Introduction
Heparanase (HPSE) is an endo-β-glucuronidase that
promotes the invasion and metastasis of tumor cells (1). HPSE has been confirmed to facilitate
the proliferation and metastasis of ovarian cancer (2), and downregulation of HPSE inhibits the
adhesive and aggressive properties of hepatocellular carcinoma
cells (3). HPSE is also an
important regulator in the tumor microenvironment, such as tumor
angiogenesis. Notably, HPSE mRNA expression is negatively
associated with the prognostic factors of patients with pancreatic
ductal adenocarcinoma. Furthermore, HPSE affects vascular
endothelial growth factor C expression and promotes the invasion of
BxPC-3 cells (4). HPSE is a key
protein influencing tumor angiogenesis and cell proliferation and
invasion in cervical cancer, potentially via the nuclear
factor-Kappa B (NF-κB) signaling pathway (5). In a thyroid carcinoma study, HPSE2
expression was upregulated and tumors exhibited a typical
combination of positive labeling for neoplastic cells and negative
immunostaining in colloid (6).
The molecular mechanisms underlying the progression
of different tumors by HPSE regulation can be multifaceted. It has
been reported that HCCLM3 cells with high HPSE expression exhibit
higher transendothelial migration (TEM) rates (7). Furthermore, downregulation of HPSE or
inhibition of its activity suppresses TEM in HCC cells. The role of
HPSE in modulating autophagy was established in normal and
malignant cells, thereby conferring growth advantages, as well as
resistance to chemotherapy (7).
MicroRNA (miRNA/miR)-558 facilitates the progression of gastric
cancer by directly targeting the HPSE promoter to attenuate
SMAD4-mediated repression of HPSE expression (8).
Currently, the biological function and molecular
mechanism of miR-219a-2-3p in cancer remain unclear. miR-219a-2-3p
expression is upregulated in neural stem cell-derived exosomes,
which forms in the presence of insulin growth factor-1(9). Consequently, the NF-κB pathway is
inhibited and neuroinflammation is attenuated. In addition,
miR-219a-2-3p expression is downregulated in pituitary adenomas,
which suppresses proliferation and promotes apoptosis of pituitary
adenoma cells (10). Similarly,
miR-219a-2-3p is expressed at low levels and negatively associated
with the proliferation of gastric cancer cells (11). Our preliminary bioinformatics
analysis suggested that HPSE may be a probable target of
miR-219a-2-3p (data not shown). However, the function of
miR-219a-2-3p in thyroid cancer and its regulation of HPSE are yet
to be investigated.
The present study aimed to investigate the potential
role of HPSE and miR-219a-2-3p in thyroid cancer, and to elucidate
the molecular mechanism by which miR-219a-2-3p regulates the
proliferation of thyroid cancer cells via HPSE.
Materials and methods
Thyroid cancer samples
Thyroid cancer sections from 80 patients, 30 fresh
thyroid cancer tissues and corresponding paracancerous tissues were
collected between January 2018 and June 2018 in Shengjing Hospital
of China Medical University (Shenyang, China). The inclusion
criteria were as follows: i) Patients diagnosed with thyroid cancer
for the first time; ii) no treatment accepted prior to surgery; and
iii) a pathological diagnosis of thyroid cancer. The exclusion
criteria included: i) Cases with incomplete clinicopathological
data; ii) patients who received radiotherapy or chemotherapy before
surgery; and iii) patients with a previous history of thyroid
surgery. The 80 cases comprised 21 male and 59 female patients with
a mean age of 51 years (range, 32-78 years). All samples were
surgical resections and the paracancerous tissues were collected at
least 2 cm away from the tumor edge. The sections were placed at
room temperature and the fresh tissues were preserved in liquid
nitrogen until subsequent experimentation.
Formalin-fixed paraffin-embedded (FFPE) thyroid
cancer sections were stained at room temperature with hematoxylin
for 10 min and eosin (H&E) for 2 min, and diagnosed according
to the 4th edition of World Health Organization classification of
endocrine tumors guidelines (12)
and the 8th edition of the American Joint Committee on Cancer
(AJCC)/TNM staging system of thyroid cancer (13), by two senior pathologists. The
clinicopathological characteristics of the 80 patients, including
sex, age, differentiation, tumor size, extracapsular invasion and
lymph node metastasis are presented in Table I. The present study was approved by
the Medical Research Ethics Committee of China Medical University
(ethics approval no. 2014PS47K) and written informed consent was
provided by all patients prior to the study start.
 | Table IAssociation between heparanase
expression and clinicopathological characteristics of patients with
thyroid cancer via immunohistochemical staining (n=80). |
Table I
Association between heparanase
expression and clinicopathological characteristics of patients with
thyroid cancer via immunohistochemical staining (n=80).
| Characteristic | Number of
patients | Low expression
(n=20) | High expression
(n=60) | P-value |
|---|
| Sex | | | | |
|
Male | 21 | 7 | 14 | 0.304 |
|
Female | 59 | 13 | 46 | |
| Age, years | | | | |
|
<55 | 30 | 10 | 20 | 0.182 |
|
≥55 | 50 | 10 | 40 | |
| Differentiation | | | | |
|
Well | 62 | 14 | 48 | 0.367 |
|
Poor | 18 | 6 | 12 | |
| Tumor size, cm | | | | |
|
≤2 | 37 | 14 | 23 | 0.018a |
|
>2-≤4 | 35 | 5 | 30 | |
|
>4 | 8 | 1 | 7 | |
| Extracapsular
invasion | | | | |
|
Yes | 33 | 3 | 30 | 0.006b |
|
No | 47 | 17 | 30 | |
| Lymph node
metastasis | | | | |
|
Yes | 43 | 6 | 37 | 0.014a |
|
No | 37 | 14 | 23 | |
Immunohistochemistry
A total of 80 thyroid cancer samples were fixed with
10% formaldehyde at room temperature for 10 min, embedded in
paraffin, and 4-µm thick sections were prepared. These sections
were deparaffinized and conventionally rehydrated. Following
antigen recovery, the sections were incubated with 3%
H2O2 to inhibit endogenous peroxidase
activity, followed by 5% non-immune goat serum at 37˚C for 30 min
to block the unspecific antibody binding. Sections were incubated
with rabbit polyclonal antibody specific for HPSE (1:200;
24529-1-AP; ProteinTech Group, Inc.) at 4˚C overnight. After 24 h,
tissue sections were incubated with goat anti-rabbit IgG (1:1,000;
KIT-0105R; Maxim Biotechnologies, Inc.) and streptavidin-peroxidase
(SP) complex at 37˚C for 30 min (KIT-9710; Maxim Biotechnologies,
Inc.), and subsequently stained with 3,3'-diaminobenzidine at room
temperature for a few seconds. The non-immune goat IgG (1:200
dilution; KIT-0105R; Maxim Biotechnologies, Inc.) was used as the
negative control instead of primary antibody. A total of two senior
pathologists separately evaluated the immunostained sections under
an optical microscope. Brown particles in the cytoplasm were
considered positive HPSE expression. The intensity of HPSE staining
was determined as follows: 0, negative; 1, weak and 2, intense. The
percentage of positive cells (≤50%, 1; >50%, 2) were assessed in
at least five randomly selected fields (magnification, x400) and
both values were multiplied to obtain a final score for each
section as 0, 1, 2 or 4. The sections were classified into low
expression (score ≤2, including score 0) or high expression (score
>2).
