Introduction
Prostate cancer is the second most frequently
diagnosed cancer in men, and the second largest cause of
cancer-associated mortalities globally in 2018 (1,2).
Despite improvements in therapy, the 5-year survival rate of
prostate cancer remained at 66.4% for patients diagnosed with
prostate cancer in 2012 in China (3,4).
Metastasis and invasion are the main causes of prostate
cancer-associated mortality (5).
Thus, deciphering the mechanism underlying invasive and metastatic
behavior is of great importance for the early diagnosis and
treatment of prostate cancer.
Ubiquitination and deubiquitination are core
regulatory functions in the process of post-translational protein
modification. Deubiquitinating enzymes serve an important role in
the regulation of multiple biological processes, including cell
cycle control, DNA repair, chromatin remodeling, the
epithelial-mesenchymal transition (EMT) and several signaling
pathways that are frequently dysregulated during tumor development
(6).
Ubiquitin-specific protease (USP)9X is a
deubiquitinase member of the USP family (7). USP9X has been demonstrated to remove
ubiquitin moieties, principally directing target proteins toward
proteasomal degradation, and to be involved a variety of diseases,
including malignant tumors (8). A
previous study identified this protein as a tumor suppressor with
prognostic and therapeutic relevance in pancreatic ductal
adenocarcinoma (PDA), as its depletion significantly increased
colony formation in soft agar and suppressed anoikis in metastatic
PDA (9). In clear cell renal cell
carcinoma, USP9X expression was reported to be downregulated and
associated with a poor prognosis (10). USP9X also ameliorates the oncogenic
activity of transcriptional coactivator YAP and tafazzin, which are
core components of the Hippo pathway, by deubiquitinating their
inhibitor, angiomotin (10). The
aforementioned evidence indicates that USP9X may be a potent tumor
suppressor. However, whether USP9X exerts a similar effect on human
prostate cancer remains unclear.
In the present study, the clinical significance of
USP9X in prostate cancer was investigated, and its involvement in
the biological features of prostate cancer cells and the underlying
mechanisms were explored.
Materials and methods
Samples
A total of 102 cases of prostate adenocarcinoma
tissue with no prior adjuvant treatment and 25 cases of normal
tissue were obtained from male patients attended at the Second
Affiliated Hospital of Kunming Medical University (Kunming, China)
between January 2013 and November 2016, with informed consent. The
samples were fixed in 10% neutral buffered formalin for 24 h and
embedded in paraffin. The mean age ± standard deviation was
68.1±8.67 years (range, 40–87 years). The normal tissues were
obtained from patients with benign prostate diseases. An additional
set of8 fresh specimens of tumor tissue and adjacent normal tissue
were stored at −70°C immediately following resection for subsequent
protein expression analysis. This study was conducted with the
approval of the Ethics Committee of the Second Affiliated Hospital
of Kunming Medical University.
Immunohistochemistry
The sections were deparaffinized in xylene,
rehydrated with graded alcohol, and then boiled in 0.01 M citrate
buffer (pH 6.0) for 2 min in an autoclave. Hydrogen peroxide (0.3%)
was applied to block endogenous peroxide activity, and the sections
were incubated with normal goat serum (Fuzhou Maixin Biotech Co.,
Ltd., Fuzhou, China) at 37°C for 15 min to reduce non-specific
binding. The tissue sections (4 µm thickness) were incubated with
anti-USP9X antibody (1:400; cat. no. 55054-1-AP; ProteinTech Group,
Inc., Chicago, IL, USA) overnight at 4°C. Biotinylated goat
anti-mouse or anti-rabbit serum IgG was used as a secondary
antibody and the incubation was performed at 37°C for 25 min.
Following washing, the sections were incubated at 37°C for 25 min
with streptavidin-biotin conjugated with horseradish peroxidase
(HRP) (all supplied in the Elivision™ Super HRP IHC Kit-9922;
Fuzhou Maixin Biotech Co., Ltd.), and the peroxidase reaction was
developed with 3,3′-diaminobenzidine tetrahydrochloride.
Counterstaining with hematoxylin was performed at room temperature
for 2 min and the sections were dehydrated in ethanol prior to
mounting.
Two independent blinded investigators examined all
tumor slides randomly and a third pathologist was consulted in the
case of any discrepancies. Five views were examined per slide, and
100 cells were observed per view at ×400 magnification using a
light microscope. Immunostaining of USP9X was scored on a
semiquantitative scale by evaluating the percentage of
immunoreactive tumor cells and the staining intensity. Based on
previous studies (11,12), the staining intensity was
categorized as follows: 0, negative or weak; 1, moderate; and 2,
strong. The percentage of stained tumor cells was scored as: 0, 0%;
1 1–5%; 2, 6–25%; 3, 26–75%; and 4, 76–100%. The scores of each
tumor sample were multiplied to give a final score of 0–8, and
those with a final score of 4–8 were classified as having high
USP9X expression. Tumor samples with a score <4 were considered
as having low USP9X expression.
Cell culture and reagents
All cell lines were obtained from the Cell Bank of
Type Culture Collection of Chinese Academy of Sciences (Shanghai,
China). The prostate cancer cell lines, LNCaP, DU145 and PC-3, were
maintained in RPMI-1640 medium, minimum essential medium and F-12K
medium (Thermo Fisher Scientific, Inc., Waltham, MA, USA),
supplemented with 10% fetal bovine serum (FBS; Thermo Fisher
Scientific, Inc.) at 37°C in 5% CO2. The human prostate
epithelial RWPE-1 cell line was maintained in K-SFM medium (Thermo
Fisher Scientific, Inc.). The extracellular signal-regulated kinase
(ERK) inhibitor PD98059 was purchased from Selleck Chemicals
(Houston, TX, USA) and the cells were treated at the concentration
of 10 µM.
Transfection
ON-TARGETplus small interfering (si)RNAs and control
siRNA (siControl) were obtained from GE Healthcare Dharmacon, Inc.
(Lafayette, CO, USA). siRNAs were transfected into cells (50 nM)
using DharmaFECT1 transfection reagent (GE Healthcare Dharmacon,
Inc.) according to the manufacturer's protocol. The sequence of the
siControl was 5′-GCGCGATAGCGCGAATATA-3′. The target sequences for
USP9X siRNA were: siRNA1, 5′-AGAAAUCGCUGGUAUAAAU-3′; siRNA2,
5′-GUACGACGAUGUAUUCUCA-3′; and siRNA3, 5′-GAAAUAACUUCCUACCGAA-3′.
siRNA3 exhibited the best efficiency, as determined by the mRNA and
protein measurements described below, and was chosen for the
knockdown experiments. Subsequent experiments were performed 55–72
h after transfection.
Reverse transcription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was isolated from cells using TRIzol
(Invitrogen; Thermo Fisher Scientific, Inc.). For qPCR, cDNA was
synthesized using the iScript™ Reverse Transcription SuperMix
(Bio-Rad Laboratories, Inc., Hercules, CA, USA) using the following
temperature protocol: 5 min at 25°C, 20 min at 46°C and 1 min at
95°C. The relative mRNA expression levels were determined from the
cDNA using SYBR Master mix on an ABI 7500 Real-Time PCR system
(both Applied Biosystems; Thermo Fisher Scientific, Inc.) using the
following thermocycling conditions: 50°C for 2 min, 95°C for 10 min
and 40 cycles of 95°C for 15 sec and 60°C for 60 sec. The primer
sequences were as follows: USP9X forward,
5′GGACTCCTGGCAAACTCACAGA3′ and reverse, 5′-TCAAACAGCCCATCTCGGTC-3′;
and GAPDH forward, 5′-CTGGGCTACACTGAGCACC-3′ and reverse,
5′-AAGTGGTCGTTGAGGGCAATG-3′. Relative expression was normalized
against GAPDH and calculated using the comparative Cq method
(2−ΔΔCq) (13). All
experiments were performed in triplicate.
