Introduction
Antibiotics are central to the management of severe
infections and have transformed human health (1,2).
However, the rise and rapid spread of antimicrobial resistance
(AMR) have made numerous infections increasingly difficult to
treat. This crisis stems from the emergence and proliferation of
resistant microorganisms and is exacerbated by inappropriate use of
antibacterial agents in both clinical and community settings
(3-6).
Numerous currently available antibiotics have failed to overcome
established resistance mechanisms, which underscores the need for
prudent antimicrobial stewardship and the discovery of new agents
(7,8). There is an urgent need for prudent
antibiotic use and the development of novel antibacterial agents to
safeguard human, animal and agricultural health (9-11).
Natural products have long been recognized as an
important source of bioactive compounds for modern medicine
(12), and their unique chemical
and structural diversity provides opportunities to develop agents
with novel mechanisms of action (10,13).
Modifications of existing natural compounds can temporarily
overcome resistance (2,14), but the discovery of entirely new
natural molecules remains essential to address the global threat of
AMR (7,15,16).
Marine organisms, and particularly sponges, have attracted
increasing attention for their ability to produce antibacterial
metabolites (17-19).
The marine ecosystem represents a promising reservoir of novel
antibiotics, and there is a need to explore less-studied habitats
to meet the demand for new therapeutics (10,20,21).
Compared with terrestrial environments, marine
ecosystems offer greater potential for discovering unique bioactive
molecules (12,22,23).
The Gulf of Aqaba is a relatively isolated deep basin in the Red
Sea and exhibits unique thermal and ecological features that
distinguish it from other deep-sea regions (24,25).
Its unusual conditions support high biodiversity and make it an
exceptional setting for exploring novel bioactive metabolites
(26,27).
As sessile organisms, marine sponges are constantly
exposed to diverse microbial communities in their aquatic habitats,
including predators, biofouling microorganisms and pathogens
(28-30).
Because they lack an innate immune system, their primary defence
strategy is the production of secondary metabolites, which act as
chemical defences and enable them to adapt to environmental
pressures (9,31-33).
Symbiotic microorganisms associated with sponges also contribute to
their defence, nutrition and metabolism (34). The compounds derived from the hosts
and symbionts include terpenoids, peptides, alkaloids, macrolides
and steroids that have shown potential as drug leads for the
treatment of diseases such as malaria, cancer and infections caused
by antibiotic-resistant pathogens (8,35,36).
Numerous sponge species have demonstrated antibacterial,
anticancer, antifungal, anti-inflammatory and antimalarial
properties, supporting their pharmaceutical relevance (37-41).
Given the escalating problem of bacterial resistance, marine
biotechnology (‘blue biotechnology’) is increasingly a focus to
discover bioactive molecules from marine organisms as sources of
novel antibacterial agents (11,13).
Several studies have demonstrated the antibacterial
activity of sponge extracts against both Gram-positive and
Gram-negative bacteria (42-44).
For instance, extracts from the marine sponge Acanthella
cavernosa exhibit inhibitory effects against Staphylococcus
aureus (S. aureus), Escherichia coli (E.
coli) and Pseudomonas aeruginosa (45). Similarly, extracts from the sponge
Callyspongia plicifera have been reported to have
antibacterial activity against S. aureus, Bacillus
subtilis and Klebsiella pneumoniae (46,47).
According to studies conducted in the Gulf of Aqaba, several
shallow-water sponge species have demonstrated notable
antibacterial properties. For example, the ethanolic crude extract
of Grayella cyathophora exhibits strong activity,
particularly against Pseudomonas aeruginosa (48). By contrast, deep-sea environments,
which are characterized by high pressure, low temperature, and the
absence of light, favour the production of unique secondary
metabolites. This further supports the rationale for investigating
deep-sea sponges as promising sources of biologically active
compounds (49,50).
A recent study selected Red Sea sponge species at
shallower depths (140-290 m) and focused primarily on antibacterial
screening (51). The present
investigation provides the first comprehensive chemical; molecular
and biological characterization of deep-sea sponges collected at
previously unexplored depths (345-362 m) from the Jordanian Gulf of
Aqaba during the OceanXplorer Jordan Expedition 2022. DNA-barcoding
(28S rRNA, GenBank accession numbers PX278188.1-PX278188.3)
confirmed that the analyzed specimens represent genetically
independent lineages relative to earlier samples. LC-MS/MS
metabolite profiling revealed a distinct secondary-metabolite
fingerprint, including Hyatellaquinone, Manoalide, Motualevic acid,
Manzamine A and Popolohuanone, which were not reported in the
previous study. Furthermore, the present study broadens the
biological scope by assessing antibacterial activity against
multidrug-resistant S. aureus (including MRSA), thereby
delivering a more complete pharmacological assessment.
Collectively, the integration of deeper-water sampling, molecular
verification, metabolomic differentiation, and expanded bioassays
establishes the present study as an independent and novel
contribution to marine-derived drug-lead discovery in the Gulf of
Aqaba.
The aim of the present study was to investigate the
antibacterial potential of three deep-sea sponges collected from
the Gulf of Aqaba, Jordan, one of the deepest and relatively
unexplored marine environments in the region. By evaluating their
bioactivity and chemical composition, it was aimed to identify safe
and biocompatible agents with potential therapeutic applications.
Special emphasis was placed on the ethanolic extract of the most
active sponge, Dactylospongia cf. elegans (D. cf.
elegans), which was chemically characterized using
LC-MS/MS.
Materials and methods
Sampling of sponge specimens
In July 2022, as part of the OceanXplorer Jordan
Expedition, three deep-sea sponge samples were collected from
Stations 1 and 2, located in the deepest bottom of the Gulf of
Aqaba, Jordan, aboard the research vessel OceanXplorer (Fig. S1). Sampling was conducted at depths
ranging from 345 to 362 m using the robotic arm on the manned
submersible. Specifically, Sponge 1 was collected at 345 m, Sponge
2 at 362 m, and Sponge 3 at 362 m, as detailed in Table I, all retrieved on 14 July 2022.
Samples were immediately placed in sterile labeled containers,
frozen at -20˚C onboard and transferred on ice to the Laboratory
for Molecular and Microbial Ecology, The University of Jordan, for
subsequent analyses. Sampling was authorized under the OceanXplorer
permit no. 9795, issued by the Aqaba Special Economic Zone
Authority. The morphological characteristics of these samples are
summarized in Table SI, and
representative morphological images are provided in Fig. S2, Fig.
S3 and Fig. S4 to enhance the
visualization and clarity of Table
SI.
 | Table ISponge sample ID and their collection
depths. |
Table I
Sponge sample ID and their collection
depths.
| Sample ID | Sponge 1 | Sponge 2 | Sponge 3 |
|---|
| Depth of the
collection area | 345 m | 362 m | 345 m |
| Station No. | Station 1 | Station 2 | Station 1 |
Molecular identification of sponge
samples by DNA barcoding
To identify the sponge species, ~25 mg of tissue was
fragmented before DNA extraction using the DNeasy Blood and Tissue
Kit (Qiagen, Inc.) following established protocols (22). The 28S ribosomal RNA gene was
selected as the molecular marker (52) with primers listed in Table II. This marker was chosen because
of its reliability in resolving taxonomic relationships within the
class Demospongiae, and its successful application in previous
sponge barcoding studies (53,54).
It is acknowledged, however, that species-level identification of
Porifera should not rely solely on a single locus. Ideally, such
assignments are corroborated by morphological traits (Table SI) and, where possible, multi-locus
data (for example, COI + 28S ± ITS). Accordingly, the authors'
species assignments were made conservatively and supported by both
molecular and morphological evidence.
 | Table IIPrimer sequences for 28S ribosomal
RNA gene. |
Table II
Primer sequences for 28S ribosomal
RNA gene.
| Primer | Sequence |
|---|
| 28F63mod
(Forward) |
5'-ACCCGCTGAAYTTAAGCATATHANTMA-3' |
| 28R1072
(Reverse) |
5'-GCTATCCTGAGGGAAACTTCGG-3' |
PCR was used to amplify partial fragments of the 28S
rRNA gene, which has proven effective in classifying sponge taxa
from underexplored habitats (12,55,56).
The PCR amplification was performed using EntiLink™ PCR Master Mix
(ELK Biotechnology, Co., Ltd.) with the following cycling
condition: An initial denaturation at 94˚C for 3 min; 35 cycles of
denaturation at 94˚C for 30 sec, annealing at 56˚C for 30 sec, and
extension at 72˚C for 83 sec; followed by a final extension at 72˚C
for 10 min. Amplification products were visualized by 1% agarose
gel electrophoresis and documented using a Gel Documentation
system.
PCR products were subsequently purified and sent to
Macrogen (South Korea) for Sanger sequencing. Sequence analysis was
performed using MEGA software and BLAST searches against the
GenBank database. A phylogenetic tree was constructed by the
neighbor-joining method based on 28S rRNA sequences from closely
related sponge taxa available in GenBank (57).
Ethanolic extraction and preparation
of the sponges' crude extracts
Freshly chopped sponges were soaked in a 70%
ethanol/water solution for storage at -20˚C. An adapted extraction
procedure was performed (14,20,58).
