Introduction
Velvet antler growth is a rapid process (1). During this period, constitutive tissues
including bone, cartilage, skin, nerves and blood vessels grow
quickly (2). Previous studies have
demonstrated that certain growth factors, including vascular
endothelial growth factor, epidermal growth factor, fibroblast
growth factor and nerve growth factor are abundant in velvet antler
and are responsible for rapid tissue growth (3–6).
Numerous studies have also revealed that velvet antler contains
amino acids (7–9), polypeptides (10,11),
proteins (9,12–15) and
phospholipids (16). However, there
have been few reports that assess the phytochemical components of
velvet antler, which are highly biologically active.
Ultra-performance liquid chromatography coupled with
electrospray ionization quadrupole time-of-flight tandem mass
spectrometry (UPLC/ESI-QTOF-MS) has been utilized to reliably
analyze the complex composition of food, herbal medicines and
biological samples (17–20). QTOF-MS analyzers offer a high mass
resolution, sensitivity and accuracy, providing accurate ion masses
to determine molecular formulas (17–20). The
present study developed a method of UPLC/QTOF-MS on a Waters Xevo
G2-XS QTOF system for the determination of the chemical components
of velvet antler. Based on accurate mass measurements, a total of
87 compounds were detected in the velvet antler and tentatively
identified using the Traditional Medicine Library (TML) of the
UNIFI platform.
Phospholipids are major constituents of biological
membranes, which maintain membrane integrity and cell homeostasis
(21,22). Glycerophospholipids (GPLs) are the
largest family of amphiphilic phospholipids (23). They are comprised of a glycerol
backbone containing fatty acid chains that are acylated at the sn-1
and sn-2 position and possess a polar head group containing a
phosphate esterified at the sn-3 position, which is attached to a
polyol or amino acid moiety (23).
The following classes of GPLs are defined by the structure of the
head group: phosphatidylserine, phosphatidylglycerol,
phosphatidylethanolamine, phosphatidylinositol, phosphatidic acid
and phosphatidylcholine (23). Each
class of GLP comprises many species that possess the same head
group but different chain lengths, number of unsaturations and
substitution positions of the esterified fatty acid (23). Various methods have been utilized for
the quantification and structural determination of individual
molecular species of phospholipids. For example, thin-layer
chromatography was used in the chemical analysis of components from
velvet antlers (24). Gas
chromatography techniques are not so generalized for determining
the components from velvet antler, due to analytical difficulty,
even following derivatization, and the long time required for these
reactions and the low limits of detection (LOD) (25). QTOF-MS has several advantages,
including a high reliability, short analysis time, a small required
sample quantity and a high accuracy in the identification and
quantification of phospholipids (26,27).
The present study determined the phospholipids
present in velvet antler, which was determined via
qualitative-quantitative analysis using the UPLC/QTOF-MS method.
Previously, Zhou and Li (26)
identified 5 phospholipids in the velvet antlers of sika deer
including sphingomyelin, phosphatidylcholine,
phosphatidylethanolamine, lysophosphatidylcholine and
lysophosphatidylethanolamine. The present study identified 16
phospholipids from velvet antler extracts, with 3 phospholipids
being quantified. Consequently, a UPLC/QTOF-MS method with
commendable validation results, a slope of the standard curve and
precision was developed and validated for the quantitative
detection of phospholipids and the identification of complex
components in velvet antler. This method may provide an
experimental basis for further pharmacological and clinical
applications of velvet antler products.
Materials and methods
Materials
Antler velvet (Cervus nippon Temminck var.
mantchurieus Sainhoe) was collected from farmed sika deer in the
Shuangyan district of Changchun (Jilin Ruikang Biotechnology Co.,
Ltd., Liaoyuan, China). The upper section was cut to a thickness of
2–3 mm, freeze dried and then powdered to 160–180 mesh (84–95 µm).
Antler velvet samples were identified at School of Pharmaceutical
Sciences in Jilin University (Changchun, China).
Mass spectrometry grade acetonitrile and methanol
were purchased from Thermo Fisher Scientific, Inc., (Waltham, MA,
USA). Phospholipid standards
(1-myristoyl-sn-glycero-3-phosphocholine, 98.5%;
dimyristoyl-sn-glycero-3-phospho choline, 98.5%; and
1-palmitoyl-2-oleoyl-sn-glycero-3-phosphocholine, 98.5%) were
purchased from Nanjing NutriHerb BioTech Co., Ltd., (Jiangsu,
China). Formic acid was purchased from Sigma-Aldrich (Merck KGaA,
Darmstadt, Germany). Leucine-enkephalin (99%) was also purchased
from Sigma-Aldrich; Merck KGaA. Deionized water was prepared using
a Milli-Q system (EMD Millipore, Billerica, MA, USA). All other
reagents were of analytical grade.
UPLC/QTOF-MS conditions
An ACQUITY UPLC I-Class System coupled to a Waters
Xevo G2-XS QTOF mass spectrometer detector (Waters UK, Elstree, UK)
was used. All chromatographic and MS equipment was purchased from
Waters Corporation (Milford, MA, USA). Chromatographic separations
were achieved using an ACQUITY UPLC® BEH C18, 1.7 µm
(2.1×50 mm; Waters Corporation) capillary column. Analytical column
chromatography was performed at 40°C. The mobile phase consisted of
a mixture of acetonitrile, water and formic acid at a flow rate of
0.4 ml min−1. Acetonitrile with 0.1% formic acid was
used as mobile phase A and water with 0.1% formic acid was used as
mobile phase B. A 90/10 mixture of water/acetonitrile was utilized
as the weak wash solvent and 50/50 water/acetonitrile was used as
the strong wash solvent for rinsing the injection needle. Prior to
running the elution, the column was equilibrated to 35%. The
elution gradient program was 35–82% A from 0–3 min, 82% A from 3–7
min, 35–82% A from 7–8 min and 35% A from 8–9 min.
MS experiments were performed using a Waters Xevo
G2-XS QTOF mass spectrometer connected to the ACQUITY UPLC I-Class
System via an electrospray ionization (ESI) interface. Atmospheric
pressure ionization was performed in positive ion, negative ion and
sensitivity analyzer modes for QTOF-MS data acquisition. A wide
mass range (m/z 100–1200) was selected for the acquisition of
accurate mass precursor and fragment ion data. The corona voltage,
sampling cone voltage, source temperature and desolvation
temperature was 3.0 kV, 40 V, 100°C and 350°C, respectively.
Nitrogen (20±2°C; 10 psi) was used for desolvation and the cone gas
flow rate was 800 and 50 l h−1, respectively. Argon was
used as the collision gas and the collision energy was 15–45 V for
high energy ionizations. Data were acquired and analyzed using
MassLynx™ NT 4.1 software (Waters Corporation). Analyses were
performed in full scan mode and the scan time was set to 0.2 sec.
To ensure for mass accuracy and reproducibility of the optimized MS
conditions, leucine-enkephalin (m/z 554.2615 in negative mode and
m/z 556.2771 in positive mode) was used as a reference (lock mass)
at a concentration of 200 pg/ml and a flow rate of 10 µl/min. The
reference was injected into the MS instrument every 10 sec. The
instrument was calibrated using sodium formate solution as the
calibration standard to achieve mass accuracies of <0.5 mDa.
