Introduction
Primary liver cancer originates in the liver and
comprises mainly hepatocellular carcinoma (HCC) and intrahepatic
cholangiocarcinoma (ICC), which account for approximately 90% and
5–15% of liver cancer cases, respectively (1). Hepatitis B virus (HBV), HCV, and
aflatoxins have been identified as major causal factors that act
individually and synergistically in the development of liver cancer
(2,3). Other genetic factors such as genetic
polymorphisms or gene mutations in tumor-associated genes may also
play important roles in the development of liver cancer (4).
Neurofibromatosis type 2 is a devastating autosomal
dominant disorder that includes schwannomas, meningiomas, and
ependymomas (5). Neurofibromatosis
type 2 with mutations in the neurofibromin 2 (NF2) gene is
characterized by enhanced cancer predisposition. The NF2
gene encodes Merlin, a cytoskeletal protein that also functions as
a microtubule stabilizing protein (6). NF2 mutations have been
identified in the majority of sporadic and NF2-associated
schwannomas (7). NF2 is a
regulator of the Hippo pathway, which controls organ size and
regulates cell proliferation, motility, survival, and signaling
pathways. Genetic mutations in NF2 are frequently identified
in tumors of the central nervous system; however, their presence in
cancers of other organs has seldom been reported (8).
Recently, an animal study reported that
liver-specific deletion of the NF2 tumor-suppressor gene in
the developing or adult mouse results in the progressive expansion
of hepatic progenitor/oval cells throughout the liver without
affecting differentiated hepatocytes (8). All surviving mice eventually developed
both cholangiocellular and hepatocellular carcinomas, suggesting
that Nf2−/− progenitors can be a cell of origin for
these tumors (8). Drvarov et
al also proposed that the development of primary liver cancer
may be directly regulated by the NF2 gene (9). Although these studies indicated that
loss of function of the NF2 gene may be associated with the
development of liver cancers, its role in human primary liver
cancer remains unclear (8–11).
To explore the role of NF2 in human liver
carcinogenesis, we investigated the effects of molecular
alterations of NF2/Merlin in human primary liver cancer. In
addition, we analyzed the potential association of
NF2/Merlin with Yes-associated protein (YAP), another Hippo
pathway gene downstream of NF2, in the development of
primary liver cancer.
Materials and methods
Tissue samples and cell line
A total of 144 patients with primary liver cancer
were enrolled in this study, including 106 HCCs and 38 ICCs. The
HCC patients included 92 men (86.8%) and 14 women (13.2%) aged
22–80 years with a mean age of 51.6 years, while the ICC patients
included 31 men (81.6%) and 7 women (18.4%) aged 30–81 years with a
mean age of 54 years. The patients underwent surgical treatment at
the Liver Research Center, Beijing Friendship Hospital, Capital
Medical University, Department of Hepatology, Tianjin Infectious
Disease Specialty Hospital and the Minimally Invasive Hepatobiliary
Cancer Center, Beijing You-an Hospital between May 2011 and
December 2015. All tumor samples were fixed in buffered formalin
and embedded in paraffin; five matched pairs of frozen tumors were
also included in the analysis (Table
I).
 | Table I.Clinicopathological parameters of
investigated cases. |
Table I.
Clinicopathological parameters of
investigated cases.
| Subjects | PLC | HCC | ICC | P-value |
|---|
| No. of cases | 144 | 106 | 38 | – |
| Age (years) |
|
|
|
|
|
Mean | 52.2 | 51.6 | 54.0 | – |
|
Range |
22–81 |
22–80 | 30–81 |
|
| Sex |
|
|
|
|
|
Male | 123 | 92 | 31 | 0.30 |
|
Female | 21 | 14 | 7 |
|
| Tumor stage |
|
|
|
|
| Stage
I | 72 | 55 | 17 | 0.29 |
|
>Stage I | 72 | 51 | 21 |
|
| HBV DNA |
|
|
|
|
|
Positive | 115 | 93 | 22 | <0.001 |
|
Negative | 29 | 13 | 16 |
|
|
Differentiation |
|
|
|
|
|
Well | 22 | 18 | 4 | 0.54 |
|
Moderate | 73 | 54 | 19 |
|
|
Poor | 49 | 34 | 15 |
|
Two hundreds and nineteen whole blood samples from
patients with HCC and 194 whole blood samples, from healthy
controls from the Beijing Friendship Hospital and Tianjin
Infectious Disease Specialty Hospital, were enrolled to analyze the
genetic susceptibility of the NF2 single nucleotide
polymorphism (SNP).
All patients provided informed consent and approval
for the use of their clinical materials for research purposes. The
study protocol was approved by the Clinical Research Ethics
Committee of Beijing Friendship Hospital. The HCC cell line HepG2
was purchased from the National Platform of Experimental Cell
Resource for Sci-Tech (Beijing, China).
Single-stranded conformational
polymorphism (SSCP) analysis and direct DNA sequencing for NF2
mutations
SSCP analysis was performed to prescreen for
mutations in the NF2 gene. Primers for NF2 were
designed with the online software Primer3 (Table II). Extraction of genomic DNA from
paraffin sections and PCR-SSCP analysis for NF2 were
performed as described previously (12,13).
Genomic DNA of the whole blood was extracted using the QIAamp DNA
Mini kit (Qiagen, Hilden, Germany). Electrophoresis was performed
using apparatus DYCZ-20E (Beijing Liu-Yi Biotechnology, Beijing,
China) at 45 W for 4.5–5.5 h at room temperature with cooling by
fan. Gels were silver stained as previously described (12,13).
Samples exhibiting mobility shifts on SSCP analysis were
subsequently re-amplified with the same primers as used for SSCP,
and the PCR products were sequenced using a Big Dye Terminator
cycle sequencing kit (ABI Prism; Applied Biosystems, Foster City,
CA, USA) in an ABI 3730 DNA sequencer (Applied Biosystems).
Mutations identified were verified by sequencing a second product
of amplification on both strands. PolyChen-2 software (http://genetics.bwh.harvard.edu) was used to
predict the functional consequence of the identified individual
missense mutations.
 | Table II.Primers for SSCP and direct DNA
sequencing for the NF2 gene. |
Table II.
