Introduction
Moyamoya disease (MMD) is characterized by the
progressive occlusion of bilateral carotid forks, which are
associated with moyamoya vessels formation at the base of the brain
(1,2). MMD tends to be more prevalent in Asian
countries. Since the etiology of MMD remains poorly understood, the
criteria for the diagnosis of MMD are mainly based on
characteristic angiographic findings. Ischemic attack and
hemorrhage are two of the most common presentations of MMD. In
particular, the hemorrhagic subtype is associated with a poor
clinical course, which is found in 20% patients with MMD (3), but only 50% patients experience
adequate recovery after the first hemorrhagic event (4). There is no clinical scale for
stratifying the occurrence of ischemic MMD (iMMD) and hemorrhagic
MMD (hMMD), nor is there a tool for the prediction of rebleeding or
hemorrhagic transformation. Previous studies have suggested that
different molecular mechanisms underlie the two subtypes.
RNA expression studies offer unique opportunities
for understanding the pathogenesis of a wide variety of
neurological diseases (5,6). Several studies suggest that non-coding
RNA, including microRNAs (miRNAs) and long non-coding RNAs
(lncRNAs), are associated with hemorrhagic neurological diseases
(7-10).
However, it remains unclear whether altered expression of such
non-coding RNAs directly results in hemorrhage in cerebrovascular
diseases. Circular RNAs (circRNAs) form another class of stable,
single-stranded, non-coding RNAs that are formed by back-splicing
events through exon or intron circularization (11-13).
Previous studies have reported that circRNAs can regulate gene
expression on transcriptional or post-transcriptional levels by
functioning as miRNA sponges (11,14).
In addition, stroke has been reported to alter the expression of
circRNAs with possible functional implications in poststroke
pathophysiology (15). Since
circRNA-based research is an emerging area of investigation, to the
best of our knowledge, circRNA expression is abnormally expressed
in MMD (16). No study to date has
investigated the circRNA expression profile in hMMD.
Therefore, the present study focused on the
potential differences in the circRNA expression profile between
hMMD and iMMD. circRNA dysregulation may serve a role in the
hemorrhage of MMD. CircRNAs may become potential future biological
targets and prognostic indicators for hemorrhage in MMD.
Materials and methods
Patient selection and sample
collection
In total, adult patients diagnosed with MMD
presenting with hemorrhage or transient ischemic attack were
recruited from Beijing Tiantan Hospital (Beijing, China) between
March and July 2016. The diagnosis of MMD adhered to the guidelines
established by the Research Committee on Moyamoya Disease of the
Ministry of Health, Labor, and Welfare of Japan (17). Inclusion criteria were: Age ≥18
years; diagnosis of definite MMD per Research Committee guidelines;
presenting with either hemorrhage or transient ischemic attack; no
prior surgical revascularization; no other cerebrovascular
conditions; no systemic diseases that could affect RNA expression;
informed consent provided. Exclusion criteria encompassed pediatric
patients, those with quasi-MMD (moyamoya syndrome) and individuals
with history of other cerebrovascular conditions, hypertension or
diabetes to mitigate potential confounding factors. Patients with
conditions known to influence RNA expression were excluded,
including autoimmune diseases, chronic infections, malignancies,
chronic inflammatory conditions, recent major surgeries, and any
use of immunosuppressive or anti-inflammatory medications. Finally,
12 patients (5 males and 7 females, age 22-50 years old) were
included in microarray analysis.
Whole venous blood samples (12 patients with MMD for
microarray and 22 patients for validation, 3 ml each) were obtained
from the patients with MMD 2 weeks post-symptom onset, prior to any
revascularization procedures. Blood samples were collected between
March and July 2016 with patients' written informed consent for
biobanking and future research use. The specific circRNA analysis
protocol was reviewed and approved by the Ethics Committee Review
Board of Beijing Tiantan Hospital (approval no. KYSQ2020-161-01)
prior to conducting the molecular studies.
RNA extraction and microarray
analysis
Total RNA was isolated from blood samples using
TRIzol® reagent (Thermo Fisher Scientific, Inc.)
following the manufacturer's protocol. RNA quantity and integrity
were assessed using a NanoDrop ND-1000 spectrophotometer (Thermo
Fisher Scientific, Inc.) and agarose gel electrophoresis,
respectively. Sample labeling and circRNA array hybridization were
performed according to the manufacturer's protocol (Arraystar,
Inc.). CircRNA enrichment, amplification and fluorophore-labeled
cRNA synthesis were performed using the Super RNA Labeling kit
(Arraystar, Inc.), followed by purification with the RNeasy Mini
kit (Qiagen GmbH). The labeled cRNAs were hybridized onto the
Arraystar Human circRNA Array (6x7K; Arraystar, Inc.) for 17 h at
65˚C in an Agilent Hybridization Oven (Agilent Technologies, Inc.).
Post-hybridization, arrays were washed, fixed and scanned using a
G2505C scanner (Agilent Technologies, Inc.). Raw data extraction
was conducted using Feature Extraction v.11.0.1.1 software (Agilent
Technologies, Inc.). Subsequent data processing, including quantile
normalization, was performed using R v.3.3 software (R Foundation
for Statistical Computing; https://www.r-project.org). Differentially expressed
circRNAs were defined as those with fold changes ≥2.00 and
P<0.05. Hierarchical clustering was employed to visualize
distinct circRNA expression patterns among samples.
Reverse transcription-quantitative
(RT-q) PCR
To validate the microarray data, RT-qPCR was
conducted. Cells were harvested at approximately 80% confluence
(approximately 1x106 cells/ml) for RNA extraction. Total
RNA was reverse transcribed into cDNA using Superscript III reverse
transcriptase (Invitrogen; Thermo Fisher Scientific, Inc.)
according to the manufacturer's instructions. CircRNA expression
levels were quantified using a ViiA 7 real-time PCR system (Applied
Biosystems; Thermo Fisher Scientific, Inc.) with SYBR Green Master
Mix (Applied Biosystems; Thermo Fisher Scientific, Inc.) following
the manufacturer's protocol. Divergent primers were designed to
specifically amplify circRNAs and differentiate them from their
linear isoforms (Table I). The PCR
cycling conditions were as follows: initial denaturation at 95˚C
for 10 min, followed by 40 cycles of denaturation at 95˚C for 15
sec, annealing at 60˚C for 30 sec, and extension at 72˚C for 30
sec. β-actin served as the internal control. The expression level
of each circRNA was calculated as a fold change using the
2-ΔΔCq method (18).
 | Table IPrimers used for reverse
transcription-quantitative PCR. |
Table I
Primers used for reverse
transcription-quantitative PCR.
| Gene | Primer |
|---|
| β-actin | Forward: 5'
GTGGCCGAGGACTTTGATTG3' |
| | Reverse: 5'
CCTGTAACAACGCATCTCATATT3' |
|
hsa_circRNA_103572 | Forward: 5'
ATGTGGAAAATTTCCTAGAAGC 3' |
| | Reverse: 5'
AGGTCTGTCATCACTCTGAGGT 3' |
|
hsa_circRNA_103574 | Forward: 5'
GTACCTAAATTAACAATGGCGA 3' |
| | Reverse: 5'
AAGGGGTGAAGCATGACCT 3' |
|
hsa_circRNA_029937 | Forward: 5'
GGCCATAGGAAAAGGATACAG 3' |
| | Reverse: 5'
CTCTAGGTCCCAAGAATTTACC 3' |
|
hsa_circRNA_104293 | Forward: 5'
GCACAGATCTGATTCTGAACGT 3' |
| | Reverse: 5'
TCATTGGATATGTCCTGATAGTCC 3' |
|
hsa_circRNA_025016 | Forward: 5'
TATTCCCTTTCCAGAAGATGAT 3' |
| | Reverse: 5'
CATAGTTGGAACCAGGTTGG 3' |
|
hsa_circRNA_091419 | Forward: 5'
CGTGTTTTCCTCTCTGAATCTG 3' |
| | Reverse: 5'
TCGCTTAATCCTGAAAGTCTTG 3' |
|
hsa_circRNA_060184 | Forward: 5'
ACCCGCCATGGGAGTGTG 3' |
| | Reverse: 5'
GGGGCTTCCAGCAGTGCT 3' |
Bioinformatics analysis and
circRNA/miRNA gene network construction
The microarray data are in the supplementary tables.
