Introduction
Epithelial ovarian cancer has been reported to be
the second most common gynecological cancer and accounts for nearly
half of all deaths associated with gynecological pelvic
malignancies (1). Although
improvement of the quality of cytoreductive surgery as well as the
development of novel drugs and new chemotherapy regimens for
ovarian carcinoma therapy have been made, prognosis remains poor
for as many as 50% of the patients who will experience recurrence
and die of secondary disease within 5 years after diagnosis
(2). Thus, it is needed to identify
some novel molecular mechanisms involved in ovarian cancer
initiation and development.
microRNA (miRNA) belongs to a class of endogenously
expressed, non-coding small RNAs, which have emerged as key
post-transcriptional regulators of gene expression (3). In mammals, miRNAs have been predicted
to control the activity of approximately 30% of all protein-coding
genes, and have been shown to participate in the regulation of
almost every cellular process investigated so far (4,5).
Recently, dysregulation of miRNAs has been found to be involved in
a variety of human diseases including cancer (6,7). Up to
date, many researches have studied miRNA expression in ovarian
carcinoma by using various gene expression profiling approaches
(8). By an integrative genomic
approach, Zhang et al showed that out of 35 miRNAs
deregulated between ovarian carcinoma and the normal controls
(immortalized ovarian surface epithelial cells), 31 (88.6%) were
downregulated in cancer compared to non-cancer tissues (9). By miRNA microarrays, 36 miRNAs were
found to be deregulated between normal ovarian cells and epithelial
ovarian tumors, with miR-199a*, miR-214, miR-200a, and
miR-100 being the most highly differentially expressed candidates
(10). In particular, miR-100 was
found to be downregulated in 76% of tumors. In another research, a
subset of 37 miRNAs was found to be overexpressed in all epithelial
ovarian cancer subtypes and 21 were underexpressed and the
validated downregulated genes included miR-100, miR-210, miR-22 and
miR-222 (11). Although miR-100 was
reported to be significantly downregulated in EOC tissues, the
correlation of miR-100 expression with clinicopathological factors
or prognosis of patients with EOC and its functional roles in EOC
remain unclear.
In this study, our aim was to determine the
expression of miR-100 in 98 EOC tissues and corresponding adjacent
normal epithelial tissues. Then, the clinicopathological or
prognostic significance of miR-100 expression in human EOCs was
statistically analyzed. Next, a miR-100 inhibitor or precusor was
transiently transfected into EOC cell lines, and the effect of
miR-100 expression on the growth of EOC cells was analyzed.
Finally, whether polo-like kinase 1 (PLK1) was a target regulated
by miR-100 expression was also determined.
Materials and methods
Patients and tissue samples
A total of 98 fresh surgical tissue samples of EOC
and 15 adjacent normal epithelial tissues were collected at the
Jiangsu Province Hospital between 2002 and 2004, after informed
consent had been obtained. An independent pathologist assigned
histopathology and tumor grade according to International
Federation of Gynecology and Obstetrics (FIGO) criteria. A
gynecologic oncologist reviewed tumor stage and residual disease.
Normal tissues were obtained from tumor-free participants that had
undergone oophorectomy. Namely, they underwent surgery for a total
hysterectomy, bilateral salpingo-oophorectomy, partial omentectomy,
appendectomy, and pelvic and para-aortic lymphadenectomies. The
clinicopathological variables of patients were shown in Table I. All tissue samples were
snap-frozen in liquid nitrogen, which were transferred to 500 ml
TRIzol solution (Invitrogen, Carlsbad, CA, USA) immediately after
harvesting in order to avoid mRNA degradation. The samples were
stored in a biobank at −80°C until processed.
 | Table IAssociation of miR-100 expression with
clinicopathological variables of EOC patients. |
Table I
Association of miR-100 expression with
clinicopathological variables of EOC patients.
| Low miR-100
expression (n=50) | High miR-100
expression (n=48) | |
|---|
|
|
| |
|---|
| Variables | n (%) | n (%) | P-value |
|---|
| Age (years) | | | 0.155 |
| ≥50 | 36 (72.0) | 28 (58.3) | |
| <50 | 14 (28.0) | 20 (41.7) | |
| Histological
type | | | 0.486 |
| Serous | 16 (32.0) | 11 (22.9) | |
| Mucinous | 21 (42.0) | 20 (41.7) | |
| Others | 13 (26.0) | 17 (35.4) | |
| Histological
grade | | | 0.849 |
| G1 | 25 (50.0) | 22 (45.8) | |
| G2 | 10 (20.0) | 9 (18.8) | |
| G3 | 15 (30.0) | 17 (35.4) | |
| FIGO stage | | | 0.001 |
| I/II | 14 (28.0) | 29 (60.4) | |
| III/IV | 36 (72.0) | 19 (39.6) | |
| Residual tumor
diameter (cm) | | | 0.366 |
| <1.0 | 20 (40.0) | 15 (31.3) | |
| ≥1.0 | 30 (60.0) | 33 (68.7) | |
| Ascites | | | 0.279 |
| No | 22 (44.0) | 16 (33.3) | |
| Yes | 28 (56.0) | 32 (66.7) | |
| Serum CA125 level
(U/l) | | | 0.001 |
|
<5.0×105 | 15 (30.0) | 30 (62.5) | |
|
≥5.0×105 | 35 (70.0) | 18 (37.5) | |
| Lymph node
involvement | | | 0.014 |
| No | 21 (42.0) | 32 (66.7) | |
| Yes | 29 (58.0) | 16 (33.3) | |
Cell culture
The EOC cell line (SKOV-3) was purchased from the
Shanghai Institute of Cell Biology (Shanghai, China). All cell
lines were cultured in RPMI-1640 (Gibco-BRL) medium supplemented
with 10% fetal bovine serum (FBS), 100 U/ml penicillin, and 100
μg/ml streptomycin in humidified air at 37°C with 5%
CO2.
TaqMan real-time quantitative RT-PCR
assay
Real-time qRT-PCR analysis of mature miRNA was
performed using the TaqMan microRNA Assay kit (Applied Biosystems,
Foster City, CA) as previously described (12).
