Introduction
Melanoma is a highly malignant skin tumor derived
from melanocytes and its incidence has been rapidly increasing
worldwide (1). Melanoma is not
sensitive to radiotherapy, chemotherapy or biological immunotherapy
and metastasizes at an early clinical stage (2). Therefore, it is crucial to understand
the molecular mechanisms underlying the development of melanoma.
Although researchers worldwide have performed extensive research on
gene mutations, epigenetics, immune abnormalities and the tumor
microenvironment of melanoma, its precise mechanism remains poorly
understood. Studies on non-coding RNA, notably circular RNA
(circRNA), may provide a breakthrough for the explanation of the
mechanism of melanoma and the identification of early intervention
targets and biomarkers for early diagnosis.
Circular RNAs (circRNAs) have been reported as a
novel type of non-coding RNAs, which form covalently-closed
continuous loops without 5′ to 3′ polarity or polyadenylated tails
(3,4). They mainly arise from exons or introns
and are differentially generated by back-splicing or lariat-introns
(5). circRNAs are characterized by
stable structure, high abundance and tissue-specific expression.
These characteristics imply that circRNAs hold a great potential as
novel clinical diagnostic and prognostic markers and as new
treatment targets of diseases (6–8).
Increasing evidence has revealed that circRNAs specifically serve
as miRNA sponges, regulate alternative splicing and modulate the
expression of parental genes (9–11). It
has been proved that circRNA-CDR1 contains miR-7 binding sites and
exerts a negative regulatory effect in miR-7 as an endogenous miRNA
sponge and an upregulator of miR-7-targeted genes in the nervous
system (12,13). In esophageal squamous cell
carcinoma, cir-ITCH was downregulated in cancerous tissue and
functioned as a sponge of miR-7, miR-17 and miR-214. Therefore, it
enhanced the level of ITCH which negatively regulates the
Wnt/β-catenin signaling pathway (14).
To date, a study concerning the expression of
circRNAs in melanoma has not been reported. Therefore, in the
present study, we performed the profiling of circRNA expression in
normal melanocytes and melanoma cells with different invasive
abilities using circRNA microarray analysis. Furthermore, we
investigated differentially expressed circRNAs, as well as the
changes in melanoma cell proliferation and invasion after
circRNA-siRNA transfection. In addition, we analyzed the features
of the circRNA-expression profiling and predicted miRNAs
competitively binding to circRNAs. Finally, we provided important
experimental evidence for clarifying the occurrence, the
development, the invasion and metastasis of melanoma and for
exploring the non-coding RNA regulatory networks related to the
malignant phenotype of melanoma.
Materials and methods
Cell culture and reagents
The human melanocytes HM, the low-metastatic
melanoma cell line WM35 and the high-metastatic melanoma cell line
WM451 were purchased from the American Type Culture Collection
(ATCC; Manassas, VA, USA). The cells were incubated with Dulbecco's
modified Eagle's medium (DMEM; Gibco; Thermo Fisher Scientific,
Inc., Waltham, MA, USA) containing 10% fetal bovine serum (FBS) in
a 5% CO2 incubator at 37°C. The reagents in the present
study included an RNA-free extraction kit (Invitrogen Life
Technologies, Carlsbad, CA, USA), an Opti-MEM I Reduced-Serum
medium (Invitrogen Life Technologies), Transwell 6-well plates,
(8-µm pore size; Corning Incorporated, Corning, NY, USA), Matrigel
basement membrane (Becton, Dickinson and Company, Franklin Lakes,
NJ, USA), RNase-free glycogen in RNase water (Invitrogen Life
Technologies), Arraystar Human circRNA Array (8×15 K; Arraystar,
Rockville, MD, USA) including 5,396 circRNAs, GoldView dye (SBS
Genetech Co., Ltd., Shanghai, China), 2X PCR Master Mix (Arraystar)
and SuperScript™ III Reverse Transcriptase (Invitrogen
Life Technologies). The experiments were conducted by Kangchen
Biotech Co., Ltd. (Shanghai, China).
RNA extraction
Total RNA was extracted from the cells using the RNA
extraction kit according to the manufacturer's instructions and
examined by 1% agarose gel electrophoresis. Both the 28S and 18S
bands were clear and the brightness of the 28S band was roughly
two-fold higher than that of the 18S band, indicating the integrity
of total RNA. Ultraviolet spectrophotometry was used to determine
the concentration and purity of RNA. The D260/D280 nm ratio was
1.8–2.1 These results indicated that the purity of total RNA was
suitable for subsequent circRNA microarray analysis and
quantitative fluorescence PCR.
circRNA microarray
For analyzing the acquired array images, the Agilent
Feature Extraction software (version 11.0.1.1; Agilent
Technologies, Inc., Santa Clara, CA, USA) was performed. Following
the instructions of the Arraystar Super RNA Labeling kit
(Arraystar), total RNA of each sample was amplified with random
primers and reverse transcribed into fluorescent-labeled cRNA.
Subsequently, fluorescent-labeled cRNA was hybridized on an
Arraystar Human circRNA Array (8×15 K; Arraystar) and then
incubated in an Agilent Hybridization oven at 65°C for 17 h. After
washing, the samples were scanned on an Agilent scanner
(G2505C).
Data collection and analysis of
circRNA microarray
Fluorescence intensity was scanned in a microarray
scanner and loaded to the Agilent Feature Extraction software to
read and analyze the original data. The R software package (limma
package, Cytoscape http://www.cytoscape.org/ 3.3.2, R environment
https://www.r-project.org/ reversion
3.42) was used for quantile normalization and subsequent data
processing. Differentially expressed circRNAs between groups was
screened by fold change and P-value. A value of P<0.05 was
considered to indicate a statistically significant difference.
Cluster analysis revealed differential circRNA expression using
Cluster 3.0 software (R programming language package: Gplots).
Prediction for circRNA-miRNA-mRNA
network
The circRNA-microRNA interaction was predicted using
Arraystar miRNA target-prediction software (Arraystar) and GCBI
(https://www.gcbi.com.cn/gclib/html/index) based on
TargetScan and miRanda software packages. A site with a higher
matching score was noted. The map of circRNA-miRNA interaction
network was illustrated using Cytoscape 3.01 (Cytoscape Consortium
San Diego, CA, USA).
Real-time quantitative fluorescence
PCR
Five differentially expressed circRNAs were verified
by real-time quantitative fluorescence PCR. Reverse transcription
reaction was carried out according to the instructions of the
SuperScript™ III Reverse Transcriptase reverse
transcription kit (Invitrogen Life Technologies). The
ViiA™ 7 Real-Time PCR system was used for quantitative
detection of PCR. The PCR primers were designed and synthesized by
Invitrogen Biotechnology Co., Ltd. (Shanghai, China). The primer
sequences and related information are listed in Table I. GAPDH served as an internal
standard for normalization. The PCR conditions were as follows:
95°C for 10 min, 95°C for 10 sec and 60°C for 60 sec, for a total
of 40 cycles. The 2−ΔΔCt value refers to the relative
expression level of circRNAs. The experiments were performed in
triplicate.
 | Table I.The primers for qRT-PCR. |
Table I.
The primers for qRT-PCR.
| Gene | Primer
sequence | Length (bp) |
|---|
| GAPDH | F:
5′GGGAAACTGTGGCGTGAT3′ |
|
|
| R:
5′GAGTGGGTGTCGCTGTTGA3′ | 299 |
| circRNA0000082 | F:
5′CGGATTAGAAACCTCGACACC3′ |
|
|
| R:
5′TGGACCACAGGAGCATCATT3′ | 270 |
| circRNA0016418 | F:
5′CTCCGACCCAAGTGAGAAGC3′ |
|
|
| R:
5′CAGCCTGTAGTTTGGGACC3′ | 124 |
| circRNA0023988 | F:
5′TGGTGGTGGTGCTATTCCTC3′ |
|
|
| R:
5′TCTCCTGGTTCTCCTGCTTG3′ | 265 |
| circRNA0008157 | F:
5′GAATTCTAAGCAGCACAACATCA3′ |
|
|
| R:
5′GGGTCCATGTCTTTGCCTCT3′ | 161 |
| circRNA0030388 | F:
5′TGGGACATCCATCAGATAAGAA3′ |
|
|
| R:
5′CTGTAGTGGGAGGCAGTGTTT3′ | 169 |
Transfection
The cells were transfected with five differentially
expressed circRNAs which were verified using qRT-PCR as follows: 24
h before transfection, 5×104 cells were seeded in 2 ml
of basal medium containing FBS. The cells were ready for
transfection when they reached 70% confluency. Lipofectamine™ 2000
(5 µl) was diluted in 250 µl of Opti-MEM® I, mixed
gently and incubated at room temperature for 5 min, and then 7.5 µl
siRNA-circRNAs was diluted in 250 µl of Opti-MEM® I,
mixed gently and incubated at room temperature for 5 min. Diluted
siRNA-circRNAs and Lipofectamine™ 2000 were mixed together gently
and the complexes were incubated at room temperature for 20 min.