Accurate mass screening of the
constituents of velvet antler
The UNIFI 1.8 informatics platform (Waters
Corporation) was utilized to integrate data acquisition, data
mining, library searching and to generate a report (27,28). The
raw data were imported and screened against the TML and a
customized phospholipid library produced in the current study. A
natural product analytical workflow within UNIFI was used to
analyze the chromatographic and mass spectral data of the velvet
antler extract components utilizing various in-built tools,
including the customized library and filters.
The TML of the UNIFI software contains 6,415
compounds and their associated data. A customized library was also
created in the current study that comprised 45 phospholipids with
detailed metadata (including the molecular structure and compound
name) based on their chemical structure. Compound screening was
performed by setting a mass tolerance of 2 mDa, a retention time
cutoff of ±0.2 min, counts >1,000 and a minimum of 5
fragmentations, and a mean of 10 false detects per sample analysis.
To demonstrate the validity of these results three standards were
utilized: 1-myristoyl-sn-glycero-3-phosphocholine (MPC),
1,2-dimyristoyl-sn-glycero-3-phosphocholine (DPC) and 1-palmitoyl-
2-oleoyl-sn-glycero-3-phosphocholine (POPC). Their retention times
and accurate masses were compared with the sample of velvet antler
extract.
Extraction of velvet antler
An ultrasound-assisted extraction (25°C; 40 kHz) of
100 g velvet antler powder was performed with 3×150 ml ethanol for
10 min each time. The solid was filtered in each step and the pore
size was 30–50 µm. The filtered and pooled liquid phases were
concentrated to 3 ml under a reduced pressure at −20°C.
Subsequently, 6 ml acetone was added to generate a white
precipitate. Then the mixture was centrifuged at 2,012 × g at 20°C
for 10 min, the supernatant was removed, and the precipitate was
stored at −20°C until analysis. The precipitate was dissolved in
methanol to achieve the concentration of 40 mg/ml, and 2 ml
solution was filtered by 0.22-µm microporous filter membrane and
put into automatic sampling bottle prior injection of 5 µl
methanolic solution into the UPLC system. Internal standards with
were added to the antler powder to allow commenting on the
extraction procedure. The mean recovery rates were in the range of
90–110%.
Qualitative determination of MPC, DPC
and POPC in EVA samples by UPLC/QTOF-MS/MS
The MS data of the 3 phospholipid standards were
initially assessed using either ESI or the atmospheric pressure
chemical ionization (APCI) mode. ESI was selected as the ionization
mode for the present experiments as it provides greater analyte
responses than those achieved with APCI. Furthermore, high
ionization efficiency was observed under ESI conditions when
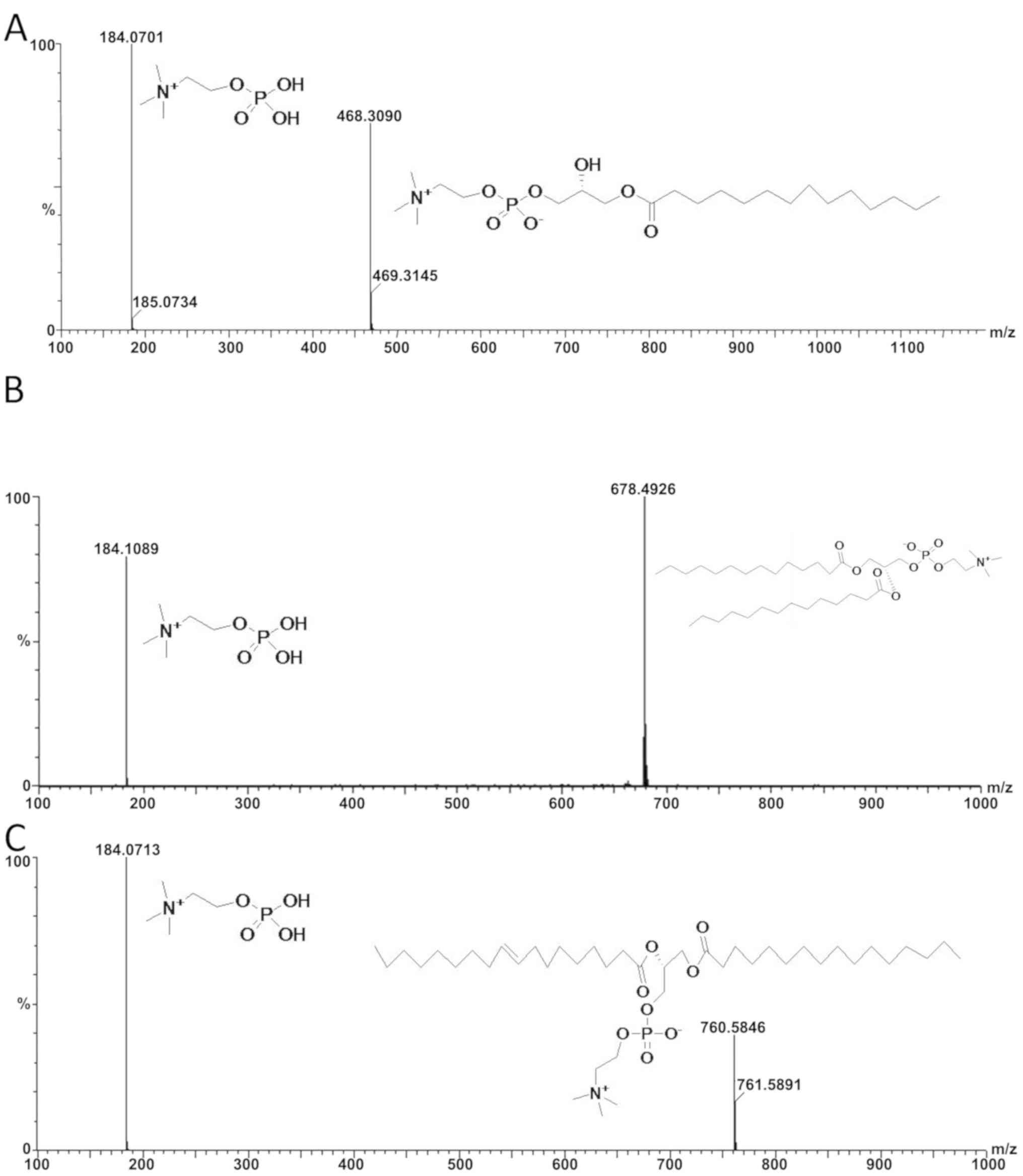
monitoring the signal in positive ion mode. Following instrument
parameter optimization to achieve the highest sensitivity and
lowest background noise for the protonated molecules of MPC, DPC
and POPC, the ion transition (m/z) 468.30→184.07 was selected for
the quantification of MPC, m/z 678.49→184.10 for DPC and m/z
760.58→184.07 for POPC (Fig. 1).
Other UPLC and MS conditions were the same as those
aforementioned.
