Introduction
In adults, hair follicles (HFs) undergo cycles of
growth, quiescence and regeneration (1). Human dermal papilla cells (DPCs) are
mesenchymal cells located in the hair bulb of HFs, and are
associated with the development and periodic growth of HFs
(2–6). Additionally, DPCs serve an important
role in the formation of HFs (7,8).
Previous studies have demonstrated that the introduction of
exogenous DPCs into the follicles induced the formation of new HFs
in mice (9–12). Furthermore, DPCs have been
investigated as potential cell therapy for hair loss (13,14).
However, previous studies demonstrated that DPCs lose their
stemness and inductive ability during in vitro culture
(7,12).
According to previous reports, DPCs rapidly lose
their inductive ability when cultured in AmnioMAX™-C100 medium
(Thermo Fisher Scientific, Inc.) in vitro, and require the
addition of chemical factors such as bone morphogenic protein 6 and
Wnt3a to maintain their hair-inductive properties (15). The Wnt signaling pathway was
demonstrated to be involved in the maintenance of the intrinsic
properties in hair follicle morphogenesis and cycling of cultured
DPCs in vitro (16). The Wnt
signaling pathway is associated with the repair of several tissues,
including skin and HFs, and was demonstrated to serve a regulatory
role in morphogenesis during embryogenesis, the growth of various
tissues, the maintenance of stem cells and the occurrence of tumors
(17–19). Among the Wnt family, Wnt family
member 10B (WNT10B) was demonstrated to be associated with the
proliferation and the maintenance of DPCs in vitro (20–24).
Moreover, adenovirus-mediated WNT10B overexpression was shown to
induce HF regeneration in vivo (25–29).
WNT10B is one of the earliest and most determinate markers during
the embryonic stage of HF substrate formation (7,11,12) and
promotes the differentiation of epithelial cells and the
development of HFs (30).
Furthermore, WNT10B promoted the development of HFs during
long-term in vitro culture (4,7,11–12,29,31–32).
However, the mechanisms linking WNT10B, HF formation and the
inductive ability of DPCs have not been fully elucidated. As the
downstream target genes of the Wnt signaling pathway, the
β-catenin/TCF/LEF transcription family are expressed in epithelial
and mesenchymal cells at the early morphogenetic stages of HF
development and in post-natal HF stem cells (33). Additionally, β-catenin activity
regulates the regeneration of hair (15,20,33–37).
The aim of the current study was to investigate the
effect of WNT10B on the DPC transcriptome using mRNA sequencing.
Progress in this area of research may facilitate the development of
novel and effective therapeutic approaches for HF regeneration
dysfunction in alopecia.
Materials and methods
Cells and reagents
HFDPCs isolated from human dermis originating from
the lateral scalp were purchased from PromoCell GmbH. Cells were
cultured in DMEM (Invitrogen; Thermo Fisher Scientific, Inc.)
containing 10% fetal bovine serum (FBS; Gibco; Thermo Fisher
Scientific, Inc.; 10099-14-FBS) and 1% Penicillin-Streptomycin
Solution (E607011, Sangon Biotech Co., Ltd.), at 37°C and 5%
CO2. Human recombinant WNT10B protein was purchased from
R&D Systems (cat. no. 7196-WN). The WNT10B treatment conditions
used in the present study were as previously described (38). WNT10B protein was reconstituted in
PBS containing 0.1% bovine serum albumin (BSA; Sangon Biotech Co.,
Ltd.; A602440) to form a solution of 10 µg/ml and used at a final
concentration of 1 µg/ml. The expression of the Wnt target gene
β-catenin in DPCs was increased following treatment with 1 µg/ml
WNT10B protein for 3 days at 37°C, which was in agreement with the
results of a previous study (25)
indicating that the culture conditions were appropriate.
For RNA-sequencing (RNA-seq), the DPCs were divided
into two groups: i) The experimental group, in which DPCs were
cultured in DMEM supplemented with 10% FBS and 1 µg/ml recombinant
WNT10B for 3 days at 37°C; and ii) the control group, in which DPCs
were cultured in DMEM containing 10% FBS and 1 µg/ml BSA for 3 days
at 37°C.
RNA extraction and sequencing
Total RNA from WNT10B-treated and control DPCs was
extracted using TRIzol™ reagent (Invitrogen; Thermo Fisher
Scientific, Inc.) following the manufacturer's instructions. The
total RNA quantity and purity were analyzed using Bioanalyzer 2100
(Agilent Technologies, Inc.) and RNA 6000 Nano LabChip Kit (Agilent
Technologies, Inc.). RNA samples with an RNA integrity number >
7.0 were used for subsequent experimentation. The Illumina TruSeq
RNA Sample Preparation kit (Illumina, Inc.) was used to construct
sequencing libraries. Sequencing was performed using an Illumina
HiSeq 2500 Sequencing system (Illumina, Inc.) by Shanghai
YingBiotech Co., Ltd. Each group was analyzed in triplicate.
Mapping and identification of
differentially expressed genes (DEGs)
Prior to read mapping, the raw sequencing data were
analyzed using FAST-QC, which assessed the nucleotide quality
distribution, PCR duplication rate, position-specific sequencing
quality, k-mer frequency and GC content. The clean reads were
mapped to the human genome (GRCH37). Then the aligned clean read
number was further normalized to reads per kilo of per million
mapped reads (RPKM) with RSEM software (version 1.2.3).
Bioconductor DESeq2 version 1.12.3 (https://www.rdocumentation.org/packages/DESeq2) was
used to identify DEGs using a fold-change (FC) >2 for
significant upregulation or downregulation and a false discovery
rate (FDR) <0.05. A volcano plot was drawn according to the
analysis of the DEGs.
Gene Ontology (GO) term and Kyoto
Encyclopedia of Genes and Genomes (KEGG) pathway analysis
GO (www.geneontology.org) analysis was performed to
identify the biologic implications of the DEGs. Fisher's exact test
was used to identify the significant GO terms with FDR-adjusted
P-values. KEGG pathway analysis was performed to identify
biologically important pathways associated with the DEGs. Fisher's
exact test was used to select the significant pathways based on
P-values (P<0.05) and FDR (FDR<0.27).
Reverse-transcription quantitative PCR
(RT-qPCR)
To verify the results obtained from RNA-seq, seven
highly expressed and enriched DEGs with high FCs (FC >2 or FC
<0.5), including ribosomal protein L17 (RPL17), ribosomal
protein L39 (RPL39), Rho associated coiled-coil containing protein
kinase 2 (ROCK2), cytoplasmic FMR1 interacting protein 2 (CYFIP2),
fibroblast growth factor 14 (FGF14), protein phosphatase 2
regulatory subunit B′ε (PPP2R5E) and vascular endothelial growth
factor B (VEGFB), were selected for RT-qPCR. Total RNA was
extracted from HFDPCs with TRIzol™ reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocol.