Primers for SSCP and direct DNA
sequencing for the NF2 gene.
|
| Sequence |
|
|---|
|
|
|
|
|---|
| Exon | Forward | Reverse | Product size
(bp) |
|---|
| 1 |
GGTCCCGGGCCTGAGC |
CTCGACTGTCACCGCAGCAG | 179 |
| 2 |
TCCCCATTGGTTTGTTATTG |
AGCCCCACCAGTTTCATC | 182 |
| 3 |
CACAGGAGGAAGTGCCAATA |
GGGGTAGCCTTGACTGATG | 199 |
| 4 |
GTGAGGCCATCTGTTGTG |
TTAACGCCCAGGAAAAATAC | 205 |
| SNP43608
A>Ca |
|
|
|
| 5 |
CTCTCCCTTTCTTCTTTCC |
GGTTAGCTTTCTTTTAGACCA | 154 |
| 6 |
TAAAAGTGGCAAACAATACC |
GCCCATAAAGGAATGTAAAC | 191 |
| 7 |
GACAGTGTCTTCCGTTCTCC |
GGCCCTCACTCAGTCTCTG | 158 |
| 8 |
GGCGCTTACAGTAGCTGTTCTT |
CACACATGTCCTACCTCCTTGTC | 184 |
| 9 |
GGCTGTCGGACTGAAACTGT |
GCGCCAAGTGAGATACCATT | 150 |
| 10 |
AACCTTTTTGTCTGCTTCTG |
CCAGGACTGACCACACAG | 194 |
| 11 |
TCGAGCCCTGTGATTCAATGACT |
AACCCCAGCCCCTCAGAAAT | 190 |
| 12-1 |
ATCTGGGCGGGAGAACAG |
ATTTCCTGCTCAGCCTCTGC | 162 |
| 12-2 |
CCGAGGAGGAGGCAAAACTTC |
CAGCCTCCTCGCCAGTCTG | 207 |
| 13 |
CCCTCTTCTGTGAAGCTGACA |
CCGGGAGGAAAGAGAACATC | 208 |
| 14 |
CCAAGCTCCTAATCCGAAAT |
GGCACAGGGGGCTACATA | 180 |
| 15 |
TGATGCATGATACCCTCTTG |
GAGACCCTGGGTACCTTTTTA | 204 |
| 16 |
CCCTCTCAGCTTCTTCTCTGC |
CCCTATGGATGGCTCTCTTGA | 170 |
| SNP 8240
G>C |
AGAATATTCGCCGTGTGTCC |
TGAGTGGCTACAGTGGCAGT | 227 |
| SNP 99632
C>T |
TGCAATTGCCTTGAACTACG |
GCTTCTCAGGGCTTCAGTGT | 228 |
Immunohistochemistry (IHC)
Sections (4-µm) were cut for IHC. After
deparaffinization of the slides, antigen retrieval was performed in
antigen unmasking solution (Vector H-3300) with microwaving for 15
min, keeping the solution boiling, and endogenous peroxidase
activity was blocked with 0.3% H2O2 in
methanol for 30 min followed by treatment with 5% skimmed milk in
phosphate buffered saline (PBS)-0.1% bovine serum albumin for at
least 1 h at room temperature to block nonspecific staining.
Immunohistochemical staining was performed using
antibodies against Merlin (Santa Cruz Biotechnology, sc331) and YAP
(Abcam, ab52771) at 4°C overnight. Secondary antibody (Vector,
MP-7401) was used at 37°C for 1 h, and visualization of
antigen-antibody reactions was achieved with 3,3′-diaminobenzidine
(Vector, SK-4100). Tissue structures were visualized by
counterstaining with hematoxylin.
IHC scoring for Merlin and YAP was based on the
strength and distribution of staining by two independent observers
blinded to the clinical data (14).
The IHC reaction was scored by multiplying the percentage of
positive tumor cells (PP: 1, <10% positive tumor cells; 2,
10–49%; 3, ≥50% positive tumor cells) by their prevalent degree of
staining (SI: 0, negative; 1, weak; 2, strong staining).
Immunoreactive scores (IRS = PP × SI) ranging from 3 to 6 were
defined as IHC++, whereas IRS value <3 was defined as IHC 0 to
+. Cases with Merlin IHC level ++ were defined as having increased
Merlin expression. Cases with YAP IHC level ++ were defined as
having increased YAP expression (14). When there was a discordance between
the observers, a pathologic peer review was undertaken to determine
a consensus score.
Western blot analysis
Western blot analysis was performed as previously
described (14,15). Briefly, frozen tissues or cell
cultures were lysed and clarified by centrifugation. The protein
concentration was determined using a BCA kit (Pierce, Rockford, IL,
USA). Each protein extract (80 µg) was loaded onto a 10% SDS-PAGE
gel and transferred to a nitrocellulose membrane. The membrane was
probed with the primary antibody against Merlin (Santa Cruz
Biotechnology, sc331), with β-actin (Sigma, St. Louis, MO, USA) as
the loading control. The membrane was then incubated with
species-specific secondary horseradish peroxidase-conjugated
antibodies (Sigma). Protein bands were revealed by
chemiluminescence (Pierce) and detected on X-ray film (Kodak,
Rochester, NY, USA).
Real-time PCR for YAP
amplification
For the detection of YAP gene amplification,
real-time PCR was performed with the Homo sapiens actin, H
(HACTIN) sequence as the reference. The sequences of primers
were as follows: 5′-GTTTGGATGATGGATGCCATT-3′ (forward) and
5′-ATGCTGTGACATGAAGCATCTGA-3′ (reverse) for YAP and
5′-GCAAAGACCTGTACGCCAACA-3′ (forward) and 5′-TGCATCCTGTCGGCAATG-3′
(reverse) for HACTIN. qPCR was performed using the
SYBR® Premix Ex Taq™ in an ABI PRISM 7500 detection
system according to the manufacturer's instructions. For the
reactions, annealing was carried out at 65°C, and 40 cycles of
amplification were performed. Data were normalized using
HACTIN as a reference gene. The relative levels of
expression of the target gene among the different samples were
calculated accordingly.
The gene copy numbers of the samples were calculated
according to the following formula: ∆Ct = [Ct (target) - Ct
(reference)] and ∆∆Ct = [∆Ct(tumor) - ∆Ct (normal)]. The relative
gene copy numbers were calculated using the expression 2 ×
2−∆∆Ct, with a ∆∆Ct ratio 2 × 2−∆∆Ct >2.88
considered to indicate amplification (16).
