Introduction
Magnetic resonance imaging (MRI) is a widely adopted
method used to conduct clinical tumor examination. Specifically,
diffusion MRI is utilized for the early diagnosis of tumors and
brain infarctions (1-3).
The aim of the current study was to evaluate the potential of
polyethylene glycol (PEG) phantoms for use as a standard phantom
for restricted diffusion in MRI.
In a previous study, Matsuya et al (4) revealed that PEG phantoms may be used as
the standard phantom for apparent diffusion coefficient (ADC). ADC,
which indicates the average area that a water molecule moves per
second, is calculated using diffusion-weighted MR images. In the
body, there are two types of water-molecule diffusion (5). The first is free diffusion, which is
defined as water-molecule movement in free space without the
restriction of barriers. Free diffusion in solution is affected by
temperature, viscosity and various other factors. The second type
is restricted diffusion, which is defined as water-molecule
movement that is impeded by membrane structures of cells in the
body (6). ADC accounts for both free
and restricted diffusion in MR images.
In recent years, new MR imaging techniques that
represent restricted diffusion have been developed. One example of
this is diffusion kurtosis imaging (DKI) (7) which has been developed and has been
revealed to be as useful as ADC maps in certain clinical trials. A
new technique for imaging restricted diffusion has recently been
proposed; the ADC subtraction method (ASM) (8).
The current study aimed to evaluate the usability of
polyethylene glycol (PEG) phantoms as standard phantoms for
restricted diffusion in MRI. It was revealed that PEG phantoms may
have the potential to be used as the standard phantoms, not only
for ADC, but also for restricted diffusion in DKI and ASM. The
current study details these phantoms and provides formulae for
calculating PEG concentrations that enable us to create phantoms
with arbitrary ADC, DKI and ASM values mimicking various tissues
and tumors.
Materials and methods
Polyethylene glycol (PEG) phantom
The phantom (4)
comprised: i) PEG (P3640-500G; Sigma-Aldrich; Merck KGaA) as a
diffusion modifier; ii) NaN3 (Katayama Chemical Industry
Co., Ltd.) as an antiseptic; and iii) distilled water.
The phantom solution was heated and diluted using
distilled water to achieve concentrations of 20, 40, 60, 80, 100
and 120 mM with 0.03% w/w NaN3. The solution was
transferred to microcuvettes (Halbmikro 1.5 ml; Greiner
Labortechnik Manufacturing Ltd.) and used as the PEG phantoms.
Subsequently, the cuvettes were installed in a phantom container
(length, 12.5 cm; width, 10.5 cm; and height 9.5 cm) and were
immersed in physiological saline (0.9% NaCl).
Bio-phantom
A bio-phantom (9) was
prepared using Jurkat cells (Bio Resource Center) to ensure that
the range of ADC, DKI and ASM values fall within the range of those
of the PEG phantoms. Cells were cultured in 10% fetal bovine serum
(Filtron Pty Ltd.), 1% penicillin-streptomycin-neomycin (Gibco;
Thermo Fisher Scientific, Inc.) and RPMI-1640 medium (pH 7.4;
Gibco; Thermo Fisher Scientific, Inc.). The incubation was carried
out at 37˚C with 5% CO2. The number of cells with a
diameter >8 µm was counted using an electric cell counter
(Beckman Coulter, Inc.) prior to bio-phantom preparation, as the
diameter of the majority of Jurkat cells is >8 µm (mean, 9.6
µm). After measuring the cell number, the cell solution was
condensed to ~0.89 ml, transferred to a micro-cuvette (Halbmikro
1.5 ml; Greiner Labortechnik Manufacturing Ltd.), and centrifuged
at 161 x g for 5 min. Thereafter, the supernatant was removed and
the resulting bio-phantom was composed of pellet-like cells
(density, ~1-8x108 cells/ml).
MRI device, image analysis software
and statistical analysis software
A 3.0T MRI device (MAGNETOM Prisma VE11C; Siemens
AG) was used which had a 20-channel head/neck coil. ImageJ 1.52a
(National Institute of Health) was used for image analysis.
Diffusional Kurtosis Estimator (DKE) version 2.6 was used for DKI
image analysis.
Phantom-heating device
The phantom's temperature was adjusted to a
temperature close to that of the human body (~37˚) by installing
the phantom container in a heating device (constructed in-house)
using ethylene-vinyl acetate copolymer. This heating device was
connected to a circulating thermostatic chamber (Therm-Mate BF-41;
Yamato Scientific Co., Ltd.).
Temperature measurement during
MRI
An optic fiber thermometer (FLUOROPTICTM
m3300; Luxtron Co.) was installed in the microcuvette in the
phantom container and was utilized to monitor the phantom's
temperature in real time.
