many
domains of the protein, predominantly in the C-terminus. Following
database analysis, it was indicated that the most critical site
mutation E4177K/X4177 occurs at the C-terminus. The higher the
value of the vertical axis, the higher the mutation rate.
Therefore, it is suggested that since C-terminus had the highest
mutation rate, it may have the greatest impact on the function of
HUWE1 (Fig. 6) and the highest
mutation frequencies were revealed in neuroendocrine prostate
cancer (Fig. 7).
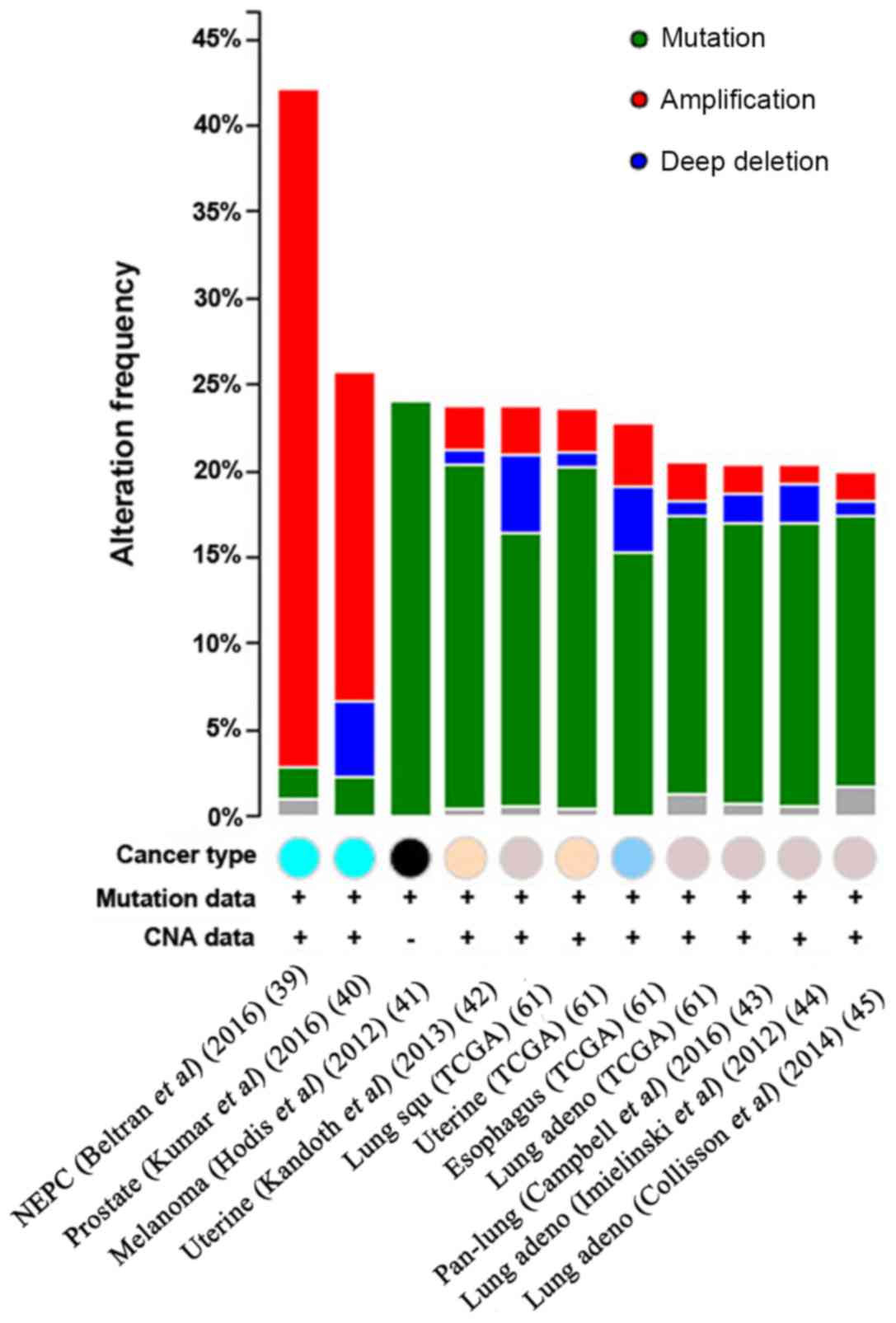 | Figure 7.CNAs of the HUWE1 gene in different
cancer subtypes. The alteration frequency of HUWE1 was determined
using the cBioPortal database. Cancer types containing >100
samples were selected and only alteration frequencies >20% are
presented. The different colored circles represent different types
of tumors. The alterations included amplifications (red), deletions
(blue), multiple alterations (grey) or mutations (green). HUWE1, E3
ubiquitin-protein ligase; NEPC, neuroendocrine prostate cancer;
Prostate, prostate cancer; Melanoma, Skin cutaneous melanoma
cancer; Uterine, Uterine corpus endometrial carcinoma cancer; Lung
squ, Lung squamous cell carcinoma cancer; Esophagus, Esophageal
carcinoma cancer; Lung adeno, Lung adenocarcinoma cancer; Pan-lung,
Pan-lung cancer; TCGA, The Cancer Genome Atlas; CNA, copy number
alteration. |
 | Table III.Alteration frequency of a six-gene
signature (RBM10, UBA1, JADE3, TAB3, CTPS2 and SLC9A7) in different
cancer types. |
Table III.