All raw relevant datasets are in the supplementary materials. The
parent linear mRNAs of differentially expressed circRNAs were
subjected to Gene Ontology (GO) analysis (http://www.geneontology.org) to elucidate the
functional enrichment of these coding genes. Pathway analyses were
conducted based on the Kyoto Encyclopedia of Genes and Genomes
(KEGG; http://www.genome.ad.jp/kegg/).
A circRNA/miRNA gene network was constructed for the
differentially expressed circRNAs identified from microarray and
RT-qPCR validation experiments. CircRNA/miRNA interactions were
predicted using the Arraystar miRNA target prediction software
(version 1.0, Arraystar, Inc.; https://www.arraystar.com), which integrates
TargetScan and miRanda algorithms (19). miRNA target gene analysis was
performed using miRTarBase (20).
All miRNA gene targets were experimentally validated with strong
evidence (western blotting or Reporter assay). The circRNA/miRNA
gene network was visualized using Cytoscape 2.8.2 (https://cytoscape.org).
Statistical analysis
All data are presented as the mean ± standard error.
Statistical comparisons were performed using paired t-tests or
independent t-tests as appropriate. All statistical analyses were
conducted using R v.3.3 software (R Foundation for Statistical
Computing; https://www.r-project.org). P<0.05
was considered to indicate a statistically significant
difference.
Results
Overview of circRNA profiles
A total of 12 patients with MMD, namely 6 hMMD and 6
age and sex matched patients with iMMD, were enrolled into the
present study (Table SI). The
expression profiles of human circRNAs were obtained by microarray
analysis (Table SII).
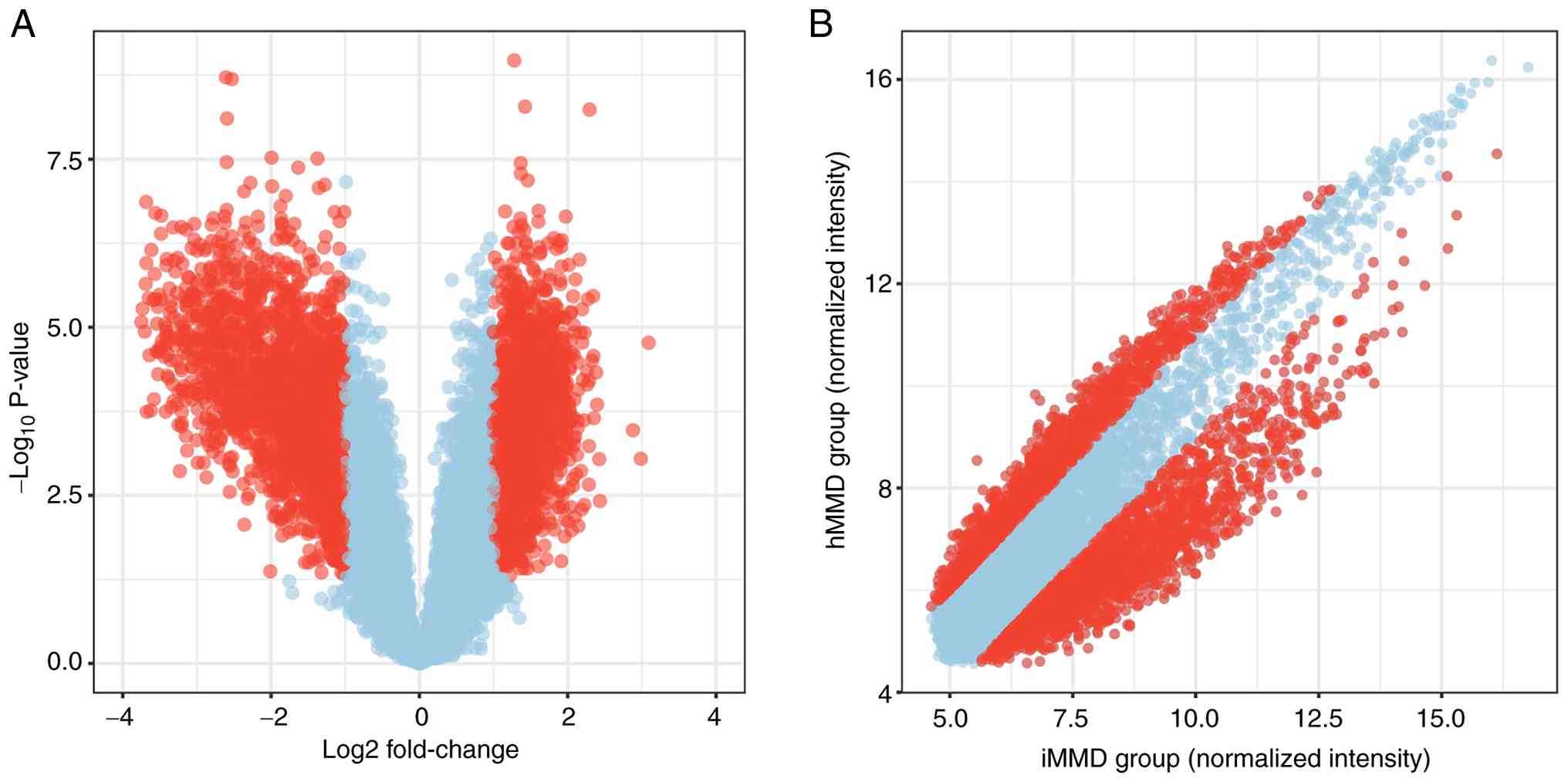
Differentially expressed circRNAs with statistical significance
(fold changes ≥2.0 and P<0.05) between hMMD and iMMD groups were
identified using a volcano plot (Fig.
1A) and scatter plot (Fig. 1B).
