Introduction
Cervical cancer is the fourth leading cause of
cancer-related mortality in women worldwide (1). Widespread employment of screening
programs has reduced the incidence and mortality associated with
this cancer. However, it remains a major public health issue,
particularly concerning advanced cases (1). Accordingly, recent research has
focused on identifying tumor-specific markers capable of predicting
the biological behavior of cervical cancers with respect to cell
motility and invasion (2).
Non-coding RNAs (ncRNAs) are potential key factors
influencing gene regulation and cancer cell phenotypes (3,4).
Recently, more than 3,000 human long intervening coding RNAs and
most long ncRNAs (lncRNAs) have been found to correlate the
activity associated with DNA-binding proteins, including
chromatin-modifying complexes (5)
that epigenetically modify the expression of various genes
(6). Additionally, transcription of
lncRNAs modulates gene expression in response to external oncogenic
stimuli or DNA damage (7).
Human lncRNA, steroid receptor RNA activator (SRA),
was shown to co-activate several human sex hormone receptors and
was found to be highly associated with several types of cancers
including ovarian and breast cancer (8–11). In
addition, lncRNA SRA functionally interacts with several proteins,
including nuclear receptors, such as the retinoic acid receptor, as
well as proteins involved in myogenic differentiation (12,13).
Although lncRNA SRA is also correlated with the progression of
other malignancies, its role in cervical cancer has not yet been
fully elucidated.
The present study investigated the expression and
molecular function of SRA in cervical cancer cell lines and
elucidated the role of SRA in the metastatic progression of
cervical cancer.
Materials and methods
Cell lines
Squamous cell cervical carcinoma (SiHa) cells and
epidermoid cell cervical carcinoma (CaSki; originally derived from
a small-bowel mesentery metastasis) cells and human cervical cancer
cell lines [American Type Culture Collection (ATCC) Manassas, VA,
USA], were maintained in Dulbecco's modified Eagle's medium
(Gibco-BRL, Gaithersburg, MD, USA) containing 10% (v/v) fetal
bovine serum and penicillin/streptomycin. Cells were maintained at
37°C in an atmosphere of 95% air and 5% CO2. Only cells
that had been passaged <20 times were utilized in the
experiments.
Quantitative reverse-transcription
polymerase chain reaction (qRT-PCR)
We extracted total RNA using TRIzol reagent
(Invitrogen, Carlsbad, CA, USA) according to the manufacturers
instructions. Total RNA (2 µg) was reverse transcribed into
first-strand cDNA using a reverse transcription reagent kit
(Invitrogen). The cDNA template was amplified by qRT-PCR using the
SYBR-Green Real-Time PCR kit (Toyobo Co., Ltd., Osaka, Japan) on an
ABI StepOnePlus Real-Time PCR system (Applied Biosystems, Foster
City, CA, USA). We used U6 as the internal standard for all
quantifications. We analyzed the relative gene expression using the
2−ΔΔCt method, and the results are expressed as the
extent of change relative to control values. All qRT-PCR
experiments were repeated at least three times. Primers used for
the PCR reactions are listed in Table
I.
 | Table I.Primer sequences used in the present
study. |
Table I.
Primer sequences used in the present
study.
|
| Primer
sequence |
|
|---|
|
|
|
|
|---|
| Gene | Forward (5–3) | Reverse (5–3) | Product size
(bp) |
|---|
| SRA |
CTCCCTTCTTACCACCACCA |
TGCAGATACACAGGGAGCAG | 217 |
| MMP-9 |
CGCTACCACCTCGAACTTTG |
GCCATTCACGTCGTCCTTAT | 196 |
| MMP-2 |
GGATGATGCCTTTGCTCG |
ATAGGATGTGCCCTGGAA | 487 |
| VEGF |
TTGCTGCTCTACCTCCAC |
AAATGCTTTCTCCGCTCT | 419 |
| E-cadherin |
ATTCTGATTCTGCTGCTCTTG |
AGTAGTCATAGTCCTGGTCCT | 421 |
| β-catenin |
TGCAGTTCGCCTTCACTATG |
ACTAGTCGTGGAATGGCACC | 162 |
| N-cadherin |
CCCAAGACAAAGAGACCCAG |
GCCACTGTGCTTACTGAATTG | 140 |
| Vimentin |
TGGATTCACTCCCTCTGGTT |
GGTCATCGTGATGCTGAGAA | 111 |
| Snail |
GAGGCGGTGGCAGACTAG |
GACACATCGGTCAGACCAG | 178 |
| NOTCH1 |
GCCGCCTTTGTGCTTCTGTTC |
CCGGTGGTCTGTCTGGTCGTC | 300 |
| HES1 |
TCAACACGACACCGGATAAA |
TCAGCTGGCTCAGACTTTCA | 111 |
| P300 |
GACCCTCAGCTTTTAGGAATCC |
TGCCGTAGCAACACAGTGTCT | 301 |
Small interfering RNA (siRNA)
transfection
We obtained siRNA for SRA lncRNA and
negative-control siRNA (siNC) from Genolution Pharmaceuticals
(Seoul, Korea). Cells (5×104 cells/well) were seeded
into 6-well plates and transfected with 10 nM siRNA in
phosphate-buffered saline using the G-Fectin kit (Genolution
Pharmaceuticals) according to the manufacturers instructions. The
siRNA-transfected cells were used in the in vitro assays 48
h after transfection. The target sequence for the siSRA was,
5′-CTCCCTTCTTACCACCACCA-3′. Experiments were repeated at least
three times.
Plasmid constructs and generation of
stable cell lines
Full-length human SRA-transcript cDNA was amplified
by PCR and inserted into the pLenti6/V5-D-TOPO vector using the
ViraPower lentiviral expression system (Invitrogen) according to
the manufacturers instructions. We then transfected the plasmid
into 293FT cells for packaging, with the resultant lentivirus used
to infect the desired cell lines. Stably SRA-transfected cells were
selected in medium containing blasticidin (Invitrogen).
Cell proliferation assay
Cell Counting Kit-8 (CCK-8; Dojindo Laboratories,
Kumamoto, Japan) was utilized to evaluate cell proliferation. Cells
(2×103 cells/well) were seeded into 96-well
flat-bottomed plates containing 100 µl of complete medium per well.
The cells were incubated overnight to allow for cell attachment and
recovery, and subsequently transfected with siNC or siSRA for 24,
48, 72 or 96 h. An aliquot of 10 µl of CCK-8 solution was
supplemented into each well, followed by incubation for 2 h.
Absorbance was measured at 450 nm to estimate the number of viable
cells in each well. The assay was performed in triplicate.