Approximately 5 µg total RNA from each sample was used for RT,
which was performed using PrimeScript™ RT Master mix (Takara
Biotechnology Co., Ltd.) following the manufacturer's instructions.
qPCR was subsequently performed using the One Step TB Green™
PrimeScript™ PLUS RT-PCR kit (Takara Biotechnology Co., Ltd) and
the 7500 real-time PCR system (Applied Biosystems; Thermo Fisher
Scientific, Inc.). The qPCR program was: 95°C for 10 min, followed
by 45 cycles of 95°C for 15 sec and 60°C for 60 sec. Gene
expression was quantified using was the 2−ΔΔCq method
(39) and normalized to the
expression of the internal control GAPDH. Each reaction was
performed in triplicate. The primers used for qPCR are presented in
Table I.
 | Table I.Primer sequence for
reverse-transcription quantitative PCR. |
Table I.
Primer sequence for
reverse-transcription quantitative PCR.
|
| Primer
sequences |
|---|
|
|
|
|---|
| Gene | Forward | Reverse |
|---|
| RPL17 |
5′-AGCCTGAGGTGATCTGTGAAAAT-3′ |
5′-CGAGTGTTATTTCGTGGGGTT-3′ |
| RPL39 |
5′-GCCTTCTAAGCTCGTTCTTCCG-3′ |
5′-CGAGCAGCGGAGTCAAGAACA-3′ |
| ROCK2 |
5′-GCAGAAGTGGGTTAGTCGGTTG-3′ |
5′-GGCAGTTAGCTAGGTTTGTTTGG-3′ |
| CYFIP2 |
5′-CCTTAAACCAGCCACTACCTCTC-3′ |
5′-TCTGTATTCTGCACTCATCCGC-3′ |
| FGF14 |
5′-TGCTGGATTGCTTTTCGCC |
5′-GCTGGGGATCAGTTGGGTTCT-3′ |
| PPP2R5E |
5′-TGTCCTCAGCACCAACTACTCCT-3′ |
5′-CAAGATACCTTTTAGCAGCGGC-3′ |
| VEGFB |
5′-GATCCGGTACCCGAGCAGT-3′ |
5′-TTAGGTCTGCATTCACACTGGC-3′ |
Statistical analysis
All data are expressed as mean ± standard error of
the mean. The statistical analyses were performed using GraphPad
Prism software (version 6; GraphPad Software, Inc.). A two-tailed
Student's t-test was used to evaluate statistical significant
differences. P<0.05 was considered to indicate a statistically
significant difference.
Results
Analysis of transcription sequencing
of WNT10B-treated DPCs
In order to identify the differential expression of
mRNA in DPCs following WNT10B treatment, DPCs were divided into the
experimental and control groups. An mRNA library was constructed
for each group and subjected to Illumina mRNA deep sequencing. As
presented in Table II, 95.37% of
51.5 million reads from the WNT10B-treated group and 93.78% of 52.6
million reads from the control group remained following filtering
and quality control.
 | Table II.Analysis of the data generated. |
Table II.
Analysis of the data generated.
| Sample | Total reads | High quality | Reads filter % | Clean reads | Mapped reads | Mapped rate % |
|---|
| DPC-WNT10B-1 |
5.74×107 |
5.44×107 |
9.49×10−1 |
4.05×107 |
3.86×107 |
9.52×10−1 |
| DPC-WNT10B-2 |
5.21×107 |
5.00×107 |
9.58×10−1 |
7.51×107 |
7.16×107 |
9.54×10−1 |
| DPC-WNT10B-3 | 4.5
×107 |
4.30×107 |
9.54×10−1 |
4.13×107 |
3.95×107 |
9.54×10−1 |
| DPC-NC-1 |
4.08×107 |
3.83×107 |
9.39×10−1 |
5.64×107 |
5.45×107 |
9.65×10−1 |
| DPC-NC-2 |
7.52×107 |
7.10×107 |
9.44×10−1 |
5.18×107 |
5.00×107 |
9.64×10−1 |
| DPC-NC-3 |
4.20×107 |
3.93×107 |
9.38×10−1 |
4.45×107 |
4.31×107 |
9.69×10−1 |
Hierarchical clustering of global gene
expression
RPKM analysis was performed to evaluate the
differential mRNA expression between WNT10B-treated and control
DPCs. Using FC >2 and P<0.05 as the cut-off criteria, a total
of 1,525 DEGs were identified, of which 1,074 were upregulated and
451 were downregulated following treatment with WNT10B (Table SI). The 3 WNT10B treatment and 3
control samples were used for hierarchical clustering and to
construct a volcano plot. The WNT10B-treated DPCs were easily
distinguished from the control cells, suggesting that there was a
significant difference in gene expression between the two groups
(Fig. 1).