Cell culture and transfection
Human normal Nthy-ori 3-1 thyrocytes and the thyroid
cancer cell lines, B-CPAP, TPC-1, KTC-1 and K1, were purchased from
the Cell Bank of Type Culture Collection of The Chinese Academy of
Sciences and maintained in RPMI-1640 medium (Gibco; Thermo Fisher
Scientific, Inc.) supplemented with 10% fetal bovine serum (BioInd,
Inc.), at 37˚C with 5% CO2. The B-CPAP and K1 cell lines
were authenticated via short tandem repeat profiling, provided by
the Cell Line Authentication Service from Shanghai Blowing
Biotechnology Co., Ltd (http://www.biowing.com.cn/). miR-219a-2-3p mimic and
miR-negative control (NC) were purchased from Shanghai GeneChem
Co., Ltd. The sequence of miR-219a-2-3p was
5'-AGAAUUGUGGCUGGACAUCUGU-3' and the NC sequence was
5'-ACGUCGUCACCGGCUAGCAGCAC-3'. Cells were transfected with 50 nM
miR-219a-2-3p or miR-NC using Lipofectamine® 3000
(Invitrogen; Thermo Fisher Scientific, Inc.) at 37˚C for 24 h. The
plasmids for HPSE and the empty vector were purchased from OBiO
Technology (https://www.obiosh.com). The plasmids
were transfected into B-CPAP or TPC-1 cells using
Lipofectamine® 3000 (L3000008; Invitrogen; Thermo Fisher
Scientific, Inc.) 24 h after miR-219a-2-3p or miR-NC transfection
at 37˚C for another 24 h. 48 h later, cells were applied for
subsequent experimentation. All experiments were performed in
triplicate.
Reverse transcription-quantitative
(RT-q)PCR
Total RNA from fresh thyroid tumors and
corresponding paracancerous tissues or cultured thyroid cancer
cells was extracted using the RNApure kit (Aidlab; (http://www.aidlab.cn), and RNA concentration and
purity was determined by UV spectroscopy. The RNA was then reverse
transcribed into cDNA at 25˚C for 5 min and 42˚C for 1 h using the
GoScript Reverse Transcription System (Promega Corporation). qPCR
was subsequently performed using the GoTaq qPCR Master Mix (Promega
Corporation) in the Roche LightCycler 480 Real-Time PCR instrument.
The following primer sequences were used for qPCR: HPSE forward,
5'-AGTGGGTGTGGGTGATTTCC-3' and reverse, 5'-GGCTCCTGGGTGAAGAAGTC-3';
GAPDH forward, 5'-CAGGAGGCATTGCTGATGAT-3 and reverse,
5'-GAAGGCTGGGGCTCATTT-3'; miR-219a-2-3p forward,
5'-GTCCAGAATTGTGGCTGGAC-3' and reverse, 5'-GCAGGGTCCGAGGTATTC-3';
and U6 forward, 5'-CTCGCTTCGGCAGCACA-3' and reverse,
5'-AACGCTTCACGAATTTGCGT-3'. The length of PCR products were 193
base pairs (bp), 152, 22 and 94 bp, respectively. The following
thermocycling conditions were used for qPCR: 40 cycles with an
initial denaturation at 95˚C for 2 min, amplification at 95˚C for
15 sec and annealing at 60˚C for 1 min. Relative expression levels
were calculated using the 2-ΔΔCq (14) method and normalized to the internal
reference gene GAPDH (for HPSE) or U6 (for miR-219a-2-3p).
Western blotting
The cultured cells were lysed using RIPA containing
protease inhibitor (G6521; Promega Corporation) and centrifuged at
22,000 x g at 4˚C for 30 min for supernatant extraction. The
protein content was determined using a BCA assay kit (Beyotime
Institute of Biotechnology). Total protein (50 µg) was separated by
12% SDS-PAGE and then transferred to a PVDF membrane. The membranes
were incubated with 5% non-fat milk powder at room temperature for
2 h to block nonspecific antibody binding. The membranes were then
incubated with the following primary antibodies: Rabbit anti-HPSE
(1:1,000; cat. no. 24529-1-AP; ProteinTech Group, Inc.), mouse
anti-cyclin D1 (1:500; cat. no. TA804673; ZSGB-BIO) and anti-GAPDH
(1:500; cat. no. TA-08; ZSGB-BIO) overnight at 4˚C. Following the
primary incubation, membranes were incubated with goat anti-rabbit
(1:4,000; cat. no. ZB-2301) or anti-mouse IgG (1:2,000; cat. no.
ZB-2305) secondary antibodies at room temperature for 2 h, which
were both purchased from ZSGB-BIO. Protein bands were developed
using electrochemiluminescence (WBKLS0100; MilliporeSigma) and
analyzed using ImageJ software (version 1.52v; National Institutes
of Health). The ratio of IODtarget protein and
IODGAPDH of the same specimen was calculated as the
relative expression level of target protein.