Imaging conditions
Table I lists the
imaging conditions for DKI and ASM. In DKI, single shot-echo planar
imaging (SS-EPI) was used in three sequences; DKI-1, -2 and -3.
 | Table IImaging conditions of DKI and
ASM. |
Table I
Imaging conditions of DKI and
ASM.
| | DKI | ASM |
|---|
| Parameters | DKI-1 | DKI-2 | DKI-3 | RESOLVE-basic | RESOLVE-modify |
|---|
| Diffusion time
(msec) | 28.9 | 28.9 | - | 39.3 | 46.0 |
| δ (msec) | 13.8 | 13.8 | - | 5.6 | 15.6 |
| Δ (msec) | 33.5 | 33.5 | - | 41.2 | 51.2 |
| b-value
(sec/mm²) | 0, 500, 1,000 | 0, 500, 1,000 | 0 | 0, 500, 1,000 | 0, 500, 1,000,
10,000 |
| Imaging time
(min:sec) | 6:24 | 6:24 | 1:12 | 13:28 | 19:06 |
| TE (msec) | 75 | 75 | 75 | 86 | 106 |
| TR (msec) | 6,000 | 6,000 | 6,000 | 8,000 | 8,000 |
| ES (msec) | 0.93 | 0.93 | 0.93 | 0.56 | 0.56 |
| FOV (mm) | 120 | 120 | 120 | 120 | 120 |
| Matrix | 82x82 | 82x82 | 82x82 | 224x224 | 224x224 |
| BW (Hz/pixel) | 1,220 | 1,220 | 1,220 | 399 | 399 |
| Averages | 1 | 1 | 9 | 2 | 2 |
| Segments | 1 | 1 | 1 | 7 | 7 |
| Slice thickness
(mm) | 5 | 5 | 5 | 5 | 5 |
| Slice number | 5 | 5 | 5 | 1 | 1 |
| Phase
direction | AP | AP | AP | AP | AP |
In ASM, two types of readout segmentation of long
variable echo-train (RESOLVE) sequences were used; RESOLVE-basic
and -modify. By changing the number of b-values, two types of DWI
were obtained for ASM. For RESOLVE-basic sequences the b-values
were set to different points; 0, 500 and 1,000 sec/mm2,
and for RESOLVE-modify the b-values were set to 4 points: 0, 500,
1,000 and 10,000 sec/mm2 (Table I). As the number of b-values varied,
the δ (motion probing gradient (MPG) pulse duration) and Δ (MPG
pulse spacing) of both sequences changed. In the equation used to
calculate b-values (equation 1), ‘Δ-δ/3’ is termed the effective
diffusion time and represents the time during which diffusion
phenomena are observed.
Equation 1:
b=γ2G2δ2 x (Δ-δ/3)
In the above equation, γ represents the gyromagnetic
ratio of protons and G is the gradient magnetic field strength. The
10,000 sec/mm2 b-value for RESOLVE-modify was used to
lengthen the effective diffusion time. The effective diffusion
times of RESOLVE-basic and RESOLVE-modify were set to 39.3 and 46.0
msec, respectively. Imaging of the PEG phantom was performed 60
times for ADC and ASM and 45 times for DKI.
Image processing of DKI. The DKI analysis
software DKE (version 2.6) is available on the website of the
Medical University of South Carolina (https://medicine.musc.edu/departments/centers/cbi/dki/dki-data-processing).
The DWIs obtained by the imaging of DKI-1, -2 and -3 were processed
using DKE to produce a mean kurtosis (MK) image (equation 2), which
is a mean value of the spatial direction. By interpolated
processing, the voxel size of the MK image becomes 1.0x1.0x1.0
mm.
Equation 2: S=S0 x exp (-b x
ADC+b2 x ADC2 x MK/6)
In equation 2, S represents signal intensity and
S0 is the signal intensity when the b-value=0
sec/mm2. The b-values used are exhibited in Table I.
Image processing of ASM
The ADC values (ADCb and ADCm)
were calculated for RESOLVE-basic using the b-values: 0, 500 and
1,000 sec/mm2 and for RESOLVE-modify using the above 3
b-values from 0 to 1,000 sec/mm2 only. The 10,000
sec/mm2 b-value for RESOLVE-modify was not used to
calculate the ADCm value. The formula used to calculate
ASM is as follows:
Equation 3:
ASM=|ADCb-ADCm|/(ADCb)3
Image evaluation
For MK images, MK values were determined from 1
region of interest (ROI) of 2x2 pixels in the PEG phantom and from
ROIs of the same size in ≤6 areas per image of the physiological
saline portion inside the phantom container.
Regarding ADCb and ASM, the signal
intensity was determined from one ROI of 6x6 pixels selected in the
PEG phantom, and from ROIs of the same size in ≤6 areas for each
image of the physiological saline portion inside the phantom
container. For DWI, signal intensity was determined from ROIs of
6x6 pixels in the images of RESOLVE-basic and RESOLVE-modify of
each b-value. Each signal intensity value was logarithmically
transformed. Subsequently, the ADC value for each ROI was
calculated using the inverse of the slope and the ASM value was
calculated using equation 3.