Statistical analysis
The Chi-square test (for expected values >5) and
the Fisher's exact test (for expected values ≤5) were used to
determine the molecular associations using SAS v9.2 software (SAS
Institute, Inc., Cary, NC, USA). P<0.05 was considered to
indicate a statistically significant result for all tests.
Results
NF2 mutations and single nucleotide
polymorphisms in HCC and ICC
A total of 144 cases of primary liver cancer,
including 106 cases of HCC and 38 cases of ICC, were screened for
NF2 mutations. Since NF2 mutations have been
identified in exons 1–15, but less identified in exons 16–17, and
due to the very large size of exon 17 (17), we screened only mutation in exon
1–16 of the NF2 gene.
SSCP followed by direct sequencing of the 16 exons
of NF2 revealed a total of four missense NF2 point
mutations in 2 of 106 (1.9%) HCCs and 2 of 38 (5.3%) ICCs.
(Table III, Fig. 1A), distributed among exons 10, 11,
and 15 (Fig. 1B), including one
G→C, one C→T, one A→G, and one G→A transition (Table III). In addition, two splicing
variants and one synonymous mutation: i.e. IVS7+36 G→C, IVS16+24
C→T and E89E (GAA→GAG) were identified in three ICC cases,
respectively. As predicted by PolyChen-2 software, the mutation
Q319H and H562R were predicted to be probably damaging, while the
L339F and E355K were predicted to be benign mutations but with
minor impairments of function with scores of 0.441 and 0,031,
respectively.
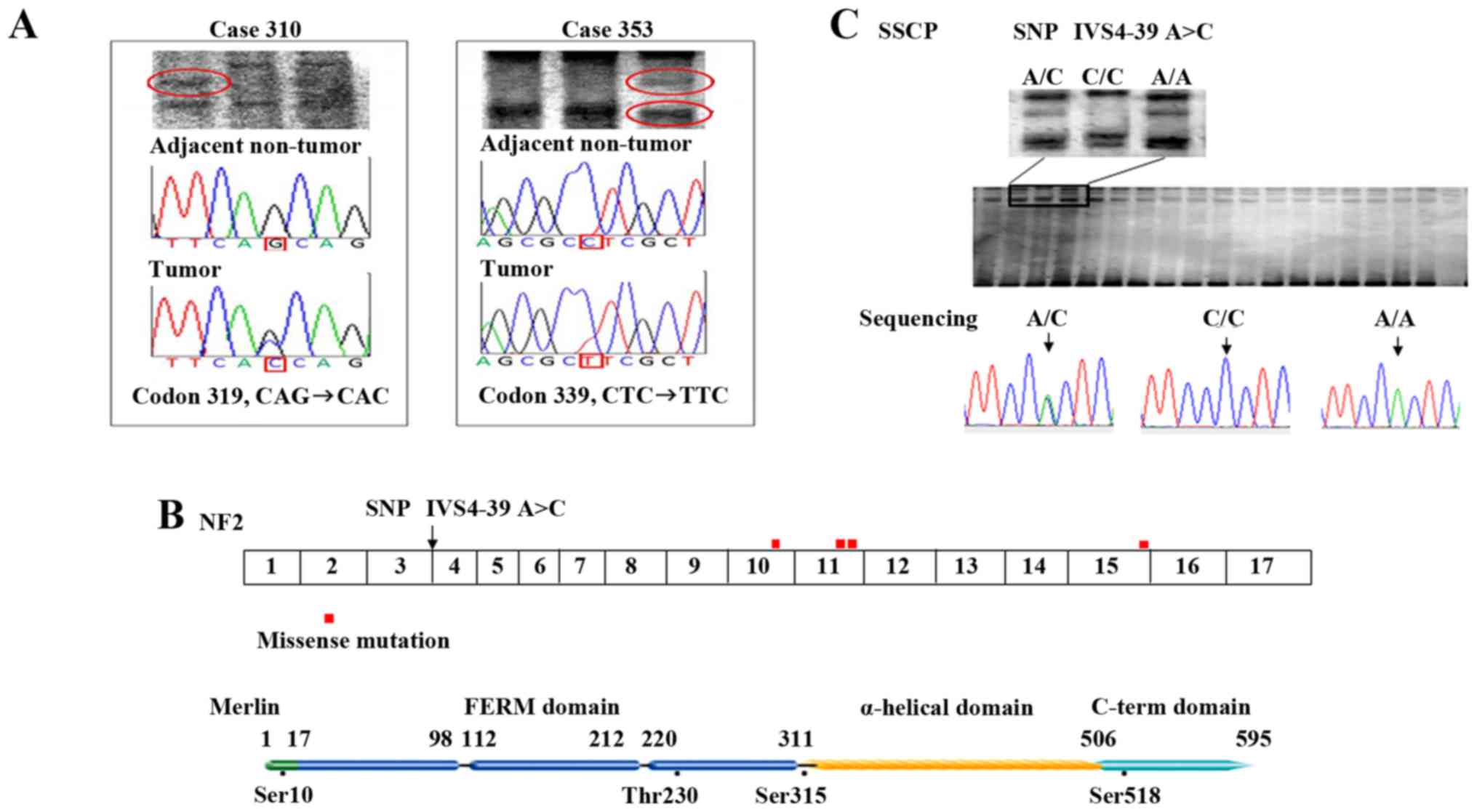 | Figure 1.NF2 missense mutations
identified in primary liver cancer cases and allele frequency of
NF2 IVS4-39 A/A. (A) Representative SSCP and direct
sequencing of the NF2 gene in HCC. Right panel, mutation at
codon 319 (CAG→CAC, Gln→His) in an HCC case (case 310). Left panel,
mutation at codon 339 (CTC→TTC, Leu→Phe) in an HCC case (case 353).
The reverse complement sequence is given. The mutations were not
identified in the corresponding adjacent non-tumor tissue,
indicating that the mutations are somatic. (B) Distribution of
NF2 mutations in the primary liver cancer cases. Mutation
are located preferentially in exon 10 to 11 (the joint part of FERM
domain and α-helical domain), but also in other functional domains
(close to phosphorylation site, Ser518/Ser315). (C) SSCP analysis
and direct sequencing of SNP IVS4-39 A>C, with three patterns of
bands identified by SSCP electrophoresis, corresponding to genotype
A/C, C/C, and A/A. HCC, hepatocellular carcinoma; SSCP,
single-stranded conformational polymorphism; SNP, single nucleotide
polymorphism. |
 | Table III.Identified NF2 missense
mutations in liver cancer cases and the allele frequency of SNP
IVS4-39 A/A. |
Table III.