Initially, the sponges blended while immersed in the ethanol
solution. Subsequently, the mixture was heated at 55-60˚C for 2 h
to optimally extract hydrophilic and hydrophobic compounds with
ethanol solution. The resulting solution was then filtered, and the
filtrate was concentrated using a rotary evaporator.
The concentrated extract was further processed by
lyophilization (freeze drying), resulting in a powder (59). A concentration of 25 mg/ml was
achieved by dissolving 25 mg of extracted powder in 1 ml of
Dimethyl Sulfoxide (DMSO) to create the stock solution. This liquid
was vortexed and filtered through a 0.45-µm nylon syringe filter
before being placed in a Falcon tube for storage. The stock
solution was then diluted into various quantities (5, 10, 15 and 20
mg/ml) using sterile distilled water.
Antibacterial effect of sponges'
ethanolic extracts
The antibacterial efficacy of ethanolic extract was
assessed using six bacterial strains, including S. aureus
(ATCC 29213), Staphylococcus epidermidis (S.
epidermidis; ATCC 51625) and Bacillus pumilus (isolate)
as Gram-positive bacteria, and Klebsiella aerogenes
(isolate) and E. coli (ATCC 25922) as Gram-negative
bacteria. In addition, methicillin-resistant S. aureus
(MRSA; ATCC 1026) was included as a clinically relevant resistant
strain.
The agar well diffusion method was used to evaluate
the antibacterial activity of the extracts (60). Briefly, Muller-Hinton Broth (MHB)
was inoculated with the bacterial strains and incubated at 37˚C
overnight. The bacterial culture density was adjusted to the 0.5
McFarland turbidity standard (≈1.5x108 CFU/ml) (61), and the inoculum was spread onto
Muller-Hinton agar plates. Wells (8 mm) were created in the agar
and filled with different extract concentrations (5, 10, 15 and 20
mg/ml). Plates were left at room temperature for 1-2 h to allow
pre-diffusion before incubation at 37˚C for 24 h.
After incubation, antibacterial activity was
determined by measuring the inhibition zones around the wells
(52,62). Gentamycin (10 µg) served as the
positive control for all bacteria (63,64),
and vancomycin (30 µg) was used as the MRSA-specific positive
control (65). Furthermore, 80%
DMSO was used as the negative control, as it was required to ensure
complete dissolution of the crude sponge extracts for accurate
antimicrobial testing. To exclude any solvent-related effects, pure
80% DMSO was tested against all bacterial strains and showed no
antibacterial activity. In addition, Sponge 1 extract, despite
being dissolved in 80% DMSO, exhibited no detectable antibacterial
effect, further supporting the lack of interference from the
solvent itself (66). All
experiments were performed in triplicate.
Determination of MBC and MIC
The MIC of the sponge extracts was determined using
the standard 96-well microdilution method (67). Each well contained 100 µl of
bacterial culture (~6.0 log10 CFU/ml) obtained from an overnight
culture. Following the protocol of Balouiri et al (68), serial dilutions of the extracts were
prepared in MHB. The concentration ranges were selected based on
the results of the agar well diffusion assay. For Sponge 2,
concentrations of 5, 4, 3, 2, 1, 0.5, 0.25 and 0.125 mg/ml were
tested against S. aureus and MRSA, while 10-1 mg/ml
dilutions were tested against S. epidermidis. Negative
controls (80% DMSO and un-inoculated MHB) and positive control
(bacterial suspension only) were included to ensure accuracy
(65,69). The plates were sealed and incubated
at 37˚C for 24 h. Similarly, MBC was determined as described
previously (67). The MBC assay was
designed to distinguish between bactericidal and bacteriostatic
effects of the sponge extracts. Samples from wells with
concentrations at or above the MIC were subcultured and evenly
spread onto fresh agar plates (70). The lack of bacterial colonies on the
agar after 24 h at 37˚C indicated the MBC, defined as the lowest
extract concentration capable of completely eliminating the tested
microorganism (71).
LC-MS-MS analysis of sponge
extracts
The most bioactive sponge extract's chemical makeup
was assessed utilizing LC-MS-MS. Smart Labs Group conducted this
analysis using a Shimadzu LC system that comprised the following
parts: SIL-30AC (autosampler), CBM-20A (control bus module),
LCMS-8030 (Triple Quadrupole Mass Spectrometer), LC-30AD (liquid
chromatograph), and CTO-30A (column oven). A total of ~30 mg of the
sponge extract was fully extracted using methanol (MeOH) to prepare
the sample. To purify the resulting crude extract, a solid-phase
extraction column was used. In this stage, the methanolic fraction
(100% MeOH) was preserved, whereas the aqueous fraction (100%
H2O) was discarded. A final working concentration of 2.5
mg/ml of the crude extract was used for the LC-MS-MS analysis.
Using a gradient elution protocol that lasted 15 min, the LC-MS-MS
technique moved from 100% water (containing 0.1% formic acid) to
95% acetonitrile (also containing 0.1% formic acid). Main details,
such as particular molecular ion masses (with an accuracy of <5
ppm), compound retention periods (in minutes), and MS-MS daughter
ion patterns for structural clarification, were all supplied by the
ensuing LC-MS spectra.
Statistical analysis
All data were processed and visualized using
GraphPad Prism™ software (version 10.3.0; GraphPad Software Inc.;
Dotmatics). Inhibition zone diameters obtained from the agar well
diffusion assay were expressed as the mean ± standard error of the
mean (SEM) based on at least three independent replicates. Since
the study was exploratory in nature and aimed at characterizing
biological trends rather than testing predefined hypotheses, no
formal statistical comparisons (for example, ANOVA or t-tests) were
performed. The results are therefore presented descriptively, and
MIC and MBC values are shown as representative data from replicate
experiments to illustrate reproducibility and relative potency.
Although triplicate measurements were obtained, they represent
technical replicates of the same sample and thus were not subjected
to inferential statistical testing. The study was exploratory and
descriptive in design.
Results and Discussion
Sponge identification (DNA barcoding
and phylogenetic analysis)
The 28S rRNA gene was successfully amplified from
all three sponge specimens, which produced single amplicons of
~1,000-1,300 bp and were confirmed by agarose gel electrophoresis
(Fig. S5). The purified PCR
products were sequenced, and the resulting sequences were analysed
using BLAST against the NCBI GenBank database. The BLAST results
demonstrated high similarity to known representatives of the class
Demospongiae with query coverage values of 99%. Specifically,
Sponge 1 displayed the highest similarity to Stelletta
fibrosa (90.04% identity), Sponge 2 was most similar to
Dactylospongia elegans (95.74% identity) and Sponge 3 was
most similar to Axinella polypoides (91.32% identity).
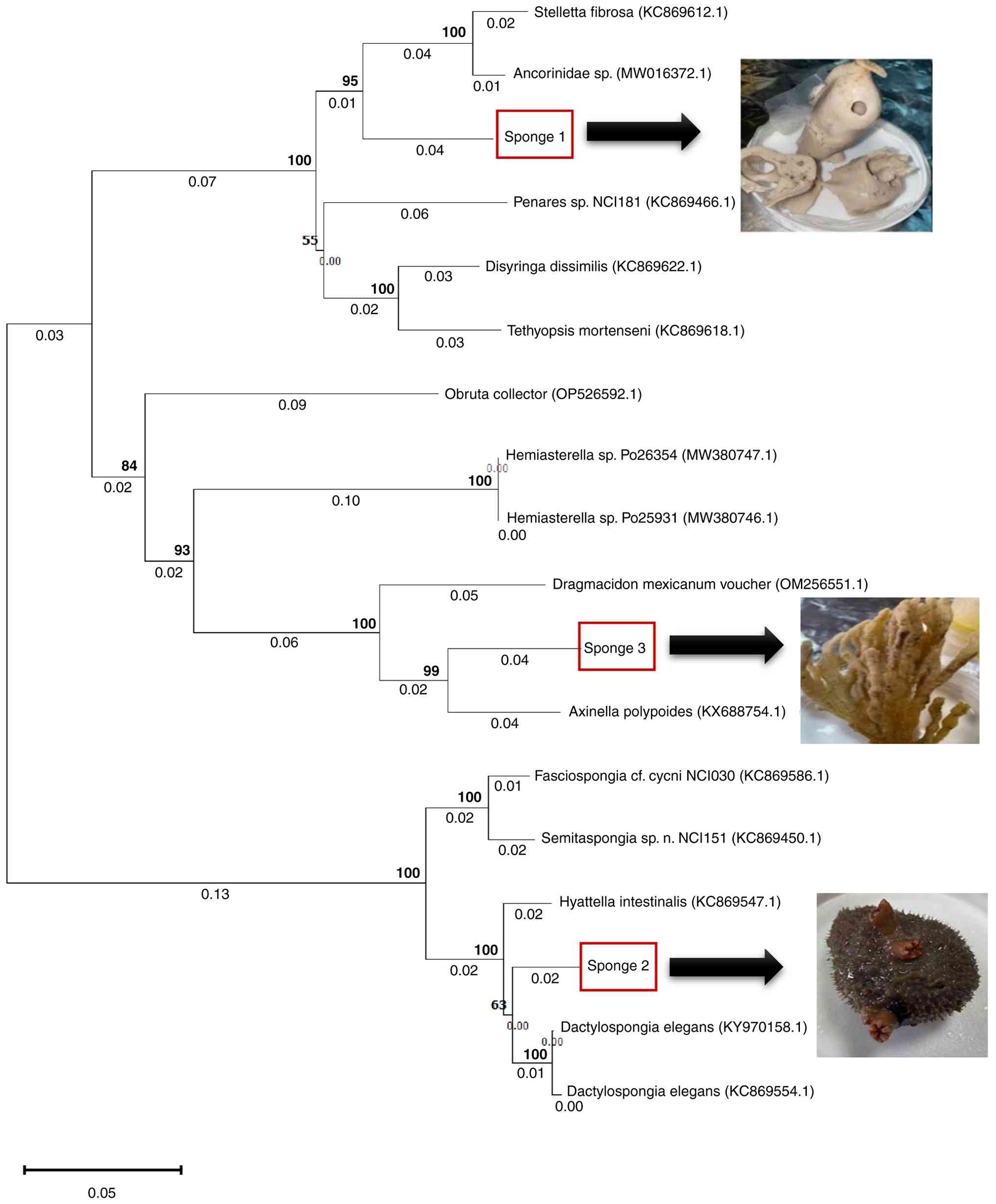
To refine taxonomic placement, a phylogenetic tree
was constructed using the neighbour-joining method with 1,000
bootstrap replicates (Fig. 1). The
tree topology supported the BLAST results and clustered Sponge 1
within the genus Stelletta (family Ancorinidae, order
Tetractinellida), with Sponge 2 grouped closely with D. elegans
(family Thorectidae, order Dictyoceratida), and Sponge 3 within the
genus Axinella (family Axinellidae, order Axinellida). Given the
limitations of single locus barcoding for Porifera, we
conservatively report the three specimens at the genus level:
Stelletta sp., Dactylospongia cf. elegans and
Axinella sp. This conservative identification is further
supported by morphological traits (Table SI). The final sequences generated
in the present study were deposited in GenBank under accession
numbers PX278187 (Stelletta sp.), PX278188 (D. cf.