The total ROIs used for the calculation of the
ADCb and ASM values was 74 for physiological saline, 6
for 20 mM, 12 for 40 mM, 6 for 60 mM, 14 for 80 mM, 6 for 100 mM
and 16 for 120 mM PEG. The ROIs used for the calculation of MK
values were 57 for physiological saline, 6 for 20 mM, 6 for 40 mM,
6 for 60 mM, 9 for 80 mM, 6 for 100 mM and 12 for 120 mM PEG.
To evaluate the same values for the bio-phantom, the
MK values were determined from three ROIs of 4x1 pixels in the
cellular section inside the bio-phantom, and from ROIs of the same
size in 6 areas of the physiological saline portion inside the
phantom container.
With regard to ADCb and ASM, the signal
intensity was determined from four ROIs of 3x3 pixels selected in
the cellular section of the bio-phantom. The signal intensity was
also determined from 8 3x3-pixel ROIs located in the physiological
saline portion of the phantom container. For DWI, signal intensity
was determined from ROIs of 3x3 pixels in the images of
RESOLVE-basic and RESOLVE-modify of each b-value. Each signal
intensity value was logarithmically transformed. Then, the ADC
value for each ROI was calculated from the inverse of the slope and
the ASM value was calculated using Equation 3. The total ROIs used
for the calculation of the ADC and ASM values was 12, 24 and 20 for
cellularities of 46,522,778, 158,267,558 and 741,367,436 cells/ml,
respectively. The ROIs used for the calculation of MK values were
9, 18 and 15 for cellularities of 46,522,778, 158,267,558 and
741,367,436 cells/ml, respectively.
Results
The mean temperature and SD inside the bio-phantom
was maintained at 37.2±0.7˚C during imaging.
ADCb, MK and ASM values of
PEG phantoms
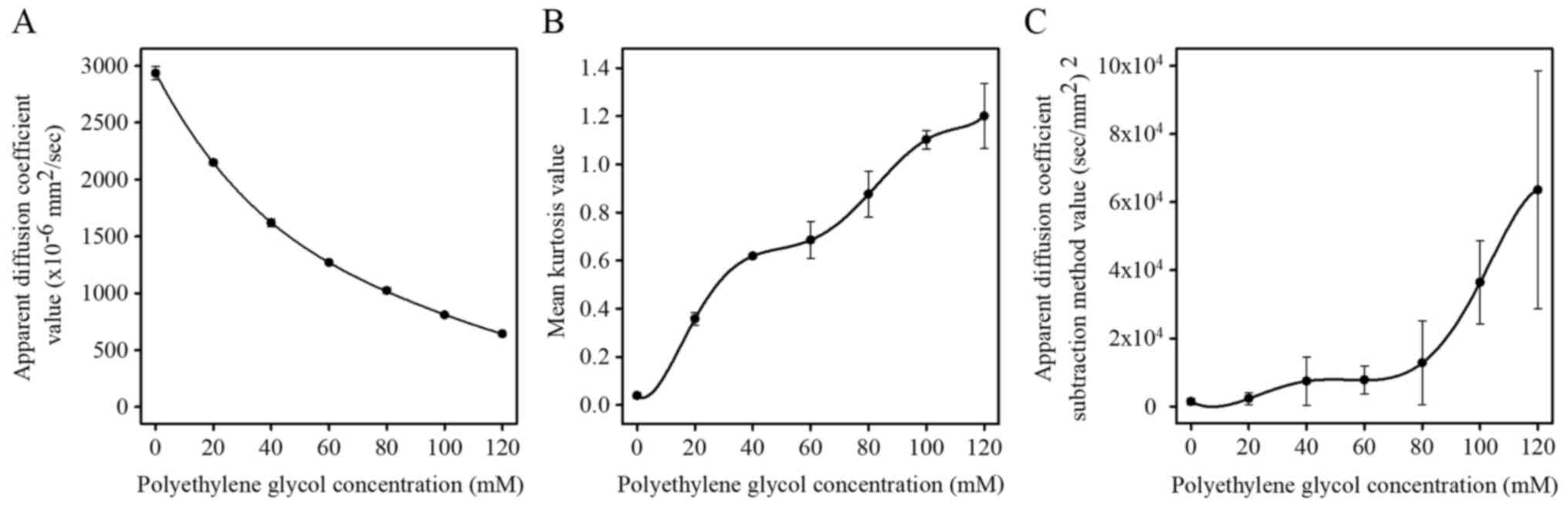
An increase in PEG concentration was associated with
a decrease in ADCb values (Fig. 1A). Conversely, MK (Fig. 1B) and ASM (Fig. 1C) values increased in line with
increasing PEG concentrations. The mean ADCb and SD
values of the 0 mM PEG phantom were 2.93±0.06x10-3
mm2/sec, and those for the 120 mM PEG phantom were
0.64±0.01x10-3 mm2/sec. MK values
corresponding to the 0 mM and 120 mM PEG phantom were 0.04±0.01 and
1.20±0.14, respectively. The ASM calculations for the 0 mM and 120
mM PEG concentrations yielded 1,412±965 and 63,544±34,965
(sec/mm2)2, respectively.