Method validation of phospholipid
quantitative detection
The method of quantitative phospholipid detection
was fully validated according to the guidelines set by the US Food
and Drug Administration. Specificity was tested by inspecting the
solvent used in each validation run for interfering peaks. The
calibration curve was determined by plotting the peak area vs. the
corresponding concentration of injected standards. The limit of
quantitation (LOQ) was the concentration that exhibited an
identifiable and reproducible analyte peak (response) with a
precision of 10% and an accuracy of 90–110%. Additionally, the
analyte response at the LOQ should be at least ten times the
response of the blank sample. Intra- and inter-day precision and
accuracy were determined from 6 replicates of the QC samples
analyzed on the same day and on 3 different days. At each
concentration, acceptable precision (repeatability) and accuracy
were defined as the relative standard deviation (RSD) of <10%
and a relative error within ±10%. The sample recovery was
calculated at three different concentrations by comparing the peak
areas of the sample and the peaks in samples spiked with standard.
Short- and long-term stability were investigated by reanalyzing the
quality control batches following storage at −20°C for 30 days and
at room temperature for 12 h, respectively.
Results
Characterization of velvet antler
complex constituents
A total of 87 compounds in velvet antler were
identified or tentatively characterized. These included: 1 lignan,
30 terpenoids (including 20 triterpenes), 39 steroids, 8 alkaloids,
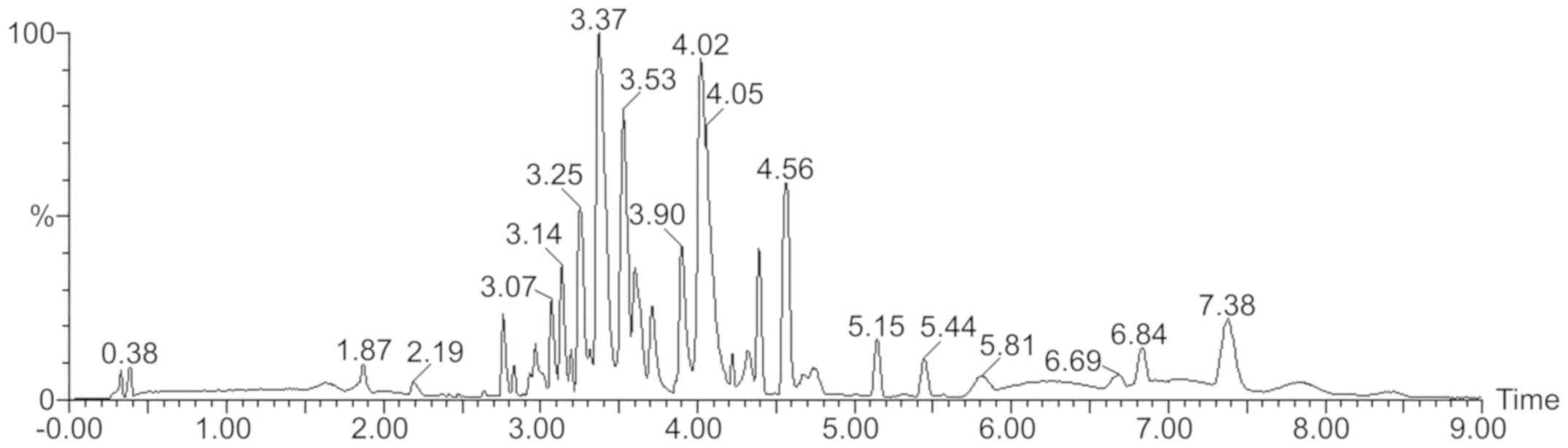
4 organic acids and 5 esters (Table
I; Fig. 2). The compounds were
identified based on accurate mass measurements, tandem MS
behaviors, database matching and comparison to reference standards,
considering all data reported in the literature. The advantages of
using UPLC for the analysis of samples with complex components
(including enhance separation efficiency and higher peak capacity)
are fully demonstrated here, for example via the shorter
chromatographic peaks (Fig. 2).
Enhanced separation efficiency (sharper chromatographic peaks) and
higher peak capacity were observed in the analysis at 10 min. The
raw data includes the molecular weight of the compounds and the
respective fragment ion information, which may be matched to the
TML for in-depth ingredient analysis and structural identification
(Data not shown).
 | Table I.Results of complex constituent
accurate mass screening in velvet antler. |
Table I.
Results of complex constituent
accurate mass screening in velvet antler.
| Category | Formula | Observed m/z | Mass error
(mDa) | Observed retention
time (min) | Responds | Adducts | Identified |
|---|
| Lignans |
| 1 |
C29H28O9 | 559.1316 | −4.9 | 5.14 | 185584 | +K, +Na | Schisantherin
D |
| Alkaloids |
| 2 |
C27H43NO4 | 468.3084 | 0.0 | 2.77 | 806437 | +Na | Caussin alkali
nitrogen oxide |
| 3 |
C27H45NO5 | 502.2938 | 0.9 | 3.08 | 238334 | +K | Pingbeimine A |
| 4 |
C28H47NO3 | 468.3447 | −0.1 | 3.24 | 43744 | +Na | Pingbeinine |
| 5 |
C28H47NO2 | 452.3496 | −0.3 | 3.35 | 120928 | +Na | sevcoridinine |
| 6 |
C27H43NO3 | 452.3095 | −4.0 | 3.81 | 79749 | +Na | peiminine |
| 7 |
C41H80NO8P | 768.5459 | −5.5 | 3.92 | 42787 | +Na |
Phosphatidylethanolamine (PE) |
| 8 |
C29H48O7 | 509.3451 | −2.2 | 3.99 | 335087 | +H | Rhapontisterone
C |
| 9 |
C10H14N2O5 | 281.0521 | −1.3 | 4.22 | 18316 | +K | Thymine DNA
nucleotides |
| Organic acids |
| 10 |
C22H34O4 | 363.2504 | −2.5 | 3.79 | 2273591 | +H | 4-OH-ginkgolic
acid |
| 11 |
C18H30O2 | 301.2118 | −2.0 | 5.31 | 17789 | +Na | γ-Linolenate |
| 12 |
C20H32O2 | 327.2271 | −2.3 | 5.50 | 22385 | +Na, +K | Arachidonic
