Introduction
Gastric cancer is one of the most common types of
tumor and is the fourth most common cause of cancer-related
mortality worldwide. Although the incidence of gastric cancer has
been markedly reduced in certain developed countries over the past
few decades, an estimated one million new patients are diagnosed
annually (1). The tumorigenesis of
gastric cancer is a complicated process that involves the
deregulation of a variety of genes (2,3).
Despite recent developments in the treatment of gastric cancer, the
prognosis of patients with advanced gastric cancer remains poor.
Therefore, investigation of the underlying molecular mechanism of
gastric cancer progression is urgently required.
microRNAs (miRNAs/miRs) are a group of small
non-coding RNAs, which are known to function by negatively
regulating the expression of downstream genes through base pairing
with the 3′-untranslated region (3′-UTR) of the corresponding mRNAs
(4,5). Increasing evidence indicates that
miRNAs are critical in the proliferation, apoptosis, migration,
invasion and metabolism of tumor cells (6). miRNA deregulation has been detected in
a number of different types of cancer, including breast, lung,
prostate and gastric cancer (7–10).
miRNAs may function as oncogenes or tumor suppressor genes,
depending on the target genes regulated. For example, miR-22, miR-7
and miR-101 have been found to be downregulated in tumors and
function as tumor suppressors (11–13),
whereas miR-21 and miR-17 have been observed to be upregulated in
tumors and function as oncogenes (14,15).
The role of miR-183 in tumors is controversial; for instance,
miR-183 has been found to be downregulated and inhibit cell
migration and invasion in breast cancer and osteosarcoma (16,17);
conversely, miR-183 has been revealed to be overexpressed and
promote tumor progression in synovial sarcoma, rhabdomyosarcoma and
colon cancer (18). However, the
biological role of miR-183 in gastric cancer remains unclear. The
aim of the present study was to investigate the biological role and
clinical significance of miR-183 in gastric cancer in order to
explore its potential application as a prognostic marker and
therapeutic target for gastric cancer patients.
Materials and methods
Human specimens
All experimental procedures were approved by the
Institutional Review Board of Wuhan University of Science and
Technology (Wuhan, China). Human gastric cancer tissues (n=65) and
matched adjacent normal tissues were obtained from patients who
underwent surgery at Tianyou Hospital of Wuhan University of
Science and Technology between June 2007 and March 2011. None of
the patients had received any treatment prior to resection. Written
informed consent was obtained from the patients. The
clinicopathological data were collected from medical records.
Cell culture
HEK293 human embryonic kidney cells, GES-1
immortalized normal gastric mucosa cells, and AGS, SGC7901, MKN28,
MGC803 and HGC27 gastric cancer cells were purchased from the
American Type Culture Collection (Rockville, MD, USA) and cultured
according to the manufacturer’s instructions. All transfections
were performed using Lipofectamine 2000 (Invitrogen Life
Technologies, Carlsbad, CA, USA).
RNA isolation and reverse transcription
quantitative polymerase chain reaction (RT-qPCR) analysis
Total RNA was isolated from the tissues or cells
using TRIzol reagent (Invitrogen Life Technologies) according to
the manufacturer’s instructions. miR-183 expression levels were
measured using a mirVana qRT-PCR miRNA Detection kit (Ambion Inc.,
Austin, TX, USA). U6 small RNA served as an internal reference.
Bmi-1 mRNA expression levels were measured using a SYBR Premix Ex
Taq™ kit (Takara Bio, Inc., Shiga, Japan), and β-actin expression
levels served as an endogenous control. RT-qPCR was conducted using
an Applied Biosystems 7,900 Fast Real-Time PCR system (Applied
Biosystems, Foster City, CA, USA). cDNA was synthesized from 200 ng
of total RNA, using a high-capacity cDNA Reverse Transcription Kit
(Applied Biosystems) in a total volume of 20 μl per reaction. The
sequences of the primers were as follows: Forward,
5′-GCTTCAAGATGGCCGCTTG-3′ and reverse, 5′-TTCTCGTTGT
TCGATGCATTTC-3′ for Bmi-1; and forward, 5′-TGGATCAGCAAGCAGGAGTA-3′
and reverse, 5′-TCGGCCACATTGTGAACTTT-3′. Quantitative PCR was
performed as follows: heating at 95°C for 5 min, followed by 95°C
for 15 sec, 60C for 15 sec and 72C for 32 sec for 40 cycles. The
data were analyzed using the 2−ΔΔCt method.