ADCb, MK and ASM values of
bio-phantoms
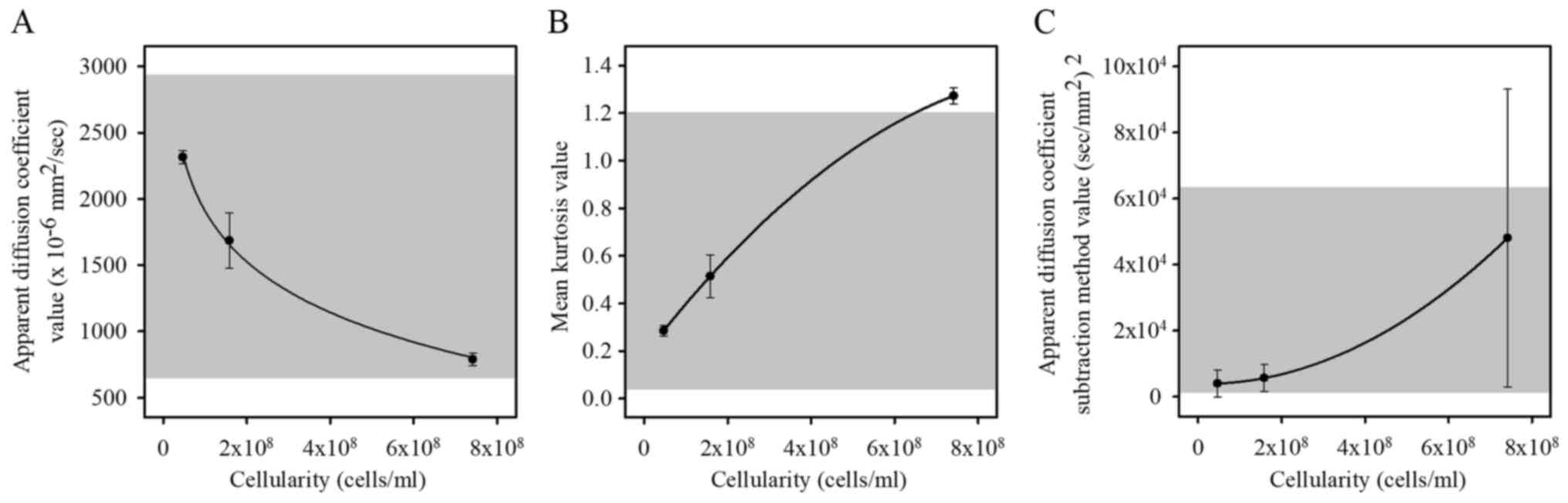
The final cellularity of the bio-phantom ranged from
0.47x108-7.41x108 cells/ml. The mean
ADCb and SD values for the bio-phantoms were
2.31±0.05x10-3 mm2/sec for the
low-cellularity phantom and 0.79±0.05x10-3
mm2/sec for the high-cellularity phantom (Fig. 2A). MK values ranged between 0.29±0.02
and 1.27±0.03 (Fig. 2B). ASM values
ranged between 3,987±3,991 and 48,039±45,071
(sec/mm2)2 (Fig.
2C). The values for the bio-phantoms mostly lie within the
range of the values produced for the PEG phantoms. Fig. 2 indicates the correlation between
cellularity and ADCb, MK, and ASM values; moreover, all
data (except for the highest MK cellularity values) were
ascertained to be within the range of the PEG phantom results.
Fig. 3 displays
ADCb, MK and ASM images of PEG and bio-phantoms. The
ADCb and ASM images provide a higher resolution and less
distortion compared with the MK image.