Dual-luciferase reporter assay
The TargetScan database (http://www.targetscan.org) revealed that miR-219a-2-3p
shares complementary binding sequences with HPSE. TargetScan
predicts HPSE as a potential target of any miRNAs by searching for
the presence of conserved 8mer sites that match the seed region,
and miR-219a-2-3p (position 1603-1610 of HPSE 3'-UTR) was selected
in the present study. Partial HPSE sequences containing predictive
miR-219a-2-3p binding sites or mutated sites [HPSE-wild-type (WT)
and HPSE-mutant (MUT)] were customized by Shanghai GenePharma, Co.,
Ltd. B-CPAP or TPC-1 cells were transfected with miR-219a-2-3p
mimic or miR-NC, with WT or MUT luciferase reporter, using
Lipofectamine® 3000 (Invitrogen; Thermo Fisher
Scientific, Inc.). The sequence of miR-219a-2-3p was
5'-AGAAUUGUGGCUGGACAUCUGU-3' and the NC sequence was
5'-ACGUCGUCACCGGCUAGCAGCAC-3'. Following incubation at 37˚C for 48
h, cells were harvested, and luciferase activities were detected
using the Dual-Lumi™ Luciferase Reporter Gene Assay kit (RG089S;
Beyotime Institute of Biotechnology). Renilla luciferase
activity was used as an internal control.
EdU incorporation assay
The kFlour555 Click-iT EdU imaging kit (Nanjing
KeyGen Biotech, Co., Ltd.) was used to assess cell proliferation.
Staining was performed as follows: Cells were cultured with 20
µmol/l of diluted EdU at 37˚C for 2 h. Cells were fixed with 4%
paraformaldehyde at room temperature for 10 min, and neutralized
with a 2 mg/ml glycine solution. Cells were washed with PBS,
permeated with 0.5% Triton X-100 and subsequently incubated with
prepared Click-iT reaction cocktail at room temperature for 30 min
in the dark. DAPI was used to counterstain the nuclei at room
temperature for 30 min, and stained cells were counted in five
randomly selected fields using an inverted fluorescent microscopy
(magnification, x200). Cell proliferative rates were represented as
red/blue.
Statistical analysis
Statistical analysis was performed using SPSS 16.0
software (SPSS, Inc.). Data are representative of three independent
experiments and presented as the mean ± standard deviation. The
χ2 test was used to assess the association between HPSE
expression and the clinicopathological characteristics of patients
with thyroid cancer. Paired-samples t-test was used to compare
differences between cancer tissues and paracancerous tissues.
Spearman's rank correlation coefficient was used to assess the
correlation between HPSE mRNA and miR-219a-2-3p expression levels
in thyroid cancer tissues and cell lines. Unpaired t-test was used
to compare differences between two groups, while one-way ANOVA and
LSD (3 groups) or Tukey's (>3 groups) post hoc tests were used
to compare differences between multiple groups. P<0.05 was
considered to indicate a statistically significant difference.
Results
HPSE expression in thyroid cancer
specimens via immunohistochemistry analysis
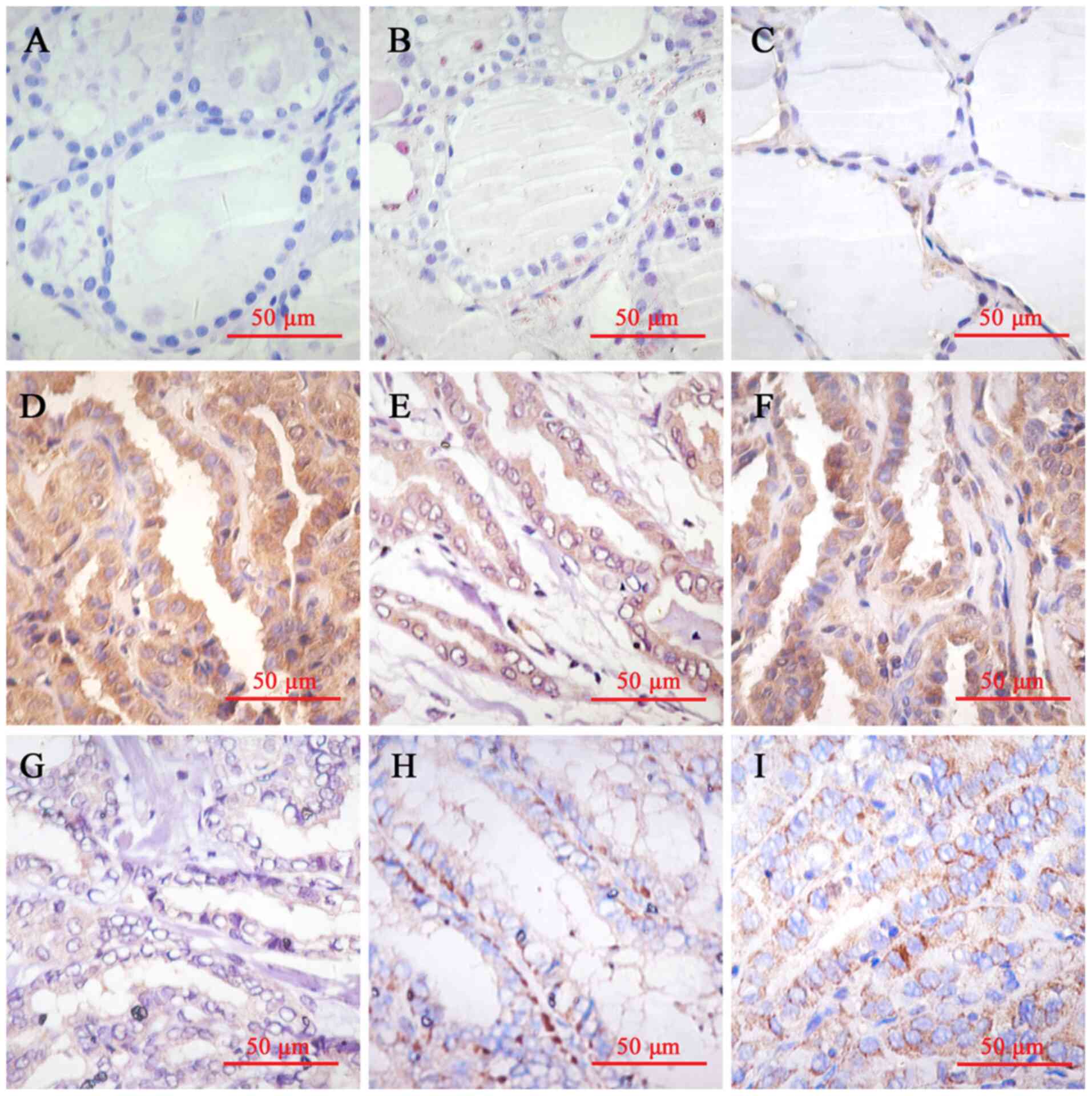
The thyroid follicular epithelial cells exhibited
weak HPSE expression in the cytoplasm. HPSE immunostaining was also
observed in the cytoplasm of thyroid cancer cells. Among the
specimens, 60 (75.0%) were considered high expression (scores
>2), while 20 were considered low expression (scores ≤2). Given
that HPSE expression was associated with tumor growth and
metastasis, the association between HPSE expression and the
clinicopathological characteristics of patients with thyroid cancer
was assessed. As presented in Table
I, HPSE expression was significantly associated with tumor size
(P=0.018), extracapsular invasion (P=0.006) and lymph node
metastasis (P=0.014). However, no significant differences were
observed between HPSE expression and sex, age or differentiation
(P>0.05). HPSE expression in normal thyroid follicular
epithelium (Fig. 1A and B), paracancerous tissues (Fig. 1C) and thyroid cancer tissues
(Fig. 1D-I) is presented in
Fig. 1. The corresponding H&E
images of Fig. 1 are presented in
Fig. S1.