Western blot assay
Total cellular protein extract was isolated from
harvested cells, protein concentration was determined, and western
blot analysis was carried out as previously described (13). The antibodies used were anti-PLK1
and anti-GAPDH (Santa Cruz Biotechnology, Inc., Santa Cruz,
CA).
Plasmid of construction
Two pairs of primers were used to generate PLK1
fragment based on the published PLK1 sequence (GenBank Accession
no. NM_005030). The sequences of primers for the fragment were:
sense 5′-ATGA GTGCTGCAGTGACTGCAGGGAAG-3′ antisense 5′-AGC
TATTAGGAGGCCTTGAGACGG-3′. Reverse transcription was carried out
using the EOC cell RNA as a template. Reverse transcription PCR
products were cloned into the pcDNA3.1 vector (Invitrogen) with the
sequences and orientations confirmed from both ends to generate the
recombinant vector pcDNA/PLK1.
Transfection of oligonucleotides or
plasmids
Molecules of dsRNAs that mimic endogenous
hsa-miR-100 (pre-miR-100) and single-strand miR-100 inhibitor
(anti-miR-100), designed to inhibit endogenous hsa-miR-100, were
purchased from Ambion (Austin, TX, USA). The siRNAs were designed
and synthesized by GenePharma (Shanghai, China). PLK1 siRNA
(siRNA/PLK1) was synthesized as follows: sense
5′-AAGGGCGGCUUUGCCAAGUGC-3′; negative control siRNA (siRNA/control)
were synthesized as follows: sense 5′-UUCUCCGAACGUGUCACGUTT-3′.
siRNAs, miRNAs and plasmid transfections were performed using
Lipofectamine 2000 (Invitrogen). In brief, cells were plated in
6-well plates to 50% confluence. For each well, 5 μl siRNA or miRNA
(20 μM) or 5 μg plasmid was added into 250 μl Opti-MEM medium, 5 μl
of Lipofectamine 2000 into 250 μl Opti-MEM medium, and then mixed
with siRNA or miRNA or plasmid with Lipofectamine 2000. The mixture
was added to the cells and incubated for 6 h before replacing the
medium.
Cell proliferation assay
Cell proliferation was determined by the MTT assay.
In brief, the cells were transfected with miRNA mimics or
inhibitors after 48 h. The MTT solution in PBS was added to attain
a final concentration of 0.5 mg/ml, and incubation was continued
for 4 h. Finally, an equal volume of a lysis buffer containing 50%
dimethylformamide and 20% sodium dodecyl sulfate (pH 4.8) was
added. The mixtures were kept overnight and then the amount of MTT
formazan present was quantified by determining its absorbance at
490 nm using an ELISA plate reader (Wallac).
Luciferase reporter assays
The 3′-UTR of human PLK1 (GenBank Accession no.
NM_005030) was amplified from human genomic DNA and individually
cloned into the pGL3-Basic vector (Ambion) by directional cloning.
Seed regions were mutated to remove all complementarity to
nucleotides 2–8 of miR-100 by using the QuickChangeXL mutagenesis
kit (Stratagene, La Jolla, CA). HEK-293 cells were co-transfected
with 0.4 mg of firefly luciferase reporter vector and 0.02 mg of
the control vector containing Renilla luciferase, pRL-SV40
(Promega), using Lipofectamine 2000 (Invitrogen) in 24-well plates.
Each transfection was carried out in four wells. For each well, 50
nM of pre-miR-100 (anti-miR-100) or a negative control pre-miR-NC
(anti-miR-100) was co-transfected with the reporter constructs.
Luciferase assays were performed 24 h after transfection using the
Dual Luciferase Reporter Assay system (Promega). Firefly luciferase
activity was normalized to Renilla luciferase activity.
Statistical analysis
Statistical analysis was performed using SPSS 17.0.
For continuous variables, data are expressed as mean ± standard
deviation (SD). The relationship between miR-100 expression and
clinicopathological factors was analyzed using the Student’s
t-test. Survival probabilities were calculated by the product limit
method of Kaplan-Meier. Differences between the groups were tested
using the log-rank test. The results were analyzed for the endpoint
of 5 years overall survival (OS). Survival times of patients still
alive were censored with the last follow-up date. The Cox
proportional hazards regression model was used to assess the
independence of different prognostic factors. It was considered
significant when the P-value was <0.05.