Western blot analysis
Total protein was extracted using Pierce lysis
buffer (Thermo Fisher Scientific, Inc.). Protein quantification was
performed using the Bradford method. A total of 50 µg sample
protein was separated by 7–15% SDS-PAGE, which was transferred to
polyvinylidene fluoride (PVDF) membranes (Merck KGaA, Darmstadt,
Germany). The incubation of primary antibodies was performed
overnight at 4°C. The antibodies used were against the following
proteins: USP9X (1:1,000, cat. no. 55054-1-AP; ProteinTech Group,
Inc.), phosphorylated (p-)ERK (cat. no. 4037), ERK (cat. no. 4695),
p-protein kinase B (p-AKT; cat. no. 4060), AKT (cat. no. 4685),
cyclin D1 (cat. no. 2978), vimentin (cat. no. 5741),
p-transcription factor p65 (cat. no. 3033), p-dynamin-related
protein 1 (p-DRP1; cat. no. 3455), DRP1 (cat. no. 8570), p65 (cat.
no. 4282) (all 1:1,000; Cell Signaling Technology, Inc., Danvers,
MA, USA), matrix metalloproteinase 9 (MMP9; 1:1,000; cat. no.
ab38898; Abcam, Cambridge, UK), E-cadherin (1:1,500, cat. no.
610181; Becton, Dickinson and Company, Franklin Lakes, NJ, USA) and
β-actin (1:2,000; cat. no. sc-47778; Santa Cruz Biotechnology,
Inc., Dallas, TX, USA). Following incubation with HRP-coupled
anti-mouse or rabbit IgG antibody (1:1,000 dilution; cat. nos. 7076
and 7074, respectively; Cell Signaling Technology, Inc.) at 37°C
for 2 h. The target proteins on the PVDF membrane were visualized
using the Pierce ECL kit (Thermo Fisher Scientific, Inc.) and
captured using the MicroChemi imaging system (DNR Bio-Imaging
Systems, Ltd., Neve Yamin, Israel). The western blot intensity was
analyzed using ImageJ software version 1.48 (National Institutes of
Health, Bethesda, MD, USA).
Cell invasion assay
A cell invasion assay was performed using a 24-well
Transwell chamber with a pore size of 8 µm (Costar; Corning, Inc.,
Corning, NY, USA). The inserts were coated with Matrigel (Becton,
Dickinson and Company). At 48 h post-transfection, the cells were
trypsinized and 3×105 cells in 100 µl serum-free medium
were transferred to the upper Matrigel chamber and incubated for 16
h. Medium supplemented with 10% FBS was added to the lower chamber
as the chemoattractant. Following incubation, the non-invaded cells
on the upper membrane surface were removed with a cotton tip, and
the cells that passed through the filter were fixed with 4%
paraformaldehyde and stained with hematoxylin for 2 min at room
temperature. Images were captured using alight microscope (Olympus
Corporation, Tokyo, Japan) at ×400 magnification. All experiments
were performed in triplicate.
Scratch wound-healing assay
To observe cell migration, cells were seeded at a
density of ~80% confluence as a monolayer, and scratched with a
1-ml pipette tip. The detached cells were washed away and the
remaining cells were cultured for 24 h in corresponding medium
(RPMI-1640 medium for the LNCaP cells and F-12K medium for the PC-3
cells) without FBS to slow down the growth rate. Images of the cell
monolayer were taken using a light microscope (Olympus Corporation;
×200 magnification) and the width of the scratch wound was measured
using ImageJ software version 1.48. All experiments were performed
in triplicate.
Cell cycle analysis
For cell cycle analysis, cells were fixed in cold
70% ethanol for ≥1 h at 4°C. The cells were then washed with PBS,
pelleted at 3,500 × g at 4°C for 10 min and the supernatant was
discarded. The cells were subsequently resuspended in 500 µl PBS
containing 10 µg/ml RNase A and 20 µg/ml propidium iodide (PI), and
incubated at room temperature in the dark for 30 min. The cell
cycle was analyzed by flow cytometry using the FACSCalibur™ flow
cytometer and FloJo software version 10 (both Becton, Dickinson and
Company).
Apoptosis analysis
Cell apoptosis detection was performed with Annexin
V/PI double staining, using the FITC Annexin V Apoptosis Detection
kit I (Becton, Dickinson and Company). At 48 h post-transfection,
the cells were harvested, washed in chilled PBS and resuspended in
250 µl binding buffer. Annexin V-fluorescein isothiocyanate and PI
solutions were added to the cell suspension and incubated for 30
min at room temperature. The cells were analyzed using the
FACSCalibur flow cytometer and FlowJo software version 10.
Mitochondrial membrane potential
The mitochondrial membrane potential (Δψm) was
detected using JC-1 staining. Briefly, cells were harvested, washed
with PBS and incubated with 5 µM JC-1 (Cell Signaling Technology,
Inc.) for 30 min at 37°C in the incubator. The cells were then
washed and analyzed using the FACSCalibur flow cytometer and FlowJo
software version 10.
MTT assay
To determine the proliferation rate of cells, 5,000
cells per well were plated in 96-well plates in medium containing
10% FBS. MTT solution (20 µl, 5 mg/ml) was added into each well and
incubated for 1 hat 37°C. Dimethylsulfoxide was added into each
well to dissolve the formazan and the dye was measured at 490 nm
using a microplate reader (Thermo Fisher Scientific, Inc.).
Proliferation assay
Proliferation was assessed using the CyQUANT™ cell
proliferation assay (Invitrogen; Thermo Fisher Scientific, Inc.).
The cells were plated in 96-well black fluorescence microtitre
plates, at a density of 5,000 cells/well. After 24 h of
transfection at 37°C, the medium was discarded, and the plates
frozen at −80°C until use. On the day of the analysis, the plates
with the adherent cells were thawed and incubated with the CyQUANT
dye for 5 min in the dark. The fluorescence was measured on a
fluorescence microplate reader (BioTek Instruments, Inc., Winooski,
VT, USA) with the excitation set at 480 nm and emission at 520
nm.
Mitochondrial function
To determine the subcellular distribution of
mitochondria, the cells were treated with 50 nM MitoTracker™ Red
(Invitrogen; Thermo Fisher Scientific, Inc.) at 37°C for 30 min to
stain the mitochondria and cell cytosol. The mean mitochondrial
length was determined by measuring 10 individual mitochondria from
cells obtained by fluorescence microscopy using the cellSens
software version 1.16 (Olympus Corporation).
Statistical analysis
SPSS version 16.0 for Windows (SPSS Inc., Chicago,
IL, USA) was used for all statistical analyses. The χ2
test was used to examine associations between USP9X expression
levels and patient clinicopathological factors. Paired Student's
t-test was used for the comparison of the intensity of USP9X in
normal and cancer tissues. Analysis of variance, followed by a
least-significant-difference post hoc test was performed for the
comparison of USP9X expression among the cell lines and comparison
of protein change in cells treated with USP9X siRNA and PD98059.
Unpaired Student's t-test was used to perform the remaining
comparisons. The P-values were based on a two-sided statistical
analysis and P<0.05 was considered to indicate statistically
significant differences.