Identified NF2 missense
mutations in liver cancer cases and the allele frequency of SNP
IVS4-39 A/A.
| NF2 missense
mutations |
|---|
|
|---|
| Case no. | Age/Sex | Tumor type | NF2
mutation |
|---|
| 310 | 50/M | HCC | Codon 319, CAG→CAC,
Gln→His |
| 353 | 43/M | HCC | Codon 339, CTC→TTC,
Leu→Phe |
| 276 | 47/M | ICC | Codon 355, GAG→AAG,
Glu→Lys |
| 312 | 61/M | ICC | Codon 562, CAC→CGC,
His→Arg |
|
| SNP
IVS4-39 |
|
| Alleles | HCC | Health
controls |
χ2 | P-value |
|
| C/C+A/C | 146 (66.67%) | 154 (79.38%) | 7.74 | 0.005401 |
| A/A | 73
(33.33%) | 40
(20.62%) |
|
|
Three specific SNPs of the NF2 gene were
chosen for analysis based on their intragenic location, the
validation status in ethnically diverse population, and a
preliminary analysis showing their polymorphic nature in the
Chinese population (18,19). There were no significant differences
between HCC and healthy controls in the rare allele frequency of
the SNPs 8240 G>C (promoter region, G/G: 74.18% vs. 76.65%,
P=0.66) and 99632 C>T (3′ non-coding region, C/C: 10.00% vs.
13.57%, P=0.33), except IVS4-39 A>C identified by the present
study, of which the allele (A/A) frequency was significantly higher
in tumors than in controls (Table
III and Fig. 1C).
Merlin expression is increased in
tumor tissues compared with that in adjacent non-tumor tissues of
HCC
Of the 144 cases of primary liver cancer with
paraffin sections, 131 (94 HCCs and 37 ICCs) were available for
analysis of Merlin IHC expression. In the 94 cases of HCC and 37
cases of ICC, adjacent non-tumor tissue was available for 80 HCCs
and 31 ICCs. The IHC analysis showed that 26 of 80 HCCs or 10 of 31
ICCs (32.5 or 32.3%) had increased Merlin expression in tumors
compared with that in adjacent tissues. In 49 of 80 HCCs or 21 of
31 ICCs (61.3 or 67.7%), the expression levels of Merlin were low
in both tumors and adjacent tissues. None of the ICCs showed
increased Merlin expression in adjacent non-tumor tissues compared
with that in tumor tissues, and increased Merlin expression was not
observed in both adjacent non-tumor tissues and tumor tissues of
the ICCs. However, the results showed that the rate of increased
Merlin expression was significant higher in tumor tissues than in
adjacent non-tumor tissues of HCC (P=0.013, Table IV), suggesting that Merlin may play
a role as an oncogene in HCC development.
 | Table IV.Merlin expression in tumor and
adjacent non-tumor tissue in HCC. |
Table IV.
Merlin expression in tumor and
adjacent non-tumor tissue in HCC.
|
| Non-tumor (IHC
++) | Non-tumor (IHC
0-+) | Total | χ2 | P-value |
|---|
| Tumor (IHC ++) | 9 | 17 | 26 | 6.16 | 0.013 |
| Tumor (IHC
0-+) | 5 | 49 | 54 |
|
|
| Total | 14 | 66 | 80 |
|
|
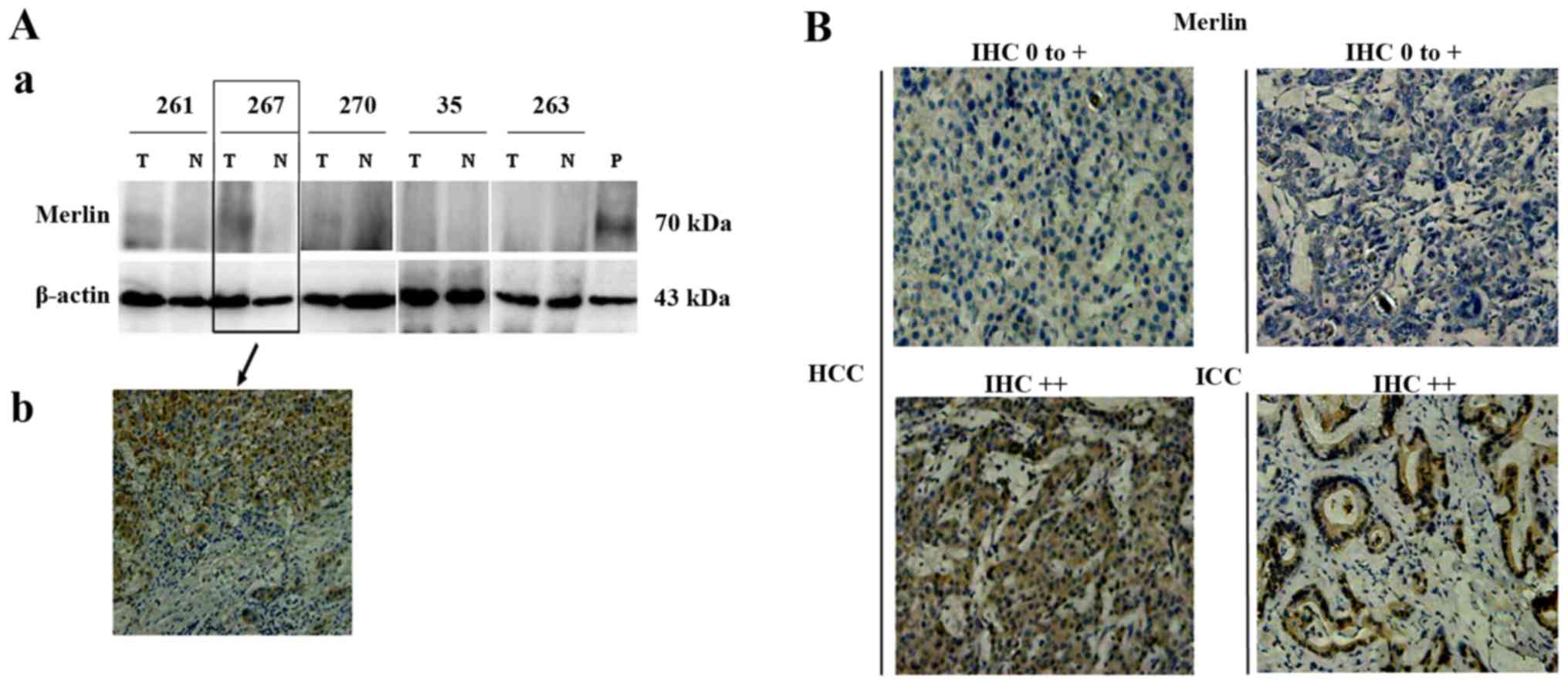
Western blot analysis of five pairs of
representative HCC cases with available IHC data (cases 261, 267,
and 270, IHC ++; case 35 and 263, IHC 0-+) showed consistent
results with those of IHC regarding Merlin expression, confirming
that Merlin levels were higher in tumors than in adjacent non-tumor
tissues of HCC (Fig. 2A).