Matrigel invasion assay
We performed a Matrigel invasion assay using the BD
Biocoat Matrigel invasion chamber (pore size, 8-µm; 24-wells; BD
Biosciences, Bedford, MA, USA) according to the manufacturers
protocol. Cells (5×104) were seeded in the upper
chamber, which contained serum-free medium, and complete medium was
added to the lower chamber. The Matrigel-invasion chamber was
incubated at 37°C under 5% CO2 for 48 h. Non-invading
cells were removed from the upper chamber using cotton-tipped
swabs. Cells that had invaded through the pores onto the lower side
of the filter were stained (Diff-Quik; Sysmex, Kobe, Japan) and
counted using a hemocytometer. The assay was repeated at least
three times.
Wound-healing migration assay
We evaluated cell migration using a wound-healing
assay. Approximately 5×105 cells were seeded into 6-well
culture plates containing serum-enriched medium and allowed to grow
to 90% confluence in complete medium. The serum-containing medium
was eliminated, and cells were serum-starved for 24 h. When the
cells reached 100% confluence, an artificial homogenous wound was
made by scratching the monolayer using a sterile 200-µl pipette
tip. Following the scratching, cells were washed with serum-free
medium. Images of cells migrating into the wound were captured at
0, 24 and 48 h using a microscope. Three independent experiments
were performed in triplicate.
Western blot analysis
We used radioimmunoprecipitation assay buffer to
extract proteins, and a Pierce BCA protein assay kit (both from
Thermo Fisher Scientific, Waltham, MA, USA) was used to measure
protein concentration. Proteins were boiled with 2X sample buffer,
subsequently resolved on 10% sodium dodecyl sulfate-polyacrylamide
gels, and transferred electrophoretically to polyvinylidene
difluoride membranes (Millipore, Billerica, MA, USA). After
blocking with 5% non-fat dried milk in 1X Tris-buffered saline
containing 0.1% Tween-20 (pH 7.6) at room temperature for 1 h, the
membranes were incubated with primary antibodies at 4°C overnight
under continual agitation. The following primary antibodies were
used: rabbit anti-human vascular endothelial growth factor (VEGF)
(1:500), rabbit anti-human matrix metalloproteinase (MMP)-2 (1:500)
(both from Abcam, Cambridge, UK, USA), rabbit anti-human MMP-9
(1:1,000), rabbit anti-human E-cadherin (1:1,000), rabbit
anti-human β-catenin (1:1,000) (all from Cell Signaling Technology,
Danvers, MA, USA), mouse anti-human vimentin (1:1,000;
Sigma-Aldrich, St. Louis, MO, USA), mouse anti-human Snail
(1:1,000), rabbit anti-human SOX-2 (1:1,000), rabbit anti-human
Nanog (1:1,000), rabbit anti-human Oct4 (1:1,000) (all from Cell
Signaling Technology), and mouse anti-human β-actin (1:5,000;
Sigma-Aldrich). We visualized the proteins using an enhanced
chemiluminescence system (Amersham, Little Chalfont, UK), and band
intensities were quantified using a luminescent image analyzer
(LAS-4000 Mini; Fujifilm, Uppsala, Sweden).
Statistical analysis
IBM SPSS version 23 for Windows (SPSS, Inc.,
Chicago, IL, USA) was used for statistical analysis. The
Kolmogorov-Smirnov test was used to validate standard
normal-distributional assumptions. Statistical significance was
determined using the Fisher's exact-test, Pearson's Chi-square test
and Student's t-test. Mean differences were considered significant
at P<0.05. Results are presented as the means ± standard
deviation.
Results
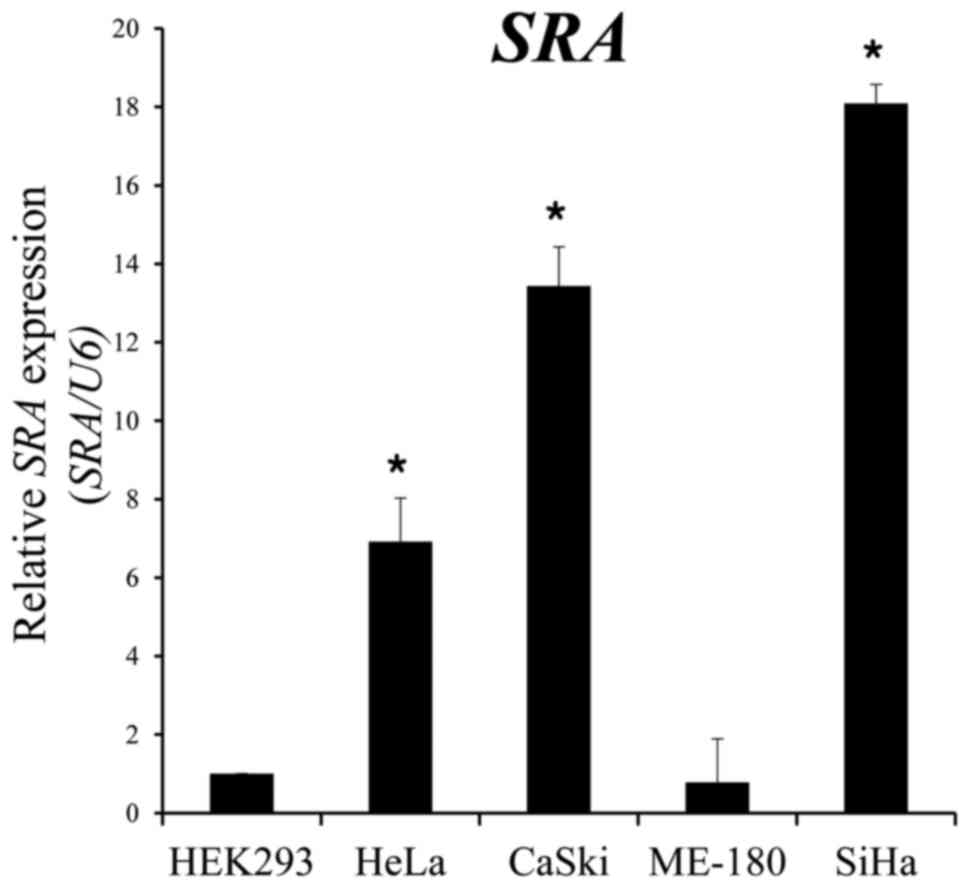
SRA expression is elevated in cervical
cancer cells
We performed qRT-PCR to estimate the expression
levels of SRA in five different cell lines, one of which was
derived from human embryonic kidney cells (HEK293), whereas the
other four were derived from human cervical cancers. We observed
that SRA expression levels were significantly higher in epithelioid
cervical carcinoma (HeLa), CaSki and SiHa cells as compared with
levels observed in epidermoid cervical carcinoma cells (ME-180)
(Fig. 1).