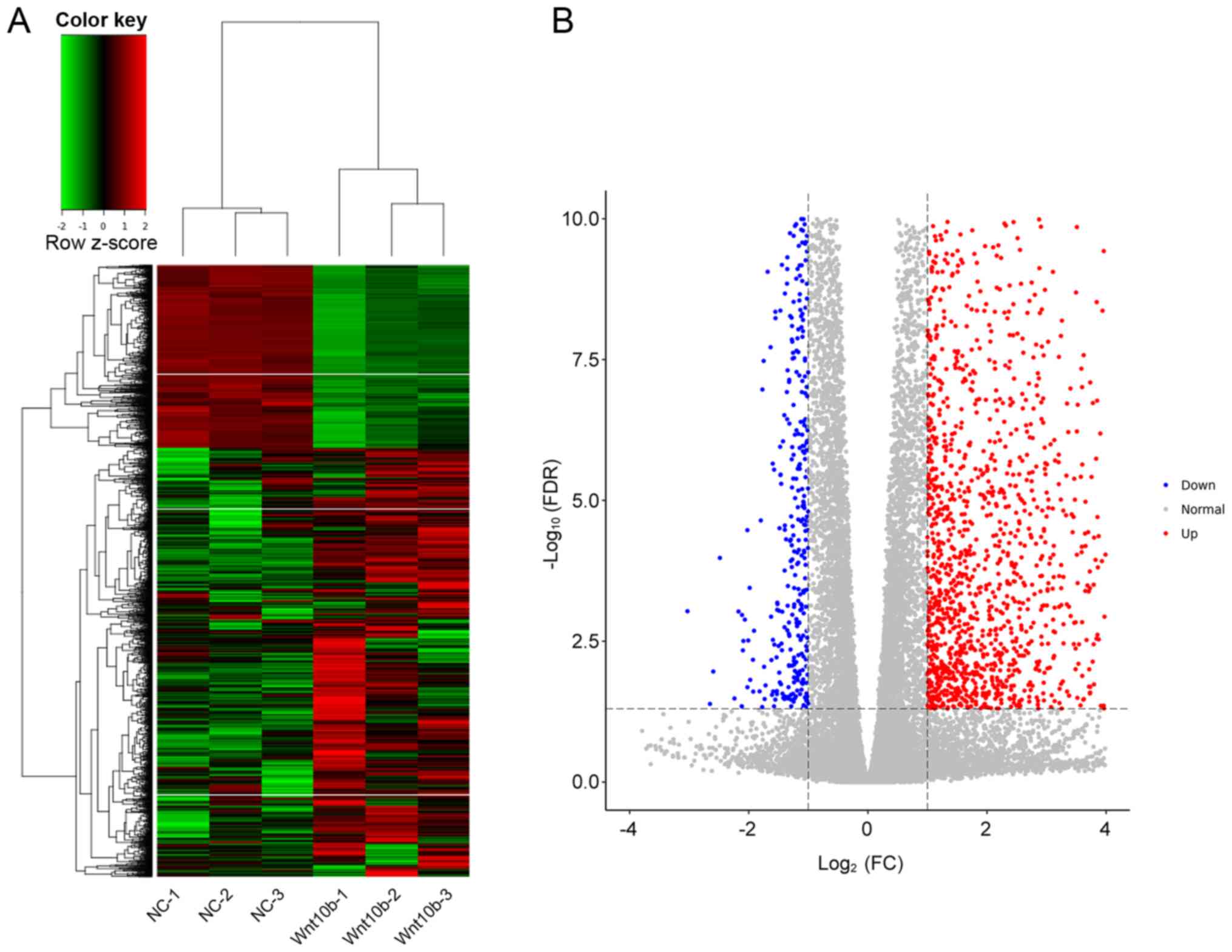 | Figure 1.Gene expression in WNT10B-treated and
control DPCs. (A) The heat map displays gene expression changes in
the 3 WNT10B-treated and 3 control DPCs samples. Red, black, and
green represent increased, unchanged, and decreases expression,
respectively. (B) Volcano plot of upregulated and downregulated
differentially expressed genes between WNT10B-treated and control
DPCs. WNT10B, Wnt family member 10B; DPCs, dermal papilla cells;
FC, fold-change; FDR, false discovery rate; NC, negative
control. |
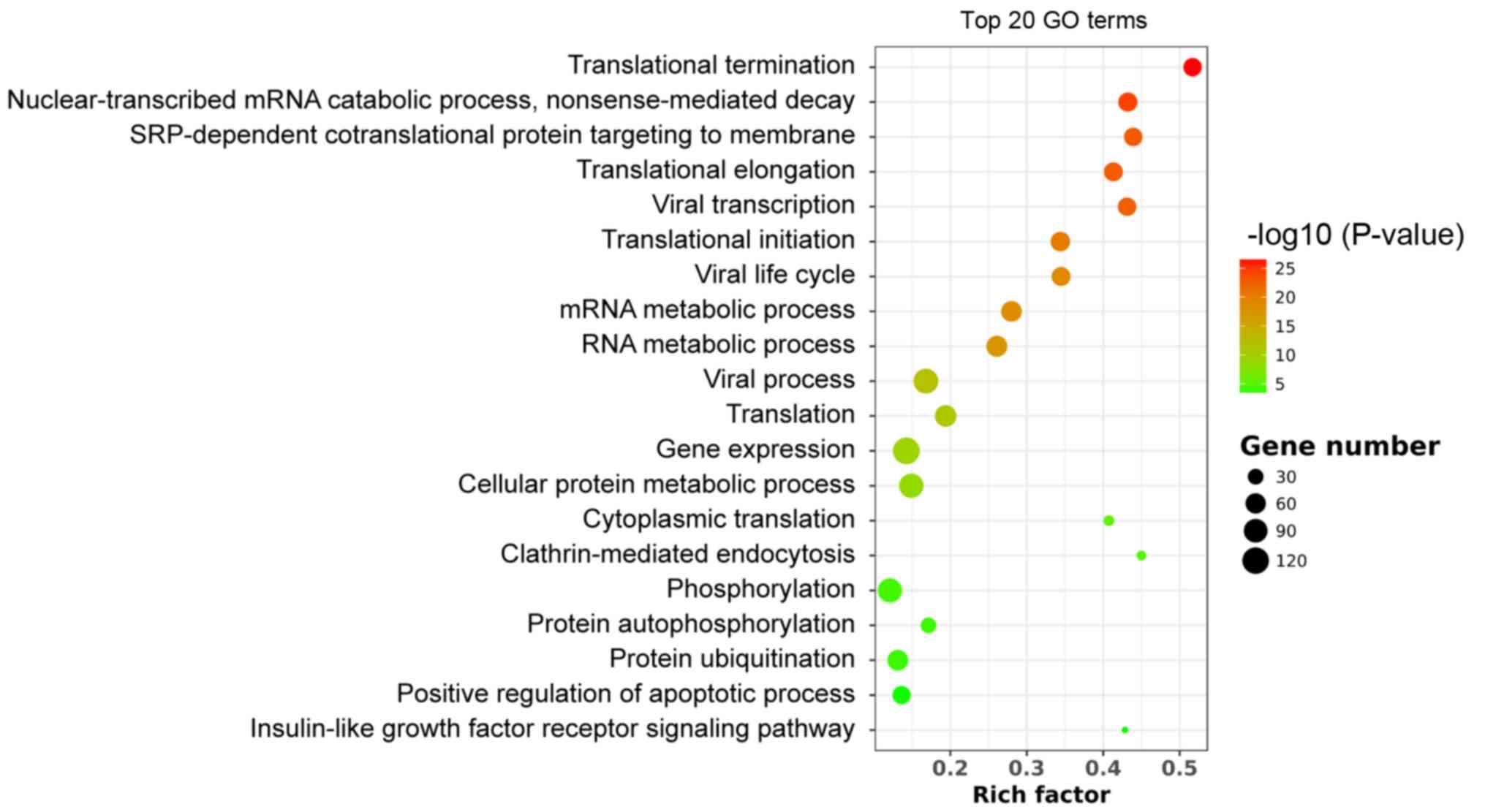
GO analysis
GO enrichment includes biological process, cellular
component (CC) and molecular function (MF). Significantly enriched
GO terms were associated with translational initiation, elongation
and termination (Fig. 2). The
‘structural constituent of ribosome’ in the MF analysis and the
‘cytosolic large ribosomal subunit’ in the CC analysis indicated
that WNT10B treatment may influence RNA translation and protein
synthesis in DPCs, which subsequently affect HF induction. Several
upregulated genes were enriched in the term ‘stem cell
maintenance’. These included dicer 1 ribonuclease III, APC
regulator of WNT signaling pathway, NIPBL cohesin loading factor,
notch receptor 2, cell division cycle 73, replication timing
regulatory factor 1, mediator complex subunit 12 and bone
morphogenetic protein receptor type 1A (Table SIII).
