Introduction
Intervertebral disc degeneration is a leading cause
of chronic low back pain, which imposes a substantial clinical and
socioeconomic burden (1). Low
proliferative viability and increased apoptosis of human nucleus
pulposus cells (NPCs) are two common features of intervertebral
disc degeneration. Notably, nucleus pulposus therapies are being
developed to treat intervertebral disc degeneration, and clinical
trials are advancing (2).
Hypoxia-inducible factor-1α (HIF-1α) is a crucial
player in the progression of nucleus pulposus growth. A previous
study revealed that HIF-1α regulates metabolic flux by coordinating
glycolysis and mitochondrial TCA cycle interactions, thereby
controlling the overall biosynthetic capacity of NPCs (3). Moreover, it was previously observed
that the alteration of HIF-1α concentration affects NPC
proliferation and apoptosis. Therefore, recombinant HIF-1α protein
may be a potential preventive medicine and therapeutic
application.
Phosphatase and tensin homolog (PTEN) has been
extensively studied in various diseases, such as neoplastic and
orthopedic diseases (4,5). PTEN regulates intervertebral disc
degeneration, which is an orthopedic disease. Liu et al
(6) found that PTEN affects NPC
proliferation during intervertebral disc degeneration (6). Notably, PTEN inhibitors such as
VO-OHpic are being studied as a potential therapy for
intervertebral disc erosion (7).
Indeed, PTEN is a critical factor in the development of
degenerating intervertebral discs.
MicroRNA(miR)-32-5p is a potential diagnostic target
in cancer (8). Previous studies
reported that the long non-coding RNA (lncRNA) SNHG5/miR-32 axis
regulates gastric cancer cell proliferation and migration by
targeting KLF4(9). Wu et al
(10) found that miR-32 regulates
PTEN expression and promotes growth, migration and invasion in
colorectal carcinoma cells. However, the involvement of miR-32-5p
in intervertebral disc degeneration has not been investigated.
Further, the mechanism underlying HIF-1α regulation by the
miR-32-5p/PTEN axis has not yet been reported. In the present
study, the differential expression of miR-32-5p and PTEN was
investigated by analyzing microarray data. The influence of HIF-1α
and the role of miR-32-5p or PTEN on NPCs in vitro was also
studied. Overall, the findings suggest that the maintenance of the
miR-32-5p/PTEN axis is critical for HIF-1α expression in NPCs
during intervertebral disc homeostasis.
Materials and methods
Differentially expressed mRNA analysis
by raw microarray data mining
Raw microarray data (GSE34095_RAW.tar) was
downloaded from the NCBI Gene Expression Omnibus database
(ncbi.nlm.nih.gov/geo). Three samples
were from NPCs of young patients with non-degenerative disc tissues
(GSM841717, GSM841719 and GSM841721), and three samples were taken
from NPCs from elderly patients with disc degeneration (GSM841718,
GSM841720 and GSM841722). An Affymetrix Human Genome U133A Array
(cat. no. GPL96; Affymetrix; Thermo Fisher Scientific, Inc.) was
analyzed using the Affymetrix Transcriptome Analysis Console
(version 1.3). Differentially expressed mRNA was identified by
|fold change (FC)|>1.5 and P<0.05.
Differentially expressed miRNA data
mining using online GEO2R analysis
The GSE116726 dataset contained 3 NPC samples from
healthy control tissues (GSM3259525, GSM3259526 and GSM3259527) and
3 samples of NPCs from patients with degenerative intervertebral
disc (GSM3259528, GSM3259529 and GSM3259530). The differentially
expressed miRNA were screened using the DataSet analysis tool
(GEO2R; ncbi.nlm.nih.gov/geo/geo2r). Differentially expressed
miRNA were identified by|FC|>2 and P<0.05, shown in Table I.
 | Table IDifferentially expressed miRNA data
mining using online GEO2R analysis (|logFC|>2 and
P<0.05). |
Table I
Differentially expressed miRNA data
mining using online GEO2R analysis (|logFC|>2 and
P<0.05).
| miRNA_ID | adj.P.Val | P-value | -lgP | logFC |
|---|
| hsa-miR-32-5p |
2.37x10-17 |
1.17x10-20 | 19.93181414 | -2.81 |
| hsa-miR-187-3p |
2.37x10-17 |
1.85x10-20 | 19.73282827 | 2.52 |
| hsa-miR-2682-3p |
3.64x10-17 |
4.91x10-20 | 19.30891851 | 2.01 |
| hsa-miR-141-3p |
7.80x10-14 |
4.55x10-16 | 15.3419886 | 2.52 |
| hsa-miR-223-5p |
8.89x10-14 |
5.53x10-16 | 15.25727487 | 2.41 |
| hsa-miR-1273h-3p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-130b-5p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-15a-3p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-1911-5p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-27a-5p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-339-5p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
| hsa-miR-379-3p |
1.35x10-13 |
1.21x10-15 | 14.91721463 | 2 |
|
hsa-miR-4715-5p |
1.42x10-13 |
1.82x10-15 | 14.73992861 | 2.01 |
| hsa-miR-492 |
1.42x10-13 |
1.82x10-15 | 14.73992861 | 2.01 |
|
hsa-miR-302c-5p |
3.38x10-13 |
6.84x10-15 | 14.1649439 | 2.01 |
| hsa-miR-3164 |
4.48x10-13 |
1.09x10-14 | 13.9625735 | 2.01 |
|
hsa-miR-516a-3p |
4.48x10-13 |
1.09x10-14 | 13.9625735 | 2.01 |
|
hsa-miR-1288-5p |
4.48x10-13 |
1.10x10-14 | 13.95860731 | 2.01 |
|
hsa-miR-4482-3p |
9.23x10-13 |
3.18x10-14 | 13.49757288 | 2.01 |
| hsa-miR-127-5p |
3.44x10-9 |
1.31x10-10 | 9.882728704 | 2.4 |
|
hsa-miR-3606-3p |
8.20x10-9 |
3.19x10-10 | 9.496209317 | 2 |
| hsa-miR-373-3p |
5.31x10-8 |
2.40x10-9 | 8.619788758 | 2.01 |
| hsa-miR-597-3p |
1.68x10-7 |
8.50x10-9 | 8.070581074 | 2.15 |
| hsa-miR-4434 |
4.44x10-7 |
2.75x10-8 | 7.560667306 | 2 |
| hsa-miR-518b |
4.52x10-7 |
2.83x10-8 | 7.548213564 | 2.04 |
|
hsa-miR-6810-3p |
4.84x10-7 |
3.09x10-8 | 7.510041521 | 2 |
