Introduction
Oral squamous cell carcinoma (OSCC) is the commonest
type of oral tumor, accounting for 90% of all oral malignancies and
38% of head and neck tumors; >500,000 patients are newly
diagnosed with OSCC every year worldwide (1). Surgical resection, chemo-radiotherapy
and the latest emerging molecular targeted therapy are common
treatments for OSCC, but the 5-year survival rate of OSCC patients
remains <50% due to the high degree of malignancy, postoperative
recurrence and metastasis (2,3). Finding
new effective therapeutic targets and exploring molecular
mechanisms are particularly important for improving the prognosis
of patients with OSCC.
In recent years, with the widespread development of
second-generation sequencing technology, the importance of long
non-coding RNAs (lncRNAs) in many human diseases has been
recognized by researchers. Accumulating research data indicate that
lncRNAs participate in the occurrence and development of tumors at
several levels (4,5). Some lncRNAs have been shown to be
abnormally expressed in OSCC and participate in the development of
OSCC (6). lncRNAs including FTH1P3,
GTF2IRD2P1 and PDIA3F may be responsible for OSCC progression and
metastasis by targeting the matrix metallopeptidase family and
interleukin 8(6). Metastasis
associated lung adenocarcinoma transcript 1 (MALAT1) has been found
to be upregulated in OSCC in a recent study, and the growth of OSCC
in nude mice is greatly inhibited by downregulation of
MALAT1(7).
The present study detected the expression of
MCM3AP-AS1 in OSCC and investigated the underlying mechanisms of
MCM3AP-AS1 in OSCC. It identified a potential effective therapeutic
target for OSCC and laid a new theoretical foundation for the
targeted therapy of OSCC.
Materials and methods
Patients and tissues
Between July 2015 and April 2018, 36 patients (age,
47-54 years; mean age, 55±7.3 years) with OSCC volunteered to join
the study and all patients provided written informed consent. The
present study was approved by the Ethics Committee of Zaozhuang
Municipal Hospital.
All patients had undergone surgical resection, and
information on their clinicopathological features, such as age,
sex, clinical stage and lymph node metastasis, was recorded. During
surgery, normal tissues and tumor tissues were obtained and
immediately stored in liquid nitrogen (Table I).
 | Table IExpression of MCM3AP-AS1 and its
relationship with the clinicopathological features of patients with
oral squamous cell carcinoma. |
Table I
Expression of MCM3AP-AS1 and its
relationship with the clinicopathological features of patients with
oral squamous cell carcinoma.
| Parameter | Low (n=19) | High (n=17) | P-value |
|---|
| Age, years | | | 0.7388 |
|
≥60 | 11 | 8 | |
|
<60 | 8 | 9 | |
| Sex | | | 0.7317 |
|
Male | 12 | 12 | |
|
Female | 7 | 5 | |
| Clinical stage | | | 0.0228a |
|
I/II | 12 | 4 | |
|
III/IV | 7 | 13 | |
| Lymph node
metastasis | | | 0.0.0058b |
|
No | 11 | 2 | |
|
Yes | 8 | 15 | |
Cell lines and culture
In the present study, human oral keratinocytes (HOK
cells; Shanghai Key Laboratory of Stomatology) and OSCC cells
(SCC-9, SCC-15 and CAL-27; American Type Culture Collection) were
maintained in sterile culture flasks containing 2.5 ml complete
DMEM. Penicillin (1%), streptomycin (1%) and fetal bovine serum
(FBS; 10%; Gibco; Thermo Fisher Scientific, Inc.) were dissolved in
the DMEM to produce complete DMEM. The flasks were placed in a 5%
CO2 incubator at 37˚C and 95% humidity. The complete
DMEM was changed every 3 days.
Cell transfection
SCC-15 and CAL-27 cells suspended in serum-free DMEM
(2x105 cells/ml) were subjected to transfection with
small interfering (si)-MCM3AP-AS1 (100 nM; si-MCM3AP-AS1 group;
5'-GCTGCTAATGGCAACACTGA-3') and negative control (NC; si-NC group;
5'-TTCTCCGAACGTGTCACGTTT-3'). siRNA was purchased from Guangzhou
Geneseed Biotech Co., Ltd. miR-363-5p mimic (100 nM;
5'-UUUAACGUAGCACUAGGUGGGC-3'), miR-363-5p inhibitor (100 nM
5'-AAAUUGCAUCGUGAUCCACCCG-3') and the negative control
(5'-ACAUCUGCGUAAGAUUCGAGUCUA-3') were provided by RiboBio Co., Ltd.
All transfections were performed using Lipofectamine®
2000 (Thermo Fisher Scientific, Inc.). Cells successfully
transfected were screened 48 h after transfection and further
cultured in complete DMEM in the 5% CO2 incubator at
37˚C and 95% humidity.