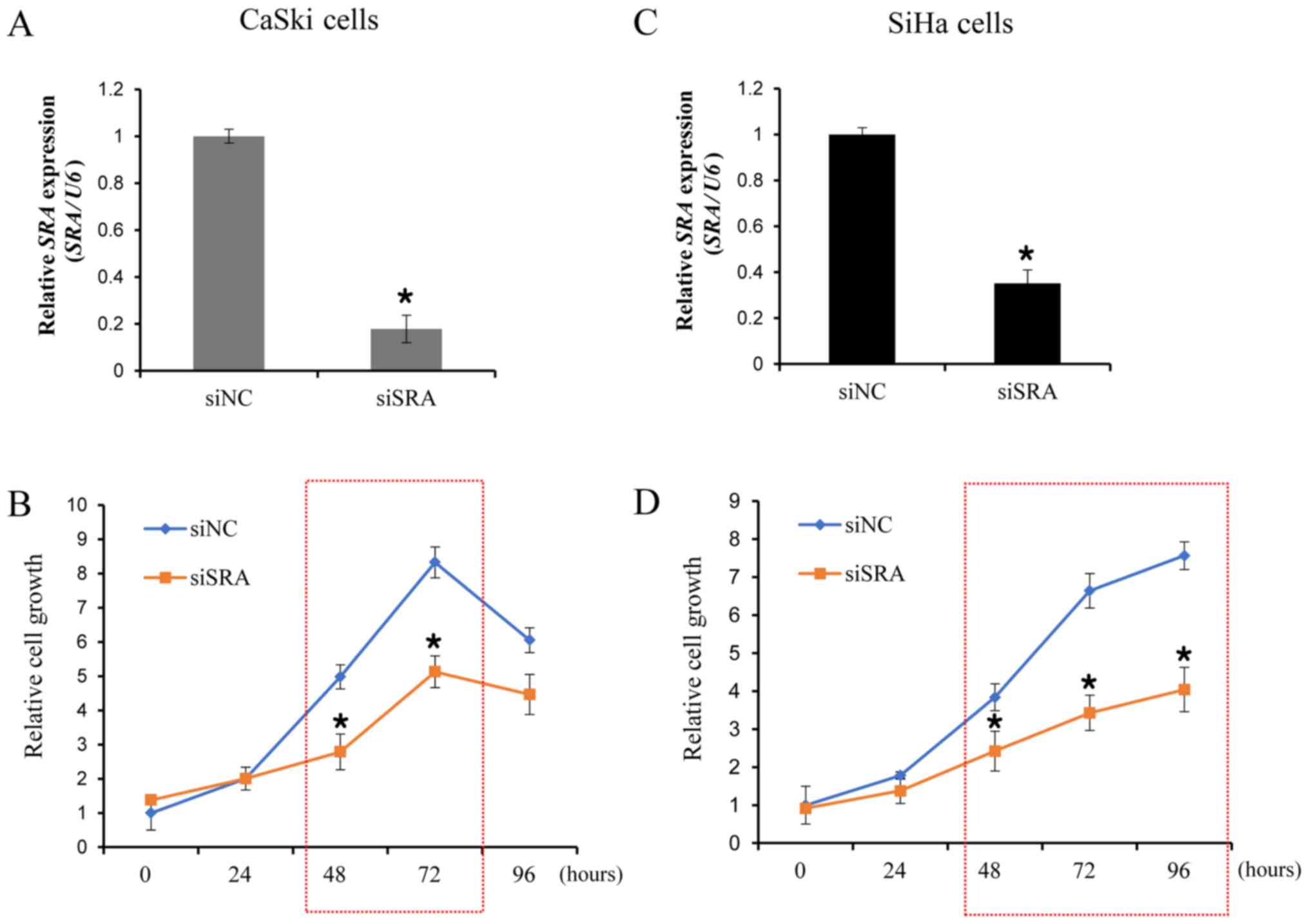
Knockdown of SRA decreases cell
proliferation in cervical cancer cells
To examine the functional role of SRA in cervical
cancer cells, siRNA was utilized to downregulate SRA expression in
the CaSki and SiHa cells, which showed significantly higher levels
of SRA expression. We evaluated the knockdown efficiency of siSRAs,
and verified that siSRA transfection was more efficient in
silencing SRA as compared with siNC (Fig. 2A and C). We then performed a CCK-8
assay to determine the effect of SRA knockdown on cell
proliferation. Our results showed that siRNA-mediated knockdown of
SRA in CaSki and SiHa cells substantially decreased cell
proliferation (Fig. 2B and D).
These findings indicated that SRA is involved in the proliferation
of cervical cancer cells.