Western blot analysis
Total cell lysate was extracted from the cells and
tissues using RIPA reagent (Beyotime Institute of Biotechnology,
Shanghai, China). Proteins were separated using SDS-PAGE and
transferred to polyvinylidene difluoride membranes (Sterlitech
Corporation, Kent, WA, USA). The membranes were then blocked with
5% skimmed milk in Tris-buffered saline with Tween 20 buffer (TBST;
10 mmol/l Tris-HCl, 150 mmol/l NaCl and 0.1% Tween 20; pH 8.0) for
1 h at room temperature then incubated with primary antibodies,
including monoclonal anti-human antibodies Bmi-1 (1:500, Santa Cruz
Biotechnology, Inc., Santa Cruz, CA, USA) and polyclonal rabbit
antibody GAPDH (1:1,000; Sigma-Aldrich, St. Louis, MO, USA)
overnight at 4°C. Following three washes with TBST, the membranes
were incubated with a goat anti-rabbit secondary antibody (Boster
Systems, Inc., Pleasanton, CA, USA) for 1 h. The bands were then
visualized using the enhanced chemiluminescence system (Amersham
Pharmacia Biotech, Amersham, UK) following the manufacturer’s
instructions.
Oligonucleotide transfection
The hsa-miR-183 mimic and negative control (NC)
oligonucleotides were obtained from Guangzhou Ribobio Co., Ltd.
(Guangzhou, China). SGC7901 and AGS cells were plated in a six-well
plate the day prior to transfection. The cells were transfected
with hsa-miR-183 mimic or NC (50 nmol/l) using Lipofectamine 2000.
After 24 h, the cells were collected to perform in vitro
assays.
Cell proliferation assay
At 24 h post-transfection, the cells were seeded in
96-well plates (2×103 cells/well). The viability of the
cells was examined by MTT assay (Sigma-Aldrich), conducted daily
for five days.
Colony formation assay
For the colony formation assay, 500 transfected
cells were placed in a six-well plate and cultured for 14 days
using RPMI 1640 medium (Gibco-BRL, Carlsbad, CA, USA) containing
10% fetal bovine serum. Cell colonies were fixed with methanol and
then stained with 0.1% crystal violet. Colonies were observed using
a microscope and colonies containing >50 cells were counted
(IX71, Olympus Corporation, Tokyo, Japan).
Cell invasion assay
The cell invasive capacity was assessed with
specialized Transwell chambers (8-μm pore; BD Biosciences, Franklin
Lakes, NJ, USA). The transfected cells (5×104
cells/well) were added to the upper chamber of the inserts, which
was coated with a Matrigel mix; 500 μl fetal bovine serum was added
to the lower chamber as a chemoattractant. After 24 h, any
non-invading cells on the upper surface were removed with swabs and
the cells that had migrated to the lower side of the membrane were
fixed with methanol, stained with 0.1% crystal violet and
air-dried. Images of the cells were then captured. The number of
invading cells was evaluated in five fields using microscopy. The
mean of triplicate assays for each experimental condition was
analyzed.
Vector construction and dual-luciferase
reporter assay
For the luciferase assays, the potential miR-183
binding site in the 3′-UTR of Bmi-1 mRNA was predicted by
TargetScan (www.targetscan.org) and miRanda
(www.microrna.org). Wild-type (wt) and mutant (mt)
Bmi-1 mRNA 3′-UTRs were synthesized and cloned into the Xba
I site of a pGL3 basic vector (Promega Corporation, Madison, WI,
USA) downstream of the luciferase stop codon, and the resulting
vectors were termed pGL3-wt-Bmi-1 and pGL3-mt-Bmi-1, respectively.
The HEK293 cells were cultured in 24-well plates and co-transfected
with pGL3-Control (0.4 mg), pGL3-wt-Bmi-1 (0.4 mg) or pGL3-mt-Bmi-1
(0.4 mg), plus pRL-TK luciferase reporters (25 ng/well) and
pcDNA-miR-183 (20 nm) or pcDNA-miR-NC (20 nm) using Lipofectamine
2000. After 48 h, the cells were collected and luciferase activity
was assessed using a Dual-Luciferase Reporter Assay kit (Promega
Corporation).
Statistical analysis
The data are expressed as the mean ± standard
deviation. Statistical significance was analyzed using Student’s
t-test (two-tailed) or the χ2 test. All statistical
analyses were performed using GraphPad Prism 5.0 (GraphPad
Software, Inc., La Jolla, CA, USA) and SPSS 13.0 (SPSS, Inc.,
Chicago, IL, USA) software packages. The Kaplan-Meier method with
log-rank test was used to analyze the prognostic significance.
P<0.05 was considered to indicate a statistically significant
difference.