CCK-8 assay
A total of 100 µl complete DMEM was added into
96-well plates containing ~1x103 cells/well. Cells were
cultured in a 5% CO2 incubator at 37˚C and 95% humidity
for 24, 48 and 72 h respectively. At each time point, cells of each
group were removed from the incubator and 10 µl CCK8 reaction
solutions (Dojindo Molecular Technologies, Inc.) was added into
each well according to the manufacturer's protocol. After 4 h
incubation at room temperature, the optical density (OD) of each
well was determined at 450 nm wavelength using a microplate reader.
Relative proliferation of cells in each well was calculated and
OSCC cells transfected with si-NC served as control.
Transwell experiment
Transwell chambers (8 µm pore; Costar; Corning Inc.)
were placed in 24-well plates and 300 µl serum-free DMEM cell
suspensions (1x105 cells/ml) were added into the upper
chamber. The lower chamber contained 500 µl complete DMEM. The
24-well plates were maintained in the 5% CO2 incubator
at 37˚C and 95% humidity for 24 h and then cells that failed to
migrate were gently scraped off using cotton swabs. Cells that had
migrated through the membrane were fixed with formaldehyde for 30
min at 25˚C and stained with crystal violet (0.1%) for 30 min at
25˚C. For the invasion assay, Matrigel (BD Bioscience) was
pre-plated for 30 min at 37˚C in the upper chambers and the other
procedures were performed as described above. Cells that had
successfully migrated and invaded were counted in six random fields
of view under a routine light microscope.
Luciferase reporter gene assay
The binding site of MCM3AP-AS1 and miR-363-5p was
located in 3'-untranslated region (UTR) through analysis using
TargetScan version 7.1 (http://www.targetscan.org/vert_71/). The 3'-UTR
sequence of MCM3AP-AS1 was amplified and cloned into the pGL3
luciferase reporter vectors. The binding site was mutated to obtain
MCM3AP-AS1 mutant type (Mut) using the Quick-change Site-Directed
Mutagenesis kit (Agilent Technologies, Inc.). SCC-15 cells of the
miR-363-5p group and miR-NC group were seed in six-well plates, and
pGL3 luciferase reporter vectors containing the MCM3AP-AS1-Mut and
MCM3AP-AS1-wild-type (WT) were used to transfect SCC-15 cells of
these two groups. Cells were incubated in the 5% CO2
incubator at 37˚C, 95% humidity for 48 h, and the luciferase
activities of each well were determined by Dual-Luciferase reporter
assay kit (Promega Corporation) and normalized to Renilla
luciferase activity.
Reverse transcription-quantitative
(RT-q) PCR
Total RNA was extracted from tissues and cells by
using TRIzol® (Invitrogen; Thermo Fisher Scientific,
Inc.) and the procedure was performed according to the
manufacturer's protocols. The concentration of each total RNA
sample was measured using spectrophotometer. Each RNA sample was
subjected to reverse transcription to obtain cDNA at 42˚C for 30
min using the PrimeScript RT Reagent kit (Takara Bio, Inc.)
according to the manufacturer's protocol. qPCR was performed using
SYBR® Premix Ex Taq™ kit (Takara
Biotechnology Co., Ltd.) on an ABI7300 PCR detection system. The
thermocycling conditions were as follows: 40 cycles of 95˚C for 5
min, 95˚C for 10 sec, and 60˚C for 45 sec. The relative expression
of MCM3AP-AS1 and miR-363-5p was normalized to U6 using the
2-ΔΔCq method (8). MCM3AP-AS1 expression was normalized to
GAPDH and miR-363-5p expression was normalized to U6. Primer
sequences were as follows: MCM3AP-AS1 forward,
5'-GCTGCTAATGGCAACACTGA-3' and reverse, 5'-AGGTGCTGTCTGGTGGAGAT-3';
miR-363-5p forward, 5'-GCAGCAACTAGAAACG-3' and reverse,
5'-GCACTCATGCCCATTCATCC-3'; U6 forward, 5'-GTGGACCGCACAAGCTCGCT-3'
and reverse, 5'-TTGTTGAACGGCACTGTGTATAGCA-3'); and GAPDH forward,
5'-CAGGAGGCATTGCTGATGAT-3' and reverse,
5'-GAAGGCTGGGGCTCATTT-3'.
Statistical analysis
Data were processed using SPSS version 18.0 software
(SPSS, Inc.) and are expressed as the mean ± standard deviation.
Student's t-test was applied to perform comparisons between two
groups. One-way ANOVA followed by Tukey's test was used to analyze
datasets with more than two groups. Pearson correlation analysis
was used to analyze the correlation between MCM3AP-AS1 and
miR-363-5p expression. The association between MCM3AP-AS1 levels
and the clinical features of patients was analyzed using the
χ2 test. P<0.05 was considered to indicate a
statistically significant difference.