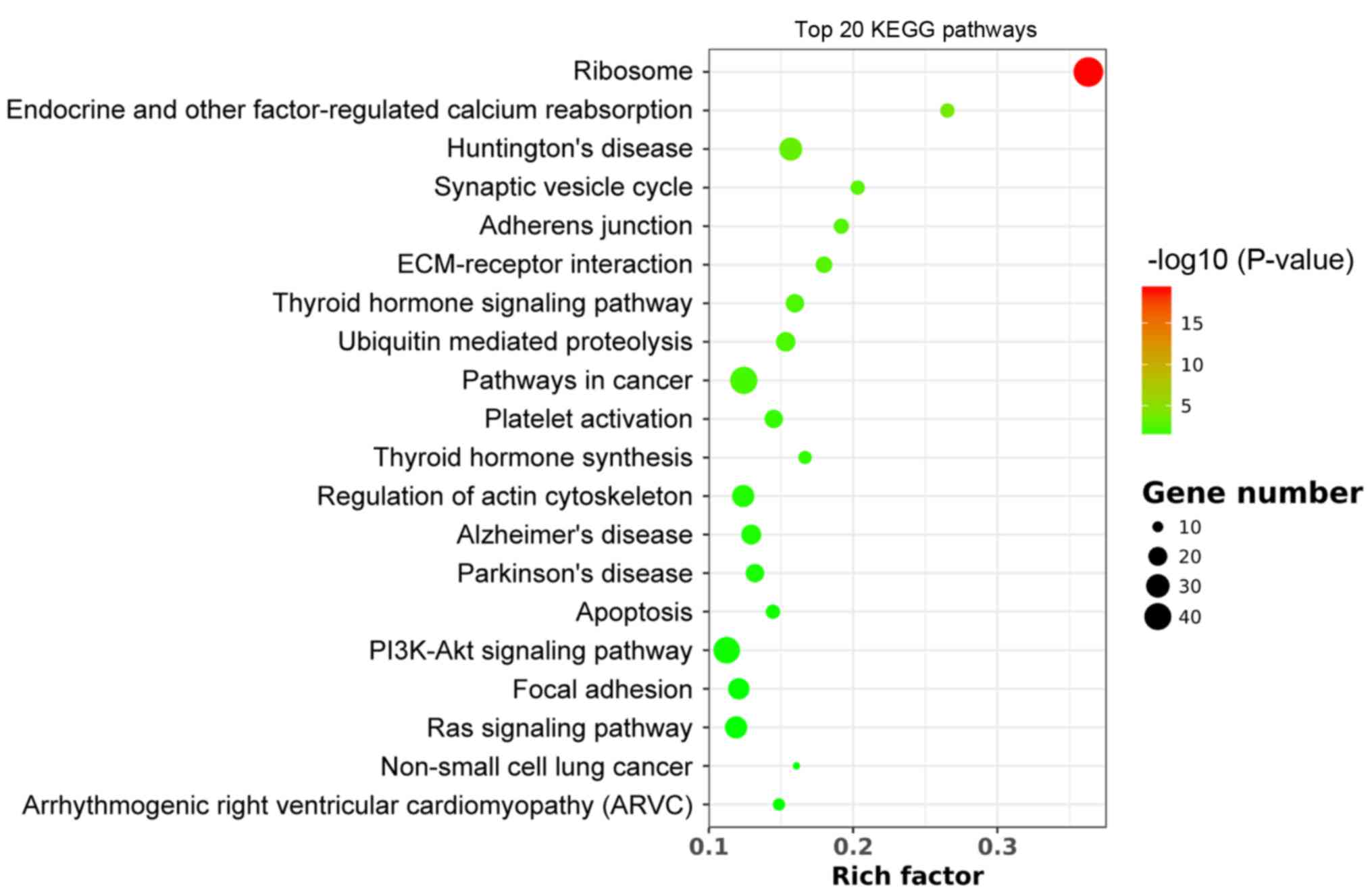
KEGG pathway analysis
KEGG pathway analysis revealed that DEGs were
enriched in a total of 21 pathways following WNT10B treatment.
Among those pathways, the ‘ribosome’ was identified as the most
enriched (Fig. 3), suggesting that
WNT10B may influence protein synthesis, which was consistent with
the GO analysis.
In addition, KEGG pathway analysis revealed that the
upregulated DEGs were significantly enriched in the ‘PI3K-Akt
signaling pathway’ (Table SII).
Therefore, WNT10B may activate the signaling PI3K/Akt pathway and
maintain the HF inductive proprieties of DPCs.
Validation of RNA-seq results
RT-qPCR was performed to verify the expression
levels of the DEGs. DEGs with high FPKM and high FCs (FC>2 or
FC<0.5) were selected, including RPL17, RPL39, ROCK2, CYFIP2,
FGF14, PPP2R5E and VEGFB. The RT-qPCR results were highly
consistent with RNA-seq data (Fig.
4).