The siRNA-circRNAs-Lipofectamine™ 2000 transfection complexes were
added in 6-well plates and mixed by gently rocking the plate back
and forth. The plates were incubated at 37°C in a CO2
incubator for 24–72 h. The transfected cells were observed under a
fluorescence microscope (CKX53; Olympus, Tokyo, Japan) to assess
the transfection efficiency.
Cell proliferation assay
After transfection, the cell density in each group
was adjusted to 1×105 cells/ml and the cells were seeded
in 96-well plates in a volume of 100 µl per well, and placed at
37°C in a CO2 incubator for 24 h. The WM35 cells were
divided into four groups: i) negative control group, ii)
si-circ0023988, iii) si-circ0008157 and iv) si-circ0030388 treated
group. The WM451 cells were divided into three groups: i) negative
control, ii) si-circ0000082 and iii) si-circ0016418 group. Methyl
thiazolyl tetrazolium (MTT) (50 µl, 1 mg/ml) was added to each well
and the plates were incubated for 4 h at 37°C, until MTT was
reduced to formazan. The supernatant was aspirated and 150 µl DMSO
was added to each well, and then the plates were placed on a
horizontal shaker and shaked for 10 min to dissolve the formazan.
The optical density (OD) value of each well was detected using an
enzyme-linked immunosorbent assay (ELISA) reader at 570 nm. Cells
without treatment served as a control and the blank well was used
for zero adjustment. The cell survival rate were calculated using
the following formula: Cell survival rate = (OD value of the
experiment group/OD value of the control group) × 100%.
Cell invasion assay
The upper chamber surface of the basement membrane
of the Transwell chambers was coated with 50 mg/l Matrigel 1:8
dilution and air dried at 4°C. Diluted Matrigel (60–80 µl, 3.9
µg/µl) was added to the polycarbonate membrane in the upper chamber
and placed at 37°C for 30 min to allow the Matrigel to polymerize.
The cells were digested and after terminating the digestion, the
cells were centrifuged and then the culture medium was discarded.
After washing with PBS for 1–2 times, the cells were resuspended in
serum-free media containing bovine serum albumin (BSA). The cell
density was adjusted to 5×104/ml. One milliliter of
medium-containing FBS was added to the lower chambers of 6-well
plates, cell supernatant was added to the upper chambers and after
72 and 24-h incubation for the WM451 and the WM35 cells
respectively, the membranes were removed. The cells on the Matrigel
and upper chambers were removed with a cotton swab, the membrane
was removed and fixed with 95% alcohol for 15–20 min and stained
with hematoxylin for 10 min. The cells were counted and imaged
under an inverted microscope (CKX53; Olympus).
Statistical analysis
The data are presented as the mean ± SD as analyzed
using SPSS 13.0 (SPSS, Inc., Chicago, IL, USA) and deemed to have
statistical significance at P<0.05 using the Student's t-test,
one-way analysis of variance (ANOVA) or χ2 test.
Results
Differentially expressed circRNAs
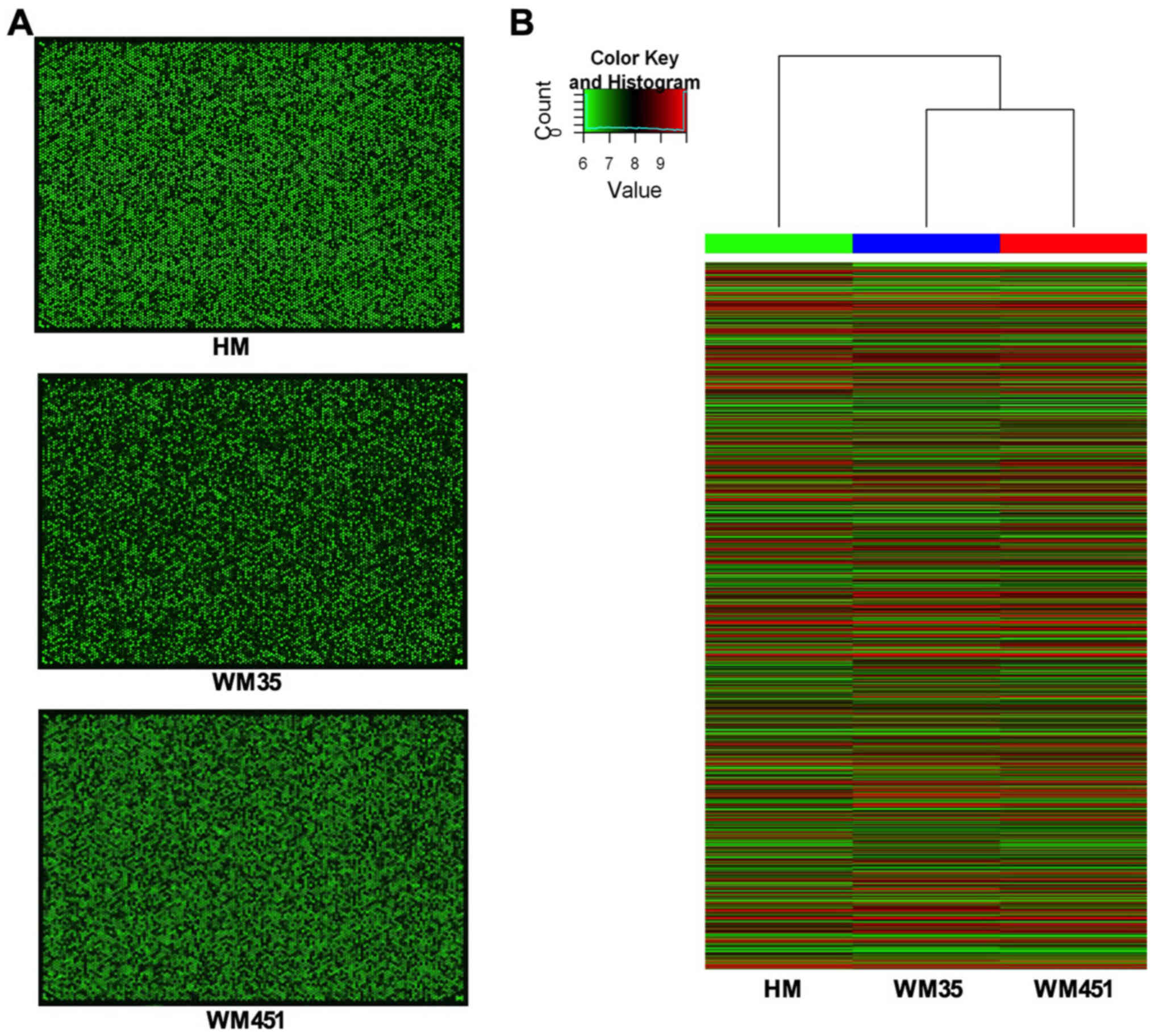
Genome scan (Fig.
1A) and cluster analysis (Fig.
1B) were used to identify abnormal expression of circRNAs among
the HM, WM35 and WM451 cells. Approximately 5,396 circRNAs were
detected using the Arraystar Human circRNA Array data analysis. The
results of the heat maps are displayed in Fig. 1B. The data obtained from circRNA
microarray analysis were normalized. Approximately 5,396 circRNAs
were detected by the Arraystar Human circRNA Array data. The
results of the genome scans and differentially expressed circRNAs
among the HM, WM35 and WM451 cells are displayed in Fig. 1. Data obtained from circRNA
microarray analysis were normalized. circRNAs with expression
levels that increased 2-fold were upregulated and circRNAs with
expression levels that decreased 2-fold were downregulated.
Different fold changes of circRNA expression according to chip
analyses are summarized in Table
II. CircRNA microarray analysis results demonstrated that
compared with HM cells, 797 circRNAs were upregulated and 969
circRNAs were downregulated in WM35 cells, while 307 circRNAs were
upregulated and 312 circRNAs were downregulated in WM451 cells.
Only five circRNAs (circ0001056, circ0000082, circ0016418,
circ0008602 and circ0033496) were upregulated as well as four
circRNAs (circ0023988, circ0008157, circ0030388 and circ0023990)
were downregulated both in WM35 and WM451 cells. Compared with the
WM35 cells, 977 circRNAs were upregulated and 757 circRNAs were
downregulated in the WM451 cells (Table II). In WM35 vs. HM cells, WM451 vs.