Increased Merlin expression is
associated with the degree of differentiation of tumor cells
Merlin expression was mainly localized in the
cytoplasm of tumor cells (Fig. 2B),
and upregulated in 20.2% (19/94) of the HCCs and 45.9% (17/37) of
the ICCs. There was no correlation between Merlin expression and
the clinicopathological characteristics of HCC (Table V). However, of the 37 ICC cases, 3
(13.6%), 7 (31.8%), and 12 (54.5%) with IHC scores of 3–6, 1–2, and
0, respectively, had well and medium differentiated tumors, whereas
9 (60.0%), 1 (6.7%), and 5 (33.3%) cases with IHC scores of 3–6,
1–2, and 0, respectively, had poorly differentiated ICCs,
indicating that low Merlin expression was associated with a worse
degree of differentiation of tumor cells (P=0.009; Table V). These results indicated that
alterations in Merlin expression are associated with the degree of
malignancy of ICC cells and that Merlin may play a tumor-suppressor
role.
 | Table V.Association between Merlin IHC
expression and clinical parameters in HCC and ICC. |
Table V.
Association between Merlin IHC
expression and clinical parameters in HCC and ICC.
|
|
|
| Merlin IHC
scoring |
|
|
|---|
|
|
|
|
|
|
|
|---|
| Tumor type | Parameters | No. | 0 (n=12) n (%) | 1-2 (n=8) n
(%) | 3-6 (n=17) n
(%) | χ2 | P-value |
|---|
| HCC | Tumor stage |
|
|
|
|
|
|
|
| TNM
I | 54 | 2 (3.7) | 42 (77.8) | 10 (18.5) | 0.356 | 0.837 |
|
| >TNM
I | 40 | 2 (5.0) | 29 (72.5) | 9
(22.5) |
|
|
|
| HBV
infectiona |
|
|
|
|
|
|
|
|
Positive | 80 | 4 (5.0) | 61 (76.3) | 15 (18.8) | 0.504 | 0.777 |
|
|
Negative | 9 | 0 (0) | 7
(77.8) | 2
(22.2) |
|
|
|
|
Differentiation |
|
|
|
|
|
|
|
|
Well+moderate | 61 | 3 (4.9) | 46 (75.4) | 12 (19.7) | 0.205 | 0.903 |
|
|
Poor | 33 | 1 (3.0) | 25 (75.8) | 7
(21.2) |
|
|
| ICC | Tumor stage |
|
|
|
|
|
|
|
| TNM
I | 16 | 5 (31.3) | 4 (25.0) | 7
(43.7) | 0.191 | 0.909 |
|
| >TNM
I | 21 | 7 (33.3) | 4 (19.1) | 10 (47.6) |
|
|
|
| HBV
infectionb |
|
|
|
|
|
|
|
|
Positive | 20 | 7 (35.0) | 5 (25.0) | 8 (40.0) | 0.944 | 0.624 |
|
|
Negative | 16 | 4 (25.0) | 3 (18.8) | 9 (56.3) |
|
|
|
|
Differentiation |
|
|
|
|
|
|
|
|
Well+moderate | 22 | 3 (13.6) | 7
(31.8) | 12 (54.5) | 9.394 | 0.009 |
|
|
Poor | 15 | 9 (60.0) | 1 (6.7) | 5
(33.3) |
|
|
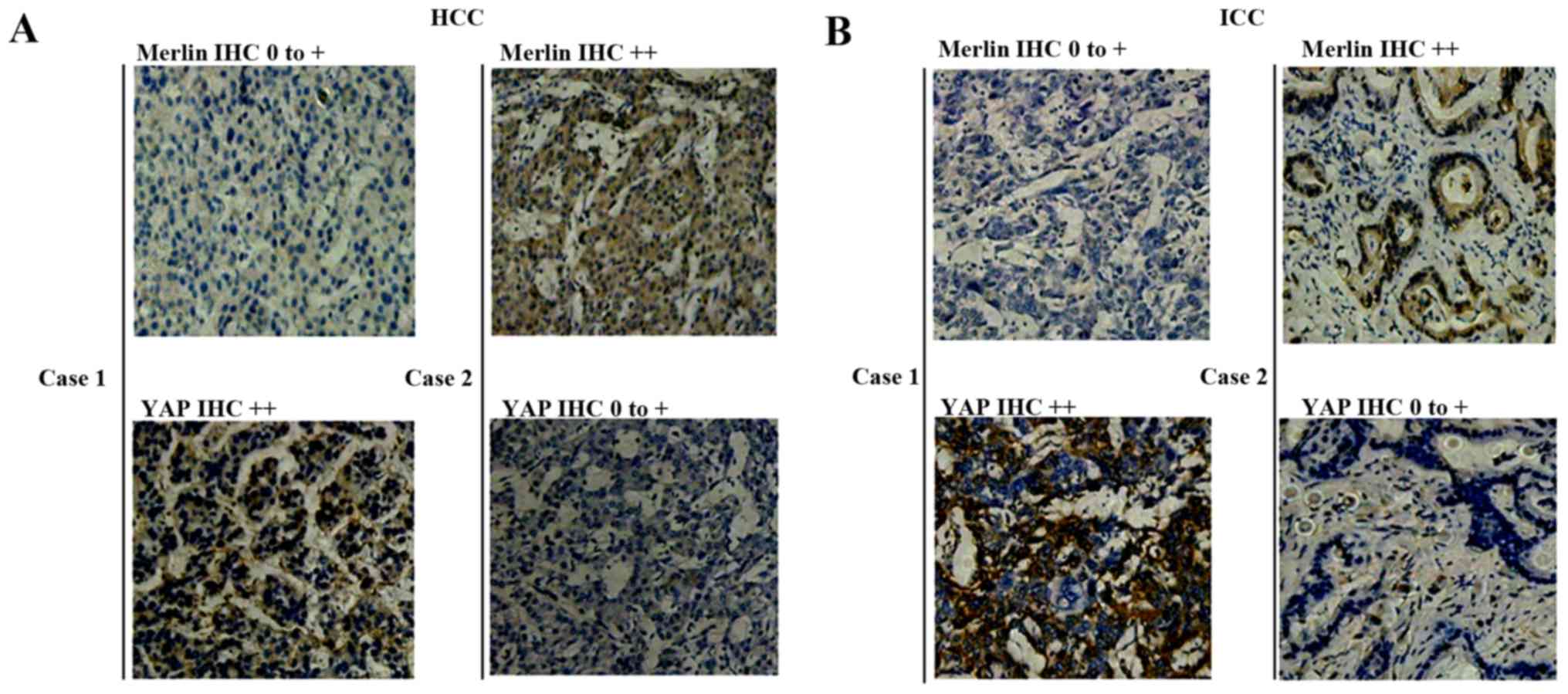
Merlin expression is significantly
negatively correlated with YAP expression in tumor cells
YAP, which acts downstream of NF2/Merlin, was
analyzed by IHC and gene amplification to examine the association
between the two genes. Similar to Merlin, the YAP protein was
expressed predominantly in the cytoplasm of tumor cells (Fig. 3A and B). YAP was upregulated in 48
of the 94 HCC cases (51.1%) and in 27 of the 37 ICC cases (73.0%)
analyzed. No correlation was observed between Merlin and YAP
expression in HCCs, although a tendency towards a negative
correlation was observed (P=0.165, Table VI, Fig.
3A). However, a significant negative correlation was observed
between Merlin and YAP expression in ICC (P=0.016, Table VI, Fig.
3B), suggesting that they may play antagonistic roles in
tumorigenesis.
 | Table VI.Association of Merlin and YAP protein
expression in HCC and ICC. |
Table VI.
Association of Merlin and YAP protein
expression in HCC and ICC.
| Tumor type | Merlin/Yap
expression | YAP IHC ++ | YAP IHC 0-+ | Total | χ2 | P-value |
|---|
| HCC | Merlin IHC ++ | 7 | 12 | 19 | 1.93 | 0.165 |
|
| Merlin IHC 0-+ | 41 | 34 | 75 |
|
|
|
| Total | 48 | 46 | 94 |
|
|
| ICC | Merlin IHC ++ | 9 | 8 | 17 |
| 0.011a |
|
| Merlin IHC 0-+ | 18 | 2 | 20 |