elegans) and PX278189 (Axinella sp.).
Antibacterial activity
The antibacterial activity of the three ethanolic
sponge extracts was evaluated against six clinically relevant
bacterial strains, including Gram-positive strains (S. aureus,
B. pumilus, S. epidermidis and MRSA and Gram-negative
strains (E. coli and K. aerogenes). As shown in
Fig. 2, only the extract of D.
cf. elegans exhibited marked antibacterial activity, which was
limited to Gram-positive bacteria. The inhibition zones ranged from
7 to 21 mm in a concentration-dependent manner against S.
aureus, MRSA and S. epidermidis (Table III). By contrast, no inhibitory
effect was observed against B. pumilus or the Gram-negative
strains (E. coli and K. aerogenes).
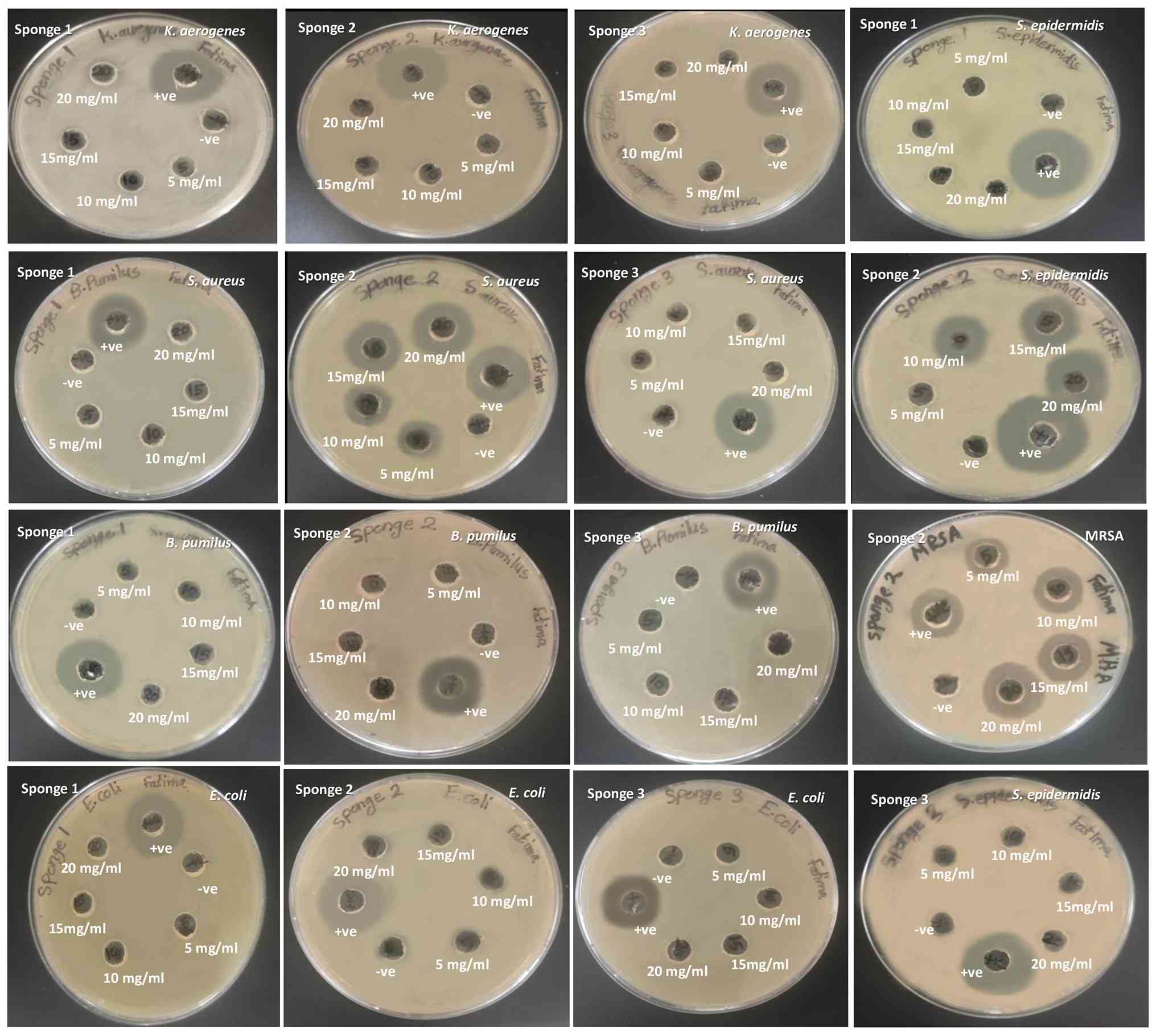 | Figure 2Results of the agar well diffusion
assay for antibacterial activity against: Escherichia coli,
Staphylococcus epidermidis, Klebsiella aerogenes,
Staphylococcus aureus, Bacillus pumilus and
methicillin-resistant Staphylococcus aureus. Where
Stelletta sp. (Sponge 1), D. cf. elegans (Sponge 2),
Axinella sp. (Sponge 3) with concentrations (5, 10, 15, 20
mg/ml). Gentamycin (10 µg): Positive control for all bacteria,
Vancomycin (30 µg): MRSA-specific positive control, and 80% DMSO:
Negative control. MRSA, methicillin-resistant Staphylococcus
aureus. |
 | Table IIIInhibition zone diameters (mm) for
different bacterial strains treated with sponge 2 ethanolic
extracts in different concentrations. |
Table III
Inhibition zone diameters (mm) for
different bacterial strains treated with sponge 2 ethanolic
extracts in different concentrations.
| Mean of the
inhibition zone diameter (mm) for sponge 2 |
|---|
| Bacterial
strains | 5 mg/ml | 10 mg/ml | 15 mg/ml | 20 mg/ml | Positive
control | Negative
control |
|---|
| E. coli | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 19.67±1.20 | 0.00±0.00 |
| S.
aureus | 11.00±0.58 | 15.33±0.88 | 17.67±1.85 | 21.00±1.53 | 20.00±3.00 | 0.00±0.00 |
| K.
aerogenes | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 19.67±0.88 | 0.00±0.00 |
| B.
pumilus | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 0.00±0.00 | 18.33±0.88 | 0.00±0.00 |
| S.
epidermidis | 12.66±2.33 | 7.00±3.51 | 14.00±0.58 | 17.33±0.33 | 27.00±0.58 | 0.00±0.00 |
| MRSA | 12.33±0.33 | 15.33 ± 0.33 | 17.00±0.58 | 18.33±0.33 | 15.67±0.33 | 0.00±0.00 |
The extracts of Stelletta sp. and Axinella
sp. also failed to show any detectable antibacterial effect.