Results
miR-183 is significantly downregulated in
gastric cancer cell lines and tissues
miR-183 expression levels were measured in the
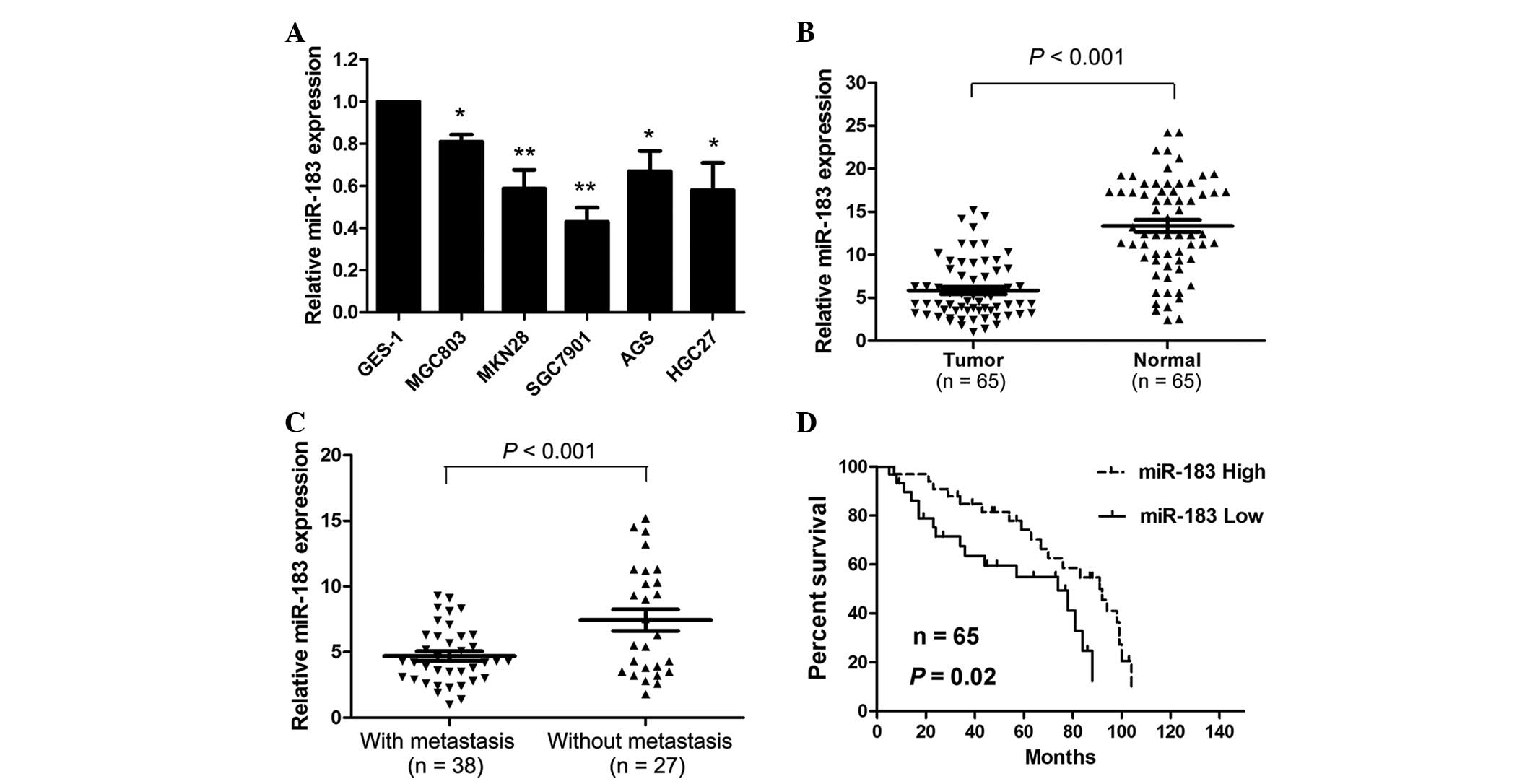
gastric cancer tissues and cell lines. miR-183 was significantly
downregulated in the gastric cancer cell lines compared with the
GES-1 normal gastric epithelial cell line (Fig. 1A). Of all six gastric cancer cell
lines, SGC7901 exhibited the lowest levels of miR-183 expression,
whereas MGC803 possessed the highest levels of miR-183
expression.
To examine the clinicopathological significance of
miR-183 expression in gastric cancer patients, miR-183 expression
levels were measured in 65 gastric cancer tissue samples and
adjacent normal tissue specimens. miR-183 expression levels were
significantly reduced in the tumor tissues compared with the normal
tissues (P<0.001; Fig. 1B).