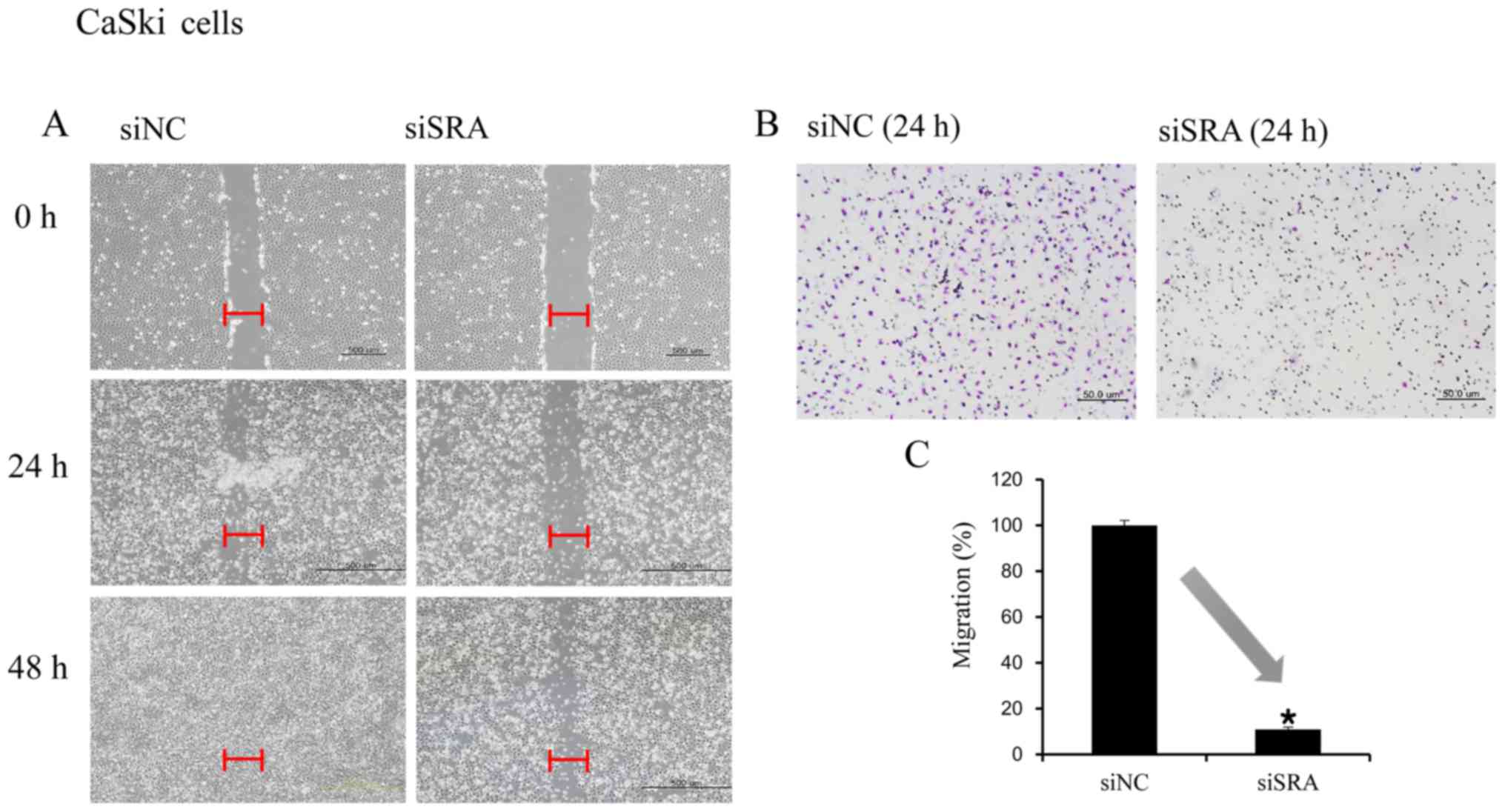
SRA knockdown inhibits cervical cancer
cell migration and invasion
Wound-healing and Matrigel-invasion assays were
performed to determine whether SRA plays a role in cervical cancer
cell migration and invasion. In the wound-healing assay,
siRNA-mediated knockdown of SRA inhibited cell migration in the
CaSki and SiHa cells (Fig. 3A and
D). Moreover, SRA-knockdown inhibited cell invasion in the
CaSki and SiHa cells according to the results of the
Matrigel-invasion assay (Fig. 3B and
E). The extent of inhibition upon SRA knockdown was quantified
by qRT-PCR (Fig. 3C and F).
SRA knockdown inhibits MMP-9, MMP-2
and VEGF expression in cervical cancer cells
To elucidate the molecular mechanisms underlying the
inhibitory effect of SRA knockdown on cell migration and invasion,
we explored the relationship between SRA expression and MMP-2,
MMP-9 and VEGF expression. We observed that siSRA treatment
decreased levels of MMP-2, MMP-9 and VEGF mRNA
in the CaSki cells (Fig. 4A) as
well as MMP-2, MMP-9 and VEGF protein levels (Fig. 4B). These findings indicated that SRA
promoted cervical cancer cell migration and invasion via
upregulation of MMP-9, MMP-2 and VEGF expression.
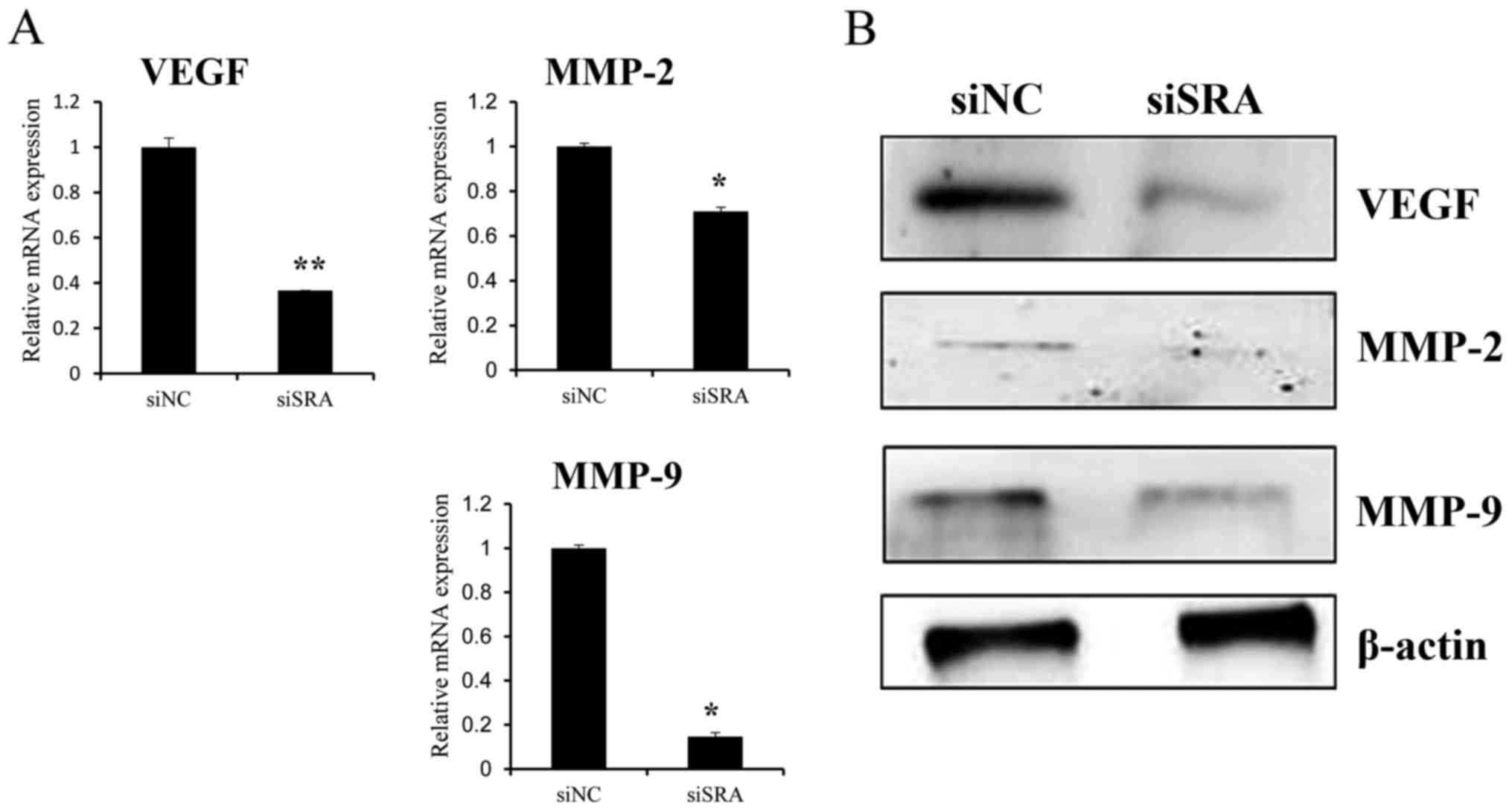 | Figure 4.SRA knockdown decreases MMP-9, MMP-2
and VEGF expression in cervical cancer cells. (A) VEGF,
MMP-2 and MMP-9 mRNA expression levels were analyzed
by qRT-PCR. (B) Protein lysates were obtained from siSRA- and
siNC-transfected CaSki cells 48-h post-transfection. VEGF, MMP-2,
and MMP-9 protein levels were analyzed by western blotting;
*P<0.05 vs. siNC. MMP, matrix metalloproteinase; qRT-PCR,
quantitative reverse transcription polymerase chain reaction; siNC,
negative-control siRNA; siSRA, SRA-specific siRNA; SRA, steroid
receptor RNA activator; VEGF, vascular endothelial growth
factor. |
SRA knockdown downregulates expression
of genes related to epithelial-mesenchymal transition (EMT) and the