This absence of activity may be attributed to differences in the
secondary metabolite composition of these sponges, ecological
variation, or the presence of compounds with specificity toward
microbial taxa that were not included in the present panel. The MIC
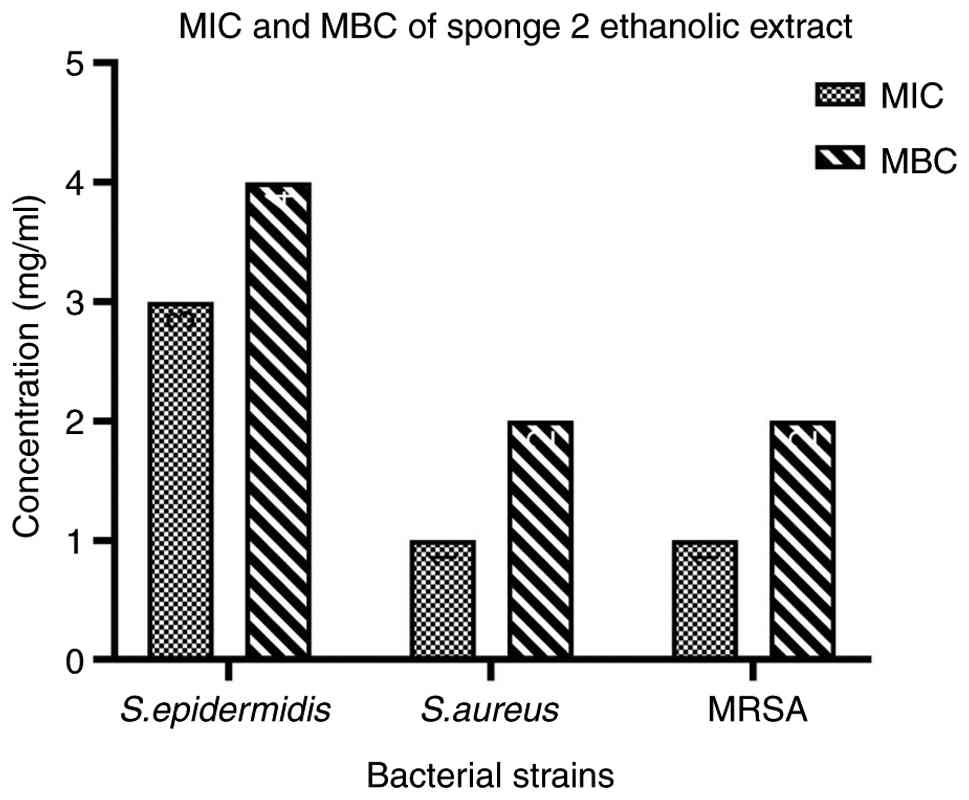
and MBC assays further supported the diffusion results. The
ethanolic extract of D. cf. elegans exhibited MIC and MBC
values of 1 and 2 mg/ml against Gram-positive bacteria,
respectively (Fig. 3), which
confirmed its potent bactericidal potential. All inhibition-zone
data are presented as the mean ± SEM with 95% confidence intervals
from three independent replicates, while MIC and MBC values are
reported descriptively according to standard practice in
antimicrobial susceptibility testing.
These findings align with previous investigations on
sponge-derived metabolites. For instance, sesquiterpene quinones
from Acanthella cavernosa and diterpenoids from Haliclona
sp. displayed selective activity against Gram-positive
bacteria, but often at higher MIC ranges (2-8 mg/ml) (72,73).
Similarly, extracts from D. elegans have been reported to
yield structurally diverse compounds with broad-spectrum activity,
including against resistant strains of S. aureus (74). Mechanistic studies have shown that
drimane meroterpenoids, such as pelorol, inhibit bacterial
dihydrofolate reductase, a key enzyme in folate metabolism and DNA
synthesis in pathogens (75,76).
This proposed mechanism may explain the pronounced activity of
Dactylospongia-derived metabolites observed in the present
study.
The strong bioactivity of Dactylospongia cf.
elegans is also consistent with its metabolite composition,
which was confirmed by LC-MS/MS analysis (Table IV). Compounds such as
bolinaquinone, dactyloquinone, gallic acid and caffeic acid were
detected in the extract, which are metabolites that have been
extensively reported for their antibacterial properties (77-79).
Their co-occurrence in the ethanolic extract suggests a possible
synergistic contribution to the selective activity against
Gram-positive pathogens.
 | Table IVBioactive compounds were detected in
the ethanolic extract of sponge 2 by LC-MS-MS. |
Table IV
Bioactive compounds were detected in
the ethanolic extract of sponge 2 by LC-MS-MS.
| Compounds | Molecular
formula | Molecular weight
(g/mol) | % | RT |
|---|
| Gallic acid |
C7H6O5 | 170.12 | 7 | 1.3 |
| Dactyloquinone |
C22H28O4 | 356.5 | 6.2 | 3 |
| Chromazonarol |
C21H30O2 | 314.5 | 5.1 | 2.35 |
|
Manoalide |
C25H36O5 | 416.5 | 5.1 | 5.4 |
| Caffeic acid |
C9H8O4 | 180.16 | 5 | 1.4 |
| Bolinaquinone |
C22H30O4 | 358.5 | 5 | 3.6 |
|
Mamanuthaquinone |
C22H30O4 | 358.5 | 4.2 | 3.7 |
| δ-humulene |
C15H24 | 204.35 | 4.1 | 1.6 |
| Ergosterol |
C28H44O | 396.6 | 4.1 | 5.1 |
| Linoleic acid |
C18H32O2 | 280.4 | 3.6 | 1.9 |
| Ilimaquinone |
C22H30O4 | 358.5 | 3.3 | 3.22 |
| Pelorol |
C23H32O4 | 372.5 | 3.3 | 4.5 |
|
Hyatellaquinone |
C12H14O4 | 222.24 | 3.2 | 1.7 |
| Ellagic Acid |
C14H6O8 | 302.19 | 3.2 | 2.1 |
|
Cyclospongiaquinone |
C22H30O4 | 358.5 | 3.1 | 4 |
|
indole-3-carbaldehyde |
C9H7NO | 145.16 | 3 | 1.2 |
| Petasitolone |
C15H24O2 | 236.35 | 2.8 | 1.75 |
| Ocimene |
C10H16 | 136.23 | 2.2 | 1.1 |
| Scopoletin |
C10H8O4 | 192.17 | 2.2 | 1.45 |
|
Isospongiaquinone |
C22H30O4 | 358.5 | 2.1 | 3.5 |
| Lectin |
C26H28O6 | 436.5 | 2.1 | 5.7 |
| Dysideamine |
C21H29NO3 | 343.5 | 2 | 2.8 |
| Stelletin |
C30H38O4 | 462.6 | 1.7 | 6.3 |
| Nakijiquinone
D |
C25H35NO6 | 445.5 | 1.6 | 6 |
| Ferulic acid |
C10H10O4 | 194.18 | 1.5 | 1.5 |
| Clathric acid |
C20H30O2 | 302.5 | 1.5 | 2.2 |
| Motualevic
acid |
C16H23Br2NO2 | 421.2 | 1.3 | 5.5 |
|
Smenospongimine |
C22H31NO3 | 357.5 | 1.2 | 3.1 |
| Limonene |
C10H16 | 136.23 | 1.1 | 1 |
| p-coumaric
acid |
C9H8O3 | 164.16 | 1.1 | 1.25 |
| Catechin |
C15H14O6 | 290.27 | 1 | 2 |
| Smenospongine |
C21H29NO3 | 343.5 | 1 | 2.6 |
| Dactyltronic
acid |
C21H30O5 | 362.5 | 1 | 4.15 |
| Popolohuanone |
C42H57NO3 | 623.9 | 1 | 6.7 |
| Chlorogenic
acid |
C16H18O9 | 354.31 | 0.9 | 2.9 |
|
Luffariellolide |
C25H38O3 | 386.6 | 0.8 | 4.8 |
| Rutin |
C27H30O16 | 610.5 | 0.5 | 6.5 |
| Dictyoceratin
A |
C23H32O4 | 372.5 | 0.3 | 4.35 |
| Squalene |
C30H50 | 410.7 | 0.3 | 5.3 |
| Manzamine A |
C36H44N4O | 548.8 | 0.3 | 6.4 |
LC-MS/MS-based chemical profiling of
D. cf. elegans extract
LC-MS/MS analysis of the ethanolic extract of D.
cf. elegans (Sponge 2) revealed a chemically diverse profile
comprising ~40 metabolites (Table
IV, Fig. S6). These included
terpenoids (for example, limonene, squalene, δ-humulene), alkaloids
(for example, manzamine A, smenospongine), quinones (for example,
hyatellaquinone, bolinaquinone, ilimaquinone and dactyloquinone),
and phenolic acids (for example, gallic, caffeic, ferulic,
p-coumaric and chlorogenic). Compounds such as gallic acid (7%),
dactyloquinone (6.2%), bolinaquinone (5.0%), chromazonarol (5.1%),
manoalide (5.1%) and δ-humulene (4.1%) were detected in relatively
high abundance. Interestingly, phenolic compounds such as gallic
and caffeic acid, which are commonly associated with terrestrial
plants, were also present. The occurrence of gallic acid and
chromazonarol in marine sponges has been attributed to
sponge-associated symbionts or the uptake of dissolved organic
matter (78,80,81).
Several of the metabolites detected in this extract
are well recognized for their antibacterial properties. Gallic acid
and caffeic acid disrupt bacterial membranes, interfere with
metabolism, inhibit biofilm formation, and have potent effects
against S. aureus and S. epidermidis (82-85).