Results
Expression of MCM3AP-AS1 increases in
OSCC tissues and cells
Tumor tissues and normal tissues were collected from
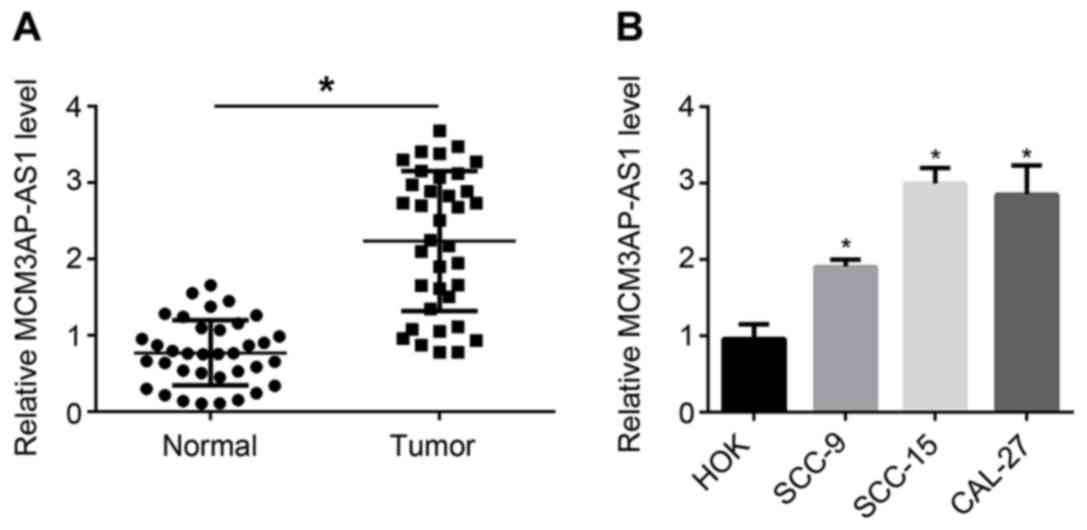
36 patients with OSCC. It can be seen from Fig. 1A that, compared with the relative
MCM3AP-AS1 expression in normal tissues, it was evidently increased
in tumor tissues (P<0.05). In addition, results from in
vitro experiments indicated a higher relative MCM3AP-AS1
expression in OSCC cells (including SCC-9, SCC-15 and CAL-27 cell
lines) compared with that in HOK cells (P<0.05). Therefore, the
expression of MCM3AP-AS1 was upregulated in OSCC tissues and
cells.
A high MCM3AP-AS1 level is associated
with poor prognosis of OSCC patients
The clinicopathological features of 36 patients with
OSCC were recorded, including age, sex, clinical stage and lymph
node metastasis. These samples were divided into MCM3AP-AS1 high
expression and low expression groups according to the median value
of MCM3AP-AS1. The relationship between MCM3AP-AS1 expression
levels and clinicopathological features of the patients was
explored (Table I). It was
identified that the MCM3AP-AS1 expression level had no obvious
relationship with the age or sex of the patients. However, clinical
stage and lymph node metastasis were significantly associated with
the expression level of MCM3AP-AS1. Compared with patients with