NOTCH-signaling pathway in cervical cancer cells
Since EMT is crucial to cell migration and invasion,
we examined whether SRA is required for EMT. EMT was monitored
using qRT-PCR and western blotting following siRNA-mediated
knockdown of SRA in CaSki cells. SRA knockdown resulted in
increased E-cadherin expression and decreased β-catenin and
vimentin expression (Fig. 5A and
B). Additionally, the EMT-mediating transcription factor Snail
and Twist were downregulated in the siSRA-transfected cells
relative to levels observed in the siNC-transfected cells (Fig. 5A and B). These results suggested
that SRA may be involved in the dysregulation of EMT-related genes
in cervical cancer cells, thereby promoting migration and
invasion.
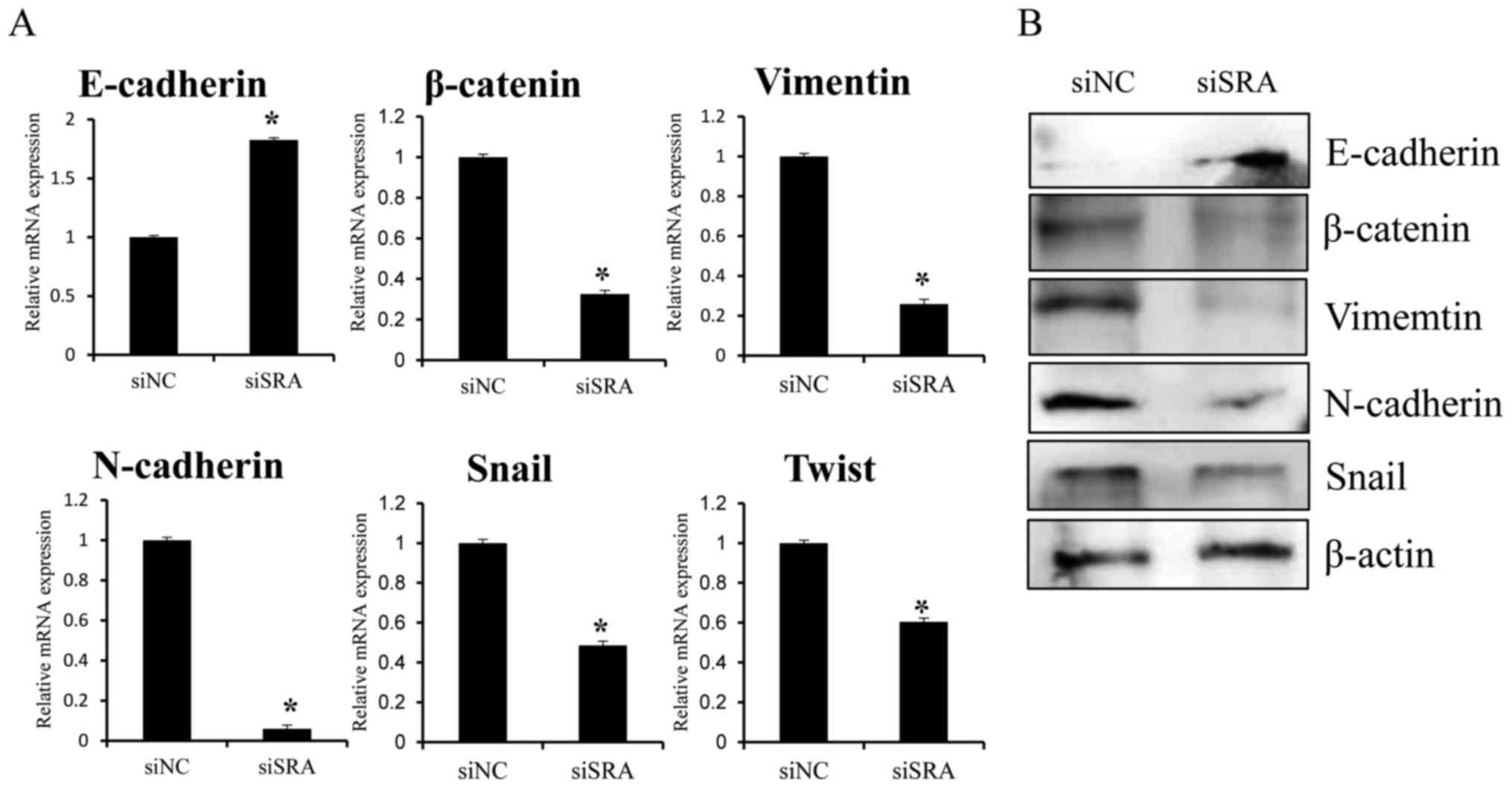 | Figure 5.Effect of SRA knockdown on
EMT-related genes in CaSki cells. CaSki cells were transfected with
siSRA and siNC for 48 h. E-cadherin, β-catenin,
vimentin, N-cadherin, Snail and Twist
mRNA expression and protein levels were analyzed by (A) qRT-PCR and
(B) western blotting, respectively; *P<0.05 vs. siNC. CaSki,
epidermoid cell cervical carcinoma cells; EMT,
endothelial-to-mesenchymal transition; qRT-PCR, quantitative
reverse transcription polymerase chain reaction; siNC,
negative-control siRNA; siSRA, SRA-specific siRNA; SRA, steroid
receptor RNA activator. |
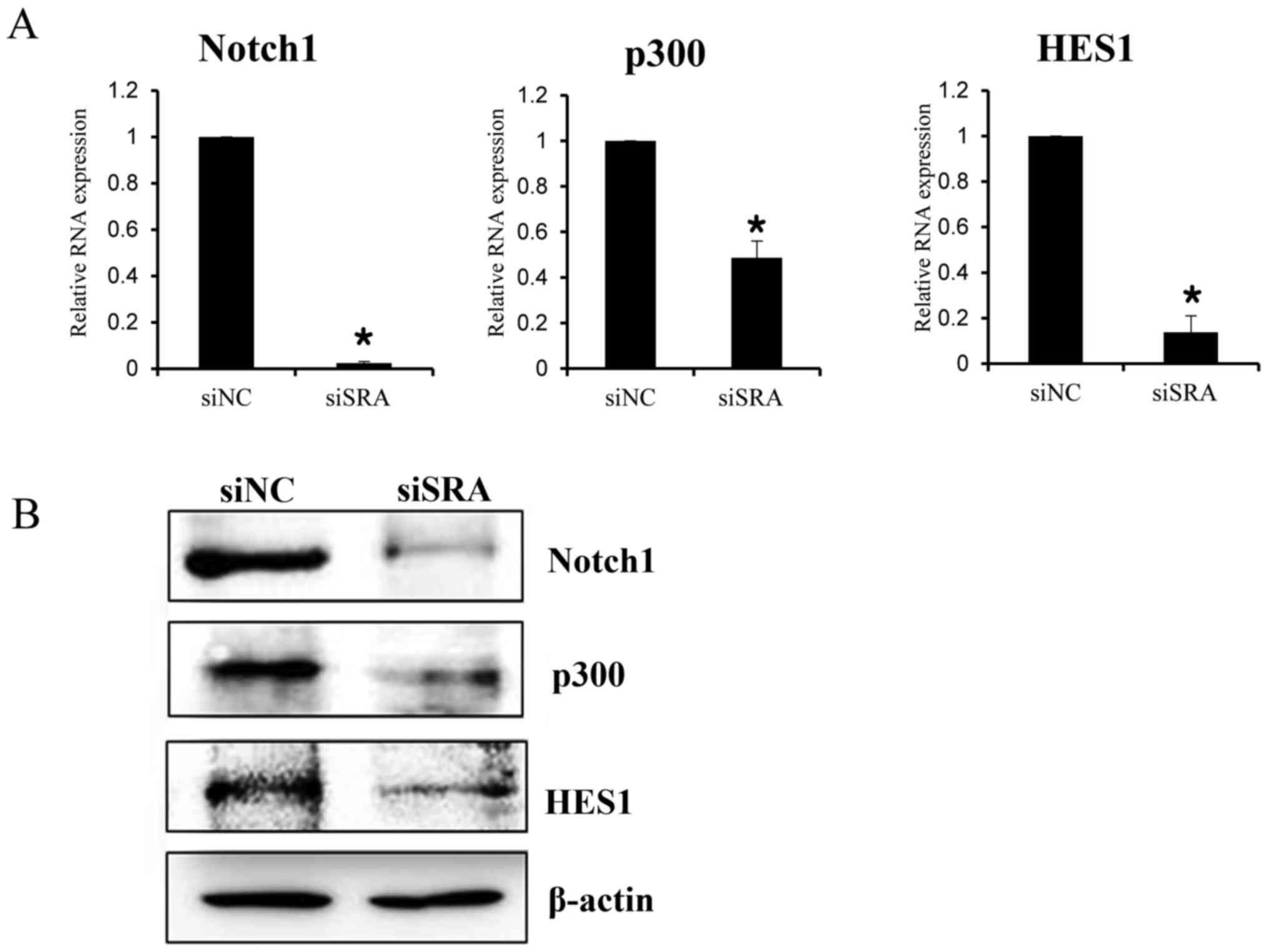
To elucidate the mechanism by which SRA promotes a
malignant phenotype in cervical cancer cells, we assessed the
status of important signaling cascades controlled by NOTCH in the
SRA-knockdown cells. SRA knockdown in the CaSki cells resulted in
downregulation of NOTCH1, HES1 and p300 expression [both mRNA
(Fig. 6A) and protein (Fig. 6B)]. These data indicated that SRA
promoted tumor growth in vitro via the EMT and NOTCH
signaling pathways.
Discussion
lncRNAs are transcripts of >200 nucleotides