Results
USP9X is downregulated in prostate
cancer tissues
To assess the protein expression and distribution of
USP9X in prostate cancer, immunohistochemistry was performed on 102
prostate adenocarcinoma tissue samples and 25 normal prostate
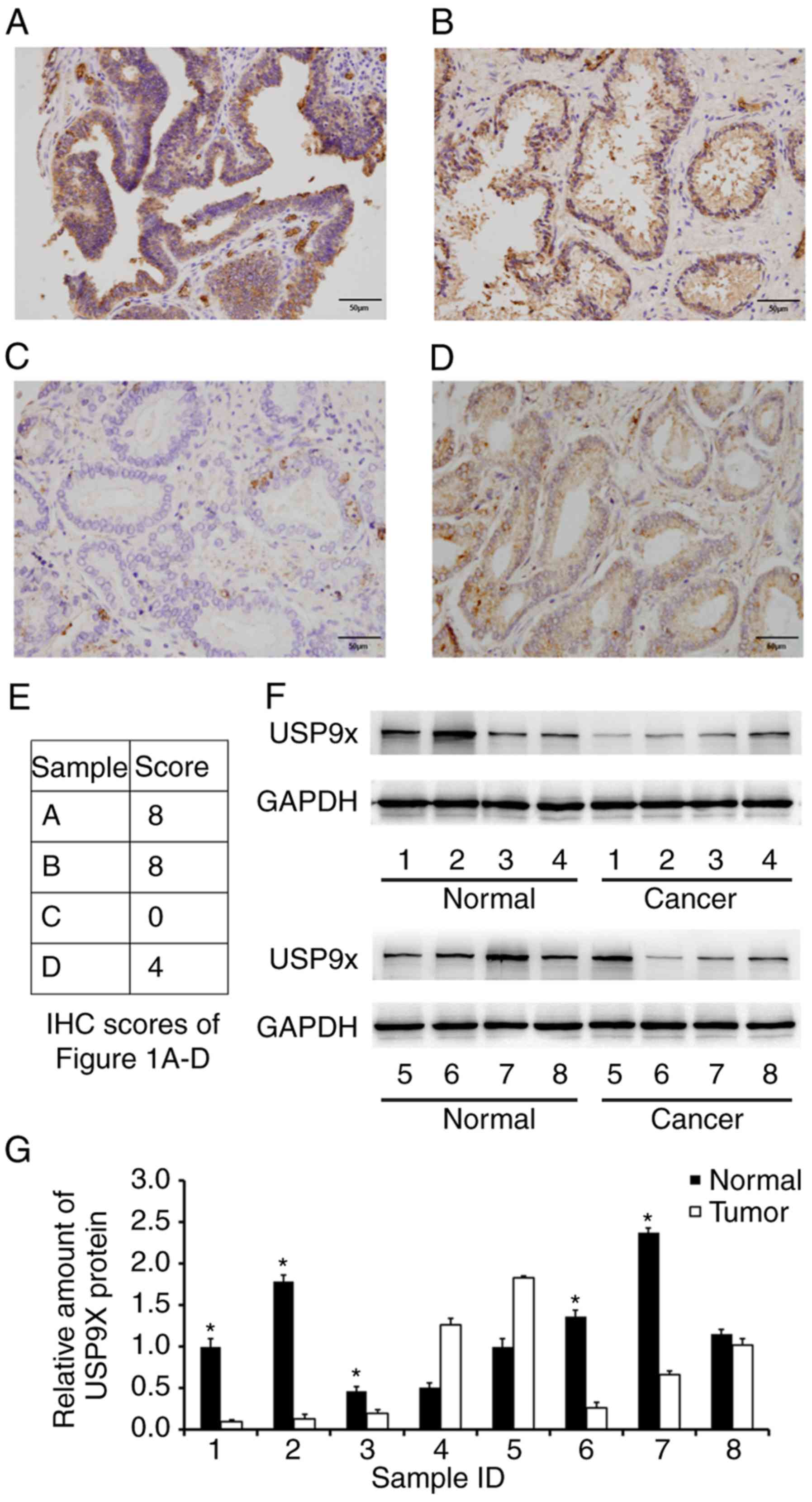
tissue samples from different individuals. As displayed in Fig. 1A-D, USP9X was mainly localized in
the cytoplasm with sporadic nuclear distribution. Low USP9X protein
expression (downregulation) was demonstrated in 54.9% (56/102
samples) of the prostate cancer cases, whereas only 8% (2/25
samples) of the normal tissues exhibited low USP9X expression. The
immunostaining scores of the images in Fig. 1A-D are indicated in Fig. 1E. Statistical analysis revealed that
low USP9X expression was associated with higher Gleason scores
(14) (>7; P=0.0046) and higher
T stage (15) (T1-T2 vs. T3-T4,
P=0.0138), indicating that downregulation of USP9X is linked with
poor differentiation and local invasion. No statistical difference
was found between USP9X expression and either age (P=0.7074),
tumor-node-metastasis stage (16)
(P=0.3103), distal metastasis (P=0.6715) or nodal status (16) (P=0.7045) (Table I). Additionally, USP9X protein
expression levels were analyzed in 8 cases of paired prostate
cancer/normal tissues using western blotting, and the overall
downregulation of USP9X in cancer tissues compared with normal
tissues at the protein level was confirmed in 5 out of 8 cases
(Fig. 1F and G).
 | Table I.Distribution of USP9X status in
prostate cancer according to the clinicopathological
characteristics of the patients. |
Table I.
Distribution of USP9X status in
prostate cancer according to the clinicopathological
characteristics of the patients.
|
|
| USP9X expression,
n |
|
|---|
|
|
|
|
|
|---|
|
Characteristics | Patients, n | Negative/low | High | P-value |
|---|
| Age, years |
|
|
| 0.7074 |
|
<65 | 33 | 19 | 14 |
|
|
≥65 | 69 | 37 | 32 |
|
|
Tumor-node-metastasis stage |
|
|
| 0.3103 |
|
I–II | 52 | 26 | 26 |
|
|
III–IV | 50 | 30 | 20 |
|
| Gleason score |
|
|
| 0.0046a |
| ≤7 | 40 | 15 | 25 |
|
|
>7 | 62 | 41 | 21 |
|
| T stage |
|
|
| 0.0138a |
|
T1-T2 | 62 | 28 | 34 |
|
|
T3-T4 | 40 | 28 | 12 |
|
| Lymph node
metastasis |
|
|
| 0.7045 |
|
Absent | 76 | 41 | 35 |
|
|
Present | 26 | 15 | 11 |
|
| Metastasis |
|
|
| 0.6715 |
|
Absent | 71 | 38 | 33 |
