Introduction
Retinoblastoma (RB), which is a common and harmful
intraocular malignant tumor in children that threatens life
(1). RB incidence is between
1/15,000 and 1/20,000, and is more common in infants and young
children, with the majority of cases occurring before the age of
six (1). Chemical volume reduction
is one of the principal methods of conservative treatment. Local
treatment mainly includes laser photocoagulation, transpupillary
thermotherapy, photodynamic therapy, cryotherapy and scleral
application radiotherapy (2).
However, the aforementioned methods lack specificity, and exhibit
destructive injuries, low safety, serious systemic and/or local
adverse effects and other problems, such as tumor implantation and
induction of secondary malignant tumors (2). With the development of tumor molecular
biology and genetic engineering technology, coupled with the
anatomical advantages of eyeballs as target tissues in the field of
RB therapy, gene therapy, as a new type of biological therapy, has
demonstrated good application prospects (3,4).
Tumor suppressor candidate 3 (TUSC3) is located on
chromosome 8p22, and is widely expressed in human tissues, such as
the brain, heart, lung and liver (5-8).
Due to its decreased expression in tumor cells such as colon,
breast, liver, pancreatic and rectal cancer, TUSC3 is considered to
be a tumor suppressor gene (9-14).
TUSC3 is an intrinsic membrane protein that catalyzes the process
of endoplasmic reticulum N-glycosylation, which is a major
post-translational modification mechanism in cells, and serves a
critical role in the folding, regulation and stabilization of
proteins (15). Insufficient
glycosylation has been indicated to cause endoplasmic reticulum
stress, result in genomic damage mutations and cause cancer
(16-18).
Silencing TUSC3 in prostate and ovarian cancer has been revealed to
promote tumor cell growth, metastasis and invasiveness (7,8).
Concurrently, the low expression of TUSC3 in cancer cells may
indicate a poor prognosis and a higher possibility of metastasis
(6,19-22).
TUSC3 has been indicated to serve a role in a number of malignant
tumors including prostate cancer, ovarian cancer, lung cancer and
glioma. Therefore, the expression and role of TUSC3 in
retinoblastoma cells requires additional elucidation.
MicroRNAs (miRs/miRNAs), which are a class of small
endogenous non-coding RNAs that are ~22 nucleotides in length,
regulate gene expression at the post-transcriptional level via
binding to the 3'-untranslated region (3'-UTR) of target mRNAs
(23-25).
miRNAs have been identified to serve critical roles in regulating
cell proliferation, differentiation and apoptosis (26-28).
A number of studies have indicated the role of miRNAs in tumors
(29,30). miR-320a, which is an extensively
studied miRNA, has been reported to serve a critical role in
diabetic retinopathy (31) and
atherosclerosis (32). Moreover,
miR-320a has been investigated in several types of cancer, such as
lung cancer (33), papillary
thyroid cancer (34), osteosarcoma
(35) and hepatocellular carcinoma
(36). Moreover, miR-320a has been
reported to be upregulated in retinoblastoma tissues (37); however its role and mechanism in
retinoblastoma remain to be elucidated.
Bioinformatics analysis revealed direct interaction
sites between miR-320a and TUSC3. Therefore, it was hypothesized
that miR-320a may serve a role in retinoblastoma cells via
regulating TUSC3. The aim of the present study was to explore the
role of miR-320 in retinoblastoma cells and analyze its molecular
mechanism of function to provide novel insights for the treatment
of retinoblastoma.
Materials and methods
Cell culture
The human normal retinal vascular endothelial cell
line ARPE-19 and the retinoblastoma cell lines Y79 and WERI-Rb-1
were obtained from American Type Culture Collection. The cells were
cultured in DMEM (Gibco; Thermo Fisher Scientific, Inc.)
supplemented with 10% FBS (Gibco; Thermo Fisher Scientific, Inc.)
and 1% penicillin/streptomycin in a humidified incubator at 37˚C
with 5% CO2.
Reverse transcription-quantitative PCR
(RT-qPCR)
RNA extraction from ARPE-19, Y79 and WERI-Rb-1 cells
was performed using TRIzol® reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's
instructions. cDNA was reverse transcribed from RNA using the
HiScript II Q RT SuperMix (Vazyme Biotech Co., Ltd.). The following
temperature conditions for RT were as follows: 70˚C for 5 min, 37˚C
for 5 min and 42˚C for 60 min. Subsequently, qPCR was performed
using ChamQ Universal SYBR® qPCR Master Mix (Vazyme
Biotech Co., Ltd.) according to the manufacturer's instructions.
The following thermocycling conditions were used for the qPCR:
Initial denaturation at 95˚C for 10 min; 40 cycles of denaturation
at 95˚C for 10 sec, annealing at 60˚C for 20 sec and extension at
72˚C for 34 sec. GAPDH for mRNA and U6 for miRNA were used as the
internal controls. The primer sequences used for the PCR were
listed as follows: GAPDH forward, 5'-CTTTGGTATCGTGGAAGGACTC-3' and
reverse, 5'-GTAGAGGCAGGGATGATGTTCT-3'; U6 forward,
5'-GCTTCGGCAGCACATATACTAAAAT-3' and reverse,
5'-CGCTTCACGAATTTGCGTGTCAT-3'; miR-320a forward,
5'-GTTGGATCCGGCGTTTCCTTCCGACATG-3' and reverse,
5'-GCTGAATTCGTCCACTGCGGCTGTTCC-3'; TUSC3 forward,
5'-GGCTCAGTTTGTGGCAGAATC-3' and reverse,
5'-CATCGCCTTTCGAAGTTGCT-3'. The relative gene expression levels
were analyzed using the 2-ΔΔCq method (38). All experiments were performed in
triplicate.