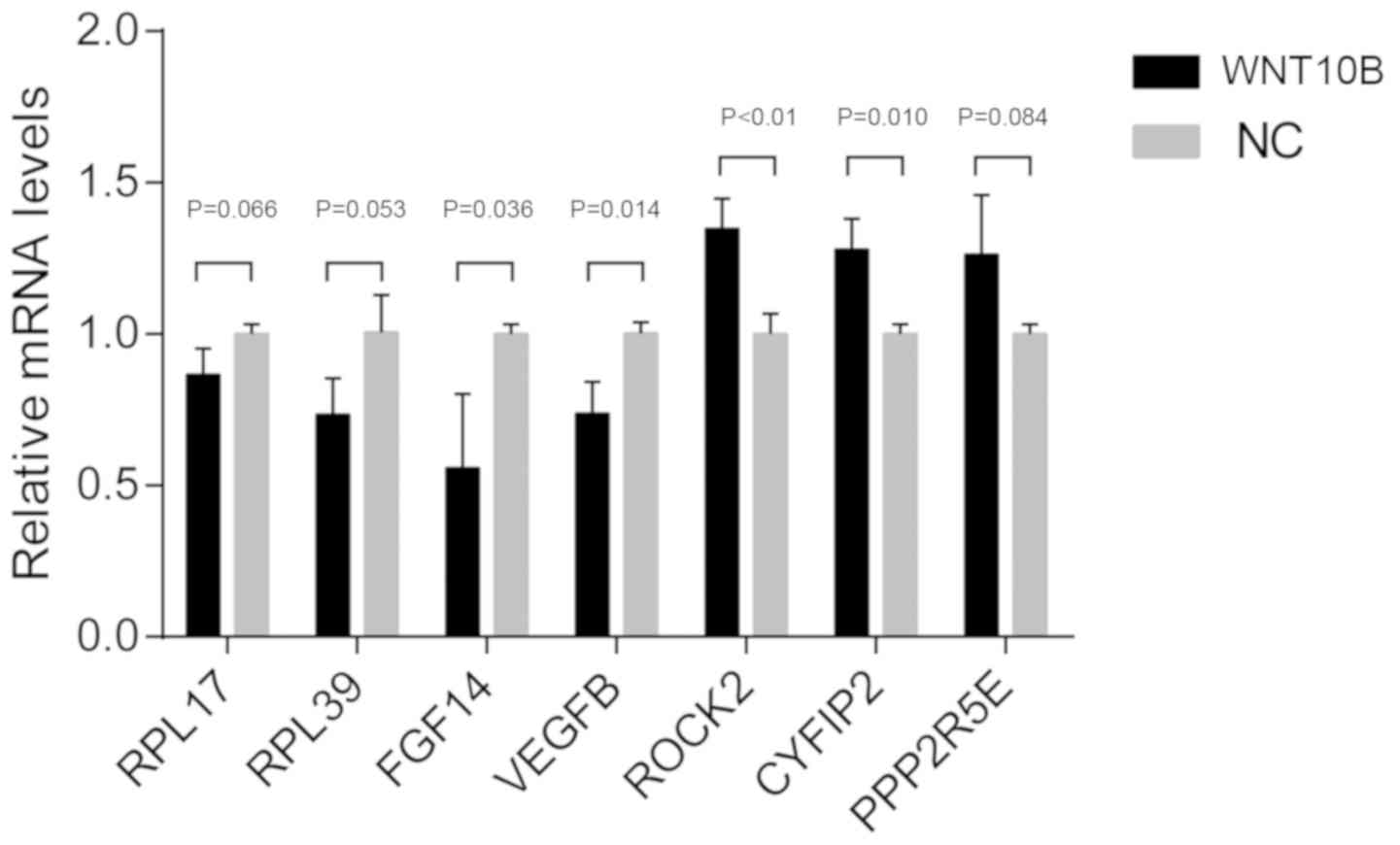 | Figure 4.RT-qPCR to confirm RNA sequencing
results from WNT10B-treated and control dermal papilla cells. A
total of 7 genes were selected for the qRT-PCR analysis, including
RPL17, RPL39, ROCK2, CYFIP2, FGF14, PPP2R5E and VEGFB. The data are
presented as the mean ± SEM of three independent experiments.
RT-qPCR, reverse-transcription quantitative PCR; WNT10B, Wnt family
member 10B; RPL17, ribosomal protein L17; RPL39, ribosomal protein
L39; ROCK2, Rho associated coiled-coil containing protein kinase 2;
CYFIP2, cytoplasmic FMR1 interacting protein 2; FGF14, fibroblast
growth factor 14; PPP2R5E, protein phosphatase 2 regulatory subunit
B′ε; VEGFB, vascular endothelial growth factor B; NC, negative
control. |
Discussion
Human HFDPCs are a type of mesenchymal cell isolated
from the DPCs of HFs (5,6,40–41).
According to reports, DPCs serve important roles in the
dermal-epidermal interactions regulating hair regeneration and in
the hair growth cycle (5,42–43).
DPCs have been investigated as potential cell therapy due to their
HF inductive ability (44,45). However, this ability rapidly
diminishes when DPCs are cultured in vitro (8,12,46),
which limits the potential for application in alopecia therapy.
The Wnt signaling pathway is the most important
regulator of DPC behavior (47,48).
Previous studies have revealed that WNT10B serves a vital role in
the proliferation and maintenance of DPCs and promotes the growth
of HFs and the differentiation of mouse HF melanocytes (7,11,18,20).
However, the mechanisms underlying these processes remain unclear.
The present study used GO term and KEGG pathway analyses to explore
the potential mechanisms based on mRNA-seq data. Results revealed
1,073 upregulated and 451 downregulated DEGs between WNT10B-treated
and control DPCs in vitro. GO and KEGG results suggested
that WNT10B significantly downregulated all the three steps
(initiation, elongation and termination) of the translation process
and as well as the ribosome pathway In the majority of adults,
progenitor cells such as hematopoietic progenitor cell are
quiescent and undergo lower levels of protein synthesis (49), suggesting that the level of protein
synthesis may be related to stemness and properties of HFDPCs.
Additionally, KEGG pathway analysis revealed that WNT10B
upregulated the PI3K/Akt signaling pathway, which is known to
regulate various cellular processes, including proliferation,
metabolism, transcription, protein synthesis, growth and survival
(49). A previous study revealed
that deletion of the PI3K/Akt signaling pathway antagonist
phosphatase and tensin homolog increased protein synthesis and
depleted hematopoietic stem cells (50), suggesting that translational
regulation and lower rates of ribosome biogenesis may maintain the
properties of HFDPCs. The results of the present study suggested
that WNT10B treatment may downregulate the protein synthesis
rate.
The PI3K/Akt signaling pathway promotes the
development, proliferation and differentiation of adult stem cells,
particularly neural stem cells (51). As the downstream effector of Wnt
signaling pathway, β-catenin regulates the proliferation of HFSCs
through the PI3K/Akt signaling pathway (52,53).
Furthermore, the activation of the PI3K/Akt signaling pathway
triggers the expression of growth factors and promotes DPC-mediated
hair growth (54–56). The activation of the PI3K/Akt
signaling pathway may therefore be essential for DPCs to perform
their normal functions.