|
|
Present | 31 | 18 | 13 |
|
USP9X depletion facilitates invasion
and migration in prostate cancer cells
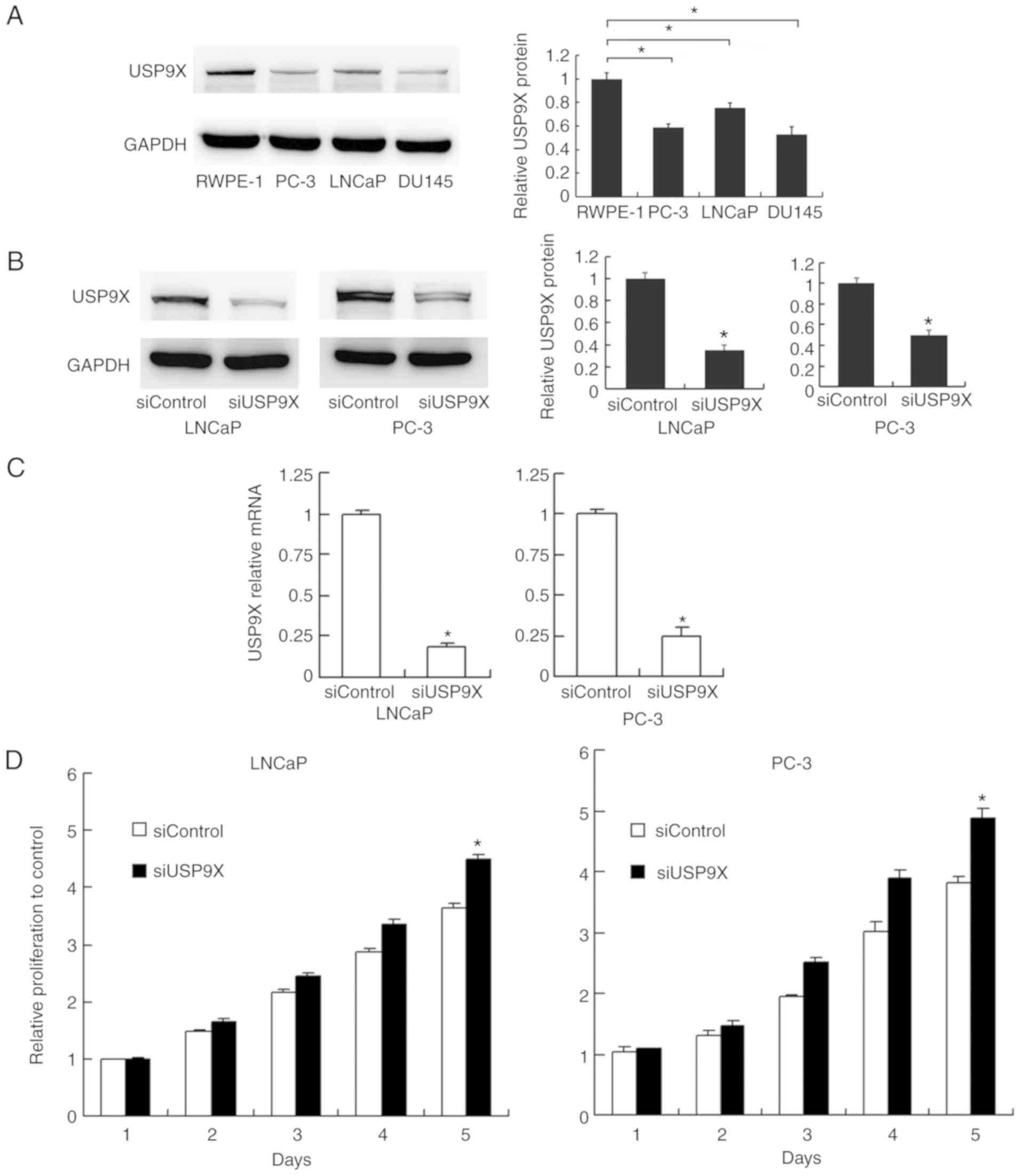
The levels of USP9X were examined by western
blotting in normal prostate RWPE-1 cells and prostate cancer PC-3,
LNCaP and DU145 cells. The expression of USP9X was higher in the
normal RWPE-1 cells compared with that in the cancer cells
(Fig. 2A). To characterize the
impact of USP9X on the biological function of prostate cancer,
USP9X silencing was performed in the prostate cancer PC-3 and LNCaP
cells using siRNA. The siRNA knockdown efficiency was confirmed by
western blotting (Fig. 2B) and
RT-qPCR analysis (P<0.001 vs. siControl; n=3; Fig. 2C). Subsequently, an MTT assay and
cell cycle analysis were performed to determine the involvement of
USP9X in cell proliferation. The cell proliferation assay
demonstrated that USP9X depletion increased the rate of cell
proliferation in the prostate cancer cell lines only on day 5
(LNCaP, 21% increase; and PC-3, 28% increase; P<0.001 vs.
siControl of the same day; n=3; Fig.
2D), which was not considered to be a notable overall effect.
The cell cycle analysis demonstrated that USP9X knockdown did not
cause any changes in the percentage distribution of cells in each
phase (Fig. 2E). The change in the
rate of cell apoptosis was then assessed. The Annexin V/PI staining
revealed that USP9X silencing downregulated the apoptotic rate
(P<0.001 vs. siControl; n=3; Fig.
2F). These results indicate that USP9X partially inhibits cell
proliferation in prostate cancer cells.
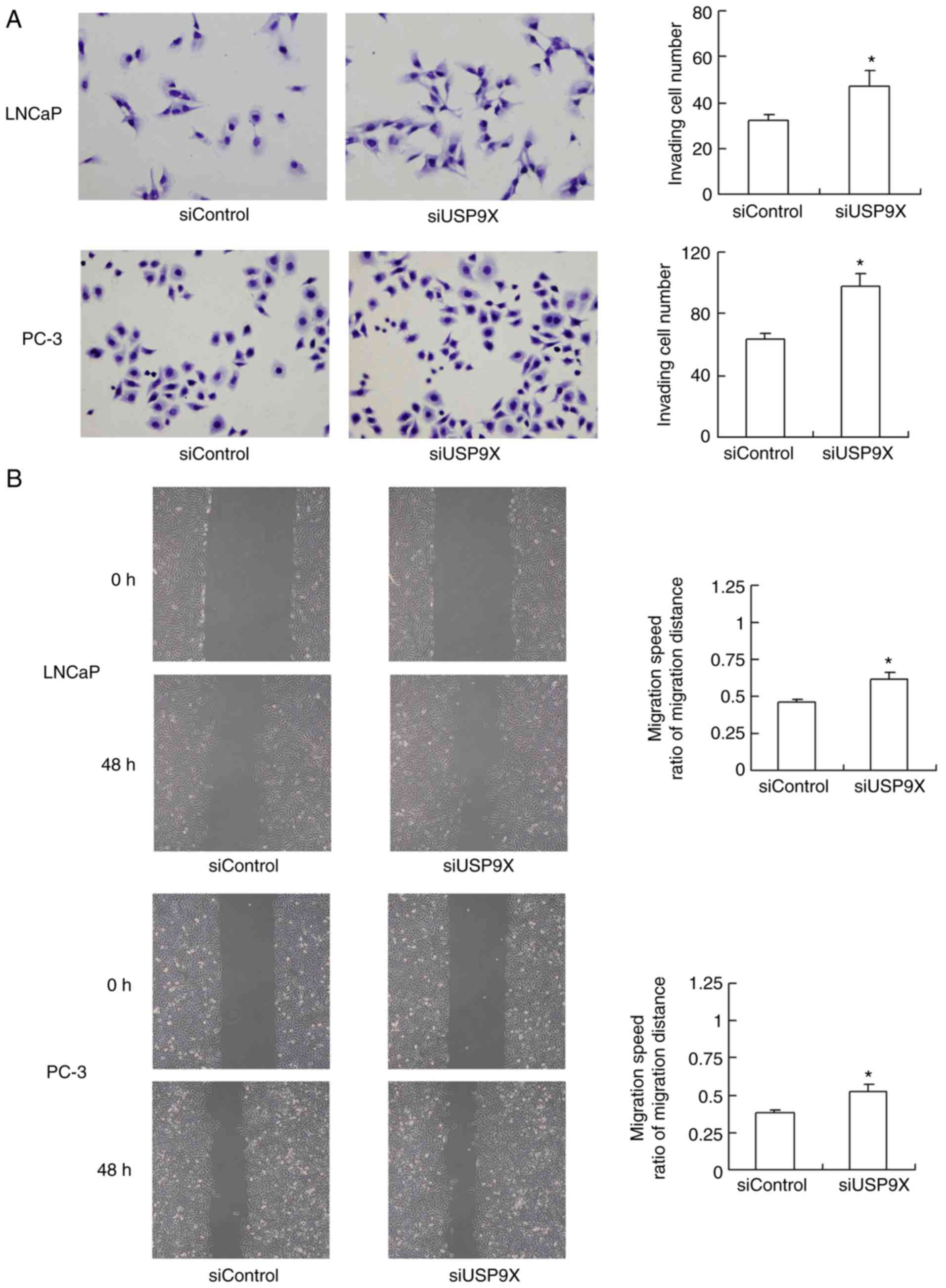
The effect of USP9X silencing on cell invasion and
migration was also evaluated. As demonstrated in Fig. 3A, USP9X-knockdown significantly
increased the invasive ability of the LNCaP and PC-3 cells (47 and
56% increase, respectively; P<0.05 vs. siControl). The migration
of the cells was assessed using a wound-healing assay (Fig. 3B). USP9X-knockdown significantly
upregulated the migratory ability of the LNCaP and PC-3 cell lines
(37 and 42% increase; P<0.05 vs. siControl). The experiments
were performed in triplicate. These results indicate that USP9X
regulates the invasive ability of prostate cancer cells, and may
therefore be a tumor suppressor in prostate cancer, which is in
agreement with the immunohistochemistry results.