Alteration frequency of a six-gene
signature (RBM10, UBA1, JADE3, TAB3, CTPS2 and SLC9A7) in different
cancer types.
| Cancer type | Data source | n | Frequency, (%) | Amplification,
(%) | Deletion, (%) | Mutation, (%) | Multiple
alterations, (%) | (Refs.) |
|---|
| Neuroendocrine
prostate cancer | Beltran et
al (2016) | 107 | 42.10 | 39.30 |
|
| 0.90 | (39) |
| Prostate
adenocarcinoma | Kumar et al
(2016) | 136 | 25.70 | 19.10 | 4.40 |
2.20 |
| (40) |
| Skin cutaneous
melanoma | Hodis et al
(2012) | 121 | 24.00 |
|
| 24.00 |
| (41) |
| Uterine corpus
endometrial carcinoma | Kandoth et
al (2013) | 240 | 23.80 |
2.50 | 0.80 | 20.00 | 0.40 | (42) |
| Lung squamous cell
carcinoma | TCGA,
Provisional | 177 | 23.70 |
2.80 | 4.50 | 15.80 | 0.60 |
|
| Uterine Corpus
Endometrial carcinoma | TCGA,
Provisional | 242 | 23.60 |
2.50 | 0.80 | 19.80 | 0.40 |
|
| Esophageal
carcinoma | TCGA,
Provisional | 184 | 22.80 |
3.80 | 3.80 | 15.20 |
|
|
| Lung
adenocarcinoma | TCGA,
Provisional | 230 | 20.40 |
2.20 | 0.90 | 16.10 | 1.30 |
|
| Pan-lung
cancer | Campbell et
al (2016) | 1144 | 20.40 |
1.70 | 1.70 | 16.30 | 0.70 | (43) |
| Lung
adenocarcinoma | Imielinski et
al (2012) | 182 | 20.30 |
1.10 | 2.20 | 16.50 | 0.50 | (44) |
| Lung
Adenocarcinoma | Collisson et
al (2014) | 230 | 20.00 |
1.70 | 0.90 | 15.70 | 1.70 | (45) |
Subsequently, OncoPrint was used to query for
alterations in the RBM10, UBA1, JADE3, TAB3, CTPS2 and SLC9A7
genes. The proportion of alterations in these genes among prostate
cancer varied between 1.8 and 2.5% for individual genes (RBM10, 2%;
UBA1, 2.5%; JADE, 3.2%; TAB3, 1.7% CTPS2, 1.9%; and SLC9A7, 1.8%;
Fig. 8). The UBA1 gene demonstrated
a high level of amplification in prostate cancer.