Dual-luciferase reporter assay
TargetScan bioinformatics software version 7.2
(www.targetscan.org/vert_72) was used
to predict the potential targets of miR-320a. Binding sites between
miR-320a and the 3'-untranslated region (3'-UTR) of TUSC3 were
observed. Dual luciferase reporter assay was performed to determine
whether miR-320a directly bound to TUSC3. Wild-type (WT) and mutant
(MUT) 3'-UTR of TUSC3 were cloned into pmiR-RB-Report™ dual
luciferase reporter vector (Guangzhou RiboBio Co., Ltd.) according
to the manufacturer's instructions. 293 T cells (American Type
Culture Collection) were co-transfected with WT-TUSC3 or MUT-TUSC3
and 100 nM miR-320a mimic (5'-AAAAGCUGGGUUGAGAGGGCGA-3';
3'-UUUUCGACCCAACUCUCCCGCU-5'; Guangzhou RiboBio Co., Ltd.) or 100
nM mimic control (5'-UUCUCCGAACGUGUCACGUTT-3';
3'-TTAAGAGGCUUGCACAGUGCA-5'; Guangzhou RiboBio Co., Ltd.) using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.) at 37˚C for 48 h. At 48 h after transfection,
luciferase activity was determined using the
Dual-luciferase® Reporter Assay system (Promega
Corporation) and normalized to Renilla luciferase
activity.
Cell transfection
1 µg TUSC3-plasmid (cat no. sc-405571-ACT; Santa
Cruz Biotechnology, Inc.), 1 µg control-plasmid (cat no. sc-437275;
Santa Cruz Biotechnology, Inc.), 100 nM inhibitor control
(5'-UUGUCCUACACCUCACUCCUG-3'; Guangzhou RiboBio Co., Ltd.), 100 nM
miR-320a inhibitor (5'-UCGCCCUCUCAACCCAGCUUUU-3'; Guangzhou RiboBio
Co., Ltd.), 1 µg TUSC3-short hairpin RNA (shRNA; cat no.
sc-77535-SH; Santa Cruz Biotechnology, Inc.), 1 µg control-shRNA
(cat no. sc-108060; Santa Cruz Biotechnology, Inc.), 100 nM
miR-320a inhibitor + 1 µg control-shRNA and 100 nM miR-320a
inhibitor + 1 µg TUSC3-shRNA were transfected into Y79 and
WERI-Rb-1 cells (5x104 cells per well; 24 well plates)
using Lipofectamine® 2000 reagent (Invitrogen; Thermo
Fisher Scientific, Inc.) according to the manufacturer's protocol.
Cells without any treatment were used as the control. The
transfection efficiency was examined via RT-qPCR following 48 h of
transfection.
MTT assay
Y79 and WERI-Rb-1 cells were seeded in 96-well
plates (5x104 cells per well) and transfected with
TUSC3-plasmid, control-plasmid, miR-320a inhibitor, control
inhibitor, miR-320a inhibitor + control-shRNA or miR-320a inhibitor
+ TUSC3-shRNA for 48 h. Subsequently, 20 µl MTT solution (5 g/l;
Sigma-Aldrich; Merck KGaA) was added to each well. The plates were
incubated at 37˚C with 5% CO2 for 4 h. The culture
medium was then discarded and 150 µl DMSO (Beyotime Institute of
Biotechnology) was added to each well. The plates were gently
agitated at 37˚C for 10 min. The optical density was measured at a
wavelength of 490 nm using a multifunctional plate reader (BD
Biosciences).
Flow cytometry analysis
At 48 h after transfection, Y79 and WERI-Rb-1 cells
(106 cells) in the log phase were digested with trypsin
(0.25%) without EDTA at room temperature for 1 min, centrifuged at
1,000 x g for 5 min at 4˚C, and the supernatant was discarded. The
cell pellet was washed twice with pre-chilled PBS and then
resuspended in 195 µl pre-chilled 1X Annexin V binding buffer
(Annexin V-FITC Cell apoptosis detection kit; Beyotime Institute of
Biotechnology). Subsequently, cells were incubated with 5 µl
Annexin V-FITC and 10 µl propidium iodide for 15 min at room
temperature in the dark. To detect apoptosis, flow cytometry
(Beckman Coulter, Inc.) was performed. The data were analyzed using
CellQuest™ v5.1 software (BD Biosciences).
Western blot analysis
The expression of TUSC3 was detected via western
blotting. Proteins from ARPE-19, Y79 and WERI-Rb-1 cells were
extracted using RIPA lysis buffer (Beyotime Institute of
Biotechnology) and protein concentration was measured using a BCA
assay kit (Sigma-Aldrich; Merck KGaA) according to the
manufacturer's protocol. A total of 40 µg proteins/lane were
separated using 10% SDS-PAGE and subsequently transferred to PVDF
membranes (EMD Millipore). Following blocking with 5% skimmed milk
for 1 h at room temperature, the membranes were incubated with
primary antibodies for TUSC3 (1:1,000; cat no. ab230520; Abcam) and
GAPDH (1:1,000; cat no. ab181602; Abcam) overnight at 4˚C.
Subsequently, the membranes were incubated with a corresponding
horseradish peroxidase-conjugated secondary antibody (1:2,000; cat.
no. 7074; Cell Signaling Technology, Inc.) for 1 h at room
temperature. Protein bands were visualized using ECL Western
blotting Detection Reagents (Cytiva).
Statistical analysis
Experiments were repeated in triplicate. Data are
presented as the mean ± standard deviation of three independent
experiments. Statistical analysis was performed using GraphPad
Prism v5 software (GraphPad Software, Inc.). Statistical
differences between multiple groups were analyzed using one-way
ANOVA with a Bonferroni post hoc test, and Student's t-test was
used for comparison between two groups, as applicable. P<0.05
was considered to indicate a statistically significant
difference.