USP9X depletion upregulates MMP9 and
induces mitochondrial fission in prostate cancer cells
The aforementioned findings demonstrated that USP9X
regulates prostate cancer cell invasion/migration more than
proliferation. The mechanistic basis of USP9X-regulated invasion
and migration was explored next. The MMP proteins serve pivotal
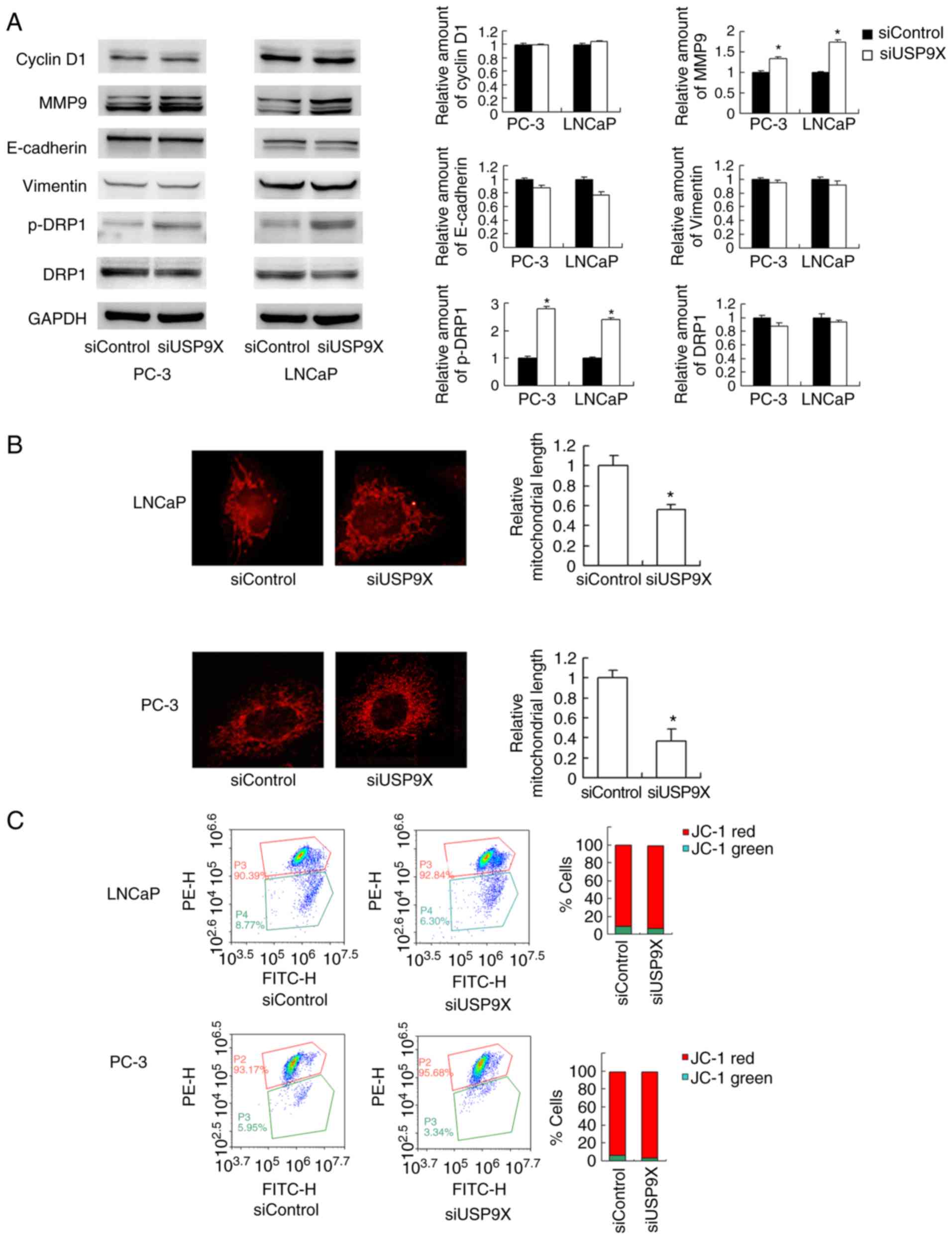
roles during the initiation of cancer invasion (17). A western blot analysis revealed that
the level of the MMP9 protein was increased in USP9X-knockdown
LNCaP and PC-3 cells compared with that in the corresponding
controls (Fig. 4A). Regarding cell
cycle- and EMT-associated proteins, no significant differences were
observed in the levels of cyclin D1, E-cadherin or vimentin
(Fig. 4A). Cancer metabolism,
particularly the mitochondrial status, has been reported to control
cell migration (18). In the
present study, changes in mitochondrial function were tested by
examining the mitochondrial dynamics. As observed in Fig. 4B, USP9X-knockdown induced
mitochondrial fission in the prostate cancer cell lines. In the
USP9X-depleted LNCaP and PC-3 cells, the mitochondria displayed
short tubules and spheres with an average length ~50% shorter than
those in the control cells (P<0.001 vs. siControl; n=3; Fig. 4B). Accordingly, the level of p-DRP1,
a member of the dynamin superfamily of GTPases that control
mitochondrial fission/fusion balance (19), was increased. These results indicate
that USP9X depletion induces prostate cancer cell
invasion/migration, likely through the regulation of MMP9 and
activation of DRP1.
JC-1 staining was used to examine the mitochondrial
membrane potential, exhibiting red fluorescence under normal
conditions that turns green when the Δψm is decreased. No
significant change in the red/green ratio was observed (Fig. 4C), suggesting that USP9X-knockdown
did not affect the mitochondrial membrane potential.
USP9X-induced effects on MMP9 and DRP1
in prostate cancer cells are dependent on the ERK signaling
pathway
To further elucidate the mechanism of MMP9 and
p-DRP1 regulation by USP9X, several signaling pathways were
screened. USP9X-knockdown upregulated p-ERK, whereas no significant
differences were observed in the levels of p-AKT or p-p65 (Fig. 5A). ERK signaling has been reported
to induce transcriptional activation of MMP9 and DRP1
phosphorylation in cancer cells (20,21).
To confirm the involvement of ERK signaling in the USP9X-induced
regulation of MMP9 and phosphorylation of DRP-1, the ERK inhibitor
PD98059 was used to block the ERK pathway, and protein expression
levels were compared in the siUSP9X vs. siControl cells. The
PD98059 treatment blocked ERK phosphorylation in the prostate
cancer cells (Fig. 5B). ERK
inhibition significantly decreased the levels of MMP9 and p-DRP1
(P<0.05). Notably, the inhibition of ERK abolished the effect of
USP9X siRNA on MMP9 and p-DRP1 levels. These data support the
conclusion that the effects of USP9X on MMP9 and p-DRP1 are
dependent on ERK signaling (Fig.
5B).