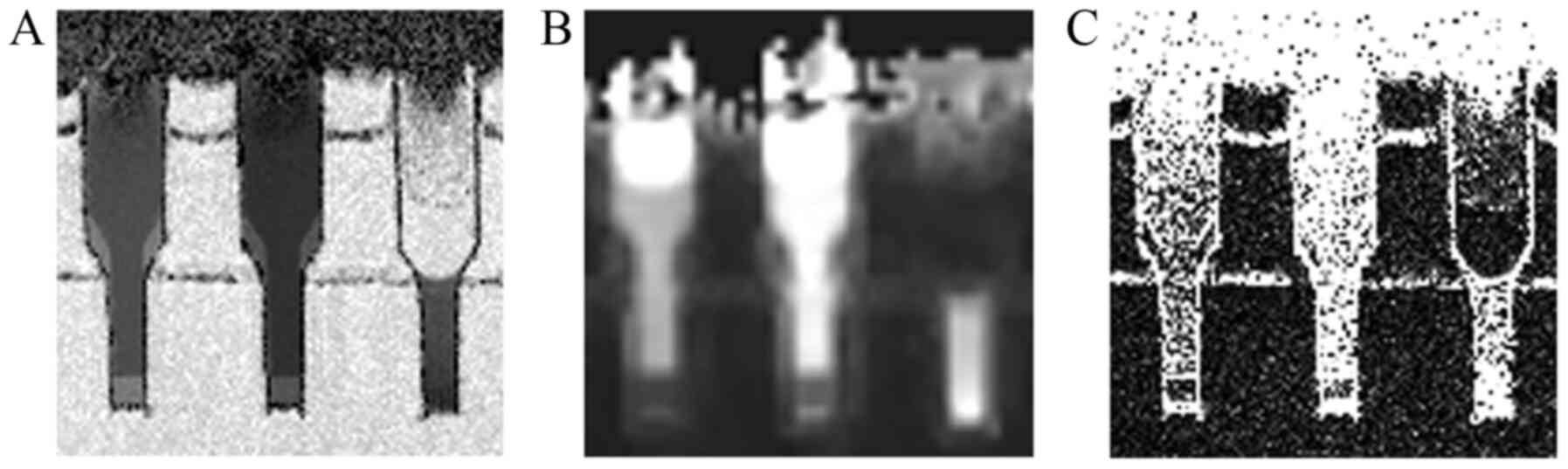 | Figure 3ADCb, MK and ASM images of
PEG and bio-phantoms. (A) ADCb image. (B) MK image. (C)
ASM image. In each figure, the left, middle and right phantoms are
80 mM PEG, 120 mM PEG and high-cellularity (741,367,436 cells/ml)
bio-phantom, respectively. ADCb and ASM images provide a
higher resolution and less distortion compared to the MK image, due
to the use of RESOLVE sequence. ADCb, apparent diffusion
coefficient calculated for readout segmentation of long variable
echo-trains-basic; MK, mean kurtosis; ASM, apparent diffusion
coefficient subtraction method; PEG, polyethylene glycol; RESOLVE,
readout segmentation of long variable echo-trains. |
Empirical formula for calculating
ADCb values of phantoms made using varying PEG
concentrations
ADCb values for PEG phantoms with
concentrations of 0, 20, 40, 60, 80, 100 and 120 mM were plotted as
in Fig. 1A. Using these data points,
a regression curve and an empirical formula were constructed
facilitating the calculation of the ADCb values from
phantoms produced using varying PEG concentrations. The correlation
factor (R2) for this approximation was 1.00. The
empirical formula to calculate the ADCb value
corresponding to any PEG concentration (x) is as follows:
Formula 1:
ADCb=a1x4+
a2x3+a3x2+a4x+a5
(a1,
0.0000066786969390685x10-6; a2, -0.0028279562
3507326000x10-6; a3,
0.484704389217086x10-6; a4,
-48.0980342756939x10-6; a5,
2,935.33497680997x10-6).
Empirical formula for calculating MK
values of phantoms made using varying PEG concentrations
MK values for PEG phantoms with concentrations of 0,
20, 40, 60, 80, 100 and 120 mM phantoms were plotted as in Fig. 1B. Using these data points, a
regression curve and an empirical formula were created for
calculating the MK values of phantoms made with varying PEG
concentrations. The correlation factor (R2) for this
approximation was 1.00. The empirical formula to calculate the MK
value corresponding to any PEG concentration (x) is as follows:
Formula 2:
MK=b1x6+b2x5+b3x4+b4x3+b5x2+b6x+b7
(b1, 0.00000000002574679427;
b2, -0.000000009958 61887106; b3,
0.00000144033983524861; b4, -0.00009462031 704821070;
b5, 0.00263877966804671000; b6, -0.0089848099
7102524000; b7, 0.03857894408353960000).
Empirical formula for calculating ASM
values of phantoms made using varying PEG concentrations
ASM values for PEG phantoms with concentrations of
0, 20, 40, 60, 80, 100 and 120 mM phantoms were plotted as in
Fig. 1C. Using these data points, a
regression curve and an empirical formula were created for
calculating the ASM values of phantoms made with varying PEG
concentrations. The correlation factor (R2) for this
approximation was 1.00. The empirical formula to calculate the ASM
value corresponding to any PEG concentration (x) is as follows:
Formula 3:
ASM=c1x6+c2x5+c3x4+c4x3+c5x2+c6x+c7
(c1, -0.0000004118023456; c2,
0.0000866210497676; c3, -0.0018022499380095;
c4, -0.5404589664540250; c5,
34.3619915451854000; c6, -423.5577461123460000;
c7, 1,411.8674192056000000).
Empirical formula for calculating
ADCb values of bio-phantoms with any cellularity
ADCb values for bio-phantoms were plotted
as in Fig. 2A. Using these data
points, a regression curve and an empirical formula were created
for calculating the ADCb values of bio-phantoms made
with any cellularity. The correlation factor (R2) for
this approximation was 1.00. The empirical formula to calculate the
ADCb value corresponding to any cellularity (y) is as
follows:
Formula 4:
ADCb=d1ln(y)+d2
(d1, -552.230079387058x10-6;
d2, 12,081.0508279336 00x10-6).
