Introduction
Cholangiocarcinoma (CCA) is a highly aggressive
cancer with a poor prognosis, especially in patients diagnosed at
an advanced or unresectable stage. Chemotherapy is a potential
option for the treatment of patients diagnosed with unresectable
CCA. Gemcitabine is widely used as a first-line treatment for
metastatic or unresectable CCA. However, clinical outcomes of
patients with CCA treated with gemcitabine remain unsatisfactory
due to the development of chemoresistance (1). Gemcitabine resistance is a complex
molecular alteration across multiple cellular processes, such as
alterations in drug uptake, metabolism, DNA repair mechanisms, and
evasion of apoptosis (2).
Therefore, understanding the molecular basis of resistance is
critical for improving therapeutic strategies in patients with
cancer. However, the molecular mechanisms underlying gemcitabine
resistance in CCA remain unclear.
At present, advances in transcriptomics and
bioinformatics have allowed researchers to explore large-scale
public datasets, especially the Gene Expression Omnibus (GEO)
(3), to uncover molecular
signatures associated with drug resistance in various cancer types.
These tools have been successfully applied to identify potential
biomarkers and therapeutic targets in pancreatic (4), bladder (5), lung (6) and bile duct cancers (7). Molecular docking is a computational
technique that predicts the preferred orientation and binding
affinity of a small molecule when bound to a target macromolecule.
This technique plays a pivotal role in the identification of small
molecules that may reverse chemoresistance in cancer cells
(8,9).
To the best of our knowledge, no comprehensive
analysis has been reported that integrates public transcriptomic
datasets from gemcitabine-resistant models to identify common
differentially expressed genes (DEGs) relevant to CCA. Therefore,
the objective of the present study was to identify potential
molecular targets associated with gemcitabine resistance in CCA
using an integrated approach, combining public transcriptomic
dataset analysis, gene expression validation in CCA cell lines, and
molecular docking to predict potential inhibitors against candidate
genes associated with gemcitabine resistance.
Materials and methods
CCA cell lines
KKU-213A (10) and
KKU-100(11) were obtained from the
Japanese Collection of Research Bioresources Cell Bank, Osaka,
Japan. Stable gemcitabine-resistant CCA cell lines (CCA-GemR) were
developed from the parental cell lines KKU-213A and KKU-100 by
continuous exposure to gradually increasing doses of gemcitabine,
as previously described (12).
Parental CCA cell lines were cultured in complete Dulbecco's
modified Eagle's medium (DMEM) containing 5 mM glucose,
supplemented with 10% fetal bovine serum, 100 U/ml penicillin, and
100 µg/ml streptomycin (all from Gibco/BRL; Thermo Fisher
Scientific, Inc.) and maintained at 37˚C in a humidified incubator
with 5% CO2. Gemcitabine-resistant CCA cell lines were
maintained under the same conditions but were additionally treated
with the IC10 concentration of gemcitabine.
The present study was approved (approval no. HE661419) by the Khon
Kaen University Ethics Committee for Human Research (Khon Kaen,
Thailand).
Determination of DEGs in stable
gemcitabine-resistant cell lines and patients with CCA
To identify common molecules related to gemcitabine
resistance in cancer cell lines, transcriptomic datasets from
stable gemcitabine-resistant cancer cell lines were retrieved from
the GEO database (https://www.ncbi.nlm.nih.gov/geo/), including
GSE116118 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE116118)
(13), GSE208659 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE208659)
(14), and GSE140077 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE140077)
(15). In addition, a
transcriptomic dataset (GSE76297, https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE76297)
(16) from 92 normal and 91 Thai
tissues from patients with CCA was also used to determine the
expression levels of candidate genes in patients with CCA. DEGs
were investigated using GEO2R (www.ncbi.nlm.nih.gov/geo/geo2r) and Galaxy (https://usegalaxy.org/) (17). DEGs were determined based on the
criteria of an adjusted P-value <0.05 and log2|FC| >0.5. Data
visualizations, including volcano plots and Venn diagrams, were
generated using RStudio version 2024.12 (https://posit.co/) and jvenn (https://jvenn.toulouse.inrae.fr/app/index.html)
(18), respectively.
Identification of Rhotekin 2
(RTKN2)-correlated genes and pathway enrichment analysis
Genes correlated with RTKN2 expression in patients
with CCA were identified using the cBioPortal platform (https://www.cbioportal.org) (19). The TCGA-CHOL dataset was selected,
and Spearman correlation analysis was performed to determine the
genes that were positively and negatively correlated with RTKN2
expression. To investigate the biological significance of
RTKN2-correlated genes, pathway enrichment analysis was performed
separately for positively and negatively correlated genes using
Enrichr (https://maayanlab.cloud/Enrichr/). Enrichment results
were ranked by adjusted P-values using the Benjamini-Hochberg
correction, and the most significantly enriched pathways (adjusted
P<0.05) were reported and visualized.
Reverse trascription-quantitative
polymerase chain reaction (RT-qPCR)
Total RNA was extracted from CCA cell lines using
TRIzol reagent (Invitrogen; Thermo Fisher Scientific, Inc.). The
RNA concentration was measured using NanoDrop™ 2000
Spectrophotometer (Thermo Fisher Scientific, Inc.). The extracted
RNA from CCA cell lines was converted to cDNA using High-capacity
cDNA Reverse Transcription Kit (cat. no. 4368814, Applied
Biosystem™; Thermo Fisher Scientific, Inc.). Using
real-time PCR, the mRNA expression levels of candidate genes
associated with gemcitabine resistance were determined in CCA cell
lines. Real-time PCR was performed using LightCycle 480 real-time
PCR system (Roach Diagnostics). Each PCR condition contained 2X
LightCycle 480 SYBR Green I Master (Roach Diagnostics), 5 mM of
primers, and 40 ng of cDNA. The amplification was initiated by
incubating at 95˚C for 5 min, followed by 40 cycles at 95˚C for 20
sec, 65˚C (RAB1B, JAG1, PACS1 and ANKS6) and 63˚C (SF3B4 and RTKN2)
for 10 sec, and 72˚C for 20 sec. Mean and standard deviation values
of the cycle of quantification threshold (Cq) and melting
temperature were calculated. The expression of each gene was
normalized with β-actin. The gene expression levels were determined
using the 2-ΔΔCq method (20). All PCR primers and expected PCR
product sizes are shown in Table
I.
 | Table IThe six common upregulated gene
primers. |
Table I
The six common upregulated gene
primers.
| Gene | Forward sequence
5'→3' | Reverse sequence
5'→3' | Product size
(bp) |
|---|
| RAB1B |
GGAATCCTACGCCAACGTGA |
GTGGTGAGGTCGCTCTTGTT | 102 |
| JAG1 |
CTACAACCGTGCCAGTGACT |
CCTTCAGGTGTGTCGTTGGA | 150 |
| PACS1 |
CCCACCACCATACATGCTGT |
AGAACAGAGCAGGGAAAGGC | 108 |
| SF3B4 |
GAACGACTTCTGGCAGCTCA |
CACAGGATTGGGAGCAGAGG | 99 |
| ANKS6 |
CTCCTGCTCACGTCCTCTTG |
TGCTGGGAAGCCACTATGTG | 98 |
| RTKN2 |
AGCCAATGGAAGCACTGTTG |
ACAAAGGTGGTTTCCGTGGT | 70 |
| ACTB |
TCGTGCGTGACATTAAGGAG |
GAAGGAAGGCTGGAAGAGT | 176 |
To evaluate the effect of gemcitabine on RTKN2
expression, parental CCA cell lines (KKU-213A and KKU-100) were
plated in 10-cm culture dishes and incubated for 24 h. The cells
were then treated with gemcitabine at concentrations corresponding
to IC0 and IC50 values. After 48 h of
incubation, total RNA was extracted and RT-qPCR was performed as
aforementioned.