stage I/II and negative lymph node metastasis, a higher proportion
of patients with stage III/IV and positive lymph node metastasis
exhibited high MCM3AP-AS1 expression (P<0.05). Thus, high
MCM3AP-AS1 expression was clearly associated with poor prognosis in
patients with OSCC.
Knockdown of MCM3AP-AS1 inhibits OSCC
cells proliferation, migration and invasion
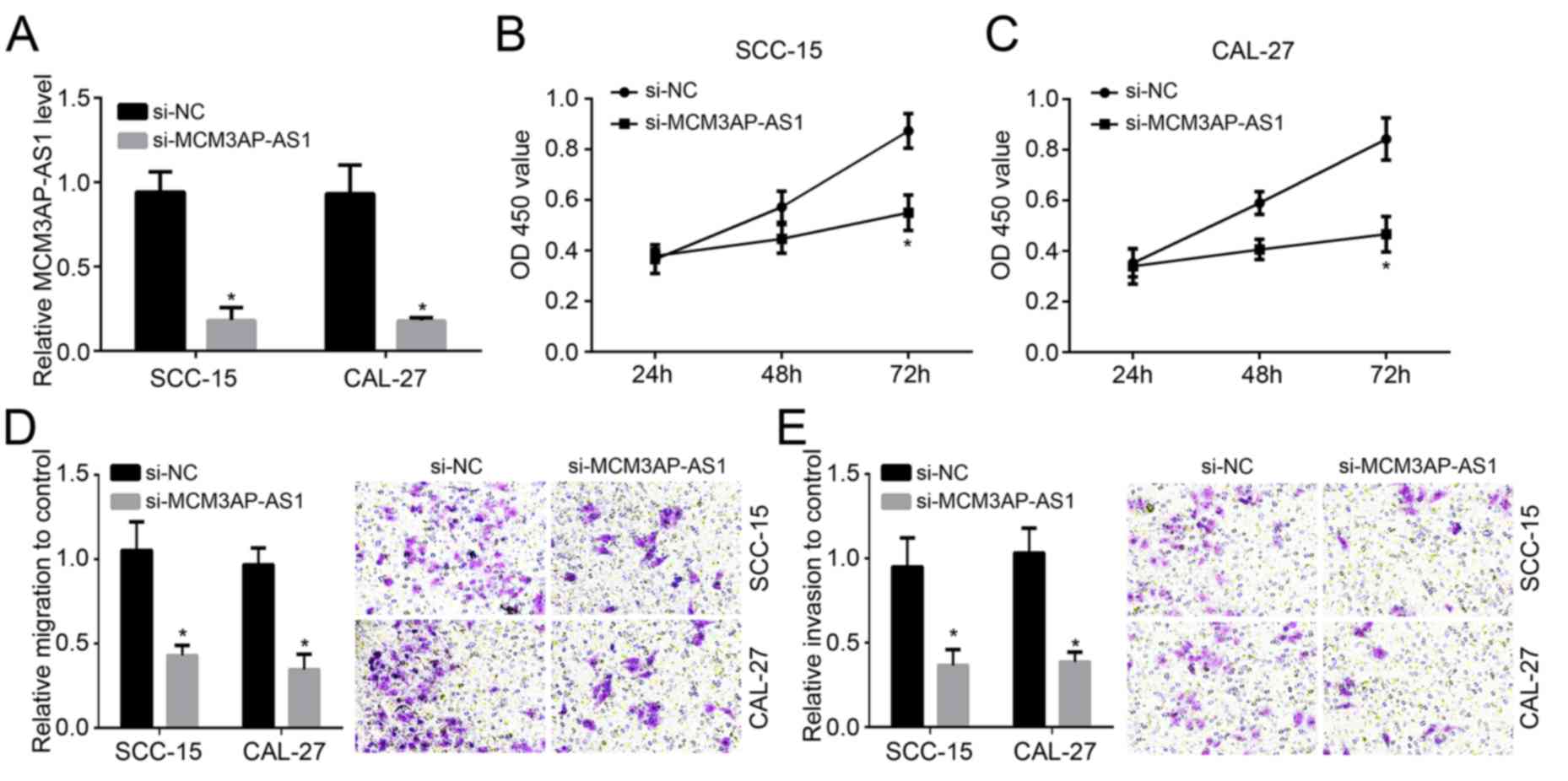
As MCM3AP-AS1 expression was highest in SCC-15 and
CAL-27 cells among all measured cell lines (Fig. 1B), these were chosen for
investigation. SCC-15 and CAL-27 cells were transfected with
MCM3AP-AS1 siRNA or MCM3AP-AS1 siRNA NC. The expression of
MCM3AP-AS1 was markedly reduced in the si-MCM3AP-AS1 group compared
with the si-NC group (P<0.05; Fig.
2A), suggesting that the transfection was successful. The
effects of MCM3AP-AS1 on OSCC cell proliferation, migration and
invasion were also demonstrated. The data indicated that compared
with the si-NC group, SCC-15 and CAL-27 cells of the si-MCM3AP-AS1
group exhibited a significantly decreased OD450 value after
transfection for 72 h (P<0.05; Fig.
2B and C). Results from the
Transwell assay indicated lower relative migration and invasion in
the si-MCM3AP-AS1 group compared with the si-NC group (P<0.05;
Fig. 2D and E). It may be that the proliferation,
migration and invasion of SCC-15 and CAL-27 cells were inhibited
after MCM3AP-AS1 knockdown.