Empirical formula for calculating MK
values of bio-phantoms with varying cellularity
MK values for bio-phantoms are plotted in Fig. 2B and the data points were used to
create a regression curve and an empirical formula for the
calculation of MK values of bio-phantoms made with varying
cellularity. The correlation factor (R2) for this
approximation was 1.00. The empirical formula to calculate the MK
value corresponding to varying cellularity (y) is as follows:
Formula 5:
MK=e1y2+e2y+e3
(e1, -0.00000000000000000108;
e2, 0.00000000226921884 624; e3,
0.18153679067533800000).
Empirical formula for calculating ASM
values of bio-phantoms with varying cellularity
The ASM values for bio-phantoms are exhibited in
Fig. 2C. Using these data points, a
regression curve and an empirical formula were created to calculate
the ASM values of bio-phantoms made with varying cellularity. The
correlation factor (R2) for this approximation was 1.00.
The empirical formula to calculate the ASM value corresponding to
varying cellularity (y) is as follows:
Formula 6: ASM =
f1y2+f2y+f3
(f1, 0.00000000000008320968;
f2, -0.0000021617730 7991540; f3,
3,907.60230188793000000000).
Discussion
In the present study, it was revealed that
polyethylene glycol (PEG) phantoms are able to mimic a wide range
of values, not only for the apparent diffusion coefficient (ADC),
but also for restricted diffusion. Formulae were constructed in
order to calculate PEG concentrations corresponding to arbitrary
ADC, mean kurtosis (MK) and ADC subtraction method (ASM) values.
The current results indicate that PEG phantoms may be used as a
standard phantom for restricted diffusion with the desired MK and
ASM values in MRI.
PEG, which is used as the base material for our
standard phantom for ADC, MK and ASM, is a safe material and used
as a base in many everyday products, including lapactics, skin
creams and cosmetic emulsifiers (4).
PEG phantom is safer than previously reported ADC standard
phantoms-gelatinous substances such as agar (10), agarose (11) and polyacrylamide (12); and liquid solution materials such as
ethanol (13), acetone (14-17),
Gd-DTPA solution (18) and cupric
sulfate solution (19). PEG phantom
is also less expensive and easier to make than the previously
reported ADC standard phantoms.
Matsumoto et al used a bio-phantom using
tumor cells to visualize restricted diffusion (9). Bio-phantoms are complicated to create,
time consuming and expensive, and impossible to use for a long
time. The PEG phantom is ready to make, cheap and safe and can be
stored for a long time.
PEG is a high molecular compound with a structure
polymerized by ethylene glycol. Sehy et al (20) revealed that injecting PEG into cells
decreased ADC, suggesting its suppressive action on water
diffusion. In a previous study, Matsuya et al (4) proposed the use of PEG phantoms as the
standard phantom for apparent diffusion coefficient (ADC).
Empirical formulae were then constructed to calculate the PEG
concentration at any measured temperature to obtain arbitrary ADC
values for 1.5T MRI. Following the formulae, any phantoms with
arbitrary ADC values can be made to mimic any tumors at any
temperature.
ADC represents free and restricted diffusion in MR
images. In recent years, new MR imaging techniques representing
restricted diffusion have been developed. Of these, DKI is often
considered the most promising and has been suggested as an
alternative to ADC maps in clinical trials (7). A typical DKI protocol consists of a
total of 60-70 images; and more data-demanding protocols are also
available. Post-DKI data processing also requires significant time,
especially in patients requiring urgent attention (e.g., critical
cases and children). However, the superiority of DKI imaging
supports developing the clinical and preclinical studies for DKI.
Recently, faster imaging techniques for DKI have been already
developed (21).
In a recent study, ASM was proposed as a new
technique for restricted diffusion (8). ASM enables the observation of
water-molecule diffusion with a space-separation resolution of 1
µm. ASM also produces a high image quality (without distortion)
using readout segmentation of long variable echo-train (RESOLVE)
sequences (22). Under the imaging
conditions of this study, the resolution of ASM was higher than
that of DKI (Fig. 3). If ASM's image
quality were set to the same as that of DKI, ASM would take a much
shorter imaging time than DKI in this study. The PEG phantom
created in the current study is the first reported standard
restricted diffusion phantom for DKI and ASM.
The MK values for the bio-phantoms predominantly
fall within the range of the values for the PEG phantoms. Although
the phantoms in the current study did not cover the upper limit for
the bio-phantoms' MK (1.27±0.03), the maximum cell density covered
by our phantom was calculated using empirical formula 5 to be
650,892,050 cells/ml.