A total of 3,607 circRNAs with expression change >2x were
identified (Table SIII). Compared
with iMMD, 1,940 circRNAs were upregulated and 1,967 circRNAs were
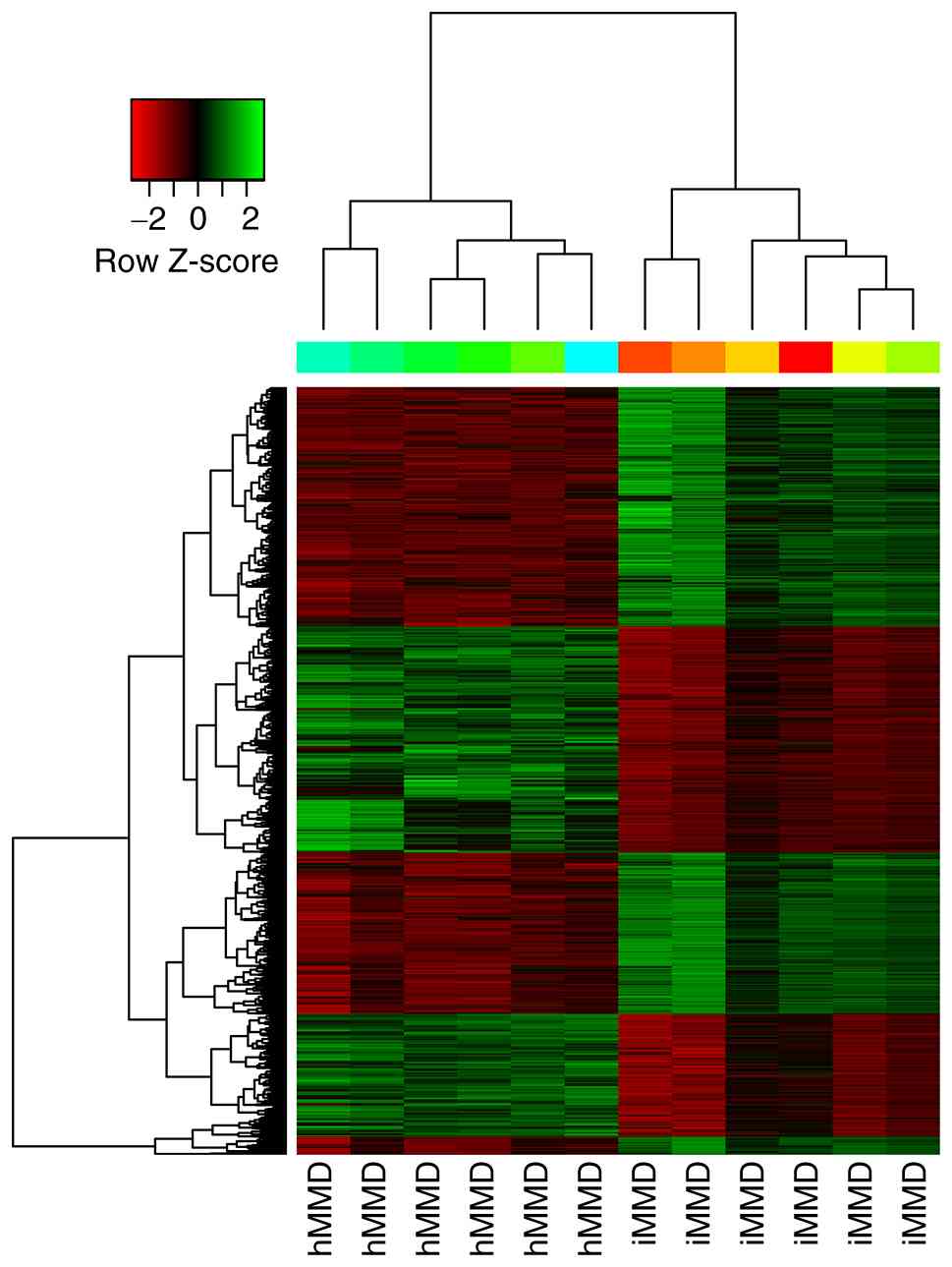
downregulated in hMMD samples. Hierarchical clustering revealed the
circRNA expression patterns between hMMD and iMMD were
significantly different (Fig. 2).
The complete dataset from the present study is available in the
supplementary materials, including all raw microarray data files
and associated documentation.
RT-qPCR validation of the microarray
data
RT-qPCR was conducted in a group of 11 hMMD and 11
iMMD samples to verify the differential expression of the candidate
circRNAs. A total of seven circRNAs were selected for RT-qPCR
validation based on: i) Magnitude of differential expression (fold
change >2), ii) statistical significance (P<0.05), and iii)
predicted interactions with MMD-relevant pathways. A total of seven
circRNAs, including four upregulated circRNAs (circRNA-025016,
circRNA-104293, circRNA-060184 and circRNA-091419) and three
downregulated circRNAs (circRNA-029937, circRNA-103574 and
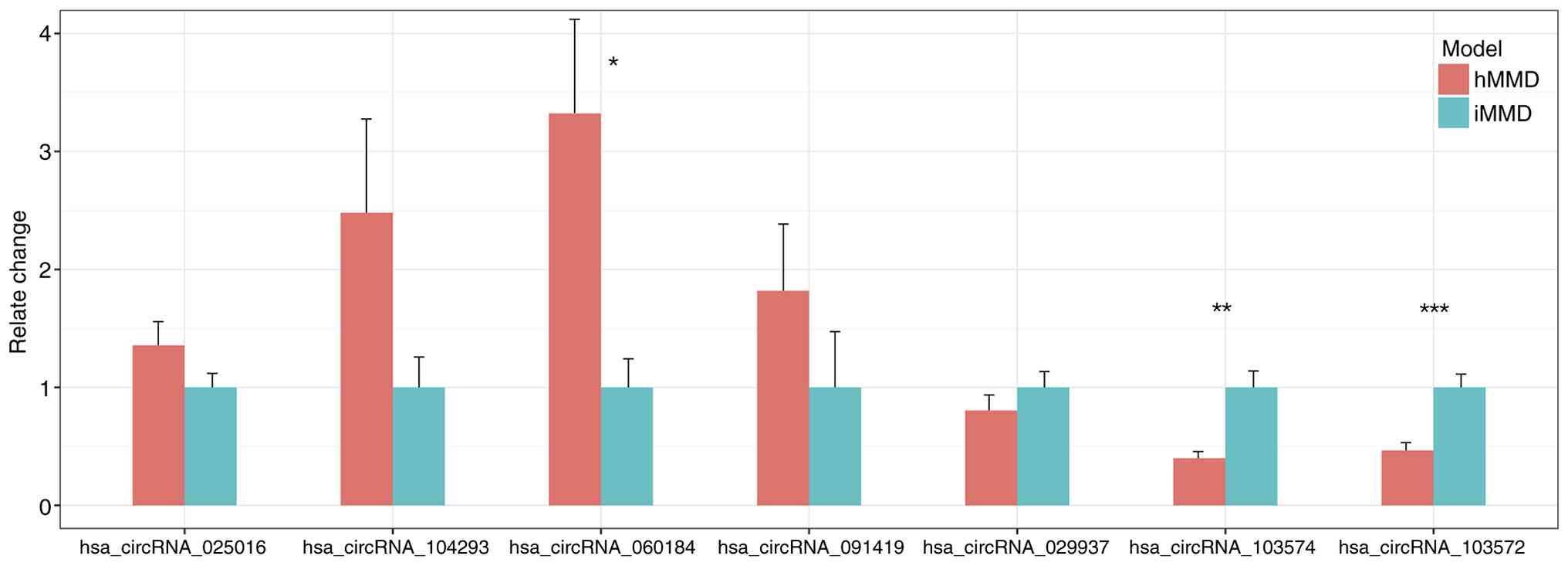
circRNA-103572), were selected for further assessment. The results
obtained from RT-qPCR were consistent with the RNA sequencing data
(Fig. 3), RT-qPCR validation
confirmed significant upregulation of circRNA-025016,
circRNA-104293, circRNA-060184 and circRNA-091419 in hMMD samples
compared to iMMD. Conversely, circRNA-029937, circRNA-103574 and
circRNA-103572 showed significant downregulation in hMMD
samples.