miR-219a-2-3p expression is negatively
correlated with HPSE mRNA expression in thyroid cancer tissues and
cell lines
HPSE was predicted as a target gene of
miR-219a-2-3p, thus, HPSE mRNA and miR-219a-2-3p expression levels
were detected in 30 fresh thyroid cancer tissues and four thyroid
cancer cell lines, and the paracancerous thyroid tissues and normal
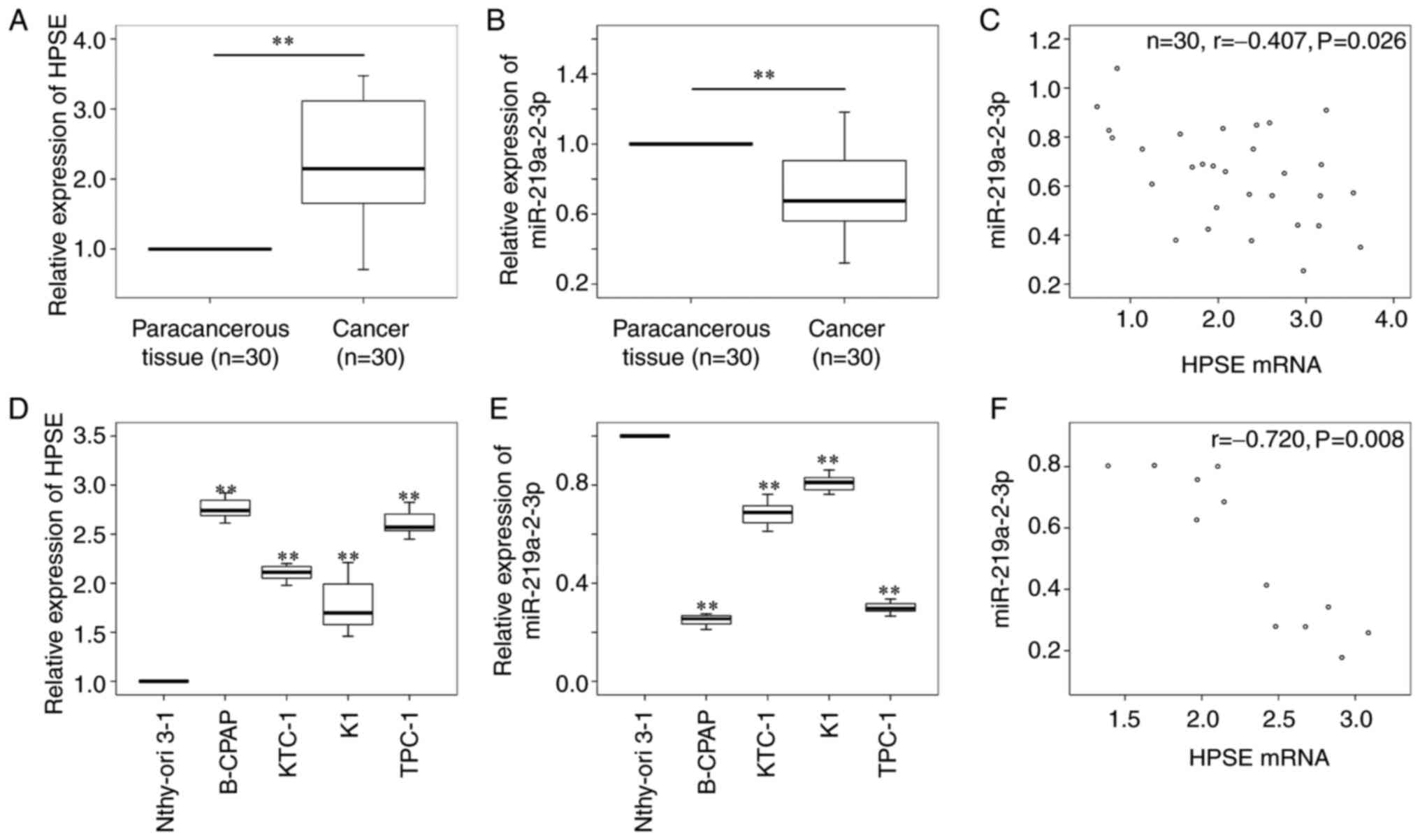
thyrocytes were used as the controls, respectively. The results
demonstrated that HPSE mRNA expression was significantly higher
(P<0.01; Fig. 2A), while
miR-219a-2-3p expression was significantly lower (P<0.01;
Fig. 2B) in thyroid cancer tissues
compared with paracancerous tissues. Furthermore, a negative
correlation was observed between HPSE mRNA and miR-219a-2-3p
expression in thyroid cancer tissues (r, -0.407; P=0.026; Fig. 2C). Similar results were presented in
thyroid cancer cell lines, whereby HPSE expression was
significantly higher (Fig. 2D,
P<0.01), while miR-219a-2-3p expression was significantly lower
(Fig. 2E, P<0.01) in thyroid
cancer cells compared with normal thyrocytes. In addition, a
negative correlation was observed between HPSE mRNA and
miR-219a-2-3p expression in thyroid cancer cells (r, -0.720;
P=0.008; Fig. 2F). B-CPAP and TPC-1
cells (lowest miR-219a-2-3p expression) were selected for
subsequent experimentation.
miR-219a-2-3p modulates HPSE
expression in B-CPAP and TPC-1 cells
Previous studies have reported the involvement of
miR-219a-2-3p in cancer development (10,11).