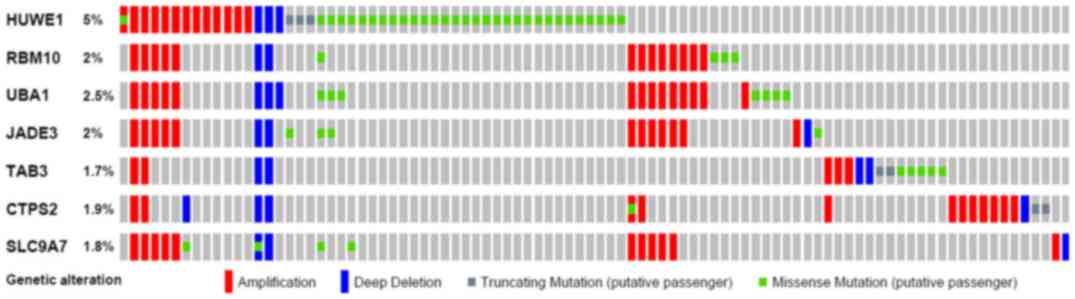 | Figure 8.Mutation of HUWE1 and associated
genes in prostate cancer. The Oncoprint feature of cBioPortal was
used to determine the copy number alteration frequency of each
individual gene. The percentages of alterations in HUWE1, RBM10,
UBA1, JADE3, TAB3, CTPS2 and SLC9A7 genes are presented for
prostate cancer. HUWE1, E3 ubiquitin-protein ligase; RBM10, RNA
binding motif protein 10; UBA1, ubiquitin-like modifier-activating
enzyme 1; SLC9A7, solute carrier family 9 member A7; CTPS2, TPP
synthase 2; TAB3, TGF-β activated kinase 1. |
A co-expression gene profile for HUWE1 in breast
cancer was generated using Oncomine (Fig. 9). HUWE1 was revealed to be
co-expressed with roundabout guidance receptor 1 (ROBO1),
ectodysplasin A (EDA), spalt like transcription factor 1 (SALL1),
p21 (RAC1) activated kinase 2 (PAK2) and glutamyl aminopeptidase
(ENPEP).
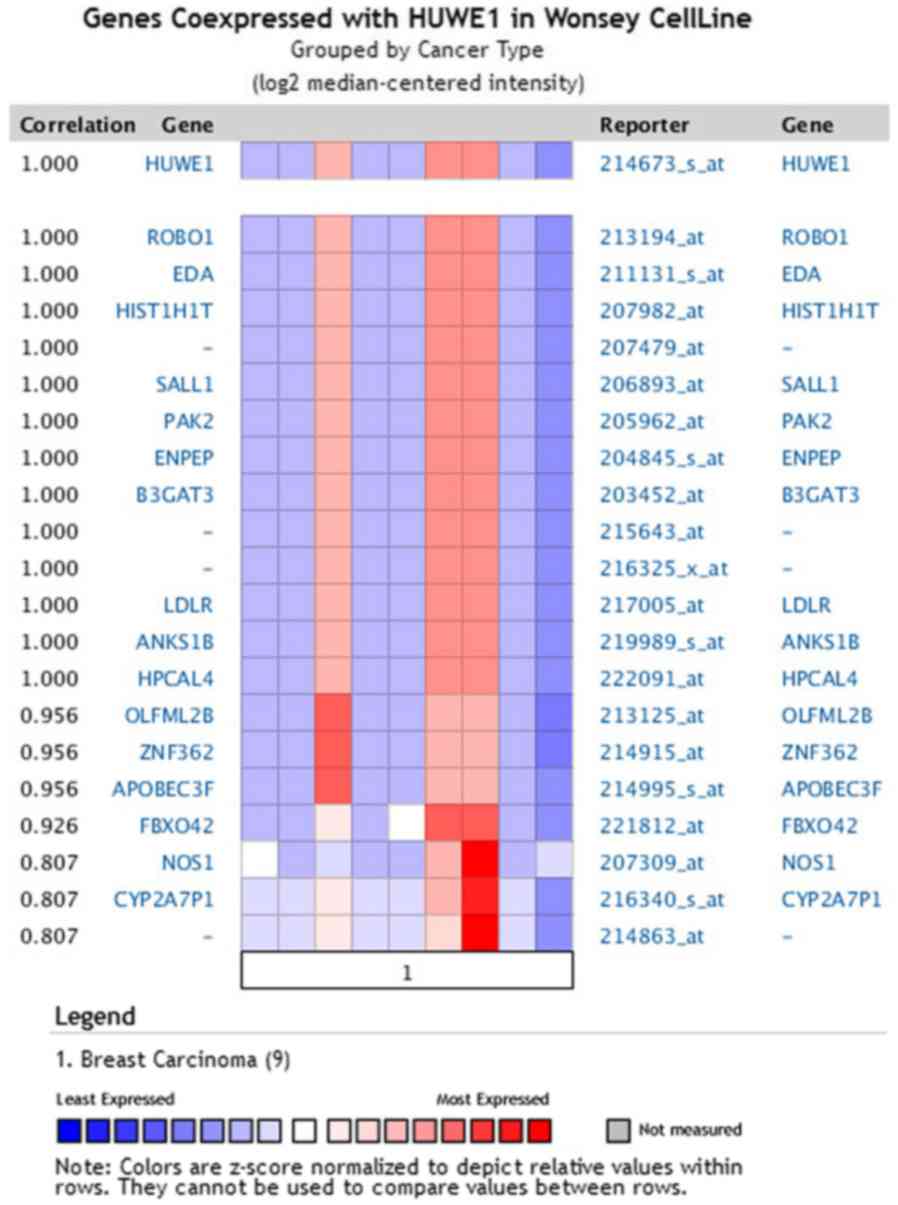 | Figure 9.HUWE1-associated genes in breast
carcinoma. HUWE1 is co-expressed in breast carcinoma tissues with
the indicated genes. HUWE1, E3 ubiquitin-protein ligase; ROBO1,
roundabout guidance receptor 1; EDA, ectodysplasin A; SALL1, spalt
like transcription factor 1; PAK2, p21 (RAC1) activated kinase 2;
ENPEP, glutamyl aminopeptidase; HIST1H1T, histone cluster 1 H1
family member T; B3GAT3, β-1,3-glucuronyltransferase 3; LDLR,
low-density lipoprotein receptor; ANKS1B, ankyrin repeat and
sterile alpha motif domain-containing protein 1B; HPCAL4,
hippocalcin like 4; OLFML2B, olfactomedin like 2B; ZNF362, zinc
finger protein 362; APOBEC3F, apolipoprotein B mRNA editing enzyme
catalytic subunit 3F; FBXO42, F-box protein 42; NOS1, nitric oxide
synthase 1; CYP2A7P1, cytochrome p450 family 2 subfamily member 7
pseudogene 1. |
Discussion
HUWE1 is a HECT E3 ligase that serves a critical
role in proteasomal degradation of several proteins and
participates in cell cycle control, DNA damage response and
tumorigenesis (1,11–22).
However, to the best of our knowledge, whether any of these
existing or unidentified substrates, including p53, MCL-1 and cdc6
ect, mediate the predominant function of ARF-BP1, also named MULE,
HUWE1, in cell proliferation remains unclear. HUWE1 was first
identified in 2005 (14); however,
it remains unclear whether it can serve as a biomarker for cancer
diagnosis or prognosis. To investigate this, the present study
selected data according to the expression of several genes with
clearly defined parameters between cancer and normal tissues. In
the Oncomine analysis, HUWE1 was revealed to be overexpressed in