MK values in clinical studies using 3T MRI have been
previously reported for various tumors; for example, MK for grade
II glioma is 0.50±0.08(23); for
rectal cancer 1.000±0.112(24); for
squamous cell carcinoma 0.917±0.144; and for olfactory
neuroblastoma 1.209±0.262(25). ADC
values for common tumors in clinical studies using 3T MRI vary
between 0.62x10-3 mm2/sec (brain lymphoma)
and 2.60x10-3 mm2/sec (neoplastic cystic
lesions in the pancreas) (26). The
current phantom covers the majority of these values and can
therefore be used to mimic most common tumor types.
The ASM values of the bio-phantoms fall completely
within the range of the values for the PEG phantoms, suggesting the
usability of the phantom in the present study for future ASM
experiments. To the best of our knowledge, this is the first paper
to present a phantom covering ASM values of biological tissue since
the advent of ASM.
There are a few limitations to this study. The
equations in the present study were constructed according to values
acquired using 3T MRI and sequences available for this experiment.
Further examination may be necessary to clarify the effect of
imaging conditions, such as the type of MRI devices and sequences
used, to ensure the reproducibility of the current results.
Furthermore, the mechanism by which PEG mimics restricted diffusion
is yet to be elucidated, although there is a possibility that PEG
molecules exert a limiting effect on the movement of water
molecules. The PEG phantom, however, may have limitations in
mimicking restricted diffusion beyond a certain point, since it
does not have a concrete restricting structure (such as a
membrane), but it was revealed to mimic restricted diffusion within
a certain range of clinically reported DKI values. Therefore, this
technique may prove useful for future experiments as long as the
truncated range is accurately accounted for.
Using the newly developed MRI phantom, instead of
imaging the actual tumor in the patient's body, a new method for
imaging restricted diffusion might be developed by imaging the
phantom with the MK value and ADC value of the tumor. Unlike
clinical trials with patients, this phantom contributes to safe and
cost-effective research. Improvement of imaging methods for future
clinical dissemination of restricted diffusion imaging is expected
worldwide and we believe this phantom will contribute to such
research.
In conclusion, it was revealed that PEG decreased
restricted diffusion and this indicates the potential of the PEG
phantom as a standard phantom for restricted diffusion MRIs.
Acknowledgements
Not applicable.
Funding
The present study was partially supported by
Grants-in-Aid for Scientific Research (grant nos. C22591335,
15K09924, and 19K0809801) from the Ministry of Health, Labour and
Welfare of Japan.
Availability of data and materials
The materials and datasets used and/or analyzed
during the current study are available from the corresponding
author on reasonable request.
Authors' contributions
AKh, YY and MK conceived and designed the study,
processed the data and wrote the article. AKh, MK, YY, IS, BOB, KH,
MB, NT, KK, KS, HI, AKu, TM, SO, SK and JA participated in
conducting the experiments. IS, AKh, KH, MB and NT edited the
article. All authors read and approved the final version of the
manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that there are no competing
interests regarding the publication of the present manuscript.
References
|
1
|
Drake-Pérez M, Boto J, Fitsiori A, Lovblad
K and Vargas MI: Clinical applications of diffusion weighted
imaging in neuroradiology. Insights Imaging. 9:535–547.
2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Stecco A, Buemi F, Iannessi A, Carriero A
and Gallamini A: Current concepts in tumor imaging with whole-body
MRI with diffusion imaging (WB-MRI-DWI) in multiple myeloma and
lymphoma. Leuk Lymphoma. 59:2546–2556. 2018.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Tang L and Zhou XJ: Diffusion MRI of
cancer: From low to high b-values. J Magn Reson Imaging. 49:23–40.
2019.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Matsuya R, Kuroda M, Matsumoto Y, Kato H,
Matsuzaki H, Asaumi J, Murakami J, Katashima K, Ashida M, Sasaki T,
et al: A new phantom using polyethylene glycol as an apparent
diffusion coefficient standard for MR imaging. Int J Oncol.
35:893–900. 2009.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Le Bihan D and Iima M: Diffusion magnetic
resonance imaging: What water tells Us about biological tissues.
PLoS Biol. 13(e1002203)2015.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Le Bihan D and Johansen-Berg H: Diffusion
MRI at 25: Exploring brain tissue structure and function.
Neuroimage. 61:324–341. 2012.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Jensen JH and Helpern JA: MRI
quantification of non-Gaussian water diffusion by kurtosis
analysis. NMR Biomed. 23:698–710. 2010.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Yoshimura Y, Kuroda M, Sugianto I,
Khasawneh A, Bamgbose BO, Hamada K, Barham M, Tekiki N, Kurozumi A,
Matsushita T, et al: Development of a novel method for visualizing
restricted diffusion using subtraction of apparent diffusion
coefficient values. Mol Med Rep. 20:2963–2969. 2019.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Matsumoto Y, Kuroda M, Matsuya R, Kato H,
Shibuya K, Oita M, Kawabe A, Matsuzaki H, Asaumi J, Murakami J, et
al: In vitro experimental study of the relationship between
the apparent diffusion coefficient and changes in cellularity and
cell morphology. Oncol Rep. 22:641–648. 2009.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Li TQ, Kim DH and Moseley ME:
High-resolution diffusion-weighted imaging with interleaved
variable-density spiral acquisitions. J Magn Reson Imaging.