It has previously been reported that Wnt signaling
increases MTOR complex 1 (mTORC1) activity. mTORC1 plays a key role
in protein synthesis and positively regulates cellular metabolism,
ATP production and lipid synthesis (57–59). Wnt
signaling may therefore promote protein synthesis by upregulating
mTORC1. However, the regulation of the Wnt signaling pathway is an
area requiring further study and may involve other signaling
pathways (60,61). Moreover, the Wnt signaling pathway
maintains stem cell pluripotency and balances progenitor
self-renewal and differentiation (62). A previous study identified several
Wnt proteins that were expressed in mouse embryonic and postnatal
skin, including WNT3, WNT3A WNT4, WNT5A, WNT6, WNT7A, WNT7B,
WNT10A, WNT10B and WNT16. The earliest and most highly expressed
Wnt ligand in mouse hair follicle development and hair cycle
induction was shown to be WNT10B (47). In agreement with the GO term
enrichment results in the present study, WNT10B is involved in
signaling pathways controlling stemness, pluripotency and cell fate
(63). Recent studies have shown
that embryonic and somatic stem cells rely on low translation rates
to maintain an undifferentiated state. By contrast, differentiation
requires increased protein synthesis. The transition from
self-renewal to differentiation relies on enhanced ribosome
biogenesis accompanied by increased protein synthesis (64). Therefore, the regulation of protein
synthesis is important for cellular differentiation. WNT10B may
maintain the stemness of human dermal papilla cells via decreasing
the rate of protein synthesis.
The results obtained in the current study may aid in
the elucidation of mechanisms of DPC activation and contribute to
the development of therapies to treat dysfunction of HF
regeneration in diseases such as alopecia.
Supplementary Material
Supporting Data
Supporting Data
Supporting Data
Acknowledgements
Not applicable.
Funding
The present study was supported by the National
Natural Science Foundation of China (grant nos. 81573057, 81703135
and 81472889).
Authors' contributions
QZ, YS and HC designed the experiments. QZ, YS, QLZ,
and RH carried out the experiments and interpreted the data. QZ and
YS prepared the sequenced samples and performed the qPCR. HC
supervised all experiments. All the authors contributed to the
manuscript.
Availability of data and materials
The datasets used and analyzed during the present
study are available from the corresponding author on reasonable
request.
Ethical approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ma X, Tian Y, Song Y, Shi J, Xu J, Xiong
K, Li J, Xu W, Zhao Y, Shuai J, et al: Msi2 maintains quiescent
state of hair follicle stem cells by directly repressing the Hh
signaling pathway. J Invest Dermatol. 137:1015–1024. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
2
|
Harel S, Higgins CA, Cerise JE, Dai Z,
Chen JC, Clynes R and Christiano AM: Pharmacologic inhibition of
JAK-STAT signaling promotes hair growth. Sci Adv. 1:e15009732015.
View Article : Google Scholar : PubMed/NCBI
|
|
3
|
Jahoda CA and Reynolds AJ: Hair follicle
dermal sheath cells: unsung participants in wound healing. Lancet.
358:1445–1448. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
4
|
Yu Z, Jiang K, Xu Z, Huang H, Qian N, Lu
Z, Chen D, Di R, Yuan T, Du Z, et al: Hoxc-dependent mesenchymal
niche heterogeneity drives regional hair follicle regeneration.
Cell Stem Cell. 23:487–500.e486. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
5
|
Driskell RR, Clavel C, Rendl M and Watt
FM: Hair follicle dermal papilla cells at a glance. J Cell Sci.
124:1179–1182. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
6
|
Balañá ME, Charreau HE and Leirós GJ:
Epidermal stem cells and skin tissue engineering in hair follicle
regeneration. World J Stem Cells. 7:711–727. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
7
|
Ouji Y, Yoshikawa M, Shiroi A and Ishizaka
S: Wnt-10b promotes differentiation of skin epithelial cells in
vitro. Biochem Biophys Res Commun. 342:28–35. 2006. View Article : Google Scholar : PubMed/NCBI
|
|
8
|
Driskell RR, Giangreco A, Jensen KB,
Mulder KW and Watt FM: Sox2-positive dermal papilla cells specify
hair follicle type in mammalian epidermis. Development.
136:2815–2823. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
9
|
Abaci HE, Coffman A, Doucet Y, Chen J,
Jacków J, Wang E, Guo Z, Shin JU, Jahoda CA and Christiano AM:
Tissue engineering of human hair follicles using a biomimetic
developmental approach. Nat Commun. 9:53012018. View Article : Google Scholar : PubMed/NCBI
|
|
10
|
Osada A and Kobayashi K: Appearance of
hair follicle-inducible mesenchymal cells in the rat embryo. Dev
Growth Differ. 42:19–27. 2000. View Article : Google Scholar : PubMed/NCBI
|
|
11
|
Ouji Y, Yoshikawa M, Moriya K and Ishizaka
S: Effects of Wnt-10b on hair shaft growth in hair follicle
cultures. Biochem Biophys Res Commun. 359:516–522. 2007. View Article : Google Scholar : PubMed/NCBI
|
|
12
|
Jahoda CA, Horne KA and Oliver RF:
Induction of hair growth by implantation of cultured dermal papilla
cells. Nature. 311:560–562. 1984. View
Article : Google Scholar : PubMed/NCBI
|
|
13
|
Jeong KH, Joo HJ, Kim JE, Park YM and Kang
H: Effect of mycophenolic acid on proliferation of dermal papilla
cells and induction of anagen hair follicles. Clin Exp Dermatol.
40:894–902. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
14
|
Park S, Shin WS and Ho J: Fructus panax
ginseng extract promotes hair regeneration in C57BL/6 mice. J
Ethnopharmacol. 138:340–344. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
15
|
Rendl M, Polak L and Fuchs E: BMP
signaling in dermal papilla cells is required for their hair
follicle-inductive properties. Genes Dev. 22:543–557. 2008.
View Article : Google Scholar : PubMed/NCBI
|
|
16
|
Ohyama M, Kobayashi T, Sasaki T, Shimizu A
and Amagai M: Restoration of the intrinsic properties of human
dermal papilla in vitro. J Cell Sci. 125:4114–4125. 2012.
View Article : Google Scholar : PubMed/NCBI
|
|
17
|
Galluzzi L, Spranger S, Fuchs E and
Lopez-Soto A: WNT signaling in cancer immunosurveillance. Trends
Cell Biol. 29:44–65. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
18
|
Nusse R and Clevers H: Wnt/β-catenin
signaling, disease, and emerging therapeutic modalities. Cell.
169:985–999. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
19
|
Siebel C and Lendahl U: Notch signaling in
development, tissue homeostasis, and disease. Physiol Rev.