Thus, it was speculated that miR-219a-2-3p participates in the
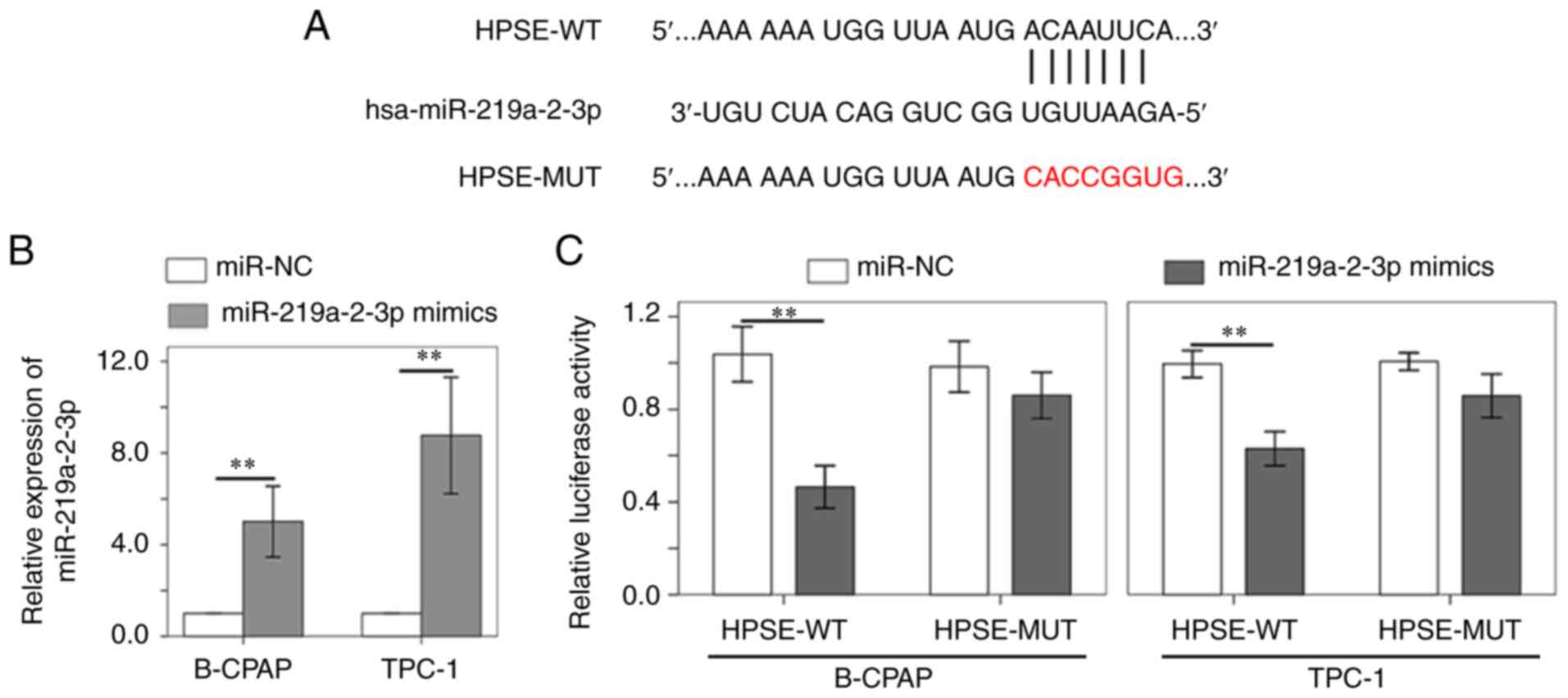
progression of thyroid cancer by targeting HPSE. The TargetScan
database revealed that miR-219a-2-3p shares complementary binding
sequences with HPSE (Fig. 3A). The
dual-luciferase reporter assay was performed to verify the
interaction between HPSE and miR-219a-2-3p. miR-219a-2-3p
expression was confirmed to be upregulated in B-CPAP and TPC-1
cells following transfection with miR-219a-2-3p mimic (Fig. 3B). The results demonstrated that
miR-219a-2-3p mimic notably decreased the luciferase activity of
HPSE reporter compared with miR-NC (Fig. 3C). However, no significant
differences were observed in the luciferase activity of HPSE-MUT
reporter (Fig. 3C).
HPSE mediates miR-219a-2-3p-induced
cyclin D1 inhibition, cell cycle arrest at
G0/G1 and cell proliferation suppression
B-CPAP and TPC-1 cells were transiently transfected
with miR-NC and miR-219a-2-3p mimic, and western blot analysis was
performed to detect the protein expression levels of HPSE and
cyclin D1. The results demonstrated that HPSE expression increased
in both B-CPAP and TPC-1 cells following transfection with HPSE
plasmids (Fig. 4A). Conversely,
HPSE and cyclin D1 expression levels significantly decreased
following transfection with miR-219a-2-3p mimic compared with the
miR-NC groups, respectively. Notably, when HPSE expression
exogenously increased in miR-219a-2-3p mimic-transfected cells,
cyclin D1 expression increased accordingly (Fig. 4B). Taken together, these results
suggest that miR-219a-2-3p regulates cyclin D1 by targeting
HPSE.