Results
miR-320a expression in retinoblastoma
cell lines
RT-qPCR was performed to examine the expression of
miR-320a in retinoblastoma cell lines Y79 and WERI-Rb-1, and human
normal retinal vascular endothelial cell line ARPE-19. As
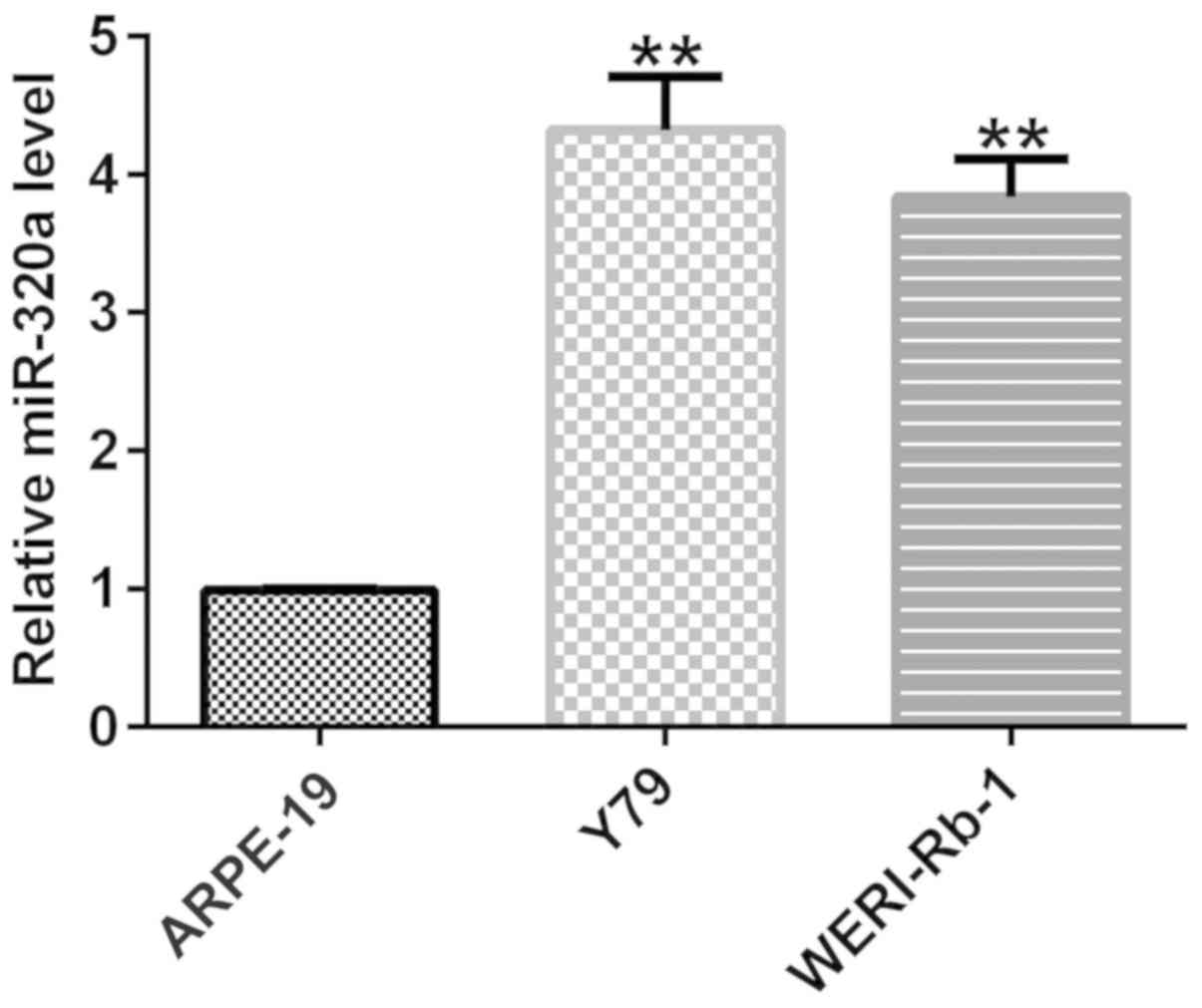
demonstrated in Fig. 1, compared
with ARPE-19 cells, the expression of miR-320a in retinoblastoma
cell lines Y79 and WERI-Rb-1 was upregulated, which is consistent
with a previous study on miR-320a (37). These results indicated that miR-320a
was expressed at a higher level in retinoblastoma cell lines Y79
and WERI-Rb-1, compared with normal retinal cells.
Target gene of miR-320a
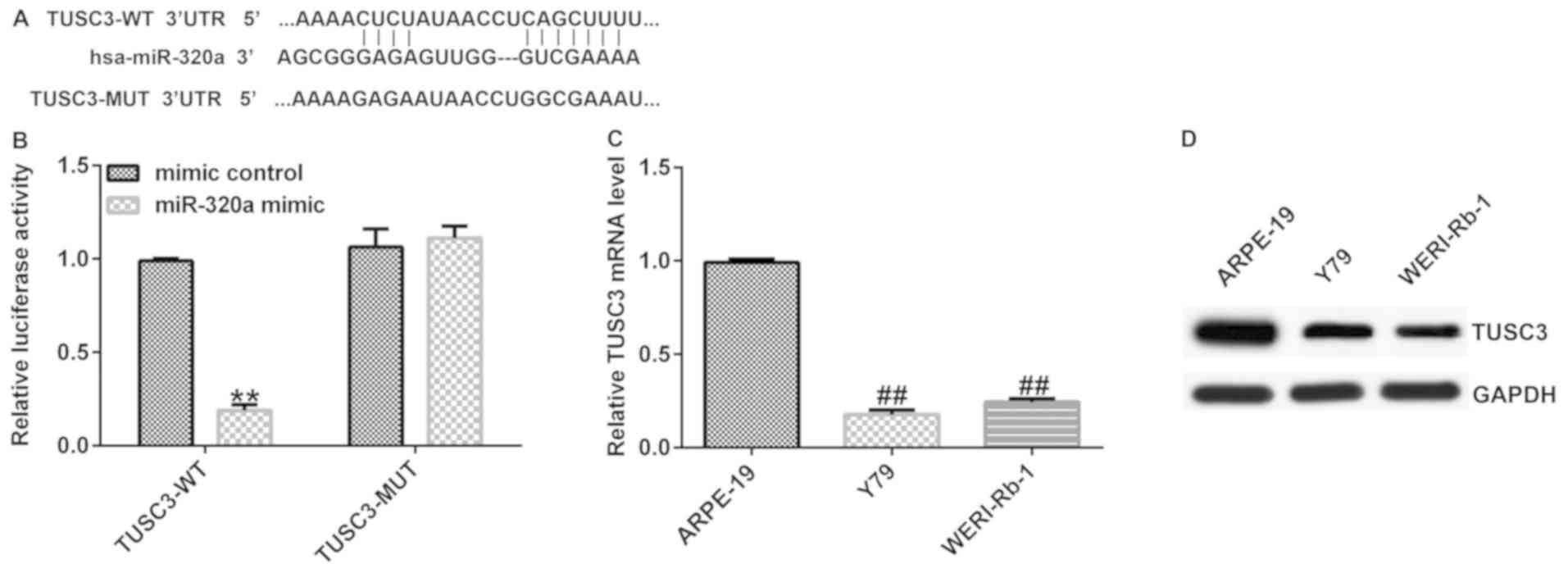
Binding sites between miR-320a and the 3'-UTR of
TUSC3 mRNA were predicted via TargetScan (Fig. 2A), which indicated that TUSC3 is a
potential target gene of miR-320a. A luciferase reporter assay
revealed that miR-320a mimic suppressed the luciferase activity
when 293 T cells were co-transfected with a reporter plasmid
containing the WT 3'-UTR and miR-320a mimic (Fig. 2B). However, the luciferase activity
of the MUT 3'-UTR was not altered. These data indicated that TUSC3
was a direct target of miR-320a.