leukemia and lung cancer, but was downregulated in brain cancer,
lymphoma, sarcoma and testicular seminoma, compared with normal
tissue. To further investigate the survival rates of patients with
different expression levels of HUWE1, the associations between the
expression levels of HUWE1 and the survival rates of patients were
analyzed using Kaplan-Meier analysis and PrognoScan. In general,
high expression levels of HUWE1 were associated with lower survival
rates for patients with ovarian cancer (30–33,46–55);
however, the results for lung cancer were not clear.
The main four causes of cancer are
somatically-acquired genetic, epigenetic, transcriptomic and
proteomic alterations in cells. These alterations occur in specific
genomic regions, which can result in tumor suppressive or oncogenic
effects (56). The cBioPortal
analysis identified cancer types with significant CNAs in the
selected HUWE1-gene signature. In total, 11 cancer studies
representing 681 samples were analyzed, which contained >20%
alteration frequency and ≥100 samples in the dataset. The
alteration frequency ranged between 20.00 and 42.10%, with the
dominance hierarchy.
The cBioPortal was used for interactive analysis and
visualization of the HUWE1-associated network. The network was
generated based on pathways and interactions from the Human Protein
Reference database, Reactome Pathway database, National Cancer
Institute Pathway Interaction database and the MSKCC Cancer Cell
Map (56,57). The generated network improves
understanding of the molecular mechanisms of HUWE1 in cancer.
The Oncomine database presents a potentially
significant list of co-expressed genes that are critical in
defining pathways. HUWE1 was identified to be co-expressed with
ROBO1 in breast carcinoma, as well as with EDA, SALL1, PAK2 and
ENPEP. This analysis may assist future studies regarding the
function of HUWE1.
However, the precise role of HUWE1 remains
controversial due to conflicting evidence. To investigate the role
of HUWE1 in various cancer types, the Oncomine platform, which
includes data from ~90,000 microarray experiments, was used to
assess gene expression in different cancer types (28,29). In
addition, the survival of patients with cancer was assessed using
Kaplan-Meier plots and the PrognoScan database (34,38).
Co-expressed data show that HUWE1 may regulate proteins and the
signaling pathways involved in these proteins, which may assist in
the examination of the molecular mechanism by which HUWE1 regulates
cellular biological functions. The function and regulatory
mechanisms of genes can be identified by STRING (36).