Results
The expression of miR-100 is
significantly downregulated in human EOC tissues
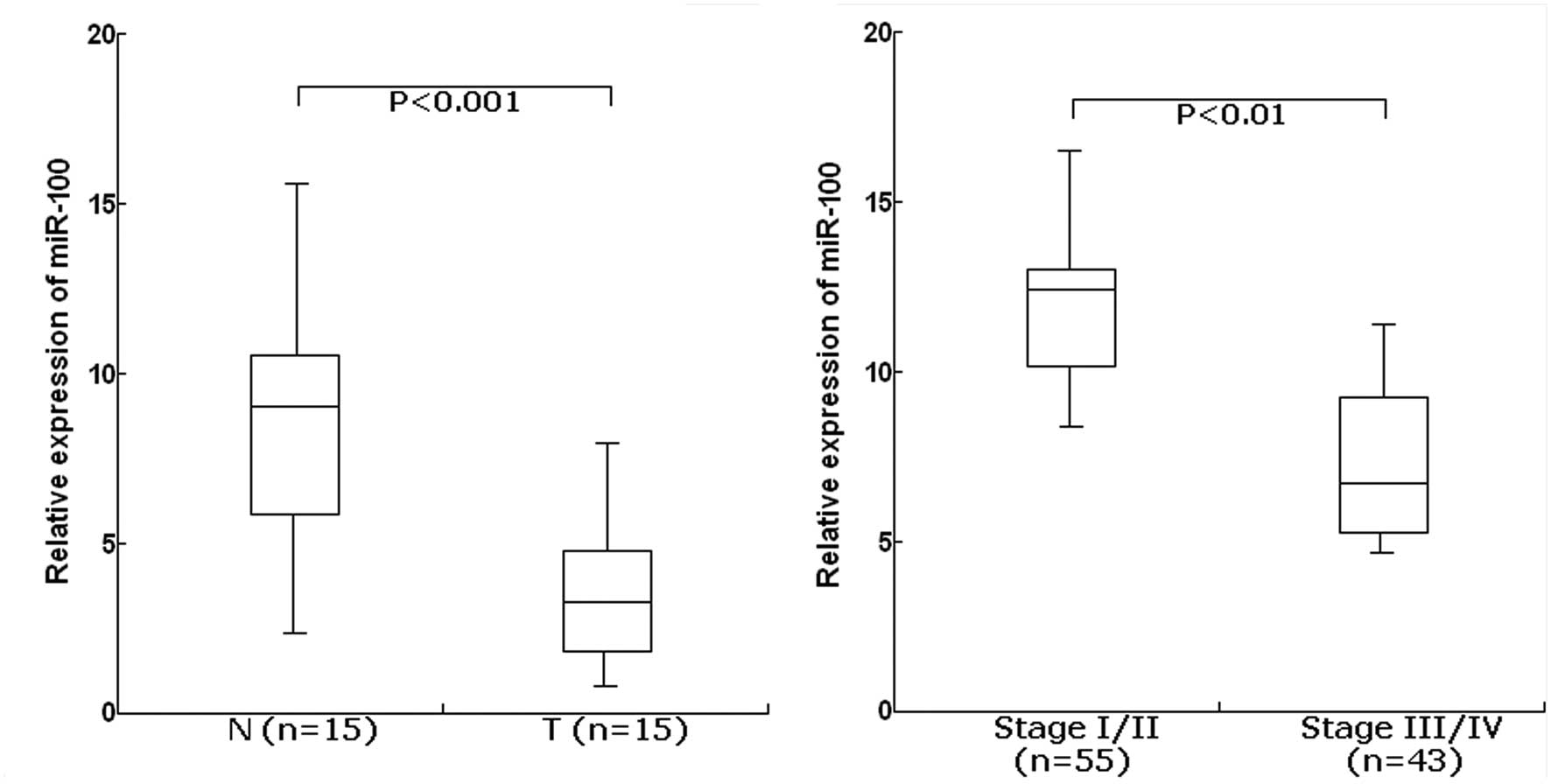
The TaqMan real-time RT-PCR assay was performed to
determine the expression of miR-100 in 15 cases of EOC and their
matched normal tissues. For the EOC tissues, the mean level of
miR-100 expression was 3.8 (range, 1.6–8.5). For the matched normal
tissues, the mean level of miR-100 expression was 8.4 (range,
2.7–16.4). The level of miR-100 was significantly lower in EOC
tissues than in adjacent normal tissues (P<0.001; Fig. 1A). Furthermore, the expression level
of miR-100 was significantly lower in EOC patients with advanced
clinical stage (III/IV) compared with those with clinical stage
(I/II) (P<0.01; Fig. 1B). From
these data, it was concluded that downregulation of miR-155 may
play a critical role in ovarian tumorigenesis.
Association of miR-100 expression with
clinicopathological characteristics in EOC
Next, the clinicopathological significance of
miR-100 expression in human EOC was analyzed. Patients with
relative miR-100 expression levels in EOC tissues less than the
median value of 0.14 formed the low expression group (n=50), while
patients with relative miR-100 expression levels in EOC tissues
≥0.14 formed the high expression group (n=48). As shown in Table I, the low level of miR-100
expression was closely correlated with advanced FIGO stage, higher
serum CA125 expression level and lymph node involvement (P=0.001,
0.001 and 0.014, respectively). However, there was no significant
correlation between miR-100 expression and other
clinicopathological variables of EOC patients including age,
histological type, histological grade, residual tumor diameter and
ascites (P=0.155, 0.486, 0.849, 0.366 and 0.279, respectively).
Interestingly, we also found that the incidence of lymph node
metastasis in EOC patients with low miR-100 level (58.0%) was
significantly higher than that in patients with high miR-100 levels
(33.3%).
Association of miR-100 expression with
prognosis of EOC patients
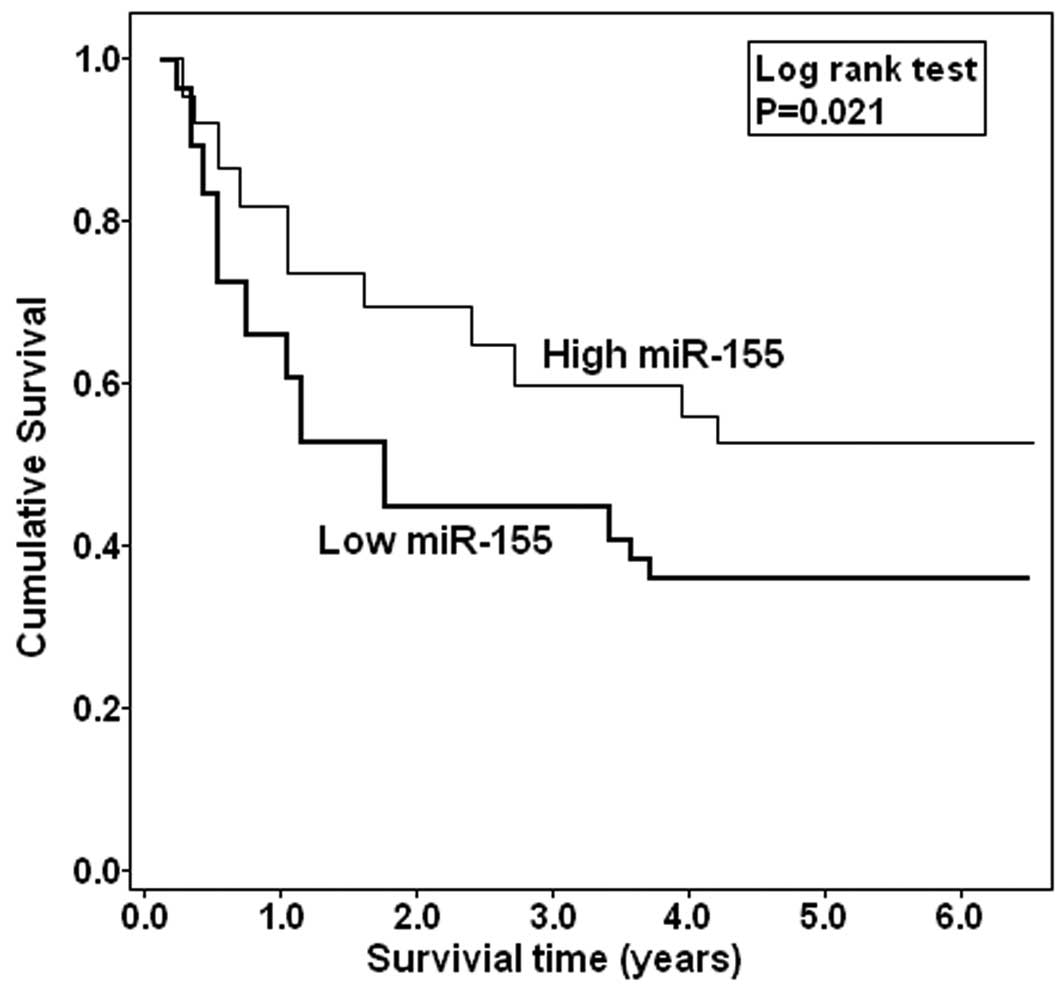
Whether miR-100 expression could affect prognosis of
patients was statistically analyzed. The Kaplan-Meier curve is
shown in Fig. 2. Those EOC patients
with low miR-100 expression were more likely to have a shorter
overall survival (P=0.021), when compared to patients with high
miR-100 expression. These data are further supported by the Cox
regression analysis presented in Table
II. By univariate analysis, the status of miR-100 expression
was correlated with poor survival of EOC patients (P=0.038).