The expression of TUSC3 in retinoblastoma cell lines
Y79 and WERI-Rb-1 and the human normal retinal vascular endothelial
cell line ARPE-19 was examined via RT-qPCR and western blotting.
The results indicated that compared with ARPE-19 cells, the mRNA
and protein expression of TUSC3 was reduced in both retinoblastoma
cell lines (Fig. 2C and D).
Effect of TUSC3 overexpression on
retinoblastoma cell viability and apoptosis
Y79 and WERI-Rb-1 cells were transfected with
TUSC3-plasmid and control-plasmid. Following 48 h of transfection,
RT-qPCR was performed to assess transfection efficiency. MTT assay
and flow cytometry were also performed to assess cell viability and
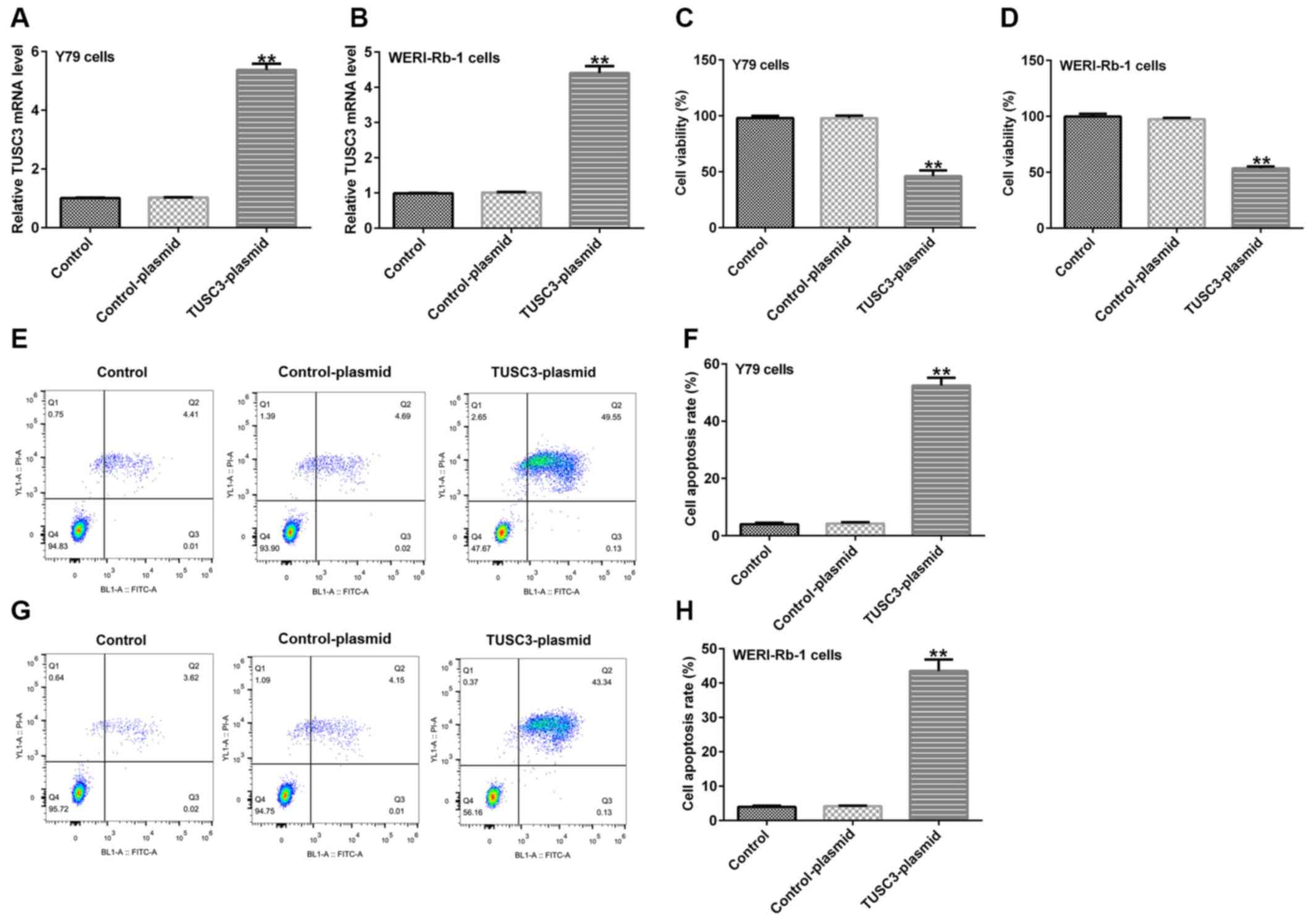
apoptosis, respectively. Compared with the control-plasmid group,
the mRNA expression of TUSC3 in Y79 and WERI-Rb-1 cells was
increased following transfection with TUSC3-plasmid (Fig. 3A and B). Moreover, the viability of Y79
(Fig. 3C) and WERI-Rb-1 (Fig. 3D) cells was reduced, while the
apoptotic rates of Y79 (Fig. 3E and
F) and WERI-Rb-1 (Fig. 3G and H) cells were increased, compared with the
control-plasmid group.
Effect of miR-320a inhibition on
retinoblastoma cell viability and apoptosis
Y79 and WERI-Rb-1 cells were transfected with
inhibitor control, miR-320a inhibitor, TUSC3-shRNA, control-shRNA,
miR-320a inhibitor + control-shRNA or miR-320a inhibitor +
TUSC3-shRNA for 48 h. RT-qPCR was performed to assess transfection
efficiency.
As presented in Fig.
4A and B, compared with the
inhibitor control group, miR-320a inhibitor reduced the expression
of miR-320a in Y79 and WERI-Rb-1 cells. Compared with the
control-shRNA group, TUSC3-shRNA reduced the mRNA expression of
TUSC3 in Y79 and WERI-Rb-1 cells (Fig.