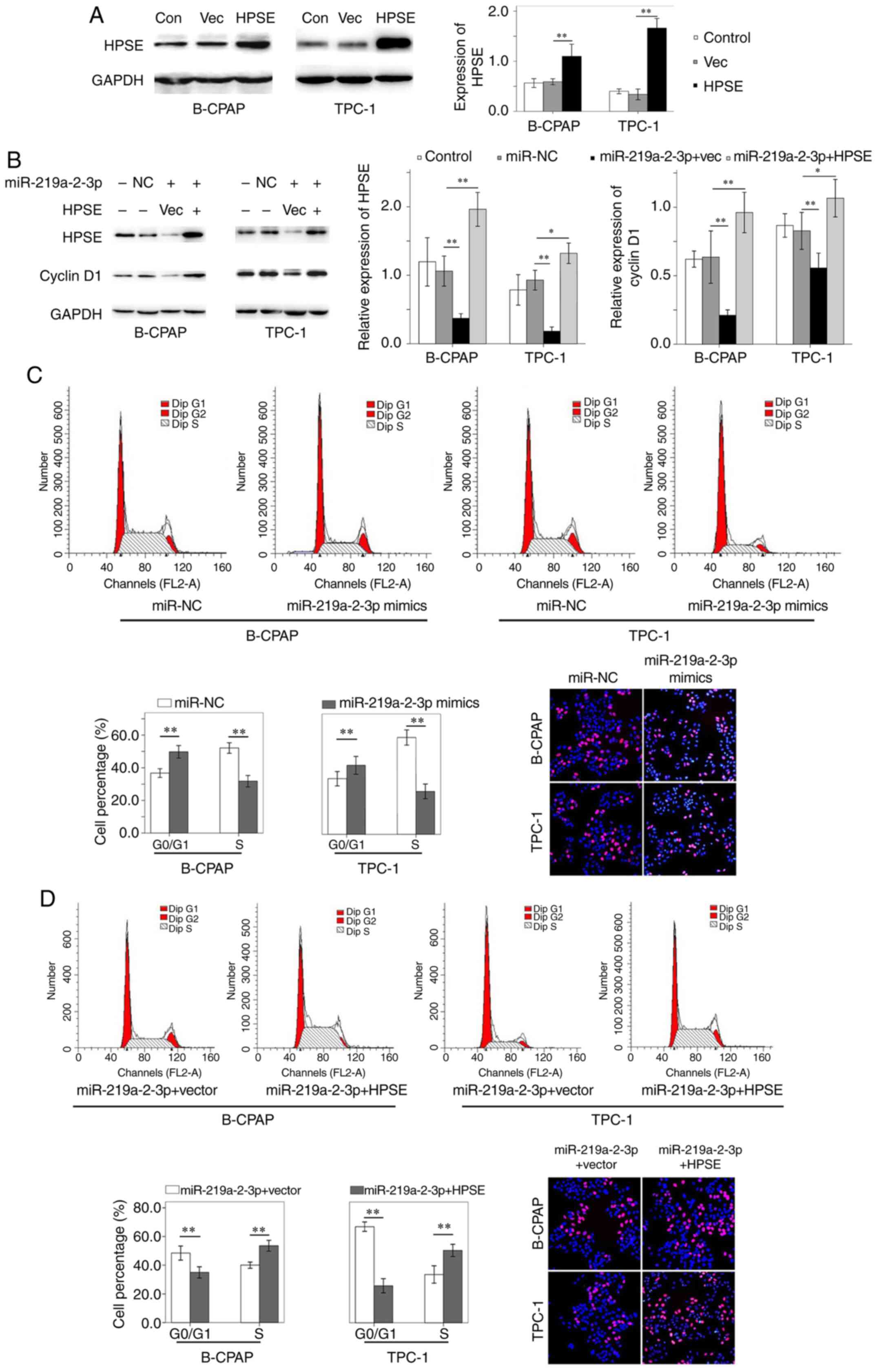 | Figure 4HPSE and cyclin D1 expression, cell
cycle and cell proliferation are regulated by miR-219a-2-3p mimic
via HPSE. (A) HPSE expression increased in B-CPAP and TPC-1 cells
following transfection with HPSE plasmids. (B) miR-219a-2-3p mimic
suppressed HPSE and cyclin D1 expression. Notably, ectopic HPSE
expression reversed the inhibitory effect of miR-219a-2-3p mimic on
cyclin D1 expression. (C) Cell cycle analysis indicated a higher
number of B-CPAP and TPC-1 cells in the G0/G1
phase and less cells in the S phase following transfection with
miR-219a-2-3p mimic. Furthermore, the number of EdU-positive B-CPAP
and TPC-1 cells decreased following transfection with miR-219a-2-3p
mimic. (D) HPSE was exogenously expressed in miR-219a-2-3p
mimic-transfected cells, which decreased the number of cells in the
G0/G1 phase and increased the number of cells
in the S phase compared with the empty vector group. In addition,
the number of proliferative cells was increased in miR-219a-2-3p
mimic-transfected cells with HPSE upregulation. Magnification,
x200. Red staining, EdU; blue staining, DAPI.
*P<0.05; **P<0.01. HPSE, heparanase;
miR, microRNA; NC, negative control; Con, control; Vec, empty
vector. |
Cell cycle analysis demonstrated a higher number of
B-CPAP and TPC-1 cells in the G0/G1 phase and
less cells in the S phase following transfection with miR-219a-2-3p
mimic compared with the miR-NC groups, respectively (Fig. 4C). However, when ectopic HPSE
expression was induced in miR-219a-2-3p mimic-transfected cells,
the percentage of cells decreased in the
G0/G1 phase and increased in the S phase
compared with the empty vector-transfected cells (Fig. 4D). Collectively, these results
suggest that miR-219a-2-3p induces cell cycle arrest at the
G0/G1 phase, which was also associated with
HPSE.
The proliferative ability of B-CPAP and TPC-1 cells
was assessed via the EdU incorporation assay. The results
demonstrated that the percentage of EdU-positive B-CPAP cells was
32.4±1.4 and 28.3±2.7% in the miR-NC and miR-219a-2-3p mimic
groups, respectively (P=0.016; Fig.
4C). In the empty vector or HPSE transfected cells
overexpressed with miR-219a-2-3p, the percentage of EdU-positive
B-CPAP cells was 23.3±2.0 and 37.2±2.3%, respectively (P<0.01;
Fig. 4D). Similarly, the percentage
of proliferative TPC-1 cells was 30.3±1.2 and 28.0±1.6% in the
miR-NC and miR-219a-2-3p mimic groups, respectively (P=0.028;
Fig. 4C). In the empty vector or
HPSE transfected cells overexpressed with miR-219a-2-3p, the
percentage of proliferative TPC-1 cells was 22.5±2.4 and 33.5±2.5%,
respectively (P<0.01; Fig. 4D).
Taken together, these results suggest that miR-219a-2-3p inhibits
the proliferative ability of thyroid cancer cells via HPSE.