Bioinformatics analysis
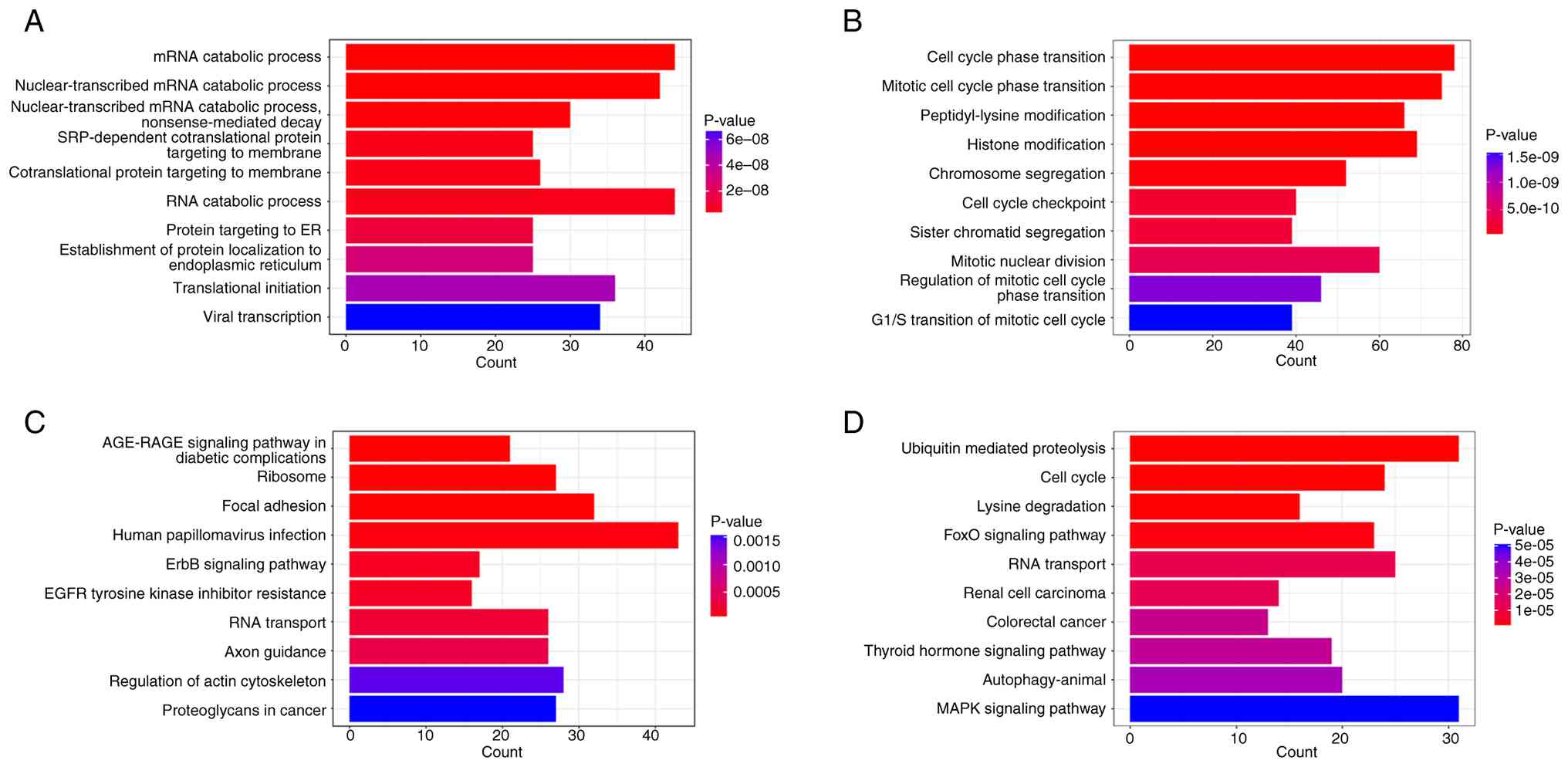
GO and KEGG analysis of differentially expressed
circRNAs was next performed (Fig.
4). The GO and KEGG pathway analyses revealed distinct patterns
between hMMD and iMMD (Fig. 4).
Fig. 4A shows upregulated circRNAs
in hMMD were enriched in mRNA catabolic processes and protein
targeting pathways. Fig. 4B
indicates downregulated circRNAs were mainly involved in cell cycle
regulation. In KEGG analysis, Fig.
4C demonstrates upregulated pathways including AGE-RAGE
signaling and focal adhesion, while Fig. 4D shows downregulated pathways with
ubiquitin-mediated proteolysis being the most significant. In GO
analysis of biological processes, ‘mRNA catabolic process’ and
‘nuclear-transcribed mRNA catabolic process’ were two of the
biological processes with the most significance in upregulated
circRNAs in hMMD samples. By contrast, ‘cell cycle phase
transition’ and ‘mitotic cell cycle phase transition’ were the
biological processes with the most significance amongst the
downregulated circRNAs in the hMMD samples (Tables II and SIV).
 | Table IITop five GO biological processes of
upregulated and downregulated target genes with most significance
in hMMD compared with iMMD. |
Table II
Top five GO biological processes of
upregulated and downregulated target genes with most significance
in hMMD compared with iMMD.
| GO term | Fold
enrichment | P-value |
|---|
| A, Upregulated
target genes |
|---|
| GO:0006402 mRNA
catabolic process | 2.979113618 | 4.34E-11 |
| GO:0000956
nuclear-transcribed mRNA catabolic process | 3.061329416 | 4.82E-11 |
| GO:0000184
nuclear-transcribed mRNA catabolic process, nonsense-mediated
decay | 3.571550985 | 5.75E-10 |
| GO:0006614
SRP-dependent cotranslational protein targeting to membrane | 3.840377404 | 3.09E-09 |
| GO:0006613
cotranslational protein targeting to membrane | 3.714413025 | 3.26E-09 |
| B, Downregulated
target genes |
| GO:0044770 cell
cycle phase transition | 2.764948269 | 4.06E-13 |
| GO:0044772 mitotic
cell cycle phase transition | 2.818120352 | 4.06E-13 |
| GO:0018205
peptidyl-lysine modification | 2.811021812 | 2.10E-11 |
| GO:0016570 histone
modification | 2.719584675 | 2.12E-11 |
| GO:0007059
chromosome segregation | 2.764948269 | 1.57E-08 |
KEGG pathway analysis demonstrated 10 enrichment
pathways in the upregulated circRNAs and 36 enriched pathways in
the downregulated circRNAs (Tables
III and SV). Among them, the
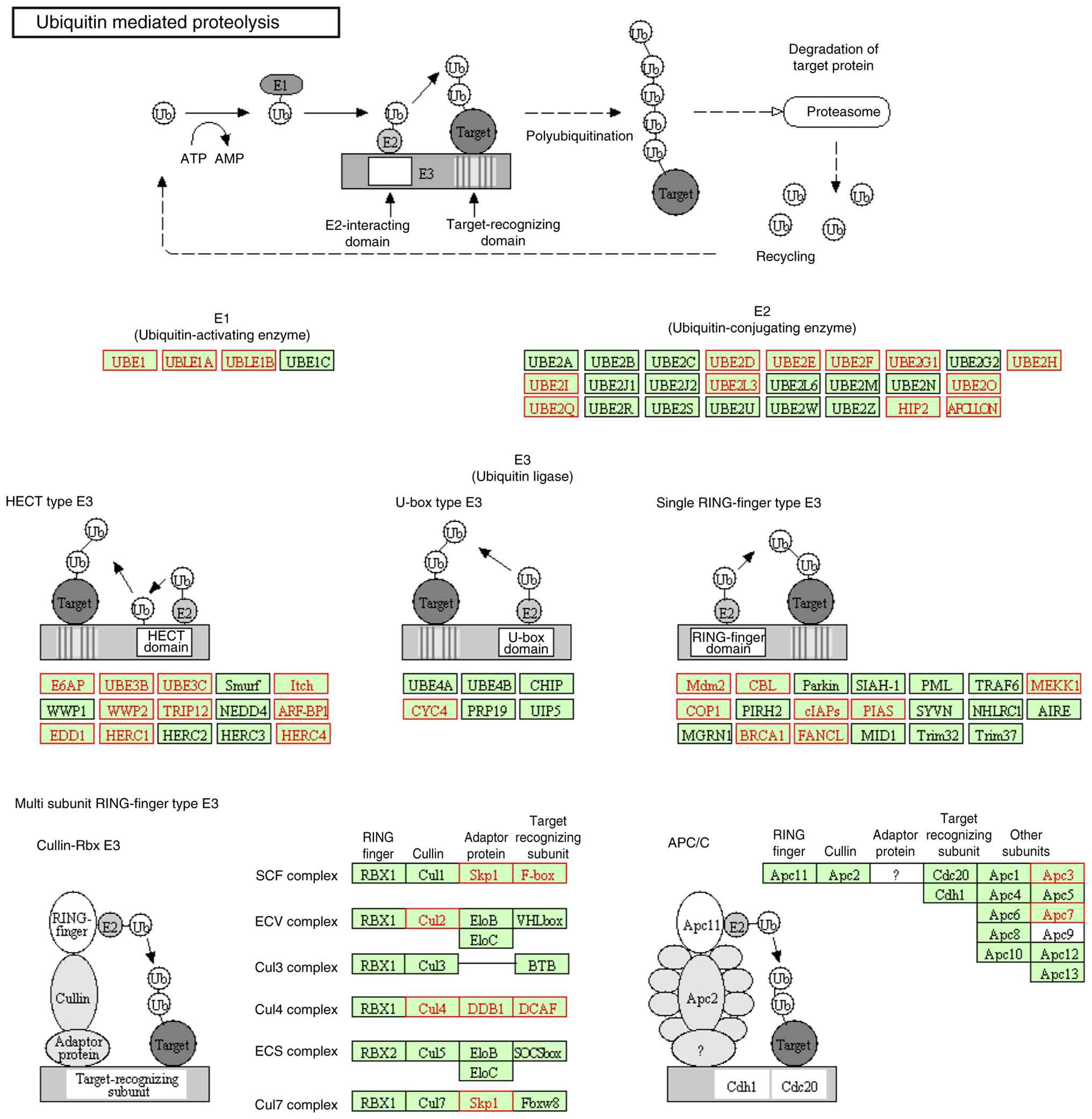
‘Ubiquitin mediated proteolysis’ (Fig.