Furthermore, miR-183 expression levels were significantly reduced
in the tissues with distant metastasis compared with the tissues
without distant metastasis (P<0.001; Fig. 1C). Subsequently, the patients were
divided into two groups, as determined by the median miR-183
expression levels in each patient. Low miR-183 expression levels
were significantly associated with increased tumor size (P=0.003),
the presence of distant metastasis (P=0.018) and increased
tumor-node-metastasis stage (P=0.019; Table I). In addition, patients with low
miR-183 levels had significantly lower survival rates compared with
patients with high miR-183 levels (P=0.020; Fig. 1D).
 | Table ICorrelation between miR-183 expression
levels and clinicopathological characteristics in gastric cancer
patients. |
Table I
Correlation between miR-183 expression
levels and clinicopathological characteristics in gastric cancer
patients.
| | miR-183
expression | |
|---|
| |
| |
|---|
| Characteristic | Patients, n | Low, n (%) | High, n (%) | P-value |
|---|
| Age, years | | | | 0.261 |
| <60 | 34 | 19 (51) | 15 (45) | |
| ≥60 | 31 | 13 (49) | 18 (55) | |
| Gender | | | | 0.371 |
| Male | 37 | 20 (62) | 17 (51) | |
| Female | 28 | 12 (38) | 16 (49) | |
| CEA level, ng/ml | | | | 0.108 |
| 0–5 | 35 | 14 (43) | 21 (63) | |
| >5 | 30 | 18 (57) | 12 (37) | |
| CA19-9 level,
U/ml | | | | 0.518 |
| 0–35 | 49 | 23 (71) | 26 (78) | |
| >35 | 16 | 9 (29) | 7 (22) | |
| Tumor size, cm | | | | 0.003 |
| ≤5 | 44 | 16 (50) | 28 (84) | |
| >5 | 21 | 16 (50) | 5 (16) | |
| Differentiation | | | | 0.321 |
| Well/moderate | 45 | 24 (75) | 21 (63) | |
| Poor | 20 | 8 (25) | 12 (37) | |
| Distant
metastasis | | | | 0.018 |
| Yes | 38 | 14 (43) | 24 (72) | |
| No | 27 | 18 (57) | 9 (28) | |
| TNM stage | | | | 0.019 |
| I/II | 14 | 3 (9) | 11 (33) | |
| III/IV | 51 | 29 (91) | 22 (67) | |
miR-183 inhibits proliferation and
invasion in gastric cancer cells
As miR-183 is downregulated in gastric cancer
tissues and cell lines, miR-183 may function as a tumor suppressor