97:1235–1294. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
20
|
Ouji Y, Ishizaka S and Yoshikawa M: Dermal
papilla cells serially cultured with Wnt-10b sustain their hair
follicle induction activity after transplantation into nude mice.
Cell Transplant. 21:2313–2324. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
21
|
Soma T, Fujiwara S, Shirakata Y, Hashimoto
K and Kishimoto J: Hair-inducing ability of human dermal papilla
cells cultured under Wnt/β-catenin signalling activation. Exp
Dermatol. 21:307–309. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
22
|
Kitagawa T, Matsuda K, Inui S, Takenaka H,
Katoh N, Itami S, Kishimoto S and Kawata M: Keratinocyte growth
inhibition through the modification of Wnt signaling by androgen in
balding dermal papilla cells. J Clin Endocrinol Metab.
94:1288–1294. 2009. View Article : Google Scholar : PubMed/NCBI
|
|
23
|
Shin H, Kwack MH, Shin SH, Oh JW, Kang BM,
Kim AA, Kim J, Kim MK, Kim JC and Sung YK: Identification of
transcriptional targets of Wnt/beta-catenin signaling in dermal
papilla cells of human scalp hair follicles: EP2 is a novel
transcriptional target of Wnt3a. J Dermatol Sci. 58:91–96. 2010.
View Article : Google Scholar : PubMed/NCBI
|
|
24
|
Ouji Y, Nakamura-Uchiyama F and Yoshikawa
M: Canonical Wnts, specifically Wnt-10b, show ability to maintain
dermal papilla cells. Biochem Biophys Res Commun. 438:493–499.
2013. View Article : Google Scholar : PubMed/NCBI
|
|
25
|
Li YH, Zhang K, Ye JX, Lian XH and Yang T:
Wnt10b promotes growth of hair follicles via a canonical Wnt
signalling pathway. Clin Exp Dermatol. 36:534–540. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
26
|
Lei M, Guo H, Qiu W, Lai X, Yang T,
Widelitz RB, Chuong CM, Lian X and Yang L: Modulating hair follicle
size with Wnt10b/DKK1 during hair regeneration. Exp Dermatol.
23:407–413. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
27
|
Ye J, Yang T, Guo H, Tang Y, Deng F, Li Y,
Xing Y, Yang L and Yang K: Wnt10b promotes differentiation of mouse
hair follicle melanocytes. Int J Med Sci. 10:691–698. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
28
|
Li YH, Zhang K, Yang K, Ye JX, Xing YZ,
Guo HY, Deng F, Lian XH and Yang T: Adenovirus-mediated Wnt10b
overexpression induces hair follicle regeneration. J Invest
Dermatol. 133:42–48. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
29
|
Zhang Y, Xing Y, Guo H, Ma X and Li Y:
Immunohistochemical study of hair follicle stem cells in
regenerated hair follicles induced by Wnt10b. Int J Med Sci.
13:765–771. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
30
|
Bai X, Lei M, Shi J, Yu Y, Qiu W, Lai X,
Liu Y, Yang T, Yang L, Widelitz RB, et al: Roles of gasdermina3 in
catagen-telogen transition during hair cycling. J Invest Dermatol.
135:2162–2172. 2015. View Article : Google Scholar : PubMed/NCBI
|
|
31
|
Carroll LS and Capecchi MR: Hoxc8
initiates an ectopic mammary program by regulating Fgf10 and Tbx3
expression and Wnt/β-catenin signaling. Development. 142:4056–4067.
2015. View Article : Google Scholar : PubMed/NCBI
|
|
32
|
Zhang T, Liu L, Fan J, Tian J, Gan C, Yang
Z, Jiao H, Han B and Liu Z: Low-level laser treatment stimulates
hair growth via upregulating Wnt10b and β-catenin expression in
C3H/HeJ mice. Lasers Med Sci. 32:1189–1195. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
33
|
Enshell-Seijffers D, Lindon C, Wu E,
Taketo MM and Morgan BA: Beta-catenin activity in the dermal
papilla of the hair follicle regulates pigment-type switching. Proc
Natl Acad Sci USA. 107:21564–21569. 2010. View Article : Google Scholar : PubMed/NCBI
|
|
34
|
Reddy S, Andl T, Bagasra A, Lu MM, Epstein
DJ, Morrisey EE and Millar SE: Characterization of Wnt gene
expression in developing and postnatal hair follicles and
identification of Wnt5a as a target of sonic hedgehog in hair
follicle morphogenesis. Mech Dev. 107:69–82. 2001. View Article : Google Scholar : PubMed/NCBI
|
|
35
|
Matheson J, Bühnemann C, Carter EJ, Barnes
D, Hoppe HJ, Hughes J, Cobbold S, Harper J, Morreau H, Surakhy M
and Hassan AB: Epithelial-mesenchymal transition and nuclear
β-catenin induced by conditional intestinal disruption of Cdh1 with
Apc is E-cadherin EC1 domain dependent. Oncotarget. 7:69883–69902.
2016. View Article : Google Scholar : PubMed/NCBI
|
|
36
|
Cheon SS, Cheah AY, Turley S, Nadesan P,
Poon R, Clevers H and Alman BA: beta-Catenin stabilization
dysregulates mesenchymal cell proliferation, motility, and
invasiveness and causes aggressive fibromatosis and hyperplastic
cutaneous wounds. Proc Natl Acad Sci USA. 99:6973–6978. 2002.
View Article : Google Scholar : PubMed/NCBI
|
|
37
|
Lin GL and Hankenson KD: Integration of
BMP, Wnt, and notch signaling pathways in osteoblast
differentiation. J Cell Biochem. 112:3491–3501. 2011. View Article : Google Scholar : PubMed/NCBI
|
|
38
|
Ouji Y, Yoshikawa M, Moriya K, Nishiofuku
M, Matsuda R and Ishizaka S: Wnt-10b, uniquely among Wnts, promotes
epithelial differentiation and shaft growth. Biochem Biophys Res
Commun. 367:299–304. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
39
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. methods. 25:402–408. 2001.
View Article : Google Scholar : PubMed/NCBI
|
|
40
|
Nilforoushzadeh MA, Zare M, Zarrintaj P,
Alizadeh E, Taghiabadi E, Heidari-Kharaji M, Amirkhani MA, Saeb MR
and Mozafari M: Engineering the niche for hair regeneration - A
critical review. Nanomedicine. 15:70–85. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
41
|
El Agha E, Kramann R, Schneider RK, Li X,
Seeger W, Humphreys BD and Bellusci S: Mesenchymal stem cells in
fibrotic disease. Cell Stem Cell. 21:166–177. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
42
|
Madaan A, Verma R, Singh AT and Jaggi M:
Review of hair follicle dermal papilla cells as in vitro screening
model for hair growth. Int J Cosmet Sci. 40:429–450. 2018.