An aim of the present study was to determine whether
CNAs of the HUWE1 network were associated with aggressive cancer
subtypes, based on both the cBioPortal and Tumorscape (37,38,58). The
role of HUWE1 in different cancer types was first analyzed. It was
revealed that HUWE1 may serve an oncogenic role in breast, brain,
central nervous system and prostate cancer types, and may serve a
tumor suppressive role in colorectal cancer and certain types of
lung cancer. In addition, the associations between HUWE1 expression
levels and patient survival rates were investigated. According to
STRING analysis, co-expression analysis demonstrated that HUWE1 was
co-expressed with ROBO1 in breast carcinoma, as well as with EDA,
HIST1S1T, SALL1, PAK2, ENPEP, B3GAT3 and ROBO1. The relationship
between the expression of HUWE1 and prognosis remains unclear. By
contrast, a high HUWE1 expression level was associated with a poor
prognosis for gastric and ovarian cancer. The cBioPortal and
Tumorscape analyses identified a HUWE1 alteration frequency of
20–42%, with the highest frequency observed in prostate cancer. In
summary, HUWE1-coexpressed proteins may be used to further
elucidate the function of HUWE1 in specific types of cancer.
The present study interpreted multidimensional
oncogenic data. The use of databases contributes to improved
understanding of cancer molecular etiology and epidemiology, which
may assist with the translation of genomic understanding into
clinical practice (59). The current
study aimed to use extensive oncogenic databases to improve
understanding regarding the molecular mechanisms mediated by HUWE1.
In conclusion, HUWE1 may serve as a target for treatment strategies
and act as a biomarker for certain cancer types.
Acknowledgements
We would like to acknowledge all of the data
contributors of the TCGA, Oncomine, Kaplan-Meier plotter,
Prognoscan, STRING and cBioPortal databases, which generously
providing cyber resources for the scientists for the exploration,
visualization and analysis of the multidimensional genomics data of
cancer.
Funding
This study was supported by grants from the National
Natural Science Foundation of China (grant nos. 31501114 and
81502035), the Foundation of Xiamen City (grant no. 3502Z20174080)
and the Foundation of Fujian Province (grant no. 2018-ZQN-87).
Availability of data and materials
The data that support the findings of this study are
available from Oncomine, Kaplan-Meier plotter, Prognoscan, STRING
and cBioPortal databases, but restrictions apply to the
availability of these data, which were used under license for the
current study, and so are not publicly available. Data are however
available from the authors upon reasonable request and with
permission of Oncomine, Kaplan-Meier plotter, Prognoscan, STRING
and cBioPortal databases.
Authors' contributions
JH and CS conceived the study. CS, TW, JZ and JC
analyzed the data. JH, CS and TW wrote and revised the manuscript.
All authors read and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Zhong Q, Gao W, Du F and Wang X:
Mule/ARF-BP1, a BH3-only E3 ubiquitin ligase, catalyzes the
polyubiquitination of Mcl-1 and regulates apoptosis. Cell.
121:1085–1095. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Schreiber A and Peter M: Substrate
recognition in selective autophagy and the ubiquitin-proteasome
system. Biochim Biophys Acta. 1843:163–181. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Rotin D and Kumar S: Physiological
functions of the HECT family of ubiquitin ligases. Nat Rev Mol Cell
Biol. 10:398–409. 2009. View
Article : Google Scholar : PubMed/NCBI
|
|
4
|
Schrader EK, Harstad KG and Matouschek A:
Targeting proteins for degradation. Nat Chem Biol. 5:815–822. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Teixeira LK and Reed SI: Ubiquitin ligases
and cell cycle control. Annu Rev Biochem. 82:387–414. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Aleidi SM, Howe V, Sharpe LJ, Yang A, Rao
G, Brown AJ and Gelissen IC: The E3 ubiquitin ligases, HUWE1 and
NEDD4-1, are involved in the post-translational regulation of the
ABCG1 and ABCG4 lipid transporters. J Biol Chem. 290:24604–24613.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Clague MJ, Liu H and Urbé S: Governance of
endocytic trafficking and signaling by reversible ubiquitylation.
Dev Cell. 23:457–467. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Scheffner M and Kumar S: Mammalian HECT
ubiquitin-protein ligases: Biological and pathophysiological
aspects. Biochim Biophys Acta. 1843:61–74. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Sun Y: Targeting E3 ubiquitin ligases for
cancer therapy. Cancer Biol Ther. 2:623–629. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Scheffner M and Staub O: HECT E3s and
human disease. BMC Biochem. 8 (Suppl 1):S62007. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Leboucher GP, Tsai YC, Yang M, Shaw KC,
Zhou M, Veenstra TD, Glickman MH and Weissman AM: Stress-induced
phosphorylation and proteasomal degradation of mitofusin 2
facilitates mitochondrial fragmentation and apoptosis. Mol Cell.