HM cells and WM451 vs. WM35 cells, four circRNAs (circ0023988,
circ0023990, circ0008157 and circ0030388) were all downregulated
(Table III).
 | Table II.Differential expression of circRNA
according to chip analyses. |
Table II.
Differential expression of circRNA
according to chip analyses.
| A, Upregulated
circRNA |
|---|
|
|---|
| Fold change | WM35 vs. HM | WM451 vs. HM | WM451 vs. WM35 |
|---|
| >20 | 0 | 0 | 2 |
| 10–20 | 0 | 5 | 32 |
| 5–10 | 2 | 26 | 126 |
| 2–5 | 795 | 276 | 817 |
|
| B, Downregulated
circRNA |
|
| >20 | 1 | 5 | 1 |
| 10–20 | 10 | 8 | 0 |
| 5–10 | 89 | 20 | 13 |
| 2–5 | 869 | 279 | 743 |
 | Table III.Differential expression of circRNAs
according to chip analyses. |
Table III.
Differential expression of circRNAs
according to chip analyses.
| A, Upregulated |
|---|
|
|---|
|
| Fold change |
|
|---|
|
|
|
|
|---|
| circRNA | WM35 vs. HM |
| WM451 vs. HM | miRNA binding
sites |
|---|
| circ0001056 | 3.45 |
| 2.13 | miR-30b-3p,
miR-619-5p, miR-665, miR-153-5p, miR-149-5p |
| circ0000082 | 2.39 |
| 4.65 | miR-30b-3p,
miR-23a-5p, miR-143-5p, miR-106b-3p, miR-508-5p |
| circ0016418 | 2.63 |
| 3.63 | miR-214-5p,
miR-153-5p, miR-657, miR-450a-2-3p, miR-605-3p |
| circ0008602 | 2.69 |
| 3.28 | miR-544a,
miR-200c-3p, miR-148a-3p, miR-362-5p, miR-17-3p |
| circ0033496 | 2.031 |
| 2.15 | miR-612,
miR-502-5p, miR-1264, miR-105-3p, miR-423-5p |
|
| B,
Downregulated |
|
| Fold
change |
|
|
|
|
|
| circRNA | WM35 vs.
HM | WM451 vs.
HM | WM451 vs.
WM35 | miRNA binding
sites |
|
| circ0023988 | –2.08 | –5.64 | –2.71 | miR-655-5p,
miR-448, miR-485-5p, miR-613, miR-103a-3p |
| circ0023990 | –2.55 | –10.12 | –3.96 | miR-485-5p,
miR-613, miR-339-5p, miR-329-5p, miR-873-5p |
| circ0008157 | –2.38 | –4.92 | –2.06 | miR-136-3p,
miR-365b-5p, miR-365a-5p, miR-329-5p, miR-335-3p |
| circ0030388 | –2.39 | –17.82 | –7.45 | miR-449b-5p,
miR-449a, miR-34a-5p, miR-34c-5p, miR-200a-3p |
Differential expression of circRNAs
verified by real-time quantitative fluorescence PCR
Five upregulated circRNAs and four downregulated
circRNAs were found by microarray analysis and qRT-PCR was used to
verify the background expression levels of the above-mentioned
circRNAs in melanoma cells. GAPDH served as an internal standard
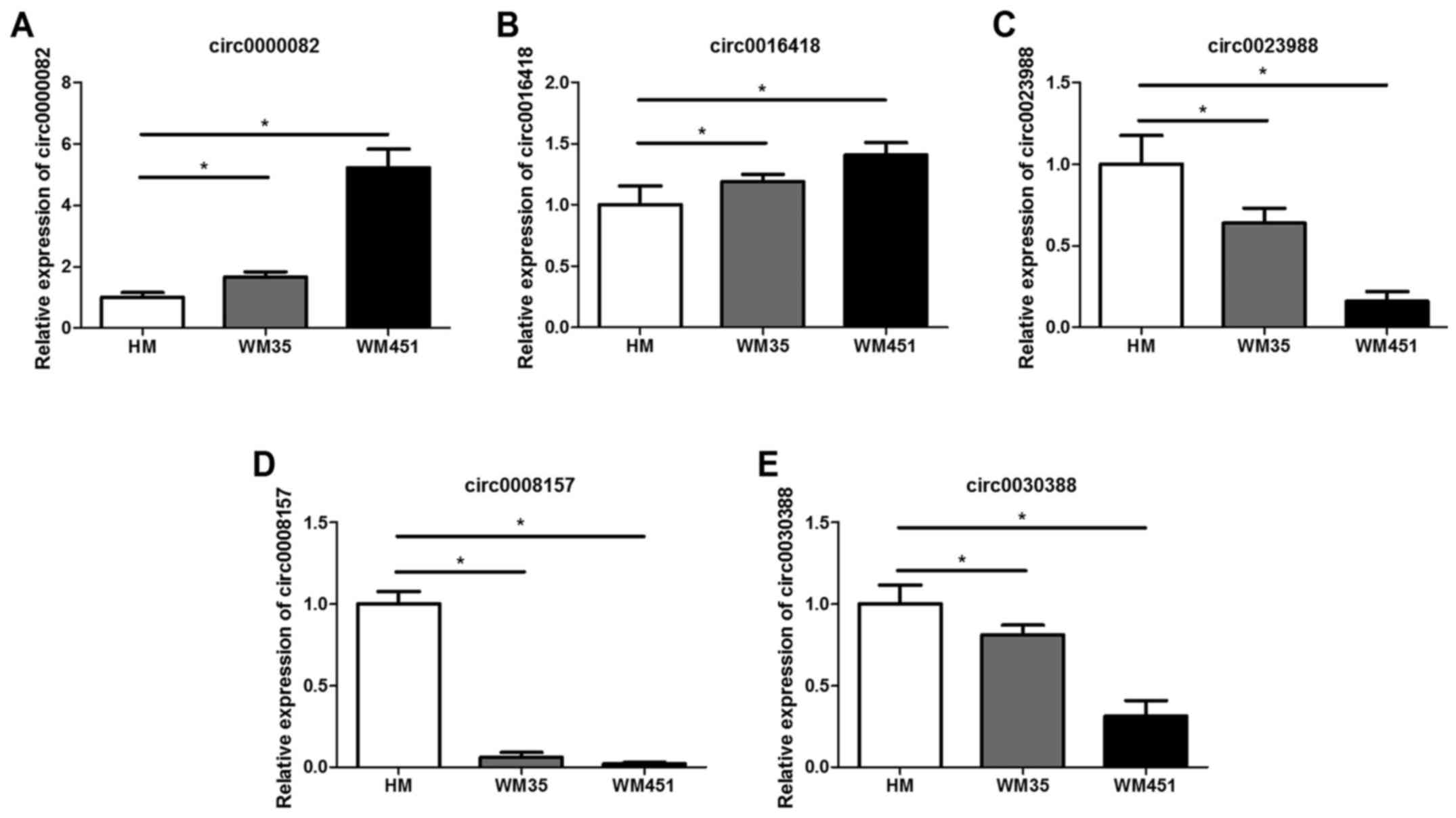
for normalization. The results revealed that circ0000082 and
circ0016418 had higher expression levels and circ0023988,
circ0008157 and circ0030388 had lower expression levels both in the
WM35 and the WM451 cells, compared with the HM cells (Fig. 2). The above-mentioned five circRNAs
were verified by real-time quantitative fluorescence PCR and the
PCR results were generally consistent with those of the circRNA
microarray, which confirmed that the microarray analysis results
were reliable (Table IV). The
other four circRNAs with low basal expression were excluded
(circ0001056, circ0033496, circ0008602 and circ0023990).
 | Table IV.Differential expression of circRNAs
according to qRT-PCR. |
Table IV.