4C and D). Moreover, miR-320a
inhibitor increased the mRNA and protein expression of TUSC3 in Y79
cells (Fig. 4E and F) and WERI-Rb-1 cells (Fig. 4G and H), compared with the inhibitor control
group, while this increase was reversed by TUSC3-shRNA.
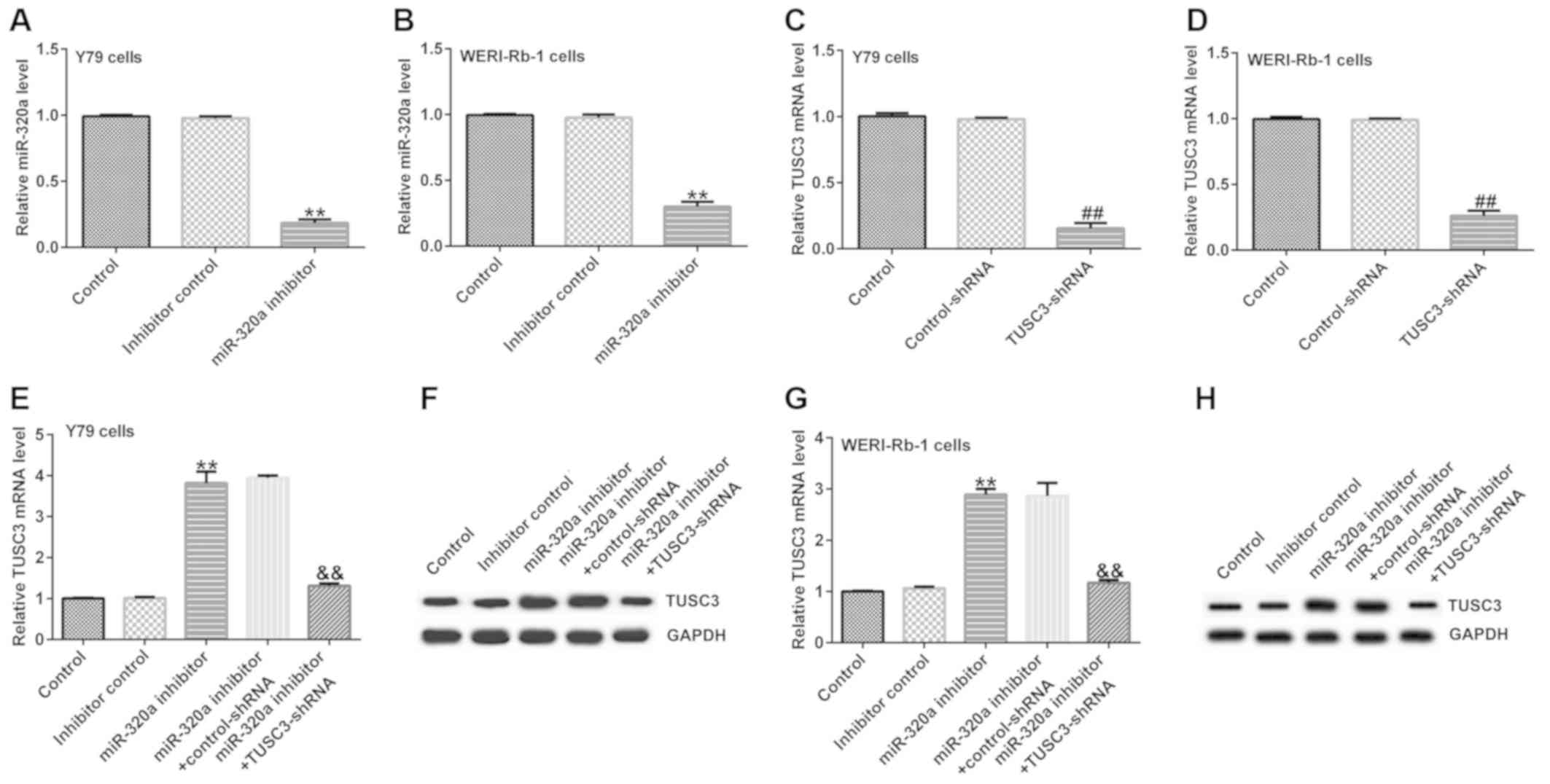 | Figure 4Expression of miR-320a and TUSC3 in
transfected cells. Y79 and WERI-Rb-1 cells were transfected with
inhibitor control, miR-320a inhibitor, TUSC3-shRNA, control-shRNA,
miR-320a inhibitor + control-shRNA or miR-320a inhibitor +
TUSC3-shRNA. Expression of miR-320a in (A) Y79 and (B) WERI-Rb-1
cells transfected with inhibitor control or miR-320a inhibitor.