Molecular docking
The amino acid sequence of human RTKN2 was retrieved
from the Uniprot website (UniProt ID: Q8IZC4; https://www.uniprot.org/), and utilized for
constructing the 3D structure using AlphaFold3(21). The protonation states of ionizable
amino acid residues of the constructed RTKN2 were assigned at pH
7.4 using the PDB2PQR web server version 3.6.1 (https://server.poissonboltzmann.org/pdb2pqr) (22,23).
The protein structure was then minimized using molecular dynamics
(MD) simulations for 100 ns using the AMBER24 software (https://ambermd.org/). The AMBER ff19SB force field
was applied to the protein (24).
The TIP3P water model (25) was
used to solvate the system. MD simulations were performed under
periodic boundary conditions in the isothermal-isobaric
(NPT) ensemble at 310 K and 1 atm.
The last MD snapshot of the RTKN2 structure
(Fig. S1) was subsequently
analyzed for the potential ligand-binding sites using the
CavityPlus 2022 web server (http://pkumdl.cn:8000/cavityplus) (26,27).
Sites 2, 5, and 9, which exhibited strong druggability (Table SI), were then selected as
ligand-binding sites for molecular docking. The coordinates (x, y,
z) of these binding sites were as follows: Site 2 (94.957, 76.997,
99.134), site 5 (113.028, 76.794, 108.707), and site 9 (45.114,
90.615, 54.979). For virtual screening, compounds (excluding those
with a molecular weight >1,000 g/mol, glycans, peptides, and
lipids) from the drug repurposing compound library plus (https://www.medchemexpress.com/screening-libraries.html,
accessed on 1st March 2025) were docked into each of the three
binding sites using AutoDock Vina version 1.2.6 (28,29).
The top three compounds exhibiting the highest binding affinities
(lowest binding energy) at each site were selected as potential
hits.
Statistical analysis
All RT-qPCR data are presented from three
independent biological experiments (triplicate in each experiment).
Statistical analysis was performed using SPSS statistics 24.0
software (IBM Corp.) and GraphPad Prism 9.2 software (Dotmatics).
To determine whether the observed expression showed a significant
difference between the two groups (resistance vs. parental), data
that demonstrated normality and homogeneity of variance were
compared using Student's t-test, while data with non-normal
distribution or unequal variance were analyzed using the
Mann-Whitney U test. For visualization, box-and-whisker plots
displaying the median and interquartile range (IQR) were used in
nonparametric cases, whereas for parametric analyses (t-test), the
mean value with 95% confidence intervals (CI) is presented.
P-values <0.05 were considered to indicate statistically
significant differences.
Results
Determination of common DEGs in stable
gemcitabine-resistant cancer cell lines
A total of three transcriptomic datasets of stable
gemcitabine-resistant cell lines and their corresponding parental
cell lines, including GSE116118, GSE208659, and GSE140077, were
retrieved from the GEO database. These datasets included cell lines
from CCA (MT-CHC01R1.5 gemcitabine resistant/MT-CHC01 parental),
gallbladder (NOZ-GemR gemcitabine resistant/NOZ parental), and
pancreatic cancers (BxPC-3-GR gemcitabine resistant/BxPC-3
parental). These cancers are commonly grouped as pancreatobiliary
cancers because they arise from foregut-derived pancreatobiliary
epithelium and share overlapping histopathological features,
clinical behavior, and poor prognosis. By integrating these three
datasets, the aim was to minimize bias arising from cell line
specificity and to identify reproducible DEGs consistently
associated with gemcitabine resistance.
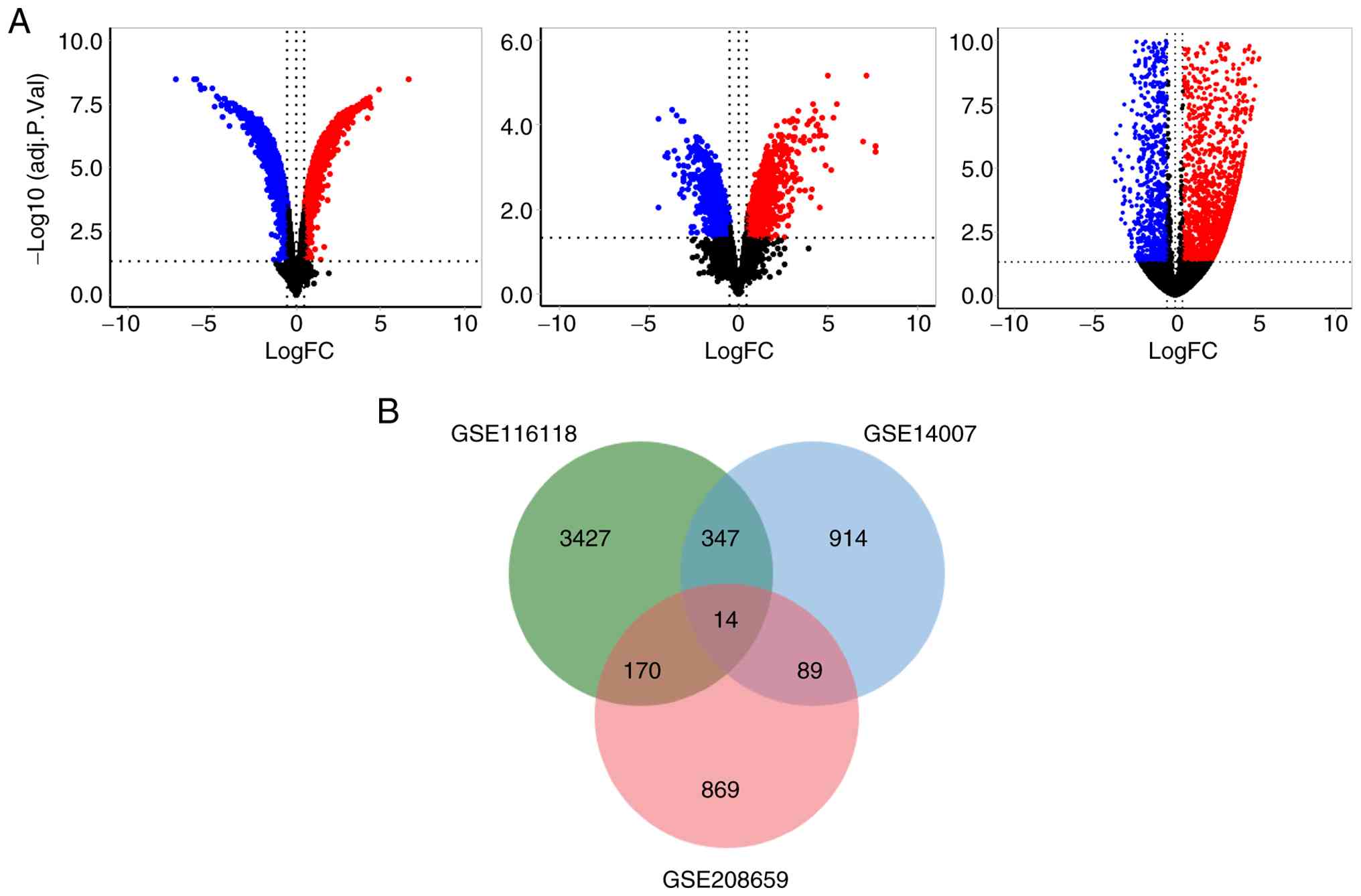
The analysis of DEGs between gemcitabine-resistant
cell lines and their corresponding parental cell lines identified a
total of 7,751, 2,730, and 2,206 DEGs in GSE116118, GSE208659, and
GSE140077, respectively. These results, including upregulated and
downregulated genes in the three stable gemcitabine-resistant
cancer cell lines, are shown in Table
II. Volcano plots in Fig. 1A
show the upregulated and downregulated DEGs. Venn diagrams were
used to determine common DEGs across the three transcriptomic
datasets from stable gemcitabine-resistant cancer cell lines. The
analysis identified 14 DEGs shared among the three stable
gemcitabine-resistant cancer cell lines (Fig. 1B). The 14 DEGs were RAB1B, JAG1,
PACS1, ANKS6, SF3B4, RTKN2, PGM2, DNAH11, NT5DC1, CDK14, IGFBP6,
RNFT1, RBBP9, and ZDHHC18. These commonly upregulated DEGs may be
associated with gemcitabine resistance in cancer cells.