Expression of miR-363-5p is directly
inhibited by MCM3AP-AS1 in OSCC cells
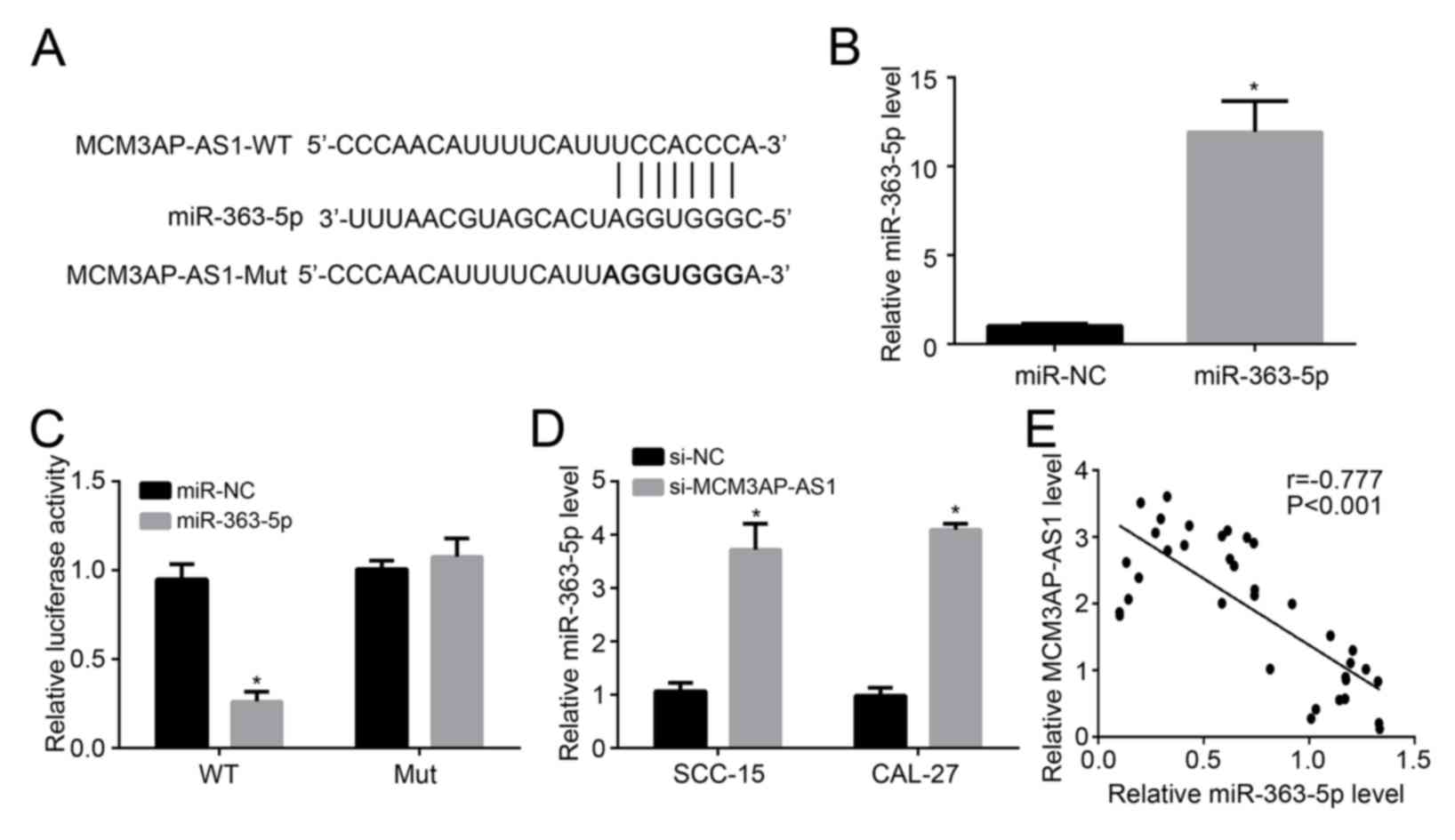
The binding site of miR-363-5p and MCM3AP-AS1 was
predicted, and MCM3AP-AS1-WT and MCM3AP-AS1-Mut sequences
containing the binding site were designed and are presented in
Fig. 3A. miR-363-5p expression in
SCC-15 cells was successfully elevated by transfection with
miR-363-5p mimic (P<0.05; Fig.
3B). The results of the luciferase reporter assay are shown in
Fig. 3C. It was demonstrated that
miR-363-5p upregulation decreased the relative luciferase activity
of SCC-15 cells loaded with MCM3AP-AS1-WT sequences (P<0.05).
However, the overexpression of miR-363-5p in SCC-15 cells had no
significant influence on the relative luciferase activity of SCC-15
cells loaded with MCM3AP-AS1-Mut sequences. miR-363-5p expression
in SCC-15 and CAL-27 cells of si-MCM3AP-AS1 group and si-NC group
was detected to further verify the regulatory relationship between
MCM3AP-AS1 and miR-363-5p. As shown in Fig. 3D, compared with si-NC group, much
higher relative miR-363-5p level was observed in the si-MCM3AP-AS1
group (P<0.05). Furthermore, in 36 OSCC tissue samples, there
was a significant negative correlation between the relative
MCM3AP-AS1 level and the relative miR-363-5p level (P<0.05;
Fig. 3E). Combining the above
results, it can be considered that miR-363-5p expression was
directly inhibited by MCM3AP-AS1 in OSCC cells.