Finally, a multivariate analysis with the Cox proportional hazards
showed that the status of miR-100 expression, along with FIGO stage
and lymph node involvement, was an independent predictor of overall
survival in EOC (RR, 2.12; 95%CI, 1.88–3.41; P=0.008).
 | Table IIUnivariate and multivariate analysis
of prognostic variables by Cox regression analysis. |
Table II
Univariate and multivariate analysis
of prognostic variables by Cox regression analysis.
| Univariate
analysis | Multivariate
analysis |
|---|
|
|
|
|---|
| Clinicopathological
variables | RR (95% CI) | P-value | RR (95% CI) | P-value |
|---|
| Age (years)
(≥50/<50) | 1.41
(0.69–1.72) | 0.215 | 1.56
(0.91–2.45) | 0.098 |
| Histological type
(Serous/non-serious) | 0.94
(0.81–1.08) | 0.156 | 2.56
(0.89–3.12) | 0.227 |
| Histological grade
(G1/G2+G3) | 2.43
(1.68–2.94) | 0.008 | 2.13
(0.75–2.70) | 0.089 |
| FIGO satge
(III+IV/I+II) | 3.08
(2.23–4.18) | 0.016 | 1.69
(1.23–2.55) | 0.005 |
| Residual tumor (cm)
(≥1.0/<1.0) | 1.48
(0.77–1.82) | 0.105 | 1.65
(0.87–1.93) | 0.274 |
| Ascites
(Yes/no) | 0.85
(0.71–1.16) | 0.476 | 1.87
(0.63–2.63) | 0.078 |
| Serum CA125
(≥5.0×105/<5.0×105 U/l) | 1.37
(0.82–1.79) | 0.123 | 2.06
(0.90–3.12) | 0.244 |
| Lymph node
involvement (Yes/no) | 2.12
(1.59–2.34) | 0.023 | 3.18
(2.18–4.03) | 0.011 |
| Expression of
miR-100 (Low/high) | 1.65
(1.16–2.73) | 0.038 | 2.12
(1.88–3.41) | 0.008 |
Effect of miR-100 expression on in vitro
proliferation of EOC cells