 | Table IITranscriptomic datasets for stable
gemcitabine-resistant cancer cell lines. |
Table II
Transcriptomic datasets for stable
gemcitabine-resistant cancer cell lines.
| Cancer type | GSE no. | Total DEGs | Upregulated
DEGs | Downregulated
DEGs | Experiment
type |
|---|
| Bile duct | GSE116118 | 7,751 | 3,958 | 3,793 | Microarray |
| Gallbladder | GSE208659 | 2,703 | 1,142 | 1,561 | Microarray |
| Pancreas | GSE140077 | 2,206 | 1,364 | 842 | RNA-seq |
Validation of genes associated with
gemcitabine resistance in CCA
To prioritize genes potentially relevant to CCA, the
expression levels of the 14 upregulated DEGs in patients with CCA
were further validated using the transcriptomic dataset (GSE76297),
which includes 92 normal and 91 CCA tissues from Thai patients. The
results indicated that six of the 14 upregulated DEGs, including
RAB1B, JAG1, PACS1, ANKS6, SF3B4, and RTKN2, were significantly
upregulated in CCA tissues compared with adjacent normal tissues
(Figs. 2A and S1).
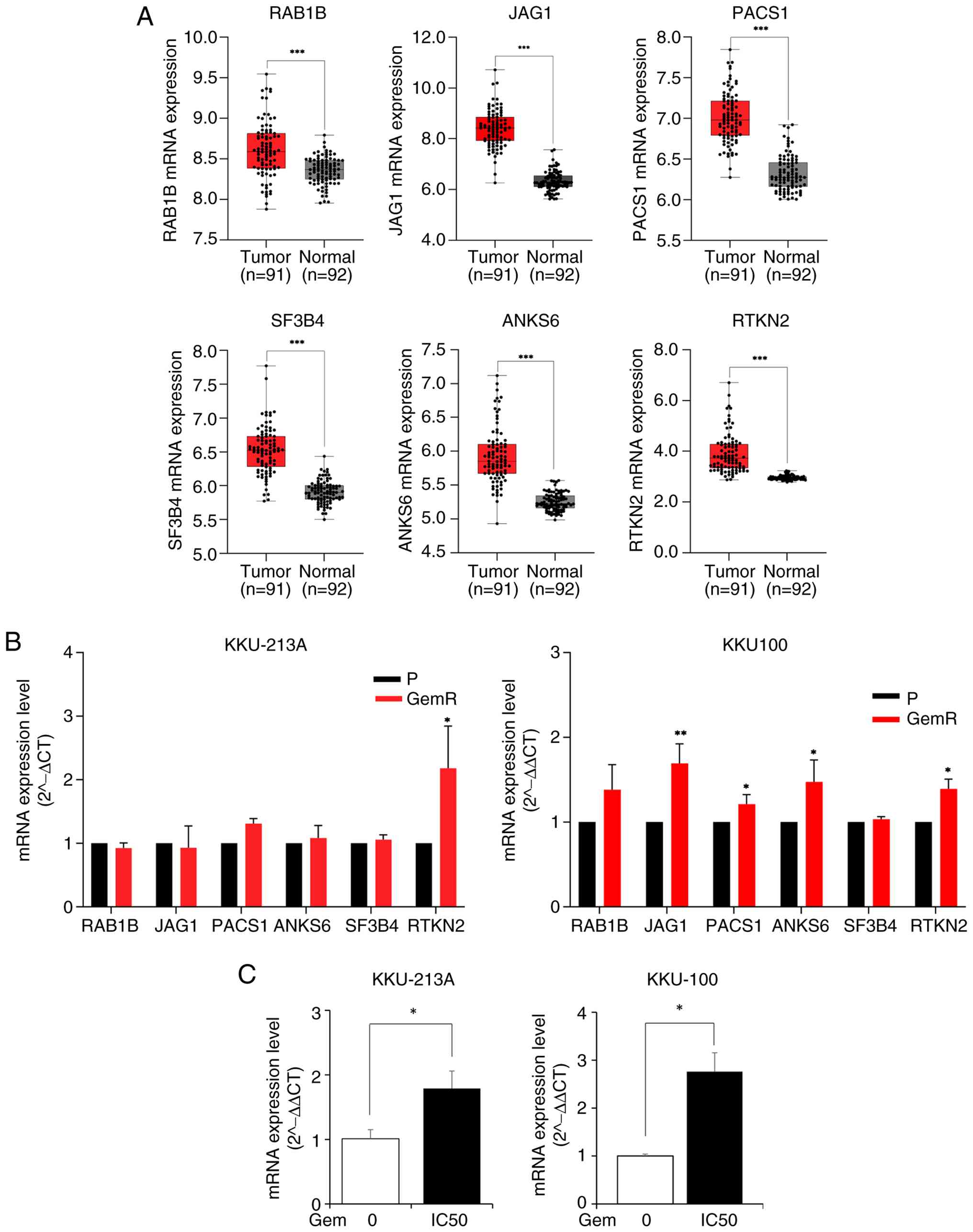 | Figure 2Validation of common upregulated
genes associated with gemcitabine resistance in CCA tissues and
cell lines. (A) Box plots show the mRNA expression levels of RAB1B,
JAG1, PACS1, ANKS6, SF3B4, and RTKN2 in CCA and normal tissues. The
Mann-Whitney U test was used to evaluate the expression differences
between CCA and normal tissue. (B) PCR analysis of six common
upregulated genes associated with gemcitabine resistance in stable
gemcitabine-resistant CCA cell lines. The data are expressed as the
mean ± SD from three independent experiments. (C) RTKN2 mRNA
expression after gemcitabine treatment in CCA cell lines. KKU-213A
and KKU-100 cells were treated with gemcitabine at IC25
and IC50 concentrations for 48 h. RTKN2 expression was
quantified by real-time PCR and normalized to β-actin (ACTB).
*P<0.05, **P<0.01 and
***P<0.001. CCA, cholangiocarcinoma; RTKN2, Rhotekin
2; P, parental; GemR, gemcitabine resistant. |
The expression levels of the six upregulated DEGs
associated with gemcitabine resistance were further validated in
two stable gemcitabine-resistant CCA cell lines using RT-qPCR. As
shown in Fig. 2B, RTKN2 was
significantly elevated in stable gemcitabine-resistant CCA cell
lines, namely KKU-213A-GemR and KKU-100-GemR, compared with their
parental CCA cell lines (KKU-213A and KKU-100). However, RAB1B,
JAG1, PACS1, ANKS6, and SF3B4 expression levels were not
significantly altered in the two stable gemcitabine-resistant CCA
cell lines.