Reduced proliferation, migration and
invasion of OSCC cells induced by MCM3AP-AS1 silencing is reversed
by downregulation of miR-363-5p
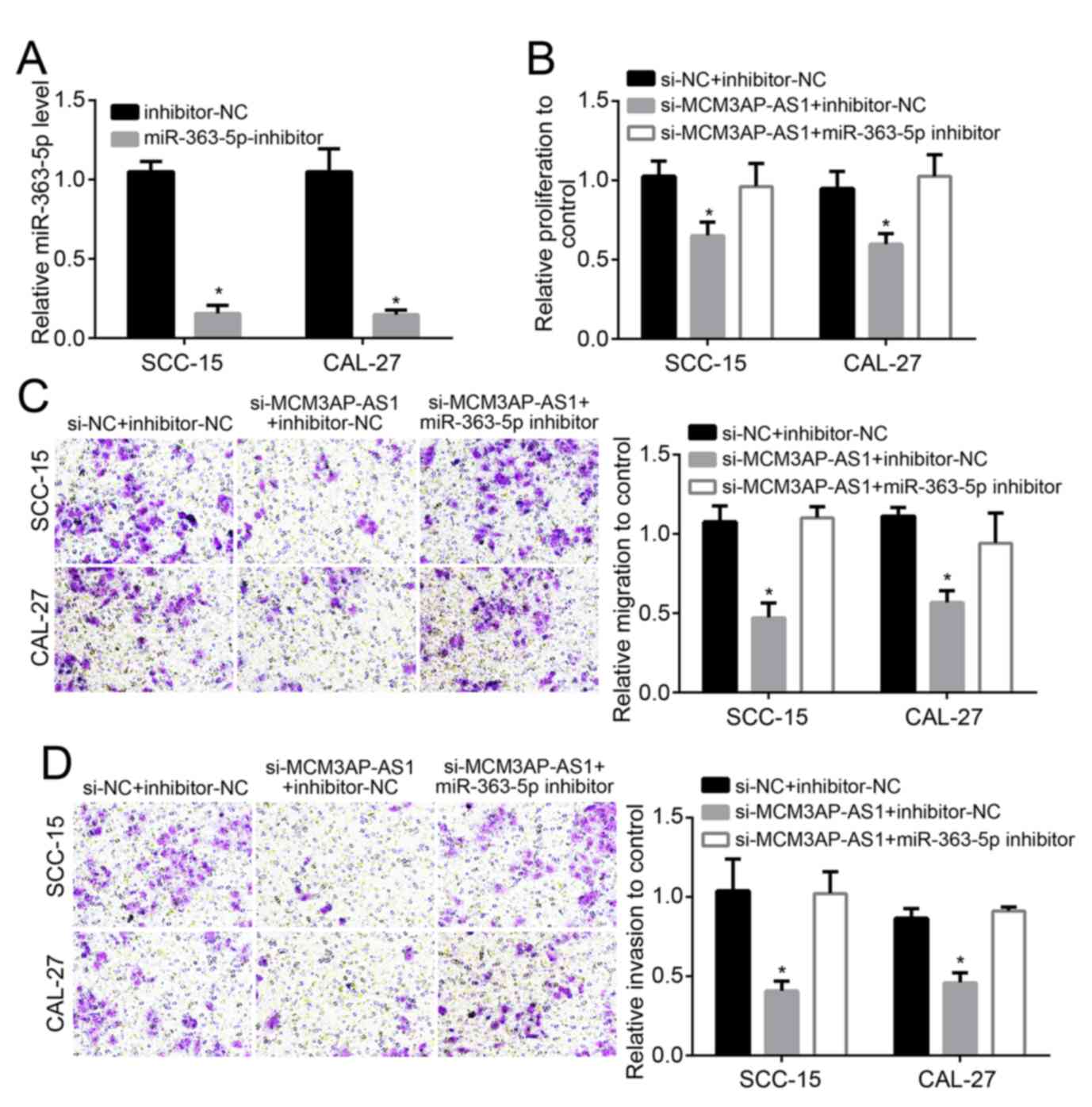
The relative miR-363-5p expression in SCC-15 and
CAL-27 cells of the miR-363-5p-inhibitor group was notably lower
compared with in the cells of inhibitor-NC group (P<0.05),
indicating that cells were successfully transfected with miR-363-5p
inhibitor (Fig. 4A). Co-transfection
was performed on SCC-15 and CAL-27 cells. Compared with the si-NC +
inhibitor-NC group, SCC-15 and CAL-27 cells of si-MCM3AP-AS1 +
inhibitor-NC group exhibited significantly lower relative
proliferation, migration and invasion (P<0.05). However, when
compared with the si-MCM3AP-AS1 + inhibitor-NC group, increased
relative proliferation, migration and invasion was found in SCC-15
and CAL-27 cells of the si-MCM3AP-AS1 + miR-363-5p inhibitor group
(P<0.05; Fig. 4B-D). This
illustrated that the reduced proliferation, migration and invasion
of OSCC cells induced by MCM3AP-AS1 silencing was reversed by
downregulation of miR-363-5p.
Discussion
OSCC is prone to lymph node metastasis and this is
the leading cause of mortality in patients (9). Finding an effective therapeutic target
for OSCC has important clinical implications for improving the
prognosis of patients with OSCC in the future. The present study
identified that MCM3AP-AS1 was an oncogene in OSCC and
downregulation of MCM3AP-AS1 could inhibit OSCC cell proliferation,
migration and invasion by enhancing miR-363-5p expression.
lncRNAs are highly conserved non-coding RNA
transcripts of >200 nucleotides in length, which have attracted
attention because of the discovery in recent years of their
association with the process of tumor formation, invasion and
metastasis (10,11). lncRNAs can fold into complex
three-dimensional structures and interact specifically with
transcription factors, histones and chromatin-modifying proteins,
thereby affecting the expression of other genes (12,13).