Differential expression of circRNAs
according to qRT-PCR.
|
| Fold change |
|
|---|
|
|
|
|
|---|
| circRNA | WM35 vs. HM | WM451 vs. HM | WM451 vs. WM35 | Expression |
|---|
| circ0000082 | 1.67 | 5.23 | 3.31 | Up |
| circ0016418 | 1.19 | 1.41 | 1.19 | Up |
| circ0023988 | 0.64 | 0.16 | 0.24 | Down |
| circ0008157 | 0.056 | 0.02 | 0.268 | Down |
| circ0030388 | 0.816 | 0.31 | 0.384 | Down |
Knockdown of the expression levels of
circRNAs
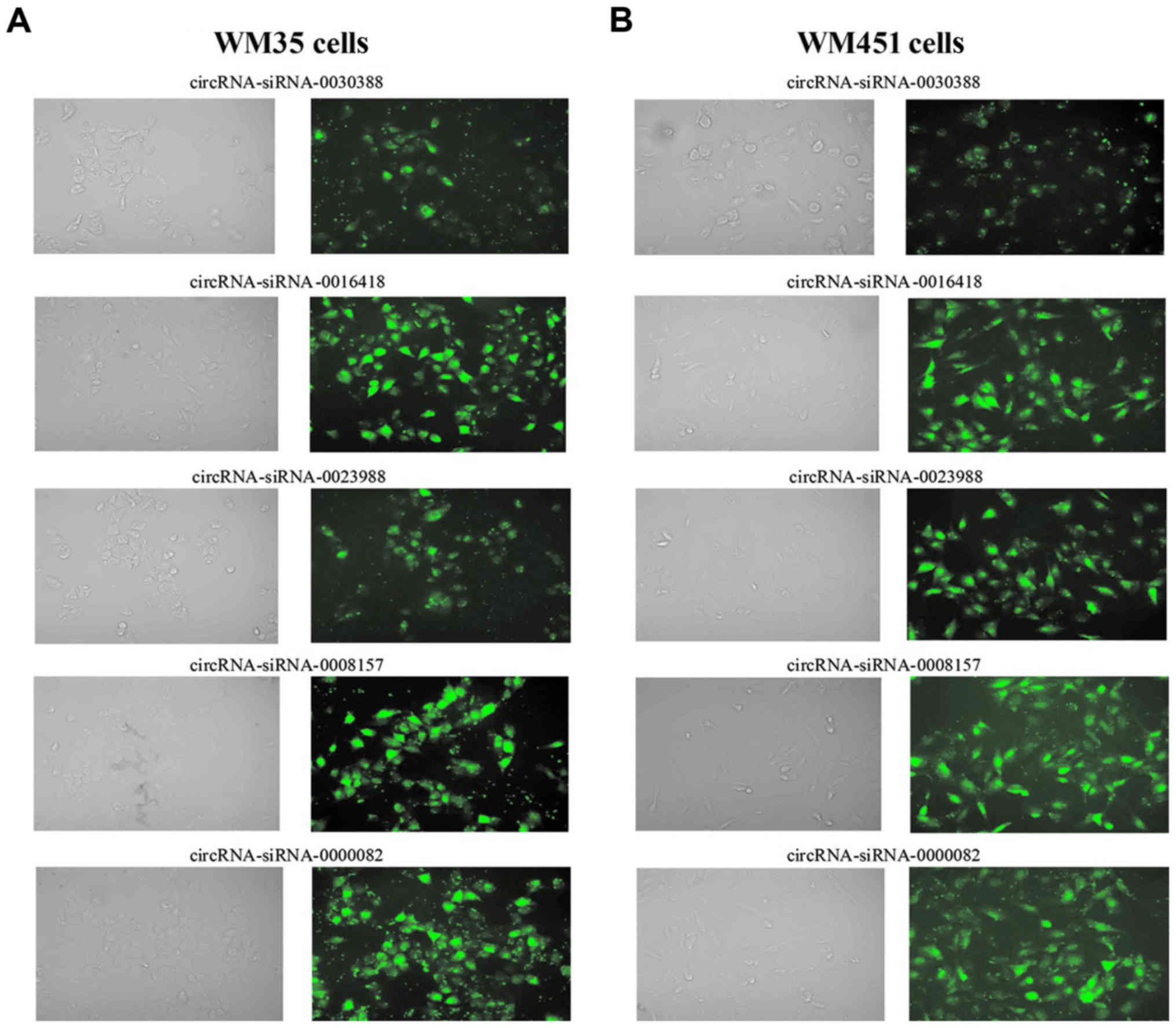
In order to interfere with circRNA expression
levels, circ0023988, circ0008157, or circ0030388 siRNAs were
transfected into WM35 cells, while circ0000082 or circ0016418 siRNA
swere transfected into WM451 cells. The highest transfection
efficiency was observed at 24 h and the transfection efficiency was
>80% by comparing the cells in bright field and fluorescent
views of the same field (Fig. 3).
Therefore, these results indicated that the transfection efficiency
was satisfactory and the cells could be used for the following
experiments.
Changes in cell proliferation after
siRNA-circRNAs transfection
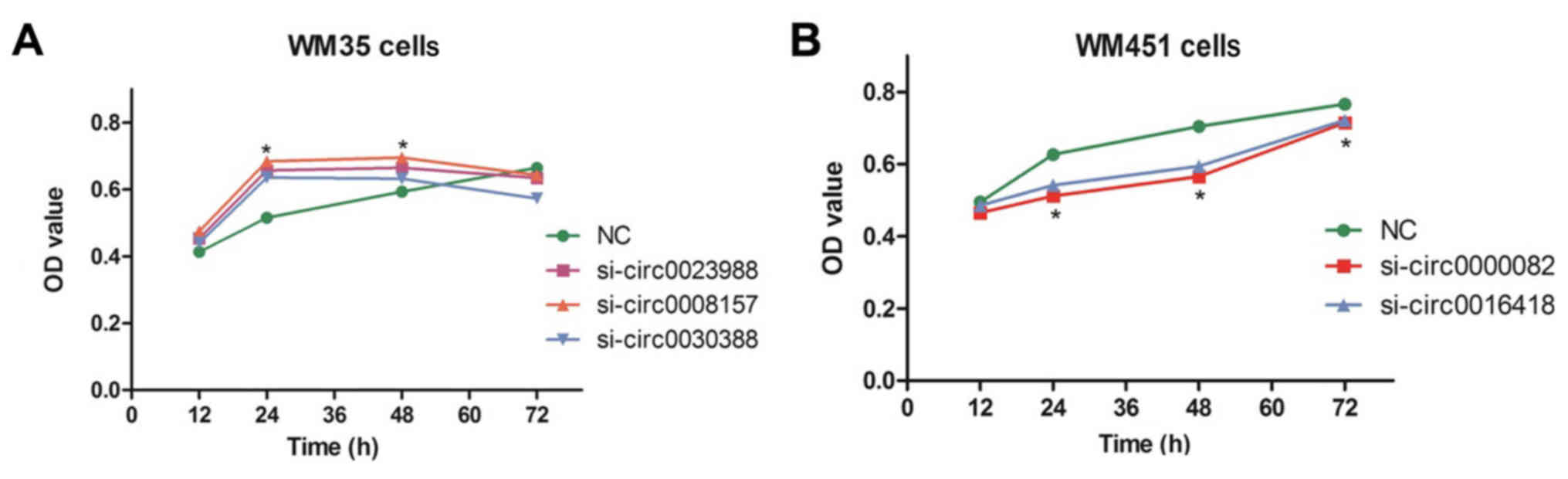
Twenty-four hours after the WM35 and WM451 cells
were transfected with siRNA-circRNAs, the cell proliferation in
each group was detected by an MTT assay. The results revealed that
at 24 and 48 h after transfection, the OD values of low-metastatic
melanoma WM35 cells in the si-circ0023988, si-circ0008157 and
si-circ0030388 groups were significantly higher than that in the
negative control group (P<0.05), indicating that silencing of
circ0023988, circ0008157 and circ0030388 promoted the proliferation
of the WM35 cells (Fig. 4A). At 24,
48 and 72 h after transfection, the OD values of high-metastatic
melanoma WM451 cells in the si-circ0000082 and si-circ0016418
groups were significantly lower than that in the negative control
group (P<0.05), indicating that silencing of circ0000082 and
circ0016418 inhibited the proliferation of the WM451 cells
(Fig. 4B).