To further verify the effect of gemcitabine
treatment on RTKN2 expression in CCA cells, parental CCA cell
lines, KKU-213A and KKU-100, were treated with gemcitabine at
IC50 concentrations for 48 h. Specifically, KKU-213A
cells were treated with 0 and 0.72 µM gemcitabine, while KKU-100
cells were treated with 0 and 0.102 µM gemcitabine. As shown in
Fig. 2C, RTKN2 mRNA expression was
significantly upregulated in cells treated with gemcitabine in both
cell lines. These findings revealed that RTKN2 is inducible by
gemcitabine exposure and may contribute to the adaptive resistance
mechanism in CCA cells.
RTKN2-related gene identification and
pathway enrichment analysis
Genes correlated with RTKN2 expression were
identified using the TCGA Cholangiocarcinoma dataset via the
cBioPortal platform. A total of 210 genes were found to be
significantly correlated with RTKN2 expression (q-value <0.05)
(Table SI). Pathway enrichment
analysis demonstrated that genes negatively associated with RTKN2
showed statistically significant enrichment in several metabolic
and mitochondrial pathways, such as ‘aerobic respiration and
respiratory electron transport’, ‘respiratory electron transport’,
‘TP53 regulates metabolic genes’, and ‘complex IV assembly’. These
findings suggest that high RTKN2 expression may be associated with
the downregulation of mitochondrial oxidative phosphorylation and
TP53-mediated metabolic control, implicating RTKN2 in metabolic
reprogramming in CCA. However, no statistically significant
pathways were identified in genes positively associated with RTKN2
(Table III).
 | Table IIIPathway enrichment analysis of
RTKN2-related genes in patients with CCA. |
Table III
Pathway enrichment analysis of
RTKN2-related genes in patients with CCA.
| A, Pathways
positively correlated to RTKN2 |
|---|
| Pathway | P-value | Adj P-value |
|---|
| Activation of HOX
genes during differentiation | 0.001 | 0.145 |
| Activation of
anterior HOX genes in hindbrain development during early
embryogenesis | 0.001 | 0.145 |
| B, Pathways
negatively correlated to RTKN2 |
| Aerobic respiration
and respiratory electron transport |
3.74x10-18 |
5.04x10-16 |
| Respiratory
electron transport |
4.80x10-16 |
3.24x10-14 |
| Metabolism |
5.45x10-10 |
2.45x10-8 |
| TP53 regulates
metabolic genes |
2.84x10-8 |
9.59x10-7 |
| Complex IV
assembly |
5.78x10-8 |
1.559x10-6 |
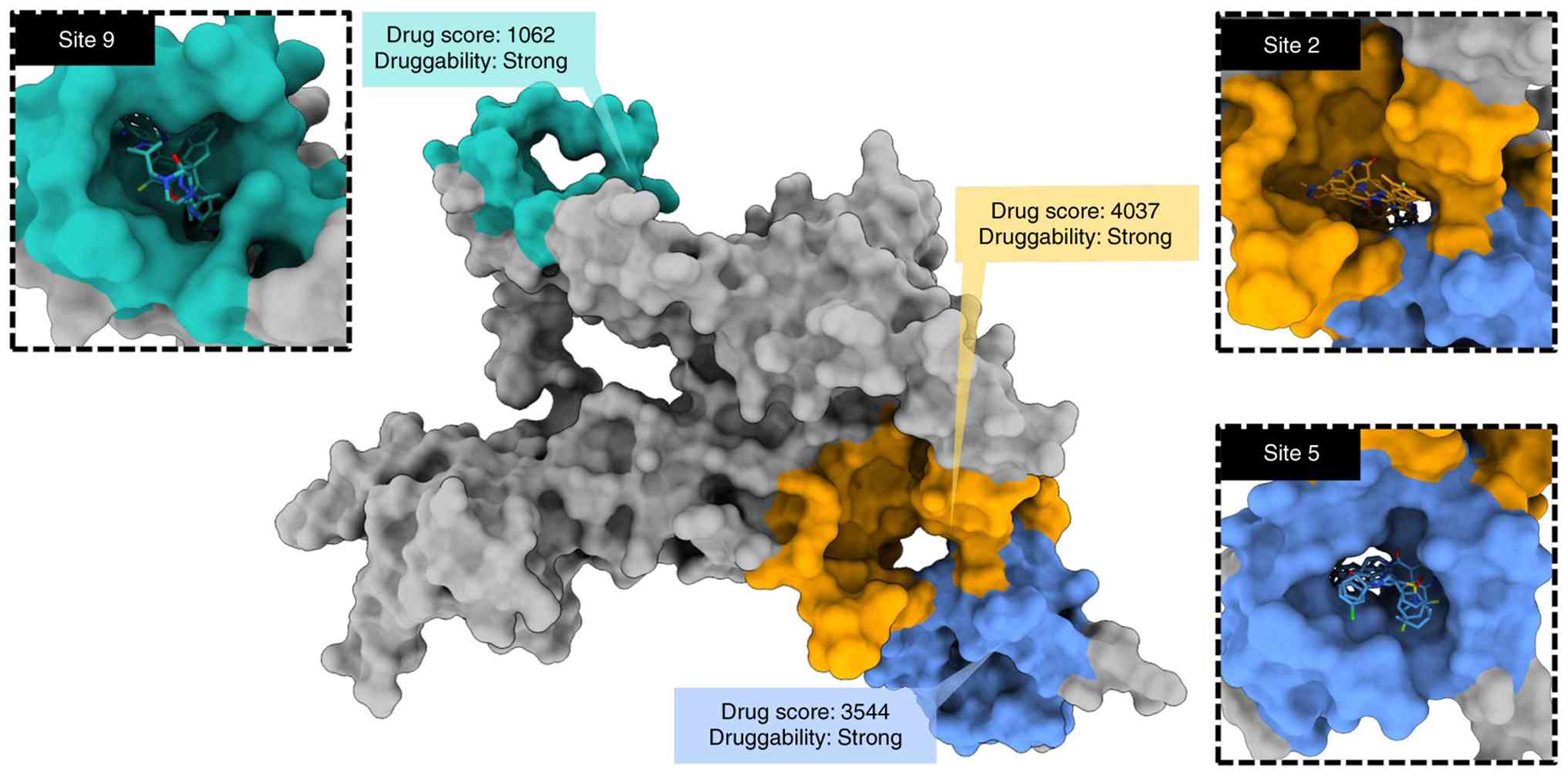
Molecular docking
The 3D structure of RTKN2 was constructed using
AlphaFold3 (Fig. S2). Due to the
lack of reported ligand-binding sites for RTKN2, the CavityPlus web
server was utilized to predict potential ligand-binding pockets on
the RTKN2 protein. The results showed that there were 31 possible
ligand-binding sites on RTKN2 (Table
SII). Among these, the binding pockets 2, 5, and 9 of RTKN2
exhibited strong druggability, with a drugscore of 4,037, 3,544,
and 1,062, respectively (Fig. 3).
Therefore, these three binding pockets of RTKN2 were selected for
further study.