Several lncRNAs have been demonstrated to participate in the
occurrence and progression of OSCC, including H1 and HAS2-AS1. In
addition to the previously confirmed lncRNAs, the role of other
lncRNAs in the initiation and progression of OSCC is still worth
exploring. Thus, through clinical data and in vitro cell
experiments, the present study identified that MCM3AP-AS1 was
prominently upregulated in OSCC and that it acted as an oncogene in
OSCC. According to the limited studies available, MCM3AP-AS1 is
involved in the development of glioblastoma and hepatocellular
carcinoma (14,15). Researchers knocked out MCM3AP-AS1 in
glioma-associated endothelial cells by lentiviral-mediated
sh-MCM3AP-AS1 transfection and noticed the decline in
proliferation, migration and invasion of glioma-associated
endothelial cells (14). Excessive
MCM3AP-AS1 expression was also found in hepatocellular carcinoma,
and this was associated with a lower 5-year survival rate of
patients (15). Following the
deletion of MCM3AP-AS1 by transfection, hepatocellular carcinoma
cell proliferation in vitro and growth in vivo were
both curbed (15). The data of the
present study were consistent with the results of the
aforementioned previous studies and, to the best of our knowledge,
this was the first study to report the effects of MCM3AP-AS1 on
OSCC.
miRNAs are an important type of regulator that
affect post-transcriptional expression levels of genes (16). miRNAs are involved in multiple
biological functions, including cell proliferation, apoptosis and
invasion, and their deregulation is closely related to a variety of
human tumors (17). Previous studies
have reported that MCM3AP-AS1 can regulate the development of
glioblastoma and hepatocellular carcinoma by directly binding to
miR-211 and miR-194-5p (14,15). The present study identified that
MCM3AP-AS1 promoted the development of OSCC by targeting inhibition
of miR-363-5p expression. miR-363 has been discovered to be
involved in human cancer. Qiao et al (18) suggested that the upregulated
expression of miR-363 can suppress the tumorigenicity and
metastasis of neuroblastoma. A previous study also noted that in
head and neck squamous cell carcinoma, miR-363 expression was
significantly reduced in the tumor tissues of patients with lymph
node metastasis and tumor cells with high invasive ability
(19). Decreased miR-363 expression
is conducive to the invasion and metastasis of head and neck
squamous cell carcinoma (19). In
gastric cancer, decreased miR-363-3p is detected in tumor tissues
and cells, and tumor cells migration and growth can be suppressed
by enhancing miR-363-3p expression (20). Thus far, there have been relatively
few studies on miR-363-5p in human tumors. Zhang et al
(21) illustrated that low
expression of miR-363-5p is correlated with high overall survival
of patients with hepatocellular carcinoma and may be a potential
prognostic indicator for hepatocellular carcinoma. In patients with
nasopharyngeal carcinoma, lncRNA FOXD2-AS1 blocks the expression of
miR-363-5p and aggravates tumorigenesis (22). The results of the present study
revealed that the miR-363-5p expression level was negatively
correlated with MCM3AP-AS1 and that silencing of MCM3AP-AS1
inhibited OSCC cell proliferation, migration and invasion by
enhancing the expression of miR-363-5p. To further verify this, the
measurement of the effects of miR-363-5p overexpression on
proliferation, migration and invasion should be performed in the
future.
The results from the present study identified that
MCM3AP-AS1 expression was markedly elevated in OSCC tissues and
cells, and increased MCM3AP-AS1 expression was associated with poor
prognosis of OSCC patients. OSCC cells proliferation, migration and
invasion were obviously inhibited after MCM3AP-AS1 knockout.
Mechanistically, MCM3AP-AS1 directly inhibited miR-363-5p
expression and enhanced the development of OSCC. Therefore,
MCM3AP-AS1 served a carcinogenic role in OSCC, which might be a
potential therapeutic target for OSCC.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the present
study are available from the corresponding author on reasonable
request.
Authors' contributions
CH and BD initiated and designed the present study,
analyzed the data, interpreted the results and wrote the
manuscript. CH and XW performed the experiments. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
Regarding the use of human samples, the protocol of
the present study was approved by the Ethics Committee of Zaozhuang
Municipal Hospital and all enrolled patients signed a written
informed consent document.