Acid |
| 13 |
C24H48O3 | 423.3202 | −3.3 | 6.52 | 10920 | +K | cerebronic
acid |
| Steroids |
| 14 |
C27H40O6 | 461.2907 | 0.9 | 2.70 | 126562 | +H | lucidenic acid
N |
| 15 |
C36H58O10 | 673.3917 | −0.5 | 3.40 | 24412 | +Na | Huangqiyiesaponin
A |
| 16 |
C30H46O7 | 541.3133 | −0.3 | 3.43 | 22203 | +Na | Picfeltarraegenin
VII |
| 17 |
C22H34O5 | 379.2455 | −2.4 | 3.45 | 20114 | +H | Tenacigenin A |
| 18 |
C28H42O6 | 497.2877 | 0.4 | 3.47 | 55767 | +Na | Methyl- lucidenic
acid Q |
| 19 |
C27H42O5 | 485.2635 | −2.8 | 3.53 | 97889 | +K | Ophiopogon
japonicus saponin |
| 20 |
C32H46O6 | 549.3169 | −1.8 | 3.53 | 46050 | +Na | Alisol J
23-acetate |
| 21 |
C35H56O9 | 643.3812 | −0.4 | 3.70 | 57942 | +Na | Cimiaceroside
B |
| 22 |
C28H46O7 | 495.3289 | −2.8 | 3.72 | 227048 | +H | Periplocoside
L |
| 23 |
C30H44O6 | 523.3036 | 0.6 | 3.82 | 48765 | +Na | Picfeltarraenone
II |
| 24 |
C29H46O7 | 507.3288 | −2.9 | 3.82 | 56876 | +H | Decumbesterone
A |
| 25 |
C27H42O6 | 463.3022 | −3.2 | 3.83 | 105652 | +H | 23, 27-2OH-
pennogenin |
| 26 |
C35H56O9 | 621.3983 | −1.4 | 4.00 | 46793 | +H, +Na | Cimidahuside G |
| 27 |
C33H52O8 | 577.3727 | −0.8 | 4.02 | 67266 | +H, +Na | Trillin |
| 28 |
C27H44O6 | 465.3188 | −2.3 | 4.04 | 774273 | +H | Periplocoside
N |
| 29 |
C28H36O4 | 437.2692 | 0.6 | 4.07 | 576968 | +H | Daturametelin
D |
| 30 |
C43H66O11 | 759.4659 | −1.9 | 4.19 | 35772 | +H |
7-O-sitosteriol-β-D-glucopyranoside-tetra
acetyl ester |
| 31 |
C36H58O9 | 635.4135 | −1.9 | 4.24 | 134111 | +H |
25-O-methyl-cimigenol xyloside |
| 32 |
C38H62O10 | 679.4397 | −1.8 | 4.23 | 64727 | +H | Yesanchinoside
H |
| 33 |
C24H40O4 | 415.2829 | 1.0 | 4.30 | 58724 | +Na | Chenodeoxycholic
acid |
| 34 |
C28H46O6 | 479.3348 | −1.9 | 4.35 | 2060505 | +H | Fungal steroid
A |
| 35 |
C33H52O7 | 561.3766 | −2.0 | 4.70 | 372203 | +H | Ajugamarin C |
| 36 |
C29H44O5 | 473.3258 | −0.3 | 4.76 | 130477 | +H | Momordica acid
A |
| 37 |
C28H40O4 | 441.2991 | −0.8 | 5.31 | 53350 | +H | Momordica acid
C |
| 38 |
C30H50O3 | 481.3642 | −1.0 | 5.31 | 32662 | +Na | Curculigenin C |
| 39 |
C35H60O6 | 577.4455 | −0.8 | 5.31 | 15112 | +H | Daucosterol |
| 40 |
C41H70O11 | 761.4798 | −1.3 | 5.42 | 45206 | +Na |
β-sitosterol-3-O-gentiobioside |
| 41 |
C32H50O6 | 531.3678 | −0.2 | 5.52 | 16522 | +H | Alisol F
24-acetate |
| 42 |
C35H58O8 | 629.4002 | −2.2 | 5.54 | 20278 | +Na | Curculigosaponin
B |
| 43 |
C33H54O7 | 585.3733 | −2.8 | 5.58 | 14707 | +Na | Tribuloside F |
| 44 |
C33H52O9 | 593.3648 | −3.6 | 5.96 | 46057 | +H |
Spirostan-12-one,3-hydroxy--3-O-β-D--galactose |
| 45 |
C27H44O3 | 439.3174 | −0.9 | 6.00 | 39914 | +Na, +K | Tigogenin |
| 46 |
C30H50O4 | 497.3579 | −2.3 | 6.02 | 24851 | +Na | Curculigenin A |
| 47 |
C39H56O4 | 589.4206 | −4.6 | 6.27 | 27283 | +H |
11-O-p-Coumarylnepeticin |
| 48 |
C35H56O9 | 621.3980 | −1.8 | 6.69 | 28395 | +H |
Cimigenolxyloside |
| 49 |
C24H40O4 | 393.2979 | −2.1 | 6.97 | 92484 | +H | Deoxycholic
acid |
| 50 |
C35H60O7 | 593.4429 | 1.7 | 7.52 | 47272 | +H |
7-IKshusterol-3-glucoside |
| 51 |
C27H46O | 409.3415 | −2.6 | 7.81 | 16049 | +Na | Cholesterol |
| 52 |
C31H52O5 | 505.3905 | 1.8 | 7.83 | 23108 | +H | Alisol A
25-methoxyl |
| Triterpenes |
| 53 |
C31H48O7 | 555.3303 | 1.1 | 3.72 | 83437 | +Na |
Phytolaccagenin |
| 54 |
C40H64O11 | 743.4318 | −2.3 | 3.75 | 17152 | +Na |
3-O-β-D-xylopyranoside-(1→2)-α-L-arabinopyranoside-oleanolic
acid |
| 55 |
C36H56O9 | 655.3799 | −1.7 | 3.77 | 30610 | +Na |
Deglucose-chikusetsu saponin IVa |
| 56 |
C34H52O8 | 611.3539 | −1.5 | 3.79 | 37775 | +Na | Quinatoside A |
| 57 |
C36H58O9 | 657.3973 | 0.0 | 3.96 | 74016 | +Na | Eclalbasaponin
II |
| 58 |
C30H46O6 | 503.3350 | −1.7 | 4.29 | 261873 | +H | Phytolaccic
acid |