TUSC3 mRNA expression in (C) Y79 and (D) WERI-Rb-1 cells
transfected with control-shRNA or TUSC3-shRNA. (E) mRNA and (F)
protein expression of TUSC3 in Y79 cells transfected with inhibitor
control, miR-320a inhibitor, miR-320a inhibitor + control-shRNA or
miR-320a inhibitor + TUSC3-shRNA. (G) mRNA and (H) protein
expression of TUSC3 in WERI-Rb-1 cells transfected with inhibitor
control, miR-320a inhibitor, miR-320a inhibitor + control-shRNA or
miR-320a inhibitor + TUSC3-shRNA. The data are presented as the
mean ± standard deviation. **P<0.01 vs. inhibitor
control; ##P<0.01 vs. control-shRNA;
&&P<0.01 vs. miR-320a inhibitor +
control-shRNA. miR-320a, microRNA-320a; TUSC3, tumor suppressor
candidate 3; shRNA, short hairpin RNA. |
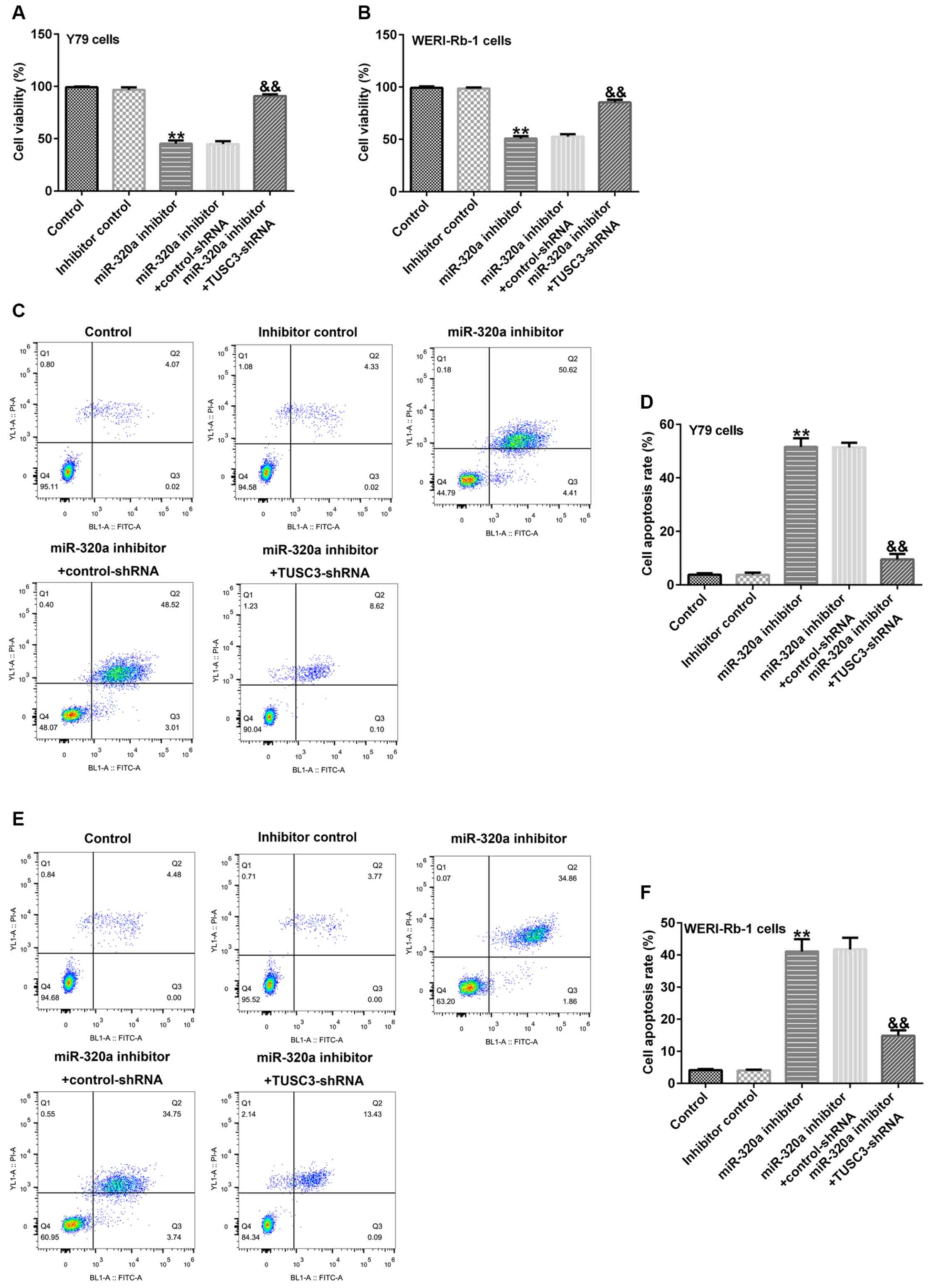
Subsequent analysis indicated that compared with the
inhibitor control group, miR-320a inhibitor reduced the cell
viability (Fig. 5A and B) and induced apoptosis (Fig. 5C-F) in Y79 and WERI-Rb-1 cells,
while these alterations were reversed by TUSC3-shRNA.
Discussion
The potential biomarkers and therapeutic targets of
tumors have provided insight into the clinical treatment of
retinoblastoma (39,40). In recent years, the relationship
between miRNAs and retinoblastoma has been extensively studied
(41,42). Different miRNAs have been indicated
to exhibit distinct expression levels in diverse tumor tissues, and
may exert both oncogenic and antitumor effects (40,43).
Gao et al (44) revealed
that compared with the placental samples from healthy control
subjects, miR-320a expression was enhanced in the placental
specimens of patients with pre-eclampsia and excessive miR-320a
expression was indicated to suppress the trophoblast invasion;
however it did not affect the trophoblast migration or
proliferation. Yong et al (45) presented evidence that ectopic
expression of DiGeorge syndrome critical region gene 5 inhibited
proliferation and migration, and promoted fluorouracil resistance
in pancreatic ductal adenocarcinoma cells, while its mechanism of
action was associated with miR-320a. The present study focused on
the investigation of the role of miR-320a in retinoblastoma
cells.
The expression of miR-320a and TUSC3 in
retinoblastoma and their mechanism of action require additional
elucidation. The present study was performed based on the results
of previous research (37).
Consistently with a previous study (37), the results of the present study
indicated that miR-320a expression was upregulated in
retinoblastoma cells compared with normal retinal cells. TUSC3,
which is a well-known tumor suppressor gene, was revealed to be a
direct target of miR-320a, and was indicated to be negatively
regulated by miR-320a. Furthermore, the expression of TUSC3 was
downregulated in retinoblastoma cells compared with normal retinal
cells. Subsequent analyses indicated that TUSC3 overexpression
reduced retinoblastoma cell viability and induced cell apoptosis.
Moreover, miR-320a downregulation inhibited the viability of
retinoblastoma cells and induced cell apoptosis. The effects of
miR-320a inhibitor on retinoblastoma cells were reversed by
TUSC3-shRNA. However, apoptosis-related proteins were not examined
in the current study, and the efficiency of TUSC3 knock-down and
upregulation were only detected via RT-qPCR. These were the
limitations of the current study, and require additional
investigation.
In conclusion, the present study demonstrated that
in human retinoblastoma cells, inhibition of miR-320a prevented
cell growth via targeting TUSC3. miR-320a may be a novel potential
target for retinoblastoma treatment.
Acknowledgements
Not applicable.
Funding
No funding was received.
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
LK designed the current study, collected the data,
performed statistical analysis, interpreted the data and prepared
the manuscript preparation. YS, MC and YD collected the data and
performed statistical analysis. ZL designed the current study,
collected the data and prepared the manuscript. All authors read
and approved the final manuscript.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Wong JR, Tucker MA, Kleinerman RA and
Devesa SS: Retinoblastoma incidence patterns in the US
surveillance, epidemiology, and end results program. JAMA
Ophthalmol. 132:478–483. 2014.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Riquelam I, Tapia O, Leal P, Sandoval A,
Varga MG, Letelier P, Buchegger K, Bizama C, Espinoza JA, Peek RM,
et al: miR-101-2, miR-125b-2 and miR-451a act as potential tumor
suppressors in gastric cancer through regulation of the
PI3K/AKT/mTOR pathway. Cell Oncol (Dordr). 39:23–33.