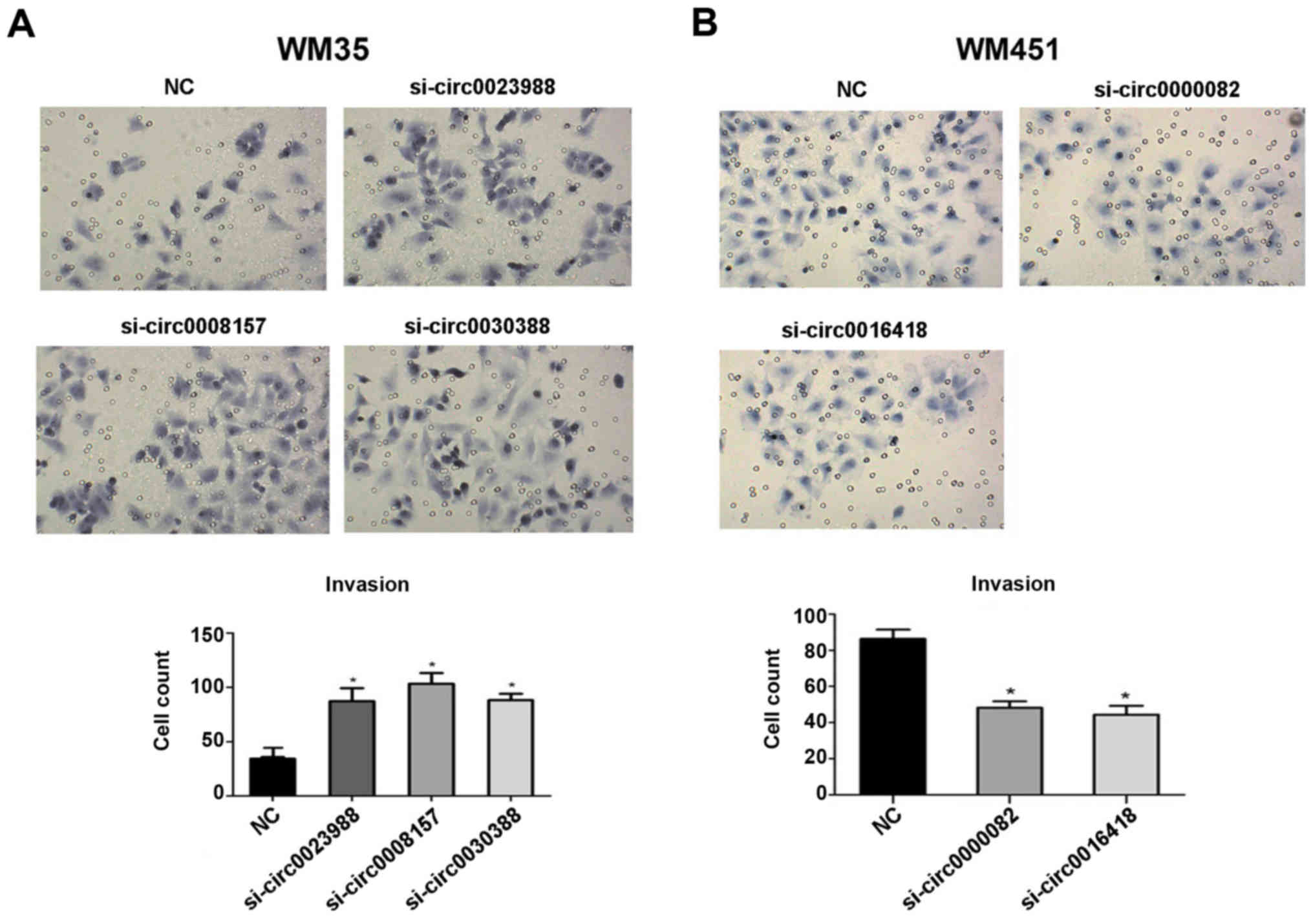
Changes in cell invasion after
siRNA-circRNA transfection
The transfected WM35 and WM451 cells were subjected
to invasion assays for 24 and 72 h, respectively. The cells that
passed through the basement membrane were counted by light
microscope (CKX53; Olympus) at ×200 magnification. The cell numbers
in the si-circ0023988, si-circ0008157 and si-circ0030388 groups
were 87.3±12,103.3±10 and 88.3±5.7, respectively, which were
significantly higher than these in the negative control group
(34.3±1.5) (P<0.05, Fig. 5A),
indicating that the invasion ability of low-metastatic melanoma
WM35 cells was enhanced by silencing of circ0023988, circ0008157
and circ0030388. Compared with the negative control group, the
WM451 cell numbers in the si-circ0000082 and si-circ0016418 groups
were lower (48.3±3.5, 44.3±5 vs. 86.3±5.1) (P<0.05, Fig. 5B). These results indicated that
silencing of circ0000082 and circ0016418 decreased the invasion
capability of high-metastatic melanoma WM451 cells and that
circ0000082 and circ0016418 have a promoting effect on the invasion
of WM451 cells.
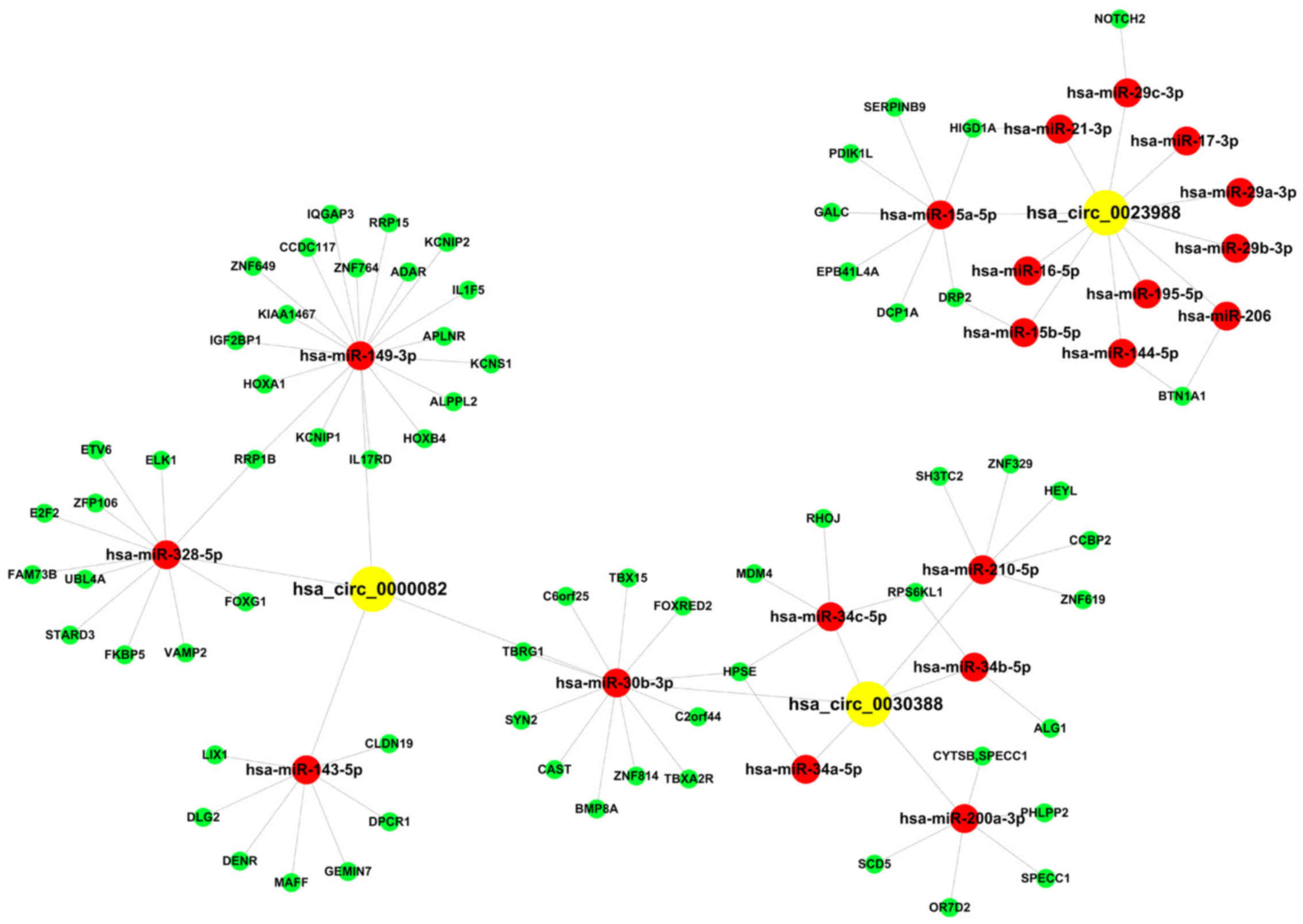
Competing endogenous RNA (ceRNA)
analysis of circ0000082, circ0023988 and circ0008157
In accordance with the results of real-time
quantitative fluorescence PCR, ceRNAs of apparently differentially
expressed circ0000082, circ0023988 and circ0008157 were found by
mutually targeted MRE enrichment analysis (MuTaME, https://cm.jefferson.edu/rna22/Precomputed/) and then
a regulatory network of circRNA-miRNA-mRNA was constructed using
Cytoscape 3.01 (Fig. 6). Prediction
results (Table V) revealed that
circ0000082 possibly competitively bound to 64 miRNAs and impacted
the expression of 425 target genes. Furthermore, circ0023988
possibly competitively bound to 78 miRNAs and impacted the
expression of 33 target genes. In addition, circ0008157 possibly
competitively bound to 111 miRNAs and impacted the expression of 52
target genes. Subsequently, combining with the confirmed
melanoma-related miRNAs, we selected miRNAs that impacted the
occurrence and development of melanoma, competitively binding to
circ0000082, circ0023988 and circ0030388 and the possible impacted
target genes (Fig. 6). The results
are displayed in Table VI.
 | Table V.Predicted ceRNAs for circRNAs. |
Table V.