Manzamine A impairs protein synthesis in Gram-positive bacteria,
while quinones such as ilimaquinone and hyatellaquinone generate
reactive oxygen species that compromise membrane integrity
(86,87). Therefore, the selectivity of the
extract for Gram-positive bacteria in our assays may be explained
by these mechanisms, which also explain the absence of activity
against Gram-negative bacteria due to their impermeable outer
membrane (73).
In addition to phenolics and quinones, other
metabolites identified in the extract have reported antibacterial
activities. Linoleic acid disrupts bacterial membranes with MIC
values as low as 0.01 mg/ml (87).
Indole-3-carbaldehyde inhibits bacterial growth and biofilm
formation by interfering with signalling pathways (88), while lectins prevent adhesion and
biofilm development (89).
Ergosterol also exerts antibacterial effects by compromising
membrane integrity metabolism (90). Notably, no studies to date have
examined the antibacterial potential of several compounds detected
in our extract, including bolinaquinone, dactyloquinone, manoalide
and chromazonarol, which underscores the novelty of these
findings.
The ecological setting of the sponge may also have
shaped its secondary metabolite profile. Dactylospongia
specimens were collected from a coral-rich reef with high
biodiversity and low anthropogenic impact, where they are exposed
to intense microbial competition, UV radiation, and other stressors
that are known to upregulate biosynthetic gene clusters (80,91).
Such conditions may explain the abundance of terpenoids (for
example, δ-humulene) and quinones, which are compounds that are
typically linked to chemical defence strategies. Some identified
metabolites, such as pelorol and smenospongimine, are rarely
reported in Dactylospongia species (92,93),
and their detection here underscores the chemical novelty of this
extract. Together with known classes (sterols, pregnanes,
sesterterpenes), these compounds extend the pharmacological
repertoire of Dactylospongia, which has been associated with
antibacterial, anticancer, cytotoxic and anti-inflammatory
properties (94).
It is noteworthy that D. cf. elegans (Sponge
2) exhibited unusually high bioactivity compared with the other
sponges analysed in this study and even compared with previous
studies on conspecifics from non-reef habitats. The specimens were
physically associated with coral structures in a reef-rich site of
the Gulf of Aqaba. Coral-associated sponges have been shown to host
distinct microbiomes and metabolomes compared with free-living
conspecifics (95,96). The ecological interactions,
including microbial symbiosis, nutrient exchange, and exposure to
coral-derived dissolved organic matter, can significantly alter the
sponge's chemical output (97-99).
The unusually high abundance of compounds such as
dactyloquinone, bolinaquinone, gallic acid and chromazonarol in
this extract were underrepresented in previous accounts of
Dactylospongia sp. (98,99)
and supports the hypothesis that reef proximity and environmental
complexity are crucial drivers of metabolomic diversity. Similar
findings have been reported for sponges from the Pacific and
Caribbean, where coral-associated sponges biosynthesize higher
levels of cytotoxic sesterterpenes and brominated alkaloids than
reef-margin species (100). These
ecological insights provide a plausible explanation for the
enhanced antibacterial profile observed in the present study and
reinforce the concept of sponges as holobionts with pharmacological
potential that is shaped by both their taxonomy and their biotic
environment.
Although numerous of the compounds detected are
individually known for antibacterial effects, the overall
bioactivity of the extract may also result from synergistic
interactions. The combination of phenolic acids and quinones, for
example, could potentiate the antibacterial potency against
Gram-positive pathogens beyond the activity of each metabolite
alone. Therefore, future studies should focus on purification,
isolation and testing of individual compounds, as well as
combinatorial assays to evaluate their synergy. In summary,
LC-MS/MS profiling revealed a rich array of secondary metabolites
in the ethanolic extract of Dactylospongia sp., including
phenolic acids, quinones, terpenoids and alkaloids. These findings
provide a strong chemical rationale for the selective antibacterial
activity observed in this sponge, particularly against
Gram-positive bacteria, and highlight its promise as a source of
novel bioactive agents.
In conclusion, the present study highlighted the
ethanolic extract of Dactylospongia sp. as a promising
source of antibacterial compounds with selective activity against
Gram-positive bacteria, including S. aureus, S.
epidermidis and MRSA. LC-MS/MS profiling revealed a chemically
diverse metabolite composition, including phenolic acids, quinones,
terpenoids and alkaloids, several of which are well known for their
antimicrobial activity. Importantly, some compounds identified in
this extract, such as bolinaquinone, dactyloquinone, manoalide and
chromazonarol, have not been previously reported for antibacterial
effects, which underscores the novelty of these findings.
Future studies should focus on the purification and
isolation of individual compounds, mechanistic investigations, and
evaluation of synergistic interactions among metabolites such as
gallic and caffeic acids with sponge-derived quinones. Toxicity
profiling and validation in vivo are also necessary to fully
assess their therapeutic potential. Collectively, these findings
emphasize the value of Dactylospongia sp. in the search for
novel antimicrobial agents and support the broader concept that
marine sponges, particularly those associated with coral reef
environments, represent important reservoirs for bioactive
metabolites that could help to address the urgent global challenge
of antibiotic resistance.
Supplementary Material
Sampling locations of marine sponges
from the Gulf of Aqaba, Jordan, where (1-8) indicates the stations
of sample collection sites.
Morphological view of Stelletta
sp. (corresponding to specimen 1 in Table SI). The image shows the external
morphology and surface texture used to support taxonomic
identification.
Morphological view of
Dactylospongia cf. elegans (corresponding to specimen 2 in
Table SI). The image illustrates
the sponge's characteristic massive form and color pattern as
recorded during sampling.
Morphological view of Axinella
sp. (corresponding to specimen 3 in Table SI). The figure displays the
branching morphology and skeletal structure features relevant to
its identification.
Agarose gel electrophoresis of 28S
rRNA PCR products in three sponge samples: This demonstrates the
PCR amplification patterns of the 28S rRNA gene in three sponge
samples. The wells were loaded with 3 μl of PCR products.
Lane 1 is the negative control, Lane 2 is Sponge 1, Lane 3 is
Sponge 2, Lane 4 is Sponge 3, and Lane 5 is the 1 Kb molecular
marker to ascertain size.
Chromatogram of sponge 2 ethanolic
extract: 40 peaks were found, each representing a different
molecule that eluted at different retention durations. The
chromatogram's intricacy suggests a varied chemical makeup that
most likely includes a range of secondary metabolites with possible
bioactive characteristics. Subsequent chemical identification and
biological activity evaluation are based on this profile.
Morphological traits of the examined
sponge specimens (corresponding images are shown separately in
Fig. S2, Fig. S3 and Fig. S4).
Acknowledgements
The present study benefited from the OceanXplorer
Jordan Expedition 2022, which provided access to its research
vessel for sample collection. The authors also relied on the
support of the Aqaba Special Economic Zone Authority (ASEZA), which
helped coordinate fieldwork in Aqaba and supplied the environmental
data essential to this research.
Funding
Funding: The present study was supported by the Deanship of
Scientific Research at the University of Jordan supported this
research for graduate students (grant no. 401).
Availability of data and materials
The data generated in the present study may be found
in the NCBI GenBank database under accession numbers PX278187.1,
PX278188.1 and PX278189.1 or at the following URL: https://www.ncbi.nlm.nih.gov/nuccore/PX278187.1,
https://www.ncbi.nlm.nih.gov/nuccore/PX278188.1 and
https://www.ncbi.nlm.nih.gov/nuccore/PX278189.1. The
data generated in the present study may be requested from the
corresponding author.
Authors' contributions
FFA, RAA, MMDA, MZ and OHA conceptualized the study
and developed the methodology. FFA conducted the formal analysis.
AAD conducted the LC-MS/MS analysis. MMDA and MZ were responsible
for project administration. MMDA, OHA, AAD and MZ supervised the
study. FFA and RAA wrote the original draft. MMDA, OHA and MZ
wrote, reviewed and edited the manuscript. All authors read and
approved the final version of the manuscript. FFA and RAA confirm
the authenticity of all the raw data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Naser AY, Aboutaleb R, Khaleel A,
Alsairafi ZK, Alwafi H, Qadus S, Itani R, El-Dahiyat F, Awaisu A,
Awwad O, et al: Knowledge, attitude, and practices of pharmacy
students in 7 Middle Eastern countries concerning antibiotic
resistance: A cross-sectional study. Medicine.
103(e39378)2024.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Anteneh YS, Yang Q, Brown MH and Franco
CM: Antimicrobial activities of marine Sponge-associated bacteria.
Microorganisms. 9(171)2021.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Lakshmanan A, Govindasamy C, Sivakumar AS,
Hussein-Al-Ali SH, Ramesh M and Lakshmanan H: Anticancer and
antimicrobial potential of zinc/sodium alginate/polyethylene
glycol/d-pinitol nanocomposites against osteosarcoma MG-63 cells.
Green Proces Synthesis. 12(20230124)2023.
|
|
4
|
U.S. Centers for Disease Control:
Antibiotic resistance threats in the United States, 2019. Atlanta,
GA, US Department of Health and Human Services, CDC, 2019.
|
|
5
|
Okada BK and Seyedsayamdost MR: Antibiotic
dialogues: Induction of silent biosynthetic gene clusters by
exogenous small molecules. FEMS Microbiol Rev. 41:19–33.