47:547–557. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Zhou X, Li TT, Feng X, Hsiang E, Xiong Y,
Guan KL and Lei QY: Targeted polyubiquitylation of RASSF1C by the
Mule and SCFβ-TrCP ligases in response to DNA damage. Biochem J.
441:227–236. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Zhang J, Kan S, Huang B, Hao Z, Mak TW and
Zhong Q: Mule determines the apoptotic response to HDAC inhibitors
by targeted ubiquitination and destruction of HDAC2. Genes Dev.
25:2610–2618. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Chen D, Kon N, Li M, Zhang W, Qin J and Gu
W: ARF-BP1/Mule is a critical mediator of the ARF tumor suppressor.
Cell. 121:1071–1083. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Adhikary S, Marinoni F, Hock A, Hulleman
E, Popov N, Beier R, Bernard S, Quarto M, Capra M, Goettig S, et
al: The ubiquitin ligase HectH9 regulates transcriptional
activation by Myc and is essential for tumor cell proliferation.
Cell. 123:409–421. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Hall JR, Kow E, Nevis KR, Lu CK, Luce KS,
Zhong Q and Cook JG: Cdc6 stability is regulated by the Huwe1
ubiquitin ligase after DNA damage. Mol Biol Cell. 18:3340–3350.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Herold S, Hock A, Herkert B, Berns K,
Mullenders J, Beijersbergen R, Bernards R and Eilers M: Miz1 and
HectH9 regulate the stability of the checkpoint protein, TopBP1.
EMBO J. 27:2851–2861. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Yang Y, Do H, Tian X, Zhang C, Liu X, Dada
LA, Sznajder JI and Liu J: E3 ubiquitin ligase Mule ubiquitinates
Miz1 and is required for TNFalpha-induced JNK activation. Proc Natl
Acad Sci USA. 107:13444–13449. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Parsons JL, Tait PS, Finch D, Dianova II,
Edelmann MJ, Khoronenkova SV, Kessler BM, Sharma RA, McKenna WG and
Dianov GL: Ubiquitin ligase ARF-BP1/Mule modulates base excision
repair. EMBO J. 28:3207–3215. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Liu Z, Miao D, Xia Q, Hermo L and Wing SS:
Regulated expression of the ubiquitin protein ligase,
E3(Histone)/LASU1/Mule/ARF-BP1/HUWE1, during spermatogenesis. Dev
Dyn. 236:2889–2898. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Zhao X, Heng JI, Guardavaccaro D, Jiang R,
Pagano M, Guillemot F, Iavarone A and Lasorella A: The HECT-domain
ubiquitin ligase Huwe1 controls neural differentiation and
proliferation by destabilizing the N-Myc oncoprotein. Nat Cell
Biol. 10:643–653. 2008. View
Article : Google Scholar : PubMed/NCBI
|
|
22
|
Qi CF, Kim YS, Xiang S, Abdullaev Z,
Torrey TA, Janz S, Kovalchuk AL, Sun J, Chen D, Cho WC, et al:
Characterization of ARF-BP1/HUWE1 Interactions with CTCF, MYC, ARF
and p53 in MYC-Driven B Cell Neoplasms. Int J Mol Sci.
13:6204–6219. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Faiola F, Liu X, Lo S, Pan S, Zhang K,
Lymar E, Farina A and Martinez E: Dual regulation of c-Myc by p300
via acetylation-dependent control of Myc protein turnover and
coactivation of Myc-induced transcription. Mol Cell Biol.
25:10220–10234. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Vervoorts J, Lüscher-Firzlaff JM, Rottmann
S, Lilischkis R, Walsemann G, Dohmann K, Austen M and Lüscher B:
Stimulation of c-MYC transcriptional activity and acetylation by
recruitment of the cofactor CBP. EMBO Rep. 4:484–490. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Bernassola F, Karin M, Ciechanover A and
Melino G: The HECT family of E3 ubiquitin ligases: Multiple players
in cancer development. Cancer Cell. 14:10–21. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Myant KB, Cammareri P, Hodder MC, Wills J,
Von Kriegsheim A, Győrffy B, Rashid M, Polo S, Maspero E, Vaughan
L, et al: HUWE1 is a critical colonic tumour suppressor gene that
prevents MYC signalling, DNA damage accumulation and tumour
initiation. EMBO Mol Med. 9:181–197. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Peter S, Bultinck J, Myant K, Jaenicke LA,
Walz S, Müller J, Gmachl M, Treu M, Boehmelt G, Ade CP, et al:
Tumor cell-specific inhibition of MYC function using small molecule
inhibitors of the HUWE1 ubiquitin ligase. EMBO Mol Med.