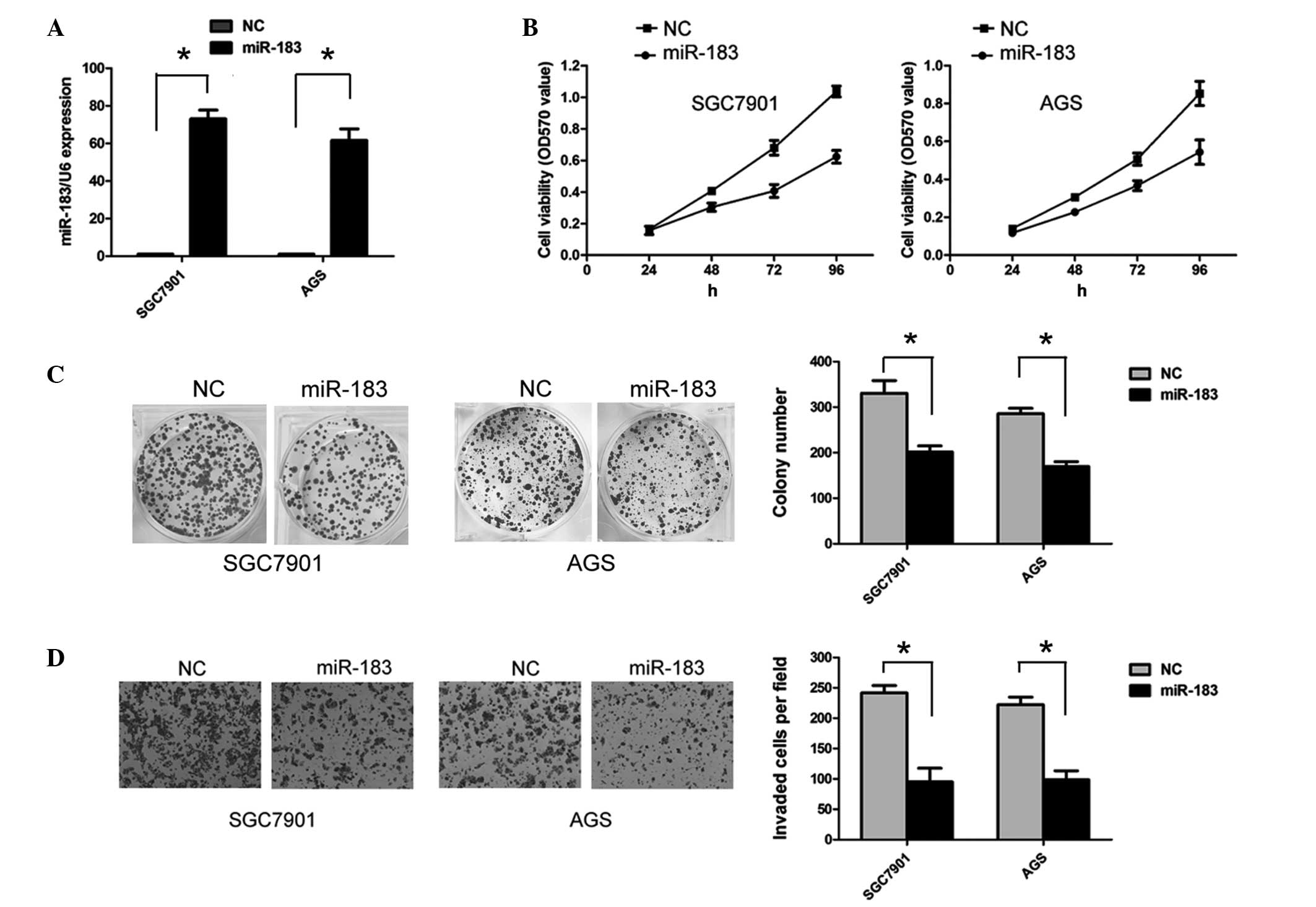
in gastric cancer. SGC7901 and AGS cells were transfected with
miR-183 mimics to overexpress miR-183. The efficiency of
transfection was confirmed by RT-qPCR (Fig. 2A). As shown in Fig. 2B, the ectopic expression of miR-183
significantly inhibited SGC7901 and AGS cell viability (P<0.05).
In addition, ectopic miR-183 expression significantly reduced the
numbers of SGC7901 and AGS cell colonies that formed compared with
the negative control (both P<0.05; Fig. 2C). To assess the effect of miR-183
on the invasive abilities of gastric cancer cells, a Transwell
assay was performed. Overexpression of miR-183 significantly
suppressed the invasive ability of the SGC7901 and AGS cells (both
P<0.05; Fig. 2D).
Bmi-1 is a direct downstream target of
miR-183 in gastric cancer cells
To analyze the underlying mechanism of miR-183 in
gastric cancer, TargetScan (www.targetscan.org) and miRanda (www.microrna.org) were used to search for potential
targets of miR-183 in gastric cancer. Bmi-1 was predicted as a
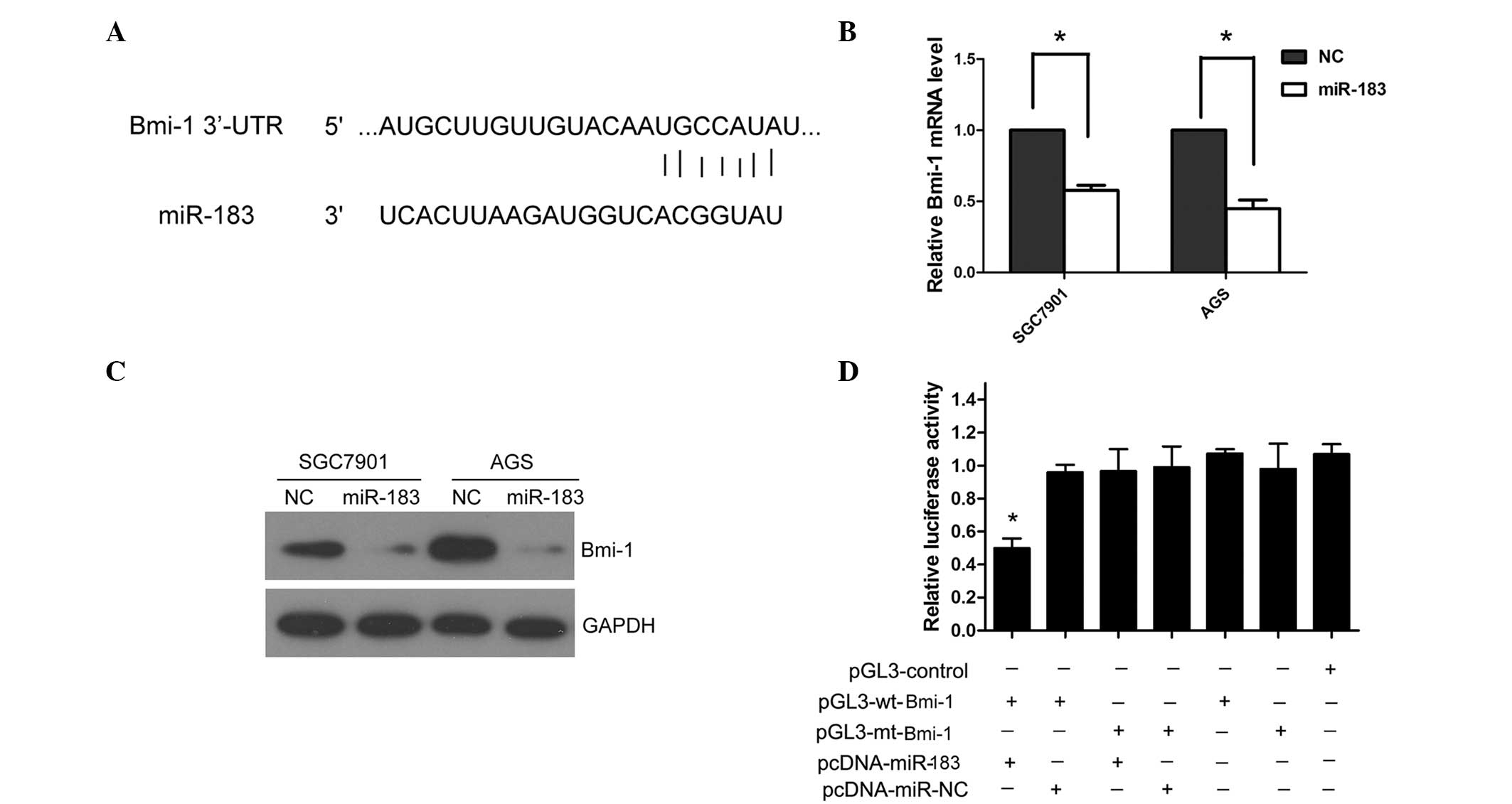
potential target of miR-183 by TargetScan and miRanda. The 3′-UTR
of Bmi-1 contains the binding site for the seed region of miR-183
(Fig. 3A). To investigate whether
miR-183 regulates Bmi-1 expression in the cellular environment, the
SGC7901 and AGS cells were transfected with miR-183 mimics to
overexpress miR-183. The ectopic expression of miR-183
significantly reduced the Bmi-1 mRNA and protein levels in the
SGC7901 and AGS cells (P=0.027; Fig. 3B
and C). To further confirm that Bmi-1 is the direct target of
miR-183 in gastric cancer, human Bmi-1 3′-UTR, containing either
the wt or mt miR-183 binding sequence was cloned downstream of the
firefly luciferase reporter gene in the pGL3 vector. HEK293 cells
were transfected with either of the two types of reporter plasmid,
plus either miR-183 mimics or NC vector. The luciferase activity of
the reporter in the vector containing the Bmi-1 3′-UTR wt was
significantly reduced by miR-183 mimics (P=0.009); however, the