2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Farha MA and Brown ED: Strategies for
target identification of antimicrobial natural products. Nat Prod
Rep. 33:668–680. 2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Al-Zereini WA, Al-Trawneh IN, Al-Qudah MA,
TumAllah HM, Abudayeh ZH and Hijazin T: Antibacterial, antioxidant,
and cytotoxic activities of Syzygium aromaticum (L.) Merr.
& Perry essential oil with identification of its chemical
constituents. Z Naturforsch C J Biosci. 78:105–112. 2023.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Bibi F, Faheem M, Azhar EI, Yasir M, A
Alvi SA, A Kamal MA, Ullah I and Naseer MI: Bacteria from marine
sponges: A source of new drugs. Curr Drug Metab. 18:11–15.
2017.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Antimicrobial Resistance Collaborators.
Global burden of bacterial antimicrobial resistance in 2019: A
systematic analysis. Lancet. 399:629–655. 2022.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Fair RJ and Tor Y: Antibiotics and
bacterial resistance in the 21st century. Perspect Medicin Chem.
6:25–64. 2014.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Altuğ G, Çiftçi Türetken PS, Kalkan S and
Topaloğlu B: The distribution and antibacterial activity of marine
sponge-associated bacteria in the Aegean Sea and The Sea of
Marmara, Turkey. Curr Microbiol. 78:2275–2290. 2021.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Erpenbeck D, Gholami A, Hesni MA, Ranjbar
MS, Galitz A, Eickhoff B, Namuth L, Schumacher T, Esmaeili HR,
Wörheide G and Teimori A: Molecular biodiversity of Iranian shallow
water sponges. Systemat Biodiversity. 18:192–202. 2020.
|
|
13
|
Devasahayam G, Scheld WM and Hoffman PS:
Newer antibacterial drugs for a new century. Expert Opin Investig
Drugs. 19:215–234. 2010.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Bashari MH, Arsydinilhuda FZ, Ilhamsyah
RS, Nugrahani AD, Nurdin RA, Kartika A, Huda F, Abdurahman M, Putri
T, Qomarilla N, et al: The ethanol extract of marine sponge Aaptos
suberitoides suppress cell viability, cell proliferation and cell
migration in HER2-positive breast cancer cell line. Asian Pac J
Cancer Prev. 22:25–32. 2021.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Carnovali M, Ciavatta ML, Mollo E, Roussis
V, Banfi G, Carbone M and Mariotti M: Aerophobin-1 from the marine
sponge Aplysina aerophoba modulates
osteogenesis in zebrafish larvae. Marine Drugs.
20(135)2022.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Dyshlovoy SA, Fedorov SN, Svetashev VI,
Makarieva TN, Kalinovsky AI, Moiseenko OP, Krasokhin VB, Shubina
LK, Guzii AG, von Amsberg G and Stonik VA: 1-O-Alkylglycerol ethers
from the marine sponge guitarra abbotti and their cytotoxic
activity. Mar Drugs. 20(409)2022.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Gong KK, Tang XL, Zhang G, Cheng CL, Zhang
XW, Li PL and Li GQ: Polyhydroxylated steroids from the South China
Sea soft coral Sarcophyton sp. and their cytotoxic and
antiviral activities. Mar Drugs. 11:4788–4798. 2013.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Manivasagan P, Venkatesan J, Sivakumar K
and Kim SK: Pharmaceutically active secondary metabolites of marine
actinobacteria. Microbiol Res. 169:262–278. 2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Tajima H, Wakimoto T, Takada K, Ise Y and
Abe I: Revised structure of cyclolithistide A, a cyclic
depsipeptide from the marine sponge Discodermia japonica. J
Nat Prod. 77:154–158. 2014.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Bashari MH, Huda F, Tartila TS, Shabrina
S, Putri T, Qomarilla N, Atmaja H, Subhan B, Sudji IR and Meiyanto
E: Bioactive compounds in the ethanol extract of marine sponge
Stylissa carteri demonstrates potential anti-cancer activity in
breast cancer cells. Asian Pac J Cancer Prev. 20:1199–1206.
2019.PubMed/NCBI View Article : Google Scholar
|
|
21
|
González-Castillo A, Carballo JL and
Bautista-Guerrero E: Genomics and phylogeny of the proposed phylum
‘Candidatus Poribacteria’ associated with the excavating
sponge Thoosa mismalolli. Antonie Van Leeuwenhoek.
114:2163–2174. 2021.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Harper LR, Neave EF, Sellers GS,
Cunnington AV, Arias MB, Craggs J, MacDonald B, Riesgo A and
Mariani S: Optimized DNA isolation from marine sponges for natural
sampler DNA metabarcoding. Environmental DNA. 5:438–461. 2023.
|
|
23
|
Gomes NG, Dasari R, Chandra S, Kiss R and
Kornienko A: Marine invertebrate metabolites with anticancer
activities: Solutions to the ‘Supply Problem’. Mar Drugs.
14(98)2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Wooster MK, Voigt O, Erpenbeck D, Wörheide
G and Berumen ML: Sponges of the Red Sea. Coral Reefs Red Sea.
91–122. 2019.doi:10.1007/978-3-030-05802-9_6.
|
|
25
|
Al-Taani AA, Rashdan M, Nazzal Y, Howari
F, Iqbal J, Al-Rawabdeh A, Al Bsoul A and Khashashneh S: Evaluation
of the Gulf of Aqaba coastal water, Jordan. Water.
12(2125)2020.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Moore S and Squires D: Governing the
depths: Conceptualizing the politics of deep sea resources. Global
Environ Politics. 16:101–109. 2016.
|
|
27
|
Sengupta S, Gildor H and Ashkenazy Y:
Depth-dependent warming of the Gulf of Eilat (Aqaba). Climatic
Change. 177(107)2024.
|
|
28
|
Helber SB, Hoeijmakers DJ, Muhando CA,
Rohde S and Schupp PJ: Sponge chemical defenses are a possible
mechanism for increasing sponge abundance on reefs in Zanzibar.
PLoS One. 13(e0197617)2018.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Keyvani-Ghamsari S, Rahimi M and Khorsandi
K: An update on the potential mechanism of gallic acid as an
antibacterial and anticancer agent. Food Sci Nutr. 11:5856–5872.
2023.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Liu T, Wu S, Zhang R, Wang D, Chen J and
Zhao J: Diversity and antimicrobial potential of Actinobacteria
isolated from diverse marine sponges along the Beibu Gulf of the
South China Sea. FEMS Microbiol Ecol. 95(fiz089)2019.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Calcabrini C, Catanzaro E, Bishayee A,
Turrini E and Fimognari C: Marine sponge natural products with
anticancer potential: An updated review. Mar Drugs.
15(310)2017.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Ramírez GA, Bar-Shalom R, Furlan A, Romeo
R, Gavagnin M, Calabrese G, Garber AI and Steindler L: Bacterial
aerobic methane cycling by the marine sponge-associated microbiome.
Microbiome. 11(49)2023.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Rizzo C, Zammuto V, Lo Giudice A, Rizzo
MG, Spanò A, Laganà P, Martinez M, Guglielmino S and Gugliandolo C:
Antibiofilm activity of Antarctic sponge-associated bacteria
against Pseudomonas aeruginosa and Staphylococcus
aureus. J Marine Sci Engineering. 9(243)2021.
|
|
34
|
Unson MD, Holland N and Faulkner DJ: A
brominated secondary metabolite synthesized by the cyanobacterial
symbiont of a marine sponge and accumulation of the crystalline
metabolite in the sponge tissue. Marine Biol. 119:1–11. 1994.
|
|
35
|
Perdicaris S, Vlachogianni T and
Valavanidis A: Bioactive natural substances from marine sponges:
New developments and prospects for future pharmaceuticals. Nat Prod
Chem Res. 1:2329–6836. 2013.
|
|
36
|
Matobole RM, Van Zyl LJ, Parker-Nance S,
Davies-Coleman MT and Trindade M: Antibacterial activities of
bacteria isolated from the marine sponges Isodictya
compressa and Higginsia bidentifera collected from Algoa
Bay, South Africa. Mar Drugs. 15(47)2017.
|
|
37
|
Sipkema D, Franssen MC, Osinga R, Tramper
J and Wijffels RH: Marine sponges as pharmacy. Mar Biotechnol (NY).
7:142–162. 2005.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Rifai S, Fassouane A, El-Abbouyi A,
Wardani A, Kijjoa A and Van Soest R: Screening of antimicrobial
activity of marine sponge extracts. J Mycologie Méd. 15:33–38.
2005.
|
|
39
|
Rehnstam AS, Bäckman S, Smith DC, Azam F
and Hagström Å: Blooms of sequence-specific culturable bacteria in
the sea. FEMS Microbiol Ecol. 11:161–166. 1993.
|
|
40
|
Müller WE, Grebenjuk VA, Le Pennec G,
Schröder HC, Brümmer F, Hentschel U, Müller IM and Breter H:
Sustainable production of bioactive compounds by sponges-cell
culture and gene cluster approach: A review. Mar Biotechnol (NY).