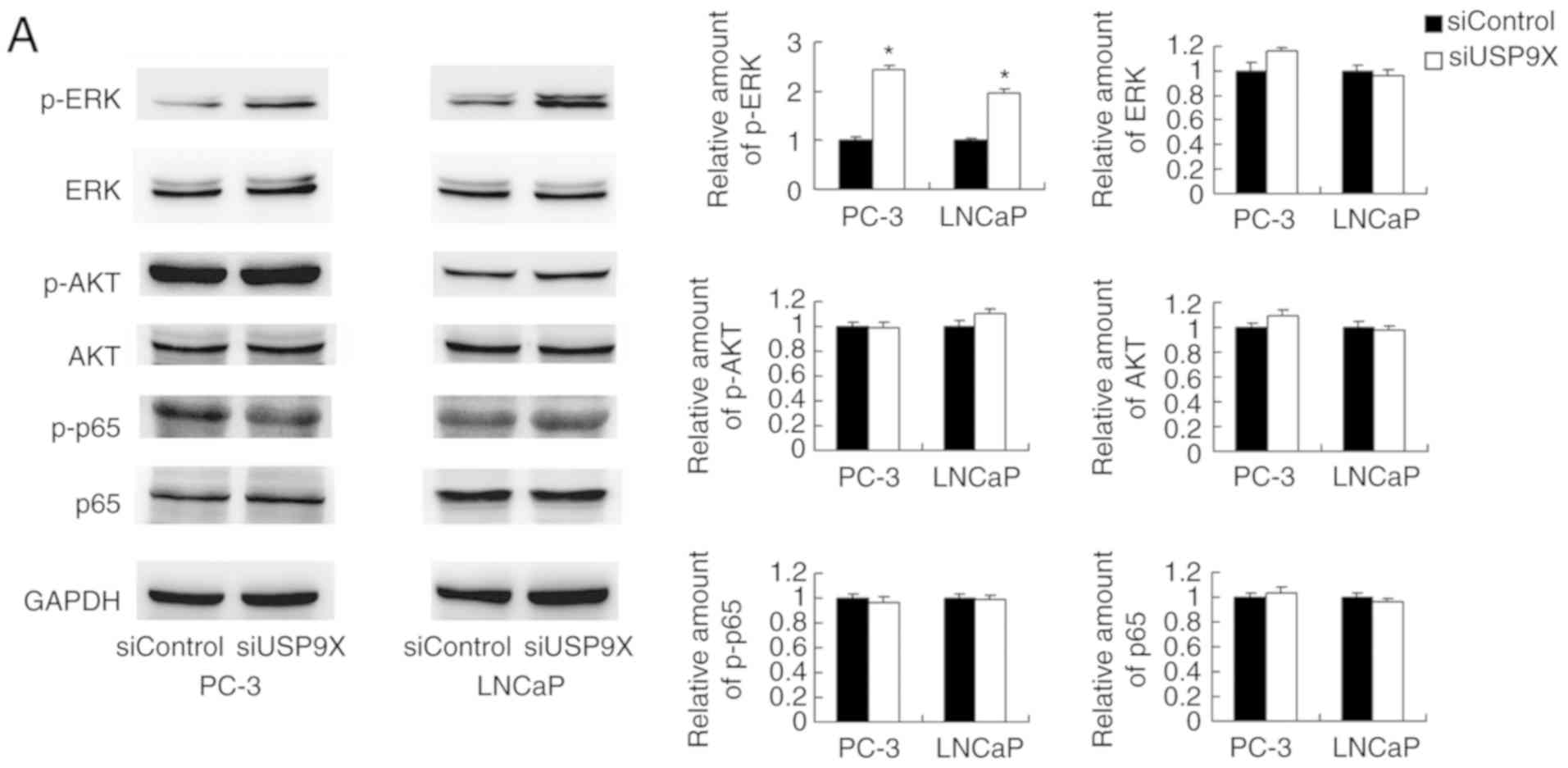 | Figure 5.USP9X regulates MMP9 and p-DRP1
through the ERK signaling pathway. (A) Western blotting
demonstrated that USP9X-knockdown upregulated the levels of p-ERK.
No changes were observed for p-AKT and p-p65. (B) PD98059 treatment
inhibited ERK phosphorylation in the prostate cancer cell lines,
and decreased MMP9 and p-DRP1 protein levels. The inhibitor PD98059
abolished the effect of USP9X silencing on the levels of MMP9 and
p-DRP1. #P<0.05, *P<0.05 vs. untreated siControl
and §P<0.05 vs. PD98059-treated siControl using
analysis of variance with a least-significant-difference post hoc
test. The data are presented as the mean ± standard deviation.
USP9X, ubiquitin-specific protease 9X; MMP9, matrix
metalloproteinase 9; DRP1, dynamin-related protein 1; p-,
phosphorylated; ERK, extracellular signal-regulated kinase; AKT,
protein kinase B; p65, transcription factor p65; siUSP9X, siRNA
targeting the USP9X gene; siControl, control siRNA. |
Discussion
USP9X is a highly conserved member of the USP family
of deubiquitinating enzymes (22)
and serves an important role in development and disease (8,23).
Previous studies have suggested that USP9X serves a
context-specific role in the progression of cancer, acting either
as a tumor suppressor or an oncoprotein (24–26).
Elevated expression of USP9X has been linked to a poor prognosis in
patients with non-small lung carcinoma (11), gastric cancer (12) and multiple myeloma (25). However, it was also reported that it
suppresses tumor growth via serine/threonine-protein kinase LATS
and the Hippo pathway (23). USP9X
may also act as a tumor suppressor by maintaining genomic stability
(23). These reports suggest that
the role of USP9X in human cancer types is context-dependent.
Therefore, it is necessary to characterize its biological function
in different types of cancer. In the present study, the USP9X
protein levels were demonstrated to be downregulated in prostate
cancer tissues using immunohistochemistry and western blotting, and
a statistical analysis revealed that its downregulation was
associated with higher Gleason scores. Since tumors with higher
Gleason scores tend to be more aggressive, the present results
indicate that there is a possible link between USP9X downregulation
and an aggressive phenotype in prostate cancer. USP9X
downregulation was associated with a more advanced T stage,
indicating its association with the local invasion of prostate
cancer. In addition, USP9X protein levels were higher in the normal
prostate RWPE-1 cells compared with those in various prostate
cancer cells. These findings demonstrate that USP9X is
downregulated in prostate cancer and acts as a potential tumor
suppressor.
The biological functions of USP9X were investigated
using siRNA treatment followed by a series of experiments,
including proliferation, cell cycle, apoptosis, invasion and
migration assays. USP9X silencing upregulated the proliferation
rate only on day 5, with no significant effect on the cell cycle
progression. However, the overall effect of USP9X silencing on
proliferation promotion was not significant. The Annexin V/PI
analysis revealed a downregulation of the apoptotic rate following
USP9X silencing, suggesting that the effect of USP9X on prostate
cancer cell growth may be due to its ability to inhibit
apoptosis.
Compared with its effect on proliferation, USP9X
siRNA led to a more notable effect on invasion and migration in the
prostate cancer cells. To determine the underlying mechanism,
proteins directly associated with invasion were investigated, and
the results revealed that MMP9 levels were increased following the
silencing of USP9X. MMP9 is a well-known collagenase involved in
the degradation of the extracellular matrix (ECM) and prostate
cancer cell invasion (27–30), explaining the increased invading
ability induced by USP9X-knockdown.
The analysis of the clinical data of patients with
prostate cancer revealed no significant association of USP9X
expression with nodal or distal metastasis. Metastasis is a
multi-step process, including the attachment and remodeling of the
ECM, migration of the tumor cell body through the remodeled matrix,
invasion through the lymph or blood vessels, survival within the
circulation and colonization/proliferation at the secondary sites
(31,32). In vitro invasion represents
one of these steps. However, multiple genetic factors contribute to
the metastasis in the clinical setting. Therefore, it is possible
that a discrepancy exists between in vitro experiments and
clinical data.
Cell migration is a complex cellular process
influenced by numerous biological mechanisms, including the actin
network, adhesion and energy metabolism. One key step in migration
is the formation of lamellipodia at the leading edge, and this
process consumes ATP produced by the mitochondria (33). Previous studies have suggested an
association between mitochondrial function and cancer
invasion/migration; for example, it was reported that increased
mitochondrial fission induced cell migration (34,35).