|
|
|
| Total | 27 | 10 | 37 |
|
|
Of the 94 cases of HCC and 37 cases of ICC,
YAP amplification was found in 33.0% (31/94) of the HCCs and
32.4% (12/37) of the ICCs. No significant correlation was observed
between NF2 mutations and YAP amplification in HCCs
and ICCs.
Discussion
The Hippo signaling pathway, also known as the
Salvador/Warts/Hippo (SWH) pathway, controls organ size in animals
through regulation of cell proliferation and apoptosis (6,8). The
involvement of the Hippo pathway in organ size determination and
tumor suppression has been confirmed in genetically engineered
mouse models. The Hippo pathway regulates growth by phosphorylating
and inactivating YAP, an evolutionarily conserved transcriptional
coactivator (8). NF2 is the
only Hippo pathway gene that is mutationally inactivated in cancer,
and YAP is a major downstream effector of the NF2 tumor
suppressor with a role in mammalian growth control. The Hippo
pathway functions in the proliferation, differentiation, and
self-renewal of tissue-specific stem cells (8).
NF2 is involved in cytoskeleton development
and is a microtubule stabilizer (6). As a Hippo pathway gene involved in the
regulation of cell proliferation, motility, survival, and signaling
pathways, genetic mutations in NF2 have been detected in
tumors of the central nervous system. However, NF2 mutations
have seldom been reported in tumors of other organs (7). Recent animal studies have shown that
the disruption of NF2 function in hepatic progenitor/oval
cells may lead to the development of primary liver cancers
including HCC and ICC (8). The
diverse roles of NF2/Merlin in the pathogenesis of cancer
have been described.
NF2 gene mutations have been identified in
the majority of sporadic and NF2-associated schwannomas
(20–23). Neurofibromatosis type 2 is a
tumor-prone disorder characterized by the development of multiple
schwannomas and meningiomas. Affected individuals inevitably
develop schwannomas, typically affecting both vestibular nerves
leading to hearing loss and deafness. The frequency of the
NF2 gene mutation is approximately 71% (12/17 cases), 24%
(21/28 cases), and 41.7% (5/12 cases) in sporadic schwannomas,
meningiomas, and ependymomas, respectively (21–23).
Recent studies have revealed that mutation in the
NF2 gene is also present in other human cancers, such as
renal cell carcinoma, breast carcinoma, malignant mesotheliomas,
lung carcinoma, colorectal carcinomas, and prostate carcinomas
(23–26). These studies suggested that
NF2 mutations may contribute to tumorigenesis in multiple
human cancers. In a study by Yoo et al, one NF2
mutation was detected in 45 HCC samples (11), whereas Kanai et al did not
detect any mutations in six exons of the NF2 gene (10). These previous studies were confined
to HCC (10,11). The present study is the first to
examine the association between NF2 mutations and the
development of ICC. Since the NF2 mutation rate in the
population is estimated to be 6.5×10−6 (27), our findings indicated that
NF2 mutations may be associated with the pathogenesis of
primary liver cancer (P<0.05), especially ICC. In addition, our
results suggested that the genotype of NF2 IVS4-39 A/A may
be a potential risk factor for development of HCC in the Chinese
population. The NF2 splicing variant IVS4-39 A>C was
close to the 5′ end of exon 4 in FERM domain, and may have
functional consequence on the splicing of NF2 mRNA. However,
further functional study of the variant would be helpful to
elucidate the exact role of the variant.