| 59 |
C28H42O5 | 459.3089 | −1.6 | 4.30 | 143388 | +H |
3β,4β,23-Trihydroxy-24,30-dinorolean-12,20(29)-dien-28-oic
acid |
| 60 |
C31H50O5 | 541.3247 | −4.3 | 4.30 | 22859 | +K | Rose acid methyl
ester |
| 61 |
C36H58O9 | 635.4139 | −1.4 | 4.39 | 22345 | +H, +Na | Soyasopogeno
B-3-glucuronide |
| 62 |
C30H50O5 | 513.3539 | −1.2 | 4.58 | 25681 | +Na | Extensor kiosk |
| 63 |
C35H56O8 | 605.4032 | −1.5 | 4.67 | 273930 | +H | Slope
acid-3-β-O-α-L-arabia glycoside |
| 64 |
C31H48O6 | 517.3511 | −1.3 | 4.74 | 223026 | +H | Phytolaccagenic
Acid |
| 65 |
C35H56O7 | 611.3931 | 1.2 | 5.03 | 230065 | +Na | RaddeanosideR0 |
| 66 |
C30H44O4 | 469.3286 | −2.6 | 6.50 | 60456 | +H |
21α-Hydroxy-3-oxo-11,13(18)-oleanadien-28-oic
acid |
| 67 |
C30H48O | 425.3767 | −1.1 | 7.00 | 12496 | +H | Alnusenone |
| 68 |
C28H34O7 | 483.2394 | 1.7 | 3.14 | 57783 | +H | Gedunin |
| 69 |
C32H48O6 | 551.3323 | −2.0 | 4.04 | 52278 | +Na | Poricoic acid
DM |
| 70 |
C43H68O15 | 825.4559 | −7.2 | 4.45 | 17042 | +H | Cimifoetiside |
| 71 |
C30H50O6 | 529.3455 | −4.5 | 4.62 | 21375 | +Na | 13β,17β-Epoxyalisol
A |
| 72 |
C30H50O5 | 513.3507 | −4.4 | 5.31 | 33473 | +Na, +H | melianotriol |
| Terpenoids |
| 73 |
C30H42O6 | 521.2857 | −1.6 | 2.96 | 64304 | +Na | 3-O-(2E,
4Z-decadienoyl)-20-O-acetylingenol |
| 74 |
C26H38O5 | 453.2615 | 0.4 | 3.44 | 71331 | +Na |
7β-(3-Ethyl-cis-crotonoyloxy)-1α-(2-methyl-butyryloxy)-3,14-dehydro-Z-notonipetranone |
| 75 |
C25H40O6 | 437.2867 | −3.1 | 3.61 | 43927 | +H |
Oriediterpenoside |
| 76 |
C21H32O4 | 349.2347 | −2.6 | 3.62 | 211816 | +H |
7β-(3-Ethyl-cis-crotonoyloxy)-14-hydroxynotonipetranone |
| 77 |
C24H38O5 | 407.2765 | −2.7 | 3.76 | 868188 | +H | Vitetrifolin D |
| 78 |
C22H34O4 | 363.2510 | −2.0 | 5.06 | 299922 | +H | Rotundifuran |
| 79 |
C20H30O4 | 335.2195 | −2.2 | 3.19 | 30342 | +H | Preleoheterin |
| 80 |
C22H32O6 | 393.2240 | −3.1 | 0.53 | 73288 | +H | Nigakihemiacetal
F |
| 81 |
C22H32O8 | 425.2148 | −2.2 | 3.96 | 44073 | +H | Nigakilactone
H |
| 82 |
C22H34O5 | 379.2456 | −2.3 | 3.19 | 33482 | +H | Tussilagone |
| Esters |
| 83 |
C22H34O4 | 363.2509 | −2.1 | 4.12 | 867964 | +H | Di-n-heptyl
phthalate |
| 84 |
C21H38O4 | 355.2828 | −1.5 | 4.30 | 65878 | +H | 2-Monolinolein |
| 85 |
C19H38O4 | 353.2667 | 0.5 | 4.70 | 513087 | +Na, +H, +K | monopalmitin |
| 86 |
C24H38O4 | 413.2660 | −0.2 | 5.31 | 97963 | +Na, +H, +K |
Di(2-ethylhexyl)phthalate |
| 87 |
C21H42O4 | 381.2974 | −0.2 | 5.95 | 263612 | +Na, +H, +K | β-Glyceryl
Monostearate |
Velvet antler phospholipid
identification
A customized library was constructed to verify the
identity of velvet antler phospholipids on the basis of previously
reported data (16,21–24). A
total of 45 phospholipids were added to the custom library with
details including compound name, chemical structure and chemical
formula. As a result, 16 phospholipids were identified or
tentatively characterized from the velvet antler extract. These
data, including retention time, formula, mass error, adducts and
compound names are presented in Table
II. The use of mass spectrometry-based approaches alone is
insufficient for the identification of complex botanical chemical
components (27). Therefore, the
present study utilized reference standards to validate these
compounds, enhancing the accuracy and reliability of the results
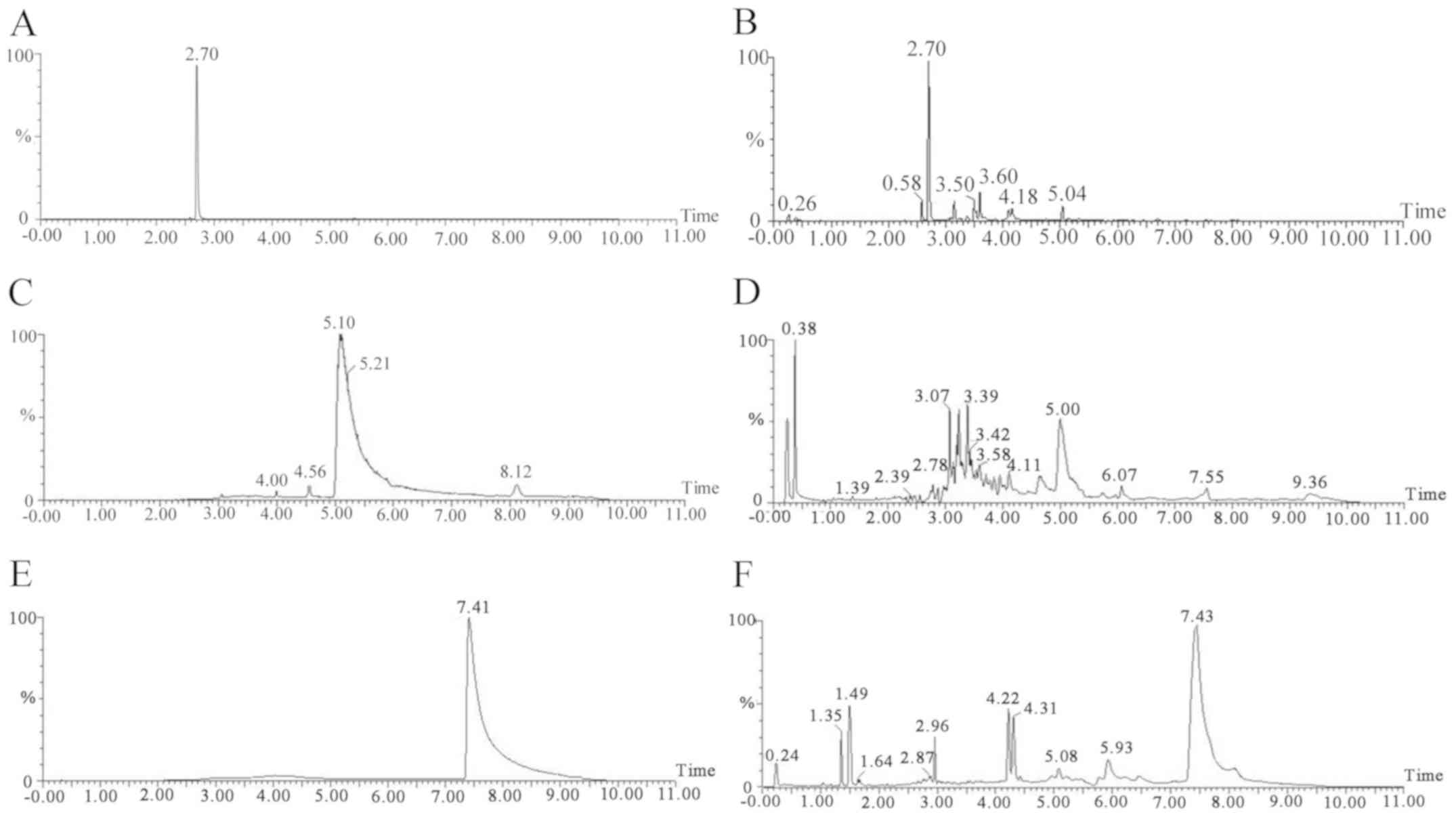
obtained. The standards of MPC, DPC and POPC were assayed under
optimized conditions and their spectra and chromatograms were
compared with those of the EVA samples. The results confirmed that
MPC, DPC and POPC were present in the EVA. The chromatograms and
spectra are presented in Figs. 1 and
3.
 | Table II.Identification of velvet antler
phospholipids. |
Table II.