In order to study the functional role of miR-100 in
ovarian tumorigenesis, SKOV-3 cells were transfected with
pre-miR-100, anti-miR-100 or scrambled precursor
(pre-miR-NC)/antisense oligonucleotide control (anti-miR-NC).
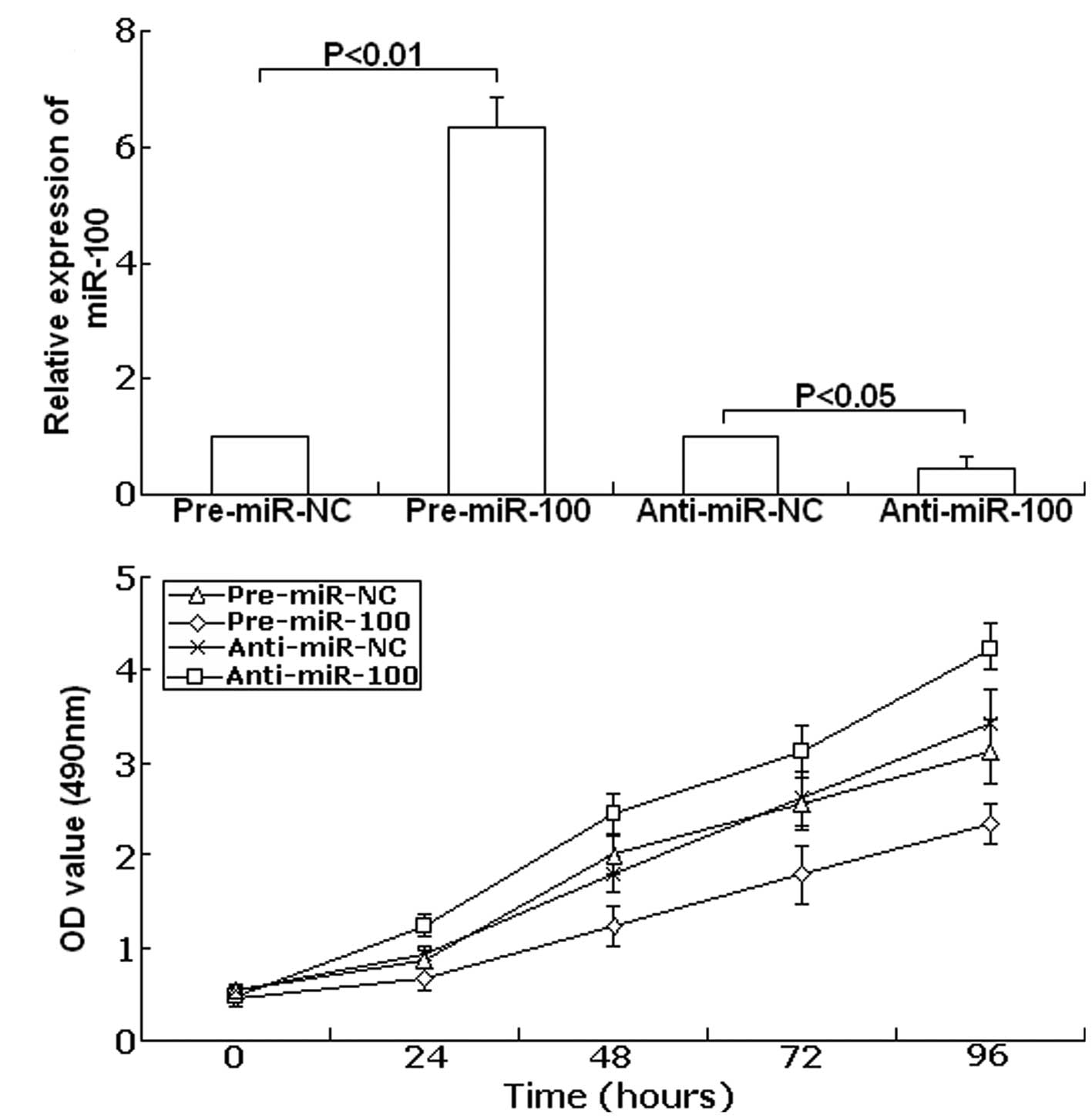
Forty-eight hours after transfection, TaqMan real-time RT-PCR assay
was performed to examine the expression of miR-100. As shown in
Fig. 3A, compared with precursor
control, the expression of miR-100 showed up-regulation to
6.34-fold in the transfected group with pre-miR-100 (P<0.01).
Meanwhile, compared with the antisense control, the expression of
miR-100 was downregulated to 45.6% (P<0.05). Next, the cell
viability was determined using the MTT assay. As shown in Fig. 3B, in SKOV-3 cells transfected with
anti-miR-100 cell viability significantly increased compared with
the anti-miR-NC-transfected cells. In contrast, upregulation of
miR-100 led to decreased cell viability compared with the
pre-miR-NC-transfected cells. These data show that cell viability
could be inhibited by upregulation of miR-100, and could be
promoted by downregulation of miR-100.
Polo-like kinase 1 (PLK1) is a target of
miR-100
In a previous study, PLK1 has been reported to be a
miR-100 target in human nasopharyngeal cancer. However, whether
PLK1 is a direct miR-100 target in EOC cells is unknown. To confirm
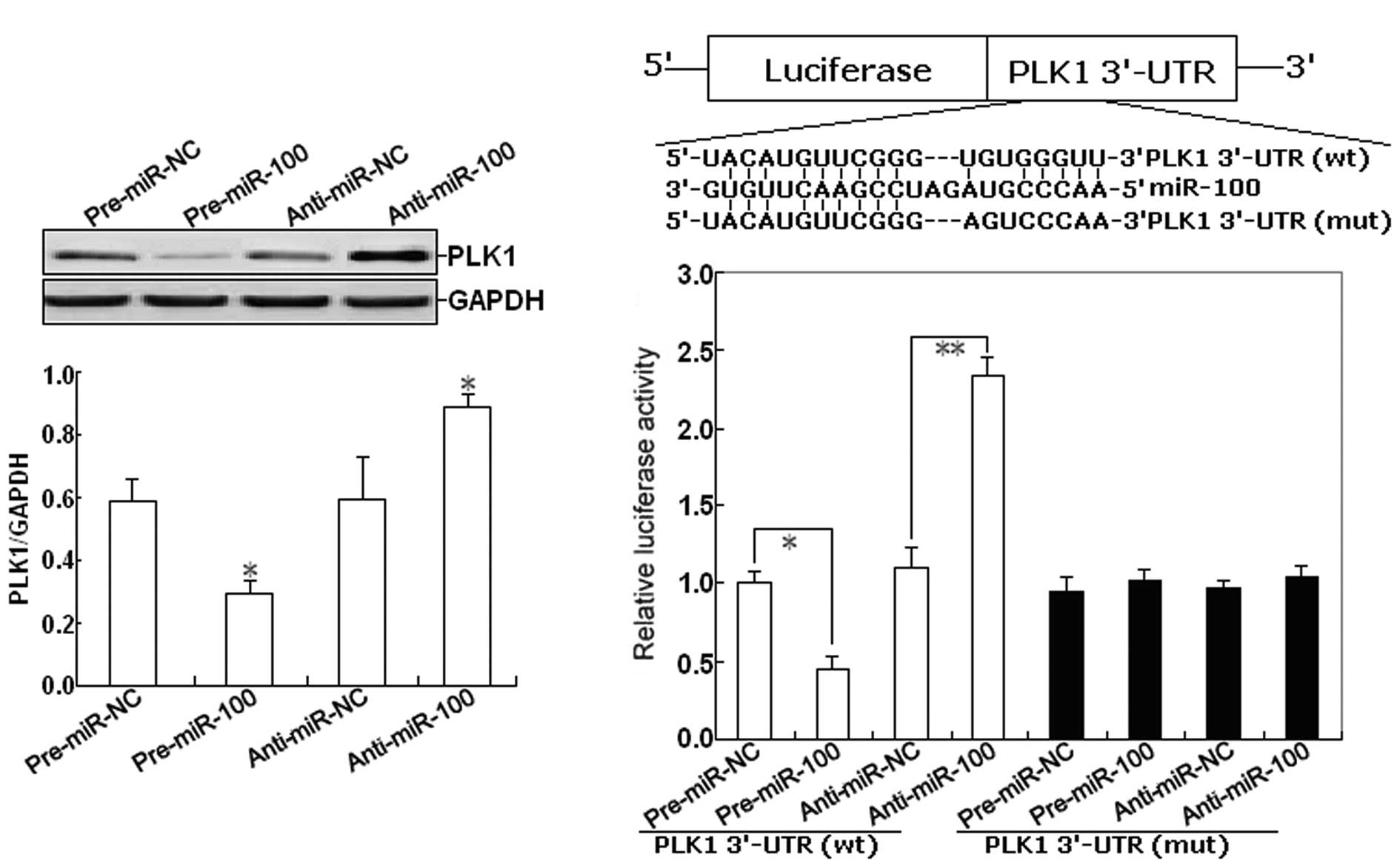
this, we firstly determined the effect of miR-100 expression on the