Discussion
HPSE expression is upregulated in different types of
human cancer, which is associated with tumor metastasis and
invasion (15,16). It has been reported that high HPSE
expression is associated with poor prognosis and contributes to the
lymphovascular invasion of breast cancer (17). In a systematic study, either HPSE
expression in cancer and/or serum HPSE concentration were reported
to act as potential biomarkers for the evaluation of surgery
effects and prognosis prediction in patients with ovarian cancer
(18). In the present study, HPSE
expression was investigated in 80 FFPE thyroid cancer samples via
immunohistochemical staining. The results demonstrated that HPSE
expression was upregulated in thyroid cancer samples, and normal
thyroid follicular epithelium exhibited negative or weak HPSE
staining, supporting its potential role in thyroid tumorigenesis.
Furthermore, the clinical significance of HPSE in thyroid cancer
was assessed. It was hypothesized that thyroid cancer with high
HPSE expression was more likely to exhibit tumor extension and
lymph node metastasis. Given that HPSE is critical in tumorigenesis
and progression of thyroid cancer (6), repressing HPSE expression may be a
promising option for inhibiting thyroid cancer. In other cancers,
downregulation of HPSE represses glioma cell proliferation, and
exogenous HPSE expression stimulates growth and activates ERK and
AKT signaling (19). In addition,
miR-429 decreases the invasive ability of gastric cancer cells by
downregulating HPSE expression (20).
miRNAs are small non-coding RNA molecules that
function in post-transcriptional silencing of gene expression via
base pairing with complementary sequences within the target mRNAs.
Bioinformatic analysis via the TargetScan database revealed that
HPSE is a regulatory target of miR-219a-2-3p. Previous studies have
reported that miR-219a-2-3p is a tumor suppressor in pituitary
adenoma (10), glioma (21) and gastric cancer (11). Mouse double minute 2 homolog (MDM2)
is a putative target of miR-219a-2-3p, based on bioinformatics
analysis in pituitary adenomas. miR-219a-2-3p decreases MDM2
expression by binding to its 3'-UTR and promoting p53 expression,
which in turn suppresses cell proliferation (10). In glioma cell lines with WT
isocitrate dehydrogenase (IDH)1/2, overexpression of
miR-219a-2-3p decreases cell proliferation and colony formation
(21). The predicted targets of
miR-219a-2-3p have been demonstrated to participate in tumor
progression by activating the Ras-ERK and PI3K-AKT pathways. In
addition, miR-219a-2-3p expression is downregulated in gastric
cancer tissues and cells. FUS expression is negatively associated
with miR-219a-2-3p. In gastric cancer cells, miR-219a-2-3p targets
FUS, which facilitates cell proliferation (11).
In the present study, miR-219a-2-3p expression was
downregulated in thyroid cancer tissues and cells, which was
negatively correlated with HPSE mRNA expression. Based on cell
cycle analysis and EdU incorporation assay, miR-219a-2-3p mimic
significantly attenuated the proliferative ability of thyroid
cancer cells, which further validates its tumor suppressive role in
thyroid cancer. In addition, miR-219a-2-3p decreased cyclin D1
expression and induced cell cycle arrest at the
G0/G1 phase, the effects of which were
reversed following exogenous HPSE expression. Taken together, these
results suggest that miR-219a-2-3p negatively regulates the
proliferation of thyroid cancer cells via HPSE. The regulatory
mechanism of miR-219a-2-3p on HPSE was further confirmed as the
3'-UTR luciferase activity of HPSE significantly decreased in
miR-219a-2-3p mimic-transfected cells, resulting in decreased HPSE
protein expression.
In conclusion, the results of the present study
suggest that HPSE is essential for the development of thyroid
cancer, and is the downstream target gene of miR-219a-2-3p.
miR-219a-2-3p decreased the proliferative ability of thyroid cancer
cells and induced cell cycle arrest at the
G0/G1 phase, likely by decreasing cyclin D1
expression, which was associated with HPSE suppression.
Furthermore, overexpression of miR-219a-2-3p suppressed the
proliferation of thyroid cancer cells by targeting HPSE and cyclin
D1, thus, the miR-219a-2-3p/HPSE/cyclin D1 axis may be a novel area
in thyroid cancer research. Further studies are required to confirm
the role of miR-219a-2-3p/HPSE and its downstream pathway in
thyroid cancer, in addition to cyclin D1 in the regulation of
thyroid cancer progression, which may provide a potential
therapeutic target to neutralize the effects of HPSE on thyroid
cancer. In addition, this in vitro analysis on the
regulation of miR-219a-2-3p/HPSE on cell proliferation requires
in vivo validation.
Supplementary Material
Corresponding hematoxylin and eosin
images of Fig. 1. (A and B) Normal
thyroid tissues. (C) Paracancerous thyroid tissues. (D) Thyroid
cancer tissues, >4 cm. (E) Thyroid cancer tissues with
extracapsular invasion. (F) Thyroid cancer tissues with lymph node
metastasis. (G) Thyroid cancer tissues, ≤2 cm. (H) Thyroid cancer
tissues without extracapsular invasion. (I) Thyroid cancer tissues
without lymph node metastasis. Magnification, x400.
Acknowledgements
Not applicable.
Funding
Funding: The present study was supported by the National Natural
Science Foundation of China (grant no. 81672644) and the 345 Talent
Project of Shengjing Hospital (grant no. M0731).
Availability of data and materials
The datasets used and/or analyzed during the current
study are available from the corresponding author on reasonable
request.
Authors' contributions
CY and ZL made substantial contributions to the
conception and design of the present study. SZ and YH performed the
experiments and drafted the initial manuscript. XC and DC analyzed
and interpreted the data. YH and CY confirmed the authenticity of
all the raw data. All authors have read and approved the final
manuscript.
Ethics approval and consent to
participate
The present study was approved by the Ethics
Committee of Shengjing Hospital of China Medical University
(Shenyang, China; ethics approval no. 2014PS47K). Written informed
consent was provided by all patients prior to the study start.
Patient consent for publication
Not applicable.
Competing interests
The authors declare that they have no competing
interests.
References
|
1
|
Masola V, Zaza G, Gambaro G, Franchi M and
Onisto M: Role of heparanase in tumor progression: Molecular
aspects and therapeutic options. Semin Cancer Biol. 62:86–98.
2020.PubMed/NCBI View Article : Google Scholar
|
|
2
|
Zheng HY, Ruan J, Zhao P, Chen SP, Pan LL
and Liu JQ: Heparanase is involved in proliferation and invasion of
ovarian cancer cells. Cancer Biomark. 15:525–534. 2015.PubMed/NCBI View Article : Google Scholar
|
|
3
|
Chen XP, Luo JS, Tian Y, Nie CL, Cui W and
Zhang WD: Downregulation of heparanase expression results in
suppression of invasion, migration, and adhesion abilities of
hepatocellular carcinoma cells. Biomed Res Int.