Identification of velvet antler
phospholipids.
| Nο. | Formula | Observed m/z | Mass error
(mDa) | Observed retention
time (min) | Adducts | Identified |
|---|
| 1 |
C45H87O13P | 905.5428 | −8.8 | 2.30 | +K |
L-alpha-Phosphatidylinositol-4,5-bisphosphate |
| 2 |
C22H46NO7P | 468.3084 | −0.1 | 2.77 | +H |
1-Myristoyl-2-hydroxy-sn-glycero-3-phosphocholine |
| 3 |
C21H40NaO7P | 459.2481 | −0.1 | 3.38 | +H |
1-Oleoyl-sn-glycero-3-lysophosphatidic
acid sodium salt |
| 4 |
C26H54NO7P | 524.3714 | 0.3 | 4.03 | +H |
β-Acetyl-γ-O-hexadecyl-L-α-phosphatidylcholine |
| 5 |
C42H78NaO10P | 835.4824 | −3.8 | 4.18 | +K |
1,2-Dioleoyl-sn-glycero-3-phospho-rac-(1-glycerol)
sodium salt |
| 6 |
C42H82NO10P | 814.5507 | −6.1 | 4.19 | +Na |
Phosphatidylserine |
| 7 |
C33H66NO8P | 674.4082 | −7.5 | 4.22 | +K |
1,2-Dimyristoyl-sn-glycero-3-phosphoethanolamine |
| 8 |
C42H81Na2O10P | 861.5002 | 0.8 | 4.32 | +K |
1,2-Distearoyl-sn-glycero-3-phospho-rac-glycerol
sodium salt |
| 9 |
C29H58NO8P | 580.3977 | 0.4 | 4.34 | +H |
1,2-Dilauroyl-sn-glycero-3-phosphoethanolamine |
| 10 |
C42H82NO8P | 760.5794 | −5.7 | 4.58 | +H |
2-Oleoyl-1-palmitoyl-sn-glycero-3-phosphocholine |
| 11 |
C36H72NO8P | 716.4543 | −8.4 | 4.60 | +K |
1,2-Dimyristoyl-sn-glycero-3-phosphocholine |
| 12 |
C42H80NO8P | 780.5484 | −2.9 | 5.26 | +Na |
L-A-phosphatidylcholine |
| 13 |
C39H69O8P | 697.4815 | 1.2 | 5.37 | +H | L-α-phosphatidic
acid |
| 14 |
C35H67Na2O8P | 693.4433 | −0.9 | 5.49 | +H |
L-β,γ-Dipalmitoyl-L-α-phosphatidicaciddisodium
salt |
| 15 |
C44H88NO8P | 790.6295 | −2.5 | 6.02 | +H |
1,2-Distearoyl-rac-glycero-3-phosphocholine |
| 16 |
C40H80NO8P | 772.5217 | −3.7 | 7.01 | +K |
(18R,21S)-24-Amino-21-
hydroxy-21-oxido-15-oxo-16,20,22-trioxa-21λ5-phosphatetracosan-18-ylicosanoate |
Quantitative analysis of MPC, DPC and
POPC in EVA
The linearity, LODs, LOQs, precision, repeatability,
stability and recovery of MPC, DPC and POPC were determined using
the optimized UPLC/QTOF-MS/MS method. The calibration curves of
each are presented in Table III.
The results demonstrated that the correlation coefficients were all
>0.9995, indicating that good linear correlations were achieved.
The RSDs of the intra-day and inter-day precisions were deemed to
be acceptable (Table IV). The
results of the repeatability and stability tests, and the mean
recovery rates were also deemed to be in the range of 90–110%,
indicating that the qualitative method was accurate, reproducible
and reliable for the assessment of MPC, DPC and POPC in the
EVA.
 | Table III.Quantitative detection of MPC, DPC
and POPC in the extract of velvet antler. |
Table III.
Quantitative detection of MPC, DPC
and POPC in the extract of velvet antler.
| Phospholipids | Calibration
curve | r2 | Content of
phospholipids in velvet antler |
|---|
| MPC |
Y=7362.7X+21137.8 | 0.9996 | 1.07±0.02 µg/g |
| DPC |
Y=39811.5X+78542.2 | 0.9992 | 7.05±0.52 ng/g |
| POPC |
Y=1653604.2X-53602.4 | 0.9998 | 18.81±0.55
ng/g |
 | Table IV.Intra- and inter-day accuracy and
precision of quality control samples. |
Table IV.
Intra- and inter-day accuracy and
precision of quality control samples.
|
|
| Inter-day
(n=6) | Intra-day
(n=3) |
|---|
|
|
|
|
|
|---|
| Samples | Concentration
(µg/ml) | Precision
(RSD%) | Accuracy (RE%) | Precision
(RSD%) | Accuracy (RE%) |
|---|
| MPC | 10 | 4.1 | −2.1 | 5.9 | 3.2 |
|
| 20 | 3.5 | −3.6 | 4.5 | 2.5 |
|
| 30 | 2.8 | −1.8 | 4.1 | −2.9 |
| DPC | 5 | 1.8 | 2.5 | 3.9 | −2.3 |
|
| 10 | 1.6 | −2.0 | 3.7 | −4.0 |
|
| 20 | 1.2 | −1.8 | 5.6 | −3.3 |
| POPC | 15 | 3.4 | −2.7 | 6.5 | −4.5 |
|
| 20 | 3.6 | 3.2 | 5.5 | −2.4 |
|
| 30 | 2.7 | −3.3 | 5.1 | −2.8 |
The aforementioned UPLC/QTOF-MS/MS analytical method
was subsequently used to quantify the three phospholipids present
in the EVA samples. Each standard was analyzed in triplicate and
each sample was analyzed once to determine the average content of
the constituents. The analytical results are presented in Table III. The results demonstrated that
there was 1.07±0.02 µg/g of MPC, 7.05±0.51 ng/g of DPC and
18.81±0.55 ng/g of POPC in the EVA.
Discussion
The chemical composition of velvet antler was
determined in the present study using the UPLC/QTOF/MS method
combined with UNIFI software for component screening. Compared with
traditional identification methods that require long and complex
purification procedures and structure identification by nuclear
magnetic resonance (NMR) and mass spectrometry, the method used in
the present study is simple, fast and easy to operate. Although the
results of component screening are based on the mass ratio of
parent and fragment ions and not NMR data, this method remains
important for the estimation of velvet antler composition and for
providing reference values, particularly for the use of velvet
antler in combination with various clinical drugs.