Similar to ezrin-radixin-moesin (ERM) proteins,
Merlin is a critical component that provides a regulated linkage
between membrane proteins and the cortical cytoskeleton, in
addition to its involvement in signal-transduction pathways
(28,29). The main functional domains of Merlin
are the FERM domain, α-helical domain, and C-terminal domain, which
contains an F-actin binding site, and the protein contains several
critical phosphorylation sites, such as Ser10, Ser315, Ser518, and
Thr230. Merlin can switch between open and closed conformations,
which determine its activity and are regulated by changes in
phosphorylation status (28,29).
Similar to other ERM proteins, the function of Merlin is modulated
by the association between the FERM domain and C-terminal domain.
Its head-to-tail folding results in a closed active conformation
upon dephosphorylation of Ser518 in the C-terminal domain, while in
the open conformation upon phosphorylation of Ser518, Merlin is
inactive and incapable of acting as a tumor-suppressor protein
(30). The NF2 mutations
identified in liver cancer are located mainly in the FERM and
C-terminal domains. In particular, mutation in Ser567, which is
proximal to the Ser518 phosphorylation site, may affect the
phosphorylation status of Ser518, as well as the association
between the FERM domain and C-terminal domain. The NF2
mutations identified in the present study may play a significant
roles in the pathogenesis of primary liver cancer.
Both the upregulation and downregulation of Merlin
in tumor tissues have been reported in different types of cancer
(20,31,32).
The results of the present study suggest that the NF2 gene
may play a dual role as an oncogene and tumor-suppressor gene, as
reported previously in other cancers (33,34).
Chen et al reported that Merlin could be overexpressed in
some ovarian cancer (OVCA) cell lines and OVCA ascites cells,
suggesting that Merlin could be an oncoprotein rather than a
tumor-suppressor protein in certain OVCA cells, and that the
functional duality of Merlin might represent a paradigm in proteome
complexity and is especially important in investigating
multifactorial diseases such as cancer (33). Similarly, it has been reported that
the activator protein-1 (AP-1) protein acts as either an oncogene
or an anti-oncogene by regulating genes involved in cell
proliferation, differentiation, apoptosis, angiogenesis and tumor
invasion depending on the cell type, cellular condition (e.g.,
differentiation stages), tumor stage and genetic background of the
tumor (34). It has a double-edged
activity; it can be anti-oncogenic by inducing apoptosis and it can
be oncogenic by signaling cell survival (34). Thus, we deduced that Merlin may act
as an oncogene in the development of HCC with a similar mechanism.
Further functional study of Merlin in HCC development would be
helpful to elucidate the underlying mechanism.
Recent reseach has shown that YAP is a key driver of
tumorigenesis and tissue overgrowth caused by loss of NF2 in
the murine liver (35). Using
conditional knockout mice, Zhang et al demonstrated that the
NF2/Merlin tumor suppressor and the YAP oncoprotein function
antagonistically to regulate liver development. While inactivation
of YAP led to loss of hepatocytes and biliary epithelial cells,
inactivation of Nf2 led to hepatocellular carcinoma and bile duct
carcinoma. The suppression of NF2 mutant liver phenotypes by
loss of YAP suggest that YAP is a major effector of the NF2
tumor suppressor in mammalian growth control (35). Consistently, the present study
identified a significant negative correlation between Merlin and
YAP expression in ICC, suggesting that they may function in an
antagonistic manner in tumorigenesis.
In summary, the present study provides initial
evidence of the involvement of NF2/Merlin in HCC and ICC,
which could be of value to improve our understanding of the
molecular pathogenesis of primary liver cancer.
Acknowledgements
The authors would like to thank Dr Xiao-Yan Shi
(Department of Pathology, Beijing Friendship Hospital, Capital
Medical University) for her critical pathological review. This
study was supported by a grant from the Beijing Natural Science
Foundation (no. 7132058), and a grant from the Nature Science
Foundation of China (no. 81650014).
References
|
1
|
Theise ND, Chen CJ and Kew MC: Liver
cancerWorld Cancer Report 2014. Stewart B and Wild C: International
Agency for Research on Cancer; Lyon: pp. 577–593. 2014
|
|
2
|
Chen JG and Zhang SW: Liver cancer
epidemic in China: Past, present and future. Semin Cancer Biol.
21:59–69. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Shiraha H, Yamamoto K and Namba M: Human
hepatocyte carcinogenesis (Review). Int J Oncol. 42:1133–1138.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Zhao Z and Huang J: Recent updates of
genetic and genomic alterations in hepatocellular carcinoma.
Hepatoma Res. 2:31–35. 2016.
|
|
5
|
Evans DG, Sainio M and Baser ME:
Neurofibromatosis type 2. J Med Genet. 37:897–904. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Chiasson-MacKenzie C, Morris ZS, Baca Q,
Morris B, Coker JK, Mirchev R, Jensen AE, Carey T, Stott SL, Golan
DE, et al: NF2/Merlin mediates contact-dependent inhibition of EGFR
mobility and internalization via cortical actomyosin. J Cell Biol.
211:391–405. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Pylkkänen L, Sarlomo-Rikala M, Wessman M,
Hämäläinen E, Sainio M, Husgafvel-Pursiainen K and Carpén O:
Chromosome 22q alterations and expression of the NF2 gene product,
merlin, in gastrointestinal stromal tumors. Hum Pathol. 34:872–879.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Benhamouche S, Curto M, Saotome I, Gladden
AB, Liu CH, Giovannini M and McClatchey AI: Nf2/Merlin controls
progenitor homeostasis and tumorigenesis in the liver. Genes Dev.
24:1718–1730. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Drvarov O and Cubero FJ: Neurofibromatosis
type 2/Merlin: Sharpening the myth of prometheus. Hepatology.
53:1767–1770. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Kanai Y, Tsuda H, Oda T, Sakamoto M and
Hirohashi S: Analysis of the neurofibromatosis 2 gene in human
breast and hepatocellular carcinomas. Jpn J Clin Oncol. 25:1–4.