Predicted ceRNAs for circRNAs.
| circRNA | Sponge miRNA
numbers | Target gene
numbers |
|---|
| circ0000082 | 64 | 425 |
| circ0023988 | 78 | 33 |
| circ0030388 | 111 | 52 |
 | Table VI.Predicted circRNA-miRNA-mRNA network
in melanoma. |
Table VI.
Predicted circRNA-miRNA-mRNA network
in melanoma.
| circRNA | miRNA | Target gene |
|---|
| circRNA0000082 | miR-30b-3p | BMP8A, C2orf44,
C6orf25, CAST, FOXRED2, HPSE, SYN2, TBRG1, TBX15, TBXA2R,
ZNF814 |
|
|
miR-143-5p[44] | CLDN19, DENR, DLG2,
DPCR1, GEMIN7, LIX1, MAFF |
|
|
miR-149-3p[45] | ADAR, ALPPL2,
APLNR, CCDC117, HOXA1, HOXB4, IGF2BP1, IL1F5, IL17RD, IQGAP3,
KCNIP2, KCNS1, KCNIP1, KIAA1467, RRP15, RRP1B, ZNF649, ZNF764 |
|
|
miR-328-5p[46] | ELK1, ETV6, E2F2,
FAM73B, FKBP5, FOXG1, RRP1B, STARD3, UBL4A, VAMP2, ZFP106 |
| circRNA0030388 | miR-30b-3p | BMP8A, CAST,
C2orf44, C6orf25, FOXRED2, HPSE, SYN2, TBRG1, TBX15, TBXA2R,
ZNF814 |
|
|
miR-34a-5p[47] | HPSE |
|
| miR-34b-5p | ALG1, RPS6KL1 |
|
| miR-34c-5p | HPSE, MDM4, RHOJ,
RPS6KL1 |
|
|
miR-200a-3p[49] | CYTSB, OR7D2,
PHLPP2, SCD5, SPECC1 |
|
| miR-210-5p | CCBP2, HEYL,
SH3TC2, ZNF329, ZNF619 |
| circRNA0023988 | miR-15a-5p | DCP1A, DRP2,
EPB41L4A, GALC, HIGD1A, PDIK1L, SERPINB9 |
|
|
miR-15b-5p[50] |
|
|
| miR-16-5p |
|
|
| miR-17-3p |
|
|
|
miR-21-3p[51] | HIGD1A |
|
|
miR-29a-3p[53] |
|
|
|
miR-29b-3p[53] |
|
|
|
miR-29c-3p[53] | NOTCH2 |
|
|
miR-144-5p[55] | BTN1A1 |
|
|
miR-195-5p[54] |
|
|
| miR-206 |
|
Discussion
CircRNAs are widely expressed in human cells and
their expression levels can be 10-fold or higher compared to their
linear isomers. Compared with traditional linear RNA (containing 5′
and 3′ends), circRNAs form a covalently-closed continuous loop and
cannot be affected by RNA exonuclease. Their expression is more
stable and circRNAs cannot be easily degraded. Thus, circRNAs have
significant superiority in the development and application of new
clinical diagnostic markers (15–17).
Abnormal expression of circRNAs is observed in many cancers, such
as the expression of circ104912 in laryngeal squamous cell cancer
(18), of circ0001649 in
hepatocellular carcinoma (19) and
circ001988 in colorectal cancer (20). Most circRNAs play a regulatory role
at the transcriptional and post-transcriptional levels and only a
few function at the transcriptional level.
Little is known about the specific biological
functions of circRNAs. Several recent breakthrough studies
confirmed that some circRNAs could be used as competing endogenous
RNAs (ceRNAs) to exert a regulatory effect on gene expression.
CircRNA molecules enrich miRNA binding sites (21), which function as miRNA sponges
(22) and play an important
regulatory role in disease by interacting with disease-related
miRNAs. Hansen et al (12)
first found that circRNA-CDR1as (also called ciRS-7) abundantly
expressed in human brain tissue, is antisense to the cerebellar
degeneration-related protein 1 transcript (CDR1as). CDR1as contains
70 miR-7 binding sites and plays a negative regulatory role in
miR-7 as an endogenous miRNA sponge. miR-7 is an important
regulatory factor for a variety of cancer-related pathways
(23–26). Thus, ciRS-7 is likely to be an
important regulatory factor in the development of nervous system
disease and cancer (27,28). In colorectal cancers, circ001569 has
been identified as a sponge of miR-145, which can inhibit tumor
development through the upregulation of the expression of miR-145
target genes E2F5, BAG4 and FMNL2 (29). In bladder carcinomas, circTCF25 acts
as a sponge for miR-103a and miR-107 which downregulate the
expression of CDK6 and promote cell proliferation and migration
(30). The above-mentioned studies
indicate that circRNAs also function as ceRNAs, reveal a new
regulation mode of circRNA on target genes by competitively binding
to miRNA and reveal that circRNAs participate in various
physiological and pathological processes.
The present study demonstrated that hundreds of
circRNAs are differentially expressed in melanoma cell lines and
normal melanocytes. Further comprehensive analysis revealed that
the expression of circ0001056, circ0000082, circ0016418,
circ0008602 and circ0033496 was upregulated in the WM35 and WM451
cells, indicating that they may play a similar role as
cancer-promoting genes in the occurrence and development of
melanoma. In WM35 vs. HM cells, WM451 vs. HM cells and WM451 vs.
WM35 cells, the expression of circ0023988, circ0023990, circ0008157
and circ0030388 was downregulated. In combination with numerous
previous studies (31–43), our prediction results of the miRNA
binding sites on the above-mentioned circRNAs using Arraystar miRNA
target prediction software and GCBI (https://www.gcbi.com.cn/gclib/html/index) based on
TargetScan and miRanda software revealed that miR-143-5p, miR-449a,
miR-657, miR-200c-3p and miR-423-5p were abnormally expressed in a
variety of tumor tissues (colorectal cancer, lung adenocarcinoma,
breast, liver and ovarian cancer). The related target genes are
involved in proliferation, apoptosis and cell cycle of various
malignant tumor cells by participating in the Wnt/β-catenin, Notch,
PI3K/AKT, NF-κB, p53, autophagy, angiogenesis and other signaling
pathways. They produce a marked effect on tumorigenesis, invasion,
metastasis and sensitivity to radiotherapy and chemotherapy
(Table VII). Based on the results
of qRT-PCR, significantly differentially-expressed circRNAs were
silenced using siRNA interference, and then transfected into WM35
and WM45 cells. Compared with the WM451 cells, the proliferation
and invasion of WM35 cells were promoted after the upregulated
circRNAs (circ0023988, circ0008157 and circ0030388) in
low-metastatic melanoma WM35 cells were silenced. Compared with the
WM35 cells, the silencing of the upregulated circRNAs (circ0000082
and circ0016418) in high-metastatic melanoma WM451 cells inhibited
the proliferation and invasion of WM451 cells, confirming that
circ0030388, circ0016418, circ0023988, circ0008157 and circ0000082
were closely related to the proliferation and invasion of melanoma
cells. The ceRNAs of circ0000082, circ0023988 and circ0008157 were
further analyzed using MuTaMe. Many predicted miRNAs and target
genes have been verified to be strongly associated with the
proliferation, invasion and metastasis of melanoma in previous
studies (44–55) (Table
VI). Although, the present study has identified the abnormally
expressed circRNAs in MM cell lines, confirmed the adverse or
positive effect on tumor development of five specific circRNAs and
predicted the potential biological functions by previously reported
circRNA-miRNA-mRNA networks, several limitations still exist. Only
cell viability and invasion abilities of the identified circRNAs
were evaluated, therefore it would be better to investigate other
deeper functions in a future study, such as the regulation on cell
material and energy metabolism, tumor metastasis or immune
function. In addition, for the predicted circRNA targets, including
miRNAs and mRNAs, more research is warranted to clarify in detail
the underlying mechanisms of such circRNAs. Subsequently, on the
basis of predicted miRNA, we will combine a microarray of mRNA
sequencing before and after the predicted circRNA silencing and
attempt to examine changes of the target genes. We may reveal the
exact regulatory mechanisms of the circRNA-miRNA-mRNAs network
which influence the occurrence and development of melanoma.