2016.PubMed/NCBI View Article : Google Scholar
|
|
3
|
McEvoy JD and Dyer MA: Genetic and
epigenetic discoveries in human retinoblastoma. Crit Rev Oncog.
20:217–225. 2015.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Gudiseva HV, Berry JL, Polski A, Tummina
SJ and O'Brien JM: Next-generation technologies and strategies for
the management of retinoblastoma. Genes (Basel).
10(1032)2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Kong L, Zhang P, Li W, Yang Y, Tian Y,
Wang X, Chen S, Yang Y, Huang T, Zhao T, et al: KDM1A promotes
tumor cell invasion by silencing TIMP3 in non-small cell lung
cancer cells. Oncotarget. 7:27959–27974. 2016.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Fan X, Zhang X, Shen J, Zhao H, Yu X, Chen
Y, Zhuang Z, Deng X, Feng H, Wang Y and Peng L: Decreased TUSC3
promotes pancreatic cancer proliferation, invasion and metastasis.
PLoS One. 11(e0149028)2016.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Horak P, Tomasich E, Vaňhara P,
Kratochvílová K, Anees M, Marhold M, Lemberger CE, Gerschpacher M,
Horvat R, Sibilia M, et al: TUSC3 loss alters the ER stress
response and accelerates prostate cancer growth in vivo. Sci Rep.
4(3739)2014.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Kratochvílová K, Horak P, Ešner M, Souček
K, Pils D, Anees M, Tomasich E, Dráfi F, Jurtíková V, Hampl A, et
al: Tumor suppressor candidate 3 (TUSC3) prevents the
epithelial-to-mesenchymal transition and inhibits tumor growth by
modulating the endoplasmic reticulum stress response in ovarian
cancer cells. Int J Cancer. 137:1330–1340. 2015.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Li P, Zheng X, Shou K, Niu Y, Jian C, Zhao
Y, Yi W, Hu X and Yu A: The iron chelator Dp44mT suppresses
osteosarcoma's proliferation, invasion and migration: In vitro and
in vivo. Am J Transl Res. 8:5370–5385. 2016.PubMed/NCBI
|
|
10
|
Li YG, Liang NX, Qin YZ, Ma DJ, Huang CJ,
Liu L and Li SQ: Effects of RNAi-mediated TUSC3 silencing on
radiation-induced autophagy and radiation sensitivity of human lung
adenocarcinoma cell line A549 under hypoxic condition. Tumour Biol.
37:16357–16365. 2016.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Ohno Y, Koyama H, Yoshikawa T, Takenaka D,
Seki S, Yui M, Yamagata H, Aoyagi K, Matsumoto S and Sugimura K:
Three-way comparison of whole-body MR, coregistered whole-body FDG
PET/MR, and Integrated whole-body FDG PET/CT imaging: TNM and stage
assessment capability for non-small cell lung cancer patients.
Radiology. 275:849–861. 2015.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Scagliotti G, Kang JH, Smith D, Rosenberg
R, Park K, Kim SW, Su WC, Boyd TE, Richards DA, Novello S, et al:
Phase II evaluation of LY2603618, a first-generation CHK1
inhibitor, in combination with pemetrexed in patients with advanced
or metastatic non-small cell lung cancer. Invest New Drugs.
34:625–635. 2016.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Chung KW and Kim SW and Kim SW: Gene
expression profiling of papillary thyroid carcinomas in Korean
patients by oligonucleotide microarrays. J Korean Surg Soc.
82:271–280. 2012.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Yang L, Chen Y, Cui T, Knösel T, Zhang Q,
Albring KF, Huber O and Petersen I: Desmoplakin acts as a tumor
suppressor by inhibition of the Wnt/β-catenin signaling pathway in
human lung cancer. Carcinogenesis. 33:1863–1870. 2012.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Yu X, Zhai C, Fan Y, Zhang J, Liang N, Liu
F, Cao L, Wang J and Du J: TUSC3: A novel tumour suppressor gene
and its functional implications. J Cell Mol Med. 21:1711–1718.
2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Pils D, Horak P, Vanhara P, Anees M, Petz
M, Alfanz A, Gugerell A, Wittinger M, Gleiss A, Auner V, et al:
Methylation status of TUSC3 is a prognostic factor in ovarian
cancer. Cancer. 119:946–954. 2013.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Zhao Y, Schetter AJ, Yang GB, Nguyen G,
Mathé EA, Li P, Cai H, Yu L, Liu F, Hang D, et al: microRNA and
inflammatory gene expression as prognostic marker for overall
survival in esophageal squamous cell carcinoma. Int J Cancer.
132:2901–2909. 2013.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Li Q, Yokoshi M, Okada H and Kawahara Y:
The cleavage pattern of TDP-43 determines its rate of clearance and
cytotoxicity. Nat Commun. 6(6183)2015.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Yu X, Zhang K, Liu F, Zhang J, Zhai C, Cao
L, Song X, Wang Y, Li B, Sun H and Du J: Tumor suppressor candidate
3 as a novel predictor for lymph node metastasis in lung cancer
patients. Oncol Lett. 12:5099–5105. 2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Gu Y, Pei X, Ren Y, Cai K, Guo K, Chen J,
Qin W, Lin M, Wang Q, Tang N, et al: Oncogenic function of TUSC3 in
non-small cell lung cancer is associated with Hedgehog signalling
pathway. Biochim Biophys Acta Mol Basis Dis. 1863:1749–1760.
2017.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Yuan J, Yu X, Wang A, Li Y, Liu F, Wang Y,
Sun S, Bing X, Liu Y and Du J: Tumor suppressor candidate 3: A
novel grading tool and predictor of clinical malignancy in human
gliomas. Oncol Lett. 15:5655–5661. 2018.PubMed/NCBI View Article : Google Scholar
|
|
22
|
Duppel U, Woenckhaus M, Schulz C, Merk J
and Dietmaier W: Quantitative detection of TUSC3 promoter
methylation-a potential biomarker for prognosis in lung cancer.
Oncol Lett. 12:3004–3012. 2016.PubMed/NCBI View Article : Google Scholar
|
|
23
|
Bartel DP: MicroRNAs: Genomics,
biogenesis, mechanism, and function. Cell. 116:281–297.