|
hsa-miR-4697-3p |
5.64x10-7 |
3.67x10-8 | 7.435333936 | 2.01 |
|
hsa-miR-6827-3p |
5.64x10-7 |
3.68x10-8 | 7.434152181 | 2.02 |
|
hsa-miR-7846-3p |
6.16x10-7 |
4.05x10-8 | 7.392544977 | 2.22 |
|
hsa-miR-6728-3p |
8.10x10-7 |
5.64x10-8 | 7.248720896 | 2.03 |
| hsa-miR-15b-3p |
8.41x10-7 |
5.92x10-8 | 7.227678293 | 2.1 |
|
hsa-miR-6759-3p |
9.89x10-7 |
7.12x10-8 | 7.147520006 | 2.02 |
| hsa-miR-410-5p |
1.08x10-6 |
8.05x10-8 | 7.09420412 | -2.49 |
|
hsa-miR-376a-5p |
1.08x10-6 |
8.05x10-8 | 7.09420412 | -2.5 |
|
hsa-miR-6799-3p |
1.08x10-6 |
8.06x10-8 | 7.093664958 | 2.03 |
|
hsa-miR-6511b-3p |
1.08x10-6 |
8.10x10-8 | 7.091514981 | 2.02 |
|
hsa-miR-6801-3p |
1.17x10-6 |
8.94x10-8 | 7.048662481 | 2.03 |
| hsa-miR-21-5p |
1.28x10-6 |
9.90x10-8 | 7.004364805 | 2.35 |
|
hsa-miR-518e-5p |
1.59x10-6 |
1.28x10-7 | 6.89279003 | 2.03 |
| hsa-miR-627-5p |
1.64x10-6 |
1.34x10-7 | 6.872895202 | 2.02 |
|
hsa-miR-378a-5p |
1.99x10-6 |
1.69x10-7 | 6.772113295 | -2.46 |
| hsa-miR-486-5p |
2.64x10-6 |
2.43x10-7 | 6.614393726 | -2.77 |
|
hsa-miR-6830-5p |
7.26x10-6 |
8.67x10-7 | 6.061980903 | 2.02 |
| hsa-miR-338-3p |
8.41x10-6 |
1.03x10-6 | 5.987162775 | 2.41 |
|
hsa-miR-146a-5p |
8.71x10-6 |
1.07x10-6 | 5.970616222 | 2.38 |
| hsa-miR-455-5p |
8.91x10-6 |
1.10x10-6 | 5.958607315 | 2.52 |
| hsa-miR-520b |
1.00x10-5 |
1.26x10-6 | 5.899629455 | 2.22 |
|
hsa-miR-6507-3p |
1.69x10-5 |
2.32x10-6 | 5.634512015 | 2.12 |
| hsa-miR-1268b |
2.27x10-5 |
3.31x10-6 | 5.480172006 | 2.01 |
| hsa-miR-563 |
2.34x10-5 |
3.50x10-6 | 5.455931956 | 2.37 |
| hsa-miR-29c-3p |
3.28x10-5 |
5.21x10-6 | 5.283162277 | 2.58 |
| hsa-miR-192-5p |
4.48x10-5 |
7.67x10-6 | 5.115204636 | 2.37 |
| hsa-miR-634 |
6.42x10-5 |
1.21x10-5 | 4.91721463 | 2.01 |
| hsa-miR-34a-5p |
6.46x10-5 |
1.23x10-5 | 4.910094889 | 2.46 |
|
hsa-miR-518a-5p |
7.81x10-5 |
1.57x10-5 | 4.804100348 | 2.33 |
| hsa-miR-610 |
7.86x10-5 |
1.59x10-5 | 4.798602876 | 2.44 |
| hsa-miR-590-5p |
8.15x10-5 |
1.66x10-5 | 4.779891912 | -2.34 |
| hsa-miR-139-5p |
3.78x10-4 |
1.02x10-4 | 3.991399828 | 2.42 |
|
hsa-miR-1306-3p |
4.07x10-4 |
1.11x10-4 | 3.954677021 | 2.06 |
|
hsa-miR-125b-1-3p |
1.89x10-3 |
6.30x10-4 | 3.200659451 | 2.25 |
| hsa-miR-204-5p |
7.03x10-3 |
2.69x10-3 | 2.57024772 | -2.26 |
Target gene prediction
The TargetScan database (targetscan.org) was used to predict microRNA target
genes. Base-pairing and minimum free energy for miR-138 binding to
the target sequences were predicted using the RNAhybrid software
(version 2.1; bibiserv.techfak.uni-bielefeld.de/rnahybrid).
Cell culture
Human nucleus pulposus cells (HNPCs) were purchased
from ScienCell Research Laboratories, Inc. Cells were maintained in
Nucleus Pulposus Cell Medium (ScienCell Research Laboratories,
Inc.) under constant oxygenated culture conditions (37˚C; 20%
oxygen; 5% carbon dioxide; 75% nitrogen) or hypoxic culture
conditions (37˚C; 1% oxygen; 5% carbon dioxide; 94% nitrogen).
Hypoxic cell culture conditions were generated by a hypoxia
incubator (HF100; Heal Force). Cells were grown in constant oxygen
or hypoxic conditions for 24-72 h, according to the experimental
requirements. Recombinant HIF-1α protein (0,0.5,1,2,5 µg/ml; cat.
no. H00003091-P01; Abnova,) was added at the beginning of the
normoxic or hypoxic experiments and maintained at 37˚C
for 24 h.
Transfection
The miR-32-5p mimics or inhibitor, and their
negative controls (scramble), as well as the PTEN small interfering
(si)RNA or PTEN overexpression plasmid (ov-PTEN), and their
negative controls (scrambled) were synthesized by Ribobio Co., Ltd.
According to the manufacturer's instructions, 2x105
human nucleus pulposus cells (HNPC) were seeded in a 6-well plate
and incubated overnight at 37˚C with 5% CO2. Then, HNPC
cells were transfected with miRNA mimics or inhibitors (100 nM),
si-PTEN or PTEN overexpression plasmid (50 nM) at 37˚C using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.). Negative control (NC) mimic was used as the
negative control of miR-32-5p mimic, and NC inhibitor was used as
the negative control of miR-32-5p inhibitor. Si-NC (100 nM) was
used as the negative control of si-PTEN, and ov-NC (100 nM) was
used as the negative control of ov-PTEN. After 4 h, the mixture was
replaced with a fresh medium containing 10% FBS (Gibco; Thermo
Fisher Scientific, Inc.). After 24 h, the cells were collected and
centrifuged at 300 x g for 5 min at 37˚C. Then,
TRIzol® reagent (Invitrogen; Thermo Fisher Scientific,
Inc.) was added for reverse transcription-quantitative PCR
(RT-qPCR) detection. Sequences of si-PTEN, miRNA mimics or
inhibitors, including the negative controls (scramble), were as
follows: si-PTEN, 5'-CAATATTGATGATGTAGTAAG-3'; si-NC,
5'-UUCUCCGAACGUGUCAGUTT-3'; miR-32-5p mimics,
5'-UAUUGCACAUUACUAAGUUGCA-3'; miR-32-5p mimics negative control,
5'-CUAAUUAUCCUGAAGAUCUUAG-3'; miR-32-5p inhibitor,
5'-TGCAACTTAGTAATGTGCAATA-3'; and miR-32-5p inhibitor negative
control, 5'-CAGUACUUUUGUGUAGUACAA-3'.
RT-qPCR
According to the manufacturer's instructions, total
RNA was extracted from HNPCs using TRIzol® reagent
(Invitrogen; Thermo Fisher Scientific, Inc.). The extracted RNA was
reverse transcribed into cDNA using the PrimeScript RT reagent kit
(Invitrogen; Thermo fisher Scientific, Inc.) in accordance with the
manufacturers protocol. qPCR reactions were performed using SYBR
Premix ExTaq II kits (Takara Biotechnology Co., Ltd.) on a 7500
Real-Time PCR System (Applied Biosystems; Thermo Fisher Scientific,
Inc.). The PCR thermocycling conditions were as follows: 95˚C for 5
min, followed by 40 cycles of 95˚C for 15 sec and 60˚C for 1 min.