lacking protein-coding capability (7). Numerous lncRNAs are capped, spliced
and polyadenylated similar to their protein-coding counterparts and
exhibit tissue-specific expression patterns (14). Furthermore, lncRNAs are essential
for the regulation of chromatin structure, gene expression and
translational control (15). The
mounting list of functionally characterized lncRNAs implies that
these transcripts are critical to various physiological processes
(16). Therefore, these findings
suggest that modified lncRNA expression may affect cancer
development and progression (17).
However, little is known concerning the regulatory roles of lncRNAs
and their relevance to malignant diseases.
Recently, lncRNAs have become the focus of intense
research due to their critical roles in malignant processes,
including tumorigenesis, drug resistance and metastasis (18–20).
The present study explored the molecular function of SRA expression
in cervical cancer cell lines. Our findings showed that
downregulated SRA expression was correlated with decreased cell
growth, migration and invasion in cervical cancer cells. This
effect of SRA on tumor progression may be mediated by genes
involved in cell migration, invasion, EMT and the NOTCH signaling
pathway, as well as genes that encode VEGF, MMP-9, MMP-2,
E-cadherin, β-catenin, vimentin, Snail, NOTCH1, p300 and HES1. Our
findings suggested that SRA could potentially represent a novel
biomarker and therapeutic target for cervical cancer.
We discovered that knockout of lncRNA SRA expression
decreased cervical cancer cell proliferation, migration and
invasion. Specifically, the expression levels of MMP-2, MMP-9 and
VEGF were significantly lower in cells with siRNA-mediated SRA
knockdown. MMP-2 and MMP-9 degrade basement-membrane collagen at
the site of local invasion, thereby promoting tumor-cell invasion
and metastasis, which in turn leads to decreased survival rates in
various types of cancers (21,22).