To identify potential small-molecule inhibitors of
RTKN2, docking-based virtual screening from 5,396 compounds
available in the drug repurposing compound library plus was
performed on three possible binding sites of RTKN2 using AutoDock
Vina. As shown in Fig. 3 and
Table IV, the potential screened
inhibitors for the binding site 2 included MK-3207 (-10.49
kcal/mol), phthalocyanine (-11.40 kcal/mol), and orforglipron
(-11.47 kcal/mol). The best-scoring compounds for the binding
pocket 5 included NKP608 (-10.94 kcal/mol), tegatrabetan (-11.10
kcal/mol), and MK-3207 (-11.31 kcal/mol). The best-scoring
compounds for the binding pocket 9 included umbralisib (-10.62
kcal/mol), MK-3207 (-10.66 kcal/mol), and vepdegestrant (-10.91
kcal/mol). Notably, MK-3207 was found to be a potential inhibitor
at all three binding sites of RTKN2. Taken together, the docking
findings revealed the top three candidate compounds at each
ligand-binding site, with binding affinities ranging from -10.49 to
-11.47 kcal/mol, suggesting them as potential RTKN2 inhibitors.
 | Table IVBinding affinity of the top three
screened compounds at each binding site of RTKN2. |
Table IV
Binding affinity of the top three
screened compounds at each binding site of RTKN2.
| Compound name | Binging affinity
(kcal/mol) | Binding site |
|---|
| MK-3207 | -10.49 | 2 |
| Phthalocyanine | -11.40 | 2 |
| Orforglipron | -11.47 | 2 |
| NKP608 | -10.94 | 5 |
| Tegatrabetan | -11.10 | 5 |
| MK-3207 | -11.31 | 5 |
| Umbralisib | -10.62 | 9 |
| MK-3207 | -10.66 | 9 |
| Vepdegestrant | -10.91 | 9 |
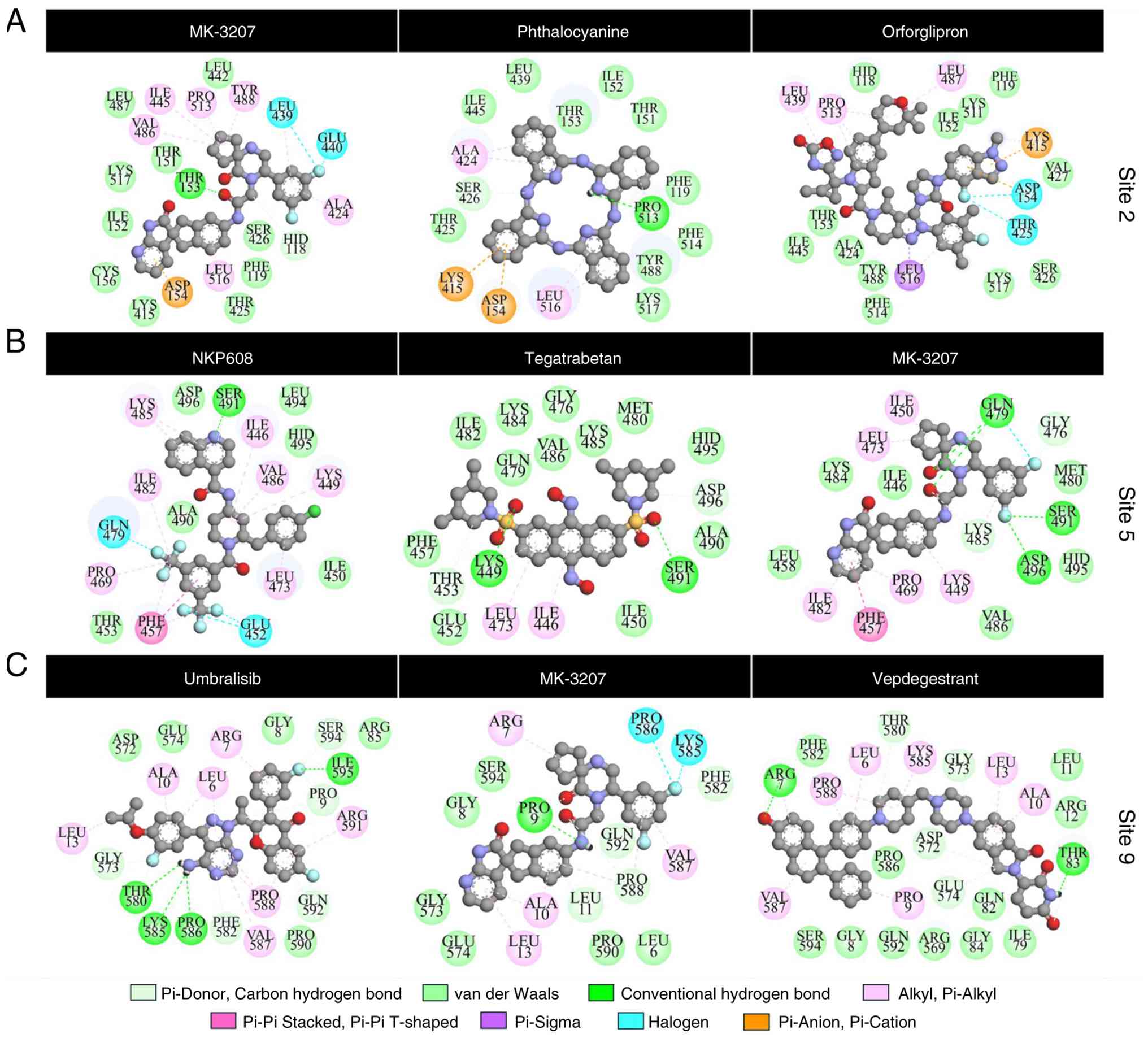
The top-ranking compounds exhibited multiple
favorable interactions with RTKN2, including hydrogen bonds,
halogen interactions, π interactions, electrostatic attractions,
and, in particular, van der Waals forces. Comprehensive 2D
interaction profiles for each ligand-binding site (Fig. 4) revealed specific interactions
between the functional groups of the ligand and key amino acid
residues of RTKN2. The Asp154 and Lys415 residues at the binding
site 2 of RTKN2 were found to electrostatically form π-anion and
π-cation interactions with the aromatic and alkyl moieties of
MK-3207, phthalocyanine, and orforglipron. In addition, the
fluorine atoms of MK-3207 and orforglipron formed halogen
interactions with i) Leu439 and Glu440 and ii) Asp154 and Thr425
residues, respectively. For ligand-binding site 5, instead of
forming halogen interactions, the fluorine atoms of MK-3207
potentially formed hydrogen bonds with Gln479, Ser491, and Asp496
residues. By contrast, the fluorine atoms of NKP608 formed halogen
forces with Glu452 and Gln479 residues. The Ile446, Lys449, Phe457,
Pro469, Leu473, and Ile482 residues were found to be hotspots for
the binding of NKP608, tegatrabetan, and MK-3207 via several π
interactions. In the case of binding site 9, the aromatic moieties
of all screened ligands were stabilized by alkyl/π-alkyl
interactions and van der Waals forces. Notably, the fluorine atoms
of MK-3207 formed halogen interactions with Lys585 and Pro586
residues. Moreover, this compound potentially formed a hydrogen
bond with Pro9 residue. The amino group and the fluorine atom of
umbralisib potentially formed hydrogen bonds with Thr580, Lys585,
Pro586, and Ile595 residues. Similarly, vepdegestrant formed two
hydrogen bonds with Arg7 and Thr83 residues. These results suggest
that the identified compounds hold promise as lead candidates for
the development of RTKN2-targeted inhibitors. By potentially
disrupting RTKN2 function, these compounds may offer a novel
therapeutic approach to overcome gemcitabine resistance in CCA.
Discussion
Gemcitabine resistance remains a major issue in the
treatment of unresectable CCA. Therefore, identification of
molecules associated with gemcitabine resistance in CCA is urgently
needed. In the present study, transcriptomic profiling of three
independent datasets from stable gemcitabine-resistant cancer cell
lines revealed thousands of DEGs. Notably, 14 genes were commonly
upregulated in all three cancer types. These shared DEGs possibly
represent core molecular mechanisms contributing to gemcitabine
resistance in cancer cells.