luciferase activity of the Bmi-1 3′-UTR mt was not significantly
affected (Fig. 3D).
Discussion
miRNAs have been found to be frequently dysregulated
in gastric cancer, and certain miRNAs have been reported to be
correlated with clinical characteristics and outcome. For example,
the upregulation of miR-221 has been associated with an adverse
clinical prognosis (19), and
downregulation of miR-7 has been shown to be responsible for an
aggressive tumor phenotype and inhibiting tumor metastasis
(20). Furthermore, increasing
evidence indicates that miRNAs are important in gastric cancer
progression by regulating cell proliferation, apoptosis, migration
and invasion (21–23). Thus, identifying the specific miRNAs
involved in gastric cancer progression may provide novel
therapeutic targets in gastric cancer treatment.
In the present study, miR-183 was demonstrated to be
significantly reduced in gastric cancer tissues and cell lines
compared with adjacent normal tissues and non-cancer cell lines,
respectively. miR-183 has previously been found to be downregulated
in osteosarcoma; thus, miR-183 may be able to inhibit cell
migration and invasion (24).
Conversely, miR-183 has been reported to be overexpressed in
prostate and colorectal cancer, stimulating tumor progression and
metastasis (25,26). These results indicate that miR-183
may exert distinct functions in different tumor types. This may be
due to the genetic heterogeneity and the varying target genes in
the different tumor types. In the present study, downregulation of
miR-183 was found to be associated with aggressive
clinicopathological characteristics and a poor prognosis in gastric
cancer. Contrary to these results, a previous study found higher
miR-183 expression levels to be a risk factor associated with
shorter survival times in lung cancer patients (27). These results demonstrate that one
miRNA may exert varying roles in different cancer types. However,
further investigation is required to confirm whether miR-183 is a
common predictor of survival rates in tumor patients.