5) and ‘Cell cycle’ pathways were the top enriched pathways
with the most significance. Fig. 5
shows the ubiquitin-mediated proteolysis pathway from the KEGG
database. The red boxes highlight differentially expressed genes
between hMMD and iMMD, including ubiquitin-activating enzyme (E1),
ubiquitin-conjugating enzymes (E2), and ubiquitin ligases (E3).
This pathway's dysregulation in hMMD suggests altered protein
degradation may contribute to vessel wall instability. The
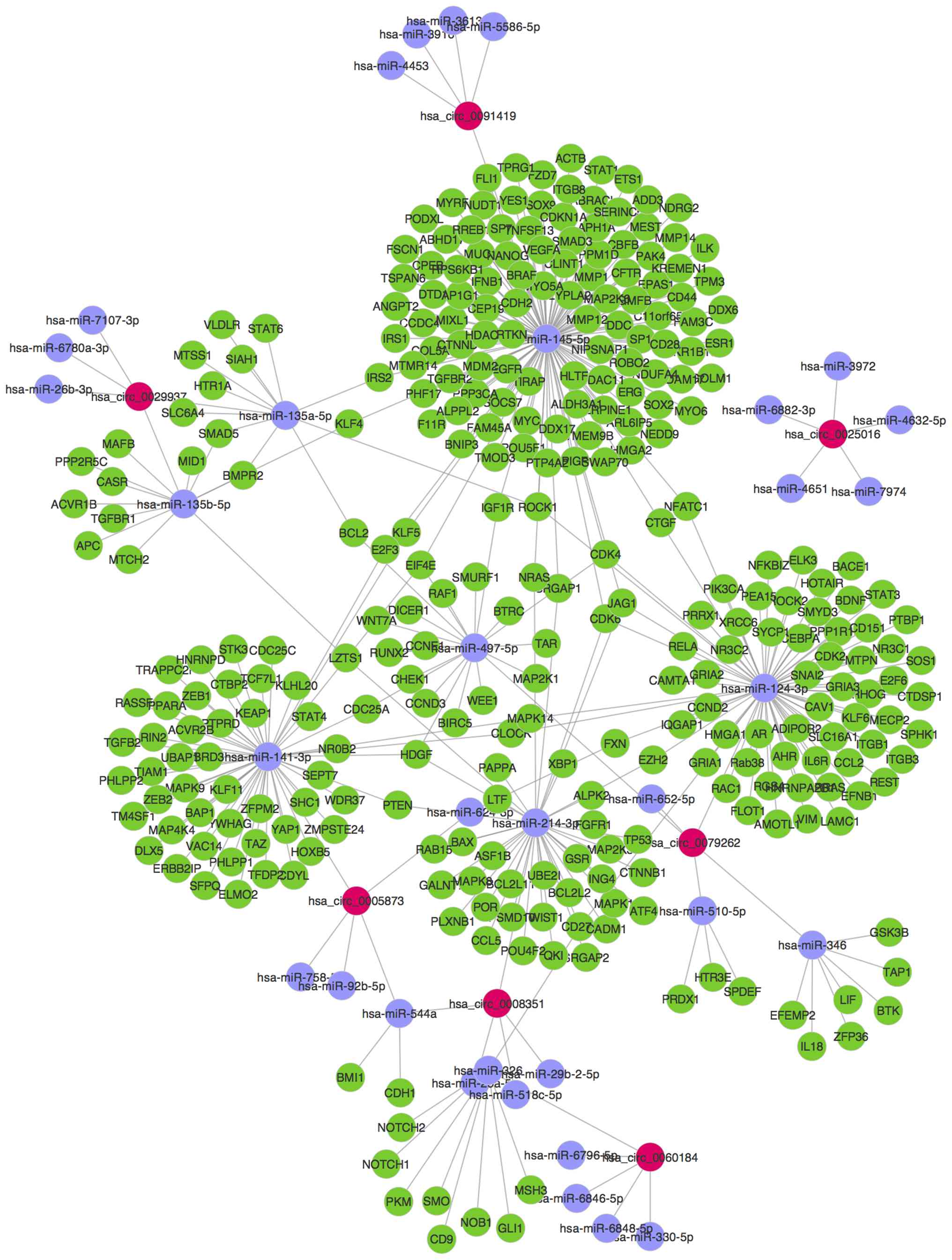
validated circRNAs, 34 predicted miRNAs and 308 target genes were
next used to construct the circRNA/miRNA network (Fig. 6).
 | Table IIIKEGG pathways of upregulated and
downregulated target genes with most significance in hMMD compared
with iMMD. |
Table III
KEGG pathways of upregulated and
downregulated target genes with most significance in hMMD compared
with iMMD.
| KEGG term | Fold
enrichment | P-value |
|---|
| A, Upregulated
target genes |
|---|
| hsa04933 AGE-RAGE
signaling pathway in diabetic complications | 2.914065180 | 6.42E-06 |
| hsa03010
Ribosome | 2.408564077 | 1.44E-05 |
| hsa04510 Focal
adhesion | 2.209083152 | 1.51E-05 |
| hsa05165 Human
papillomavirus infection | 1.863478364 | 4.39E-05 |
| hsa04012 ErbB
signaling pathway | 2.715598947 | 1.00E-4 |
| B, Downregulated
target genes |
| hsa04120 Ubiquitin
mediated proteolysis | 3.950900593 | 2.02E-11 |
| hsa04110 Cell
cycle | 3.379438385 | 1.02E-07 |
| hsa00310 Lysine
degradation | 4.735032313 | 1.13E-07 |
| hsa04068 FoxO
signaling pathway | 3.042347940 | 1.33E-06 |
| hsa03013 RNA
transport | 2.552694686 | 1.23E-05 |
Discussion
In the present study, the circRNA expression of hMMD
and iMMD were comprehensively profiled by microarray analysis.