Velvet antler has been demonstrated to exhibit
various anti-osteoporosis (29–31)
anti-fatigue (32,33), anti-inflammatory (34,35) and
anti-cancer (36) effects, which are
commonly associated with the chemical components of velvet antler.
The identification velvet antler chemical components determined in
the present study may facilitate further assessment into its
bioactivity and functional mechanism.
The qualitative and quantitative detection of
phospholipids in velvet antler was performed in the current study,
which revealed that 16 phospholipids were present. The content of
MPC in velvet antler was highest among the phospholipids and thus
likely contributes to its biological effects, particularly that of
anti-oxidation (37).
The phospholipids in velvet antler have been
reported to have various biological actives, for example
proliferation activity on spleen cells, and they are the subject of
increasing research interest throughout the world (38). Herein, a systematic, sensitive
bioanalytical UPLC/QTOF-MS/MS assay was developed to determine the
content of phospholipids in velvet antler; this method will
facilitate the quality control of velvet antler and can be widely
applied in clinical settings.
The analysis of phospholipids in velvet antler using
UPLC/QTOF-MS/MS has been demonstrated to be a suitable strategy for
biomarker discovery (18,21,23). A
total of 16 phospholipids in the EVA samples were identified and
three of these compounds were quantified. The current data revealed
that the content of phospholipids was low in the EVA samples,
hindering their detection by certain commonly used methods,
including high performance liquid chromatography with UV detection.
The method of UPLC/QTOF-MS/MS described in the current study
adequately addressed this problem as this quantitative method
exhibited great advantages in terms of ease of sample preparation,
excellent recovery and high sensitivity. To the best of our
knowledge, the lignans, alkaloids, organic acids, steroids and
terpenoids identified in velvet antler were detected and
tentatively characterized for the first time using the UNIFI
platform. However, further pharmacological studies are required to
explore the associations between velvet antler components and
bioactivity. This may advance its application in clinical
settings.
Acknowledgements
Not applicable.
Funding
The present study was supported by the Science and
Technology Development Program of Jilin Province (grant nos.
20150204037YY and 20160101072JC).
Availability of data and materials
All data generated or analyzed during the present
study are included in this published article.
Authors' contributions
JHL designed all experiments, organized data and
wrote this manuscript. LQZ determined the components of velvet
antler using UPLC/QTOF/MS and analyzed the data. JW took off
components from velvet antler with the method of ultrasound
assisted extraction. TL screened the components of velvet antler
with UNIFI platform and analyzed the data. PYL designed all
experiments and performed data analysis. YHW took off components
from velvet antler with the method of ultrasound assisted
extraction and determined the components of velvet antler using
UPLC/QTOF/MS. MY screened the components of velvet antler with
UNIFI platform and analyzed the data. JPL designed experiments
based on the references and analyzed the data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
Glossary
Abbreviations
Abbreviations:
|
UPLC/QTOF-MS
|
ultra-performance liquid
chromatography- quadrupole-time-of-flight mass spectrometry
|
|
MPC
|
1-myristoyl-sn-glycero-3-phosphocholine
|
|
DPC
|
dimyristoyl-sn-glycero-3-phosphocholine
|
|
POPC
|
1-palmitoyl-
2-oleoyl-sn-glycero-3-phosphocholine
|
|
EVA
|
extract of velvet antler
|
|
TML
|
Traditional Medicine Library
|
|
GPLs
|
glycerophospholipids.
|
References
|
1
|
Price JS, Allen S, Faucheux C, Althnaian T
and Mount JG: Deer antlers: A zoological curiosity or the key to
understanding organ regeneration in mammals? J Anat. 207:603–618.
2005. View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Clark DE, Li C, Wang W, Martin SK and
Suttie JM: Vascular localization and proliferation in the growing
tip of the deer antler. Anat Rec A Discov Mol Cell Evol Biol.
288:973–981. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Clark DE, Lord EA and Suttie JM:
Expression of VEGF and pleiotrophin in deer antler. Anat Rec A
Discov Mol Cell Evol Biol. 288:1281–1293. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Barling PM, Lai AK and Nicholson LF:
Distribution of EGF and its receptor in growing red deer antler.
Cell Biol Int. 29:229–236. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Lai AK, Hou WL, Verdon DJ, Nicholson LF
and Barling PM: The distribution of the growth factors FGF-2 and
VEGF, and their receptors, in growing red deer antler. Tissue Cell.
39:35–46. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Pita-Thomas W, Fernández-Martos C, Yunta
M, Maza RM, Navarro-Ruiz R, Lopez-Rodríguez MJ, Reigada D,
Nieto-Sampedro M and Nieto-Diaz M: Gene expression of axon growth
promoting factors in the deer antler. PLoS One. 5:e157062010.
View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Jeon B, Kim S, Lee S, Park P, Sung S, Kim
J and Moon S: Effect of antler growth period on the chemical
composition of velvet antler in sika deer (Cervus nippon). Mamm
Biol. 74:374–380. 2009. View Article : Google Scholar
|
|
8
|
Wang ZY, Shi SR, Shi YJ, Zhang J and Zhou
QY: A comparison of methods to determine amino acid availability of
feedstuffs in cecectomized ganders. Poult Sci. 87:96–100. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Pita-Thomas W, Nieto-Sampedro M, Maza RM
and Nieto-Diaz M: Factors promoting neurite outgrowth during deer
antler regeneration. J Neurosci Res. 88:3034–3047. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Zhou QL, Liu YQ, Wang Y, Guo YJ and Wang
BX: A comparison of chemical composition and bioactivity of
polypeptides from velvet antlers of Cervus nippon Temminck and
Cervus elaphus Linnaeus. Zhongguo Zhong Yao Za Zhi. 26:699–702.
2001.(In Chinese). PubMed/NCBI
|
|
11
|
Wang H, Lin Z, Liu Q, Cai MJ, Xu L and
Zhang XZ: Preparation of velvet antlers small peptides and
stimulating effects on osteosarcoma cell proliferation. Chem J
Chinese Univ. 29:1791–1796. 2008.
|
|
12
|
Pita-Thomas W, Barroso-Garcia G, Moral V,
Hackett AR, Cavalli V and Nieto-Diaz M: Identification of axon
growth promoters in the secretome of the deer antler velvet.
Neuroscience. 340:333–344. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
13
|
Nieto-Diaz M, Pita-Thomas DW,
Munoz-Galdeano T, Martinez-Maza C, Navarro-Ruiz R, Reigada D, Yunta
M, Caballero-Lopez MJ, Nieto-Sampedro M and Martinez-Maza R: Deer
antler innervation and regeneration. Front Biosci (Landmrk Ed).
17:1389–1401. 2012. View
Article : Google Scholar
|
|
14
|
Nieto-Diaz M, Pita-Thomas W, Maza RM,
Yunta M, Lopez-Rodríguez MJ, Navarro-Ruiz R, Reigada D,
Fernandez-Martos CM and Nieto-Sampedro M: Factors promoting axon
growth in the deer antler. Anim Prod Sci. 51:351–354. 2011.