6:1525–1541. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Rhodes DR, Yu J, Shanker K, Deshpande N,
Varambally R, Ghosh D, Barrette T, Pandey A and Chinnaiyan AM:
ONCOMINE: A cancer microarray database and integrated data-mining
platform. Neoplasia. 6:1–6. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Rhodes DR, Kalyana-Sundaram S, Mahavisno
V, Varambally R, Yu J, Briggs BB, Barrette TR, Anstet MJ,
Kincead-Beal C, Kulkarni P, et al: Oncomine 3.0: Genes, pathways,
and networks in a collection of 18,000 cancer gene expression
profiles. Neoplasia. 9:166–180. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Murat A, Migliavacca E, Gorlia T, Lambiv
WL, Shay T, Hamou MF, de Tribolet N, Regli L, Wick W, Kouwenhoven
MC, et al: Stem cell-related ‘self-renewal’ signature and high
epidermal growth factor receptor expression associated with
resistance to concomitant chemoradiotherapy in glioblastoma. J Clin
Oncol. 26:3015–3024. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Bredel M, Bredel C, Juric D, Harsh GR,
Vogel H, Recht LD and Sikic BI: Functional network analysis reveals
extended gliomagenesis pathway maps and three novel MYC-interacting
genes in human gliomas. Cancer Res. 65:8679–8689. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sun L, Hui AM, Su Q, Vortmeyer A,
Kotliarov Y, Pastorino S, Passaniti A, Menon J, Walling J, Bailey
R, et al: Neuronal and glioma-derived stem cell factor induces
angiogenesis within the brain. Cancer Cell. 9:287–300. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Finak G, Bertos N, Pepin F, Sadekova S,
Souleimanova M, Zhao H, Chen H, Omeroglu G, Meterissian S, Omeroglu
A, et al: Stromal gene expression predicts clinical outcome in
breast cancer. Nat Med. 14:518–527. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Gyorffy B, Lánczky A and Szállási Z:
Implementing an online tool for genome-wide validation of
survival-associated biomarkers in ovarian-cancer using microarray
data from 1287 patients. Endocr Relat Cancer. 19:197–208. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Mizuno H, Kitada K, Nakai K and Sarai A:
PrognoScan: A new database for meta-analysis of the prognostic
value of genes. BMC Med Genomics. 2:182009. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Franceschini A, Szklarczyk D, Frankild S,
Kuhn M, Simonovic M, Roth A, Lin J, Minguez P, Bork P, von Mering C
and Jensen LJ: STRING v9. 1: Protein-protein interaction networks,
with increased coverage and integration. Nucleic Acids Res 41
(Database Issue). D808–D815. 2013.
|
|
37
|
Gao J, Aksoy BA, Dogrusoz U, Dresdner G,
Gross B, Sumer SO, Sun Y, Jacobsen A, Sinha R, Larsson E, et al:
Integrative analysis of complex cancer genomics and clinical
profiles using the cBioPortal. Sci Signal. 6:pl12013. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Beltran H, Prandi D, Mosquera JM, Benelli
M, Puca L, Cyrta J, Marotz C, Giannopoulou E, Chakravarthi BV,
Varambally S, et al: Divergent clonal evolution of
castration-resistant neuroendocrine prostate cancer. Nat Med.
22:298–305. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Kumar A, Coleman I, Morrissey C, Zhang X,
True LD, Gulati R, Etzioni R, Bolouri H, Montgomery B, White T, et
al: Substantial interindividual and limited intraindividual genomic
diversity among tumors from men with metastatic prostate cancer.
Nat Med. 22:369–378. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Hodis E, Watson IR, Kryukov GV, Arold ST,
Imielinski M, Theurillat JP, Nickerson E, Auclair D, Li L, Place C,
et al: A landscape of driver mutations in melanoma. Cell.
150:251–263. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Cancer Genome Atlas Research Network, ;
Kandoth C, Schultz N, Cherniack AD, Akbani R, Liu Y, Shen H,
Robertson AG, Pashtan I, Shen R, et al: Integrated genomic
characterization of endometrial carcinoma. Nature. 497:67–73. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Campbell JD, Alexandrov A, Kim J, Wala J,
Berger AH, Pedamallu CS, Shukla SA, Guo G, Brooks AN, Murray BA, et
al: Distinct patterns of somatic genome alterations in lung
adenocarcinomas and squamous cell carcinomas. Nat Genet.