As miR-183 expression was downregulated in the
gastric cancer tissues and cell lines in the present study,
analysis was then focused on whether miR-183 affects the biological
behavior of gastric cancer cells. Notably, ectopic miR-183
expression significantly reduced gastric cancer cell proliferation,
colony formation and invasive ability. Previous studies have
indicated that miR-183 promotes or inhibits tumor progression and
metastasis in various types of tumor. For instance, miR-183
promotes cell invasion in prostate and colorectal cancer (25,26),
whereas it inhibits tumor progression in other types of tumor,
including osteosarcoma and breast cancer (17,16).
miRNAs are considered to exert effects by regulating
target mRNAs. Previously, Tanaka et al (28) reported that miR-183 targets
isocitrate dehydrogenase 2 in glioma cells, and Zhu et al
(24) revealed that Ezrin is a
target of miR-183 in osteosarcoma. In the present study, several
downstream target mRNAs of miR-183 were identified, including
Bmi-1, ITGB1 and PDCD4. However, only Bmi-1 was confirmed as a
target of miR-183 in the gastric cancer cells. Subsequent
experiments demonstrated that ectopic miR-183 expression reduced
the endogenous Bmi-1 mRNA and protein levels. A luciferase activity
assay revealed that miR-183 overexpression reduced Bmi-1 3′-UTR wt
luciferase activity, but not that of the mt. In addition, Bmi-1
protein expression was upregulated in the gastric cancer cell lines
and tissues (data not shown). These results demonstrate that
miR-183 regulates Bmi-1 in gastric cancer cells.
Bmi-1 has been reported as an oncogene that controls
cell proliferation and invasion (29). Furthermore, Bmi-1 is also involved
in the self-renewal of cancer stem cells, which may result in tumor
initiation (30,31). In the present study, Bmi-1 was found
to be significantly overexpressed in the gastric cancer tissues and
cell lines compared with the adjacent normal tissues and GES-1
normal gastric epithelial cells, respectively. In accordance with
these findings, several other studies have found that Bmi-1 is
upregulated in gastric cancer and is associated with decreased
survival rates in patients (32,33).
These results further indicate that miR-183 functions as a tumor
suppressor by targeting Bmi-1.
In conclusion, the present study demonstrated that
miR-183 is significantly downregulated in gastric cancer tissues
and cell lines. miR-183 expression levels were also associated with
clinicopathological parameters and survival rate in gastric cancer
patients. Ectopic miR-183 expression resulted in inhibition of
gastric cancer cell proliferation, colony formation and invasion.
Further investigation revealed that Bmi-1 was a potential target of
miR-183. Therefore, miR-183 may serve as a predictor for prognosis
and as a therapeutic target in gastric cancer patients.
References
|
1
|
Jemal A, Bray F, Center MM, et al: Global
cancer statistics. CA Cancer J Clin. 61:69–90. 2011.
|
|
2
|
Houghton J, Stoicov C, Nomura S, et al:
Gastric cancer originating from bone marrow-derived cells. Science.
306:1568–1571. 2004.
|
|
3
|
Zheng L, Wang L, Ajani J and Xie K:
Molecular basis of gastric cancer development and progression.
Gastric Cancer. 7:61–77. 2004.
|
|
4
|
Lewis BP, Burge CB and Bartel DP:
Conserved seed pairing, often flanked by adenosines, indicates that
thousands of human genes are microRNA targets. Cell. 120:15–20.
2005.
|
|
5
|
Lee Y, Ahn C, Han J, et al: The nuclear
RNase III Drosha initiates microRNA processing. Nature.
425:415–419. 2003.
|
|
6
|
Lu J, Getz G, Miska EA, et al: MicroRNA
expression profiles classify human cancers. Nature. 435:834–838.
2005.
|
|
7
|
Ma L, Teruya-Feldstein J and Weinberg RA:
Tumour invasion and metastasis initiated by microRNA-10b in breast
cancer. Nature. 449:682–688. 2007.
|
|
8
|
Yanaihara N, Caplen N, Bowman E, et al:
Unique microRNA molecular profiles in lung cancer diagnosis and
prognosis. Cancer Cell. 9:189–198. 2006.
|
|
9
|
Schaefer A, Jung M, Mollenkopf HJ, et al:
Diagnostic and prognostic implications of microRNA profiling in
prostate carcinoma. Int J Cancer. 126:1166–1176. 2010.
|
|
10
|
Kim YK, Yu J, Han TS, et al: Functional
links between clustered microRNAs: suppression of cell-cycle
inhibitors by microRNA clusters in gastric cancer. Nucleic Acids
Res. 37:1672–1681. 2009.
|
|
11
|
Zhang J, Yang Y, Yang T, et al:
microRNA-22, downregulated in hepatocellular carcinoma and
correlated with prognosis, suppresses cell proliferation and
tumourigenicity. Br J Cancer. 103:1215–1220. 2010.
|
|
12
|
Friedman JM, Jones PA and Liang G: The
tumor suppressor microRNA-101 becomes an epigenetic player by
targeting the polycomb group protein EZH2 in cancer. Cell Cycle.
8:2313–2314. 2009.
|
|
13
|
Kefas B, Godlewski J, Comeau L, et al:
microRNA-7 inhibits the epidermal growth factor receptor and the
Akt pathway and is down-regulated in glioblastoma. Cancer Res.