expression of PLK1 protein in EOC cells. As shown in Fig. 4A, western blot assay indicated that
pre-miR-100 could induce the decreased expression of PLK1 protein,
but anti-miR-100 could induce the increased expression of PLK1
protein in EOC cells (P<0.05). To further confirm the target
specificity between miR-100 and its potential target, PLK1, we
carried out a luciferase reporter assay with a vector containing
the putative PLK1 3′-untranslated region (UTR) target site
downstream of the luciferase reporter gene. The base pairing
between miR-100 and the wild-type (wt) or mutant (mut) target site
in the 3′-UTR of PLK1 mRNA is shown in Fig. 4B. Next, SKOV-3 cells were
co-transfected with PLK1-3′-UTR luciferase reporter
(wt-PLK1-3′-UTR) and pre-miR-100 or anti-miR-100. Transfections
with control vector were performed in parallel. As shown in
Fig. 4C, pre-miR-100 or
anti-miR-100 significantly decreased or respectively increased the
activity of the Luc-PLK1-3′-UTR reporter (P<0.01). However,
transfection of the mut-PLK1-3′-UTR reporter did not affect
reporter activity (P>0.05).
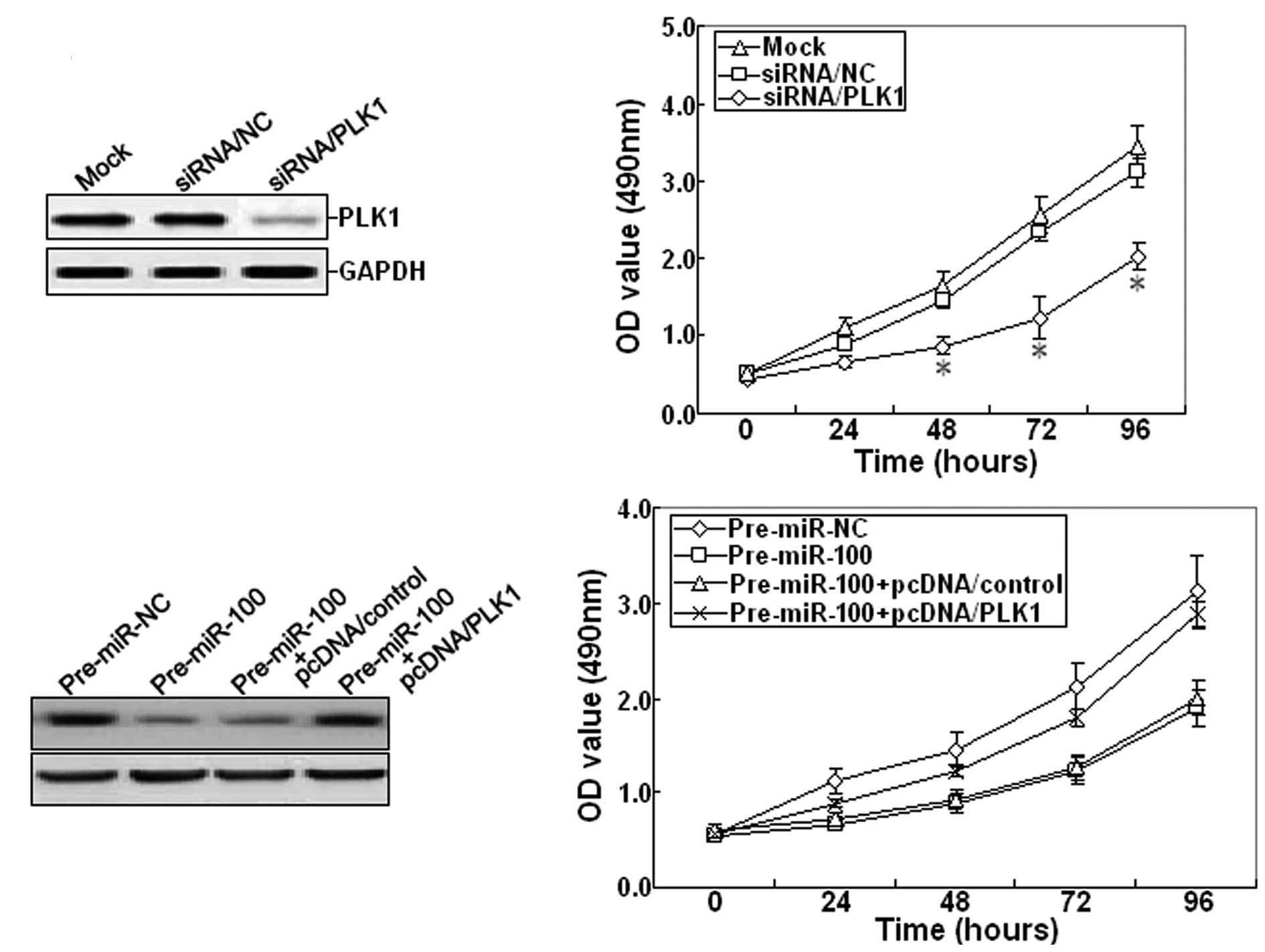
Furthermore, 48 h after siRNA/PLK1 was transfected
into SKOV-3 cells, RT-PCR and western blot assays showed that
siRNA/PLK1 could significantly inhibit the expression of PLK1 mRNA
and protein (Fig. 5A). The cell
viability was also determined. The growth of siRNA/PLK1-transfected
SKOV-3 cells was inhibited compared with mock or
siRNA/control-transfected cells (Fig.