Compared with iMMD samples, a total of 3,607 significantly
differentiated circRNAs in hMMD were detected. Subsequently, the
differentially expressed circRNAs were identified to be involved in
several biological processes and signaling pathways, such as ‘mRNA
catabolic process’ and ‘ubiquitin mediated proteolysis’, according
to GO analysis and KEGG pathway analysis. In addition, a network
map of circRNA/miRNA gene interactions was also constructed for the
validated significantly differentiated circRNAs. These results
suggested that there was a significant difference in the circRNA
expression profile between the iMMD and hMMD samples. Several key
circRNAs may show promise as candidate biomarkers for hemorrhage in
MMD.
CircRNAs have been recently found to be pervasively
transcribed in the genome (11,21).
It was previously reported that circRNAs can reverse the inhibitory
effects of miRNAs on their target mRNAs by directly binding to
miRNAs through miRNA response elements (22). In addition, previous studies
suggested that circRNAs are enriched in the brain and may
participate in regulating synaptic function and neural plasticity
(23,24). Dysregulated circRNAs have been
reported to be associated with several human diseases, including
neurological disease, cardiovascular system diseases and cancers
(25-27).
Circular antisense non-coding RNA in the INK4 locus (cANRIL) was
documented to influence INK4/ADP ribosylation factor expression and
increase the risk of atherosclerotic vascular disease (28). This finding suggests that circRNAs
are involved in the development of atherosclerotic cerebrovascular
disease (23). The mechanism of MMD
remains poorly understood. However, genetic and environmental
factors were considered to be vital in the development of the
vascular stenosis and MMD vessel formations (29).
A previous genome-wide association study has shown
Ring Finger 213 (RNF213) to be an important MMD susceptibility gene
(30). Vasculogenesis and
angiogenesis, which require endothelial cell proliferation and
migration, form the two central processes involved in the
development of biological revascularization (31). The arteriogenesis process, which
refers to the formation of collateral circulation, is typically
activated by the increased fluid shear stress generated by the
pressure difference between perfusion territories (32). The proliferation of endothelial and
smooth muscle cells may lead to aberrant angiogenesis (29). The associated changes in circulating
endothelial/smooth muscle progenitor cells (33), angiogenesis (33-35)
and caveolin (36), may also be
involved. The pathogenesis of MMD had also been associated with
non-coding RNAs in previous studies. miRNAs, which are small
non-coding RNAs ~23 nucleotides in length, can negatively regulate
the expression of proteins by altering their gene expression
through post-transcriptional repression or mRNA degradation
(37). miRNAs have been reported to
serve an important role in the regulation of proliferation and
aging of various tissues. A previous microarray study on miRNAs
profiles in serum from patients with MMD suggested that elevated
serum levels of miRNAs are associated with RNF213(38). Another study previously revealed
that the increased expression of miRNA Let-7c in patients with MMD
may also contribute to MMD pathogenesis by targeting RNF213
expression (39). In addition,
annexin 1, which is expressed in endothelial and smooth muscle
cells (SMC), is a gene target of miRNA-196a to mediate apoptosis
and inhibition of cell proliferation (29).
Although there is notable heterogeneity in clinical
symptoms depending on the age of onset and ethnicity (40-42),
patients with MMD typically present with ischemic and/or
hemorrhagic stroke. These two subtypes may have pathogenic
differences. The proposed pathophysiologic mechanism for hemorrhage
in MMA is long-term hemodynamic stress to collateral vessels
(43). Theoretically, impaired
perfusion results in hemodynamic stress on the vessel wall and
facilitates dilation or micro-aneurysm formation in collateral
vessels. Dilatation and abnormal branching of the anterior
choroidal artery and/or posterior communicating artery are viable
predictors of hemorrhage in adult patients with MMD (44). In addition, one previous study
suggested that by using 7T time-of-flight magnetic resonance
angiography, ventricular micro-aneurysms in MMD angiopathy
collateral vessels can be detected (45). The non-coding RNAs have been
suggested to be involved in other hemorrhagic cerebral vascular
diseases. A previous study suggests that lncRNAs may contribute to
the pathogenesis of cerebral aneurysms by regulating loss of the
contractile SMC phenotype (46).
The distinction between hemorrhagic and ischemic presentations of
MMD has important clinical implications. Zhao et al
(47) previously demonstrated that
patients with hMMD show distinct imaging characteristics and
collateral patterns compared to iMMD, suggesting different
pathophysiological mechanisms. The circRNA findings of the present
study provided molecular support for this, revealing divergent
expression patterns between subtypes. The identified circRNA
networks may help explain the different propensities for hemorrhage
compared with ischemia (16). The
different circRNA expression may serve a role in different vessel
formation in different MMD subgroups.
In addition, circRNAs may serve roles in the
pathogenesis of hMMD by regulating SMC proliferation and TGF-β
signaling. As shown in the network (Fig. 6), circRNA-0005873 may serve a role
in the expression of TGF-β by regulating miRNA-141-3p. Alterations
in normal TGF-β signaling have been implicated in the
pathophysiology of several vascular disorders, including
atherosclerosis and primary pulmonary hypertension (48). Another study suggested that TGF-β is
one of the underlying factors contributing to the development of
thoracic aortic aneurysm (49). As
shown in the network, pappalysin-1 (PAPPA) was the direct target
gene of miRNA-141. A previous study has also suggested that
miRNA-141 can inhibit vascular SMC proliferation through targeting
PAPPA (50).
The ubiquitin mediated proteolysis pathway was
detected as the top significant pathway according to KEGG analysis.
These results suggested that circRNAs may serve important roles in
proteolysis processes in hemorrhage. Proteases, including thrombin
and MMPs, were previously reported to have complex functions in the
brain under both normal and pathological conditions (49). MMPs are endopeptidases that can
degrade components of the extracellular matrix. The MMPs serve an
important role in normal and atherosclerotic blood vessels by being
involved in plaque disruption (51). Increased vascular MMP2 or MMP9
expression is involved in the pathogenesis of spontaneous
intracranial hemorrhage in patients with cerebral amyloid
angiopathy (52). Abdominal aortic
aneurysm expansion is likely to result from increasing proteolysis
related to increasing MMP-9 expression (53).
Proteolysis has been associated with the rupture of
abdominal aortic aneurysms. A previous study has shown that Smad7
can interact with the heteromeric TGF-β receptor complex and
recruits the E3 ubiquitin-ligases Smurf1 and Smurf2, targeting the
receptors for degradation to terminate the signaling response
(54). These factors may work in
concert to modify and direct the response to signals through this
complex pathway. Additionally, MMPs have been reported to serve a
significant role in regulating angiogenesis, the process of new
blood vessel formation (55). An
enhanced understanding of the molecular mechanisms involved in
hemorrhage in MMD may potentially lead to novel therapeutic
strategies against this potentially lethal condition. While acute
vascular events such as stroke can alter circRNA expression
patterns, emerging evidence suggests circRNAs may also play
causative roles in vascular pathology. The findings in the present
study of distinct circRNA profiles between hMMD and iMMD suggested
these molecules could be both markers and mediators of disease
progression. Further mechanistic studies are needed to fully
elucidate the complex interplay between circRNAs and vascular
remodeling in MMD.
The present study had several limitations that
warrant consideration. All samples were acquired from a single
ethnic group in mainland China. Consequently, different circRNA
signatures may exist across diverse ethnic groups. A limitation of
the present study is the relatively small sample size. While it
detected significant differences between groups, larger cohorts
will be needed to validate these findings and establish clinical
utility of circRNA biomarkers. In addition, the functions of
circRNAs in MMD were analyzed based on bioinformatics predictions.