68:3566–3572. 2008.
|
|
14
|
Marquez RT, Wendlandt E, Galle CS, Keck K
and McCaffrey AP: MicroRNA-21 is upregulated during the
proliferative phase of liver regeneration, targets Pellino-1, and
inhibits NF-kappaB signaling. Am J Physiol Gastrointest Liver
Physiol. 298:G535–G541. 2010.
|
|
15
|
Sun H, Li QW, Lv XY, et al: MicroRNA-17
post-transcriptionally regulates polycystic kidney disease-2 gene
and promotes cell proliferation. Mol Biol Rep. 37:2951–2958.
2010.
|
|
16
|
Lowery AJ, Miller N, Dwyer RM and Kerin
MJ: Dysregulated miR-183 inhibits migration in breast cancer cells.
BMC Cancer. 10:5022010.
|
|
17
|
Zhao H, Guo M, Zhao G, et al: miR-183
inhibits the metastasis of osteosarcoma via downregulation of the
expression of Ezrin in F5M2 cells. Int J Mol Med. 30:1013–1020.
2012.
|
|
18
|
Sarver AL, Li L and Subramanian S:
MicroRNA miR-183 functions as an oncogene by targeting the
transcription factor EGR1 and promoting tumor cell migration.
Cancer Res. 70:9570–9580. 2010.
|
|
19
|
Liu K, Li G, Fan C, et al: Increased
Expression of MicroRNA-221 in Gastric Cancer and Its Clinical
Significance. J Int Med Res. 40:467–474. 2012.
|
|
20
|
Zhao X, Dou W, He L, et al: MicroRNA-7
functions as an anti-metastatic microRNA in gastric cancer by
targeting insulin-like growth factor-1 receptor. Oncogene.
32:1363–1372. 2013.
|
|
21
|
Li Z, Cao Y, Jie Z, et al: miR-495 and
miR-551a inhibit the migration and invasion of human gastric cancer
cells by directly interacting with PRL-3. Cancer Lett. 323:41–47.
2012.
|
|
22
|
Carvalho J, van Grieken NC, Pereira PM, et
al: Lack of microRNA-101 causes E-cadherin functional deregulation
through EZH2 up-regulation in intestinal gastric cancer. J Pathol.
228:31–44. 2012.
|
|
23
|
Gao P, Xing AY, Zhou GY, et al: The
molecular mechanism of microRNA-145 to suppress invasion-metastasis
cascade in gastric cancer. Oncogene. 32:491–501. 2013.
|
|
24
|
Zhu J, Feng Y, Ke Z, et al:
Down-regulation of miR-183 promotes migration and invasion of
osteosarcoma by targeting Ezrin. Am J Pathol. 180:2440–2451.
2012.
|
|
25
|
Ueno K, Hirata H, Shahryari V, et al:
microRNA-183 is an oncogene targeting Dkk-3 and SMAD4 in prostate
cancer. Br J Cancer. 108:1659–1667. 2013.
|
|
26
|
Zhou T, Zhang GJ, Zhou H, Xiao HX and Li
Y: Overexpression of microRNA-183 in human colorectal cancer and
its clinical significance. Eur J Gastroenterol Hepatol. 26:229–223.
2014.
|
|
27
|
Zhu W, Liu X, He J, et al: Overexpression
of members of the microRNA-183 family is a risk factor for lung
cancer: a case control study. BMC Cancer. 11:3932011.
|
|
28
|
Tanaka H, Sasayama T, Tanaka K, et al:
MicroRNA-183 upregulates HIF-1alpha by targeting isocitrate
dehydrogenase 2 (IDH2) in glioma cells. J Neurooncol. 111:273–283.
2013.
|
|
29
|
Kang MK, Kim RH, Kim SJ, et al: Elevated
Bmi-1 expression is associated with dysplastic cell transformation
during oral carcinogenesis and is required for cancer cell
replication and survival. Br J Cancer. 96:126–133. 2007.
|
|
30
|
He S, Iwashita T, Buchstaller J, et al:
Bmi-1 over-expression in neural stem/progenitor cells increases
proliferation and neurogenesis in culture but has little effect on
these functions in vivo. Dev Biol. 328:257–272. 2009.
|
|
31
|
Douglas D, Hsu JH, Hung L, et al: BMI-1
promotes ewing sarcoma tumorigenicity independent of CDKN2A
repression. Cancer Res. 68:6507–6515. 2008.
|
|
32
|
Liu JH, Song LB, Zhang X, et al: Bmi-1
expression predicts prognosis for patients with gastric carcinoma.
J Surg Oncol. 97:267–272. 2008.
|
|
33
|
Lu H, Sun HZ, Li H and Cong M: The
clinicopathological significance of Bmi-1 expression in
pathogenesis and progression of gastric carcinomas. Asian Pac J
Cancer Prev. 13:3437–3441. 2012.
|