5B). Next, pcDNA/PLK1 vector and pre-miR-100 were
co-transfected into the SKOV-3 cell line. Western blot analysis
indicated that pcDNA/PLK1 could reverse the decreased expression of
PLK1 protein induced by pre-miR-100 (Fig. 5C). Meanwhile, overexpression of PLK1
could also reverse the inhibitory growth induced by pre-miR-100
(Fig. 5D). Therefore, it was
concluded that miR-100 may function as a tumor suppressor in EOC by
targeting PLK1.
Discussion
Recently, much research effort has been intensely
focused on the roles of dysregulated miRNA expression in various
human cancer types including human EOC (14,15).
Many studies have shown that abberant miRNAs are asscociated with
proliferation, apoptosis, metastasis, and chemoresistance of tumor
cells (16–18). Thus, identification of the important
miRNA expression signatures in EOC development and progression will
be helpful to find potential diagnostic and prognostic markers for
EOC diagnosis and treatment.
Up to date, there have been many published research
studies about the correlation of miRNAs with EOC. Lu et al
reported that hypermethylation of let-7a-3 in epithelial ovarian
cancer was associated with low insulin-like growth factor-II
expression and favorable prognosis (19). Additionally, the beneficial impact
of the addition of paclitaxel on EOC survival has been
significantly linked to let-7a levels, and it has been shown that
miRNAs such as let-7a may be useful markers for the selection of
chemotherapeutic agents in EOC management (20). Lou et al reported that
microRNA-21 could promote the cell proliferation, invasion and
migration abilities in ovarian epithelial carcinomas through
inhibiting the expression of PTEN protein (21). In other studies, several other
microRNAs have been reported to be associated with chemotherapy
resistance, such as miR-214, miR-130a, miR-27a and miR-451
(22–24). Although miR-100 was reported to be
downregulated in human EOC by other research groups, the
clinicopathological or prognostic significance of miR-100 in EOC is
still unknown. In nasopharyngeal cancer, underexpressed miR-100 was
found to lead to PLK1 overexpression, which in turn contributes to
NPC progression (25). However, in
prostate cancer, it was reported that a high level of miR-100 was
related to biochemical recurrence of localized prostate cancer in
patients treated with radical prostatectomy (26). Recently, Zheng and his group
reported that miR-100 could regulate cell differentiation and
survival by targeting RBSP3, a phosphatase-like tumor suppressor in
acute myeloid leukemia (27). From
the above studies, it may be concluded that miRNA oncogenes and
tumor suppressors show different patterns in different tumor
types.
In the present study, we firstly showed that the
mean level of miR-100 expression in EOC tissues was significantly
higher than that in the matched normal tissues. Also, the
expression level of miR-100 was significantly lower in EOC patients
with advanced clinical stage (III/IV) compared with those with
clinical stage I/II. Meanwhile, we showed that low miR-100
expression was closely correlated with advanced FIGO stage, high
serum CA125 level and lymph node involvement. Furthermore, patients
with low miR-100 expression showed poorer survival than those with
high miR-100 expression. A multivariate analysis with the Cox
proportional hazards showed that the status of miR-100 expression
was an independent predictor of overall survival in EOC. Then, we
analyzed the effect of miR-100 expression on the growth of EOC
cells and its possible mechanisms. Overexpression of miR-100 could
induce growth suppression in human EOC cells, while downregulation
of miR-100 could promote growth of EOC cells. PLK1 is a
serine/threonine kinase that functions to regulate many stages of
mitosis (28). The overexpression
of PLK1 has been found in many human cancers, including ovarian
carcinoma (29). It has been
reported that the overexpression of PLK1 could affect growth,
apoptosis, metastasis and chemo-or radioresistance in human cancers
(30,31). Weichert et al showed that
PLK1 is a novel independent prognostic marker in ovarian carcinomas
(29). Additionally, silencing of
Chk1 and PLK1 could enhance radiation-or cisplatin-induced
cytotoxicity in human ovarian cancer cells (32). In our study, we show that
overexpression of miR-100 could inhibit the expression of PLK1
protein in EOC cells. So, it can be demonstrated that miR-100
negatively regulated PLK1 at the post-translational level, acting
as a tumor suppressor in the EOC. In vitro luciferase assay
further suggested that PLK1 is the target gene of miR-100.
Importantly, siRNA-mediated downgulation of PLK1 could recapitulate
the tumor suppressor function of miR-100, and overexpression of
PLK1 could partly rescue the reduced cellular proliferation
observed upon miR-100 upregulation in SKOV-3 cells, demonstrating
that PLK1 is an important functional target of miR-100 in this
model.
In conclusion, this study suggests that miR-100 is
downregulated in human EOC and low miR-100 expression may be a poor
prognostic factor. Also, miR-100 can significantly inhibit growth
of EOC cells by targeting PLK1. Thus, this miR-100/PLK1 signaling
pathway may provide therapeutic targets for human EOCs. This study
has several limitations. Firstly, the number of tissue samples is
small, and further investigation of a larger case population is
needed to confirm the prognostic significance of miR-100 expression
in EOC. Future investigations using patient samples are needed to
further support the function of miR-100 in EOC.
Acknowledgements
The authors are grateful to Mr. Fengyu Zhang and Ms.
Lan Zhao for their excellent technical assistance.
References
|
1
|
Jemal A, Siegel R, Ward E, Hao Y, Xu J and
Thun MJ: Cancer statistics, 2009. CA Cancer J Clin. 59:225–249.
2009. View Article : Google Scholar
|
|
2
|
Armstrong DK: Relapsed ovarian cancer:
challenges and management strategies for a chronic disease.