In the present study, USP9X silencing induced mitochondrial fission
in prostate cancer cells, with a concomitant increase in DRP1
phosphorylation. The production of ATP by mitochondria is also
important for cancer cell migration and invasion. During cell
migration, the energy demands in different regions of the cell
change. Under these circumstances, the mitochondria are cleaved by
DRP1 into smaller segments due to the increased energy requirements
(35,36). Mitochondrial fission directs the
mitochondria to localize in neuronal areas that are predicted to
have higher ATP consumption (37).
It has also been reported that DRP1 is involved in cancer invasion
and migration (38–40). Thus, based on the present findings,
it is proposed that USP9X downregulation promotes invasion and
migration through the induction of MMP9 and mitochondrial fission,
which, to the best of our knowledge, has not been reported in other
types of cancer.
To further elucidate how USP9X induces MMP9 and
p-DRP1, several upstream signaling pathways were tested, and ERK
signaling was revealed to be upregulated following the silencing of
USP9X. The association between ERK and MMP9 has been demonstrated
in various types of cells, including prostate cancer cell lines
(41). ERK activation may also
induce DRP1 phosphorylation and mitochondrial fission, which
further promotes cancer cell invasion and drug resistance (40,42).
The present findings further confirmed the association between ERK
and MMP9/p-DRP1 using an inhibitor of the ERK pathway. The role of
USP9X in cancer invasion/migration has scarcely been examined. To
date, to the best of our knowledge, only a single study is
available that suggests that miR-26b induces EMT through the
downregulation of USP9X (43). In
the present study, EMT markers, including E-cadherin and vimentin,
were examined in prostate cancer cells, and no significant changes
were observed in their levels. Therefore, EMT does not appear to
serve a role in USP9X-regulated prostate cancer cell invasion and
migration. These data suggest that USP9X inhibits prostate cancer
invasion through the inhibition of ERK/MMP9/DRP1 signaling.
Two studies have reported on the role of USP9X
inhibitors in cancer. In one study, the USP9X inhibitor WP1130
resulted in a decrease in the tumor growth in prostate cancer mouse
xenograft models (44).
Furthermore, USP9X inhibitor ABT-737 disrupted the interaction
between USP9X and induced myeloid leukemia cell differentiation
protein Mcl-1, and enhanced the antitumor activity of gemcitabine
(45). However, the effects of
WP1130 and ABT-737 on USP9X are not specific. WP1130 induces rapid
proteasomal-dependent degradation of the c-Myc proto-oncogene
protein. Additionally, it regulates the stability of
tyrosine-protein kinase JAK2. The compound directly inhibits the
deubiquitinating activity of USP9X, USP5, USP14, and ubiquitin
carboxyl-terminal hydrolase isozymes L1 and L5. ABT-737 is a BH3
mimetic inhibitor of apoptosis regulator Bcl-2 and Bcl-2-like
proteins 1 and 2. Furthermore, these reports mainly focused the
role of USP9X on tumor growth. By contrast, the present results
demonstrated that USP9X has a marked effect on invasion, and an
involvement in cell proliferation, in prostate cancer cells.
In conclusion, the results of the present study
suggest that USP9X is downregulated in prostate cancer and
functions as an inhibitor of tumor cell invasion and migration,
possibly through the regulation of the ERK/MMP9 and ERK/DRP1
signaling pathways.
Acknowledgements
Not applicable.
Funding
This study was supported by the Joint Project of
Science and Technology, Department of Yunnan and Kunming Medical
University, Kunming, China (grant no. 2017FE468), the National
Natural Science Foundation of China (grant nos. 81660422 and
81660423) and the Doctor Newcomer Award Project (grant no.
60116090706).
Availability of data and materials
All data generated or analyzed during this study are
included in this published article.
Authors' contributions
JZ, HW and BH performed the experiments, evaluated
the data, wrote the manuscript and prepared the figures. JW
designed the experiments. TL evaluated the data and wrote the
manuscript. YZ and JC performed the surgery, evaluated the
experiments and provided tissue samples. HZ and ZY performed and
evaluated the experiments. All authors reviewed the manuscript. All
authors read and approved the manuscript and agree to be
accountable for all aspects of the research in ensuring that the
accuracy or integrity of any part of the work are appropriately
investigated and resolved.
Ethics approval and consent to
participate
This study was approved by the Ethics Committee of
the Second Affiliated Hospital of Kunming Medical University,
Kunming, China. Informed written consent was obtained from all
participants.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Bray F, Ferlay J, Soerjomataram I, Siegel
RL, Torre LA and Jemal A: Global cancer statistics 2018: GLOBOCAN
estimates of incidence and mortality worldwide for 36 cancers in
185 countries. CA Cancer J Clin. 68:394–424. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Karantanos T, Corn PG and Thompson TC:
Prostate cancer progression after androgen deprivation therapy:
Mechanisms of castrate resistance and novel therapeutic approaches.
Oncogene. 32:5501–5511. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Zeng H, Chen W, Zheng R, Zhang S, Ji JS,
Zou X, Xia C, Sun K, Yang Z, Li H, et al: Changing cancer survival
in China during 2003-15: A pooled analysis of 17 population-based
cancer registries. Lancet Glob Health. 6:e555–e567. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Hudson SV, O'Malley DM and Miller SM:
Achieving optimal delivery of follow-up care for prostate cancer
survivors: Improving patient outcomes. Patient Relat Outcome Meas.
6:75–90. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Goodwin JF, Kothari V, Drake JM, Zhao S,
Dylgjeri E, Dean JL, Schiewer MJ, McNair C, Jones JK, Aytes A, et
al: DNA-PKcs-mediated transcriptional regulation drives prostate
cancer progression and metastasis. Cancer Cell. 28:97–113. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Fraile JM, Quesada V, Rodriguez D, Freije
JM and López-Otín C: Deubiquitinases in cancer: New functions and
therapeutic options. Oncogene. 31:2373–2388. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Salmena L and Pandolfi PP: Changing venues
for tumour suppression: Balancing destruction and localization by
monoubiquitylation. Nat Rev Cancer. 7:409–413. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Oosterkamp HM, Hijmans EM, Brummelkamp TR,
Canisius S, Wessels LF, Zwart W and Bernards R: USP9X
downregulation renders breast cancer cells resistant to tamoxifen.
Cancer Res. 74:3810–3820. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pérez-Mancera PA, Rust AG, van der Weyden
L, Kristiansen G, Li A, Sarver AL, Silverstein KA, Grützmann R,
Aust D, Rümmele P, et al: The deubiquitinase USP9X suppresses
pancreatic ductal adenocarcinoma. Nature. 486:266–270. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Thanh Nguyen H, Andrejeva D, Gupta R,
Choudhary C, Hong X, Eichhorn PJ, Loya AC and Cohen SM:
Deubiquitylating enzyme USP9× regulates hippo pathway activity by
controlling angiomotin protein turnover. Cell Discov. 2:160012016.
View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Wang Y, Liu Y, Yang B, Cao H, Yang CX,
Ouyang W, Zhang SM, Yang GF, Zhou FX, Zhou YF, et al: Elevated
expression of USP9X correlates with poor prognosis in human
non-small cell lung cancer. J Thorac Dis. 7:672–679.
2015.PubMed/NCBI
|
|
12
|
Fu X, Xie W, Song X, Wu K, Xiao L, Liu Y
and Zhang L: Aberrant expression of deubiquitylating enzyme USP9X
predicts poor prognosis in gastric cancer. Clin Res Hepatol
Gastroenterol. 41:687–692. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2−ΔΔCT method. Methods. 25:402–408. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Partin AW, Kattan MW, Subong EN, Walsh PC,
Wojno KJ, Oesterling JE, Scardino PT and Pearson JD: Combination of
prostate-specific antigen, clinical stage, and Gleason score to
predict pathological stage of localized prostate cancer. A
multi-institutional update. JAMA. 277:1445–1451. 1997. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Fine SW: Evolution in prostate cancer
staging: Pathology updates from AJCC 8th edition and opportunities
that remain. Adv Anat Pathol. 25:327–332. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Braunhut BL, Punnen S and Kryvenko ON:
Updates on grading and staging of prostate cancer. Surg Pathol
Clin. 11:759–774. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Egeblad M and Werb Z: New functions for
the matrix metalloproteinases in cancer progression. Nat Rev
Cancer. 2:161–174. 2002. View
Article : Google Scholar : PubMed/NCBI
|
|
18
|
Jia D, Park JH, Jung KH, Levine H and
Kaipparettu BA: Elucidating the metabolic plasticity of cancer:
Mitochondrial reprogramming and hybrid metabolic states. Cells.