The expression of 14 commonly upregulated DEGs was
validated using transcriptomic data from patients with CCA. RTKN2,
RAB1B, JAG1, PACS1, ANKS6, and SF3B4 were significantly
overexpressed in CCA tissues compared with adjacent normal tissues.
RTKN2 emerged as a promising candidate for further investigation.
RTKN2 was the only gene that remained significantly upregulated in
two stable gemcitabine-resistant CCA cell lines (KKU-213A-GemR and
KKU-100-GemR), suggesting a potential role in mediating gemcitabine
resistance in CCA.
RTKN2 is a Rho-GTPase effector protein. This protein
plays a significant oncogenic role in various cancers by promoting
tumor cell proliferation, invasion, metastasis, and resistance to
apoptosis. Overexpression of RTKN2 has been reported in several
cancers, such as hepatocellular carcinoma (30), colon cancer (31), osteosarcoma (32), and non-small cell lung cancer
(33). High RTKN2 expression has
been associated with unfavorable clinical outcomes of patients with
cancer, including advanced TNM stage, lymphatic metastasis, and
shorter overall survival (33).
Knockdown of RTKN2 was shown to suppress proliferation and induce
apoptosis in cancer cells. Several studies have highlighted RTKN2
as a key regulator that may contribute to chemoresistance via
multiple molecular mechanisms. RTKN2 could promote cell survival by
activating the NF-κB signaling pathway, which leads to the
upregulation of anti-apoptotic genes such as BCL-2 and suppression
of pro-apoptotic factors such as Bax (33-35).
Silencing RTKN2 was demonstrated to reduce β-catenin
phosphorylation and nuclear localization, suppressing Wnt/β-catenin
signaling (35). Activation of
Wnt/β-catenin signaling can enhance the expression of cell
survival, and drug efflux associated genes, which contribute to
resistance against chemotherapy. Wnt/β-catenin signaling
contributes to chemoresistance in CCA by upregulating drug efflux
pumps such as P-glycoprotein (36).
To explore the functional relevance of RTKN2 in CCA,
a correlation analysis based on gene expression was performed using
transcriptomic data from the TCGA. Genes negatively correlated with
RTKN2 expression were significantly enriched in mitochondrial and
metabolic pathways, including aerobic respiration, respiratory
electron transport, and TP53-regulated metabolic processes. These
results indicated that high RTKN2 expression may potentially
contribute to metabolic reprogramming in gemcitabine-resistant
cancer cells. This finding is consistent with previous studies
showing that resistant cancer cells often exhibit a metabolic shift
from OXPHOS to glycolysis (Warburg effect), which supports survival
under stress and enhances chemotherapeutic drug resistance
(37,38).
Molecular docking was employed to screen for
potential small-molecule inhibitors of RTKN2. Compounds such as
NKP608, tegatrabetan, umbralisib, vepdegestrant, and MK-3207
exhibited strong binding affinities to RTKN2 binding pockets.
Several studies have demonstrated the antitumor activity of these
compounds in pre-clinical and clinical studies. NKP608 is a
selective antagonist of the neurokinin-1 (NK-1) receptor. NKP608
has been shown to inhibit proliferation, migration, and invasion of
human colorectal cancer cells by suppressing the Wnt/β-catenin
signaling pathway (39).
Tegatrabetan is a small molecule drug that targets cancer through
multiple mechanisms, primarily by disrupting the Wnt/β-catenin
signaling pathway. Tegatrabetan binds to the N-terminal domain of
the scaffold protein transducin β-like 1, disrupting its
interaction with β-catenin (40).
This dislocation reduces nuclear β-catenin levels and inhibits
transcription of genes that promote cancer progression.
Tegatrabetan has demonstrated anticancer activity in preclinical
studies, including osteosarcoma (41), and multiple myeloma (42). Umbralisib is a selective inhibitor
targeting the delta isoform of phosphoinositide 3-kinase and casein
kinase-1ε, thereby inhibiting the PI3K/AKT signaling pathway
involved in tumor cell proliferation and survival (43). Umbralisib was approved by the FDA in
2021 for relapsed or refractory marginal zone lymphoma and
follicular lymphoma (44).
Vepdegestrant is a novel oral compound that leverages PROteolysis
TArgeting Chimera (PROTAC) technology to target the estrogen
receptor. This compound has shown promise in both monotherapy and
combination therapy in several cancers. Vepdegestrant has
demonstrated substantial tumor growth inhibition in MCF7 orthotopic
xenograft models (45) and has been
evaluated in several clinical trials (46-49).
In the present study, MK-3207 demonstrated the greatest potential
as an RTKN2 inhibitor, as it was able to bind to three druggable
binding sites on the RTKN2 protein. MK-3207 is a highly selective
antagonist of the calcitonin gene-related peptide receptor. MK-3207
showed efficacy and tolerability in a clinical trial involving
patients with migraines (50).
However, there is currently no clear evidence demonstrating the
antitumor activity of MK-3207.
These findings suggest that compounds with
established clinical activity or anticancer mechanisms may exert
additional therapeutic benefits through RTKN2 inhibition. Their
strong binding affinities and mechanistic relevance highlight the
potential for repurposing or further development as RTKN2-targeted
therapies to overcome gemcitabine resistance in CCA.
Taken together, the findings of the present study
suggest that RTKN2 may play a key role in gemcitabine resistance in
CCA, potentially through metabolic reprogramming involving
suppression of mitochondrial activity and TP53-related pathways.
Molecular docking supports the druggability of RTKN2. Several
compounds, including NKP608, tegatrabetan, umbralisib,
vepdegestrant, and MK-3207, demonstrate high predicted binding
affinities and favorable interactions with RTKN2 binding pockets.
Targeting RTKN2 using repurposed inhibitors identified via virtual
screening may represent a novel therapeutic strategy to restore
gemcitabine sensitivity in CCA.
A limitation of the present study is that the
functional effects of RTKN2 and MK-3207 on gemcitabine-resistant
CCA cell lines were not experimentally validated. In future
studies, the effects of MK-3207 treatment in both parental and
gemcitabine-resistant CCA cell lines will be evaluated, and RTKN2
knockdown experiments using siRNA will be performed to verify its
functional role in gemcitabine resistance. These investigations
will be essential to confirm the mechanistic role of RTKN2 and to
assess the therapeutic potential of MK-3207 as a candidate
inhibitor.
Supplementary Material
mRNA expression levels of PGM2,
DNAH11, NT5DC1, CDK14, IGFBP6, RNFT1, RBBP9, and ZDHHC18 in
cholangiocarcinoma and normal tissues. *P<0.05,
**P<0.01 and ***P<0.001.
Superimposed structures of RTKN2
constructed from AlphaFold3 (blue) and MD simulation (final
snapshot, orange). RTKN2, Rhotekin 2.
Genes correlated with RTKN2 in
cholangiocarcinoma.
Outputs of the CAVITY module
(CavityPlus 2022 web server)using the MD-minimized RTKN2
structure.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by a grant from the
Fundamental Fund of Khon Kaen University, under the National
Science, Research and Innovation Fund (NSRF), Thailand (grant no.
203258).
Availability of data and materials
The data generated in the present study may be
requested from the corresponding author.
Authors' contributions
SK and WS conceived and designed the study. SK, PM,
PMI, US, and WS acquired the data and performed the investigation.
SK, PM, PMI, and US analyzed and interpreted the data and prepared
the figures. WLI, RD, SW, and WS contributed to the interpretation
of the results and critically revised the manuscript for important
intellectual content. SK drafted the original manuscript. WL, RD,
SW, and WS reviewed and edited the manuscript. WS supervised the
study, acquired funding, and administered the project. SK and WS
confirm the authenticity of all the raw data. All authors read and
approved the final manuscript.