View Article : Google Scholar : PubMed/NCBI
|
|
43
|
Kim OY, Cha HJ, Ahn KJ, An IS, An S and
Bae S: Identification of microRNAs involved in growth arrest and
cell death in hydrogen peroxide-treated human dermal papilla cells.
Mol Med Rep. 10:145–154. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
44
|
Kim J, Shin JY, Choi YH, Jang M, Nam YJ,
Lee SY, Jeon J, Jin MH and Lee S: Hair growth promoting effect of
hottuynia cordata extract in cultured human hair follicle dermal
papilla cells. Biol Pharm Bull. 42:1665–1673. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
45
|
Zhang X, Xiao S, Liu B, Miao Y and Hu Z:
Use of extracellular matrix hydrogel from human placenta to restore
hair-inductive potential of dermal papilla cells. Regen Med.
2019.(Epub ahead of print). View Article : Google Scholar
|
|
46
|
Shim JH, Lee TR and Shin DW: Novel in
vitro culture condition improves the stemness of human dermal
stem/progenitor cells. Mol Cells. 36:556–563. 2013. View Article : Google Scholar : PubMed/NCBI
|
|
47
|
Jansson L, Kim GS and Cheng AG: Making
sense of Wnt signaling-linking hair cell regeneration to
development. Front Cell Neurosci. 9:662015. View Article : Google Scholar : PubMed/NCBI
|
|
48
|
Myung PS, Takeo M, Ito M and Atit RP:
Epithelial Wnt ligand secretion is required for adult hair follicle
growth and regeneration. J Invest Dermatol. 133:31–41. 2013.
View Article : Google Scholar : PubMed/NCBI
|
|
49
|
Buszczak M, Signer RA and Morrison SJ:
Cellular differences in protein synthesis regulate tissue
homeostasis. Cell. 159:242–251. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
50
|
Signer RA, Magee JA, Salic A and Morrison
SJ: Haematopoietic stem cells require a highly regulated protein
synthesis rate. Nature. 509:49–54. 2014. View Article : Google Scholar : PubMed/NCBI
|
|
51
|
Peltier J, O'Neill A and Schaffer DV:
PI3K/Akt and CREB regulate adult neural hippocampal progenitor
proliferation and differentiation. Dev Neurobiol. 67:1348–1361.
2007. View Article : Google Scholar : PubMed/NCBI
|
|
52
|
Du KT, Deng JQ, He XG, Liu ZP, Peng C and
Zhang MS: MiR-214 regulates the human hair follicle stem cell
proliferation and differentiation by targeting EZH2 and
Wnt/β-catenin signaling way in vitro. Tissue Eng Regen Med.
15:341–350. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
53
|
Gentile P, Scioli MG, Bielli A, De Angelis
B, De Sio C, De Fazio D, Ceccarelli G, Trivisonno A, Orlandi A,
Cervelli V and Garcovich S: Platelet-rich plasma and micrografts
enriched with autologous human follicle mesenchymal stem cells
improve hair re-growth in androgenetic alopecia. Biomolecular
pathway analysis and clinical evaluation. Biomedicines. 7:E272019.
View Article : Google Scholar : PubMed/NCBI
|
|
54
|
Woo H, Lee S, Kim S, Park D and Jung E:
Effect of sinapic acid on hair growth promoting in human hair
follicle dermal papilla cells via Akt activation. Arch Dermatol
Res. 309:381–388. 2017. View Article : Google Scholar : PubMed/NCBI
|
|
55
|
Feutz AC, Barrandon Y and Monard D:
Control of thrombin signaling through PI3K is a mechanism
underlying plasticity between hair follicle dermal sheath and
papilla cells. J Cell Sci. 121:1435–1443. 2008. View Article : Google Scholar : PubMed/NCBI
|
|
56
|
Zhang H, Nan W, Wang S, Zhang T, Si H,
Yang F and Li G: Epidermal growth factor promotes proliferation and
migration of follicular outer root sheath cells via wnt/β-catenin
signaling. Cell Physiol Biochem. 39:360–370. 2016. View Article : Google Scholar : PubMed/NCBI
|
|
57
|
Yin G, Liang Y, Wang Y, Yang Y, Yang M,
Cen XM and Xie QB: mTOR complex 1 signalling regulates the balance
between lipid synthesis and oxidation in hypoxia lymphocytes.
Biosci Rep. 37:BSR201604792017. View Article : Google Scholar : PubMed/NCBI
|
|
58
|
Chen JJ and Zhang S: Heme-regulated
eIF2alpha kinase in erythropoiesis and hemoglobinopthies. Blood.
20190019152019.
|
|
59
|
Lee G, Zheng Y, Cho S, Jang C, England C,
Dempsey JM, Yu Y, Liu X, He L, Cavaliere PM, et al:
Post-transcriptional regulation of de novo lipogenesis by
mTORC1-S6K1-SRPK2 signaling. Cell. 171:1545–1558. e1518. 2017.
View Article : Google Scholar : PubMed/NCBI
|
|
60
|
Kumar D, Nitzan E and Kalcheim C: YAP
promotes neural crest emigration through interactions with BMP and
Wnt activities. Cell Commun Signal. 17:692019. View Article : Google Scholar : PubMed/NCBI
|
|
61
|
Massey J, Liu Y, Alvarenga O, Saez T,
Schmerer M and Warmflash A: Synergy with TGFβ ligands switches WNT
pathway dynamics from transient to sustained during human
pluripotent cell differentiation. Proc Natl Acad Sci USA.
116:4989–4998. 2019. View Article : Google Scholar : PubMed/NCBI
|
|
62
|
Kahn M: Wnt signaling in stem cells and
cancer stem cells: A tale of two coactivators. Prog Mol Biol Transl
Sci. 153:209–244. 2018. View Article : Google Scholar : PubMed/NCBI
|
|
63
|
Wend P, Wend K, Krum SA and
Miranda-Carboni GA: The role of WNT10B in physiology and disease.
Acta Physiol (Oxf). 204:34–51. 2012. View Article : Google Scholar : PubMed/NCBI
|
|
64
|
Sanchez CG, Teixeira FK, Czech B, Preall
JB, Zamparini AL, Seifert JR, Malone CD, Hannon GJ and Lehmann R:
Regulation of ribosome biogenesis and protein synthesis controls
germline stem cell differentiation. Cell Stem Cell. 18:276–290.
2016. View Article : Google Scholar : PubMed/NCBI
|