Moreover, tumor angiogenesis plays a decisive role in tumor growth,
invasion and metastasis (23), and
the angiogenic factor, VEGF, a major target of many anticancer
medications, plays a pivotal role in tumor angiogenesis and
increases the ability of malignant cells to migrate and invade
Matrigel membranes (24). Our
findings suggested that SRA stimulated oncogenic activity in
cervical cancer cells by promoting aggressive and metastatic
characteristics via upregulating MMP-2, MMP-9 and VEGF
expression.
We found that knockout of lncRNA SRA in cervical
cancer cells reduced cell migration and invasion, possibly by
preventing EMT induction. We analyzed the following EMT-related
genes in the present study: E-cadherin, β-catenin,
vimentin, N-cadherin, Snail and Twist.
EMT is a well-characterized process by which epithelial cells lose
their cell polarity and cell-to-cell adhesion and acquire migratory
and invasive properties to become mesenchymal stem cells (25), processes that occur during
metastasis in many carcinomas (25). Loss of E-cadherin is considered a
crucial event in EMT, whereas N-cadherin promotes transendothelial
migration, which causes diminished intercellular connection between
two adjacent endothelial, thereby allowing cancer cells to slip
through (26). Additionally,
β-catenin weakens cell-to-cell adhesion between epithelial cells,
and promotes a more mobile and loosely associated mesenchymal
phenotype (27). Furthermore,
enhanced expression of transcription factors, such as Snail and
Twist, is associated with loss of intercellular adhesion (28), and vimentin constitutes a major
component of the cytoskeleton in mesenchymal cells, with its
upregulation promoting EMT induction (29).
In the present study, we also investigated whether
the NOTCH signaling pathway was compromised upon SRA knockdown. A
clear relationship was detected between decreased levels of NOTCH1,
HES1 and p300 proteins and SRA knockdown in CaSki cells. The NOTCH
signaling pathway represents a highly conserved cell-signaling
mechanism (30) that contributes to
tumor progression, including invasion, metastasis, EMT and
angiogenesis (31,32). NOTCH1 is a regulatory transmembrane
receptor that plays a crucial developmental role in cell-fate
determination and pattern formation (30). Several studies reported that NOTCH1
induces anoikis resistance, inhibits p53 activity, upregulates Myc
expression, and abrogates the growth-inhibitory effects of TGF-β in
cervical cancer cells (33–35). Additionally, HES1 influences the
maintenance of stem and progenitor cells as a member of the
NOTCH-signaling pathway (36) and
p300, a transcriptional co-activator in the NOTCH1 pathway,
functions as a histone acetyltransferase and regulates
transcription via chromatin remodeling. Furthermore, p300 also
plays a crucial role in cell proliferation and differentiation
(37).
Based on our results, we hypothesized that lncRNA
SRA functions as a key regulator of various signaling mechanisms
involved in EMT establishment and NOTCH signaling. Our data provide
evidence, for the first time, that lncRNA SRA promotes EMT and
NOTCH signaling in cervical cancer cells, potentially contributing
to cervical cancer growth, invasion and migration. The clinical
impact of lncRNA SRA expression and its potential therapeutic value
in the treatment of advanced cervical cancer patients needs to be
evaluated in future studies.
In conclusion, these findings showed that lncRNA SRA
is correlated with motility and invasiveness in cervical cancer
cells. Furthermore, lncRNA SRA promoted cervical cancer progression
by inducing cell migration and invasion via upregulation of MMP-2,
MMP-9, VEGF, as well as genes related to EMT and the NOTCH
signaling pathway. Therefore, lncRNA SRA constitutes a potential
therapeutic target and prognostic marker for cervical cancer.
Acknowledgements
The present study was supported by the Basic Science
Research Program through the National Research Foundation of Korea
(NRF) funded by the Ministry of Education, Science and Technology
(grant nos. NRF-2015R1A2A2A01008162 and
NRF-2015R1C1A2A01053516).
Glossary
Abbreviations
Abbreviations:
|
CCK-8
|
Cell Counting Kit-8
|
|
EMT
|
epithelial-mesenchymal transition
|
|
lncRNA
|
long non-coding RNA
|
|
qRT-PCR
|
quantitative reverse
transcription-polymerase chain reaction
|
|
siNC
|
negative-control siRNA
|
|
siRNA
|
small interfering RNA
|
|
SRA
|
steroid receptor RNA activator
|
References
|
1
|
Kodama J, Seki N, Masahiro S, Kusumoto T,
Nakamura K, Hongo A and Hiramatsu Y: Prognostic factors in stage
IB-IIB cervical adenocarcinoma patients treated with radical
hysterectomy and pelvic lymphadenectomy. J Surg Oncol. 101:413–417.
2010.PubMed/NCBI
|
|
2
|
Noordhuis MG, Fehrmann RS, Wisman GB,
Nijhuis ER, van Zanden JJ, Moerland PD, Ver Loren van Themaat E,
Volders HH, Kok M, ten Hoor KA, et al: Involvement of the TGF-beta
and beta-catenin pathways in pelvic lymph node metastasis in
early-stage cervical cancer. Clin Cancer Res. 17:1317–1330. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Perez DS, Hoage TR, Pritchett JR,
Ducharme-Smith AL, Halling ML, Ganapathiraju SC, Streng PS and
Smith DI: Long, abundantly expressed non-coding transcripts are
altered in cancer. Hum Mol Genet. 17:642–655. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Guttman M, Donaghey J, Carey BW, Garber M,
Grenier JK, Munson G, Young G, Lucas AB, Ach R, Bruhn L, et al:
lincRNAs act in the circuitry controlling pluripotency and
differentiation. Nature. 477:295–300. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Rinn JL, Kertesz M, Wang JK, Squazzo SL,
Xu X, Brugmann SA, Goodnough LH, Helms JA, Farnham PJ, Segal E, et
al: Functional demarcation of active and silent chromatin domains
in human HOX loci by non-coding RNAs. Cell. 129:1311–1323. 2007.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Ponting CP, Oliver PL and Reik W:
Evolution and functions of long non-coding RNAs. Cell. 136:629–641.
2009. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Hung T, Wang Y, Lin MF, Koegel AK, Kotake
Y, Grant GD, Horlings HM, Shah N, Umbricht C, Wang P, et al:
Extensive and coordinated transcription of non-coding RNAs within
cell-cycle promoters. Nat Genet. 43:621–629. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Liu C, Wu HT, Zhu N, Shi YN, Liu Z, Ao BX,
Liao DF, Zheng XL and Qin L: Steroid receptor RNA activator:
Biologic function and role in disease. Clin Chim Acta. 459:137–146.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Yan R, Wang K, Peng R, Wang S, Cao J, Wang
P and Song C: Genetic variants in lncRNA SRA and risk of breast
cancer. Oncotarget. 7:22486–22496. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Hussein-Fikret S and Fuller PJ: Expression
of nuclear receptor coregulators in ovarian stromal and epithelial
tumours. Mol Cell Endocrinol. 229:149–160. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Cooper C, Guo J, Yan Y,
Chooniedass-Kothari S, Hube F, Hamedani MK, Murphy LC, Myal Y and
Leygue E: Increasing the relative expression of endogenous
non-coding steroid receptor RNA activator (SRA) in human breast
cancer cells using modified oligonucleotides. Nucleic Acids Res.