Patient consent for publication
All patients within this study provided consent for
the publication of their data.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Hema KN, Smitha T, Sheethal HS and
Mirnalini SA: Epigenetics in oral squamous cell carcinoma. J Oral
Maxillofac Pathol. 21:252–259. 2017.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Mourad M, Jetmore T, Jategaonkar AA,
Moubayed S, Moshier E and Urken ML: Epidemiological trends of head
and neck cancer in the united states: A SEER population study. J
Oral Maxillofac Surg. 75:2562–2572. 2017.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Nakashima C, Yamamoto K, Fujiwara-Tani R,
Luo Y, Matsushima S, Fujii K, Ohmori H, Sasahira T, Sasaki T,
Kitadai Y, et al: Expression of cytosolic malic enzyme (ME1) is
associated with disease progression in human oral squamous cell
carcinoma. Cancer Sci. 109:2036–2045. 2018.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Alessandro F and Irene B: Long non-coding
RNAs: New players in cell differentiation and development. Nat Rev
Genet. 15:7–21. 2014.PubMed/NCBI View
Article : Google Scholar
|
|
5
|
Marín-Béjar O, Mas AM, González J,
Martinez D, Athie A, Morales X, Galduroz M, Raimondi I, Grossi E,
Guo S, et al: The human lncRNA LINC-PINT inhibits tumor cell
invasion through a highly conserved sequence element. Genome Biol.
18(202)2017.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Zhang S, Tian L, Ma P, Sun Q, Zhang K,
Wang G, Liu H and Xu B: Potential role of differentially expressed
lncRNAs in the pathogenesis of oral squamous cell carcinoma. Arch
Oral Biol. 60:1581–1587. 2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Chang SM and Hu WW: Long non-coding RNA
MALAT1 promotes oral squamous cell carcinoma development via
microRNA-125b/STAT3 axis. J Cell Physiol. 233:3384–3396.
2018.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Abdulaziz MA, Amin AK, Elrouby DH and
Shaker OG: Lymphangiogenesis in oral squamous cell carcinoma:
Correlation with VEGF-C expression and lymph node metastasis. Int J
Dent. 2017(7285656)2017.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Xiao H, Zhang F, Zou Y, Li J, Liu Y and
Huang W: The function and mechanism of long non-coding RNA-ATB in
cancers. Front Physiol. 9(321)2018.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Fan S, Fan C, Liu N, Huang K, Fang X and
Wang K: Downregulation of the long non-coding RNA ZFAS1 is
associated with cell proliferation, migration and invasion in
breast cancer. Mol Med Rep. 17:6405–6412. 2018.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Bonasio R and Shiekhattar R: Regulation of
transcription by long noncoding RNAs. Annu Rev Genet. 48:433–455.
2014.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Zhang L, Wang H, Xu M, Chen F, Li W, Hu H,
Yuan Q, Su Y, Liu X, Wuri J and Yan T: Long noncoding RNA HAS2-AS1
promotes tumor progression in glioblastoma via functioning as a
competing endogenous RNA. J Cell Biochem. 121:661–671.
2020.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yang C, Zheng J, Xue Y, Yu H, Liu X, Ma J,
Liu L, Wang P, Li Z, Cai H and Liu Y: The effect of
MCM3AP-AS1/miR-211/KLF5/AGGF1 axis regulating glioblastoma
angiogenesis. Front Mol Neurosci. 10(437)2018.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Wang Y, Yang L, Chen T, Liu X, Guo Y, Zhu
Q, Tong X, Yang W, Xu Q, Huang D and Tu K: A novel lncRNA
MCM3AP-AS1 promotes the growth of hepatocellular carcinoma by
targeting miR-194-5p/FOXA1 axis. Mol Cancer. 18(28)2019.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Behbahani GD, Ghahhari NM, Javidi MA,
Molan AF, Feizi N and Babashah S: MicroRNA-mediated
post-transcriptional regulation of epithelial to mesenchymal
transition in cancer. Pathol Oncol Res. 23:1–12. 2017.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Yang Z, Wu L, Wang A, Tang W, Zhao Y, Zhao
H and Teschendorff AE: dbDEMC 2.0: Updated database of
differentially expressed miRNAs in human cancers. Nucleic Acids
Res. 45 (D1):D812–D818. 2017.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Qiao J, Lee S, Paul P, Theiss L, Tiao J,
Lan Q, Kong A and Dai HC: miR-335 and miR-363 regulation of
neuroblastoma tumorigenesis and metastasis. Surgery. 154:226–233.
2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Sun Q, Zhang J, Cao W, Wang X, Xu Q, Yan
M, Wu X and Chen W: Dysregulated miR-363 affects head and neck
cancer invasion and metastasis by targeting podoplanin. Int J
Biochem Cell Biol. 45:513–520. 2013.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Song B, Yan J, Liu C, Zhou H and Zheng Y:
Tumor suppressor role of miR-363-3p in gastric cancer. Med Sci
Monit. 21:4074–4080. 2015.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Zhang J, Fan J, Zhou C and Qi Y:
miR-363-5p as potential prognostic marker for hepatocellular
carcinoma indicated by weighted co-expression network analysis of
miRNAs and mRNA. BMC Gastroenterol. 17(81)2017.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Chen G, Sun W, Hua X, Zeng W and Yang L:
Long non-coding RNA FOXD2-AS1 aggravates nasopharyngeal carcinoma
carcinogenesis by modulating miR-363-5p/S100A1 pathway. Gene.
645:76–84. 2018.PubMed/NCBI View Article : Google Scholar
|