Oncologist. 7(Suppl 5): 20–28. 2002. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Bartel DP: MicroRNAs: genomics,
biogenesis, mechanism, and function. Cell. 116:281–297. 2004.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Huang Y, Shen XJ, Zou Q and Zhao QL:
Biological functions of microRNAs. Bioorg Khim. 36:747–752.
2010.PubMed/NCBI
|
|
5
|
Huang Y, Shen XJ, Zou Q, Wang SP, Tang SM
and Zhang GZ: Biological functions of microRNAs: a review. J
Physiol Biochem. 67:129–139. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Perera RJ and Ray A: MicroRNAs in the
search for understanding human diseases. BioDrugs. 21:97–104. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Calin GA and Croce CM: MicroRNA signatures
in human cancers. Nat Rev Cancer. 6:857–866. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Dahiya N and Morin PJ: MicroRNAs in
ovarian carcinomas. Endocr Relat Cancer. 17:77–89. 2010. View Article : Google Scholar
|
|
9
|
Zhang L, Huang J, Yang N, et al: microRNAs
exhibit high frequency genomic alterations in human cancer. Proc
Natl Acad Sci USA. 103:9136–9141. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Yang H, Kong W, He L, et al: MicroRNA
expression profiling in human ovarian cancer: miR-214 induces cell
survival and cisplatin resistance by targeting PTEN. Cancer Res.
68:425–433. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wyman SK, Parkin RK, Mitchell PS, Fritz
BR, et al: Repertoire of microRNAs in epithelial ovarian cancer as
determined by next generation sequencing of small RNA cDNA
libraries. PLoS One. 4:e53112009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Wang R, Wang ZX, Yang JS, Pan X, De W and
Chen LB: MicroRNA-451 functions as a tumor suppressor in human
non-small cell lung cancer by targeting ras-related protein 14
(RAB14). Oncogene. 30:2644–2658. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Rui W, Bing F, Hai-Zhu S, Wei D and
Long-Bang C: Identification of microRNA profiles in
docetaxel-resistant human non-small cell lung carcinoma cells
(SPC-A1). J Cell Mol Med. 14:206–214. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Bovicelli A, D’Andrilli G and Giordano A:
New players in ovarian cancer. J Cell Physiol. 226:2500–2504. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Li SD, Zhang JR, Wang YQ and Wan XP: The
role of microRNAs in ovarian cancer initiation and progression. J
Cell Mol Med. 14:2240–2249. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Lima RT, Busacca S, Almeida GM, Gaudino G,
Fennell DA and Vasconcelos MH: MicroRNA regulation of core
apoptosis pathways in cancer. Eur J Cancer. 47:163–174. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Dykxhoorn DM: MicroRNAs and metastasis:
little RNAs go a long way. Cancer Res. 70:6401–6406. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
van Jaarsveld MT, Helleman J, Berns EM and
Wiemer EA: MicroRNAs in ovarian cancer biology and therapy
resistance. Int J Biochem Cell Biol. 42:1282–1290. 2010.PubMed/NCBI
|
|
19
|
Lu L, Katsaros D, de la Longrais IA,
Sochirca O and Yu H: Hypermethylation of let-7a-3 in epithelial
ovarian cancer is associated with low insulin-like growth factor-II
expression and favorable prognosis. Cancer Res. 67:10117–10122.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Lu L, Schwartz P, Scarampi L, et al:
MicroRNA let-7a: a potential marker for selection of paclitaxel in
ovarian cancer management. Gynecol Oncol. 122:366–371. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Lou Y, Yang X, Wang F, Cui Z and Huang Y:
MicroRNA-21 promotes the cell proliferation, invasion and migration
abilities in ovarian epithelial carcinomas through inhibiting the
expression of PTEN protein. Int J Mol Med. 26:819–827.
2010.PubMed/NCBI
|
|
22
|
Yang H, Kong W, He L, et al: MicroRNA
expression profiling in human ovarian cancer: miR-214 induces cell
survival and cisplatin resistance by targeting PTEN. Cancer Res.
68:425–433. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Sorrentino A, Liu CG, Addario A, Peschle
C, Scambia G and Ferlini C: Role of microRNAs in drug-resistant
ovarian cancer cells. Gynecol Oncol. 2111:478–486. 2008. View Article : Google Scholar
|
|
24
|
Li Z, Hu S, Wang J, et al: MiR-27a
modulates MDR1/P-glycoprotein expression by targeting HIPK2 in
human ovarian cancer cells. Gynecol Oncol. 119:125–130. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Shi W, Alajez NM, Bastianutto C, et al:
Significance of Plk1 regulation by miR-100 in human nasopharyngeal
cancer. Int J Cancer. 126:2036–2048. 2010.PubMed/NCBI
|
|
26
|
Leite KR, Tomiyama A, Reis ST, et al:
MicroRNA-100 expression is independently related to biochemical
recurrence of prostate cancer. J Urol. 185:1118–1122. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Zheng YS, Zhang H, Zhang XJ, et al:
MiR-100 regulates cell differentiation and survival by targeting
RBSP3, a phosphatase-like tumor suppressor in acute myeloid
leukemia. Oncogene. June 6–2011.(Epub ahead of print).
|
|
28
|
Takaki T, Trenz K, Costanzo V and
Petronczki M: Polo-like kinase 1 reaches beyond
mitosis-cytokinesis, DNA damage response, and development. Curr
Opin Cell Biol. 20:650–660. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Weichert W, Denkert C, Schmidt M, et al:
Polo-like kinase isoform expression is a prognostic factor in
ovarian carcinoma. Br J Cancer. 90:815–821. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Degenhardt Y and Lampkin T: Targeting
Polo-like kinase in cancer therapy. Clin Cancer Res. 16:384–389.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Strebhardt K and Ullrich A: Targeting
polo-like kinase 1 for cancer therapy. Nat Rev Cancer. 6:321–330.
2006. View
Article : Google Scholar : PubMed/NCBI
|
|
32
|
Gao Q, Huang X, Tang D, et al: Influence
of chk1 and plk1 silencing on radiation- or cisplatin-induced
cytotoxicity in human malignant cells. Apoptosis. 11:1789–1800.
2006. View Article : Google Scholar : PubMed/NCBI
|