U6 was used to normalize miR expression and β-actin to normalize
PTEN expression. The following primers were used: miRNA-32-5p
forward, 5'-ACACTCCAGCTGGGTATTGCACATTACTAA-3'; miRNA-32-5p reverse,
5'-CTCAACTGGTGTCGTGGA-3'; U6 forward, 5'-CTCGCTTCGGCAGCACA-3'; U6
reverse, 5'-AACGCTTCACGAATTTGCGT-3'; PTEN forward,
5'-TGAGTTCCCTCAGCCGTTACCT-3'; PTEN reverse,
5'-GAGGTTTCCTCTGGTCCTGGTA-3'; β-actin forward,
5'-AGCGAGCATCCCCCAAAGTT-3'; β-actin reverse,
5'-GGGCACGAAGGCTCATCATT-3'. Relative miR-32-5p and PTEN expression
levels were calculated using the 2-ΔΔCq method (11). All PCR experiments were performed in
triplicate for each sample, and all experiments were repeated three
times.
Western blotting
Cells were harvested and lysed with ice-cold lysis
buffer (Beyotime Institute of Biotechnology) and the protein
concentration was determined using BCA protein assay kit (Nanjing
KeyGen Biotech Co., Ltd.). Denatured proteins (20 µg) were
separated by sodium dodecyl sulfate-polyacrylamide 10% gel
electrophoresis and transferred to polyvinylidene fluoride
membranes (EMD Millipore). The membrane was blocked with 5% BSA
Blocking Buffer (cat. no. SW3015; Beijing Solarbio Science &
Technology Co., Ltd.) and incubated with PTEN primary antibodies
(1:1,000; cat. no. ab267787; Abcam) at 4˚C overnight Next, the
membranes were incubated with secondary antibody Goat Anti-Rabbit
(1:10,000; cat. no. ab205718; Abcam) for 2 h at 25˚C. The bound
proteins were visualized by enhanced chemiluminescence (Thermo
Fisher Scientific, Inc.) and detected using an imaging system (DNR
Bio-Imaging Systems Ltd., Ma'ale Hahamisha). β-actin was used as a
loading control. Densitometric analysis was performed using ImageJ
software (version 1.8.0; National Institutes of Health).
Cell proliferation assay
For cell proliferation assays, single-cell
suspensions were prepared by trypsinization, and cells were seeded
onto 6-well plates with a density of 500 cells per well. After 2
weeks of culture, the cells were digested with trypsin. Then, 10 µl
cell suspension was mixed with 10 µl phenolic blue dye and added to
a counting plate. The cells were incubated at room temperature for
3 min, observed, and counted using light microscope (100x; Olympus;
Tokyo, Japan). The remaining cells were then inoculated in 96-well
plates at a density of 3x103 cells/well and cultured for
24-72 h. Optical density values were measured every 24 h using a
Cell Counting Kit-8 (cat. no. CA1210; Solarbio), according to the
manufacturer's instructions.
Cell apoptosis assay
Apoptosis assays were performed using an Annexin
V-FITC Apoptosis Detection kit (Nanjing KeyGen Biotech Co., Ltd.)
according to the manufacturer's instructions. Cells
(1x106 cells/ml) were harvested and washed twice with
cold PBS and resuspended in 500 µl binding buffer. Next, the cells
were incubated with 5 µl annexin V-FITC and 5 µl propidium iodide
in the dark for 15 min at 25˚C. Apoptosis was assessed by flow
cytometry (BD Biosciences) and using flowjo software (version
10.0.6; FlowJo LLC) to analysis. Each experiment was repeated three
times.
Luciferase reporter assay
The PTEN 3'-UTR was cloned into the psi-CHECK2
vector (Promega Corporation). Mutant reporters (binding site
mutations in PTEN) were then generated. Luciferase reporter assays
were performed by transfecting the mutated promoter reporter
plasmid and the psi-CHECK2-PTEN vector into human 293T cells using
Lipofectamine® 2000 (Invitrogen; Thermo Fisher
Scientific, Inc.). After 48 h, luciferase activity was detected
using a dual luciferase assay system (Promega Corporation),
according to the manufacturer's instructions. The ratio of firefly
to Renilla was used to normalize the firefly luciferase values and
each experiment was repeated three times.
Statistical analysis
All data are expressed as the mean ± standard
deviation (SD), and were analyzed using the SPSS version 19.0
statistical analysis package (SPSS, Inc.). For independent
two-group paired analyses, Student's t-tests were used (unpaired).
All experimental groups were compared with a control group for the
multiple comparisons, which were analyzed using oneway analysis of
variance (ANOVA) followed by Dunnett's post-hoc test. P<0.05 was
considered to indicate a statistically significant difference.
Results
Selection of miR-32-5p and PTEN as
candidate targets
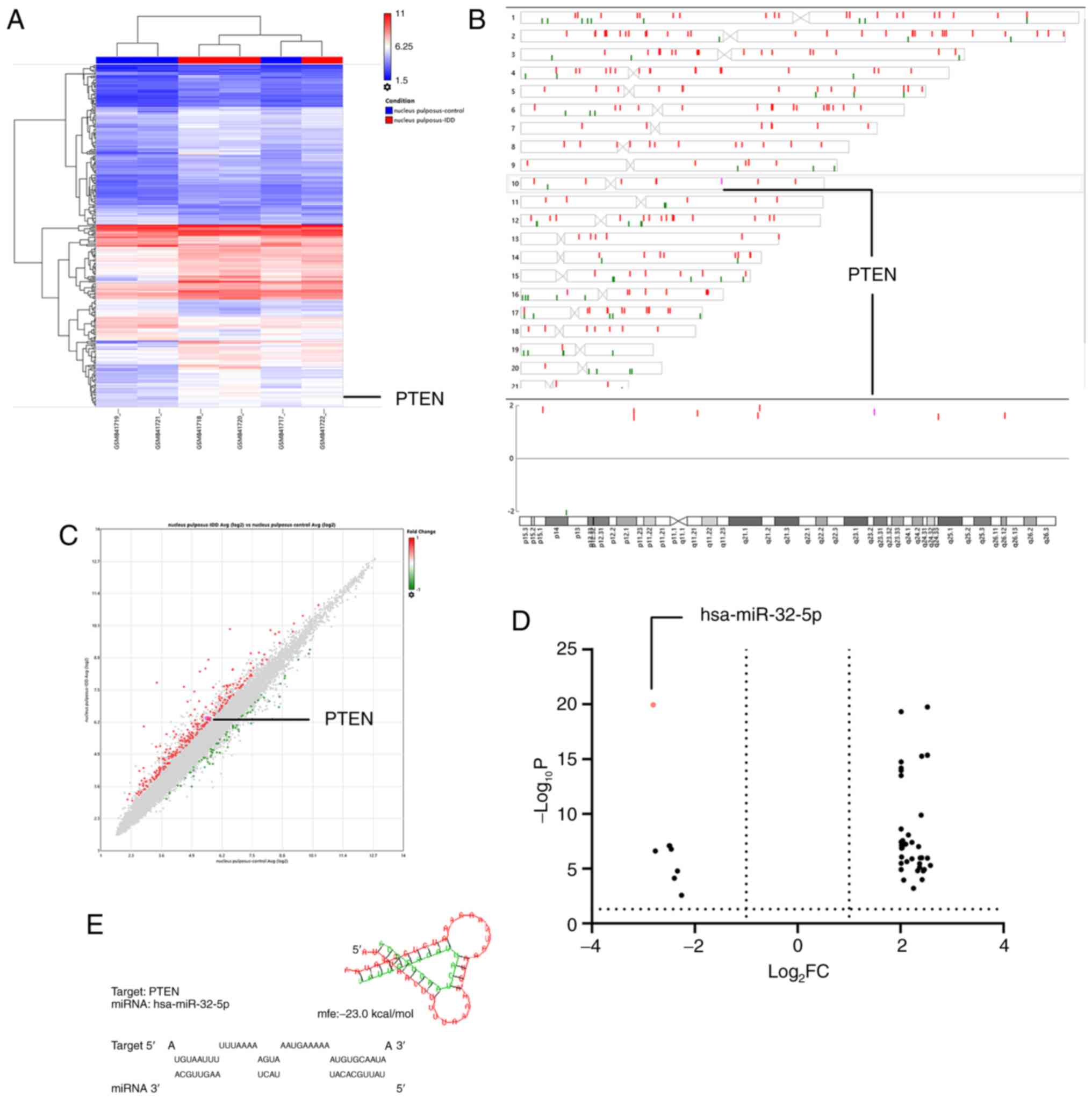
The raw microarray data mining results showed a
negative association between miRNA-32-5p and intervertebral disc
degeneration. In contrast, PTEN on chromosome 10 (q-arm) was
upregulated in NPCs of degenerative disc tissues from elderly
patients (GSE34095; Fig. 1A-C).