Ethics approval and consent to
participate
The present study was approved (approval no.
HE661419) by the Khon Kaen University Ethics Committee for Human
Research (Khon Kaen, Thailand).
Patient consent for publication
Not applicable.
Competing interests
The authors have declared that no competing interest
exists.
Use of artificial intelligence tools
During the preparation of this work, artificial
intelligence tools were used to improve the readability and
language of the manuscript, and subsequently, the authors revised
and edited the content produced by the artificial intelligence
tools as necessary, taking full responsibility for the ultimate
content of the present manuscript.
References
|
1
|
Ishimoto U, Kondo S, Ohba A, Sasaki M,
Sakamoto Y, Morizane C, Ueno H and Okusaka T: Prognostic factors
for survival in patients with advanced intrahepatic
cholangiocarcinoma treated with gemcitabine plus cisplatin as
first-line treatment. Oncology. 94:72–78. 2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Jia Y and Xie J: Promising molecular
mechanisms responsible for gemcitabine resistance in cancer. Genes
Dis. 2:299–306. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Clough E and Barrett T: The gene
expression omnibus database. Methods Mol Biol. 1418:93–110.
2016.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Xiao X, Wan Z, Liu X, Chen H, Zhao X, Ding
R, Cao Y, Zhou F, Qiu E, Liang W, et al: Screening of therapeutic
targets for pancreatic cancer by bioinformatics methods. Horm Metab
Res. 55:420–425. 2023.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Shen P, He X, Lan L, Hong Y and Lin M:
Identification of cell division cycle 20 as a candidate biomarker
and potential therapeutic target in bladder cancer using
bioinformatics analysis. Biosci Rep. 40(BSR20194429)2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Liu X, Jia Y, Shi C, Kong D, Wu Y, Zhang
T, Wei A and Wang D: CYP4B1 is a prognostic biomarker and potential
therapeutic target in lung adenocarcinoma. PLoS One.
16(e0247020)2021.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Sungwan P, Lert-Itthiporn W, Silsirivanit
A, Klinhom-On N, Okada S, Wongkham S and Seubwai W: Bioinformatics
analysis identified CDC20 as a potential drug target for
cholangiocarcinoma. PeerJ. 9(e11067)2021.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Xiao C, Yin X, Xi R, Yuan C and Ou Y:
Molecular mechanisms of reversal of multidrug resistance in breast
cancer by inhibition of P-gp by cytisine N-isoflavones derivatives
explored through network pharmacology, molecular docking, and
molecular dynamics. Int J Mol Sci. 26(3813)2025.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Zhang N, Tian X, Liu F, Jin X, Zhang J,
Hao L, Jiang S and Liu Q: Reversal of sorafenib resistance in
hepatocellular carcinoma by curcumol: Insights from network
pharmacology, molecular docking, and experimental validation. Front
Pharmacol. 16(1514997)2025.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Sripa B, Seubwai W, Vaeteewoottacharn K,
Sawanyawisuth K, Silsirivanit A, Kaewkong W, Muisuk K, Dana P,
Phoomak C, Lert-Itthiporn W, et al: Functional and genetic
characterization of three cell lines derived from a single tumor of
an Opisthorchis viverrini-associated cholangiocarcinoma patient.
Hum Cell. 33:695–708. 2020.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Sripa B, Leungwattanawanit S, Nitta T,
Wongkham C, Bhudhisawasdi V, Puapairoj A, Sripa C and Miwa M:
Establishment and characterization of an opisthorchiasis-associated
cholangiocarcinoma cell line (KKU-100). World J Gastroenterol.
11:3392–3397. 2005.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Kidoikhammouan S, Lert-Itthiporn W,
Deenonpoe R, Saengboonmee C, Obchoei S, Wongkham S and Seubwai W:
Targeting EGFR activation to overcome gemcitabine resistance in
cholangiocarcinoma. Anticancer Res. 44:5393–5404. 2024.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Varamo C, Peraldo-Neia C, Ostano P,
Basiricò M, Raggi C, Bernabei P, Venesio T, Berrino E, Aglietta M,
Leone F and Cavalloni G: Establishment and characterization of a
new intrahepatic cholangiocarcinoma cell line resistant to
gemcitabine. Cancers (Basel). 11(519)2019.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Vergara-Gómez L, Bizama C, Zhong J,
Buchegger K, Suárez F, Rosa L, Ili C, Weber H, Obreque J, Espinoza
K, et al: A novel gemcitabine-resistant gallbladder cancer model
provides insights into molecular changes occurring during acquired
resistance. Int J Mol Sci. 24(7238)2023.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Zhou J, Zhang L, Zheng H, Ge W, Huang Y,
Yan Y, Zhou X, Zhu W, Kong Y, Ding Y and Wang W: Identification of
chemoresistance-related mRNAs based on gemcitabine-resistant
pancreatic cancer cell lines. Cancer Med. 9:1115–1130.
2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Chaisaingmongkol J, Budhu A, Dang H,
Rabibhadana S, Pupacdi B, Kwon SM, Forgues M, Pomyen Y,
Bhudhisawasdi V, Lertprasertsuke N, et al: Common molecular
subtypes among Asian hepatocellular carcinoma and
cholangiocarcinoma. Cancer Cell. 32:57–70.e3. 2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Galaxy Community: The Galaxy platform for
accessible, reproducible, and collaborative data analyses: 2024
Update. Nucleic Acids Res. 52 (W1):W83–W94. 2024.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Bardou P, Mariette J, Escudié F, Djemiel C
and Klopp C: jvenn: An interactive Venn diagram viewer. BMC
Bioinformatics. 15(293)2014.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Cerami E, Gao J, Dogrusoz U, Gross BE,
Sumer SO, Aksoy BA, Jacobsen A, Byrne CJ, Heuer ML, Larsson E, et
al: The cBio cancer genomics portal: An open platform for exploring
multidimensional cancer genomics data. Cancer Discov. 2:401–404.
2012.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Abramson J, Adler J, Dunger J, Evans R,
Green T, Pritzel A, Ronneberger O, Willmore L, Ballard AJ, Bambrick
J, et al: Accurate structure prediction of biomolecular
interactions with AlphaFold 3. Nature. 630:493–500. 2024.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Jurrus E, Engel D, Star K, Monson K,
Brandi J, Felberg LE, Brookes DH, Wilson L, Chen J, Liles K, et al:
Improvements to the APBS biomolecular solvation software suite.
Protein Sci. 27:112–128. 2018.PubMed/NCBI View
Article : Google Scholar
|
|
23
|
Søndergaard CR, Olsson MHM, Rostkowski M
and Jensen JH: Improved treatment of ligands and coupling effects
in empirical calculation and rationalization of pKa values. J Chem
Theory Comput. 7:2284–2295. 2011.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Tian C, Kasavajhala K, Belfon KAA,
Raguette L, Huang H, Migues AN, Bickel J, Wang Y, Pincay J, Wu Q
and Simmerling C: ff19SB: Amino-acid-specific protein backbone
parameters trained against quantum mechanics energy surfaces in
solution. J Chem Theory Comput. 16:528–552. 2020.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Jorgensen WL, Chandrasekhar J, Madura JD,
Impey RW and Klein ML: Comparison of simple potential functions for
simulating liquid water. J Chem Phys. 79:926–935. 1983.
|
|
26
|
Wang S, Xie J, Pei J and Lai L: CavityPlus
2022 update: An integrated platform for comprehensive protein
cavity detection and property analyses with user-friendly tools and
cavity databases. J Mol Biol. 435(168141)2023.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Xu Y, Wang S, Hu Q, Gao S, Ma X, Zhang W,
Shen Y, Chen F, Lai L and Pei J: CavityPlus: A web server for
protein cavity detection with pharmacophore modelling, allosteric
site identification and covalent ligand binding ability prediction.