37:4518–4531. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Colley SM and Leedman PJ: Steroid Receptor
RNA Activator - A nuclear receptor coregulator with multiple
partners: Insights and challenges. Biochimie. 93:1966–1972. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Cooper C, Vincett D, Yan Y, Hamedani MK,
Myal Y and Leygue E: Steroid Receptor RNA Activator bi-faceted
genetic system: Heads or Tails? Biochimie. 93:1973–1980. 2011.
View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Carninci P, Kasukawa T, Katayama S, Gough
J, Frith MC, Maeda N, Oyama R, Ravasi T, Lenhard B, Wells C, et al:
RIKEN Genome Exploration Research Group and Genome Science Group
(Genome Network Project Core Group): The transcriptional landscape
of the mammalian genome. Science. 309:1559–1563. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Morris KV and Vogt PK: Long antisense
non-coding RNAs and their role in transcription and oncogenesis.
Cell Cycle. 9:2544–2547. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Dinger ME, Amaral PP, Mercer TR, Pang KC,
Bruce SJ, Gardiner BB, Askarian-Amiri ME, Ru K, Soldà G, Simons C,
et al: Long non-coding RNAs in mouse embryonic stem cell
pluripotency and differentiation. Genome Res. 18:1433–1445. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Hall PA and Russell SH: New perspectives
on neoplasia and the RNA world. Hematol Oncol. 23:49–53. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Gupta RA, Shah N, Wang KC, Kim J, Horlings
HM, Wong DJ, Tsai MC, Hung T, Argani P, Rinn JL, et al: Long
non-coding RNA HOTAIR reprograms chromatin state to promote
cancer metastasis. Nature. 464:1071–1076. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Kim HJ, Lee DW, Yim GW, Nam EJ, Kim S, Kim
SW and Kim YT: Long non-coding RNA HOTAIR is associated with
human cervical cancer progression. Int J Oncol. 46:521–530. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Kim HJ, Eoh KJ, Kim LK, Nam EJ, Yoon SO,
Kim KH, Lee JK, Kim SW and Kim YT: The long non-coding RNA
HOXA11 antisense induces tumor progression and stemness
maintenance in cervical cancer. Oncotarget. 7:83001–83016. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Curran S and Murray GI: Matrix
metalloproteinases: Molecular aspects of their roles in tumour
invasion and metastasis. Eur J Cancer. 36:1621–1630. 2000.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Biewenga P, van der Velden J, Mol BW,
Stalpers LJ, Schilthuis MS, van der Steeg JW, Burger MP and Buist
MR: Prognostic model for survival in patients with early stage
cervical cancer. Cancer. 117:768–776. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Carmeliet P: VEGF as a key mediator of
angiogenesis in cancer. Oncology. 69 Suppl 3:S4–S10. 2005.
View Article : Google Scholar
|
|
24
|
Burger RA: Role of vascular endothelial
growth factor inhibitors in the treatment of gynecologic
malignancies. J Gynecol Oncol. 21:3–11. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Chaffer CL and Weinberg RA: A perspective
on cancer cell metastasis. Science. 331:1559–1564. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ramis-Conde I, Chaplain MA, Anderson AR
and Drasdo D: Multi-scale modelling of cancer cell intravasation:
The role of cadherins in metastasis. Phys Biol. 6:0160082009.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Morin PJ: beta-catenin signaling and
cancer. BioEssays. 21:1021–1030. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Martin TA, Goyal A, Watkins G and Jiang
WG: Expression of the transcription factors snail, slug, and twist
and their clinical significance in human breast cancer. Ann Surg
Oncol. 12:488–496. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Calaf GM, Balajee AS, Montalvo-Villagra
MT, Leon M, Daniela NM, Alvarez RG, Roy D, Narayan G and
Abarca-Quinones J: Vimentin and Notch as biomarkers for breast
cancer progression. Oncol Lett. 7:721–727. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Artavanis-Tsakonas S, Rand MD and Lake RJ:
Notch signaling: Cell fate control and signal integration in
development. Science. 284:770–776. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Timmerman LA, Grego-Bessa J, Raya A,
Bertrán E, Pérez-Pomares JM, Díez J, Aranda S, Palomo S, McCormick
F, Izpisúa-Belmonte JC, et al: Notch promotes
epithelial-mesenchymal transition during cardiac development and
oncogenic transformation. Genes Dev. 18:99–115. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wang Z, Banerjee S, Li Y, Rahman KM, Zhang
Y and Sarkar FH: Down-regulation of notch-1 inhibits invasion by
inactivation of nuclear factor-kappaB, vascular endothelial growth
factor, and matrix metalloproteinase-9 in pancreatic cancer cells.
Cancer Res. 66:2778–2784. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Nair P, Somasundaram K and Krishna S:
Activated Notch1 inhibits p53-induced apoptosis and sustains
transformation by human papillomavirus type 16 E6 and E7 oncogenes
through a PI3K-PKB/Akt-dependent pathway. J Virol. 77:7106–7112.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Masuda S, Kumano K, Shimizu K, Imai Y,
Kurokawa M, Ogawa S, Miyagishi M, Taira K, Hirai H and Chiba S:
Notch1 oncoprotein antagonizes TGF-beta/Smad-mediated cell growth
suppression via sequestration of co-activator p300. Cancer Sci.
96:274–282. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Klinakis A, Szabolcs M, Politi K, Kiaris
H, Artavanis-Tsakonas S and Efstratiadis A: Myc is a Notch1
transcriptional target and a requisite for Notch1-induced mammary
tumorigenesis in mice. Proc Natl Acad Sci USA. 103:9262–9267. 2006.
View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Liu ZH, Dai XM and Du B: Hes1: A key role
in stemness, metastasis and multidrug resistance. Cancer Biol Ther.
16:353–359. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Oswald F, Täuber B, Dobner T, Bourteele S,
Kostezka U, Adler G, Liptay S and Schmid RM: p300 acts as a
transcriptional co-activator for mammalian Notch-1. Mol Cell Biol.
21:7761–7774. 2001. View Article : Google Scholar : PubMed/NCBI
|