miRNA-32-5p was downregulated in NPCs of tissues from patients with
intervertebral disc degeneration, according to the online analysis
on GEO2R of GSE116726 (Fig. 1D).
The PTEN gene sequence was examined and putative miRNA-32-5p
binding sites were identified at position 2859-2866 of the PTEN
3'-UTR. Therefore, it was hypothesized that miR-32-5p interacts
with PTEN. Thus, miR-32-5p and PTEN were chosen for subsequent
experiments.
Hypoxia induces miR-32-5p and PTEN
expression
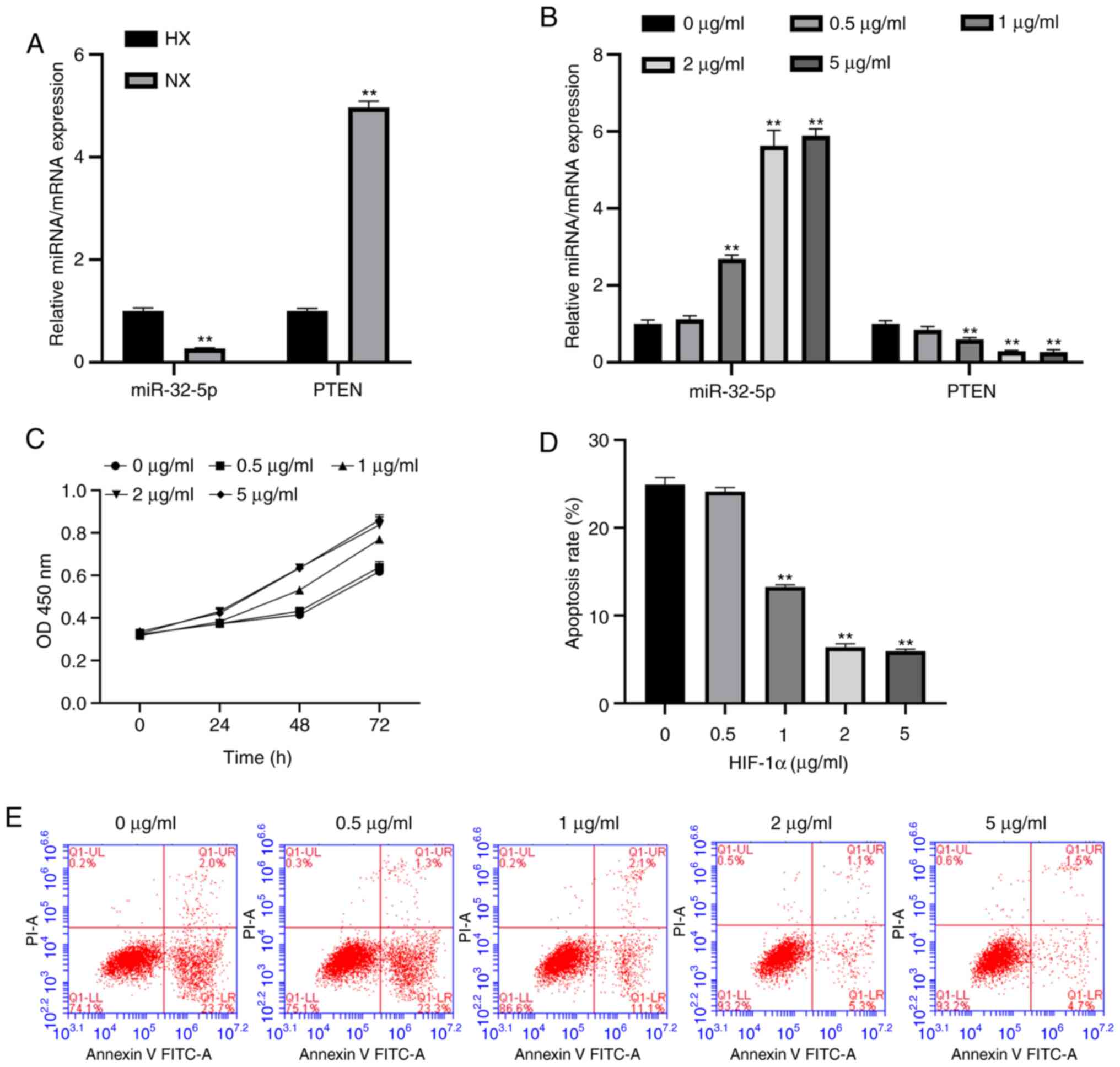
To evaluate the association between miR-32-5p, PTEN
and the NPC oxemic status, the expression levels of miR-32-5p and
PTEN were measured under hypoxic and normoxic conditions using
RT-qPCR. The results suggest that miR-32-5p expression increases
during hypoxia compared with the normoxia group, while PTEN
expression decreases (Fig. 2A).
HIF-1α promotes NPC proliferation,
inhibits apoptosis and alters miR-32-5p and PTEN expression
A previous study reported that HIF-1α is involved in
the regulation of hypoxic stress responses (3). To further investigate whether
recombinant HIF-1α protein controls hypoxic regulation, the
expression levels of miR-32-5p and PTEN were examined following
HIF-1α treatment. It was found that HIF-1α increased miR-32-5p
expression and decreased PTEN expression (Fig. 2B). In addition, HIF-1α promoted NPCs
proliferation and inhibited apoptosis (Fig. 2C-E). The optimal concentration of
HIF-1 was 2 µg/ml; this concentration will be used as the
experimental concentration for the present study.