21:468–475. 2005.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Ogura A, Maeda F, Miyai A, Hayashi K and
Hongoh T: Effect of vibration caused by time-varying magnetic
fields on diffusion-weighted MRI. Nihon Hoshasen Gijutsu Gakkai
Zasshi. 62:565–569. 2006.PubMed/NCBI View Article : Google Scholar : (In Japanese).
|
|
12
|
Hirsch JG, Bock M, Essig M and Schad LR:
Comparison of diffusion anisotropy measurements in combination with
the flair-technique. Magn Reson Imaging. 17:705–716.
1999.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Deng J, Omary RA and Larson AC: Multishot
diffusion-weighted Splice Propeller MRI of the abdomen. Magn Reson
Med. 59:947–953. 2008.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yoshikawa T, Kawamitsu H, Mitchell DG,
Ohno Y, Ku Y, Seo Y, Fujii M and Sugimura K: ADC measurement of
abdominal organs and lesions using parallel imaging technique. Am J
Roentgenol. 187:1521–1530. 2006.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Bammer R, Stollberger R, Augustin M,
Simbrunner J, Offenbacher H, Kooijman H, Ropele S, Kapeller P, Wach
P, Ebner F and Fazekas F: Diffusion-weighted imaging with navigated
interleaved echo-planar imaging and a conventional gradient system.
Radiology. 211:799–806. 1999.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Brockstedt S, Thomsen C, Wirestam R,
Holtas S and Stahlberg F: Quantitative diffusion coefficient maps
using fast spin-echo MRI. Magn Reson Imaging. 16:877–886.
1998.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Lebihan D, Breton E, Lallemand D, Aubin
ML, Vignaud J and Lavaljeantet M: Separation of diffusion and
perfusion in intravoxel incoherent motion MR imaging. Radiology.
168:497–505. 1988.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Moteki T and Ishizaka H: Evaluation of
cystic ovarian lesions using apparent diffusion coefficient
calculated from turboFLASH MR images. Br J Radiol. 71:612–620.
1998.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Kinoshita Y, Iriguchi N and Yokota A:
Study of diffusion phenomenon using an experimental magnetic
resonance system (SIS 200/400) for small animals-reliability and
apparent diffusion coefficient of normal animals. J UOEH.
17:261–269. 1995.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Sehy JV, Ackerman JJH and Neil JJ:
Apparent diffusion of water, ions, and small molecules in the
Xenopus oocyte is consistent with Brownian displacement. Magn Reson
Med. 48:42–51. 2002.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Hansen B and Jespersen SN: Recent
developments in fast kurtosis imaging. Front Phys. 5(40)2017.
|
|
22
|
Yoshimura Y, Kuroda M, Sugiantoc I,
Bamgbosec BO, Miyahara K, Ohmura Y, Kurozumi A, Matsushita T, Ohno
S, Kanazawa S and Asaumi J: The usefulness of readout-segmented
echo-planar imaging (RESOLVE) for bio-phantom imaging using 3-tesla
clinical MRI. Acta Med Okayama. 72:53–59. 2018.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Delgado A, Fahlström M, Nilsson M,
Berntsson SG, Zetterling M, Libard S, Alafuzoff I, Van Westen D,
Lätt J, Smits A and Larsson EM: Diffusion kurtosis imaging of
gliomas grades II and III-a study of perilesional tumor
infiltration, tumor grades and subtypes at clinical presentation.
Radiol Oncol. 51:121–129. 2017.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Sun Y, Xiao Q, Hu F, Fu C, Jia H, Yan X,
Xin C, Cai S, Peng W, Wang X, et al: Diffusion kurtosis imaging in
the characterisation of rectal cancer: Utilizing the most
repeatable region-of-interest strategy for diffusion parameters on
a 3T scanner. Eur Radiol. 28:5211–5220. 2018.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Xiao Z, Tang Z, Qiang J, Qian W, Zhong Y,
Wang R, Wang J, Wu L and Tang W: Differentiation of olfactory
neuroblastomas from nasal squamous cell carcinomas using MR
diffusion kurtosis imaging and dynamic contrast-enhanced MRI. J
Magn Reson Imaging. 47:354–361. 2018.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Hara M, Kuroda M, Ohmura Y, Matsuzaki H,
Kobayashi T, Murakami J, Katashima K, Ashida M, Ohno S and Asaumi
JI: A new phantom and empirical formula for apparent diffusion
coefficient measurement by a 3 tesla magnetic resonance imaging
scanner. Oncol Lett. 8:819–824. 2014.PubMed/NCBI View Article : Google Scholar
|