While this approach is widely used in non-coding RNA research,
future studies on the specific interactions of circRNAs with miRNAs
and the downstream effects on hMMD and iMMD signaling pathways are
necessary. A key limitation is the descriptive nature of the
current findings and a lack of mechanistic studies.
To conclude, to the best of the authors' knowledge,
the present study represented the first comparison of circRNA
expression profiles in hMMD and iMMD samples. These findings
expanded on the understanding of the mechanisms underlying
hemorrhage in MMD, which may provide novel insights for developing
therapeutic interventions for hemorrhagic complications of MMD.
Supplementary Material
Baseline clinical characteristics of
patients with ischemic and hemorrhagic moyamoya disease.
Microarray analysis data.
circRNA expression change >2
fold.
GO analysis BP_up
Downregulated KEGG analysis.
Acknowledgements
Not applicable.
Funding
Funding: The present study was funded by ‘13th Five-Year Plan’
National Science and Technology Supporting Plan (grant no.
2015BAI12B04), the National Natural Science Foundation of China
(grant no. 81371292) and Beijing Municipal Administration of
Hospitals' Mission Plan (grant no. SML20150501).
Availability of data and materials
The datasets generated in the current study are not
publicly available to ncbi.nlm.nih.gov due to institutional policy and
regional data sharing restrictions but may be requested from the
corresponding author. Raw data are provided in the supplementary
materials of this article.
Authors' contributions
MZ and JZ participated in study conceptualization
and methodology development. WL supervised the project, acquired
funding, and provided resources. XY performed data curation, formal
analysis and statistical analysis. QZ and YZ were responsible for
investigation, experimental work and data validation. MZ drafted
the original manuscript, while YZ and JZ reviewed and edited the
manuscript. QZ and YZ were responsible for confirming the
authenticity of all raw data. All authors read and approved the
final manuscript.
Ethics approval and consent to
participate
The patients provided written informed consent for
biobanking and future research use. The specific circRNA analysis
protocol was reviewed and approved by the Ethics Committee Review
Board of Beijing Tiantan Hospital (Beijing, China; approval no.
KYSQ2020-161-01) prior to conducting the molecular studies.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Suzuki J and Takaku A: Cerebrovascular
moyamoya disease: Disease showing abnormal net-like vessels in base
of brain. Arch Neurol. 20:288–299. 1969.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Suzuki J and Kodama N: Moyamoya disease-a
review. Stroke. 14:104–109. 1983.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Baba T, Houkin K and Kuroda S: Novel
epidemiological features of moyamoya disease. J Neurol Neurosurg
Psychiatry. 79:900–904. 2008.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Kobayashi E, Saeki N, Oishi H, Hirai S and
Yamaura A: Long-term natural history of hemorrhagic moyamoya
disease in 42 patients. J Neurosurg. 93:976–980. 2000.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Sharp FR, Xu H, Lit L, Walker W, Apperson
M, Gilbert DL, Glauser TA, Wong B, Hershey A, Liu DZ, et al: The
future of genomic profiling of neurological diseases using blood.
Arch Neurol. 63:1529–1536. 2006.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Sharp FR, Xu H, Lit L, Walker W, Pinter J,
Apperson M and Verro P: Genomic profiles of stroke in blood.
Stroke. 38 (2 Suppl):S691–S693. 2007.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Wardlaw JM, Brazzelli M, Chappell FM,
Miranda H, Shuler K, Sandercock PAG and Dennis MS: ABCD2 score and
secondary stroke prevention: Meta-analysis and effect per 1,000
patients triaged. Neurology. 85:373–380. 2015.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Amarenco P, Labreuche J and Lavallée PC:
Patients with transient ischemic attack with ABCD2 <4 can have
similar 90-day stroke risk as patients with transient ischemic
attack with ABCD2 ≥4. Stroke. 43:863–865. 2012.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Sanders LM, Srikanth VK, Blacker DJ,
Jolley DJ, Cooper KA and Phan TG: Performance of the ABCD2 score
for stroke risk post TIA: Meta-analysis and probability modeling.
Neurology. 79:971–980. 2012.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Perry JJ, Sharma M, Sivilotti ML,
Sutherland J, Symington C, Worster A, Émond M, Stotts G, Jin AY,
Oczkowski WJ, et al: Prospective validation of the ABCD2 score for
patients in the emergency department with transient ischemic
attack. CMAJ. 183:1137–1145. 2011.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Memczak S, Jens M, Elefsinioti A, Torti F,
Krueger J, Rybak A, Maier L, Mackowiak SD, Gregersen LH, Munschauer
M, et al: Circular RNAs are a large class of animal RNAs with
regulatory potency. Nature. 495:333–338. 2013.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Esteller M: Non-coding RNAs in human
disease. Nat Rev Genet. 12:861–874. 2011.PubMed/NCBI View
Article : Google Scholar
|
|
13
|
Vicens Q and Westhof E: Biogenesis of
circular RNAs. Cell. 159:13–14. 2014.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Wilusz JE and Sharp PA: Molecular biology:
A circuitous route to noncoding RNA. Science. 340:440–441.
2013.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Mehta SL, Pandi G and Vemuganti R:
Circular RNA expression profiles alter significantly in mouse brain
after transient focal ischemia. Stroke. 48:2541–2548.
2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Zhao M, Gao F, Zhang D, Wang S, Zhang Y,
Wang R and Zhao J: Altered expression of circular RNAs in Moyamoya
disease. J Neurol Sci. 381:25–31. 2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Research Committee on the Pathology and
Treatment of Spontaneous Occlusion of the Circle of Willis; Health
Labour Sciences Research Grant for Research on Measures for
Infractable Diseases: Guidelines for diagnosis and treatment of
moyamoya disease (spontaneous occlusion of the circle of Willis).
Neurol Med Chir(Tokyo). 52:245–266. 2012.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) Method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Pasquinelli AE: MicroRNAs and their
targets: Recognition, regulation and an emerging reciprocal
relationship. Nat Rev Genet. 13:271–282. 2012.PubMed/NCBI View
Article : Google Scholar
|
|
20
|
Chou CH, Chang NW, Shrestha S, Hsu SD, Lin
YL, Lee WH, Yang CD, Hong HC, Wei TY, Tu SJ, et al: miRTarBase
2016: Updates to the experimentally validated miRNA-target
interactions database. Nucleic Acids Res. 44(D1):D239–D247.
2016.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Hansen TB, Jensen TI, Clausen BH, Bramsen
JB, Finsen B, Damgaard CK and Kjems J: Natural RNA circles function
as efficient microRNA sponges. Nature. 495:384–388. 2013.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Chen Y, Li C, Tan C and Liu X: Circular
RNAs: A new frontier in the study of human diseases. J Med Genet.
53:359–365. 2016.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Shao Y and Chen Y: Roles of circular RNAs
in neurologic disease. Front Mol Neurosci. 9(25)2016.PubMed/NCBI View Article : Google Scholar
|
|
24
|
You X, Vlatkovic I, Babic A, Will T,
Epstein I, Tushev G, Akbalik G, Wang M, Glock C, Quedenau C, et al:
Neural circular RNAs are derived from synaptic genes and regulated
by development and plasticity. Nat Neurosci. 18:603–610.
2015.PubMed/NCBI View
Article : Google Scholar
|
|
25
|
Chen S, Li T, Zhao Q, Xiao B and Guo J:
Using circular RNA hsa_circ_0000190 as a new biomarker in the
diagnosis of gastric cancer. Clin Chim Acta. 466:167–171.