Hsa-miR-32-5p mimics or inhibitor and
PTEN overexpression or inhibition plasmids were successfully
constructed
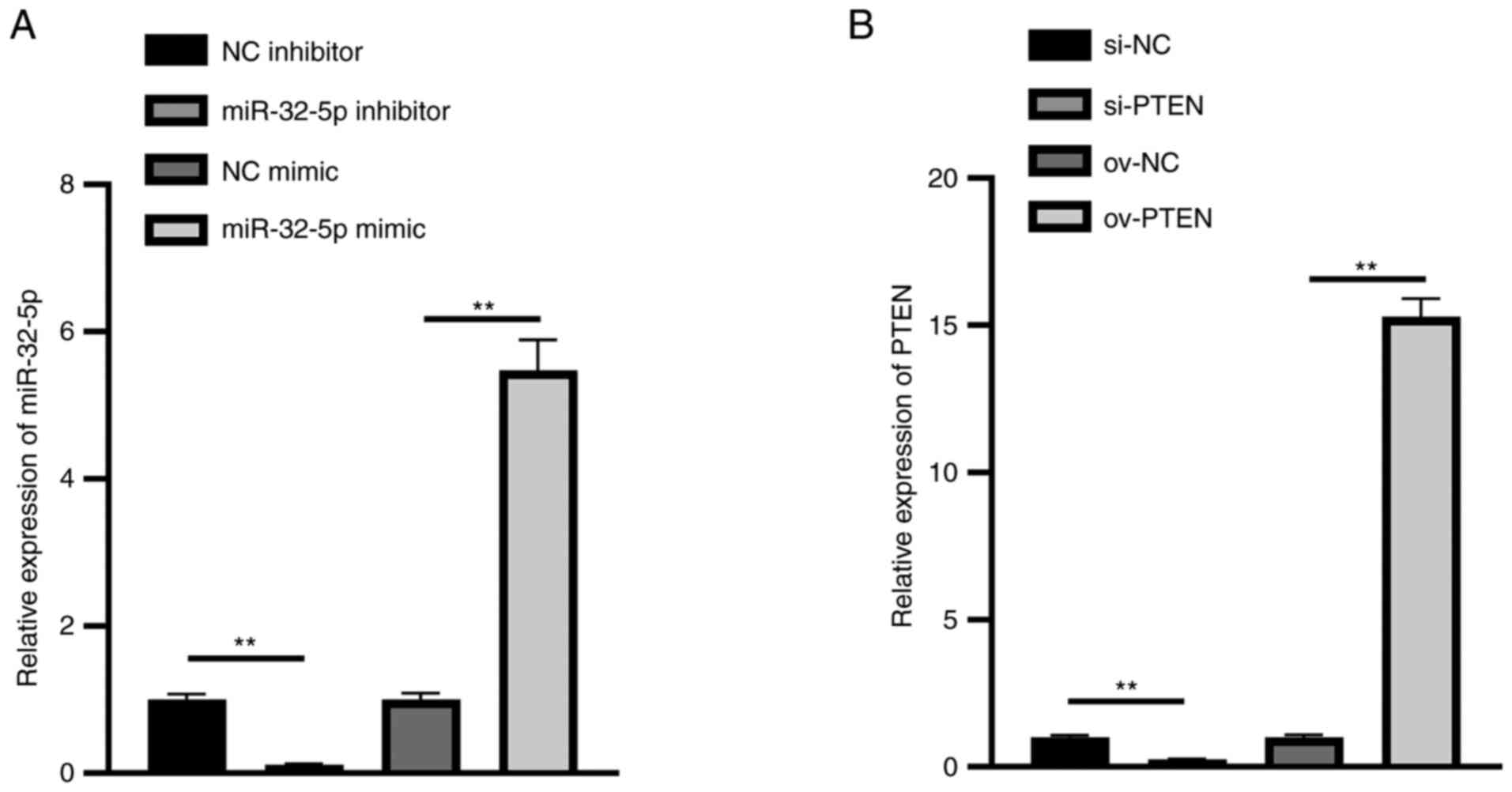
To validate Hsa-miR-32-5p mimics or inhibitor and
PTEN overexpression or inhibition plasmids, these RNAs were
transfected into NPCs. The results show that the expression of
miR-32-5p in the miR-32-5p mimics group was significantly
upregulated, while that in the miR-32-5p inhibitor group was
significantly downregulated (Fig.
3A). Meanwhile, the expression of PTEN in the ov-PTEN group was
significantly upregulated, while that in the si-PTEN group was
significantly downregulated (Fig.
3B). These results illustrate that hsa-miR-32-5p mimics or
inhibitor and PTEN overexpression or inhibition plasmids were
successfully constructed.
miR-32-5p promotes NPC proliferation
and decreases apoptosis
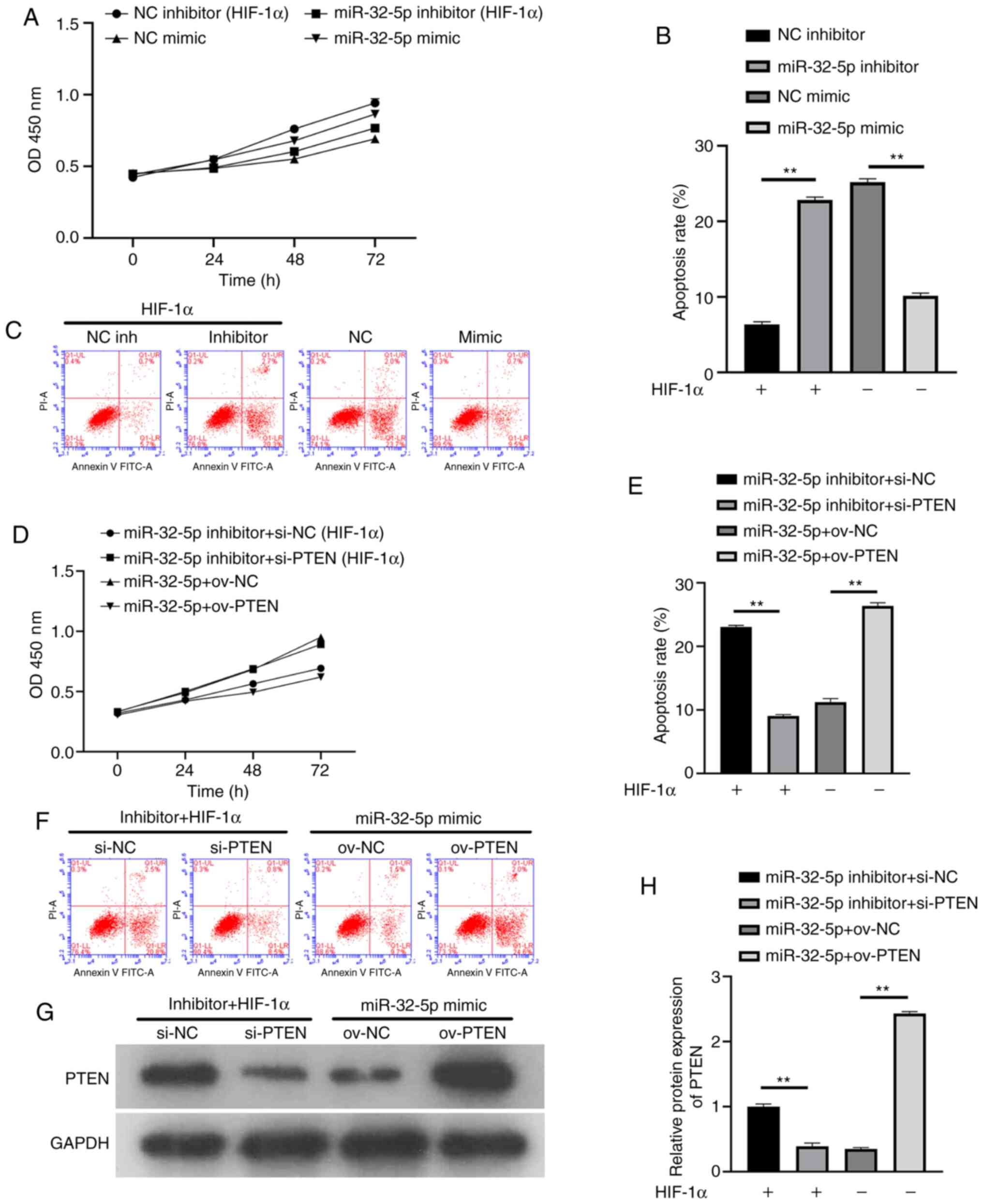
To further confirm that NPC proliferation and
apoptosis are dependent on miR-32-5p or PTEN, the effects of miRNA
or PTEN inhibition and overexpression on cell proliferation were
investigated. The results showed that miR-32-5p inhibitor, after
the addition of recombinant HIF-1α protein, inhibited cell
proliferation and increased apoptosis. In the absence of HIF-1α,
miR-32-5p overexpression promoted cell proliferation and inhibited
apoptosis (Fig. 4A-C). Following
the addition of recombinant HIF-1α protein, decreased PTEN levels
increased cell proliferation and inhibited apoptosis. In the
absence of HIF-1α, PTEN overexpression inhibited cell proliferation
and promoted apoptosis (Fig. 4D-F).