2004.PubMed/NCBI View Article : Google Scholar
|
|
24
|
Hammond SM: An overview of microRNAs. Adv
Drug Deliv Rev. 87:3–14. 2015.PubMed/NCBI View Article : Google Scholar
|
|
25
|
Ghildiyal M and Zamore PD: Small silencing
RNAs: An expanding universe. Nat Rev Genet. 10:94–108.
2009.PubMed/NCBI View
Article : Google Scholar
|
|
26
|
Soifer HS, Rossi JJ and Saetrom P:
MicroRNAs in disease and potential therapeutic applications. Mol
Ther. 15:2070–2079. 2017.PubMed/NCBI View Article : Google Scholar
|
|
27
|
Krol J, Loedige I and Filipowicz W: The
widespread regulation of microRNA biogenesis, function and decay.
Nat Rev Genet. 11:597–610. 2010.PubMed/NCBI View
Article : Google Scholar
|
|
28
|
O'Connell RM, Rao DS, Chaudhuri AA and
Baltimore D: Physiological and pathological roles for microRNAs in
the immune system. Nat Rev Immunol. 10:111–122. 2010.PubMed/NCBI View
Article : Google Scholar
|
|
29
|
Qadir MI and Faheem A: miRNA: A diagnostic
and therapeutic tool for pancreatic cancer. Crit Rev Eukaryot Gene
Expr. 27:197–204. 2017.PubMed/NCBI View Article : Google Scholar
|
|
30
|
Tutar Y: miRNA and cancer; computational
and experimental approaches. Curr Pharm Biotechnol.
15(429)2014.PubMed/NCBI View Article : Google Scholar
|
|
31
|
Prado MSG, de Jesus ML, de Goes TC,
Mendonça LSO and Kaneto CM: Downregulation of circulating miR-320a
and target gene prediction in patients with diabetic retinopathy.
BMC Res Notes. 13(155)2020.PubMed/NCBI View Article : Google Scholar
|
|
32
|
Zhang C, Yang H, Li Y, Huo P and Ma P:
LNCRNA OIP5-AS1 regulates oxidative low-density
lipoprotein-mediated endothelial cell injury via miR-320a/LOX1
axis. Mol Cell Biochem. 467:15–25. 2020.PubMed/NCBI View Article : Google Scholar
|
|
33
|
Qin H, Liu J, Du ZH, Hu R, Yu YK and Wang
QA: Circular RNA hsa_circ_0012673 facilitates lung cancer cell
proliferation and invasion via miR-320a/LIMK18521 axis. Eur Rev Med
Pharmacol Sci. 24:1841–1852. 2020.PubMed/NCBI View Article : Google Scholar
|
|
34
|
Li M, Qu L, Chen F and Zhu X: Propofol
upregulates miR-320a and reduces HMGB1 by downregulating ANRIL to
inhibit PTC cell malignant behaviors. Pathol Res Pract.
216(152856)2020.PubMed/NCBI View Article : Google Scholar
|
|
35
|
Wang Y, Yang J, Chen P, Song Y, An W,
Zhang H, Butegeleqi B and Yan J: MicroRNA-320a inhibits invasion
and metastasis in osteosarcoma by targeting cytoplasmic
polyadenylation element-binding protein 1. Cancer Med. 9:2833–2845.
2020.PubMed/NCBI View Article : Google Scholar
|
|
36
|
Wu S, Chen S, Lin N and Yang J: Long
non-coding RNA SUMO1P3 promotes hepatocellular carcinoma
progression through activating Wnt/β-catenin signalling pathway by
targeting miR-320a. J Cell Mol Med. 24:3108–3116. 2020.PubMed/NCBI View Article : Google Scholar
|
|
37
|
Zhao JJ, Yang J, Lin J, Yao N, Zhu Y,
Zheng J, Xu J, Cheng JQ, Lin JY and Ma X: Identification of miRNAs
associated with tumorigenesis of retinoblastoma by miRNA microarray
analysis. Childs Nerv Syst. 25:13–20. 2009.PubMed/NCBI View Article : Google Scholar
|
|
38
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
39
|
Zhong Y, Zhao M, Yu Y, Li Q, Wang F, Wu P,
Zhang W and Miao L: Prognostic value and therapeutic potential of
the long noncoding RNA TP73-AS1 in cancers: A systematic review and
meta-analysis. Sci Rep. 10(9053)2020.PubMed/NCBI View Article : Google Scholar
|
|
40
|
Morris LG and Chan TA: Therapeutic
targeting of tumor suppressor genes. Cancer. 121:1357–1368.
2015.PubMed/NCBI View Article : Google Scholar
|
|
41
|
Yang Y and Mei Q: miRNA signature
identification of retinoblastoma and the correlations between
differentially expressed miRNAs during retinoblastoma progression.
Mol Vis. 21:1307–1317. 2015.PubMed/NCBI
|
|
42
|
Castro-Magdonel BE, Orjuela M, Camacho J,
García-Chéquer AJ, Cabrera-Muñoz L, Sadowinski-Pine S,
Durán-Figueroa N, Orozco-Romero MJ, Velázquez-Wong AC,
Hernández-Ángeles A, et al: miRNome landscape analysis reveals a 30
miRNA core in retinoblastoma. BMC Cancer. 17(458)2017.PubMed/NCBI View Article : Google Scholar
|
|
43
|
Svoronos AA, Engelman DM and Slack FJ:
OncomiR or tumor suppressor? The duplicity of MicroRNAs in cancer.
Cancer Res. 76:3666–3670. 2016.PubMed/NCBI View Article : Google Scholar
|
|
44
|
Gao T, Deng M and Wang Q: MiRNA-320a
inhibits trophoblast cell invasion by targeting estrogen-related
receptor-gamma. J Obstet Gynaecol Res. 44:756–763. 2018.PubMed/NCBI View Article : Google Scholar
|
|
45
|
Yong S, Yabin Y, Bing Z, Chuanrong Z,
Dianhua G, Jianhuai Z, Weidong Y, Shuming W and Ling L: Reciprocal
regulation of DGCR5 and miR-320a affects the cellular malignant
phenotype and 5-FU response in pancreatic ductal adenocarcinoma.
Oncotarget. 8:90868–90878. 2017.PubMed/NCBI View Article : Google Scholar
|