2017.PubMed/NCBI View Article : Google Scholar
|
|
26
|
Wang K, Long B, Liu F, Wang JX, Liu CY,
Zhao B, Zhou LY, Sun T, Wang M, Yu T, et al: A circular RNA
protects the heart from pathological hypertrophy and heart failure
by targeting miR-223. Eur Heart J. 37:2602–2611. 2016.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Cui X, Niu W, Kong L, He M, Jiang K, Chen
S, Zhong A, Li W, Lu J and Zhang L: hsa_circRNA_103636: Potential
novel diagnostic and therapeutic biomarker in Major depressive
disorder. Biomark Med. 10:943–952. 2016.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Burd CE, Jeck WR, Liu Y, Sanoff HK, Wang Z
and Sharpless NE: Expression of linear and novel circular forms of
an INK4/ARF-associated non-coding RNA correlates with
atherosclerosis risk. PLoS Genet. 6(e1001233)2010.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Bang OY, Fujimura M and Kim SK: The
pathophysiology of moyamoya disease: An update. J Stroke. 18:12–20.
2016.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Kamada F, Aoki Y, Narisawa A, Abe Y,
Komatsuzaki S, Kikuchi A, Kanno J, Niihori T, Ono M, Ishii N, et
al: A genome-wide association study identifies RNF213 as the first
Moyamoya disease gene. J Hum Genet. 56:34–40. 2011.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Kang HS, Kim JH, Phi JH, Kim YY, Kim JE,
Wang KC, Cho BK and Kim SK: Plasma matrix metalloproteinases,
cytokines and angiogenic factors in moyamoya disease. J Neurol
Neurosurg Psychiatry. 81:673–678. 2010.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Rafat N, Beck GCh, Peña-Tapia PG,
Schmiedek P and Vajkoczy P: Increased levels of circulating
endothelial progenitor cells in patients with Moyamoya disease.
Stroke. 40:432–438. 2009.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Bedini G, Blecharz KG, Nava S, Vajkoczy P,
Alessandri G, Ranieri M, Acerbi F, Ferroli P, Riva D, Esposito S,
et al: Vasculogenic and angiogenic pathways in moyamoya disease.
Curr Med Chem. 23:315–345. 2016.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Nanba R, Kuroda S, Ishikawa T, Houkin K
and Iwasaki Y: Increased expression of hepatocyte growth factor in
cerebrospinal fluid and intracranial artery in moyamoya disease.
Stroke. 35:2837–2842. 2004.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Kim SK, Yoo JI, Cho BK, Hong SJ, Kim YK,
Moon JA, Kim JH, Chung YN and Wang KC: Elevation of CRABP-I in the
cerebrospinal fluid of patients with Moyamoya disease. Stroke.
34:2835–2841. 2003.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Frank PG, Woodman SE, Park DS and Lisanti
MP: Caveolin, caveolae, and endothelial cell function. Arterioscler
Thromb Vasc Biol. 23:1161–1168. 2003.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297.
2004.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Dai D, Lu Q, Huang Q, Yang P, Hong B, Xu
Y, Zhao W, Liu J and Li Q: Serum miRNA signature in Moyamoya
disease. PLoS One. 9(e102382)2014.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhao S, Gong Z, Zhang J, Xu X, Liu P, Guan
W, Jing L, Peng T, Teng J and Jia Y: Elevated serum microRNA Let-7c
in Moyamoya disease. J Stroke Cerebrovasc Dis. 24:1709–1714.
2015.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Hallemeier CL, Rich KM, Grubb RL Jr,
Chicoine MR, Moran CJ, Cross DT III, Zipfel GJ, Dacey RG Jr and
Derdeyn CP: Clinical features and outcome in north american adults
with Moyamoya phenomenon. Stroke. 37:1490–1496. 2006.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Kraemer M, Heienbrok W and Berlit P:
Moyamoya disease in Europeans. Stroke. 39:3193–3200.
2008.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Duan L, Bao XY, Yang WZ, Shi WC, Li DS,
Zhang ZS, Zong R, Han C, Zhao F and Feng J: Moyamoya disease in
China: Its clinical features and outcomes. Stroke. 43:56–60.
2012.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Kikuta K, Takagi Y, Nozaki K, Sawamoto N,
Fukuyama H and Hashimoto N: The presence of multiple microbleeds as
a predictor of subsequent cerebral hemorrhage in patients with
moyamoya disease. Neurosurgery. 62:104–11; discussion 111-2.
2008.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Morioka M, Hamada JI, Kawano T, Todaka T,
Yano S, Kai Y and Ushio Y: Angiographic dilatation and branch
extension of the anterior choroidal and posterior communicating
arteries are predictors of hemorrhage in adult Moyamoya patients.
Stroke. 34:90–95. 2003.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Matsushige T, Kraemer M, Schlamann M,
Berlit P, Forsting M, Ladd ME, Sure U and Wrede KH: Ventricular
microaneurysms in Moyamoya angiopathy visualized with 7T MR
angiography. AJNR Am J Neuroradiol. 37:1669–1672. 2016.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Li H, Yue H, Hao Y, Li H, Wang S, Yu L,
Zhang D, Cao Y and Zhao J: Expression profile of long noncoding
RNAs in human cerebral aneurysms: A microarray analysis. J
Neurosurg. 127:1055–1062. 2017.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Zhao M, Zhang D, Wang S, Zhang Y, Deng X
and Zhao J: The collateral circulation in moyamoya disease: A
single-center experience in 140 pediatric patients. Pediatr Neurol.
77:78–83. 2017.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Zhang Y, Chen B, Ming L, Qin H, Zheng L,
Yue Z, Cheng Z, Wang Y, Zhang D, Liu C, et al: MicroRNA-141
inhibits vascular smooth muscle cell proliferation through
targeting PAPP-A. Int J Clin Exp Pathol. 8:14401–14408.
2015.PubMed/NCBI
|
|
49
|
Jones JA, Spinale FG and Ikonomidis JS:
Transforming growth factor-beta signaling in thoracic aortic
aneurysm development: A paradox in pathogenesis. J Vasc Res.
46:119–137. 2009.PubMed/NCBI View Article : Google Scholar
|
|
50
|
Sappino AP, Madani R, Huarte J, Belin D,
Kiss JZ, Wohlwend A and Vassalli JD: Extracellular proteolysis in
the adult murine brain. J Clin Invest. 92:679–685. 1993.PubMed/NCBI View Article : Google Scholar
|
|
51
|
Shah PK, Falk E, Badimon JJ,
Fernandez-Ortiz A, Mailhac A, Villareal-Levy G, Fallon JT,
Regnstrom J and Fuster V: Human monocyte-derived macrophages induce
collagen breakdown in fibrous caps of atherosclerotic plaques.
Potential role of matrix-degrading metalloproteinases and
implications for plaque rupture. Circulation. 92:1565–1569.
1995.PubMed/NCBI
|
|
52
|
Lee JM, Yin KJ, Hsin I, Chen S, Fryer JD,
Holtzman DM, Hsu CY and Xu J: Matrix metalloproteinase-9 and
spontaneous hemorrhage in an animal model of cerebral amyloid
angiopathy. Ann Neurol. 54:379–382. 2003.PubMed/NCBI View Article : Google Scholar
|
|
53
|
McMillan WD, Tamarina NA, Cipollone M,
Johnson DA, Parker MA and Pearce WH: Size Matters: The relationship
between MMP-9 expression and aortic diameter. Circulation.
96:2228–2232. 1997.PubMed/NCBI View Article : Google Scholar
|
|
54
|
Kavsak P, Rasmussen RK, Causing CG, Bonni
S, Zhu H, Thomsen GH and Wrana JL: Smad7 binds to Smurf2 to form an
E3 ubiquitin ligase that targets the TGF beta receptor for
degradation. Mol Cell. 6:1365–1375. 2000.PubMed/NCBI View Article : Google Scholar
|
|
55
|
Sang QX: Complex role of matrix
metalloproteinases in angiogenesis. Cell Res. 8:171–177.
1998.PubMed/NCBI View Article : Google Scholar
|