To ensure the success of the transfection experiment, western
blotting was used to verify the PTEN expression (Fig. 4G and H). The results suggest that miR-32-5p or
PTEN can induce NPC proliferation and apoptosis. At the same time,
a significant negative association was demonstrated between
miR-32-5p and PTEN (P<0.01).
PTEN is a direct target of
miR-32-5p
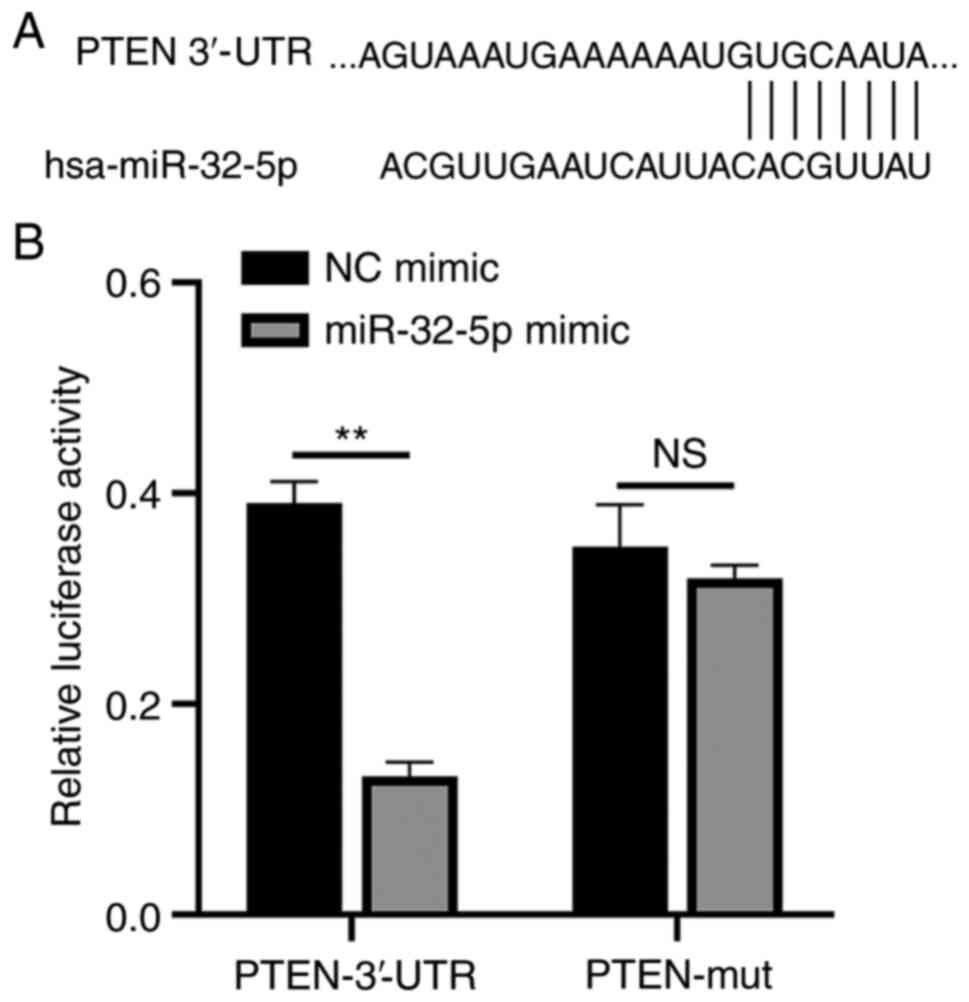
The function of miR-32-5p in NPCs was validated
using PTEN as a target. A single putative miR-32-5p recognition
site was predicted within the PTEN 3'-UTR sequence. The mechanism
through which miR-32-5p modulates PTEN expression was examined by
introducing the PTEN 3'-UTR containing one miR-32-5p binding site
downstream of a luciferase reporter (Fig. 5A). The luciferase activity of the WT
PTEN 3'-UTR construct in transfected cells was significantly lower
compared with that of cells transfected with the control construct.
A luciferase reporter containing a mutant miR-32-5p binding site at
the PTEN 3'-UTR was also constructed. The luciferase activity of
the mutant PTEN 3'-UTR construct was not suppressed upon
transfection with miR-32-5p mimics. These results indicate that
miR-32-5p selectively binds to PTEN mRNAs and that the single
recognition element identified in the PTEN 3'-UTR mRNA is
sufficient for miR-32-5p activity (Fig.
5B). Thus, miR-32-5p directly targets PTEN.