2015(241983)2015.PubMed/NCBI View Article : Google Scholar
|
|
4
|
Lv B, Zhang B, Hu XY and Zeng QD:
Heparanase regulates in vitro VEGF-C expression and its clinical
significance to pancreatic ductal cell adenocarcinoma. Oncol Lett.
11:1327–1334. 2016.PubMed/NCBI View Article : Google Scholar
|
|
5
|
Lv QY, Wu KJ, Liu FL, Wu WR, Chen YR and
Zhang W: Interleukin-17A and heparanase promote angiogenesis and
cell proliferation and invasion in cervical cancer. Int J Oncol.
53:1809–1817. 2018.PubMed/NCBI View Article : Google Scholar
|
|
6
|
Matos LL, Suarez ER, Theodoro TR, Trufelli
DC, Melo CM, Garcia LF, Oliveira OC, Matos MG, Kanda JL, Nader HB,
et al: The profile of heparanase expression distinguishes
differentiated thyroid carcinoma from benign neoplasms. PLoS One.
10(e0141139)2015.PubMed/NCBI View Article : Google Scholar
|
|
7
|
Chen XP, Jiang W, Yue CF, Zhang WJ, Tong
CG, Dai DF, Cheng B, Huang C and Lu L: Heparanase contributes to
trans-endothelial migration of hepatocellular carcinoma cells. J
Cancer. 8:3309–3317. 2017.PubMed/NCBI View Article : Google Scholar
|
|
8
|
Zheng LD, Jiao WJ, Song HJ, Qu HX, Li D,
Mei H, Chen YJ, Yang F, Li HH, Huang K and Tong QG: miRNA-558
promotes gastric cancer progression through attenuating
Smad4-mediated repression of heparanase expression. Cell Death Dis.
7(e2382)2016.PubMed/NCBI View Article : Google Scholar
|
|
9
|
Ma K, Xu HY, Zhang J, Zhao F, Liang HQ,
Sun HT, Li P, Zhang S, Wang RJ and Chen XY: Insulin-like growth
factor-1 enhances neuroprotective effects of neural stem cell
exosomes after spinal cord injury via an miR-219a-2-3p/YY1
mechanism. Aging (Albany NY). 11:12278–12294. 2019.PubMed/NCBI View Article : Google Scholar
|
|
10
|
Wang YB, Zhao JN, Zhang CC, Wang PC, Huang
CX and Peng H: MiR-219a-2-3p suppresses cell proliferation and
promotes apoptosis by targeting MDM2/p53 in pituitary adenomas
cells. Biosci Biotechnol Biochem. 84:911–918. 2020.PubMed/NCBI View Article : Google Scholar
|
|
11
|
Yang Z, Dong XH, Pu ML, Yang HW, Chang WL,
Ji FH, Liu T, Wei CQ, Zhang XF and Qiu XG:
LBX2-AS1/miR-219a-2-3p/FUS/LBX2 positive feedback loop contributes
to the proliferation of gastric cancer. Gastric Cancer. 23:449–463.
2020.PubMed/NCBI View Article : Google Scholar
|
|
12
|
Lloyd RV, Osamura RY, Klöppel G and Rosai
J (eds): WHO Classification of Tumours of Endocrine Organs. 4th
edition. IARC, Lyon, 2017.
|
|
13
|
Amin MB, Edge S, Greene F, Byrd DR,
Brookland RK, Washington MK, Gershenwald JE, Compton CC, Hess KR,
Sullivan DC, et al: AJCC Cancer Staging Manual. 8th edition.
Springer, New York, NY, 2017.
|
|
14
|
Livak KJ and Schmittgen TD: Analysis of
relative gene expression data using real-time quantitative PCR and
the 2(-Delta Delta C(T)) method. Methods. 25:402–408.
2001.PubMed/NCBI View Article : Google Scholar
|
|
15
|
Wang C, Wei YJ, Wang G, Zhou YM, Zhang JC
and Xu K: Heparanase potentiates the invasion and migration of
pancreatic cancer cells via epithelial-to-mesenchymal transition
through the Wnt/β-catenin pathway. Oncol Rep. 44:711–721.
2020.PubMed/NCBI View Article : Google Scholar
|
|
16
|
Chen XP, Cheng B, Dai DF, Wu YH, Feng ZW,
Tong CG, Wang XM and Zhao J: Heparanase induces necroptosis of
microvascular endothelial cells to promote the metastasis of
hepatocellular carcinoma. Cell Death Discov. 7(33)2021.PubMed/NCBI View Article : Google Scholar
|
|
17
|
Tang DB, Piao Y, Zhao S, Mu XD, Li S, Ma
WJ, Song Y, Wang JX, Zhao WH and Zhang QY: Expression and
correlation of matrix metalloproteinase-9 and heparanase in
patients with breast cancer. Med Oncol. 31(26)2014.PubMed/NCBI View Article : Google Scholar
|
|
18
|
Zhang W, Chan H, Wei LW, Pan ZM, Zhang JQ
and Li L: Overexpression of heparanase in ovarian cancer and its
clinical significance. Oncol Rep. 30:2279–2287. 2013.PubMed/NCBI View Article : Google Scholar
|
|
19
|
Kundu S, Xiong AQ, Spyrou A, Wicher G,
Marinescu VD, Edqvist PHD, Zhang L, Essand M, Dimberg A, Smits A,
et al: Heparanase promotes glioma progression and is inversely
correlated with patient survival. Mol Cancer Res. 14:1243–1253.
2016.PubMed/NCBI View Article : Google Scholar
|
|
20
|
Sheng N, Zhang L and Yang SF: MicroRNA-429
decreases the invasion ability of gastric cancer cell line BGC-823
by downregulating the expression of heparanase. Exp Ther Med.
15:1927–1933. 2018.PubMed/NCBI View Article : Google Scholar
|
|
21
|
Fleming J, Bell EH, Tong ZY, Grozdic I,
McElroy J, Beyer S, Fabian D, Cui TT, Popp I, Staszewski O, et al:
CSIG-21. The role of miR-219a-2-3p as a tumor suppressor in
IDH1/2-wild-type grade II/III gliomas. Neuro Oncol. 20 (Suppl
6)(vi47)2018.
|