6:105–117. 2004.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Azevedo LG, Muccillo-Baisch AL, Filgueira
Dde M, Boyle RT, Ramos DF, Soares AD, Lerner C, Silva PA and
Trindade GS: Comparative cytotoxic and anti-tuberculosis activity
of Aplysina caissara marine sponge crude extracts. Comp Biochem
Physiol C Toxicol Pharmaco. 147:36–42. 2008.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Kristiana R, Sibero MT, Farisa MY,
Ayuningrum D, Dirgantara D, Hanafi M, Radjasa OK, Sabdono A and
Trianto A: Antibacterial potential of nudibranch-associated
bacteria from Saparua and Nusa Laut Islands, Indonesia.
Biodiversitas J Biol Diversity. 20:1811–1819. 2019.
|
|
43
|
Sun H, Xie Z, Yang X, Yang B, Liao B, Yin
J and Xiao B: New insights into microbial and metabolite signatures
of coral bleaching. Sci Total Environ. 892(164258)2023.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Li H, Long J, Wang X, She J, Liu Y, Li Y
and Yang B: Bioactive secondary metabolites isolated from the soft
coral derived Penicillium sp. SCSIO 41038. Nat Prod Res.
38:2996–3003. 2023.PubMed/NCBI View Article : Google Scholar
|
|
45
|
El-Said GF: Bioaccumulation of key metals
and other contaminants by seaweeds from the Egyptian Mediterranean
Sea coast in relation to human health risk. Hum Ecol Risk
Assessment An Int J. 19:1285–1305. 2013.
|
|
46
|
Bhatnagar I and Kim SK: Immense essence of
excellence: Marine microbial bioactive compounds. Mar Drugs.
8:2673–2701. 2010.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Campos PE, Pichon E, Moriou C, Clerc P,
Trépos R, Frederich M, De Voogd N, Hellio C, Gauvin-Bialecki A and
Al-Mourabit A: New antimalarial and antimicrobial tryptamine
derivatives from the marine sponge Fascaplysinopsis reticulata. Mar
Drugs. 17(167)2019.PubMed/NCBI View Article : Google Scholar
|
|
48
|
El-Damhougy K, El-Naggar HA, Ibrahim H,
Bashar M and Abou-Senna FM: Biological activities of some marine
sponge extracts from Aqaba Gulf, Red Sea, Egypt. Int J Fisheries
Aquatic Stud. 5:652–659. 2017.
|
|
49
|
Blunt JW, Copp BR, Keyzers RA, Munro MH
and Prinsep MR: Marine natural products. Nat Product Rep.
29:144–222. 2012.
|
|
50
|
Wink M: Molecular modes of action of
cytotoxic alkaloids: From DNA intercalation, spindle poisoning,
topoisomerase inhibition to apoptosis and multiple drug resistance.
Alkaloids Chem Biol. 64:1–47. 2007.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Abuassaf RA, Al-Jamal FF, Abusara OH,
Zihlif M, Deeb AA and Al-Rshaidat MM: Evaluating the antibacterial
properties of deep-sea sponges Dactylospongia elegants,
Stelletta fibrosa, and Haliclona manglaris from the
Jordanian Gulf of Aqaba. PeerJ. 13(e19735)2025.PubMed/NCBI View Article : Google Scholar
|
|
52
|
Yang Q, Franco CM, Sorokin SJ and Zhang W:
Development of a Multilocus-based approach for sponge (phylum
Porifera) identification: Refinement and limitations. Sci Rep.
7(41422)2017.PubMed/NCBI View Article : Google Scholar
|
|
53
|
Timmers MA, Vicente J, Webb M, Jury CP and
Toonen RJ: Sponging up diversity: Evaluating metabarcoding
performance for a taxonomically challenging phylum within a complex
cryptobenthic community. Environmental DNA. 4:239–253. 2022.
|
|
54
|
Timmers PH, Widjaja-Greefkes HA, Plugge CM
and Stams AJ: Evaluation and optimization of PCR primers for
selective and quantitative detection of marine ANME subclusters
involved in sulfate-dependent anaerobic methane oxidation. Appl
Microbiol Biotechnol. 101:5847–5859. 2017.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Idan T, Shefer S, Feldstein T, Yahel R,
Huchon D and Ilan M: Shedding light on an East-Mediterranean
mesophotic sponge ground community and the regional sponge fauna.
Mediterranean Marine Sci. 19:84–106. 2018.
|
|
56
|
Erpenbeck D, Voigt O, Al-Aidaroos AM,
Berumen ML, Büttner G, Catania D, Guirguis AN, Paulay G, Schätzle S
and Wörheide G: Molecular biodiversity of Red Sea demosponges. Mar
Pollut Bull. 105:507–514. 2016.PubMed/NCBI View Article : Google Scholar
|
|
57
|
Arabeyyat Z, Sweiss M, Alajlouni A,
Al-Ajlouni Na, Mahmoud M, Shartooh S, Alsoqi F and Kteifan M:
Identification and phylogenetic analysis of marine sponges in the
Jordanian Gulf of Aqaba using DNA barcoding. Heliyon.
11(e42771)2025.PubMed/NCBI View Article : Google Scholar
|
|
58
|
Bayona LM, Videnova M and Choi YH:
Increasing metabolic diversity in marine sponges extracts by
controlling extraction parameters. Mar Drugs.
16(393)2018.PubMed/NCBI View Article : Google Scholar
|
|
59
|
Ballantine DL, Gerwick WH, Velez SM,
Alexander E and Guevara P: Antibiotic activity of lipid-soluble
extracts from Caribbean marine algae. Hydrobiologia. 151:463–469.
1987.
|
|
60
|
Magaldi S, Mata-Essayag S, De Capriles CH,
Pérez C, Colella M, Olaizola C and Ontiveros Y: Well diffusion for
antifungal susceptibility testing. Int J Infect Dis. 8:39–45.
2004.PubMed/NCBI View Article : Google Scholar
|
|
61
|
Salameh BA, Al-Hushki EH, Talib WH, Ghanem
R, Delmani FA and Mahmod AI: A new class of
pyrrolo[2,3-b]quinoxalines: Synthesis, anticancer and antimicrobial
activities. Zeitschrift für Naturforschung B. 79:303–309. 2024.
|
|
62
|
Pesewu GA, Cutler RR and Humber DP:
Antibacterial activity of plants used in traditional medicines of
Ghana with particular reference to MRSA. J Ethnopharmacol.
116:102–111. 2008.PubMed/NCBI View Article : Google Scholar
|
|
63
|
Geetha R and Roy A: In vitro evaluation of
anti bacterial activity three herbal extracts on methicillin
resistant Staphylococcus aureus [MRSA]. J Pharmaceutical Sci
Res. 5(207)2013.
|
|
64
|
Saad R, Asyikin N, Khan J, Aldahlli S,
Sultan S, Abdulhamid J, Yusuf E and Yusuf F: Determination of
minimum inhibitory concentration utilizing microtitre plate
bioassay for three Malaysian Herbal Medicines. Int J Appl
Pharmaceutical Sci Biomed Sci. 3:280–290. 2014.
|
|
65
|
Prastiyanto ME, Azizah IH, Haqi HD,
Yulianto BD, Agmala AB, Radipasari ZD, Dwi Astuti NA and Putri AR:
In-vitro antibacterial activity of the seed extract of three-member
Artocarpus towards Methicillin-Resistant Staphylococcus
aureus (MRSA). J Teknol Laboratorium. 9:128–135. 2020.
|
|
66
|
Sarker SD, Nahar L and Kumarasamy Y:
Microtitre plate-based antibacterial assay incorporating resazurin
as an indicator of cell growth, and its application in the in vitro
antibacterial screening of phytochemicals. Methods. 42:321–324.
2007.PubMed/NCBI View Article : Google Scholar
|
|
67
|
Ibrahim AI, Abul-Futouh H, Bourghli LM,
Abu-Sini M, Sunoqrot S, Ikhmais B, Jha V, Sarayrah Q, Abulebdah DH
and Ismail WH: Design and synthesis of thionated levofloxacin:
Insights into a new generation of quinolones with potential
therapeutic and analytical applications. Curr Issues Mol Biol.
44:4626–4638. 2022.PubMed/NCBI View Article : Google Scholar
|
|
68
|
Balouiri M, Sadiki M and Ibnsouda SK:
Methods for in vitro evaluating antimicrobial activity: A review. J
Pharm Anal. 6:71–79. 2016.PubMed/NCBI View Article : Google Scholar
|
|
69
|
Yasunaka K, Abe F, Nagayama A, Okabe H,
Lozada-Pérez L, López-Villafranco E, Muñiz EE, Aguilar A and
Reyes-Chilpa R: Antibacterial activity of crude extracts from
Mexican medicinal plants and purified coumarins and xanthones. J
Ethnopharmacol. 97:293–299. 2005.PubMed/NCBI View Article : Google Scholar
|
|
70
|
Majali I, Qaralleh HN, Idid SZ, Saad S,
Susanti D and Althunibat OY: Potential antimicrobial activity of
marine sponge Neopetrosia exigua. J Basic Appl Res. 1:1–13.