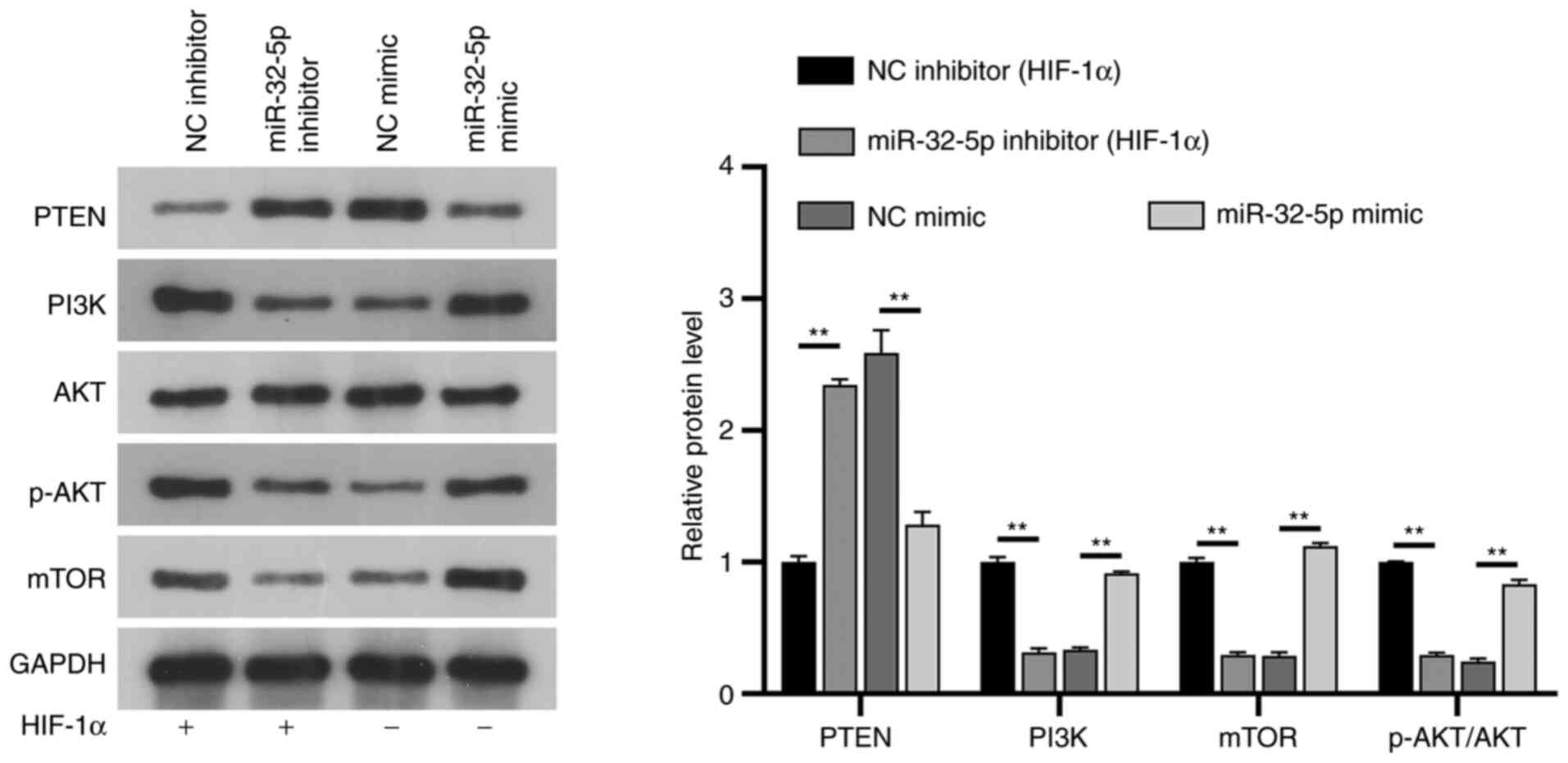
The mechanism of
miR-32-5p/PTEN/PI3K/AKT/mTOR interaction in NPCs
PTEN is a well-known inhibitor of the PI3K/AKT/mTOR
pathway in several diseases, such as autism, spinal cord injury or
prostate cancer. To further understand the potential mechanisms of
miR-32-5p and PTEN in NPCs, their association with the
PI3K/AKT/mTOR signaling pathway was investigated. The results
showed that under constant oxygen conditions, following the
addition of HIF-1α, mir-32-5p mimic could upregulate the
expressions of PI3K, phosphorylated (p)-AKT and mTOR, while the
expression of PTEN was downregulated, and the expression of AKT did
not change significantly (P>0.05). Following the addition of
HIF-1α in the miR-32-5p inhibitor group, PTEN expression was
upregulated, PI3K/p-AKT/mTOR expression was downregulated, and AKT
expression was not significantly changed compared with the NC
inhibitor group (P>0.05). Notably, the expression of
PI3K/AKT/mTOR was significantly downregulated and PTEN was
upregulated in the NC mimic group compared with the NC inhibitor
group, confirming the inhibitory effect of HIF-1α on the
PI3K/AKT/mTOR pathway. HIF-1α regulates the PI3K/AKT/mTOR signaling
pathway through miR-32-5p/PTEN (Fig.
6).
Discussion
The present study demonstrated for the first time
that miR-32-5p and PTEN induce NPC proliferation and apoptosis, and
that hypoxia induces miR-32-5p and PTEN expression. Programmed NPC
apoptosis leads to intervertebral disc degeneration (12); this previous finding combined with
the present results indicates that miR-32-5p is a key factor in the
degeneration of intervertebral discs. Several studies have focused
on the preclinical development of a microRNA-based therapy for
intervertebral disc degeneration (1,13), and
the present results corroborate those findings. The second major
finding of the present study is that HIF-1α controls the
miR-32-5p/PTEN axis to regulate cell proliferation and apoptosis in
NPCs. Together, in the hypoxic intervertebral disc, HIF-1α,
miR-32-5p and PTEN are crucial for NPC proliferation.
It was observed that PTEN was upregulated in the
nucleus pulposus from elderly patient with degenerative disc
tissues, which was consistent with a previous study (7). Interestingly, PTEN, a tumor suppressor
that is frequently mutated in several cancer types, such as breast
cancer and stomach cancer, was downregulated, which generally
inhibits tumorigenesis (14). Thus,
the question was what the major cause of high PTEN expression in
NPCs from healthy patients is? The role of hypoxia was
investigated, which showed that hypoxia induces PTEN expression.
Therefore, PTEN can have different biological functions under
different conditions or in different diseases.
The present study also showed a significant negative
association between miR-32-5p and PTEN expression. Based on the
dual role of PTEN as an inhibitor in the human body,
miR-32-5p-based therapy holds potential in intervertebral disc
degeneration treatment. Emerging evidence indicates that miR-32-5p
may suppress cancer metastasis (15). Indeed, Wang et al (8) found that miR-32-5p suppresses clear
cell renal cell carcinoma metastasis, and Li et al (16) found that HIF-1α is a key responder
to tumor hypoxia. The results of the present study clearly suggest
that miR-32-5p promotes NPC proliferation and decreases apoptosis.
Further, the present results suggest that miR-32-5p has similar
function in intervertebral disc degeneration and in cancer. In the
process of studying PTEN, it was observed that several articles
have reported that PTEN and PI3K/AKT/mTOR signaling pathway are
closely associated in different diseases. Therefore, it was
investigated whether the proliferation and apoptosis of NPC is
influenced by miR-32-5p/PTEN through the regulation of the
PI3K/AKT/mTOR signaling pathway, which was confirmed to be the case
in the present study.
Although HIF-1α is widely studied in various
diseases as a transcription factor that regulates other genes in
the nucleus (17,18), almost no literature reports its
function as a recombinant protein in vitro. Stegen et
al (19) found that HIF-1α
controls collagen synthesis and modification in chondrocytes.
Notably, HIF-1α is a therapeutic target for hepatocellular
carcinoma in the nucleus (20).
Chen and Steidl (21) demonstrated
that inhibition of HIF1α signaling is effective for myelodysplastic
syndromes therapy. In vitro recombinant HIF-1α regulates
cell proliferation and apoptosis in NPCs, similar to its role in
the nucleus. To the best of our knowledge, this is the first study
to investigate HIF-1α as a recombinant protein.