48:607–616. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Imielinski M, Berger AH, Hammerman PS,
Hernandez B, Pugh TJ, Hodis E, Cho J, Suh J, Capelletti M,
Sivachenko A, et al: Mapping the hallmarks of lung adenocarcinoma
with massively parallel sequencing. Cell. 150:1107–1120. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Comprehensive molecular profiling of lung
adenocarcinoma, . Nature. 511:543–550. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Liang Y, Diehn M, Watson N, Bollen AW,
Aldape KD, Nicholas MK, Lamborn KR, Berger MS, Botstein D, Brown PO
and Israel MA: Gene expression profiling reveals molecularly and
clinically distinct subtypes of glioblastoma multiforme. Proc Natl
Acad Sci USA. 102:5814–5819. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Haslinger C, Schweifer N, Stilgenbauer S,
Döhner H, Lichter P, Kraut N, Stratowa C and Abseher R: Microarray
gene expression profiling of B-cell chronic lymphocytic leukemia
subgroups defined by genomic aberrations and VH mutation status. J
Clin Oncol. 22:3937–3949. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Bhattacharjee A, Richards WG, Staunton J,
Li C, Monti S, Vasa P, Ladd C, Beheshti J, Bueno R, Gillette M, et
al: Classification of human lung carcinomas by mRNA expression
profiling reveals distinct adenocarcinoma subclasses. Proc Natl
Acad Sci USA. 98:13790–13795. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Garber ME, Troyanskaya OG, Schluens K,
Petersen S, Thaesler Z, Pacyna-Gengelbach M, van de Rijn M, Rosen
GD, Perou CM, Whyte RI, et al: Diversity of gene expression in
adenocarcinoma of the lung. Proc Natl Acad Sci USA. 98:13784–13789.
2001. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Storz MN, van de Rijn M, Kim YH,
Mraz-Gernhard S, Hoppe RT and Kohler S: Gene expression profiles of
cutaneous B cell lymphoma. J Invest Dermatol. 120:865–870. 2003.
View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Skotheim RI, Lind GE, Monni O, Nesland JM,
Abeler VM, Fosså SD, Duale N, Brunborg G, Kallioniemi O, Andrews PW
and Lothe RA: Differentiation of human embryonal carcinomas in
vitro and in vivo reveals expression profiles relevant to normal
development. Cancer Res. 65:5588–5598. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Luo JH, Yu YP, Cieply K, Lin F, Deflavia
P, Dhir R, Finkelstein S, Michalopoulos G and Becich M: Gene
expression analysis of prostate cancers. Mol Carcinog. 33:25–35.
2002. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Varambally S, Yu J, Laxman B, Rhodes DR,
Mehra R, Tomlins SA, Shah RB, Chandran U, Monzon FA, Becich MJ, et
al: Integrative genomic and proteomic analysis of prostate cancer
reveals signatures of metastatic progression. Cancer Cell.
8:393–406. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Detwiller KY, Fernando NT, Segal NH, Ryeom
SW, D'Amore PA and Yoon SS: Analysis of hypoxia-related gene
expression in sarcomas and effect of hypoxia on RNA interference of
vascular endothelial cell growth factor A. Cancer Res.
65:5881–5889. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Klonowska K, Czubak K, Wojciechowska M,
Handschuh L, Zmienko A, Figlerowicz M, Dams-Kozlowska H and
Kozlowski P: Oncogenomic portals for the visualization and analysis
of genome-wide cancer data. Oncotarget. 7:176–192. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Xie S, Shen C, Tan M, Li M, Song X and
Wang C: Systematic analysis of gene expression alterations and
clinical outcomes of adenylate cyclase-associated protein in
cancer. Oncotarget. 8:27216–27239. 2017.PubMed/NCBI
|
|
57
|
Stratton MR, Campbell PJ and Futreal PA:
The cancer genome. Nature. 458:719–724. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Beroukhim R, Mermel CH, Porter D, Wei G,
Raychaudhuri S, Donovan J, Barretina J, Boehm JS, Dobson J,
Urashima M, et al: The landscape of somatic copy-number alteration
across human cancers. Nature. 463:899–905. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
59
|
Hanahan D and Weinberg RA: Hallmarks of
cancer: The next generation. Cell. 144:646–674. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
60
|
National Cancer Institute: Genomic Data
Commons Data Portal. https://portal.gdc.cancer.gov/August
11–2017
|