Nucleic Acids Res. 46 (W1):W374–W379. 2018.PubMed/NCBI View Article : Google Scholar
|
|
28
|
Eberhardt J, Santos-Martins D, Tillack AF
and Forli S: AutoDock Vina 1.2.0: New docking methods, expanded
force field, and python bindings. J Chem Inf Model. 61:3891–3898.
2021.PubMed/NCBI View Article : Google Scholar
|
|
29
|
Trott O and Olson AJ: AutoDock Vina:
Improving the speed and accuracy of docking with a new scoring
function, efficient optimization, and multithreading. J Comput
Chem. 31:455–461. 2010.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Wei W, Chen H and Liu S: Knockdown of
Rhotekin 2 expression suppresses proliferation and invasion and
induces apoptosis in hepatocellular carcinoma cells. Mol Med Rep.
13:4865–4871. 2016.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Pang X, Li R, Shi D, Pan X, Ma C, Zhang G,
Mu C and Chen W: Knockdown of Rhotekin 2 expression suppresses
proliferation and induces apoptosis in colon cancer cells. Oncol
Lett. 14:8028–8034. 2017.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Wang X, Zhang L, Wang W and Wang Y, Chen
Y, Xie R, Li X and Wang Y: Rhotekin 2 silencing inhibits
proliferation and induces apoptosis in human osteosarcoma cells.
Biosci Rep. 38(BSR20181384)2018.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Ji L and Huang Y, Zhang Y, Peng A, Qin J,
Lu S and Huang Y: RTKN2 is associated with unfavorable prognosis
and promotes progression in non-small-cell lung cancer. Onco
Targets Ther. 13:10729–10738. 2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Collier FM, Loving A, Baker AJ, McLeod J,
Walder K and Kirkland MA: RTKN2 induces NF-kappaB dependent
resistance to intrinsic apoptosis in HEK cells and regulates BCL-2
genes in human CD4(+) lymphocytes. J Cell Death. 2:9–23.
2009.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Zhang X, Wang J and Li H: RTKN2 knockdown
alleviates the malignancy of breast cancer cells by regulating the
Wnt/β-catenin pathway. Sci Rep. 13(23023)2023.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Zhang GF, Qiu L, Yang SL, Wu JC and Liu
TJ: Wnt/β-catenin signaling as an emerging potential key
pharmacological target in cholangiocarcinoma. Biosci Rep.
40(BSR20193353)2020.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Liu C, Jin Y and Fan Z: The mechanism of
warburg effect-induced chemoresistance in cancer. Front Oncol.
11(698023)2021.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Bhattacharya B, Mohd Omar MF and Soong R:
The Warburg effect and drug resistance. Br J Pharmacol.
173:970–979. 2016.PubMed/NCBI View Article : Google Scholar
|
|
39
|
de Oliveira Santos LAB and de Aragão
Batista MV: Structure-based virtual screening and drug repurposing
studies indicate potential inhibitors of bovine papillomavirus E6
oncoprotein. Microbiol Immunol. 68:414–426. 2024.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Soldi R, Halder TG, Sampson S,
Vankayalapati H, Weston A, Thode T, Bhalla KN, Ng S, Rodriguez Del
Villar R, Drenner K, et al: The small molecule BC-2059 inhibits
wingless/integrated (Wnt)-dependent gene transcription in cancer
through disruption of the transducin β-Like 1-β-catenin protein
complex. J Pharmacol Exp Ther. 378:77–86. 2021.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Nomura M, Rainusso N, Lee YC, Dawson B,
Coarfa C, Han R, Larson JL, Shuck R, Kurenbekova L and Yustein JT:
Tegavivint and the β-catenin/ALDH axis in chemotherapy-resistant
and metastatic osteosarcoma. J Natl Cancer Inst. 111:1216–1227.
2019.PubMed/NCBI View Article : Google Scholar
|
|
42
|
Savvidou I, Khong T, Cuddihy A, McLean C,
Horrigan S and Spencer A: β-Catenin inhibitor BC2059 is efficacious
as monotherapy or in combination with proteasome inhibitor
bortezomib in multiple myeloma. Mol Cancer Ther. 16:1765–1778.
2017.
|
|
43
|
Fowler NH, Samaniego F, Jurczak W, Ghosh
N, Derenzini E, Reeves JA, Knopińska-Posłuszny W, Cheah CY,
Phillips T, Lech-Maranda E, et al: Umbralisib, a dual PI3Kδ/CK1ε
inhibitor in patients with relapsed or refractory indolent
lymphoma. J Clin Oncol. 39:1609–1618. 2021.
|
|
44
|
Dhillon S and Keam SJ: Umbralisib: First
approval. Drugs. 81:857–866. 2021.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Gough SM, Flanagan JJ, Teh J, Andreoli M,
Rousseau E, Pannone M, Bookbinder M, Willard R, Davenport K,
Bortolon E, et al: Oral estrogen receptor PROTAC vepdegestrant
(ARV-471) is highly efficacious as monotherapy and in combination
with CDK4/6 or PI3K/mTOR pathway inhibitors in preclinical ER+
breast cancer models. Clin Cancer Res. 30:3549–3563.
2024.PubMed/NCBI View Article : Google Scholar
|
|
46
|
Campone M, De Laurentiis M, Jhaveri K, Hu
X, Ladoire S, Patsouris A, Zamagni C, Cui J, Cazzaniga M, Cil T, et
al: Vepdegestrant, a PROTAC estrogen receptor degrader, in advanced
breast cancer. N Engl J Med. 393:556–568. 2025.PubMed/NCBI View Article : Google Scholar
|
|
47
|
Hamilton EP, Ma C, De Laurentiis M, Iwata
H, Hurvitz SA, Wander SA, Danso M, Lu DR, Perkins Smith J, Liu Y,
et al: VERITAC-2: A phase III study of vepdegestrant, a PROTAC ER
degrader, versus fulvestrant in ER+/HER2-advanced breast cancer.
Future Oncol. 20:2447–2455. 2024.PubMed/NCBI View Article : Google Scholar
|
|
48
|
Hurvitz S, Schott A, Ma C, Nanda R, Zahrah
G, Hunter N, Tan A, Telli M, Mesias JA, Jeselsohn R, et al:
Abstract PO3-05-08: Updated results from VERITAC evaluating
vepdegestrant, a PROteolysis targeting chimera (PROTAC) estrogen
receptor (ER) degrader, in ER-positive/human epidermal growth
factor receptor 2 (HER2)-negative advanced breast cancer. Cancer
Res. 84 (9 Suppl)(PO3-05-08)2024.
|
|
49
|
Iwata H, Hamilton E, Ma C, De Laurentiis
M, Hurvitz SA, Wander SA, Danso MA, Lu DR, Perkins J, Liu Y, et al:
73TiP global phase III studies evaluating vepdegestrant in estrogen
receptor (ER)+/human epidermal growth factor receptor 2
(HER2)-advanced breast cancer: VERITAC-2 and VERITAC-3. Ann Oncol.
34 (Suppl 4)(S1493)2023.
|
|
50
|
Hewitt DJ, Aurora SK, Dodick DW, Goadsby
PJ, Ge YJ, Bachman R, Taraborelli D, Fan X, Assaid C, Lines C and
Ho TW: Randomized controlled trial of the CGRP receptor antagonist
MK-3207 in the acute treatment of migraine. Cephalalgia.
31:712–722. 2011.PubMed/NCBI View Article : Google Scholar
|