There are limitations to the present study. Firstly,
other miRNAs and mRNAs and associated signaling pathways were not
analyzed in this study, despite the bioinformatics analysis. In
addition, this study was only conducted at the cellular level, and
further verification in animal models is needed in the future.
In summary, the present study findings support that
HIF-1α, as a recombinant protein, controls the
miR-32-5p/PTEN/PI3K/AKT/mTOR axis to regulate cell proliferation
and apoptosis.
Acknowledgements
Not applicable.
Funding
Funding: This study was supported by Natural Science Foundation
of Hainan Province (grant no. 819QN350).
Availability of data and materials
The microarray data (GSE34095, GSE116726) referenced
in the study are available in a public repository from the GEO
website (http://www.ncbi.nlm.nih.gov/geo). The authors declare
that all the other data supporting the findings of this study are
available within the article and its Supplementary Information
files and from the corresponding authors on reasonable request.
Authors' contributions
DZ, ML, JC and GW conceived and designed the study,
and developed the methodology. JC, WC and BW performed the
experiments and collected the data. DZ, ML, JC, WC, JL, YF and YL
analyzed the data. ML, YF, JL, YL, BW and GW interpreted the data.
DZ drafted the manuscript. All authors read and approved the final
manuscript. DZ and GW confirm the authenticity of all the raw
data.
Ethics approval and consent to
participate
Not applicable.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Ji ML, Jiang H, Zhang XJ, Shi PL, Li C, Wu
H, Wu XT, Wang YT, Wang C and Lu J: Preclinical development of a
microRNA-based therapy for intervertebral disc degeneration. Nat
Commun. 9(5051)2018.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Sloan SR Jr, Wipplinger C, Kirnaz S,
Navarro-Ramirez R, Schmidt F, McCloskey D, Pannellini T,
Schiavinato A, Härtl R and Bonassar LJ: Combined nucleus pulposus
augmentation and annulus fibrosus repair prevents acute
intervertebral disc degeneration after discectomy. Sci Transl Med.
12(eaay2380)2020.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Madhu V, Boneski PK, Silagi E, Qiu Y,
Kurland I, Guntur AR, Shapiro IM and Risbud MV: Hypoxic regulation
of mitochondrial metabolism and mitophagy in nucleus pulposus cells
is dependent on HIF-1α-BNIP3 Axis. J Bone Miner Res. 35:1504–1524.
2020.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Hamid AA, Gray KP, Shaw G, MacConaill LE,
Evan C, Bernard B, Loda M, Corcoran NM, Van Allen EM, Choudhury AD
and Sweeney CJ: Compound genomic alterations of TP53, PTEN, and RB1
tumor suppressors in localized and metastatic prostate cancer. Eur
Urol. 76:89–97. 2019.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Meng J, Zhang W, Wang C, Zhang W, Zhou C,
Jiang G, Hong J, Yan S and Yan W: Catalpol suppresses
osteoclastogenesis and attenuates osteoclast-derived bone
resorption by modulating PTEN activity. Biochem Pharmacol.
171(113715)2020.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Liu H, Huang X, Liu X, Xiao S, Zhang Y,
Xiang T, Shen X, Wang G and Sheng B: miR-21 promotes human nucleus
pulposus cell proliferation through PTEN/AKT signaling. Int J Mol
Sci. 15:4007–4018. 2014.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Lin Y, Guo W, Chen KW and Xiao ZM:
VO-OHpic attenuates intervertebral disc degeneration via PTEN/Akt
pathway. Eur Rev Med Pharmacol Sci. 24:2811–2819. 2020.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Wang M, Sun Y, Xu J, Lu J, Wang K, Yang
DR, Yang G, Li G and Chang C: Preclinical studies using miR-32-5p
to suppress clear cell renal cell carcinoma metastasis via altering
the miR-32-5p/TR4/HGF/Met signaling. Int J Cancer. 143:100–112.
2018.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Zhao L, Han T, Li Y, Sun J, Zhang S, Liu
Y, Shan B, Zheng D and Shi J: The lncRNA SNHG5/miR-32 axis
regulates gastric cancer cell proliferation and migration by
targeting KLF4. FASEB J. 31:893–903. 2017.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Wu W, Yang J, Feng X, Wang H, Ye S, Yang
P, Tan W, Wei G and Zhou Y: MicroRNA-32 (miR-32) regulates
phosphatase and tensin homologue (PTEN) expression and promotes
growth, migration, and invasion in colorectal carcinoma cells. Mol
Cancer. 12(30)2013.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Sakai D, Nakamura Y, Nakai T, Mishima T,
Kato S, Grad S, Alini M, Risbud MV, Chan D, Cheah KS, et al:
Exhaustion of nucleus pulposus progenitor cells with ageing and
degeneration of the intervertebral disc. Nat Commun.
3(1264)2012.PubMed/NCBI View Article : Google Scholar
|
|
13
|
Smith LJ, Silverman L, Sakai D, Le Maitre
CL, Mauck RL, Malhotra NR, Lotz JC and Buckley CT: Advancing cell
therapies for intervertebral disc regeneration from the lab to the
clinic: Recommendations of the ORS spine section. JOR Spine.
1(e1036)2018.PubMed/NCBI View Article : Google Scholar
|
|
14
|
Peng W, Chen JQ, Liu C, Malu S, Creasy C,
Tetzlaff MT, Xu C, McKenzie JA, Zhang C, Liang X, et al: Loss of
PTEN promotes resistance to T cell-mediated immunotherapy. Cancer
Discov. 6:202–216. 2016.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Yang D, Ma M, Zhou W, Yang B and Xiao C:
Inhibition of miR-32 activity promoted EMT induced by PM2.5
exposure through the modulation of the Smad1-mediated signaling
pathways in lung cancer cells. Chemosphere. 184:289–298.
2017.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Li H, Jia Y and Wang Y: Targeting HIF-1α
signaling pathway for gastric cancer treatment. Pharmazie. 74:3–7.
2019.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Ling S, Shan Q, Zhan Q, Ye Q, Liu P, Xu S,
He X, Ma J, Xiang J, Jiang G, et al: USP22 promotes hypoxia-induced
hepatocellular carcinoma stemness by a HIF1α/USP22 positive
feedback loop upon TP53 inactivation. Gut. 69:1322–1334.
2020.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Liu J, Zhang X, Chen K, Cheng Y, Liu S,
Xia M, Chen Y, Zhu H, Li Z and Cao X: CCR7 Chemokine
receptor-inducible inc-Dpf3 restrains dendritic cell migration by
inhibiting HIF-1α-mediated glycolysis. Immunity. 50:600–615.e15.
2019.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Stegen S, Laperre K, Eelen G, Rinaldi G,
Fraisl P, Torrekens S, Van Looveren R, Loopmans S, Bultynck G,
Vinckier S, et al: HIF-1α metabolically controls collagen synthesis
and modification in chondrocytes. Nature. 565:511–515.
2019.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Wu FQ, Fang T, Yu LX, Lv GS, Lv HW, Liang
D, Li T, Wang CZ, Tan YX, Ding J, et al: ADRB2 signaling promotes
HCC progression and sorafenib resistance by inhibiting autophagic
degradation of HIF1α. J Hepatol. 65:314–324. 2016.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Chen J and Steidl U: Inhibition of HIF1α
Signaling: A grand slam for MDS therapy? Cancer Discov.
8:1355–1357. 2018.PubMed/NCBI View Article : Google Scholar
|