Finally, we will analyze the effects of differentially expressed
circRNAs combined miRNAs and target genes on melanoma according to
the pathological type and prognosis of melanoma patients.
 | Table VII.Functions of potential miRNAs target
circRNAs. |
Table VII.
Functions of potential miRNAs target
circRNAs.
| circRNA | miRNA | Cancer | Target gene | Function |
|---|
| circRNA0001056 | miR-665 | GSRCC |
| Invasion,
metastasis, chemosensitivity |
| circRNA0000082 | miR-508-5p | gastric cancer |
ABCB1[38],
ZNRD1[38] |
chemosensitivity |
|
| miR-143-5p | CRC, gastric
cancer | COX-2 | Invasion and
metastasis |
| circRNA0016418 | miR-214-5p | HCC |
|
|
|
| miR-657 | HCC, laryngeal
carcinoma |
TLE1[39] | Proliferation |
| circRNA0008602 | miR-362-5p | HCC | CYLD | Proliferation,
clone, invasion and metastasis |
|
| miR-544a | NSCLC | CDH1 | Proliferation,
clone, invasion and metastasis |
|
| miR-200c-3p | CRC, ovarian
cancer |
|
|
|
| miR-17 | PCa, CRC, HCC,
glioblastoma[41] | TIMP3,
PTEN[40] GalNT7[40], DNA-PK MDM2 | Proliferation,
invasion and metastasis; cell cycle, angiogenesis; radiation and
chemotherapy sensitivity |
| circRNA0033496 | miR-612 | HCC |
AKT2[43] | Proliferation,
invasion and metastasis |
|
| miR-502-5p | CRC, breast
cancer |
Rab1B[42], TRAF2 | Proliferation, cell
cycle, apoptosis |
|
| miR-423-5p | PC, CRC, HCC,
gastric cancer | β-catenin,
TFF1 | Proliferation,
invasion and metastasis; cell cycle; apoptosis;
chemosensitivity |
| circRNA0023990 | miR-339-5p | NSCLC, CRC, breast
cancer |
BCL-6[32], PRL-1[31]
MDM2[33] | Proliferation,
invasion and metastasis |
| circRNA0030388 | miR-449a | NSCLC, MM, gastric
cancer, ovarian cancer, glioblastoma, endometrial cancer | c-MET lncRNA NEAT1
E2F3, NOTCH1[36] CDK6, MAZ[37] | Proliferation, cell
cycle; apoptosis; chemotherapy sensitivity |
|
| miR-34c-5p | CRC, PCa,
glioblastoma, NPC, endometrial cancer |
Notch[34], Bmf c-myc, E2F3
MAPT | cell cycle;
apoptosis; radiation and chemotherapy sensitivity |
|
| miR-200a-3p | ovarian cancer,
glioblastoma |
MGMT[35] | Chemotherapy
sensitivity |
Acknowledgements
The present study was supported by grants from the
National Natural Science Foundation of China (nos. 8130168,
81272192 and 81171882) and the Hunan Natural Science Foundation
(no. 2015JJ4053).
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Egan KM, Seddon JM, Glynn RJ, Gragoudas ES
and Albert DM: Epidemiologic aspects of uveal melanoma. Surv
Ophthalmol. 32:239–251. 1988. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Kam R, Hayungs J, Bomfeld N, Sauerwein W,
Höffken K and Seeber S: Prognosis and treatment of disseminated
uveal melanoma. Cancer. 72:2219–2223. 1993. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Qu S, Yang X, Li X, Wang J, Gao Y, Shang
R, Sun W, Dou K and Li H: Circular RNA: A new star of noncoding
RNAs. Cancer Lett. 365:141–148. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Chen LL: The biogenesis and emerging roles
of circular RNAs. Nat Rev Mol Cell Biol. 17:205–211. 2016.
View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Li J, Yang J, Zhou P, Le Y, Zhou C, Wang
S, Xu D, Lin HK and Gong Z: Circular RNAs in cancer: Novel insights
into origins, properties, functions and implications. Am J Cancer
Res. 5:472–480. 2015.PubMed/NCBI
|
|
6
|
Wang F, Nazarali AJ and Ji S: Circular
RNAs as potential biomarkers for cancer diagnosis and therapy. Am J
Cancer Res. 6:1167–1176. 2016.PubMed/NCBI
|
|
7
|
Shao Y and Chen Y: Roles of circular RNAs
in neurologic disease. Front Mol Neurosci. 9:252016. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Lu D and Xu AD: Mini review: Circular RNAs
as potential clinical biomarkers for disorders in the central
nervous system. Front Genet. 7:532016. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Li Z, Huang C, Bao C, Chen L, Lin M, Wang
X, Zhong G, Yu B, Hu W, Dai L, et al: Exon-intron circular RNAs
regulate transcription in the nucleus. Nat Struct Mol Biol.
22:256–264. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Ashwal-Fluss R, Meyer M, Pamudurti NR,
Ivanov A, Bartok O, Hanan M, Evantal N, Memczak S, Rajewsky N and
Kadener S: circRNA biogenesis competes with pre-mRNA splicing. Mol
Cell. 56:55–66. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Hansen TB, Kjems J and Damgaard CK:
Circular RNA and miR-7 in cancer. Cancer Res. 73:5609–5612. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Memczak S, Jens M, Elefsinioti A, Torti F,
Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer
M, et al: Circular RNAs are a large class of animal RNAs with
regulatory potency. Nature. 495:333–338. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Li F, Zhang L, Li W, Deng J, Zheng J, An
M, Lu J and Zhou Y: Circular RNA ITCH has inhibitory effect on ESCC
by suppressing the Wnt/β-catenin pathway. Oncotarget. 6:6001–6013.
2015.PubMed/NCBI
|
|
15
|
Jeck WR, Sorrentino JA, Wang K, Slevin MK,
Burd CE, Liu J, Marzluff WF and Sharpless NE: Circular RNAs are
abundant, conserved, and associated with ALU repeats. RNA.
19:141–157. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Suzuki H, Zuo Y, Wang J, Zhang MQ,
Malhotra A and Mayeda A: Characterization of RNase R-digested
cellular RNA source that consists of lariat and circular RNAs from
pre-mRNA splicing. Nucleic Acids Res. 34:e632006. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Suzuki H and Tsukahara T: A view of
pre-mRNA splicing from RNase R resistant RNAs. Int J Mol Sci.
15:9331–9342. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Xuan L, Qu L, Zhou H, Wang P, Yu H, Wu T,
Wang X, Li Q, Tian L, Liu M and Sun Y: Circular RNA: A novel
biomarker for progressive laryngeal cancer. Am J Transl Res.
8:932–939. 2016.PubMed/NCBI
|
|
19
|
Qin M, Liu G, Huo X, Tao X, Sun X, Ge Z,
Yang J, Fan J, Liu L and Qin W: Hsa_circ_0001649: A circular RNA
and potential novel biomarker for hepatocellular carcinoma. Cancer
Biomark. 16:161–169. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Wang X, Zhang Y, Huang L, Zhang J, Pan F,
Li B, Yan Y, Jia B, Liu H, Li S and Zheng W: Decreased expression
of hsa_circ_001988 in colorectal cancer and its clinical
significances. Int J Clin Exp Pathol. 8:16020–16025.
2015.PubMed/NCBI
|
|
21
|
Hansen TB, Wiklund ED, Bramsen JB,
Villadsen SB, Statham AL, Clark SJ and Kjems J: miRNA-dependent
gene silencing involving Ago2-mediated cleavage of a circular
antisense RNA. EMBO J. 30:4414–4422. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Valdmanis PN and Kay MA: The expanding
repertoire of circular RNAs. Mol Ther. 21:1112–1114. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Kefas B, Godlewski J, Comeau L, Li Y,
Abounader R, Hawkinson M, Lee J, Fine H, Chiocca EA, Lawler S and
Purow B: microRNA-7 inhibits the epidermal growth factor receptor
and the Akt pathway and is down-regulated in glioblastoma. Cancer
Res. 68:3566–3572. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Reddy SD, Ohshiro K, Rayala SK and Kumar
R: MicroRNA-7, a homeobox D10 target, inhibits p21-activated kinase
1 and regulates its functions. Cancer Res. 68:8195–8200. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Webster RJ, Giles KM, Price KJ, Zhang PM,
Mattick JS and Leedman PJ: Regulation of epidermal growth factor
receptor signaling in human cancer cells by microRNA-7. J Biol
Chem. 284:5731–5741. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Xiong S, Zheng Y, Jiang P, Liu R, Liu X
and Chu Y: MicroRNA-7 inhibits the growth of human non-small cell
lung cancer A549 cells through targeting BCL-2. Int J Biol Sci.