View Article : Google Scholar
|
|
15
|
Wang H, Huang YB, Gao KX, Sun H and Gao
ZL: Preparation and purification of velvet antlers peptides and its
antioxidant activities. Chem J Chinese Univ. 31:2390–2395.
2010.
|
|
16
|
Lee SR, Jeon BT, Kim SJ, Kim MH, Lee SM
and Moon SH: Effects of antler development stage on fatty acid,
vitamin and GAGS contents of velvet antler in spotted deer (Cervus
nippon). Asian Austral J Anim. 20:1546–1550. 2007. View Article : Google Scholar
|
|
17
|
Zhao M, Tao JH, Du LY, Jiang S, Qian DW
and Duan JN: UPLC-Q-TOF/MS-based metabolic profiling comparison of
two major bioactive components and Their metabolites in normal and
CKD rat plasma, urine and feces following oral administration of
fructus corni extract. J Chromatogr Sci. 55:857–865. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Tao JH, Zhao M, Wang DG, Yang C, Chen GT,
Zhao X, Pu XL and Jiang S: UPLC-Q-TOF/MS-based screening and
identification of two major bioactive components and their
metabolites in normal and CKD rat plasma, urine and feces after
oral administration of Rehmannia glutinosa Libosch extract. J
Chromatogr B Analyt Technol Biomed Life Sci. 1001:98–106. 2015.
View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Dong YF, Tang MH, Song H, Li R, Wang C, Ye
H, Qiu N, Zhang Y, Chen L and Wei Y: Characterization of metabolic
profile of honokiol in rat feces using liquid chromatography
coupled with quadrupole time-of-flight tandem mass spectrometry and
(13)C stable isotope labeling. J Chromatogr B Analyt Technol Biomed
Life Sci 953–954. 20–29. 2014. View Article : Google Scholar
|
|
20
|
Liu J, Tang MH, Lai HJ, Dong Y, Xie C, Ye
H, Ma L, Qiu N, Li Y, Cai L and Chen L: Identification of
metabolites of honokiol in rat urine using 13C stable isotope
labeling and liquid chromatography coupled with quadrupole
time-of-flight tandem mass spectrometry. J Chromatogr A.
1295:48–56. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Taguchi R, Houjou T, Nakanishi H, Yamazaki
T, Ishida M, Imagawa M and Shimizu T: Focused lipidomics by tandem
mass spectrometry. J Chromatogr B Analyt Technol Biomed Life Sci.
823:26–36. 2005. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Skwarek LC and Boulianne GL: Great
expectations for PIP: Phosphoinositides as regulators of signaling
during development and disease. Dev Cell. 16:12–20. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Zitouni M, Wewer V, Dörmann P, Abdelly C
and Ben Youssef N: Quadrupole time-of-flight mass spectrometry
analysis of glycerophospholipid molecular species in the two
halophyte seed oils: Eryngium maritimum and Cakile maritima. Food
Chem. 213:319–328. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ejsing CS, Sampaio JL, Surendranath V,
Duchoslav E, Ekroos K, Klemm RW, Simons K and Shevchenko A: Global
analysis of the yeast lipidome by quantitative shotgun mass
spectrometry. Proc Natl Acad Sci USA. 106:2136–2141. 2009.
View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Ejsing CS, Bilgin M and Fabregat A:
Quantitative profiling of long-chain bases by mass tagging and
parallel reaction monitoring. PLoS One. 10:e01448172015. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Zhou R and Li SF: Supercritical carbon
dioxide and co-scosolvent extractions of estradiol and progesterone
from antler velvet. J Food Compos Anal. 22:72–78. 2009. View Article : Google Scholar
|
|
27
|
Zhang FX, Li M, Qiao LR, Yao ZH, Li C,
Shen XY, Wang Y, Yu K, Yao XS and Dai Y: Rapid characterization of
Ziziphi Spinosae Semen by UPLC/Qtof MS with novel informatics
platform and its application in evaluation of two seeds from
Ziziphus species. J Pharm Biomed Anal. 122:59–80. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Qiu S, Yang WZ, Yao CL, Qiu ZD, Shi XJ,
Zhang JX, Hou JJ, Wang QR, Wu WY and Guo DA: Nontargeted
metabolomic analysis and ‘commercial-homophyletic’
comparison-induced biomarkers verification for the systematic
chemical differentiation of five different parts of Panax ginseng.
J Chromatogr A. 1453:78–87. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Li ZH, Zhao WH and Zhou QL: Experimental
study of velvet antler polypeptides against oxidative damage of
osteoarthritis cartilage cells. Zhongguo Gu Shang. 24:245–248.
2011.(In Chinese). PubMed/NCBI
|
|
30
|
Zhang Z, Liu X, Duan L, Li X, Zhang Y and
Zhou Q: The effects of velvet antler polypeptides on the phenotype
and related biological indicators of osteoarthritic rabbit
chondrocytes. Acta Biochim Pol. 58:297–302. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Li YJ, Kim TH, Kwak HB, Lee ZH, Lee SY and
Jhon GJ: Chloroform extract of deer antler inhibits osteoclast
differentiation and bone resorption. J Ethnopharmacol. 113:191–198.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Sui Z, Zhang L, Huo Y and Zhang Y:
Bioactive components of velvet antlers and their pharmacological
properties. J Pharmaceut Biomed. 87:229–240. 2014. View Article : Google Scholar
|
|
33
|
Wu FF, Li HQ, Jin LJ, Li X, Ma Y, You J,
Li S and Xu Y: Deer antler base as a traditional Chinese medicine:
A review of its traditional uses, chemistry and pharmacology. J
Ethnopharmacol. 145:403–415. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Suh SJ, Kim KS, Lee AR, Ha KT, Kim JK, Kim
DS, Lee YC, Kim MS, Kwon DY and Kim CH: Prevention of
collagen-induced arthritis in mice by Cervus korean TEMMINCK var.
mantchuricus Swinhoe. Environ Toxicol Phar. 23:147–153. 2007.
View Article : Google Scholar
|
|
35
|
Kim KS, Choi YH, Kim KH, Lee YC, Kim CH,
Moon SH, Kang SG and Park YG: Protective and anti-arthritic effects
of deer antler aqua-acupuncture (DAA), inhibiting dihydroorotate
dehydrogenase, on phosphate ions-mediated chondrocyte apoptosis and
rat collagen-induced arthritis. Int Immunopharmacol. 4:963–973.
2004. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Fraser A, Haines SR, Stuart EC, Scandlyn
MJ, Alexander A, Somers-Edgar TJ and Rosengren RJ: Deer velvet
supplementation decreases the grade and metastasis of
azoxymethane-induced colon cancer in the male rat. Food Chem
Toxicol. 48:1288–1292. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Zhou R and Li SF: In vitro antioxidant
analysis and characterisation of antler velvet extract. Food Chem.
114:1321–1327. 2009. View Article : Google Scholar
|
|
38
|
Landete Castillejos T, Estevez JA,
Martinez A, Ceacero F, Garcia A and Gallego L: Does chemical
composition of antler bone reflect the physiological effort made to
grow it. Bone. 40:1095–1102. 2007. View Article : Google Scholar : PubMed/NCBI
|