2019.
|
|
71
|
Bodie AR, Micciche AC, Atungulu GG,
Rothrock MJ Jr and Ricke SC: Current trends of rice milling
byproducts for agricultural applications and alternative food
production systems. Front Sustainable Food Syst. 3(47)2019.
|
|
72
|
Ebada SS, de Voogd N, Kalscheuer R, Müller
WE and Proksch P: Cytotoxic drimane meroterpenoids from the
Indonesian marine sponge Dactylospongia elegans. Phytochem
Lett. 22:154–158. 2017.
|
|
73
|
Nasiri N, Taherizadeh MR and Gozari M:
Investigating the antimicrobial activity of bacteria associated
with the marine sponge Haliclona sp. collected from the
Persian Gulf. J Marine Med. 4:215–222. 2023.
|
|
74
|
Chen B, Zhao Q, Gu YC, Lan L, Wang CY and
Guo YW: Xishaeleganins A-D, Sesquiterpenoid hydroquinones
from Xisha marine sponge Dactylospongia elegans. Mar Drugs.
20(118)2022.PubMed/NCBI View Article : Google Scholar
|
|
75
|
Omar AM, Mohammad KA, Sindi IA, Mohamed GA
and Ibrahim SR: Unveiling the efficacy of sesquiterpenes from
marine sponge Dactylospongia elegans in inhibiting
dihydrofolate reductase using docking and molecular dynamic
studies. Molecules. 28(1292)2023.PubMed/NCBI View Article : Google Scholar
|
|
76
|
Yu HB, Yin ZF, Gu BB, Zhang JP, Wang SP,
Yang F and Lin HW: Cytotoxic meroterpenoids from the marine sponge
Dactylospongia elegans. Nat Prod Res. 35:1620–1626.
2021.PubMed/NCBI View Article : Google Scholar
|
|
77
|
Hong LL, Ding YF, Zhang W and Lin HW:
Chemical and biological diversity of new natural products from
marine sponges: A review (2009-2018). Mar Life Sci Technol.
4:356–372. 2022.PubMed/NCBI View Article : Google Scholar
|
|
78
|
Kim JA, Choi SS, Lim JK and Kim ES:
Unlocking marine treasures: Isolation and mining strategies of
natural products from sponge-associated bacteria. Nat Prod Rep.
42:1195–1225. 2025.PubMed/NCBI View Article : Google Scholar
|
|
79
|
Michev A, Orsini A, Santi V, Bassanese F,
Veraldi D, Brambilla I, Marseglia GL, Savasta S and Foiadelli T: An
overview of the role of tumor necrosis Factor-alpha in
epileptogenesis and its terapeutic implications. Acta Biomed. 92
(Suppl)(e2021418)2022.PubMed/NCBI View Article : Google Scholar
|
|
80
|
Taylor MW, Radax R, Steger D and Wagner M:
Sponge-associated microorganisms: Evolution, ecology, and
biotechnological potential. Microbiol Mol Biol Rev. 71:295–347.
2007.PubMed/NCBI View Article : Google Scholar
|
|
81
|
Zhang B, Zhang T, Xu J, Lu J, Qiu P, Wang
T and Ding L: Marine sponge-associated fungi as potential novel
bioactive natural product sources for drug discovery: A review.
Mini Rev Med Chem. 20:1966–2010. 2020.PubMed/NCBI View Article : Google Scholar
|
|
82
|
Khan F, Bamunuarachchi NI, Tabassum N and
Kim YM: Caffeic acid and its derivatives: Antimicrobial drugs
toward microbial pathogens. J Agric Food Chem. 69:2979–3004.
2021.PubMed/NCBI View Article : Google Scholar
|
|
83
|
Yang GZ, Zhu JK, Yin XD, Yan YF, Wang YL,
Shang XF, Liu YQ, Zhao ZM, Peng JW and Liu H: Design, synthesis,
and antifungal evaluation of novel quinoline derivatives inspired
from natural quinine alkaloids. J Agric Food Chem. 67:11340–11353.
2019.PubMed/NCBI View Article : Google Scholar
|
|
84
|
Luís Â, Silva F, Sousa S, Duarte AP and
Domingues F: Antistaphylococcal and biofilm inhibitory activities
of gallic, caffeic, and chlorogenic acids. Biofouling. 30:69–79.
2014.PubMed/NCBI View Article : Google Scholar
|
|
85
|
Pinho E, Ferreira IC, Barros L, Carvalho
AM, Soares G and Henriques M: Antibacterial potential of
northeastern Portugal wild plant extracts and respective phenolic
compounds. Biomed Res Int. 2014(814590)2014.PubMed/NCBI View Article : Google Scholar
|
|
86
|
Dinarvand M and Spain M: Identification of
bioactive compounds from marine natural products and exploration of
Structure-Activity Relationships (SAR). Antibiotics (Basel).
10(337)2021.PubMed/NCBI View Article : Google Scholar
|
|
87
|
Mahboub N, Cherfi I, Laouini SE, Bouafia
A, Benaissa A, Alia K, Alharthi F, Al-Essa K and Menaa F: GC/MS and
LC composition analysis of essential oil and extracts from wild
rosemary: Evaluation of their antioxidant, antimicrobial, and
Anti-inflammatory activities. Biomed Chromatogr.
39(e70084)2025.PubMed/NCBI View Article : Google Scholar
|
|
88
|
Rattanaphan P, Yaikhan T, Rachniyom P,
Mittraparp-Arthorn P, Vuddhakul V and Tansila N: Altered virulence
of non-indole-producing pathogenic bacteria by indole signaling.
Songklanakarin J Sci Technol. 45:515–520. 2020.
|
|
89
|
Hasan I, Asaduzzaman A, Rajia S, Fujii Y,
Kawsar S and Ozeki Y: Antibiofilm activity of lectins from plants
and marine invertebrates: A comparative study. Trends Carbohydrate
Res. 15:56–61. 2023.
|
|
90
|
Andrade JC, Braga MFBM, Guedes GMM,
Tintino SR, Freitas MA, Quintans LJ Jr, Menezes IRA and Coutinho
HDM: Cholecalciferol, ergosterol, and cholesterol enhance the
antibiotic activity of drugs. Int J Vitam Nutr Res. 88:244–250.
2018.PubMed/NCBI View Article : Google Scholar
|
|
91
|
Devkar HU, Juyal K, Thakur NL, Kaur P,
Parmar K, Pullapanthula R and Narayanan S: Antimicrobial potential
of marine Sponge-associated bacillus velezensis and stutzerimonas
Stutzeri from the Indian Coast: A genome mining and metabolite
profiling approach. Curr Microbiol. 82(280)2025.PubMed/NCBI View Article : Google Scholar
|
|
92
|
Blunt JW, Carroll AR, Copp BR, Davis RA,
Keyzers RA and Prinsep MR: Marine natural products. Nat Product
Rep. 35:8–53. 2018.
|
|
93
|
Zhang R, Wang H, Chen B, Dai H, Sun J, Han
J and Liu H: Discovery of anti-MRSA secondary metabolites from a
marine-derived fungus Aspergillus fumigatus. Mar Drugs.
20(302)2022.PubMed/NCBI View Article : Google Scholar
|
|
94
|
Ibrahim SR, Fadil SA, Fadil HA, Hareeri
RH, Alolayan SO, Abdallah HM and Mohamed GA: Dactylospongia
elegans-A promising drug source: Metabolites, bioactivities,
biosynthesis, synthesis, and structural-activity relationship. Mar
Drugs. 20(221)2022.PubMed/NCBI View Article : Google Scholar
|
|
95
|
Cornwell BH, Fisher JL, Morgan SG and
Neigel JE: Chaotic genetic patchiness without sweepstakes
reproduction in the shore crab Hemigrapsus oregonensis.
Marine Ecol Progress Series. 548:139–152. 2016.
|
|
96
|
Yang Z, Wei X, He J, Sun C, Ju J and Ma J:
Characterization of the noncanonical regulatory and transporter
genes in atratumycin biosynthesis and production in a heterologous
host. Mar Drugs. 17(560)2019.PubMed/NCBI View Article : Google Scholar
|
|
97
|
Wang J, Xu F, He C, Zhang L, Lu L, Wang X,
Qin Z and Shen B: High quality AlN epilayers grown on nitrided
sapphire by metal organic chemical vapor deposition. Sci Rep.
7(42747)2017.PubMed/NCBI View Article : Google Scholar
|
|
98
|
Li Z: Sponge and Coral Microbiomes. In: Li
Z, (ed.). Symbiotic Microbiomes of Coral Reefs Sponges and Corals.
Dordrecht, Springer Netherlands, pp17-28, 2019.
|
|
99
|
Li J, Yang F, Wang Z, Wu W, Liu L, Wang
SP, Zhao BX, Jiao WH, Xu SH and Lin HW: Unusual anti-inflammatory
meroterpenoids from the marine sponge Dactylospongia sp.
Organic Biomolecular Chem. 16:6773–6782. 2018.PubMed/NCBI View Article : Google Scholar
|
|
100
|
Avila C, Núñez-Pons L and Moles J: From
the tropics to the poles: Chemical defense strategies in sea slugs
(Mollusca: Heterobranchia). Chemical Ecology: CRC Press, pp71-163,
2018.
|