7:doi: 10.3390/cells7030021. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Paupe V and Prudent J: New insights into
the role of mitochondrial calcium homeostasis in cell migration.
Biochem Biophys Res Commun. 500:75–86. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Zhou C, Wang Y, Peng J, Li C, Liu P and
Shen X: SNX10 plays a critical role in MMP9 secretion via
JNK-p38-ERK signaling pathway. J Cell Biochem. 118:4664–4671. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Huang CY, Chiang SF, Chen WT, Ke TW, Chen
TW, You YS, Lin CY, Chao KSC and Huang CY: HMGB1 promotes
ERK-mediated mitochondrial Drp1 phosphorylation for chemoresistance
through RAGE in colorectal cancer. Cell Death Dis. 9:10042018.
View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Wood SA, Pascoe WS, Ru K, Yamada T,
Hirchenhain J, Kemler R and Mattick JS: Cloning and expression
analysis of a novel mouse gene with sequence similarity to the
Drosophila fat facets gene. Mech Dev. 63:29–38. 1997.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Toloczko A, Guo F, Yuen HF, Wen Q, Wood
SA, Ong YS, Chan PY, Shaik AA, Gunaratne J, Dunne MJ, et al:
Deubiquitinating enzyme USP9X suppresses tumor growth via LATS
kinase and core components of the hippo pathway. Cancer Res.
77:4921–4933. 2017.PubMed/NCBI
|
|
24
|
McGarry E, Gaboriau D, Rainey MD,
Restuccia U, Bachi A and Santocanale C: The deubiquitinase USP9X
maintains DNA replication fork stability and DNA damage checkpoint
responses by regulating CLASPIN during S-phase. Cancer Res.
76:2384–2393. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Wu Y, Yu X, Yi X, Wu K, Dwabe S, Atefi M,
Elshimali Y, Kemp KT II, Bhat K, Haro J, et al: Aberrant
phosphorylation of SMAD4 Thr277-mediated USP9×-SMAD4 interaction by
free fatty acids promotes breast cancer metastasis. Cancer Res.
77:1383–1394. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Schwickart M, Huang X, Lill JR, Liu J,
Ferrando R, French DM, Maecker H, O'Rourke K, Bazan F,
Eastham-Anderson J, et al: Deubiquitinase USP9X stabilizes MCL1 and
promotes tumour cell survival. Nature. 463:103–107. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ding X, Yang DR, Xia L, Chen B, Yu S, Niu
Y, Wang M, Li G and Chang C: Targeting TR4 nuclear receptor
suppresses prostate cancer invasion via reduction of infiltrating
macrophages with alteration of the TIMP-1/MMP2/MMP9 signals. Mol
Cancer. 14:162015. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Moroz A, Delella FK, Almeida R, Lacorte
LM, Fávaro WJ, Deffune E and Felisbino SL: Finasteride inhibits
human prostate cancer cell invasion through MMP2 and MMP9
downregulation. PLoS One. 8:e847572013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Kato T, Fujita Y, Nakane K, Kojima T,
Nozawa Y, Deguchi T and Ito M: ETS1 promotes chemoresistance and
invasion of paclitaxel-resistant, hormone-refractory PC3 prostate
cancer cells by up-regulating MDR1 and MMP9 expression. Biochem
Biophys Res Commun. 417:966–971. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
London CA, Sekhon HS, Arora V, Stein DA,
Iversen PL and Devi GR: A novel antisense inhibitor of MMP-9
attenuates angiogenesis, human prostate cancer cell invasion and
tumorigenicity. Cancer Gene Ther. 10:823–832. 2003. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Zhang X, Yao X, Qin C, Luo P and Zhang J:
Investigation of the molecular mechanisms underlying metastasis in
prostate cancer by gene expression profiling. Exp Ther Med.
12:925–932. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Clarke NW, Hart CA and Brown MD: Molecular
mechanisms of metastasis in prostate cancer. Asian J Androl.
11:57–67. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Pollard TD and Cooper JA: Actin, a central
player in cell shape and movement. Science. 326:1208–1212. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Westermann B: Mitochondrial fusion and
fission in cell life and death. Nat Rev Mol Cell Biol. 11:872–884.
2010. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Zhao J, Zhang J, Yu M, Xie Y, Huang Y,
Wolff DW, Abel PW and Tu Y: Mitochondrial dynamics regulates
migration and invasion of breast cancer cells. Oncogene.
32:4814–4824. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Pal AD, Basak NP, Banerjee AS and Banerjee
S: Epstein-Barr virus latent membrane protein-2A alters
mitochondrial dynamics promoting cellular migration mediated by
Notch signaling pathway. Carcinogenesis. 35:1592–1601. 2014.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Hollenbeck PJ and Saxton WM: The axonal
transport of mitochondria. J Cell Sci. 118:5411–5419. 2005.
View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Yin M, Lu Q, Liu X, Wang T, Liu Y and Chen
L: Silencing Drp1 inhibits glioma cells proliferation and invasion
by RHOA/ROCK1 pathway. Biochem Biophys Res Commun. 478:663–668.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Zhang J, Zhang Y, Wu W, Wang F, Liu X,
Shui G and Nie C: Guanylate-binding protein 2 regulates
Drp1-mediated mitochondrial fission to suppress breast cancer cell
invasion. Cell Death Dis. 8:e31512017. View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Fu L, Dong Q, He J, Wang X, Xing J, Wang
E, Qiu X and Li Q: SIRT4 inhibits malignancy progression of NSCLCs,
through mitochondrial dynamics mediated by the ERK-Drp1 pathway.
Oncogene. 36:2724–2736. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
Kato T, Fujita Y, Nakane K, Mizutani K,
Terazawa R, Ehara H, Kanimoto Y, Kojima T, Nozawa Y, Deguchi T and
Ito M: CCR1/CCL5 interaction promotes invasion of taxane-resistant
PC3 prostate cancer cells by increasing secretion of MMPs 2/9 and
by activating ERK and Rac signaling. Cytokine. 64:251–257. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Cai J, Wang J, Huang Y, Wu H, Xia T, Xiao
J, Chen X, Li H, Qiu Y, Wang Y, et al: ERK/Drp1-dependent
mitochondrial fission is involved in the MSC-induced drug
resistance of T-cell acute lymphoblastic leukemia cells. Cell Death
Dis. 7:e24592016. View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Shen G, Lin Y, Yang X, Zhang J, Xu Z and
Jia H: MicroRNA-26b inhibits epithelial-mesenchymal transition in
hepatocellular carcinoma by targeting USP9X. BMC Cancer.
14:3932014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Wang S, Kollipara RK, Srivastava N, Li R,
Ravindranathan P, Hernandez E, Freeman E, Humphries CG, Kapur P,
Lotan Y, et al: Ablation of the oncogenic transcription factor ERG
by deubiquitinase inhibition in prostate cancer. Proc Natl Acad Sci
USA. 111:4251–4256. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang C, Cai TY, Zhu H, Yang LQ, Jiang H,
Dong XW, Hu YZ, Lin NM, He QJ and Yang B: Synergistic antitumor
activity of gemcitabine and ABT-737 in vitro and in vivo through
disrupting the interaction of USP9X and Mcl-1. Mol Cancer Ther.
10:1264–1275. 2011. View Article : Google Scholar : PubMed/NCBI
|