1995.PubMed/NCBI
|
|
11
|
Yoo NJ, Park SW and Lee SH: Mutational
analysis of tumour suppressor gene NF2 in common solid cancers and
acute leukaemias. Pathology. 44:29–32. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Huang J, Zhao YP, Li Q, Zhang JX, Yan W
and Zhang B: Association of single nucleotide polymorphisms of NBS1
gene with genetic susceptibility to primary liver cancer in a
Chinese Han population. Prog Biochem Biophys. 39:678–686. 2012.
View Article : Google Scholar
|
|
13
|
Wang Y, Hong Y, Li M, Long J, Zhao YP,
Zhang JX, Li Q, You H, Tong WM, Jia JD, et al: Mutation
inactivation of Nijmegen breakage syndrome gene (NBS1) in
hepatocellular carcinoma and intrahepatic cholangiocarcinoma. PLoS
One. 8:e824262013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Wang Y, Li M, Long J, Shi XY, Li Q, Chen
J, Tong WM, Jia JD and Huang J: Clinical significance of increased
expression of Nijmegen breakage syndrome gene (NBS1) in human
primary liver cancer. Hepatol Int. 8:250–259. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Hong Y, Long J, Li H, Chen S, Liu Q, Zhang
B, He X, Wang Y, Li H, Li Y, et al: An analysis of immunoreactive
signatures in early stage hepatocellular carcinoma. EBioMedicine.
2:438–446. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Nagaishi M, Kim YH, Mittelbronn M,
Giangaspero F, Paulus W, Brokinkel B, Vital A, Tanaka Y, Nakazato
Y, Legras-Lachuer C, et al: Amplification of the STOML3, FREM2, and
LHFP genes is associated with mesenchymal differentiation in
gliosarcoma. Am J Pathol. 180:1816–1823. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Lutchman M and Rouleau GA:
Neurofibromatosis type 2: A new mechanism of tumor suppression.
Trends Neurosci. 19:373–377. 1996. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Legoix P, Legrand MF, Ollagnon E, Lenoir
G, Thomas G and Zucman-Rossi J: Characterisation of 16 polymorphic
markers in the NF2 gene: Application to hemizygosity detection. Hum
Mutat. 13:290–293. 1999. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Sadetzki S, Flint-Richter P, Starinsky S,
Novikov I, Lerman Y, Goldman B and Friedman E: Genotyping of
patients with sporadic and radiation-associated meningiomas. Cancer
Epidemiol Biomarkers Prev. 14:969–976. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Gutmann DH, Giordano MJ, Fishback AS and
Guha A: Loss of merlin expression in sporadic meningiomas,
ependymomas and schwannomas. Neurology. 49:267–270. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Ikeda T, Hashimoto S, Fukushige S, Ohmori
H and Horii A: Comparative genomic hybridization and mutation
analyses of sporadic schwannomas. J Neurooncol. 72:225–230. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Lomas J, Bello MJ, Arjona D, Alonso ME,
Martinez-Glez V, Lopez-Marin I, Amiñoso C, de Campos JM, Isla A,
Vaquero J, et al: Genetic and epigenetic alteration of the NF2 gene
in sporadic meningiomas. Genes Chromosomes Cancer. 42:314–319.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Łaniewski-Wołłk M, Gos M, Koziarski A and
Szpecht-Potocka A: Identification of mutations in the NF2 gene in
Polish patients with neurofibromatosis type 2. J Appl Genet.
49:297–300. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Baser ME, Kuramoto L, Joe H, Friedman JM,
Wallace AJ, Gillespie JE, Ramsden RT and Evans DG:
Genotype-phenotype correlations for nervous system tumors in
neurofibromatosis 2: A population-based study. Am J Hum Genet.
75:231–239. 2004. View
Article : Google Scholar : PubMed/NCBI
|
|
25
|
Sheikh HA, Tometsko M, Niehouse L, Aldeeb
D, Swalsky P, Finkelstein S, Barnes EL and Hunt JL: Molecular
genotyping of medullary thyroid carcinoma can predict tumor
recurrence. Am J Surg Pathol. 28:101–106. 2004. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Ahronowitz I, Xin W, Kiely R, Sims K,
MacCollin M and Nunes FP: Mutational spectrum of the NF2 gene: A
meta-analysis of 12 years of research and diagnostic laboratory
findings. Hum Mutat. 28:1–12. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Evans DG, Huson SM, Donnai D, Neary W,
Blair V, Teare D, Newton V, Strachan T, Ramsden R and Harris R: A
genetic study of type 2 neurofibromatosis in the United Kingdom. I.
Prevalence, mutation rate, fitness, and confirmation of maternal
transmission effect on severity. J Med Genet. 29:841–846. 1992.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Bretscher A, Edwards K and Fehon RG: ERM
proteins and merlin: Integrators at the cell cortex. Nat Rev Mol
Cell Biol. 3:586–599. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
29
|
Cooper J and Giancotti FG: Molecular
insights into NF2/Merlin tumor suppressor function. FEBS Lett.
588:2743–2752. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Pećina-Šlaus N: Merlin, the NF2 gene
product. Pathol Oncol Res. 19:365–373. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Morrow KA, Das S, Metge BJ, Ye K, Mulekar
MS, Tucker JA, Samant RS and Shevde LA: Loss of tumor suppressor
Merlin in advanced breast cancer is due to post-translational
regulation. J Biol Chem. 286:40376–40385. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Horiguchi A, Zheng R, Shen R and Nanus DM:
Inactivation of the NF2 tumor suppressor protein merlin in DU145
prostate cancer cells. Prostate. 68:975–984. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Chen Z, Fadiel A and Xia Y: Functional
duality of merlin: A conundrum of proteome complexity. Med
Hypotheses. 67:1095–1098. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Eferl R and Wagner EF: AP-1: A
double-edged sword in tumorigenesis. Nat Rev Cancer. 3:859–868.
2003. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhang N, Bai H, David KK, Dong J, Zheng Y,
Cai J, Giovannini M, Liu P, Anders RA and Pan D: The Merlin/NF2
tumor suppressor functions through the YAP oncoprotein to regulate
tissue homeostasis in mammals. Dev Cell. 19:27–38. 2010. View Article : Google Scholar : PubMed/NCBI
|