7:805–814. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Rybak-Wolf A, Stottmeister C, Glazar P,
Jens M, Pino N, Giusti S, Hanan M, Behm M, Bartok O, Ashwal-Fluss
R, et al: Circular RNAs in the mammalian brain are highly abundant,
conserved, and dynamically expressed. Mol Cell. 58:870–885. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Peng L, Yuan XQ and Li GC: The emerging
landscape of circular RNA ciRS-7 in cancer. Oncol Rep.
33:2669–2674. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Xie H, Ren X, Xin S, Lan X, Lu G, Lin Y,
Yang S, Zeng Z, Liao W, Ding YQ and Liang L: Emerging roles of
circRNA_001569 targeting miR-145 in the proliferation and invasion
of colorectal cancer. Oncotarget. 7:26680–26691. 2016.PubMed/NCBI
|
|
30
|
Zhong Z, Lv M and Chen J: Screening
differential circular RNA expression profiles reveals the
regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in
bladder carcinoma. Sci Rep. 6:309192016. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhou C, Liu G, Wang L, Lu Y, Yuan L, Zheng
L, Chen F, Peng F and Li X: MiR-339-5p regulates the growth, colony
formation and metastasis of colorectal cancer cells by targeting
PRL-1. PLoS One. 8:e631422013. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Wu ZS, Wu Q, Wang CQ, Wang XN, Wang Y,
Zhao JJ, Mao SS, Zhang GH, Zhang N and Xu XC: MiR-339-5p inhibits
breast cancer cell migration and invasion in vitro and may be a
potential biomarker for breast cancer prognosis. BMC Cancer.
10:5422010. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Zhang C, Liu J, Wang X, Wu R, Lin M,
Laddha SV, Yang Q, Chan CS and Feng Z: MicroRNA-339-5p inhibits
colorectal tumorigenesis through regulation of the MDM2/p53
signaling. Oncotarget. 5:9106–9117. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Wu Z, Wu Y, Tian Y, Sun X, Liu J, Ren H,
Liang C, Song L, Hu H, Wang L and Jiao B: Differential effects of
miR-34c-3p and miR-34c-5p on the proliferation, apoptosis and
invasion of glioma cells. Oncol Lett. 6:1447–1452. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Berthois Y, Delfino C, Metellus P, Fina F,
Nanni-Metellus I, Al Aswy H, Pirisi V, Ouafik L and Boudouresque F:
Differential expression of miR-200a-3p and miR21 in grade II–III
and grade IV gliomas: Evidence that miR-200a-3p is regulated by
O6-methylguanine methyltransferase and promotes temozolomide
responsiveness. Cancer Biol Ther. 15:938–950. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Zhou Y, Chen Q, Qin R, Zhang K and Li H:
MicroRNA-449a reduces cell survival and enhances cisplatin-induced
cytotoxicity via downregulation of NOTCH1 in ovarian cancer cells.
Tumour Biol. 35:12369–12378. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Yao Y, Ma J, Xue Y, Wang P, Li Z, Li Z, Hu
Y, Shang X and Liu Y: MiR-449a exerts tumor-suppressive functions
in human glioblastoma by targeting Myc-associated zinc-finger
protein. Mol Oncol. 9:640–656. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Shang Y, Zhang Z, Liu Z, Feng B, Ren G, Li
K, Zhou L, Sun Y, Li M, Zhou J, et al: miR-508-5p regulates
multidrug resistance of gastric cancer by targeting ABCB1 and
ZNRD1. Oncogene. 33:3267–3276. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang L, Yang L, Liu X, Chen W, Chang L,
Chen L, Loera S, Chu P, Huang WC, Liu YR and Yen Y: MicroRNA-657
promotes tumorigenesis in hepatocellular carcinoma by targeting
transducin-like enhancer protein 1 through nuclear factor kappa B
pathways. Hepatology. 57:1919–1930. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Shan SW, Fang L, Shatseva T, Rutnam ZJ,
Yang X, Du W, Lu WY, Xuan JW, Deng Z and Yang BB: Mature miR-17-5p
and passenger miR-17-3p induce hepatocellular carcinoma by
targeting PTEN, GalNT7 and vimentin in different signal pathways. J
Cell Sci. 126:1517–1530. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Chaudhry MA, Sachdeva H and Omaruddin RA:
Radiation-induced micro-RNA modulation in glioblastoma cells
differing in DNA-repair pathways. DNA Cell Biol. 29:553–561. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Zhai H, Song B, Xu X, Zhu W and Ju J:
Inhibition of autophagy and tumor growth in colon cancer by
miR-502. Oncogene. 32:1570–1579. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Tang J, Tao ZH, Wen D, Wan JL, Liu DL,
Zhang S, Cui JF, Sun HC, Wang L, Zhou J, et al: MiR-612 suppresses
the stemness of liver cancer via Wnt/β-catenin signaling. Biochem
Biophys Res Commun. 447:210–215. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Li R, Zhang L, Jia L, Duan Y, Li Y, Wang
J, Bao L and Sha N: MicroRNA-143 targets Syndecan-1 to repress cell
growth in melanoma. PLoS One. 9:e948552014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Jin L, Hu WL, Jiang CC, Wang JX, Han CC,
Chu P, Zhang LJ, Thorne RF, Wilmott J, Scolyer RA, et al:
MicroRNA-149*, a p53-responsive microRNA, functions as an oncogenic
regulator in human melanoma. Proc Natl Acad Sci USA. 108:pp.
15840–15845. 2011; View Article : Google Scholar : PubMed/NCBI
|
|
46
|
Li JR, Wang JQ, Gong Q, Fang RH and Guo
YL: MicroRNA-328 inhibits proliferation of human melanoma cells by
targeting TGFβ2. Asian Pac J Cancer Prev. 16:1575–1579. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Liu R, Xie H, Luo C, Chen Z, Zhou X, Xia
K, Chen X, Zhou M, Cao P, Cao K and Zhou J: Identification of FLOT2
as a novel target for microRNA-34a in melanoma. J Cancer Res Clin
Oncol. 141:993–1006. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Zhou J, Xu D, Xie H, Tang J, Liu R, Li J,
Wang S, Chen X, Su J, Zhou X, et al: miR-33a functions as a tumor
suppressor in melanoma by targeting HIF-1α. Cancer Biol Ther.
16:846–855. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Elson-Schwab I, Lorentzen A and Marshall
CJ: MicroRNA-200 family members differentially regulate
morphological plasticity and mode of melanoma cell invasion. PLoS
One. 5(pii): e131762010. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Satzger I, Mattern A, Kuettler U,
Weinspach D, Voelker B, Kapp A and Gutzmer R: MicroRNA-15b
represents an independent prognostic parameter and is correlated
with tumor cell proliferation and apoptosis in malignant melanoma.
Int J Cancer. 126:2553–2562. 2010.PubMed/NCBI
|
|
51
|
Martin de Campo SE, Latchana N, Levine KM,
Grignol VP, Fairchild ET, Jaime-Ramirez AC, Dao TV, Karpa VI,
Carson M, Ganju A, et al: MiR-21 enhances melanoma invasiveness via
inhibition of tissue inhibitor of metalloproteinases 3 expression:
In vivo effects of MiR-21 inhibitor. PLoS One. 10:e01159192015.
View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Zhou J, Liu R, Wang Y, Tang J, Tang S,
Chen X, Xia K, Xiong W, Xu D, Wang S, et al: miR-199a-5p regulates
the expression of metastasis-associated genes in B16F10 melanoma
cells. Int J Clin Exp Pathol. 7:7182–7190. 2014.PubMed/NCBI
|
|
53
|
Schmitt MJ, Philippidou D, Reinsbach SE,
Margue C, Wienecke-Baldacchino A, Nashan D, Behrmann I and Kreis S:
Interferon-γ-induced activation of signal transducer and activator
of transcription 1 (STAT1) up-regulates the tumor suppressing
microRNA-29 family in melanoma cells. Cell Commun Signal.
10:412012. View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Bhattacharya A, Schmitz U, Wolkenhauer O,
Schönherr M, Raatz Y and Kunz M: Regulation of cell cycle
checkpoint kinase WEE1 by miR-195 in malignant melanoma. Oncogene.
32:3175–3183. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Sun L, Bian G, Meng Z, Dang G, Shi D and
Mi S: MiR-144 inhibits uveal melanoma cell proliferation and
invasion by regulating c-Met expression. PLoS One. 10:e01244282015.
View Article : Google Scholar : PubMed/NCBI
|